Network Analysis Reveals Seasonal Patterns of Bacterial Community Networks in Lake Taihu under Aquaculture Conditions
Abstract
1. Introduction
2. Materials and Methods
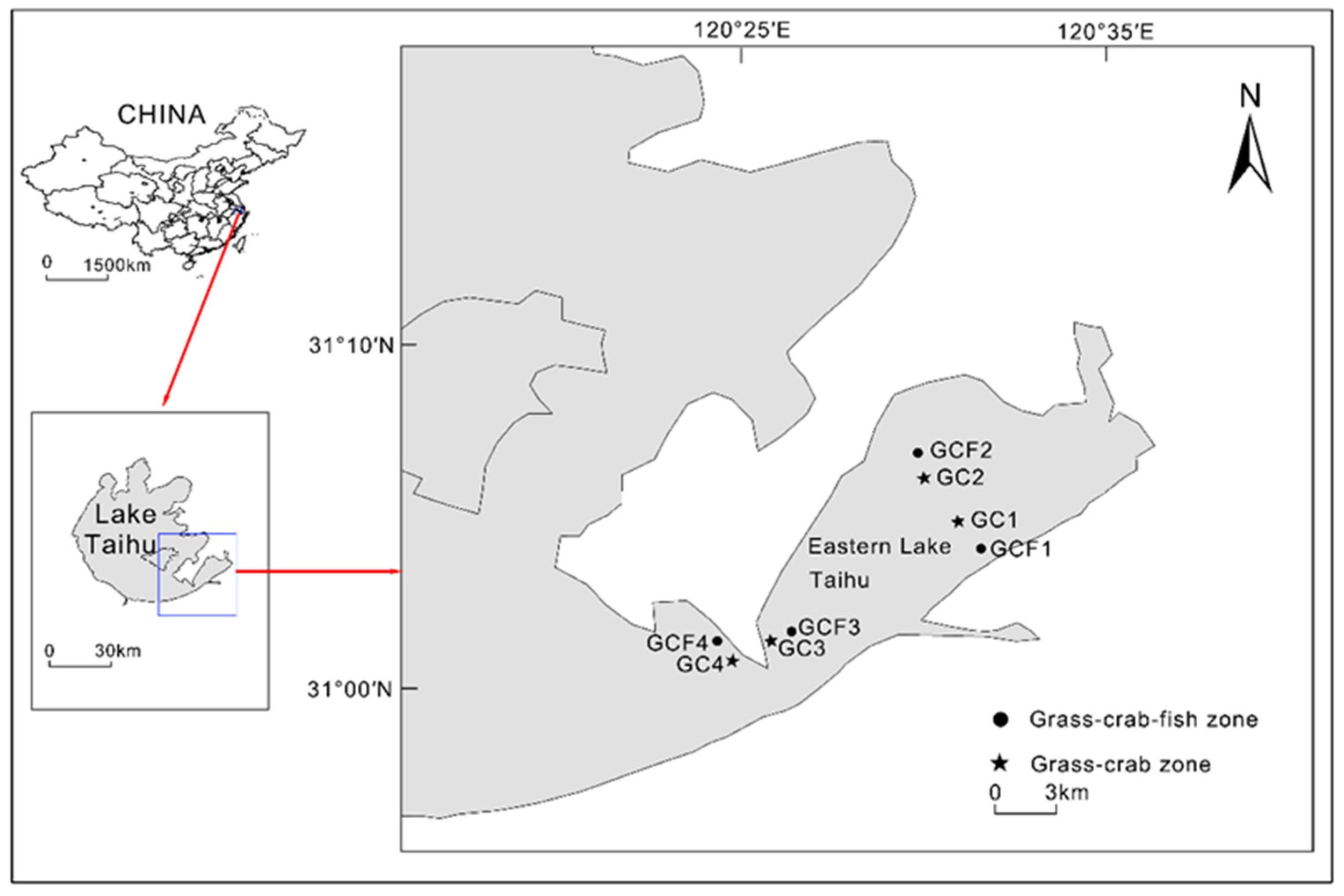
2.1. Sample Collection and Measurement of Physicochemical Variables
2.2. DNA Extraction, Polymerase Chain Reaction (PCR) Amplification and Illumina Sequencing
2.3. Sequence Data Processing and Statistical Analysis
2.4. Seasonal Network Construction and Characterization
2.5. Topological Roles of Individual Nodes
2.6. Relationships between Bacterial Networks and Environmental Variables
3. Results
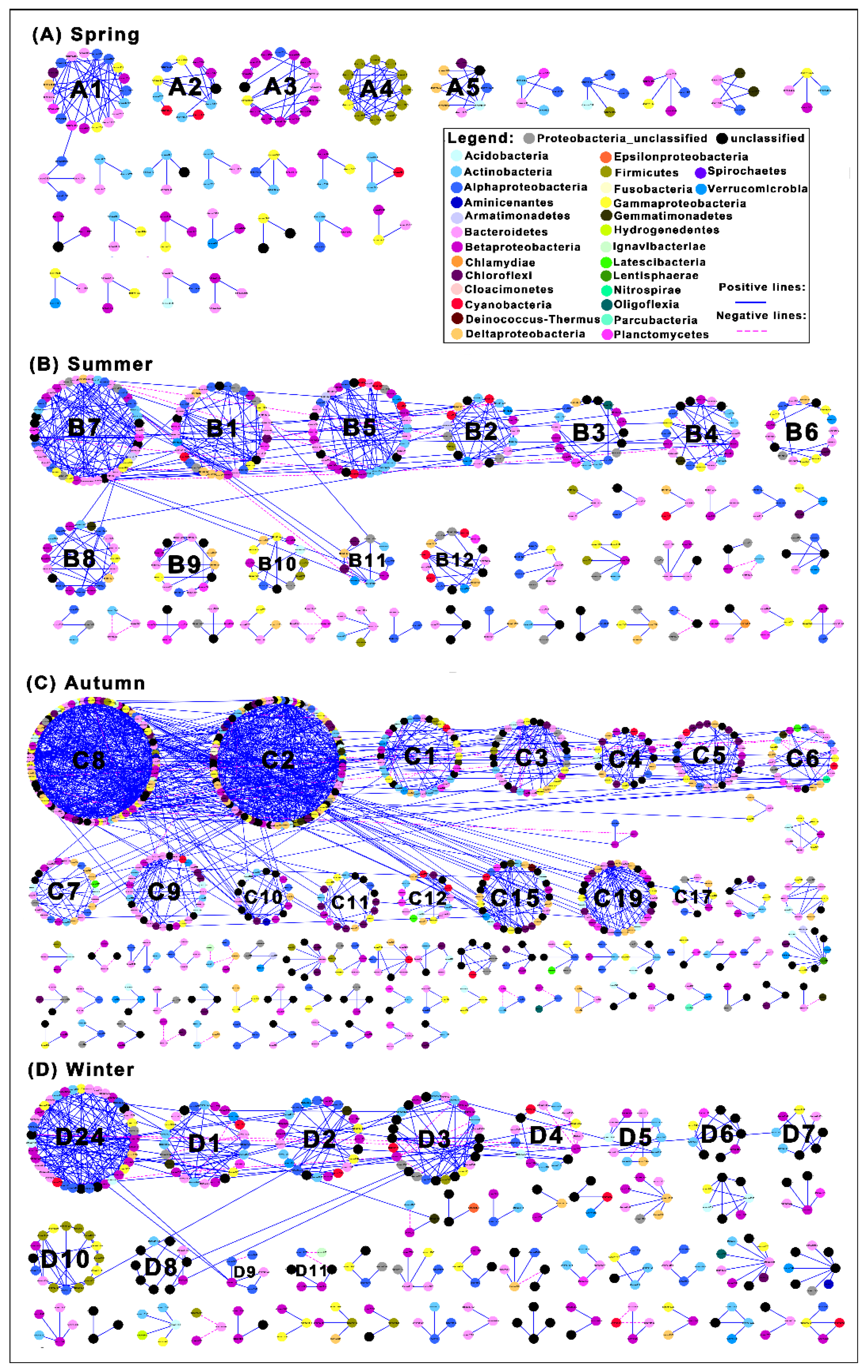
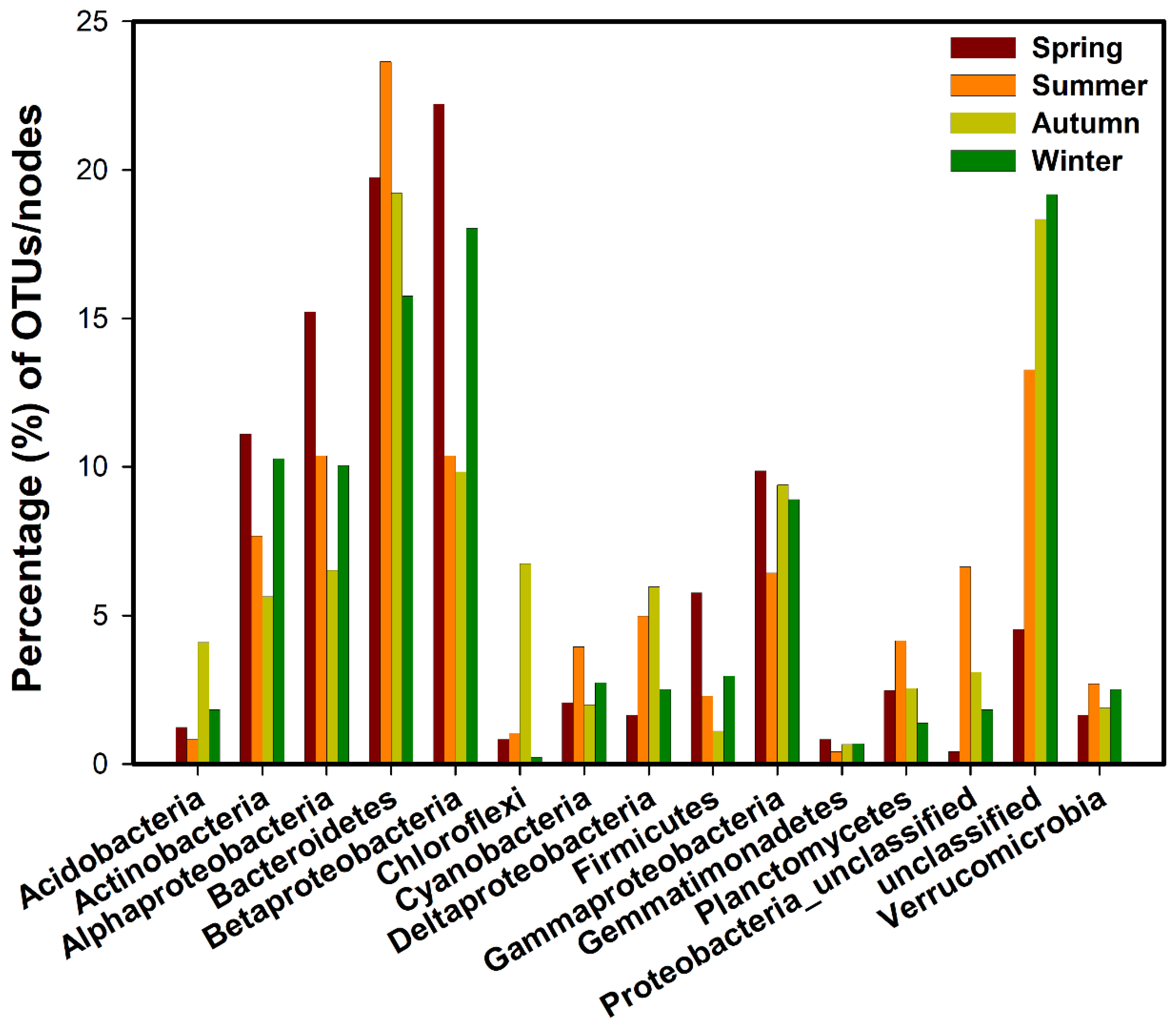
3.1. Network Structure of Bacterial Communities in Different Seasons
3.2. Co-Occurrence/Co-Exclusion Patterns in the Different Seasonal Networks
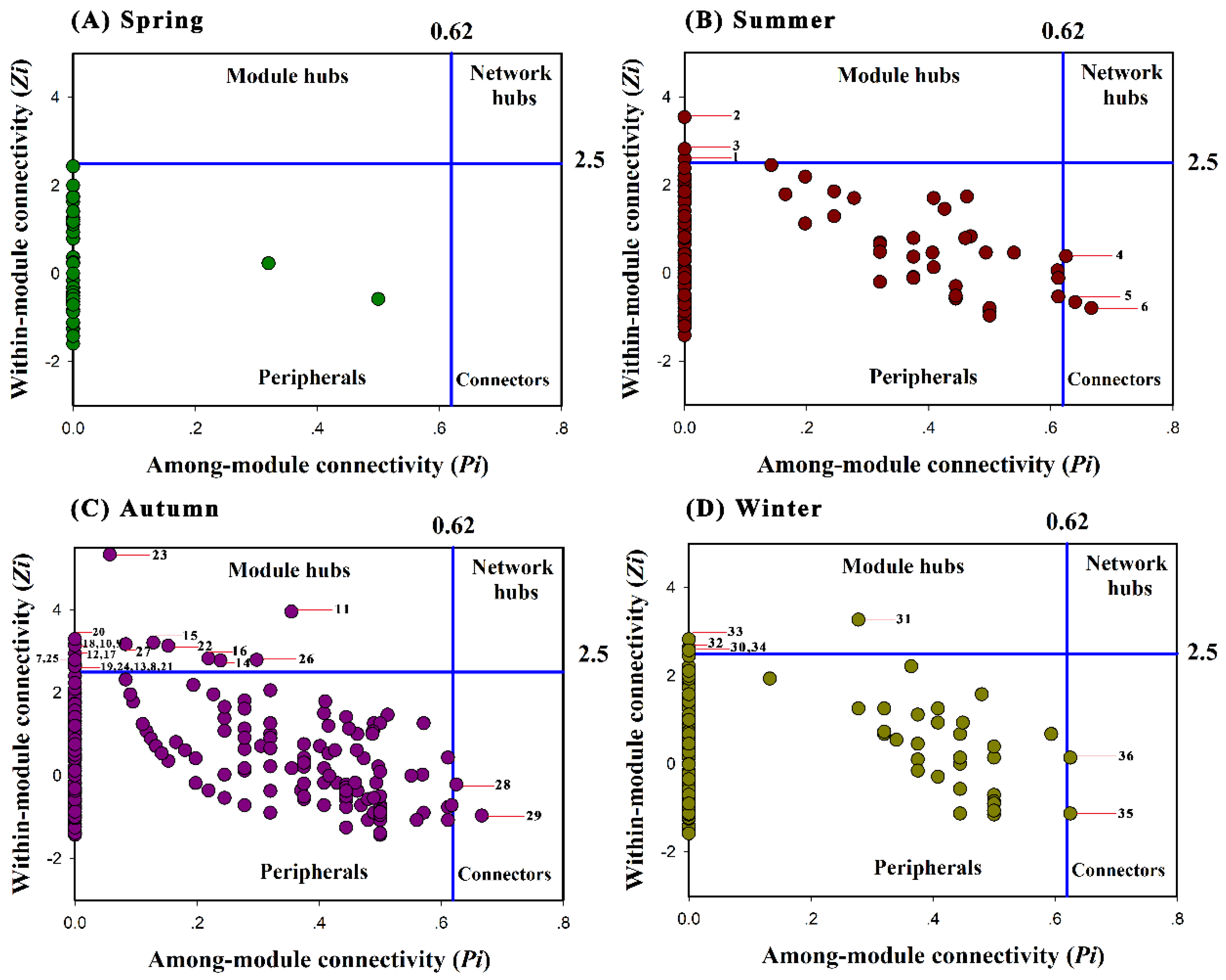
3.3. Topological Roles of Individual Nodes in Different Seasonal Networks
3.4. Relationships between Species and Environmental Variables in Different Seasons
4. Discussion
4.1. Interactions among Bacterial Taxa in the Correlation Networks Varied Remarkably between Seasons
4.2. Different Topological Roles of Individual OTUs Observed in Bacterial Community Networks
4.3. Relationships between Species and Environmental Variables Varied between Seasons
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- FAO. The State of World Fisheries and Aquaculture 2018—Meeting the Sustainable Development Goals. 2018. Available online: http://www.fao.org/3/i9540en/I9540EN.pdf (accessed on 12 July 2018).
- Cai, C.; Gu, X.; Ye, Y.; Yang, C.; Dai, X.; Chen, D.; Yang, C. Assessment of pollutant loads discharged from aquaculture ponds around Taihu Lake, China. Aquac. Res. 2013, 44, 795–806. [Google Scholar] [CrossRef]
- Fang, Y.; Li, H.; He, H. Dynamic Monitoring of Lake Reclamation in the Taihu Lake and Lake Enclosure Culture of the East Taihu Lake in Recent 30 Years. Resour. Environ. Yangtze Basin 2012, 21, 121–126. [Google Scholar]
- Qin, B.; Xu, P.; Wu, Q.; Luo, L.; Zhang, Y. Environmental issues of lake Taihu, China. In Eutrophication of Shallow Lakes with Special Reference to Lake Taihu, China; Springer: Berlin/Heidelberg, Germany, 2007; pp. 3–14. [Google Scholar]
- Wu, Y.; Xiang, Y.; Wang, J.; Zhong, J.; He, J.-Z.; Wu, Q.L. Heterogeneity of archaeal and bacterial ammonia-oxidizing communities in Lake Taihu, China. Environ. Microbiol. Rep. 2010, 2, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Yin, Q.; Yue, D.; Peng, Y.; Liu, Y.; Xiao, L. Occurrence and Distribution of Antibiotic-resistant Bacteria and Transfer of Resistance Genes in Lake Taihu. Microbes Environ. 2013, 28, 479–486. [Google Scholar] [CrossRef]
- Grossart, H.-P.; Ploug, H. Microbial degradation of organic carbon and nitrogen on diatom aggregates. Limnol. Oceanogr. 2001, 46, 267–277. [Google Scholar] [CrossRef]
- Jiao, N.; Herndl, G.J.; Hansell, D.A.; Benner, R.; Kattner, G.; Wilhelm, S.W.; Kirchman, D.L.; Weinbauer, M.G.; Luo, T.; Chen, F.; et al. Microbial production of recalcitrant dissolved organic matter: Long-term carbon storage in the global ocean. Nat. Rev. Genet. 2010, 8, 593–599. [Google Scholar] [CrossRef] [PubMed]
- Weiss, S.; Van Treuren, W.; Lozupone, C.; Faust, K.; Friedman, J.; Deng, Y.; Xia, L.C.; Xu, Z.Z.; Ursell, L.; Alm, E.J.; et al. Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. ISME J. 2016, 10, 1669–1681. [Google Scholar] [CrossRef]
- Gilbert, J.A.; Field, D.; Swift, P.; Newbold, L.; Oliver, A.; Smyth, T.; Somerfield, P.J.; Huse, S.; Joint, I. The seasonal structure of microbial communities in the Western English Channel. Environ. Microbiol. 2009, 11, 3132–3139. [Google Scholar] [CrossRef]
- Gilbert, J.A.; Steele, J.A.; Caporaso, J.G.; Steinbrück, L.; Reeder, J.; Temperton, B.; Huse, S.; McHardy, A.C.; Knight, R.; Joint, I. Defining seasonal marine microbial community dynamics. ISME J. 2012, 6, 298–308. [Google Scholar] [CrossRef]
- Zhao, D.; Shen, F.; Zeng, J.; Huang, R.; Yu, Z.; Wu, Q.L. Network analysis reveals seasonal variation of co-occurrence correlations between Cyanobacteria and other bacterioplankton. Sci. Total Environ. 2016, 573, 817–825. [Google Scholar] [CrossRef]
- Nelson, C.E. Phenology of high-elevation pelagic bacteria: The roles of meteorologic variability, catchment inputs and thermal stratification in structuring communities. ISME J. 2009, 3, 13–30. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Cao, X.; Huang, R.; Zeng, J.; Xu, H.; Wang, S.; He, X.; Yu, Z.; Shen, F. The heterogeneity of composition and assembly processes of the microbial community between different nutrient loading lake zones in Taihu Lake. Appl. Microbiol. Biotechnol. 2017, 101, 5913–5923. [Google Scholar] [CrossRef] [PubMed]
- Jones, A.C.; Hambright, K.D.; Caron, D.A. Ecological patterns among bacteria and microbial eukaryotes derived from network analyses in a low-salinity lake. Microbial Ecol. 2018, 75, 917–929. [Google Scholar] [CrossRef] [PubMed]
- Kandel, P.P.; Pasternak, Z.; Nahum, O.; Van Rijn, J.; Jurkevitch, E. Abundance, diversity and seasonal dynamics of predatory bacteria in aquaculture zero discharge systems. FEMS Microbiol. Ecol. 2014, 89, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, T. Effects of seasonal temperature variation on nitrification, anammox process, and bacteria involved in a pilot-scale constructed wetland. Environ. Sci. Pollut. R. 2015, 22, 3774–3783. [Google Scholar] [CrossRef] [PubMed]
- Denef, V.J.; Fujimoto, M.; Berry, M.A.; Schmidt, M.L. Seasonal Succession Leads to Habitat-Dependent Differentiation in Ribosomal RNA:DNA Ratios among Freshwater Lake Bacteria. Front. Microbiol. 2016, 7, 32. [Google Scholar] [CrossRef]
- Giovannoni, S.J.; Vergin, K.L. Seasonality in Ocean Microbial Communities. Science 2012, 335, 671–676. [Google Scholar] [CrossRef]
- Williams, R.J.; Howe, A.; Hofmockel, K.S. Demonstrating microbial co-occurrence pattern analyses within and between ecosystems. Front. Microbiol. 2014, 5, 358. [Google Scholar] [CrossRef]
- Ings, T.C.; Montoya, J.M.; Bascompte, J.; Blüthgen, N.; Brown, L.; Dormann, C.F.; Edwards, F.; Figueroa, D.; Jacob, U.; Jones, J.I. Ecological networks–beyond food webs. J. Anim. Ecol. 2009, 78, 253–269. [Google Scholar] [CrossRef]
- Poulin, R. Network analysis shining light on parasite ecology and diversity. Trends Parasitol. 2010, 26, 492–498. [Google Scholar] [CrossRef]
- Proulx, S.; Promislow, D.; Phillips, P. Network thinking in ecology and evolution. Trends Ecol. Evol. 2005, 20, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Bascompte, J. Networks in ecology. Basic Appl. Ecol. 2007, 8, 485–490. [Google Scholar] [CrossRef]
- Montoya, J.M.; Pimm, S.L.; Solé, R.V. Ecological networks and their fragility. Nature 2006, 442, 259–264. [Google Scholar] [CrossRef] [PubMed]
- Zeng, J.; Lin, Y.; Zhao, D.; Huang, R.; Xu, H.; Jiao, C. Seasonality overwhelms aquacultural activity in determining the composition and assembly of the bacterial community in Lake Taihu, China. Sci. Total Environ. 2019, 683, 427–435. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Magoč, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef]
- Zhao, D.; Xu, H.; Zeng, J.; Cao, X.; Huang, R.; Shen, F.; Yu, Z. Community composition and assembly processes of the free-living and particle-attached bacteria in Taihu Lake. FEMS Microbiol. Ecol. 2017, 93, fix062. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naïve Bayesian Classifier for Rapid Assignment of rRNA Sequences into the New Bacterial Taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Bittinger, K.; Bushman, F.D.; DeSantis, T.Z.; Andersen, G.L.; Knight, R. PyNAST: A flexible tool for aligning sequences to a template alignment. Bioinformatics 2010, 26, 266–267. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—Approximately Maximum-Likelihood Trees for Large Alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Subramanian, S.; Faith, J.J.; Gevers, D.; Gordon, J.I.; Knight, R.; Mills, D.A.; Caporaso, J.G. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 2013, 10, 57–59. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Zhao, D.; Huang, R.; Cao, X.; Zeng, J.; Yu, Z.; Hooker, K.V.; Hambright, K.D.; Wu, Q.L. Contrasting Network Features between Free-Living and Particle-Attached Bacterial Communities in Taihu Lake. Microb. Ecol. 2018, 76, 303–313. [Google Scholar] [CrossRef]
- Zhou, J.; Deng, Y.; Luo, F.; He, Z.; Yang, Y. Phylogenetic Molecular Ecological Network of Soil Microbial Communities in Response to Elevated CO2. mBio 2011, 2, e00122-11. [Google Scholar] [CrossRef]
- Harrell, F.E., Jr.; Dupont, C. Hmisc: Harrell Miscellaneous. R Package Version. 2008. Available online: http://ftp.auckland.ac.nz/software/CRAN/contrib/main/Descriptions/Hmisc.html (accessed on 7 September 2019).
- Cao, X.; Zhao, D.; Xu, H.; Huang, R.; Zeng, J.; Yu, Z. Heterogeneity of interactions of microbial communities in regions of Taihu Lake with different nutrient loadings: A network analysis. Sci. Rep. 2018, 8, 8890. [Google Scholar] [CrossRef]
- Junker, B.H.; Schreiber, F. Analysis of Biological Networks; Wiley-Interscience: Hoboken, NJ, USA, 2008; Volume 2, pp. 31–59. [Google Scholar]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Su, G.; Morris, J.H.; Demchak, B.; Bader, G.D. Biological Network Exploration with Cytoscape 3. Curr. Protoc. Bioinform. 2014, 47, 8.13.1–8.13.24. [Google Scholar] [CrossRef] [PubMed]
- Clauset, A.; Newman, M.E.J.; Moore, C. Finding community structure in very large networks. Phys. Rev. E 2004, 70, 066111. [Google Scholar] [CrossRef] [PubMed]
- Csardi, G.; Nepusz, T. The igraph software package for complex network research. InterJ. Complex Syst. 2006, 1695, 1–9. [Google Scholar]
- Guimerà, R.; Amaral, L.A.N. Functional cartography of complex metabolic networks. Nature 2005, 433, 895–900. [Google Scholar] [CrossRef] [PubMed]
- Olesen, J.M.; Bascompte, J.; Dupont, Y.L.; Jordano, P. The modularity of pollination networks. Proc. Natl. Acad. Sci. USA 2007, 104, 19891–19896. [Google Scholar] [CrossRef]
- Olafsen, J.A. Interactions between fish larvae and bacteria in marine aquaculture. Aquaculture 2001, 200, 223–247. [Google Scholar] [CrossRef]
- Blancheton, J.; Attramadal, K.; Michaud, L.; D’Orbcastel, E.R.; Vadstein, O. Insight into bacterial population in aquaculture systems and its implication. Aquac. Eng. 2013, 53, 30–39. [Google Scholar] [CrossRef]
- Lewinsohn, T.M.; Prado, P.I.; Jordano, P.; Bascompte, J.; Olesen, J.M. Structure in plant-animal interaction assemblages. Oikos 2006, 113, 174–184. [Google Scholar] [CrossRef]
- Mougi, A.; Kondoh, M.; Hurtgen, M.T. Diversity of Interaction Types and Ecological Community Stability. Science 2012, 337, 349–351. [Google Scholar] [CrossRef]
- Zou, L.; Pei, W.; Li, T.; He, Z.; Cheung, Y. Topological fractal networks introduced by mixed degree distribution. Phys. A Stat. Mech. Its Appl. 2007, 380, 592–600. [Google Scholar] [CrossRef]
- May, R.; McLean, A.R. Theoretical Ecology: Principles and Applications; Oxford University Press on Demand: Oxford, UK, 2007. [Google Scholar]
- Tang, X.; Chao, J.; Gong, Y.; Wang, Y.; Wilhelm, S.W.; Gao, G. Spatiotemporal dynamics of bacterial community composition in large shallow eutrophic Lake Taihu: High overlap between free-living and particle-attached assemblages. Limnol. Oceanogr. 2017, 62, 1366–1382. [Google Scholar] [CrossRef]
- Paerl, H.W.; Xu, H.; Hall, N.S.; Rossignol, K.L.; Joyner, A.R.; Zhu, G.; Qin, B. Nutrient limitation dynamics examined on a multi-annual scale in Lake Taihu, China: Implications for controlling eutrophication and harmful algal blooms. J. Freshw. Ecol. 2015, 30, 5–24. [Google Scholar] [CrossRef]
- Zhou, J.; Deng, Y.; Zhang, P.; Xue, K.; Liang, Y.; Van Nostrand, J.D.; Yang, Y.; He, Z.; Wu, L.; Stahl, D.A.; et al. Stochasticity, succession, and environmental perturbations in a fluidic ecosystem. Proc. Natl. Acad. Sci. USA 2014, 111, E836–E845. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Yang, Y.; Chen, S.; Zhao, M.; Zhu, Z.; Yang, S.; Qu, Y.; Ma, Q.; He, Z.; Zhou, J.; et al. Long-term successional dynamics of microbial association networks in anaerobic digestion processes. Water Res. 2016, 104, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Eiler, A.; Heinrich, F.; Bertilsson, S. Coherent dynamics and association networks among lake bacterioplankton taxa. ISME J. 2012, 6, 330–342. [Google Scholar] [CrossRef]
- Newton, R.J.; Jones, S.E.; Eiler, A.; McMahon, K.D.; Bertilsson, S. A Guide to the Natural History of Freshwater Lake Bacteria. Microbiol. Mol. Boil. Rev. 2011, 75, 14–49. [Google Scholar] [CrossRef] [PubMed]
- Williams, K.P.; Sobral, B.W.; Dickerman, A.W. A Robust Species Tree for the Alphaproteobacteria. J. Bacteriol. 2007, 189, 4578–4586. [Google Scholar] [CrossRef]
- Eiler, A.; Bertilsson, S. Flavobacteria Blooms in Four Eutrophic Lakes: Linking Population Dynamics of Freshwater Bacterioplankton to Resource Availability. Appl. Environ. Microbiol. 2007, 73, 3511–3518. [Google Scholar] [CrossRef]
- Yannarell, A.C.; Triplett, E.W. Geographic and Environmental Sources of Variation in Lake Bacterial Community Composition. Appl. Environ. Microbiol. 2005, 71, 227–239. [Google Scholar] [CrossRef]
- Lindström, E.S.; Langenheder, S. Local and regional factors influencing bacterial community assembly. Environ. Microbiol. Rep. 2012, 4, 1–9. [Google Scholar] [CrossRef]
- Beisner, B.E.; Peres-Neto, P.R.; Lindström, E.S.; Barnett, A.; Longhi, M.L. The Role of Environmental and Spatial Processes in Structuring Lake Communities from Bacteria to Fish. Ecology 2006, 87, 2985–2991. [Google Scholar] [CrossRef]
- Cottenie, K. Integrating environmental and spatial processes in ecological community dynamics. Ecol. Lett. 2005, 8, 1175–1182. [Google Scholar] [CrossRef] [PubMed]
- Kara, E.L.; Hanson, P.C.; Hu, Y.H.; Winslow, L.; McMahon, K.D. A decade of seasonal dynamics and co-occurrences within freshwater bacterioplankton communities from eutrophic Lake Mendota, WI, USA. ISME J. 2013, 7, 680–684. [Google Scholar] [CrossRef] [PubMed]
- Bowers, R.M.; McCubbin, I.B.; Hallar, A.G.; Fierer, N. Seasonal variability in airborne bacterial communities at a high-elevation site. Atmos. Environ. 2012, 50, 41–49. [Google Scholar] [CrossRef]
- Zhu, W.; Tan, Y.; Wang, R.; Feng, G.; Chen, H.; Liu, Y.; Li, M. The trend of water quality variation and analysis in typical area of Lake Taihu, 2010–2017. J. Lake Sci. 2018, 30, 296–305. [Google Scholar]
- Zhu, C.; Zhang, J.; Nawaz, M.Z.; Mahboob, S.; Al-Ghanim, K.A.; Khan, I.A.; Lu, Z.; Chen, T. Seasonal succession and spatial distribution of bacterial community structure in a eutrophic freshwater Lake, Lake Taihu. Sci. Total Environ. 2019, 669, 29–40. [Google Scholar] [CrossRef] [PubMed]
- Price, P.B.; Sowers, T. Temperature dependence of metabolic rates for microbial growth, maintenance, and survival. Proc. Natl. Acad. Sci. USA 2004, 101, 4631–4636. [Google Scholar] [CrossRef]
| Empirical Network | Random Network | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Groups | Number of nodes | Number of edges | Modularity | Clustering coefficient | Average path length | Network diameter | Average degree | Graph density | Modularity (SD) | Clustering coefficient (SD) | Average path length (SD) | Network diameter (SD) | |
| Positive | Negative | ||||||||||||
| Spring | 243 | 225 | 0 | 0.931 a | 0.193 a | 2.492 a | 8 a | 1.852 | 0.008 | 0.773 (0.016) | 0.007 (0.009) | 7.488 (0.573) | 18.536 (2.400) |
| Summer | 482 | 521 | 64 | 0.852 a | 0.251 a | 6.445 b | 17 b | 2.427 | 0.005 | 0.699 (0.009) | 0.005 (0.004) | 6.604 (0.166) | 15.627 (1.365) |
| Autumn | 905 | 1594 | 102 | 0.739 a | 0.177 a | 6.994 a | 22 a | 3.748 | 0.004 | 0.540 (0.004) | 0.004 (0.002) | 5.240 (0.033) | 11.322 (0.792) |
| Winter | 435 | 440 | 68 | 0.851 a | 0.196 a | 6.723 b | 17 b | 2.320 | 0.005 | 0.714 (0.010) | 0.005 (0.004) | 6.759 (0.193) | 16.230 (1.530) |
| Season | Type of Point | Number of Node | OTU ID | Number of Module | Mean Abundance (%) | Phylum/Class | Lowest Taxonomic Rank |
|---|---|---|---|---|---|---|---|
| Summer | Module hubs | 1 | denovo3272 | B8 | 0.006 | Alphaproteobacteria | g__Sphingorhabdus |
| 2 | denovo4293 | B1 | 0.013 | Unclassified | k__Bacteria | ||
| 3 | denovo10386 | B6 | 0.003 | Alphaproteobacteria | f__Sphingomonadaceae | ||
| Connectors | 4 | denovo830 | B1 | 0.014 | Chloroflexi | g__Oscillochloris | |
| 5 | denovo7514 | B2 | 0.095 | Deltaproteobacteria | g__Labilithrix | ||
| 6 | denovo12061 | B3 | 0.030 | Alphaproteobacteria | f__Acetobacteraceae | ||
| Autumn | Module hubs | 7 | denovo113 | C9 | 0.002 | Proteobacteria_unclassified | p__Proteobacteria |
| 8 | denovo1308 | C24 | 0.035 | Gammaproteobacteria | o__Chromatiales | ||
| 9 | denovo1615 | C10 | 0.002 | Lentisphaerae | g__Oligosphaera | ||
| 10 | denovo3226 | C6 | 0.009 | Bacteroidetes | g__Fluviicola | ||
| 11 | denovo3806 | C33 | 0.010 | Unclassified | k__Bacteria | ||
| 12 | denovo4788 | C14 | 0.002 | Bacteroidetes | p__Bacteroidetes | ||
| 13 | denovo6589 | C7 | 0.010 | Bacteroidetes | g__Terrimonas | ||
| 14 | denovo7064 | C23 | 0.009 | Betaproteobacteria | c__Betaproteobacteria | ||
| 15 | denovo8549 | C29 | 0.089 | Acidobacteria | o__Gp6 | ||
| 16 | denovo8853 | C8 | 0.014 | Unclassified | k__Bacteria | ||
| 17 | denovo9608 | C14 | 0.002 | Bacteroidetes | p__Bacteroidetes | ||
| 18 | denovo9867 | C9 | 0.005 | Cloacimonetes | c__Candidatus_Cloacamonas | ||
| 19 | denovo9976 | C13 | 0.003 | Chloroflexi | f__Caldilineaceae | ||
| 20 | denovo10529 | C8 | 0.005 | Unclassified | k__Bacteria | ||
| 21 | denovo10588 | C8 | 0.017 | Bacteroidetes | p__Bacteroidetes | ||
| 22 | denovo10876 | C12 | 0.005 | Gemmatimonadetes | g__Gemmatimonas | ||
| 23 | denovo10885 | C34 | 0.005 | Unclassified | k__Bacteria | ||
| 24 | denovo11435 | C8 | 0.003 | Alphaproteobacteria | f__Caulobacteraceae | ||
| 25 | denovo11533 | C9 | 0.002 | Unclassified | k__Bacteria | ||
| 26 | denovo11626 | C11 | 0.006 | Chloroflexi | f__Anaerolineaceae | ||
| 27 | denovo11710 | C23 | 0.003 | Gammaproteobacteria | c__Gammaproteobacteria | ||
| Connectors | 28 | denovo715 | C10 | 0.009 | Unclassified | k__Bacteria | |
| 29 | denovo5057 | C3 | 0.003 | Bacteroidetes | o__Cytophagales | ||
| Winter | Module hubs | 30 | denovo95 | D3 | 0.018 | Gammaproteobacteria | c__Gammaproteobacteria |
| 31 | denovo2361 | D4 | 0.315 | Betaproteobacteria | f__Methylophilaceae | ||
| 32 | denovo2444 | D16 | 0.004 | Unclassified | k__Bacteria | ||
| 33 | denovo3884 | D15 | 0.003 | Bacteroidetes | f__Chitinophagaceae | ||
| 34 | denovo13572 | D3 | 0.087 | Actinobacteria | o__Actinomycetales | ||
| Connectors | 35 | denovo3821 | D24 | 0.623 | Alphaproteobacteria | f__Rhodobacteraceae | |
| 36 | denovo9640 | D4 | 0.004 | Alphaproteobacteria | g__Caulobacter |
| Season | T (°C) | pH | DO (mg/L) | SD (m) | Chla (μg/L) | TN (mg/L) | TP (mg/L) | DOC (mg/L) | NH4+-N (mg/L) | NO3−-N (mg/L) | NO2−-N (mg/L) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Spring | 17.19 ± 0.11 | 8.84 ± 0.60 | 7.57 ± 0.96 | 1.26 ± 0.23 | 2.30 ± 2.38 | 0.80 ± 0.15 | 0.072 ± 0.045 | 7.35 ± 0.86 | 0.074 ± 0.038 | 0.165 ± 0.138 | 0.010 ± 0.004 |
| Summer | 31.60 ± 1.02 | 8.23 ± 0.40 | 4.02 ± 1.24 | 0.38 ± 0.11 | 59.05 ± 26.72 | 1.25 ± 0.54 | 0.127 ± 0.035 | 5.52 ± 1.19 | 0.021 ± 0.008 | 0.279 ± 0.078 | 0.003 ± 0.002 |
| Autumn | 19.61 ± 0.13 | 8.25 ± 0.25 | 7.43 ± 0.59 | 0.42 ± 0.10 | 7.98 ± 3.25 | 0.64 ± 0.07 | 0.031 ± 0.005 | 9.86 ± 2.28 | 0.058 ± 0.019 | 0.067 ± 0.029 | 0.004 ± 0.002 |
| Winter | 3.36 ± 0.36 | 7.96 ± 0.11 | 7.19 ± 0.79 | 0.61 ± 0.34 | 6.47 ± 2.01 | 1.16 ± 0.22 | 0.025 ± 0.006 | 11.32 ± 2.84 | 0.099 ± 0.026 | 0.456 ± 0.142 | 0.006 ± 0.004 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, Y.; Zhao, D.; Zeng, J.; Cao, X.; Jiao, C. Network Analysis Reveals Seasonal Patterns of Bacterial Community Networks in Lake Taihu under Aquaculture Conditions. Water 2019, 11, 1868. https://doi.org/10.3390/w11091868
Lin Y, Zhao D, Zeng J, Cao X, Jiao C. Network Analysis Reveals Seasonal Patterns of Bacterial Community Networks in Lake Taihu under Aquaculture Conditions. Water. 2019; 11(9):1868. https://doi.org/10.3390/w11091868
Chicago/Turabian StyleLin, Yuqing, Dayong Zhao, Jin Zeng, Xinyi Cao, and Congcong Jiao. 2019. "Network Analysis Reveals Seasonal Patterns of Bacterial Community Networks in Lake Taihu under Aquaculture Conditions" Water 11, no. 9: 1868. https://doi.org/10.3390/w11091868
APA StyleLin, Y., Zhao, D., Zeng, J., Cao, X., & Jiao, C. (2019). Network Analysis Reveals Seasonal Patterns of Bacterial Community Networks in Lake Taihu under Aquaculture Conditions. Water, 11(9), 1868. https://doi.org/10.3390/w11091868






