The Use of Algae and Fungi for Removal of Pharmaceuticals by Bioremediation and Biosorption Processes: A Review
Abstract
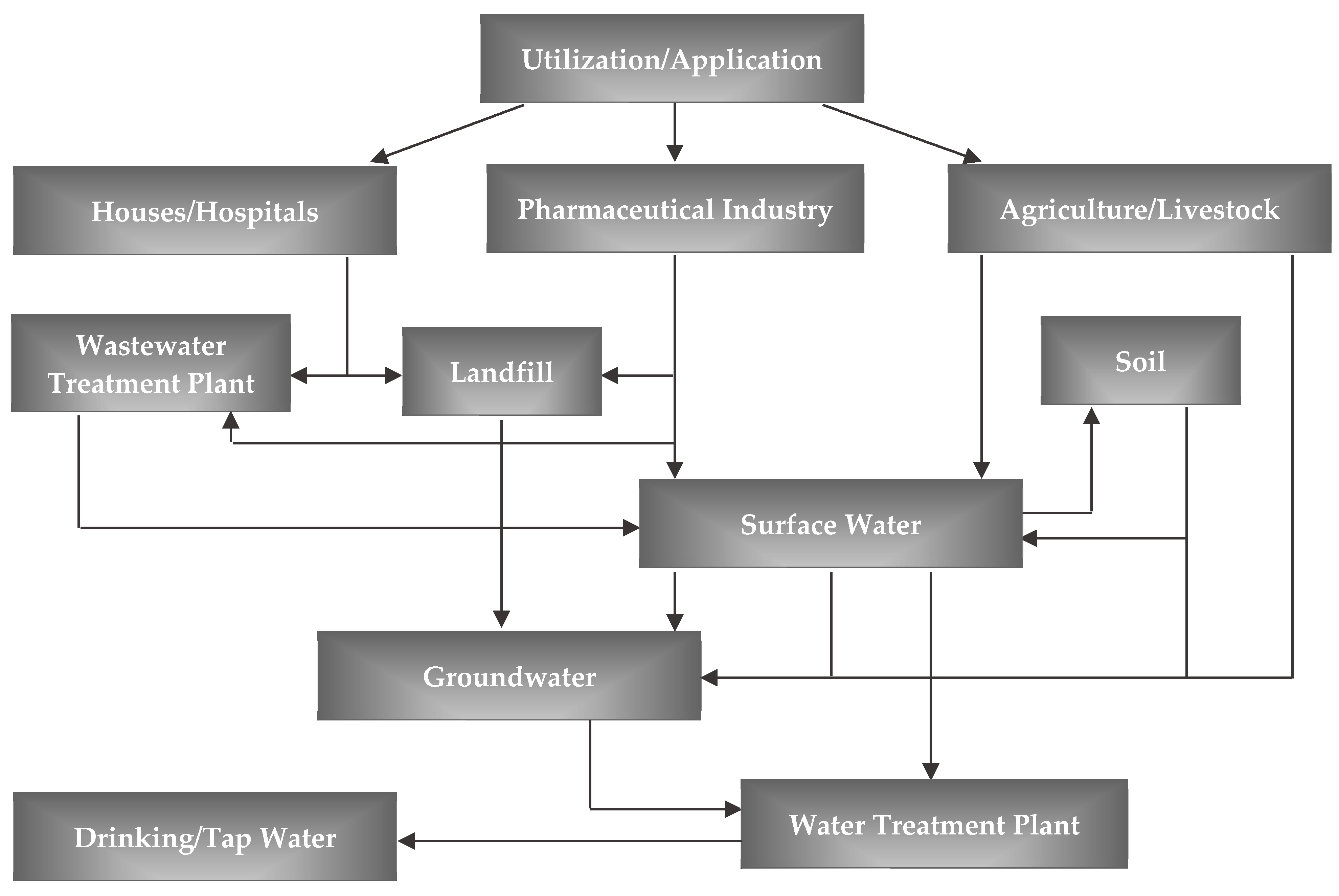
1. Introduction
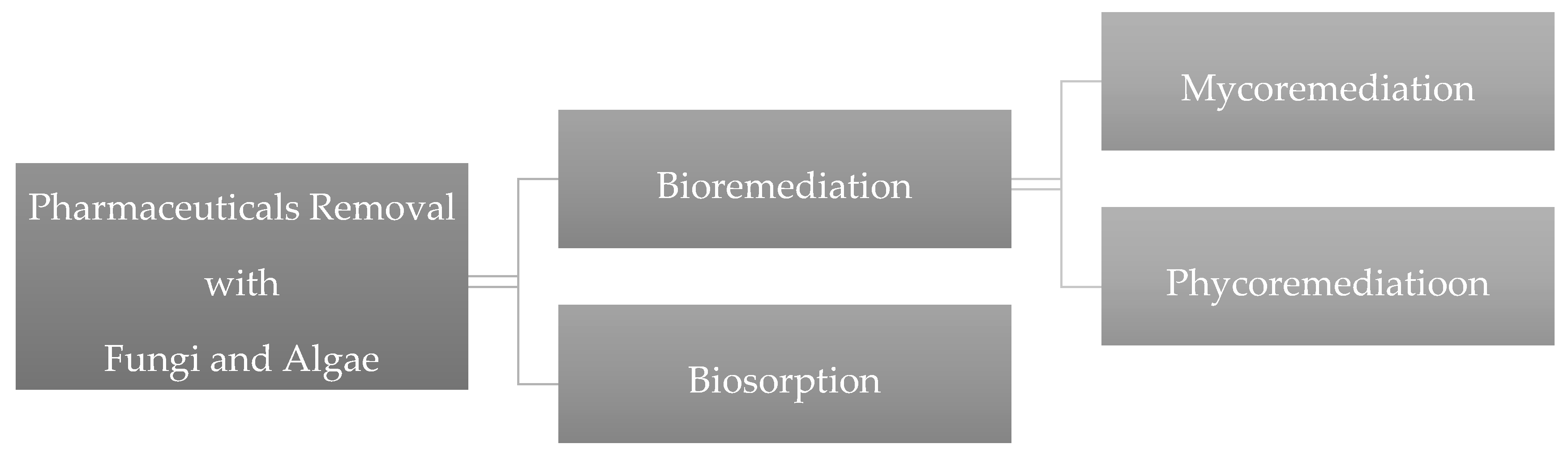
2. Bioremediation
2.1. Mycoremediation
2.1.1. Treatment Systems
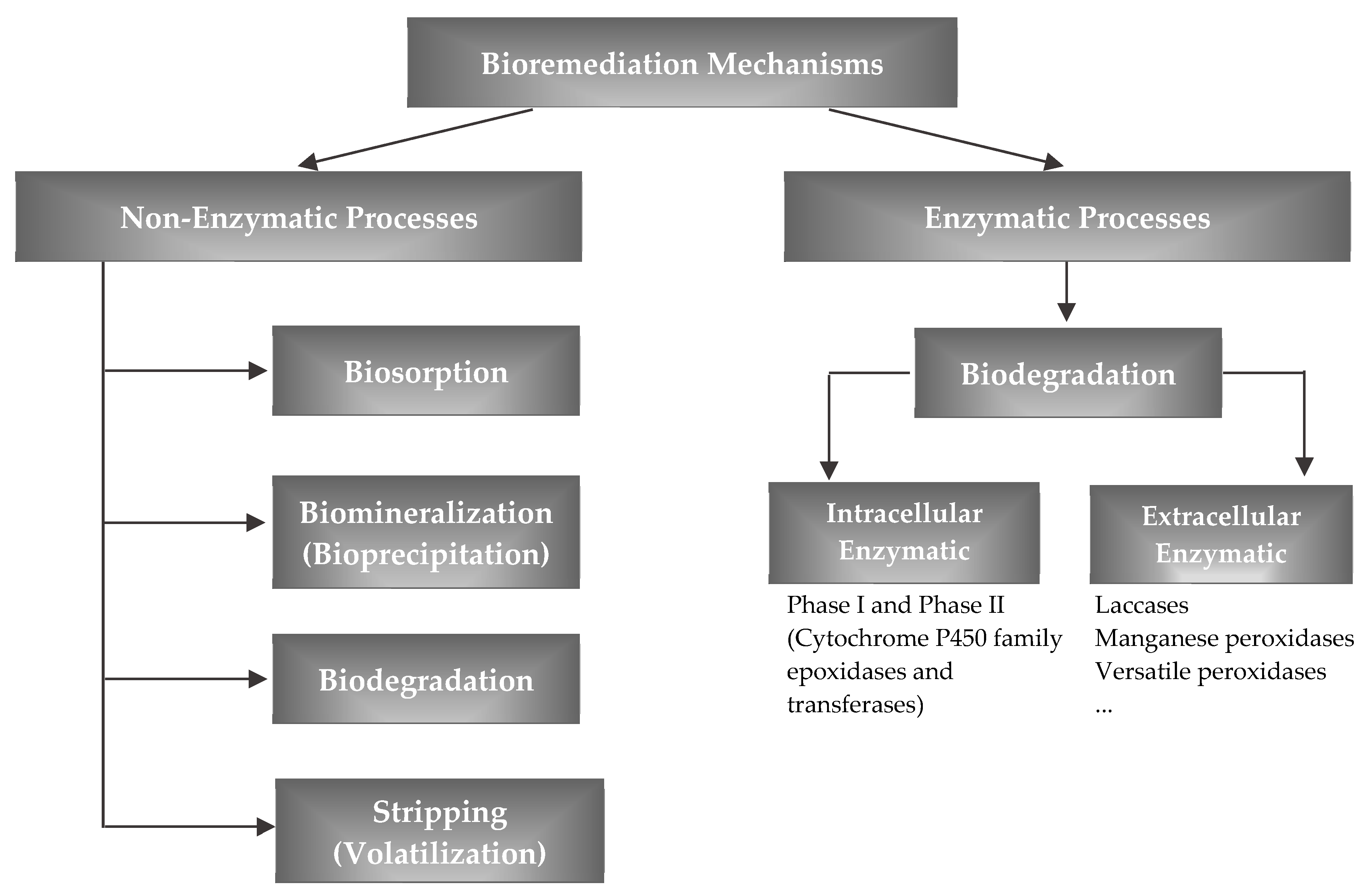
2.1.2. Mechanisms of Removal
2.1.3. Factors That Influence the Degradation Capability
2.1.4. Concluding Remarks and Future Challenges for Mycoremediation
2.2. Phycoremediation
2.2.1. Treatment Systems
2.2.2. Mechanisms of Removal
2.2.3. Factors That Influence the Degradation Capability
2.2.4. Concluding Remarks and Future Challenges
3. Biosorption
3.1. Biosorption Materials
3.2. Treatment Systems
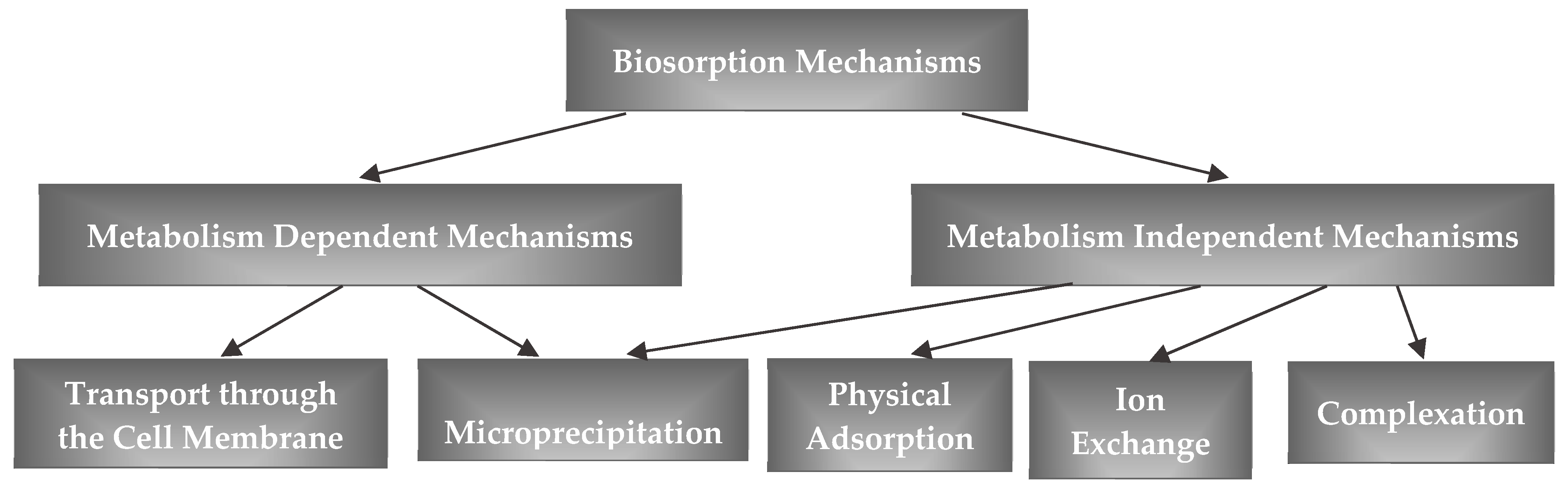
3.3. Mechanisms of Removal
3.4. Factors That Affect the Process
3.5. Biosorption Potential as a Wastewater Treatment Technology
4. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Arnold, K.E.; Brown, A.R.; Ankley, G.T.; Sumpter, J.P. Medicating the environment: Assessing risks of pharmaceuticals to wildlife and ecosystems. Philos. Trans. R. Soc. B Boil. Sci. 2014, 369. [Google Scholar] [CrossRef] [PubMed]
- Oulton, R.L.; Kohn, T.; Cwiertny, D.M. Pharmaceuticals and personal care products in effluent matrices: A survey of transformation and removal during wastewater treatment and implications for wastewater management. J. Environ. Monit. 2010, 12, 1956–1978. [Google Scholar] [CrossRef] [PubMed]
- Paíga, P.; Santos, L.H.; Amorim, C.G.; Araújo, A.N.; Montenegro, M.C.; Pena, A.; Delerue-Matos, C. Pilot monitoring study of ibuprofen in surface waters of north of Portugal. Environ. Sci. Pollut. Res. 2013, 20, 2410–2420. [Google Scholar] [CrossRef] [PubMed]
- Paíga, P.; Santos, L.H.M.L.M.; Ramos, S.; Jorge, S.; Silva, J.G.; Delerue-Matos, C. Presence of pharmaceuticals in the Lis river (Portugal): Sources, fate and seasonal variation. Sci. Total Environ. 2016, 573, 164–177. [Google Scholar] [CrossRef] [PubMed]
- Pomati, F.; Castiglioni, S.; Zuccato, E.; Fanelli, R.; Vigetti, D.; Rossetti, C.; Calamari, D. Effects of a complex mixture of therapeutic drugs at environmental levels on human embryonic cells. Environ. Sci. Technol. 2006, 40, 2442–2447. [Google Scholar] [CrossRef] [PubMed]
- Boxall, A.B.A.; Rudd, M.A.; Brooks, B.W.; Caldwell, D.J.; Choi, K.; Hickmann, S.; Innes, E.; Ostapyk, K.; Staveley, J.P.; Verslycke, T.; et al. Pharmaceuticals and personal care products in the environment: What are the big questions? Environ. Health. Perspect. 2012, 120, 1221–1229. [Google Scholar] [CrossRef] [PubMed]
- Directive 2000/60/EC of the European Parliament and of the Council of 23 October 2000 Establishing a Framework for Community Action in the Field of Water Policy. Available online: https://eur-lex.europa.eu/resource.html?uri=cellar:5c835afb-2ec6-4577-bdf8-756d3d694eeb.0004.02/DOC_1&format=PDF (accessed on 4 June 2019).
- Decision No 2455/2001/EC of the European Parliament and of the Council of 20 November 2001 Establishing the List of Priority Substances in the Field of Water Policy, and Amending Directive 2000/60/EC. Available online: http://ec.europa.eu/environment/ecolabel/documents/prioritysubstances.pdf (accessed on 4 June 2019).
- Directive 2008/105/EC of the European Parliament and of the Council of 16 December 2008 on Environmental Quality Standards in the Field of Water Policy, Amending and Subsequently Repealing Council Directives 82/176/EEC, 83/513/EEC, 84/156/EEC, 84/491/EEC, 86/280/EEC and Amending Directive 2000/60/EC of the European Parliament and of the Council. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:02008L0105-20130913&from=DE (accessed on 4 June 2019).
- Directive 2013/39/EU of the European Parliament and of the Council of 12 August 2013 Amending Directives 2000/60/EC and 2008/105/EC as Regards Priority Substances in the Field of Water Policy. Available online: http://eur-lex.europa.eu/LexUriServ/LexUriServ.do?uri=OJ:L:2013:226:0001:0017:EN:PDF (accessed on 4 June 2019).
- Commission Implementing Decision (EU) 2015/495 of 20 March 2015 Establishing a Watch list of Substances for Union-Wide Monitoring in the Field of Water Policy Pursuant to Directive 2008/105/EC of the European Parliament and of the Council (Notified Under Document C(2015) 1756). Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:32015D0495&from=PT (accessed on 4 June 2019).
- Commission Implementing Decision (EU) 2018/840 of 5 June 2018 Establishing a Watch List of Substances for Union-Wide Monitoring in the Field of Water Policy Pursuant to Directive 2008/105/EC of the European Parliament and of the Council and repealing Commission Implementing Decision (EU) 2015/495. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:32018D0840&rid=7 (accessed on 4 June 2019).
- Deziel, N. Pharmaceuticals in wastewater treatment plant effluent waters. Sch. Horiz. Univ. Minn. Morris Undergrad. J. 2014, 1, 12. [Google Scholar]
- Moller, P.; Dulski, P.; Bau, M.; Knappe, A.; Pekdeger, A.; Sommer-von Jarmerasted, C. Anthropogenic gadolinium as a conservative tracer in hydrology. J. Geochem. Explor. 2000, 69, 409–414. [Google Scholar] [CrossRef]
- Buerge, I.J.; Poiger, T.; Muller, M.D.; Buser, H.R. Caffeine, an anthropogenic marker for wastewater contamination of surface waters. Environ. Sci. Technol. 2003, 37, 691–700. [Google Scholar] [CrossRef]
- Martínez-Carballo, E.; González-Barreiro, C.; Scharf, S.; Gans, O. Environmental monitoring study of selected veterinary antibiotics in animal manure and soils in Austria. Environ. Pollut. 2007, 148, 570–579. [Google Scholar] [CrossRef]
- Kim, K.R.; Owens, G.; Kwon, S.I.; So, K.H.; Lee, D.B.; Ok, Y.S. Occurrence and Environmental Fate of Veterinary Antibiotics in the Terrestrial Environment. Water Air Soil Pollut. 2011, 214, 163–174. [Google Scholar] [CrossRef]
- Ji, K.; Kim, S.; Han, S.; Seo, J.; Lee, S.; Park, Y.; Choi, K.; Kho, Y.L.; Kim, P.G.; Park, J. Risk Assessment of chlortetracycline, oxytetracycline, sulfamethazine, sulfathiazole and erythromycin in aquatic environment. Are the current environmental concentrations safe? Ecotoxicology 2012, 21, 2031–2050. [Google Scholar] [CrossRef] [PubMed]
- Galichet, L.Y.; Moffat, A.C.; Osselton, M.D.; Widdop, B. Clarke´s Analysis of Drugs and Poisons, 3rd ed.; Pharmaceutical Press: London, UK, 2004. [Google Scholar]
- Verlicchi, P.; Al Aukidy, M.; Zambello, E. Occurrence of pharmaceutical compounds in urban wastewater: Removal, mass load and environmental risk after a secondary treatment—A review. Sci. Total Environ. 2012, 429, 123–155. [Google Scholar] [CrossRef] [PubMed]
- Saussereau, E.; Lacroix, C.; Guerbet, M.; Cellier, D.; Spiroux, J.; Goullé, J.P. Determination of levels of current drugs in hospital and urban wastewater. Bull. Environ. Contam. Toxicol. 2013, 91, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Petrovic, M.; Gonzalez, S.; Barceló, D. Analysis and removal of emerging contaminants in wastewater and drinking water. Trends Anal. Chem. 2003, 22, 685–696. [Google Scholar] [CrossRef]
- Hörsing, M.; Ledin, A.; Grabic, R.; Fick, J.; Tysklind, M.; Jansen, J.L.C.; Andersen, H.R. Determination of sorption of seventy-five pharmaceuticals in sewage sludge. Water Res. 2011, 45, 4470–4482. [Google Scholar] [CrossRef] [PubMed]
- Benotti, M.; Trenholm, R.A.; Vanderford, B.J.; Holady, H.C.; Stanford, B.D.; Snyder, S.A. Pharmaceuticals and endocrine disrupting compounds in U.S. drinking water. Environ. Sci. Technol. 2009, 43, 597–603. [Google Scholar] [CrossRef] [PubMed]
- Joss, A.; Siegrist, H.; Ternes, T.A. Are we about to upgrade wastewater treatment for removing organic micropollutants? Water Sci. Technol. 2008, 57, 251–255. [Google Scholar] [CrossRef]
- Wu, S.; Zhang, L.; Chen, J. Paracetamol in the Environment and its Degradation by Microorganisms. Appl. Microbiol. Biotechnol. 2012, 96, 875–884. [Google Scholar] [CrossRef]
- Sahu, O. Reduction of organic and inorganic pollutant from waste water by algae. Int. Lett. Nat. Sci. 2014, 8, 1–8. [Google Scholar] [CrossRef]
- Boopathy, R. Factors limiting bioremediation technologies. Bioresour. Technol. 2011, 74, 63–67. [Google Scholar] [CrossRef]
- Kartheek, B.R.; Maheswaran, R.; Kumar, G.; Banu, G.S. Biodegradation of Pharmaceutical Wastes Using Different Microbial Strains. Int. J. Pharm. Biol. Arch. 2011, 2, 1401–1404. [Google Scholar]
- Gillespie, I.M.M.; Philip, J.C. Bioremediation, an environmental remediation technology for the bioeconomy. Trends Biotechnol. 2013, 31, 329–332. [Google Scholar] [CrossRef] [PubMed]
- Misal, S.A.; Lingojwar, D.P.; Shinde, R.M.; Gawai, K.R. Purification and characterization of azoreductase from alkaliphilic strain Bacillus badius. Process Biochem. 2011, 46, 1264–1269. [Google Scholar] [CrossRef]
- Mashi, B.H. Biorremediation: Issues and Challenges. JORIND 2013, 11, 1596–8303. [Google Scholar]
- Andrade, J.A.; Augusto, F.; Jardim, I.C.S.F. Biorremediação de solos contaminados por petróleo e seus derivados. Jornal Eclética Química 2010, 35. [Google Scholar] [CrossRef]
- Gaylarde, C.C.; Bellinaso, M.L.; Manfilo, G.P. Biorremediação - aspetos biológicos e técnicos da biorremediação de xenobióticos. Biotecnol. Ciência Desenvolv. 2005, 34, 36–43. [Google Scholar]
- Alexopoulos, C.J.; Mims, C.W.; Blackwell, M. Introductory Mycology, 4th ed.; John Wiley: New York, NY, USA, 1996. [Google Scholar]
- Badia-Fabregat, M.; Lucas, D.; Gros, M.; Rodríguez-Mozaz, S.; Barceló, D.; Caminal, G.; Vicent, T. Identification of some factors affecting pharmaceutical active compounds (PhACs) removal in real wastewater. Case study of fungal treatment of reverse osmosis concentrate. J. Hazard. Mater. 2015, 283, 663–671. [Google Scholar] [CrossRef]
- Zhang, Y.; Xie, J.; Liu, M.; Tian, Z.; He, Z.; van Nostrand, J.D.; Ren, L.; Zhou, J.; Yang, M. Microbial community functional structure in response to antibiotics in pharmaceutical wastewater treatment systems. Water Res. 2013, 47, 6298–6308. [Google Scholar] [CrossRef]
- Durairaj, P.; Malla, S.; Nadarajan, S.P.; Lee, P.G.; Jung, E.; Park, H.H.; Kim, B.G.; Yun, H. Fungal cytochrome P450 monooxygenases of Fusarium oxysporum for the synthesis of ω-hydroxy fatty acids in engineered Saccharomyces cerevisiae. Microb. Cell Fact. 2015, 14, 45. [Google Scholar] [CrossRef]
- Jebapriya, G.R.; Gnanadoss, J.J. Bioremediation of textile dye using white-rot fungi: A review. Int. J. Curr. Res. Rev. 2013, 5, 1–13. [Google Scholar]
- Morel, M.; Meux, E.; Mathieu, Y.; Thuillier, A.; Chibani, K.; Harvengi, L.; Jacquot, J.P.; Gelhaye, E. Xenomic networks variability and adaptation traits in wood decaying fungi. Microb. Biotechnol. 2013, 6, 248–263. [Google Scholar] [CrossRef] [PubMed]
- Anastasi, A.; Tigini, V.; Varese, G.C. The bioremediation potential of different ecophysiological groups of fungi. In Fungi as Bioremediators; Goltapeh, E.M., Danesh, Y.R., Varma, A., Eds.; Springer-Verlag: New Delhi, India, 2013; Volume 32, pp. 29–49. [Google Scholar]
- Spina, F.; Anastasi, A.; Prigione, V.; Tigini, V.; Varese, G.C. Biological treatment of industrial wastewaters: A fungal approach. Chem. Eng. Trans. 2012, 27, 175–180. [Google Scholar]
- Crognale, S.; Federici, F.; Petruccioli, M. Enhanced separation of filamentous fungi by ultrasonic field: Possible usages in repeated batch processes. J. Biotechnol. 2002, 97, 191–197. [Google Scholar] [CrossRef]
- Chisti, Y.; Moo-Young, M. On Bioreactors. In Encyclopedia of Physical Science and Technology; Meyers, R.A., Ed.; Academic Press: San Diego, CA, USA, 2002; Volume 2, pp. 247–271. [Google Scholar]
- Zhong, J.J. Bioreactor Engineering. In Comprehensive Biotechnology; Elsevier B.V.: Amsterdam, The Netherlands, 2011; Volume 3, pp. 653–658. [Google Scholar]
- Marco-Urrea, E.; Perez-Trujillo, M.; Vicent, T.; Caminal, G. Ability of white-rot fungi to remove selected pharmaceuticals and identification of degradation products of ibuprofen by Trametes versicolor. Chemosphere 2009, 74, 765–772. [Google Scholar] [CrossRef]
- Jelic, A.; Cruz-Morato, C.; Marco-Urrea, E.; Sarra, M.; Perez, S.; Vicent, T.; Petrovic, M.; Barcelo, D. Degradation of carbamazepine by Trametes versicolor in an air pulsed fluidized bed bioreactor and identification of intermediates. Water Res. 2012, 46, 955–964. [Google Scholar] [CrossRef]
- Rodarte-Moralez, A.I.; Feijoo, G.; Moreira, M.T.; Lema, J.M. Degradation of selected pharmaceutical and personal care products (PPCPs) by white-rot fungi. World J. Microbial Biotechnol. 2011, 27, 1839–1846. [Google Scholar] [CrossRef]
- Zhang, Y.; Geißen, S.U. Elimination of carbamazepine in a nonsterile fungal bioreactor. Bioresour. Technol. 2012, 112, 221–227. [Google Scholar] [CrossRef]
- Cruz-Morató, C.; Ferrando-Climent, L.; Rodriguez-Mozaz, S.; Barceló, D.; Marco-Urrea, E.; Vicent, T.; Sarrà, M. Degradation of pharmaceuticals in non-sterile urban wastewater by Trametes versicolor in a fluidized bed bioreactor. Water Res. 2013, 47, 5200–5210. [Google Scholar] [CrossRef]
- Nguyen, L.N.; Hai, F.I.; Yang, S.; Kang, J.; Leusch, F.D.L.; Roddick, F.; Price, W.E.; Nghiem, L.D. Removal of trace organic contaminants by an MBR comprising a mixed culture of bacteria and white-rot fungi. Bioresour. Technol. 2013, 148, 234–241. [Google Scholar] [CrossRef]
- Cruz-Morató, C.; Lucas, D.; Llorca, M.; Rodriguez-Mozaz, S.; Gorga, M.; Petrovic, M.; Barceló, D.; Vicent, T.; Sarrà, M.; Marco-Urrea, E. Hospital wastewater treatment by fungal bioreactor: Removal efficiency for pharmaceuticals and endocrine disruptor compounds. Sci. Total Environ. 2014, 493, 365–376. [Google Scholar] [CrossRef] [PubMed]
- Gros, M.; Cruz-Morato, C.; Marco-Urrea, E.; Longrée, P.; Singer, H.; Sarrà, M.; Hollender, J.; Vicent, T.; Rodriguez-Mozaz, S.; Barceló, D. Biodegradation of the X-ray contrast agent iopromide and the fluoroquinolone antibiotic ofloxacin by the white rot fungus Trametes versicolor in hospital wastewaters and identification of degradation products. Water Res. 2014, 60, 228–241. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.; Hai, F.i.; Yang, S.; Kang, J.; Leusch, F.; Roddick, F.; Price, W.E.; Nghiem, L.D. Removal of pharmaceuticals, steroid hormones, phytoestrogens, UV-filters, industrial chemicals and pesticides by Trametes versicolor: Role of biosorption and biodegradation. Int. Biodeterior. Biodegration 2014, 88, 169–175. [Google Scholar] [CrossRef]
- Badia-Fabregat, M.; Lucas, D.; Pereira, M.A.; Alves, M.A.; Pennanen, T.; Fritze, H.; Rodríguez-Mozaz, S.; Barceló, D.; Vicent, T.; Caminal, G. Continuous fungal treatment of non-sterile veterinary hospital effluent: Pharmaceuticals removal and microbial community assessment. Appl. Microbiol. Biotechnol. 2016, 100, 2401–2415. [Google Scholar] [CrossRef] [PubMed]
- Bodin, H.; Daneshvar, A.; Gros, M.; Hultberg, M. Effects of Biopellets Composed of Microalgae and Fungi on Pharmaceuticals Present at Environmentally Relevant Levels in Water. Ecol. Eng. J. Ecotechnol. 2016, 91, 169–172. [Google Scholar] [CrossRef]
- Becker, D.; Rodriguez-Mozaz, S.; Insa, S.; Schoevaart, R.; Barceloó, D.; de Cazes, M.; Belleville, M.P.; Sanchez-Marcano, J.; Misovic, A.; Oehlmann, J.; et al. Removal of Endocrine Disrupting Chemicals in Wastewater by Enzymatic Treatment with Fungal Laccases. Org. Process Res. Dev. 2017, 21, 480–491. [Google Scholar] [CrossRef]
- Blánquez, P.; Guieysse, B. Continuous biodegradation of 17β-estradiol and 17α-ethynylestradiol by Trametes versicolor. J. Hazard. Mater. 2008, 150, 459–462. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Rodriguez, C.E.; García-Galán, M.J.; Blánquez, P.; Díaz-Cruz, M.S.; Barceló, D.; Caminal, G.; Vicent, T. Continuous degradation of a mixture of sulfonamides by Trametes versicolor and identification of metabolites from sulfapyridine and sulfathiazole. J. Hazard. Mater. 2012, 213–214, 347–354. [Google Scholar] [CrossRef]
- Rodriguez-Rodriguez, C.E.; Marco-Urrea, E.; Caminal, G. Degradation of Naproxen and Carbamazepine in Spiked Sludge Slurry and Solid-Phase Trametes versicolor Systems. Bioresour. Technol. 2010, 101, 2259–2266. [Google Scholar] [CrossRef]
- Marco-Urrea, E.; Perez-Trujillo, M.; Cruz-Morato, C.; Caminal, G.; Vicent, T. Degradation of the Drug Sodium Diclofenac Trametes versicolor Pellets and Identification of Some Intermediates by NMR. J. Hazard. Mater. 2010, 176, 836–842. [Google Scholar] [CrossRef]
- Marco-Urrea, E.; Perez-Trujillo, M.; Cruz-Morato, C.; Caminal, G.; Vicent, T. White-Rot Fungus-Mediated Degradation of the Analgesic Ketoprofen and Identification of Intermediates by HPLC-DAD-MS and NMR. Chemosphere 2010, 78, 474–481. [Google Scholar] [CrossRef] [PubMed]
- Becker, D.; Varela Della Giustina, S.; Rodriguez-Mozaz, S.; Schoevaart, R.; Barcelo, D.; de Cazes, M.; Belleville, M.P.; Sanchez-Marcano, J.; de Gunzburg, J.; Couillerot, O.; et al. Removal of antibiotics in wastewater by enzymatic treatment with fungal laccase—Degradation of compounds does not always eliminate toxicity. Bioresour. Technol. 2016, 219, 500–509. [Google Scholar] [CrossRef] [PubMed]
- Rodarte-Moralez, A.I.; Feijoo, G.; Moreira, M.T.; Lema, J.M. Operation of stirred tank reactors (STRs) and fixed-bed reactors (FBRs) with free and immobilized Phanerochaete chrysosporium for the continuous removal of pharmaceutical compounds. Biochem. Eng. J. 2012, 66, 38–45. [Google Scholar] [CrossRef]
- Wen, X.; Jia, Y.; Li, J. Enzymatic Degradation of Tetracycline and Oxytetracycline Crude Manganese Peroxidase Prepared Phanerochaete Chrysosporium. J. Hazard. Mater. 2010, 177, 924–928. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, R.A.; Sales, P.T.F.; Campos, L.C.; Garcia, L.C.; Valadares, T.A.; Schimidt, M.C.; Santiago, M.F. Evaluation of the use of Pycnoporus sanguineus fungus for phenolics and genotoxicity decay of a pharmaceutical effluent treatment. Ambi-Agua Taubaté 2012, 7, 41–50. [Google Scholar]
- Palli, L.; Castellet-Rovira, F.; Péerez-Trujillo, M.; Caniani, D.; Sarrá-Adroguer, M.; Gori, R. Preliminary Evaluation of Pleurotus ostreatus for the Removal of Selected Pharmaceuticals from Hospital Wastewater. Biotechnol. Prog. 2017, 33, 1529–1537. [Google Scholar] [CrossRef]
- Santos, I.J.S.; Grossman, M.; Sartoratto, A.; Ponezi, A.; Durranta, L. Degradation of the Recalcitrant Pharmaceuticals Carbamazepine and 17α-Ethinylestradiol by Ligninolytic fungi. Chem. Eng. Trans. 2012, 27, 169–174. [Google Scholar]
- Esterhuizen-Londt, M.; Schwartz, K.; Pflugmacher, S. Using aquatic fungi for pharmaceutical bioremediation: Uptake of acetaminophen by Mucor hiemalis does not result in an enzymatic oxidative stress response. Fungal Biol. 2016, 120, 1249–1257. [Google Scholar] [CrossRef]
- Buchicchio, A.; Bianco, G.; Sofo, A.; Masi, S.; Caniani, D. Biodegradation of carbamazepine and clarithromycin by Trichoderma harzianum and Pleurotus ostreatus investigated by liquid chromatography–high resolution tandem mass spectrometry (FTICR MS-IRMPD). Sci. Total Environ. 2016, 557, 733–739. [Google Scholar] [CrossRef]
- Golan-Rozen, N.; Chefetz, B.; Ben-Ari, J.; Geva, J.; Hadar, Y. Transformation of the recalcitrant pharmaceutical compound carbamazepine by Pleurotus ostreatus: Role of cytochrome P450 monooxygenase and manganese peroxidase. Environ. Sci. Technol. 2011, 45, 6800–6805. [Google Scholar] [CrossRef]
- Tran, N.H.; Urase, T.; Kusakabe, O. Biodegradation Characteristics of Pharmaceutical Substances by Whole Fungal Culture Trametes versicolor and its Laccase. J. Water Environ. Technol. 2010, 8, 125–140. [Google Scholar] [CrossRef]
- Pointing, S.B. Feasibility of bioremediation by white-rot fungi. Appl. Microbiol. Biotechnol. 2001, 57, 20–33. [Google Scholar] [PubMed]
- Wesenberg, D.; Kyriakides, I.; Agathos, S.N. White-rot fungi and their enzymes for the treatment of industrial dye effluents. Biotechnol. Adv. 2003, 22, 161–187. [Google Scholar] [CrossRef] [PubMed]
- Tortella, G.; Diez, M.; Durán, N. Fungal diversity and use in decomposition of environmental pollutants. Crit. Rev. Microbiol. 2005, 31, 197–212. [Google Scholar] [CrossRef] [PubMed]
- Buchanan, I.D.; Nicell, J.A. Kinetics of peroxidase interactions in the presence of a protective additive. J. Chem. Technol. Biotechnol. 1998, 72, 23–32. [Google Scholar] [CrossRef]
- Buchanan, I.D.; Han, Y.S. Assessment of the potential of Arthromyces ramosus peroxidase to remove phenol from industry wastewaters. Environ. Technol. 1999, 21, 545–552. [Google Scholar] [CrossRef]
- Gasser, C.A.; Hommes, G.; Schaffer, A.; Corvini, P.F. Multi-catalysis reactions: New prospects and challenges of biotechnology to valorize lignin. Appl. Microbiol. Biotechnol. 2012, 95, 1115–1134. [Google Scholar] [CrossRef]
- Kim, Y.J.; Nicell, J.A. Impact of reaction conditions on the laccase-catalyzed conversion of bisphenol A. Bioresour. Technol. 2006, 97, 1431–1442. [Google Scholar] [CrossRef]
- Sayadi, S.; Ellouz, R. Roles of lignin peroxidase and manganese peroxidase from Phanerochaete chrysosporium in the decolorization of olive mill wastewaters. Appl. Environ. Microbiol. 1995, 61, 1098–1103. [Google Scholar]
- Nguyen, L.N.; Hai, F.I.; Price, W.E.; Leusch, F.D.L.; Roddick, F.; McAdam, E.J.; Magram, A.F.; Long, D.; Nghiem, L.D. Continuous biotransformation of bisphenol A and diclofenac by laccase in an enzymatic membrane reactor. Int. Biodeterior. Biodegrad. 2014, 95, 25–32. [Google Scholar] [CrossRef]
- Nguyen, L.N.; Hai, F.I.; Price, W.E.; Leusch, F.D.; Roddick, F.; Ngo, H.H.; Guo, W.; Magram, S.F.; Nghiem, L.D. The effects of mediator and granular activated carbon addition on degradation of trace organic contaminants by an enzymatic membrane reactor. Bioresour. Technol. 2014, 167, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.N.; Hai, F.I.; Kang, J.; Leusch, F.D.L.; Roddick, F.; Magram, S.F.; Price, W.E.; Nghiem, L.D. Enhancement of trace organic contaminant degradation by crude enzyme extract from Trametes versicolor culture: Effect of mediator type and concentration. J. Taiwan Inst. Chem. Eng. 2014, 45, 1855–1862. [Google Scholar] [CrossRef]
- Nguyen, L.N.; Hai, F.I.; Dosseto, A.; Richardson, C.; Price, W.E.; Nghiem, L.D. Continuous adsorption and biotransformation of micropollutants by granular activated carbon-bound laccase in a packed-bed enzyme reactor. Bioresour. Technol. 2016, 210, 108–116. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.N.; van de Merwe, J.P.; Hai, F.I.; Leusch, F.D.L.; Kang, J.; Price, W.E.; Roddick, F.; Magram, S.F.; Nghiem, L.D. Laccase-syringaldehyde-mediated degradation of trace organic contaminants in an enzymatic membrane reactor: Removal efficiency and effluent toxicity. Bioresour. Technol. 2016, 200, 477–484. [Google Scholar] [CrossRef] [PubMed]
- Rahmani, K.; Faramarzi, M.A.; Mahvi, A.H.; Gholami, M.; Esrafili, A.; Forootanfar, H.; Farzadkia, M. Elimination and detoxification of sulfathiazole and sulfamethoxazole assisted by laccase immobilized on porous silica beads. Int. Biodeterior. Biodegrad. 2015, 97, 107–114. [Google Scholar] [CrossRef]
- Margot, J.; Bennati-Granier, C.; Maillard, J.; Blánquez, P.; Barry, D.A.; Holliger, C. Bacterial versus fungal laccase: Potential for micropollutant degradation. AMB Express 2013, 3, 63. [Google Scholar] [CrossRef] [PubMed]
- Ji, C.; Hou, J.; Wang, K.; Zhang, Y.; Chen, V. Biocatalytic degradation of carbamazepine with immobilized laccase-mediator membrane hybrid reactor. J. Membr. Sci. 2016, 502, 11–20. [Google Scholar] [CrossRef]
- Ji, C.; Hou, J.; Chen, V. Cross-linked carbon nanotubes-based biocatalytic membranes for micro-pollutants degradation: Performance, stability, and regeneration. J. Membr. Sci. 2016, 520, 869–880. [Google Scholar] [CrossRef]
- Xu, R.; Tang, R.; Zhou, Q.; Li, F.; Zhang, B. Enhancement of catalytic activity of immobilized laccase for diclofenac biodegradation by carbon nanotubes. Chem. Eng. J. 2015, 262, 88–95. [Google Scholar] [CrossRef]
- Touahar, I.E.; Haroune, L.; Ba, S.; Bellenger, J.P.; Cabana, H. Characterization of combined cross-linked enzyme aggregates from laccase, versatile peroxidase and glucose oxidase, and their utilization for the elimination of pharmaceuticals. Sci. Total Environ. 2014, 481, 90–99. [Google Scholar] [CrossRef]
- Nair, R.R.; Demarche, P.; Agathos, S.N. Formulation and characterization of an immobilized laccase biocatalyst and its application to eliminate organic micropollutants in wastewater. New Biotechnol. 2013, 30, 814–823. [Google Scholar] [CrossRef] [PubMed]
- Ding, H.; Wu, Y.; Zou, B.; Lou, Q.; Zhang, W.; Zhong, J.; Lu, L.; Dai, G. Simultaneous removal and degradation characteristics of sulfonamide, tetracycline, and quinolone antibiotics by laccase-mediated oxidation coupled with soil adsorption. J. Hazard. Mater. 2016, 307, 350–358. [Google Scholar] [CrossRef] [PubMed]
- De Cazes, M.; Belleville, M.P.; Mougel, M.; Kellner, H.; Sanchez-Marcano, J. Characterization of laccase-grafted ceramic membranes for pharmaceuticals degradation. J. Membr. Sci. 2015, 476, 384–393. [Google Scholar] [CrossRef]
- De Cazes, M.; Belleville, M.P.; Petit, E.; Llorca, M.; Rodríguez-Mozaz, S.; de Gunzburg, J.; Barceló, D.; Sanchez-Marcano, J. Design and optimization of an enzymatic membrane reactor for tetracycline degradation. Catal. Today 2014, 236, 146–152. [Google Scholar] [CrossRef]
- Llorca, M.; Rodríguez-Mozaz, S.; Couillerot, O.; Panigoni, K.; de Gunzburg, J.; Bayer, S.; Czaja, R.; Barceló, D. Identification of new transformation products during enzymatic treatment of tetracycline and erythromycin antibiotics at laboratory scale by an on-line turbulent flow liquid-chromatography coupled to a high resolution mass spectrometer LTQ-Orbitrap. Chemosphere 2015, 119, 90–98. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Sidhu, S.S.; Zhang, H.; Huang, Q. Removal of sulfadimethoxine in soil mediated by extracellular oxidoreductases. Environ. Sci. Pollut. Res. 2015, 22, 16868–16874. [Google Scholar] [CrossRef] [PubMed]
- Singh, D.; Rawat, S.; Waseem, M.; Gupta, S.; Lynn, A.; Nitin, M.; Ramchiary, N.; Sharm, K.K. Molecular modeling and simulation studies of recombinant laccase from Yersinia enterocolitica suggests significant role in the biotransformation of non-steroidal anti-inflammatory drugs. Biochem. Biophys. Res. Commun. 2016, 469, 306–312. [Google Scholar] [CrossRef]
- Li, X.; Xu, Q.M.; Cheng, J.S.; Yuan, Y.J. Improving the bioremoval of sulfamethoxazole and alleviating cytotoxicity of its biotransformation by laccase producing system under coculture of Pycnoporus sanguineus and Alcaligenes Faecalis. Bioresour. Technol. 2016, 220, 333–340. [Google Scholar] [CrossRef]
- Chen, Y.; Stemple, B.; Kumar, M.; Wei, N. Cell surface display fungal laccase as a renewable biocatalyst for degradation of persistent micropollutants bisphenol A and sulfamethoxazole. Environ. Sci. Technol. 2016, 50, 8799–8808. [Google Scholar] [CrossRef]
- Hofmann, U.; Schlosser, D. Biochemical and physicochemical processes contributing to the removal of endocrine-disrupting chemicals and pharmaceuticals by the aquatic ascomycete Phoma sp. UHH 5-1-03. Appl. Microbiol. Biotechnol 2016, 100, 2381–2399. [Google Scholar] [CrossRef]
- Shi, L.; Ma, F.; Han, Y.; Zhang, X.; Yu, H. Removal of sulfonamide antibiotics by oriented immobilized laccase on Fe3O4 nanoparticles with natural mediators. J. Hazard. Mater. 2014, 279, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Margot, J.; Copin, P.J.; von Gunten, U.; Barry, D.A.; Holliger, C. Sulfamethoxazole and isoproturon degradation and detoxification by a laccase-mediator system: Influence of treatment conditions and mechanistic aspects. Biochem. Eng. J. 2015, 103, 47–59. [Google Scholar] [CrossRef]
- Martinez, D.; Larrondo, L.F.; Putnam, N.; Gelpke, M.D.S.; Huang, K.; Chapman, J.; Helfenbein, K.G.; Ramaiya, P.; Detter, J.C.; Larimer, F.; et al. Genome sequence of lignocellulose degrading fungus Phanerochaete chrysosporium strain RP78. Nat. Biotechnol. 2004, 22, 695–700. [Google Scholar] [CrossRef] [PubMed]
- Fabbrini, M.; Galli, C.; Gentili, P. Comparing the catalytic efficiency of some mediators of laccase. J. Mol. Catal. B Enzym. 2002, 16, 231–240. [Google Scholar] [CrossRef]
- Cañas, A.I.; Camarero, S. Laccases and their natural mediators: Biotechnological tools for sustainable eco-friendly processes. Biotechnol Adv. 2010, 28, 694–705. [Google Scholar] [CrossRef]
- Hu, M.R.; Chao, Y.P.; Zhang, G.Q.; Xue, Z.Q.; Qian, S. Laccase mediator system in the decolorization of different types of recalcitrant dyes. J. Ind. Microbiol. Biotechnol. 2009, 36, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Wong, D.W. Structure and action mechanism of ligninolytic enzymes. Appl. Biochem. Biotechnol. 2009, 157, 174–209. [Google Scholar] [CrossRef]
- Morozova, O.V.; Shumakovich, G.P.; Shleev, S.V.; Yaropolov, Y.I. Laccase-mediator systems and their applications: A review. Appl. Biochem. Microbiol. 2007, 43, 523–535. [Google Scholar] [CrossRef]
- Pogni, R.; Baratto, M.C.; Sinicropi, A.; Basosi, R. Spectroscopic and computational characterization of laccases and their substrate radical intermediates. Cell Mol. Life Sci. 2015, 72, 885–896. [Google Scholar] [CrossRef]
- Weng, S.S.; Liu, S.M.; Lai, H.T. Application parameters of laccase mediator systems for treatment of sulfonamide antibiotics. Bioresour. Technol. 2013, 141, 152–159. [Google Scholar] [CrossRef]
- Kurniawati, S.; Nicell, J.A. Efficacy of mediators for enhancing the laccase-catalyzed oxidation of aqueous phenol. Enzyme Microb. Technol. 2007, 41, 353–361. [Google Scholar] [CrossRef]
- Fillat, U.; Prieto, A.; Camarero, S.; Martínez, Á.T.; Martínez, M.J. Biodeinking of flexographic inks by fungal laccases using synthetic and natural mediators. Biochem. Eng. J. 2012, 67, 97–103. [Google Scholar] [CrossRef]
- Ashe, B.; Nguyen, L.N.; Hai, F.I.; Lee, D.J.; van de Merwe, J.P.; Leusch, F.D.L.; Price, W.E.; Nghiem, L.D. Impacts of redox-mediator type on trace organic contaminants degradation by laccase: Degradation efficiency, laccase stability and effluent toxicity. Int. Biodeterior. Biodegrad. 2016, 113, 169–176. [Google Scholar] [CrossRef]
- Verma, P.; Madamwar, D. Production of lignolytic enzymes for dye decolorization by co-cultivation of white rot fungi Pleurotus ostreatus and Phanerochaete chrysosporium under solid state fermentation. Appl. Biochem. Biotechnol 2002, 102–103, 109–118. [Google Scholar] [CrossRef]
- Zhang, Y.; Geißen, S.U. In Vitro Degradation of Carbamazepine and Diclofenac Crude Lignin Peroxidase. J. Hazard. Mater. 2010, 176, 1089–1092. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, J.F.; Cabral, J.M.S. Enzyme Immobilization. In Biotechnology; Rehm, H.J., Reed, G., Eds.; VCH: Weinheim, Germany, 1987; pp. 347–404. [Google Scholar]
- Asif, M.B.; Hai, F.I.; Singh, L.; Price, W.E.; Nghiem, L.D. Degradation of Pharmaceuticals and Personal Care Products by White-Rot Fungi-a Critical Review. Curr. Pollut. Rep. 2017, 3, 88–103. [Google Scholar] [CrossRef]
- Yang, S.; Hai, F.I.; Nghiem, L.D.; Nguyen, L.N.; Roddick, F.; Price, W.E. Removal of bisphenol A and diclofenac by a novel fungal membrane bioreactor operated under non-sterile conditions. Int. Biodeterior. Biodegrad. 2013, 85, 483–490. [Google Scholar] [CrossRef]
- Suda, T.; Hata, T.; Kawai, S.; Okamura, H.; Nishida, T. Treatment of tetracycline antibiotics by laccase in the presence of 1-hydroxybenzotriazole. Bioresour. Technol. 2012, 103, 498–501. [Google Scholar] [CrossRef]
- Joutey, N.T.; Bahafid, W.; Sayel, H.; Ghachtouli, N.E. Biodegradation: Involved microorganisms and genetically engineered microorganisms. In Biodegradation—Life of Science; Chamy, R., Rosenkranz, F., Eds.; InTechOpen: London, UK, 2013; p. 305. [Google Scholar]
- Dujon, B. The yeast genome project: What did we learn? Trends Genet 1996, 12, 263–270. [Google Scholar] [CrossRef]
- Wood, V.; Gwilliam, R.; Rajandream, M.A.; Lyne, M.; Lyne, R.; Stewart, A.; Sgouros, J.; Peat, N.; Hayles, J.; Baker, S.; et al. The genome sequence of Schizosaccharomyces pombe. Nature 2002, 415, 871–880. [Google Scholar] [CrossRef]
- Martinez, M.; Bernal, P.; Almela, C.; Velez, D.; Garcia-Agustin, P.; Serrano, R. An engineered plant that accumulates higher levels of heavy metals than Thlaspi caerulescens, with yields of 100 times more biomass in mine soils. Chemosphere 2006, 64, 478–485. [Google Scholar] [CrossRef] [PubMed]
- Nam, J.M.; Fujita, Y.; Arai, T.; Kondo, A.; Morikawa, Y.; Okada, H.; Ueda, M.; Tanaka, A. Construction of engineered yeast with the ability of binding to cellulose. J. Mol. Catal. B Enzym. 2002, 17, 197–202. [Google Scholar] [CrossRef]
- Balcázar-López, E.; Méndez-Lorenzo, L.H.; Batista-García, R.A.; Esquivel-Naranjo, U.; Ayala, M.; Kumar, V.V.; Savary, O.; Cabana, H.; Herrera-Estrella, A.; Folch-Mallol, J.L. Xenobiotic compounds degradation by heterologous expression of a Trametes sanguineus laccase in Trichoderma atroviride. PLoS ONE 2016, 11, e0147997. [Google Scholar] [CrossRef] [PubMed]
- Sze, P. A Biology of the Algae, 3rd ed.; WCB/McGraw-Hill: Boston, MA, USA, 1998. [Google Scholar]
- Ruggiero, M.A.; Gordon, D.P.; Orrell, T.M.; Bailly, N.; Bourgoin, T.; Brusca, R.C.; Cavalier-Smith, T.; Guiry, M.D.; Kirk, P.M. Correction: A Higher Level Classification of All Living Organisms. PLoS ONE 2015, 10, e0130114. [Google Scholar] [CrossRef] [PubMed]
- Subashchandrabose, S.R.; Ramakrishnam, B.; Megharaj, M.; Venkateswarlu, K.; Naidu, R.R. Mixotrophic Cyanobacteria and microalgae as distinctive biological agents for organic pollutant degradation. Environ. Int. 2013, 51, 59–72. [Google Scholar] [CrossRef] [PubMed]
- Fu, W.; Chaiboonchoe, A.; Khraiwesh, B.; Nelson, D.R.; Al-Khairy, D.; Mystikou, A.; Alzahmi, A.; Salehi-Ashtiani, K. Algal Cell Factories: Approaches, Applications, and Potentials. Mar. Drugs 2016, 14, 225. [Google Scholar] [CrossRef]
- Osundeko, O.; Dean, A.P.; Davies, H.; Pittman, J.K. Acclimation of microalgae to wastewater environments involves increased oxidative stress tolerance activity. Plant Cell Physiol. 2014, 55, 1848–1857. [Google Scholar] [CrossRef]
- Cho, K.; Lee, C.H.; Ko, K. Use of phenol-induced oxidative stress acclimation to stimulate cell growth and biodiesel production by the oceanic microalga Dunaliella salina. Algal Res. 2016, 17, 61–66. [Google Scholar] [CrossRef]
- Xiong, J.Q.; Kurade, M.B.; Jeon, B.H. Biodegradation of levofloxacin by an acclimated freshwater alga Chlorella vulgaris. Chem. Eng. J. 2017, 313, 1251–1257. [Google Scholar] [CrossRef]
- Rehnstam-Holm, A.S.; Godhe, A. Genetic Engineering of Algal Species. In Biotechnology; Doelle, H.W., Ed.; Encyclopedia of Life Support Systems (EOLSS) Publishers: Oxford, UK, 2003. [Google Scholar]
- Muñoz, R.; Guieysse, B. Algal–bacterial processes for the treatment of hazardous contaminants: A review. Water Res. 2006, 40, 2799–2815. [Google Scholar] [CrossRef]
- Office of Water, Wastewater Management Fact Sheet, Energy Conservation; EPA 832-F-06-024; U.S. Environmental Protection Agency: Washington, DC, USA, 2006; p. 7.
- Oswald, W.J.; Golueke, C.G. Biological transformation of solar energy. Adv. Appl. Microbiol. 1960, 2, 223–262. [Google Scholar] [CrossRef] [PubMed]
- Polprasert, C.; Dissanayake, M.; Thanh, N. Bacterial Die-Off Kinetics in Waste Stabilization Ponds. Water Pollut. Control Fed. 1983, 55, 285–296. [Google Scholar] [CrossRef]
- Pienkos, P.; Darzins, A. The promise and challenges of microalgal-derived biofuels. Biofuels Bioprod. Biorefin. 2009, 431–440. [Google Scholar] [CrossRef]
- Becker, E.W. Micro-algae for human and animal consumption. In Micro-Algal Biotechnology; Borowitzka, M.A., Borowitzka, L.J., Eds.; Cambridge University Press: Cambridge, UK, 1988; pp. 222–256. [Google Scholar]
- Ugwu, C.U.; Aoyagi, H.; Uchiyama, H. Photobioreactors for mass cultivation of algae. Bioresour. Technol. 2008, 99, 4021–4028. [Google Scholar] [CrossRef] [PubMed]
- Chisti, Y. Biodiesel from microalgae. Biotechnol. Adv. 2007, 25, 294–306. [Google Scholar] [CrossRef] [PubMed]
- Garcia, J.; Green, B.F.; Lundquist, T.; Mujeriego, R.; Hernández-Mariné, M.; Oswald, W.J. Long term diurnal variations in contaminant removal in high rate ponds treating urban wastewater. Bioresour. Technol. 2006, 97, 1709–1715. [Google Scholar] [CrossRef] [PubMed]
- Harun, R.; Singh, M.; Forde, G.M.; Danquah, M.K. Bioprocess engineering of microalgae to produce a variety of consumer products. Renew. Sustain. Energy Rev. 2010, 14, 1037–1047. [Google Scholar] [CrossRef]
- Picot, B.; Moersidik, S.; Casellas, C.; Bontoux, J. Using diurnal variations in a high rate algal pond for management pattern. Water Sci. Technol. 1993, 28, 169–175. [Google Scholar] [CrossRef]
- Singh, S.P.; Singh, P. Effect of temperature and light on the growth of algae species: A review. Renew. Sustain. Energy Rev. 2015, 50, 431–444. [Google Scholar] [CrossRef]
- Chen, C.Y.; Yeh, K.L.; Aisyah, R.; Lee, D.J.; Chang, J.S. Cultivation, photobioreactor design and harvesting of microalgae for biodiesel production: A critical review. Bioresour. Technol. 2011, 102, 71–81. [Google Scholar] [CrossRef]
- Eriksen, N.T. The technology of microalgal culturing. Biotechnol. Lett. 2008, 30, 1525–1536. [Google Scholar] [CrossRef] [PubMed]
- Wiley, P.E.; Brenneman, K.J.; Jacobson, A.E. Improved Algal Harvesting Using Suspended Air Flotation. Water Environ. Res. 2009, 81, 702–708. [Google Scholar] [CrossRef] [PubMed]
- Akhtar, N.; Iqbal, J.; Iqbal, M. Enhancement of lead (II) biosorption by microalgal biomass immobilized onto loofa (Luffa cylindrica) sponge. Eng. Life Sci. 2004, 4, 171–178. [Google Scholar] [CrossRef]
- Mallick, N. Biotechnological potential of immobilized algae for wastewater N, P and metal removal: A review. Biometals 2002, 15, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Amaro, H.M.; Guedes, A.C.; Malcata, F.X. Advances and perspectives in using microalgae to produce biodiesel. Appl. Energy 2011, 88, 3402–3410. [Google Scholar] [CrossRef]
- Schenk, P.M.; Thomas-Hall, S.R.; Stephens, E.; Marx, U.C.; Mussgnug, J.H.; Posten, C.; Kruse, O.; Hankamer, B. Second generation biofuels: High-efficiency microalgae for biodiesel production. Bioenergy Res. 2008, 1, 20–43. [Google Scholar] [CrossRef]
- Taylor, R.L.; Rand, J.D.; Caldwell, G.S. Treatment with algae extracts promotes flocculation, and enhances growth and neutral lipid content in Nannochloropsis oculata—A candidate for biofuel production. Mar. Biotechnol. 2012, 6, 774–781. [Google Scholar] [CrossRef]
- Van Den Hende, S.; Vervaeren, H.; Saveyn, H.; Maes, G.; Boon, N. Microalgal bacterial floc properties are improved by a balanced inorganic/organic carbon ratio. Biotechnol. Bioeng. 2011, 108, 549–558. [Google Scholar] [CrossRef]
- Su, Y.; Mennerich, A.; Urban, B. Municipal wastewater treatment and biomass accumulation with a wastewater-born and settleable algal-bacterial culture. Water Res. 2011, 45, 3351–3358. [Google Scholar] [CrossRef]
- Zhou, W.; Cheng, Y.; Li, Y.; Wan, Y.; Liu, Y.; Lin, X.; Ruan, R. Novel fungal palletization-assisted technology for algae harvesting and wastewater treatment. Appl. Biochem. Biotechnol. 2012, 167, 214–228. [Google Scholar] [CrossRef]
- Zhang, J.G.; Hu, B.A. novel method to harvest microalgae via co-culture of filamentous fungi to form cell pellets. Bioresour. Technol. 2012, 114, 529–535. [Google Scholar] [CrossRef] [PubMed]
- Gultom, S.O.; Hu, B. Review of microalgae harvesting via co-pelletization with filamentous fungus. Energies 2013, 6, 5921–5939. [Google Scholar] [CrossRef]
- Wittenberg, K.M. Preservation of high-moisture hay in storage through the use of forage additives. Can. J. Anim. Sci. 1991, 71, 429–437. [Google Scholar] [CrossRef]
- Milledge, J.J.; Heaven, S.A. review of the harvesting of micro-algae for biofuel production. Rev. Environ. Sci. Biotechnol. 2013, 12, 165–178. [Google Scholar] [CrossRef]
- Olguin, E.J. Dual purpose microalgae-bacteria-based systems that treat wastewater and produce biodiesel and chemical products within a biorefinery. Biotechnol. Adv. 2012, 30, 1031–1046. [Google Scholar] [CrossRef] [PubMed]
- Matamoros, V.; Uggetti, E.; García, J.; Bayona, J.M. Assessment of the mechanisms involved in the removal of emerging contaminants by microalgae from wastewater: A laboratory scale study. J. Hazard. Mater. 2015, 301, 197–205. [Google Scholar] [CrossRef] [PubMed]
- De Godos, I.; Muñoz, R.; Guieysse, B. Tetracycline removal during wastewater treatment in high-rate algal ponds. J. Hazard. Mater 2012, 229–230, 446–449. [Google Scholar] [CrossRef]
- Harms, H.; Schlosser, D.; Wick, L.Y. Untapped potential: Exploiting fungi in bioremediation of hazardous chemicals. Nat. Rev. Microbiol. 2011, 9, 177–192. [Google Scholar] [CrossRef]
- Joss, A.; Zabczynski, S.; Gobel, A.; Hoffmann, B.; Loffler, D.; McArdell, C.S.; Ternes, T.A.; Thomsen, A.; Siegrist, H. Biological degradation of pharmaceuticals in municipal wastewater treatment: Proposing a classification scheme. Water Res. 2006, 40, 1686–1696. [Google Scholar] [CrossRef]
- Kumar, M.S.; Kabra, A.N.; Min, B.; El-Dalatony, M.M.; Xiong, J.; Thajuddin, N.; Lee, D.S.; Jeon, B.H. Insecticides induced biochemical changes in freshwater microalga Chlamydomonas mexicana. Environ. Sci. Pollut. Res. 2016, 23, 1091–1099. [Google Scholar] [CrossRef]
- Peng, F.Q.; Ying, G.G.; Yang, B.; Liu, S.; Lai, H.J.; Liu, Y.S.; Chen, Z.F.; Zhou, G.J. Biotransformation of progesterone and norgestrel by two freshwater microalgae (Scenedesmus obliquus and Chlorella pyrenoidosa): Transformation kinetics and products identification. Chemosphere 2014, 95, 581–588. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.Q.; Kurade, M.B.; Abou-Shanab, R.A.I.; Ji, M.K.; Choi, J.J.O.; Jeon, B.H. Biodegradation of carbamazepine using freshwater microalgae Chlamydomonas mexicana and Scenedesmus obliquus and the determination of its metabolic fate. Bioresour. Technol. 2016, 205, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Maes, H.M.; Maletz, S.X.; Ratte, H.T.; Hollender, J.; Schaeffer, A. Uptake, elimination, and biotransformation of 17α-ethinylestradiol by the freshwater alga Desmodesmus subspicatus. Environ. Sci. Technol. 2014, 48, 12354–12361. [Google Scholar] [CrossRef] [PubMed]
- Ding, T.; Yang, M.; Zhang, J.; Yang, B.; Lin, K.; Li, J.; Gan, J. Toxicity, degradation and metabolic fate of ibuprofen on freshwater diatom Navicula sp. J. Hazard. Mater. 2017, 330, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Thies, F.T.; Backhaus, T.; Bossmann, B.; Grimme, L.H. Xenobiotic biotransformation in unicellular green algae. Plant Physiol. 1996, 112, 361–370. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.Q.; Kurade, M.B.; Patil, D.V.; Jang, M.; Paeng, K.J.; Jeon, B.H. Biodegradation and metabolic fate of levofloxacin via a freshwater green alga, Scenedesmus obliquus in synthetic saline wastewater. Algal Res. 2017, 25, 54–61. [Google Scholar] [CrossRef]
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 632–633. [Google Scholar] [CrossRef] [PubMed]
- Matamoros, V.; Gutiérrex, R.; Ferrer, I.; García, J.; Bayona, J.M. Capability of microalgae-based wastewater treatment systems to remove emerging organic contaminants: A pilot-scale study. J. Hazard. Matter. 2015, 288, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Garduño, B.; Pintado-Herrera, M.G.; Biel-Maeso, M.; Rueda-Márquez, J.J.; Lara-Martín, P.A.; Perales, J.A.; Manzano, M.A.; Garrido-Pérez, C.; Martín-Díaz, M.L. Environmental Risk Assessment of Effluents as a Whole Emerging Contaminant—Efficiency of Alternative Tertiary Treatments for Wastewater Depuration. Water Res. 2017, 119, 136–149. [Google Scholar] [CrossRef] [PubMed]
- Hom-Diaz, A.; Jaén-Gil, A.; Bello-Laserna, I.; Rodríguez-Mozaz, S.; Vicent, T.; Barceló, D.; Blánquez, P. Performance of a Microalgal Photobioreactor Treating Toilet Wastewater: Pharmaceutically Active Compound Removal and Biomass Harvesting. Sci. Total Environ. 2017, 592, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Gentili, F.G.; Fick, J. Algal cultivation in urban wastewater: An efficient way to reduce pharmaceutical pollutants. J. Appl. Phycol. 2017, 29, 255–262. [Google Scholar] [CrossRef] [PubMed]
- Lai, K.M.; Scrimshaw, M.D.; Lester, J.N. Biotransformation and Bioconcentration of Steroid Estrogens by Chlorella Vulgaris. Appl. Environ. Microbiol. 2002, 68, 859–864. [Google Scholar] [CrossRef]
- Shi, W.; Wang, L.; Rousseau, D.P.L.; Lens, P.N.L. Removal of Estrone, 17α-Ethinylestradiol, and 17β-Estradiol in Algae and Duckweed-Based Wastewater Treatment Systems. Environ. Sci. Pollut. Res. 2010, 17, 824–833. [Google Scholar] [CrossRef] [PubMed]
- Mani, D.; Kumar, C. Biotechnological advances in bioremediation of heavy metals contaminated ecosystems: An overview with special reference to phytoremediation. Int. J. Environ. Sci. Technol. 2014, 11, 843–872. [Google Scholar] [CrossRef]
- Wang, S.; Wang, X.; Poon, K.; Wang, Y.; Li, S.; Liu, H.; Lin, S.; Cai, Z. Removal and Reductive Dechlorination of Triclosan by Chlorella Pyrenoidosa. Chemosphere 2013, 92, 1498–1505. [Google Scholar] [CrossRef]
- Chen, J.; Zheng, F.; Guo, R. Algal feedback and removal efficiency in a sequencing batch reactor algae process (SBAR) to treat the antibiotic cefradine. PLoS ONE 2015, 10, e0133273. [Google Scholar] [CrossRef] [PubMed]
- Greca, M.D.; Pinto, G.; Pistillo, P.; Pollio, A.; Previtera, L.; Temussi, F. Biotransformation of ethinylestradiol by microalgae. Chemosphere 2008, 70, 2047–2053. [Google Scholar] [CrossRef]
- Zhou, G.J.; Ying, G.G.; Liu, S.; Zhou, L.J.; Chen, Z.F.; Peng, F.Q. Simultaneous removal of inorganic and organic compounds in wastewater by freshwater green microalgae. Environ. Sci. 2014, 16, 2018–2027. [Google Scholar] [CrossRef]
- Escapa, C.; Coimbra, R.N.; Paniagua, S.; García, A.I.; Otero, M. Nutrients and pharmaceuticals removal from wastewater by culture and harvesting of Chlorella sorokiniana. Bioresour. Technol. 2015, 185, 276–284. [Google Scholar] [CrossRef]
- Santos, C.E.; de Coimbra, R.N.; Bermejo, S.P.; Pérez, A.I.G.; Cabero, M.O. Comparative Assessment of Pharmaceutical Removal from Wastewater by the Microalgae Chlorella sorokiniana, Chlorella vulgaris and Scenedesmus obliquus. In Biological Wastewater Treatment and Resource Recovery; Farooq, R., Ahmad, Z., Eds.; IntechOpen: London, UK, 2017. [Google Scholar] [CrossRef]
- Tam, N.F.Y.; Chong, A.M.Y.; Wong, Y.S. Removal of Tributyltin (TBT) by Live and Dead Microalgal Cells. Mar. Pollut. Bull. 2002, 45, 362–371. [Google Scholar] [CrossRef]
- Bai, X.; Acharya, K. Removal of Trimethoprim, Sulfamethoxazole, and Triclosan by the Green Alga Nannochloris Sp. J. Hazard. Mater. 2016, 315, 70–75. [Google Scholar] [CrossRef] [PubMed]
- De Wilt, A.; Butkovskyi, A.; Tuantet, K.; Leal, L.H.; Fernandes, T.V.; Langenhoff, A.; Zeeman, G. Micropollutant Removal in an Algal Treatment System Fed with Source Separated Wastewater Streams. J. Hazard. Mater. 2016, 304, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Dzomba, P.; Kugara, J.; Mukunyaidze, V.V.; Zaranyika, M.F. Biodegradation of tetracycline antibacterial using green algal species collected from municipal and hospital effluents. Der. Chem. Sin. 2015, 6, 27–33. [Google Scholar]
- Escapa, C.; Coimbra, R.N.; Paniagua, S.; García, A.I.; Otero, M. Removal of pharmaceuticals from industrial wastewaters by microalgae culture. In Proceedings of the International Conference on Industrial Waste and Wastewater Treatment and Valorisation, Athens, Greece, 21–23 May 2015. [Google Scholar]
- Xiong, J.Q.; Kurade, M.B.; Kim, J.R.; Roh, H.S.; Jeon, B.H. Ciprofloxacin toxicity and its co-metabolic removal in a freshwater microalgae Chlamydomonas mexicana. J. Hazard. Mater. 2017, 323, 212–219. [Google Scholar] [CrossRef] [PubMed]
- Hom-Diaz, A.; Llorca, M.; Rodríguez-Mozaz, S.; Vicent, T.; Barceló, D.; Blánquez, P. Microalgae cultivation on wastewater digestate: β-estradiol and 17α-ethynylestradiol degradation and transformation products identification. J. Environ. Manag. 2015, 155, 106–113. [Google Scholar] [CrossRef]
- Toscano, M.D.; De Maria, L.; Lobedanz, S.; Ostergaard, L.H. Optimization of a small laccase by active-site redesign. ChemBioChem 2013, 14, 1209–1211. [Google Scholar] [CrossRef]
- Gadd, G.M. Biosorption: Critical review of scientific rationale, environmental importance and significance for pollution treatment. J. Chem. Technol. Biotechnol. 2009, 84, 13–28. [Google Scholar] [CrossRef]
- Michalak, I.; Chojnacka, K.; Witek-Krowiak, A. State of the art for the biosorption process—A review. Appl. Biochem. Biotechnol. 2013, 170, 1389–1416. [Google Scholar] [CrossRef]
- Babel, S.; Kurniawan, T.A. Low-cost adsorbents for heavy metals uptake from contaminated water: A review. J. Hazard. Mater. 2003, 97, 219–243. [Google Scholar] [CrossRef]
- Dutta, M.; Duttam, N.N.; Bhattacharya, K.G. Aqueous phase adsorption of certain beta-lactam antibiotics onto polymeric resins and activated carbon. Sep. Purif. Technol. 1999, 16, 213–224. [Google Scholar] [CrossRef]
- Aitcheson, S.J.; Arnett, J.; Murray, K.R.; Zhang, J. Removal of aquaculture therapeutants by carbon adsorption. 1. Equilibrium adsorption behaviour of single components. Aquaculture 2000, 183, 269–284. [Google Scholar] [CrossRef]
- Swedish Environmental Protection Agency, Advanced Wastewater Treatment for Separation and Removal of Pharmaceutical Residues and Other Hazardous Substances-Needs, Technologies and Impacts; REPORT 6803; Naturvårdsverket: Stockholm, Sweden, 2017.
- Bailey, S.E.; Olin, T.J.; Bricka, M.; Adrian, D.D. A review of potentially low-cost adsorbents for heavy metals. Water Res. 1999, 33, 2469–2479. [Google Scholar] [CrossRef]
- Tsezos, M.; Bell, J.P. Comparison of the biosorption and desorption of hazardous organic pollutants by live and dead biomass. Water Res. 1989, 23, 561–568. [Google Scholar] [CrossRef]
- Stumm, W.; Morgan, J.J. Aquatic Chemistry: Chemical Equilibria and Rates in Natural Waters, 3rd ed.; John Wiley & Sons: New York, NY, USA, 1996. [Google Scholar]
- Gow, N.A.R.; Gadd, G.M. The Growing Fungus; Chapman and Hall: London, UK, 1995. [Google Scholar]
- Gadd, G.M. Interactions of fungi with toxic metals. New. Phytol. 1993, 124, 25–60. [Google Scholar] [CrossRef]
- Aksu, Z. Application of biosorption of the removal of organic pollutants: A review. Process Biochem. 2005, 40, 997–1026. [Google Scholar] [CrossRef]
- Gadd, G.M.; Mowll, J.L. Copper uptake by yeast-like cells, hyphae and chlamydospores of Aureobasidium pullulans. Exp. Mycol. 1985, 9, 230–240. [Google Scholar] [CrossRef]
- Crist, R.H.; Oberholser, K.; Shank, N.; Nguyen, M. Nature of bonding between metallic ions and algal cell walls. Environ. Sci. Technol. 1981, 15, 1212–1217. [Google Scholar] [CrossRef]
- Bhatti, H.N.; Hamid, S. Removal of uranium (VI) from aqueous solutions using Eucalyptus citriodora distillation sludge. Int. J. Environ. Sci. Technol. 2014, 11, 813–822. [Google Scholar] [CrossRef]
- Navarro, A.E.; Lim, H.; Chang, E.; Lee, Y.; Manrique, A.S. Uptake of Sulfa Drugs from Aqueous Solutions by Marine Algae. Sep. Sci. Technol. 2014, 49, 2175–2181. [Google Scholar] [CrossRef]
- Chu, K.H. Improved fixed bed models for metal biosorption. Chem. Eng. J. 2004, 97, 233–239. [Google Scholar] [CrossRef]
- Aksu, Z.; Gonen, F. Biosorption of phenol by immobilized activated sludge in a continuous packed bed: Prediction of breakthrough curves. Process Biochem. 2004, 39, 599–613. [Google Scholar] [CrossRef]
- Vijayaraghvan, K.; Yun, Y.S. Bacterial biosorbents and biosorption. Biotechnol. Adv. 2008, 26, 266–291. [Google Scholar] [CrossRef] [PubMed]
- Volesky, B. Sorption and Biosorption; BV Sorbex Inc.: Montreal-St. Lambert, QC, Canada, 2004. [Google Scholar]
- Adamson, A.W.; Gast, A.P. Physical Chemistry of Surfaces, 6th ed.; Wiley: New York, NY, USA, 1997. [Google Scholar]
- Kyzas, G.Z.; Deliyanni, E.A.; Matis, K.A.; Lazaridis, N.K.; Bikiaris, D.N.; Mitropoulos, A.C. Emerging nanocomposite biomaterials as biomedical adsorbents: An overview. Compos. Interfaces 2018, 25, 415–454. [Google Scholar] [CrossRef]
- Park, D.; Yun, Y.S.; Park, J.M. The past, present, and future trends of biosorption. Biotechnol. Bioproc. 2010, E15, 86–102. [Google Scholar] [CrossRef]
- Ikehata, K.; Naghashkar, N.J.; El-Din, M.G. Degradation of aqueous pharmaceuticals by ozonation and advanced oxidation processes: A review. Ozone Sci. Eng. 2006, 28, 353–414. [Google Scholar] [CrossRef]
- Aksu, Z.; Tunc, O. Application of biosorption for penicillin G removal: Comparison with activated carbon. Process Biochem. 2005, 40, 831–847. [Google Scholar] [CrossRef]
- Kapoor, A.; Viraraghavan, T. Fungal biosorption—An alternative treatment option for heavy metal bearing wastewater: A review. Bioresour. Technol. 1995, 53, 195–206. [Google Scholar]
- Eccles, H. Treatment of metal-contaminated wastes: Why select a biological process? Trends Biotechnol. 1999, 17, 462–465. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, Y.J. Biosorption isotherms, kinetics and thermodynamics. Sep. Purif. Technol. 2008, 61, 229–242. [Google Scholar] [CrossRef]
- De Rome, L.; Gadd, G.M. Use of pelleted and immobilized yeast and fungal biomass for heavy metal and radionuclide recovery. J. Ind. Microbiol. 1991, 7, 97–104. [Google Scholar] [CrossRef]
- El-Din, S.M.M.; Noaman, N.H.; Zaki, S.H. Removal of Some Pharmaceuticals and Endocrine Disrupting Compounds by The Marine Macroalgae. Pterocladia capillacea and Ulva lactuca. Egypt. J. Bot. 2017, 57, 139–155. [Google Scholar]
- Escapa, C.; Coimbra, R.N.; Nuevo, C.; Vega, S.; Paniagua, S.; García, A.I.; Calvo, L.F.; Otero, M. Valorization of Microalgae Biomass by Its Use for the Removal of Paracetamol from Contaminated Water. Water 2017, 9, 312. [Google Scholar] [CrossRef]

| Compounds | Compounds Source | Strains | Removal Mechanisms | Technologies | Reference |
|---|---|---|---|---|---|
| Ibuprofen, clofibric acid, and carbamazepine | Synthetic media | Fungi Trametes versicolor Irpex lacteus Ganoderma lucidum Phanerochaete chrysosporium | Biodegradation Adsorption | Laboratory-scale batch assays | [46] |
| Carbamazepine | Synthetic media | Fungi Trametes versicolor | Biodegradation Adsorption | Laboratory-scale batch assays Pilot-scale glass air-pulsed fluidized bioreactor (continuous and batch feed) | [47] |
| Citalopram, fluoxetine, sulfamethoxazole, diclofenac, ibuprofen, naproxen, carbamazepine, and diazepam | Synthetic media | Fungi Bjerkandera sp. R1 Bjerkandera adusta Phanerochaete chrysosporium | Biodegradation | Laboratory-scale batch assays | [48] |
| Carbamazepine | Synthetic media | Fungi Phanerochaete chrysosporium | Biodegradation Biosorption | Pilot-scale plate bioreactor (continuous and batch feed) | [49] |
| Naproxen, ibuprofen, acetaminophen, salicylic acid, ketoprofen, codeine, erythromycin, metronidazole, ciprofloxacine, azithromycin, cefalexine, propranolol, carbamazepine, 10,11-epoxycarbamazepine, 2-hydroxycarbamazepine, acridone, and citalopram | Urban wastewater | Fungi Trametes versicolor | Biodegradation | Pilot-scale air-fluidized bioreactor (batch feed) | [50] |
| Carbamazepine, ibuprofen, clofibric acid, ketoprofen, metronidazole, triclosan, 17-α-ethinylestradiol, 17-β-estradiol-17-acetate, estrone, estriol, 17-β-estradiol, gemfibrozil, amitriptyline, primidone, salicylic acid, diclofenac, naproxen | Synthetic media | Fungi Trametes versicolor | Biodegradation Biosorption | Laboratory-scale batch assays Pilot-scale fungus-augmented membrane bioreactor (continuous feed) | [51] |
| Acetaminophen, ibuprofen, ketoprofen, naproxen, salicylic acid, codeine, phenazone, dexamethasone, diclofenac, piroxicam | Hospital wastewater | Fungi Trametes versicolor | Biodegradation | Pilot-scale glass air pulsed fluidized bioreactor (batch feed) | [52] |
| X-ray contrast agent iopromide and antibiotic ofloxacin | Hospital wastewater | Fungi Trametes versicolor | Biodegradation | Laboratory-scale batch assays Pilot-scale glass air-pulsed fluidized bioreactor (batch feed) | [53] |
| Metronidazole, salicylic acid, primidone, amitriptyline, carbamazepine, ketoprofen, naproxen, ibuprofen, gemfibrozil, diclofenac, triclosan, estriol, estrone, 17-α-ethinylestradiol, 17-β-estradiol, 17-β-estradiol-17-acetate | Synthetic media | Fungi Trametes versicolor | Biodegradation Biosorption | Laboratory-scale batch assays | [54] |
| Cefalexin, ciprofloxacin, metronidazole, trimethoprim, tetracycline, ketoprofen, acridone, carbamazepine, a carbamazepine metabolite, ciprofloxacin, metronidazole and its hydroxilated metabolite, β-blocker carazolol, diazepam, naproxen, cephalexin, tetracyclin, sertraline, paroxetine, gemfibrozil, amlodipine, furosemide, dimetridazole, azythromycin, ronidazole, olanzapine, piroxicam, β-blockers metoprolol | Veterinary hospital wastewater | Fungi Trametes versicolor | Biodegradation | Pilot-scale glass air-pulsed fluidized bioreactor (continuous and batch feed) | [55] |
| Acetaminophen, carbamazepine, diclofenac, metoprolol, naproxean, ranitidine, and sulfamethoxazole | Synthetic media | Fungi Aspergillus niger Algae Chlorella vulgaris | Biodegradation | Laboratory-scale batch assays | [56] |
| Estrone, 17β-estradiol, 17α-ethinyl-estradiol, and estriol | WWTP wastewater | Fungi Trametes versicolor Myceliophthora thermophila | Biodegradation Adsorption | Laboratory-scale batch assays | [57] |
| 17β-estradiol and 17α-ethinylestradiol | Synthetic media | Fungi Trametes versicolor | Biodegradation | Laboratory-scale batch assays Pilot-scale glass air-fluidized bioreactor (continuous feed) | [58] |
| Sulfapyridine, sulfapyridine, and sulfamethazine | Synthetic media | Fungi Trametes versicolor | Biodegradation Biosorption | Laboratory-scale batch assays Pilot-scale air-pulsed fluidized-bed bioreactor (continuous feed) | [59] |
| Naproxen and carbamazepine | Synthetic media | Fungi Trametes versicolor | Biodegradation | Laboratory-scale batch assays | [60] |
| Sodium diclofenac | Synthetic media | Fungi Trametes versicolor | Biodegradation | Laboratory-scale batch assays | [61] |
| Ketoprofen | Synthetic media | Fungi Trametes versicolor | Biodegradation | Laboratory-scale batch assays | [62] |
| Diclofenac, ibuprofen, naproxen, carbamazepine, and diazepam | Synthetic media | Fungi Phanerochaete chrysosporium | Biodegradation Adsorption | Pilot-scale stirred tank reactor and fixed-bed reactor (continuous feed) | [64] |
| Tetracycline and oxytetracycline | Synthetic media | Fungi Phanerochaete chrysosporium | Biodegradation | Laboratory-scale batch assays | [65] |
| Phenolic compounds | Pharmaceutical industry wastewater | Pycnoporus sanguineus | Biodegradation | Laboratory-scale batch assays | [66] |
| Diclofenac, ketoprofen and atenolol | Hospital wastewater | Fungi Pleurotus ostreatus | Biodegradation Adsorption | Pilot-scale air-pulsed fluidized-bed bioreactor (continuous and batch feed) Laboratory-scale batch assays | [67] |
| 17α-ethinylestradiol and carbamazepine | Synthetic media | Fungi Pleurotus sp. P1 Pleurotus ostreatus BS (unidentified) basidiomycete strain BNI | Biodegradation Adsorption | Laboratory-scale batch assays | [68] |
| Acetaminophen | Synthetic media | Fungi Mucor hiemalis | Bioconcentration | Laboratory-scale batch assays | [69] |
| Carbamazepine and clarithromycin | Synthetic media | Fungi Trichoderma harzianumPleurotus ostreatus | Biodegradation | Laboratory-scale batch assays | [70] |
| Carbamazepine | Synthetic media | Fungi Pleurotus ostreatus | Biodegradation | Laboratory-scale batch assays | [71] |
| Clofibric acid, gemfibrozil, ibuprofen, fenoprofen, ketoprofen, naproxen, diclofenac, indomethacin, propyphenazone, and carbamazepine | Synthetic media | Fungi Trametes versicolor | Biodegradation | Laboratory-scale batch assays | [72] |
| Levofloxacin | Synthetic media | Algae Chlorella vulgaris | Biodegradation Bioaccumulation | Laboratory-scale batch assays | [133] |
| Acetaminophen, ibuprofen, ketoprofen, naproxen, carbamazepine, diclofenac, and triclosan | WWTP wastewater | Algae Lessonia nigrescens Bory Macrocystis integrifolia Bory | Biodegradation Photodegradation Biosorption | Pilot-scale high-rate algal ponds | [148] |
| Tetracycline | Synthetic media | Algae Chlorella vulgaris | Photodegradation Biosorption | Laboratory-scale batch assays Pilot-scale high rate algal ponds (batch feed) | [164] |
| Carbamazepine | Synthetic media | Algae Pseudokirchneriella subcapitata (and crustacean Thamnocephalus and cnidarian Hydra attenuata) | Bioaccumulation | Laboratory-scale batch assays | [166] |
| Progesterone and norgestrel | Synthetic media | Algae Scenedesmus obliquus Chlorella pyrenoidosa | Biodegradation Biosorption | Laboratory-scale batch assays | [168] |
| Carbamazepine | Synthetic media | Algae Chlamydomonas mexicana Scenedesmus obliquus | Biodegradation Adsorption Bioaccumulation | Laboratory-scale batch assays | [169] |
| 17α-Ethynylestradiol | Synthetic media | Algae Desmodesmus subspicatus | Biotransformation Bioconcentration | Laboratory-scale batch assays | [170] |
| Analgesic and antiinflammatories, lipid regulators and antihypertensive, psychiatric drugs and stimulant, antibiotics, and others | WWTP wastewater | Algae Coelastrum sp. | Biodegradation | Pilot-scale photobiotreatment microalgae and multi-barrier treatment | [176] |
| Acetaminophen, ibuprofen, naproxen, salicylic acid, ketoprofen, codeine, azithromycin, erythromycin, ciprofloxacin, ofloxacin, atenolol, lorazepam, alprazolam, paroxetine, hydrochlorothiazide, furosemide, and diltiazem | Domestic wastewater | Algae Undefined microalgae | Biodegradation | Pilot-scale tubular photobioreactor | [177] |
| Alfuzosin, alprazolam, atenolol, atracurium, azelastine, biperiden, bisoprolol, bupropion, carbamazepin, cilazapril, ciprofloxacin, citalopram, clarithromycine, clemastine, clindamycine, clonazepam, clotrimazol, codeine, cyproheptadine, desloratidin, dicycloverin, diltiazem, diphenhydramin, eprosartan, fexofenadine, flecainide, fluconazole, flupetixol, haloperidol, hydroxyzine, ibersartan, loperamide, memantin, metoprolol, miconazole, mirtazapine, nefazodon, orphenadrin, pizotifen, ranitidine, risperidone, roxithromycine, sertraline, sotalol, sulfamethoxazol, terbutalin, tramadol, trihexyphenidyl, trimetoprim, venlavafaxin, and verapamil | WWTP wastewater | Algae Green algal species (Tetradesmus dimorphus and Dictyosphaerium, between them) | Biodegradation | Real-scale photobioreactor | [178] |
| Estradiol, estrone, estriol and hydroxyestrone) and synthetic steroid estrogens (estradiol valerate, estradiol, and ethinylestradiol | Synthetic media | Algae Chlorella vulgaris | Biotransformation Bioconcentration | Laboratory-scale batch assays | [179] |
| 17α-ethinylestradiol, estrone, and 17β-estradiol | Synthetic media | Algae Anabaena cylindrical Chlorococcus Spirulina platensis Chlorella Scenedesmus quadricauda Anaebena (and duckweed Lemna) | Biodegradation Adsorption | Laboratory-scale batch assays Pilot-scale plug flow reactor (continuous feed) | [180] |
| Triclosan | Synthetic media | Algae Chlorella Pyrenoidosa | Biodegradation Biosorption | Laboratory-scale batch assays | [182] |
| Cefradinegree | Synthetic media | Algae Chlorella pyrenoidosa | Biodegradation | Pilot-scale batch-sequencing reactor algae process (batch feed) | [183] |
| Ethinylestradiol | Synthetic media | Algae Ankistrodesmus braunii Chlorella ellipsoidea Chlorella pyrenoidosa Chlorella vulgaris Scenedesmus communis Scenedesmus obliquus Scenedesmus quadricauda Scenedesmus vacuolatus Selenastrum capricornutum | Biotransformation | Laboratory-scale batch assays | [184] |
| 17α-boldenone, 17β-boldenone, 4-hydroxy-androst-4-ene-17-dione, androsta-1,4-diene-3,17-dione, 4-androstene-3,17-dione, carbamazepine, ciprofloxacin, clarithromycin, climbazole, clofibric acid, diclofenac, enrofloxacin, erythromycin–H2O, estrone, fluconazole, gemfibrozil, ibuprofen, lincomycin, lomefloxacin, norfloxacin, ofloxacin, paracetamol, progesterone, roxithromycin, salicylic acid, salinomycin, sulfadiazine, sulfadimethoxine, sulfameter, sulfamethazine, sulfamethoxzole, sulfamonomethoxine, sulfapyridine, testosterone, triclosan, trimethoprim, and tylosin | WWTP wastewater | Algae Chlamydomonas reinhardtii Scenedesmus obliquus Chlorella pyrenoidosa Chlorella vulgaris | Biodegradation | Laboratory-scale batch assays | [185] |
| Salicylic acid and paracetamol | Synthetic media | Algae Chlorella sorokiniana | Biodegradation | Pilot-scale reactor (batch and semicontinuous feed) | [186] |
| Paracetamol, salicylic acid, and diclofenac | Synthetic media | Algae Chlorella sorokiniana Chlorella vulgaris Scenedesmus obliquus | Biodegradation | Pilot-scale bubbling column photobioreactor (batch and semicontinuous feed) | [187] |
| Tributyltin | Synthetic media | Algae Chlorella miniata C. sorokiniana Scenedesmus dimorphus S. platydiscus | Biodegradation Adsorption Absorption | Laboratory-scale batch assays | [188] |
| Trimethoprim, sulfamethoxazole, and triclosan | Synthetic media | Algae Nannochloris Sp. | Biodegradation Adsorption | Laboratory-scale batch assays | [189] |
| Diclofenac, ibuprofen, paracetamol, metoprolol, carbamazepine and trimethoprim, estrone, 17β-estradiol, and ethinylestradiol | Synthetic media | Algae Chlorella sorokiniana | Biodegradation Photolysis Biosorption | Laboratory-scale batch assays | [190] |
| Oxytetracycline, doxycycline, chlortetracycline, and tetracycline | Synthetic media | Algae Haematoloccus pluvialis Chlorella sp. Selenastrum capricornutum Pseudokirchneriella subcapitata | Biodegradation | Laboratory-scale batch assays | [191] |
| Salicylic acid or paracetamol | Pharmaceutical industry wastewater | Algae Chlorella sorokiniana | Biodegradation | Pilot-scale bubbling column photobioreactor (batch and semicontinuous feed) | [193] |
| Ciprofloxacin | Synthetic media | Algae Chlamydomonas mexicana | Biodegradation | Laboratory-scale batch assays | [194] |
| β-estradiol and 17α-ethinylestradiol | WWTP anaerobic sludge | Algae Selenastrum capricornutum Chlamydomonas reinhardtii | Biodegradation Photodegradation Adsorption | Laboratory-scale batch assays | [195] |
| Compounds | Compounds Source | Strains | Removal Mechanisms | Technologies | Reference |
|---|---|---|---|---|---|
| Ibuprofen, clofibric acid, and carbamazepine | Synthetic media | Fungi Trametes versicolor Irpex lacteus Ganoderma lucidum Phanerochaete chrysosporium | Biodegradation Adsorption | Laboratory-scale batch assays | [46] |
| Carbamazepine | Synthetic media | Fungi Trametes versicolor | Biodegradation Adsorption | Laboratory-scale batch assays Pilot-scale glass air-pulsed fluidized bioreactor (continuous and batch feed) | [47] |
| Carbamazepine | Synthetic media | Fungi Phanerochaete chrysosporium | Biodegradation Biosorption | Pilot-scale plate bioreactor (continuous and batch feed) | [49] |
| Carbamazepine, ibuprofen, clofibric acid, ketoprofen, metronidazole, triclosan, 17-α-ethinylestradiol, 17-β-estradiol-17-acetate, estrone, estriol, 17-β-estradiol, gemfibrozil, amitriptyline, primidone, salicylic acid, diclofenac, naproxen | Synthetic media | Fungi Trametes versicolor | Biodegradation Biosorption | Laboratory-scale batch assays Pilot-scale fungus-augmented membrane bioreactor (continuous feed) | [51] |
| Metronidazole, salicylic acid, primidone, amitriptyline, carbamazepine, ketoprofen, naproxen, ibuprofen, gemfibrozil, diclofenac, triclosan, estriol, estrone, 17-α-ethinylestradiol, 17-β-estradiol, 17-β-estradiol-17-acetate | Synthetic media | Fungi Trametes versicolor | Biodegradation Biosorption | Laboratory-scale batch assays | [54] |
| Estrone, 17β-estradiol, 17α-ethinylestradiol, and estriol | WWTP wastewater | Fungi Trametes versicolor Myceliophthora thermophila | Biodegradation Adsorption | Laboratory-scale batch assays | [57] |
| Sulfapyridine, sulfapyridine, and sulfamethazine | Synthetic media | Fungi Trametes versicolor | Biodegradation and biosorption | Laboratory-scale batch assays Pilot-scale air-pulsed fluidized bed bioreactor (continuous feed) | [59] |
| Diclofenac, ibuprofen, naproxen, carbamazepine, and diazepam | Synthetic media | Fungi Phanerochaete chrysosporium | Biodegradation Biosorption | Pilot-scale continuous stirred tank reactor and fixed-bed reactor | [64] |
| Diclofenac, ketoprofen, and atenolol | Hospital wastewater | Fungi Pleurotus ostreatus | Biodegradation Adsorption | Pilot-scale air-pulsed fluidized-bed bioreactor (continuous and batch feed) Laboratory-scale batch assays | [67] |
| 17α-ethinylestradiol and carbamazepine | Synthetic media | Fungi Pleurotus sp. P1 Pleurotus ostreatus BS (unidentified) basidiomycete strain BNI | Biodegradation Adsorption | Laboratory-scale batch assays | [68] |
| Tetracycline | Synthetic media | Algae Chlorella vulgaris | Photodegradation Biosorption | Laboratory-scale batch assays Pilot-scale high-rate algal ponds (batch feed) | [164] |
| Progesterone and norgestrel | Synthetic media | Algae Scenedesmus obliquus Chlorella pyrenoidosa | Biodegradation Biosorption | Laboratory-scale batch assays | [168] |
| Carbamazepine | Synthetic media | Algae Chlamydomonas Mexicana Scenedesmus obliquus | Biodegradation Adsorption Bioaccumulation | Laboratory-scale batch assays | [169] |
| Acetaminophen, ibuprofen, ketoprofen, naproxen, carbamazepine, diclofenac, and triclosan | WWTP wastewater | Algae Lessonia nigrescens Bory Macrocystis integrifolia Bory | Biodegradation Photodegradation Biosorption | Pilot-scale high-rate algal ponds | [175] |
| 17α-ethinylestradiol, estrone, and 17β-estradiol | Synthetic media | Algae Anabaena cylindrical Chlorococcus Spirulina platensis, Chlorella Scenedesmus quadricauda Anaebena (and duckweed Lemna) | Biodegradation Adsorption | Laboratory-scale batch assays Pilot-scale plug flow reactor (continuous feed) | [180] |
| Triclosan | Synthetic media | Algae Chlorella Pyrenoidosa | Biodegradation Biosorption | Laboratory-scale batch assays | [182] |
| Tributyltin | Synthetic media | Algae Chlorella miniata Chlorella sorokiniana Scenedesmus dimorphus Scenedesmus platydiscus | Biodegradation Adsorption Absorption | Laboratory-scale batch assays | [188] |
| Trimethoprim, sulfamethoxazole, and triclosan | Synthetic media | Algae Nannochloris Sp. | Biodegradation Adsorption | Laboratory-scale batch assays | [189] |
| Diclofenac, ibuprofen, paracetamol, metoprolol, carbamazepine and trimethoprim, estrone, 17β-estradiol, and ethinylestradiol | Synthetic media | Algae Chlorella sorokiniana | Biodegradation Photolysis Biosorption | Laboratory-scale batch assays | [190] |
| β-estradiol and 17α-ethynylestradiol | WWTP anaerobic sludge | Algae Selenastrum capricornutum Chlamydomonas reinhardtii | Biodegradation Photodegradation Adsorption | Laboratory-scale batch assays | [194] |
| Sulfamethoxazole and sulfacetamide | Synthetic media | Algae Lessonia nigrescens Bory Macrocystis integrifolia Bory | Adsorption | Laboratory-scale batch assays | [211] |
| Penicillin G | Synthetic media | Fungi Rhizopus arrhizus | Biosorption | Laboratory-scale batch assays | [211] |
| Chloramphenicol, acetyl salicylic acid, clofibric acid | Synthetic media | Algae Pterocladia capillacea Ulva lactuca | Biosorption | Laboratory-scale batch assays | [225] |
| Paracetamol | Synthetic media | Algae Alga Synechocystis sp. | Biosorption | Laboratory-scale batch assays Pilot-scale continuous bubbling column photobioreactor (continuous feed) | [226] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silva, A.; Delerue-Matos, C.; Figueiredo, S.A.; Freitas, O.M. The Use of Algae and Fungi for Removal of Pharmaceuticals by Bioremediation and Biosorption Processes: A Review. Water 2019, 11, 1555. https://doi.org/10.3390/w11081555
Silva A, Delerue-Matos C, Figueiredo SA, Freitas OM. The Use of Algae and Fungi for Removal of Pharmaceuticals by Bioremediation and Biosorption Processes: A Review. Water. 2019; 11(8):1555. https://doi.org/10.3390/w11081555
Chicago/Turabian StyleSilva, Andreia, Cristina Delerue-Matos, Sónia A. Figueiredo, and Olga M. Freitas. 2019. "The Use of Algae and Fungi for Removal of Pharmaceuticals by Bioremediation and Biosorption Processes: A Review" Water 11, no. 8: 1555. https://doi.org/10.3390/w11081555
APA StyleSilva, A., Delerue-Matos, C., Figueiredo, S. A., & Freitas, O. M. (2019). The Use of Algae and Fungi for Removal of Pharmaceuticals by Bioremediation and Biosorption Processes: A Review. Water, 11(8), 1555. https://doi.org/10.3390/w11081555








