The Impact of a Fish Cannery Wastewater Discharge on the Bacterial Community Structure and Sanitary Conditions of Marine Coastal Sediments
Abstract
1. Introduction
2. Materials and Methods
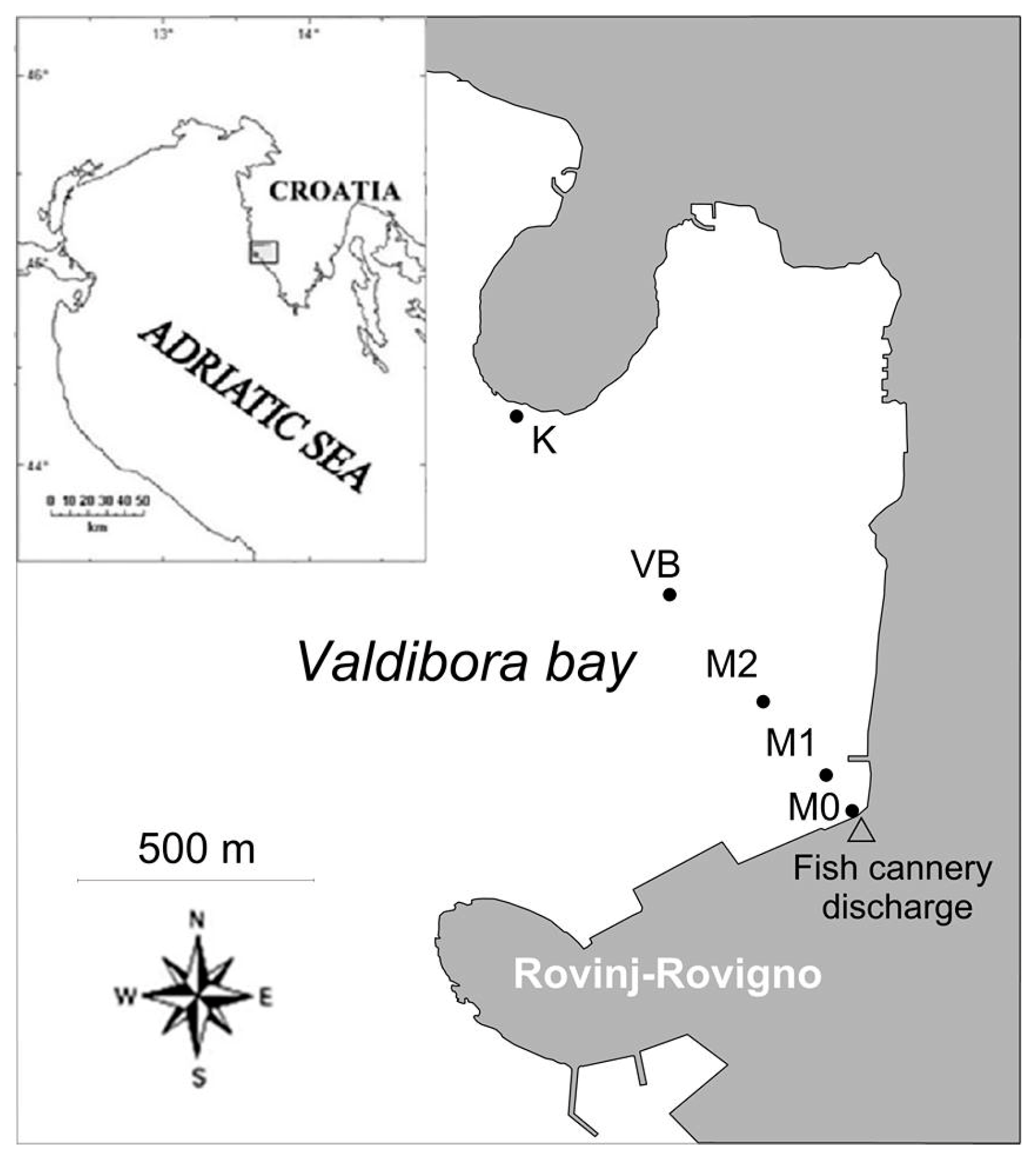
2.1. Study Location and Sampling
2.2. Redox Potential (Eh), Granulometric Composition and Permeability
2.3. Sanitary Quality of Sediments and Wastewaters
2.4. Prokaryotic Abundance (PA)
2.5. Total Lipid and Sterol Analysis
2.6. Extraction and Sequencing of DNA
3. Results
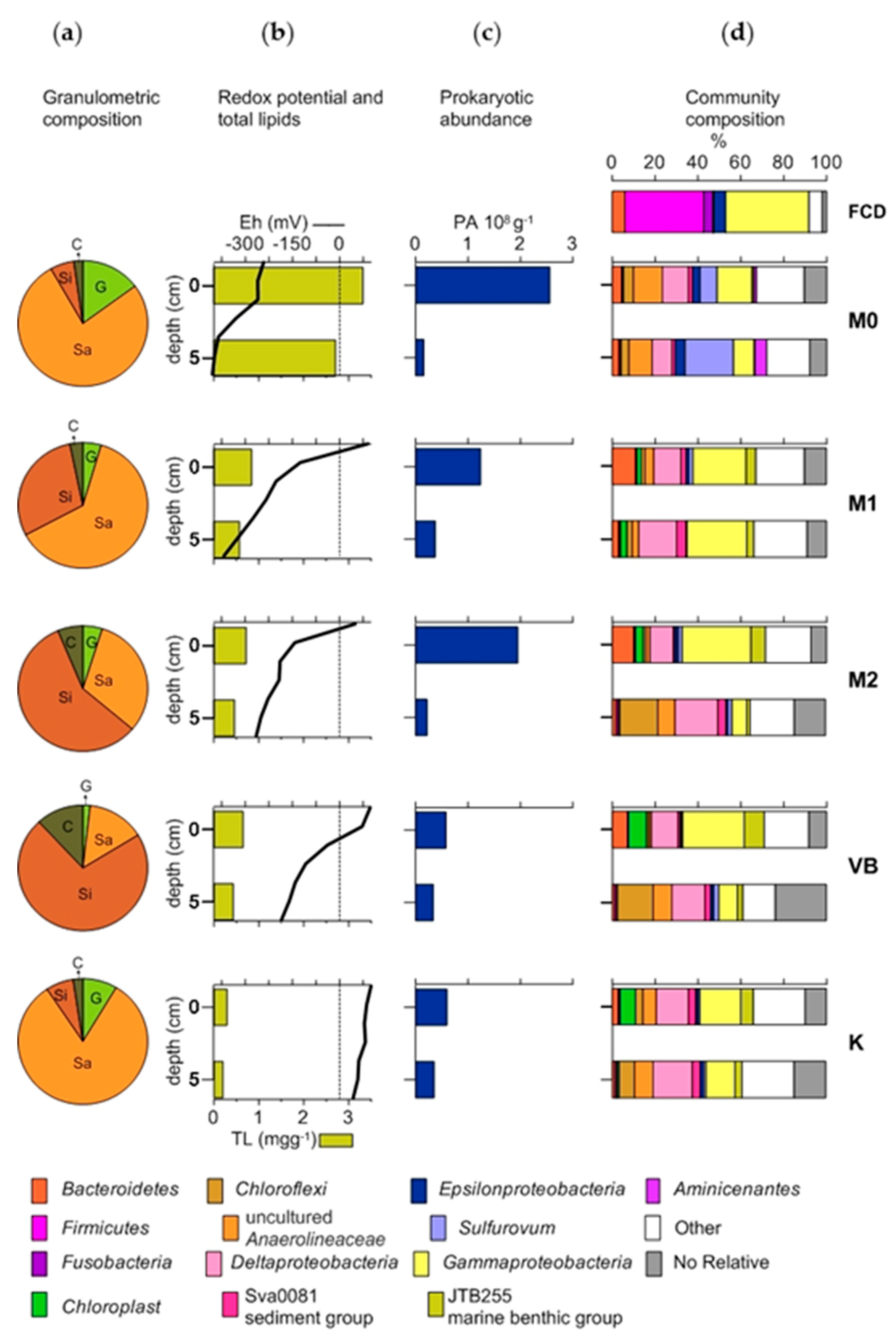
3.1. Granulometric Composition, Permeability and Redox Potential (Eh)
3.2. Granulometric Composition, Permeability and Redox Potential (Eh)
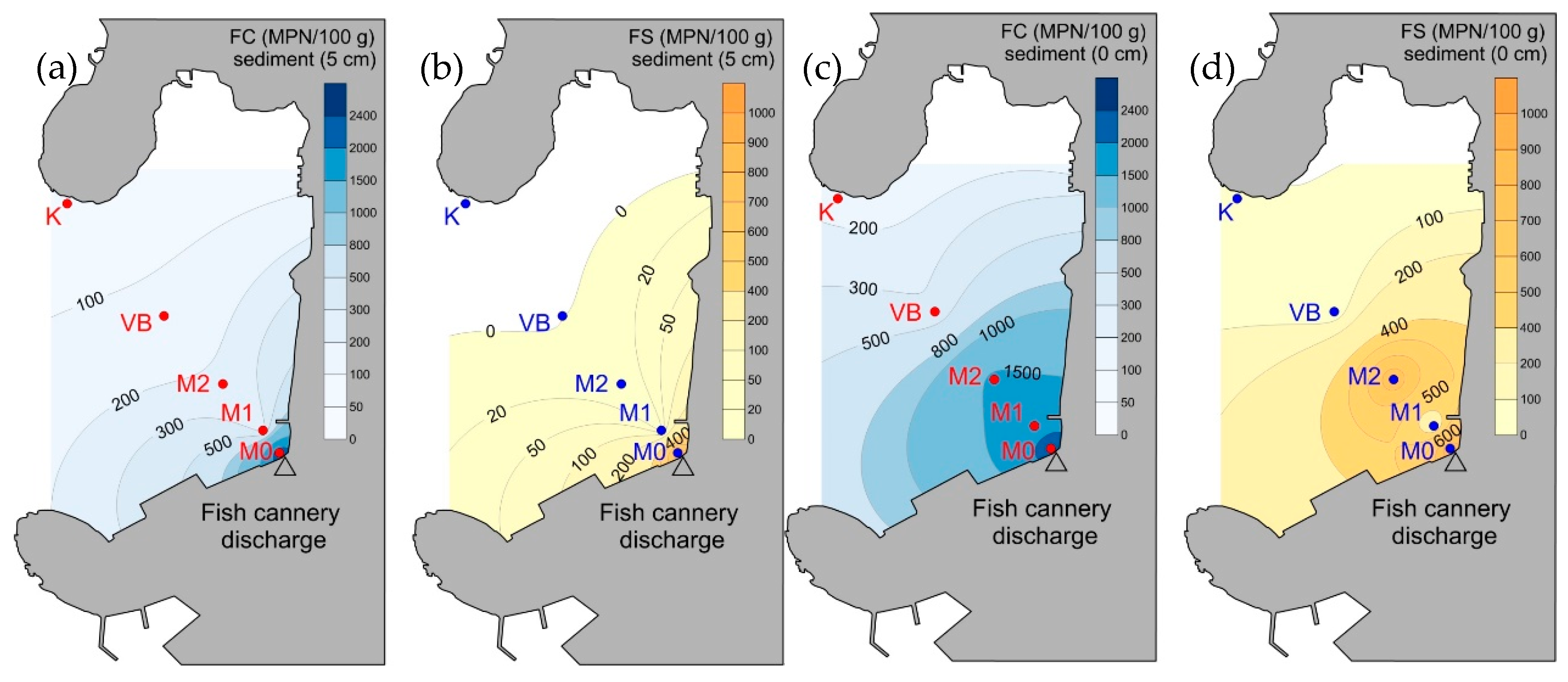
3.3. Prokaryotic Abundance and Faecal Indicator Bacteria
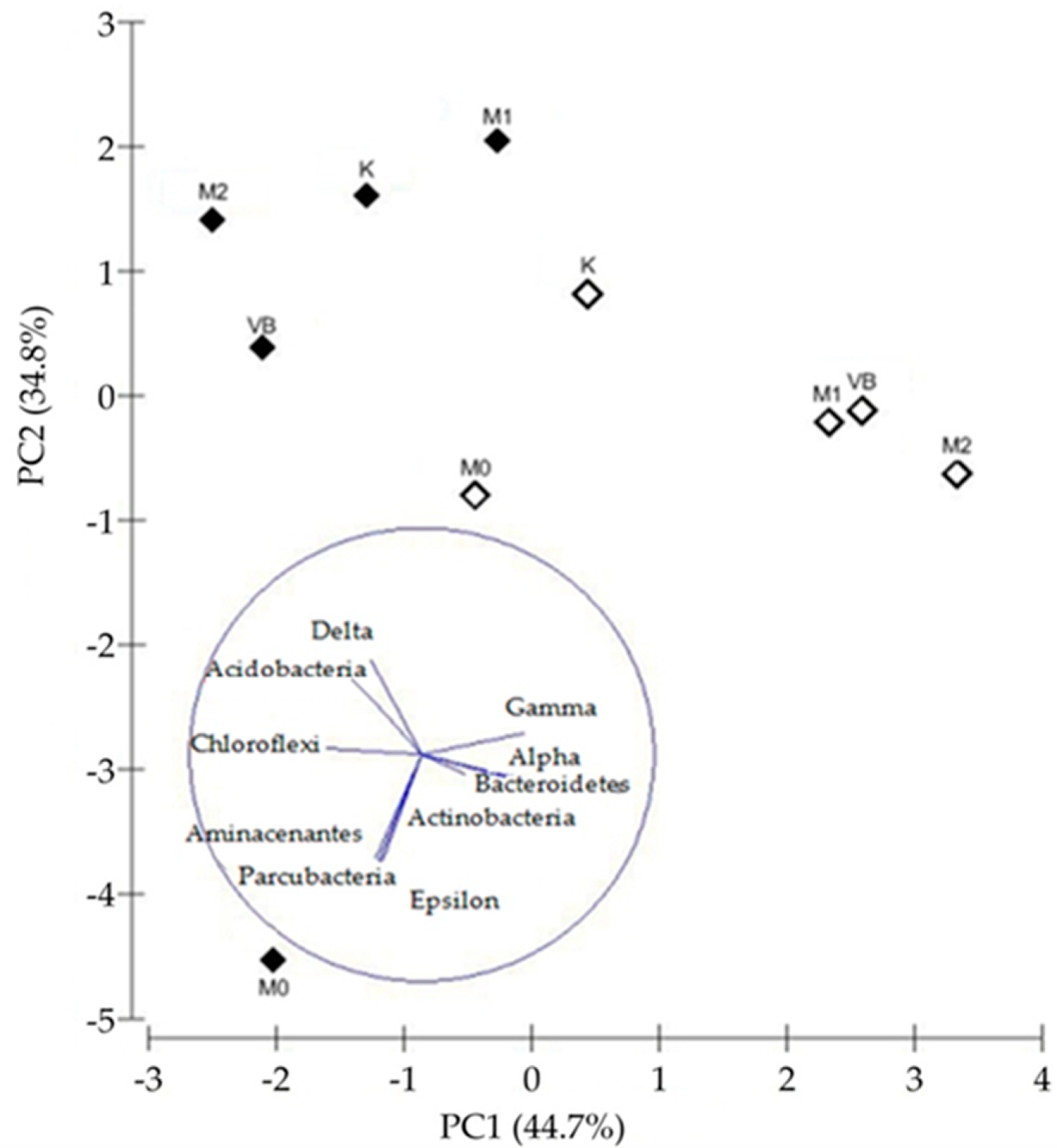
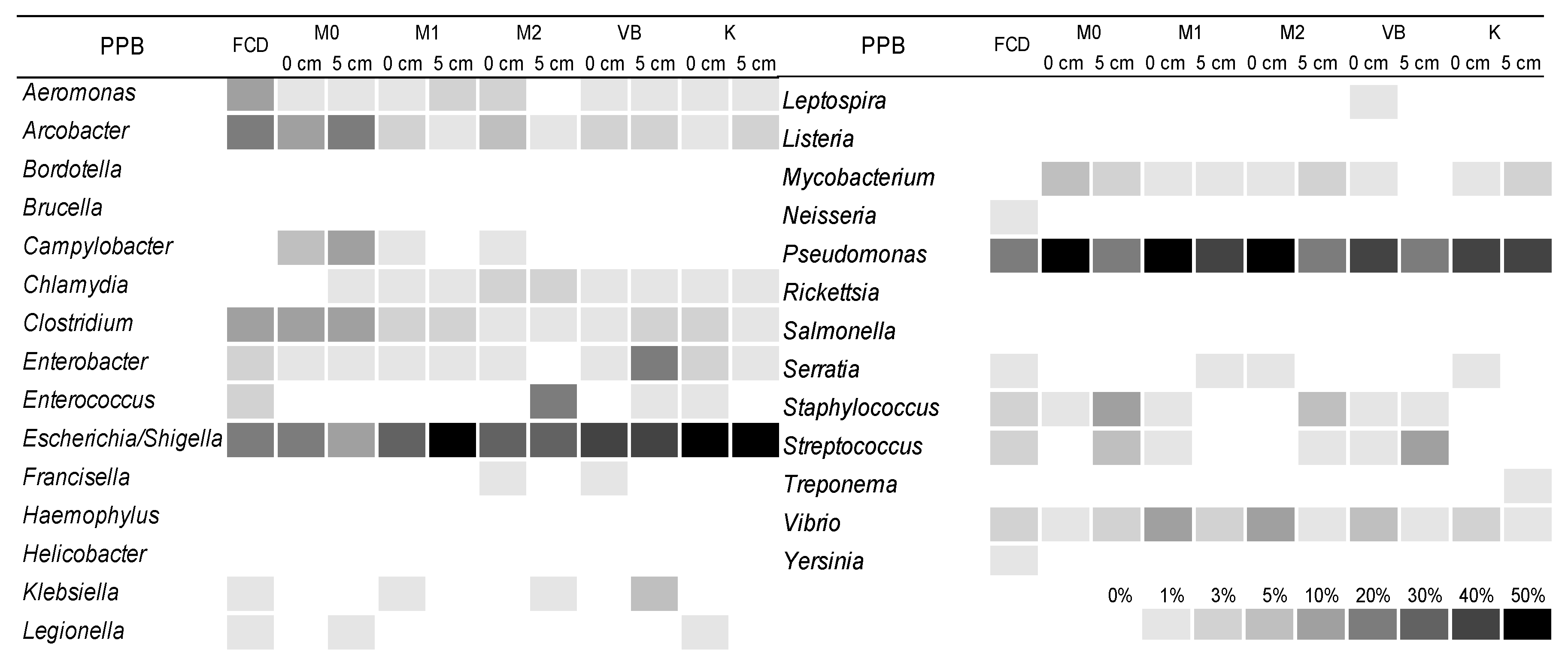
3.4. The Structure of Bacterial Community
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- El-Kholy, R.; El Saadi, A.; Abdin, A.E.D. Pollution and Measures towards De-Pollution of the Mediterranean Sea. In Integrated Water Resources Management in the Mediterranean Region: Dialogue towards New Strategy; Springer: Dordrecht, The Netherlands, 2012; pp. 175–194. [Google Scholar]
- Militon, C.; Hamdi, O.; Michotey, V.; Fardeau, M.L.; Ollivier, B.; Bouallagui, H.; Hamdi, M.; Bonin, P. Ecological significance of Synergistetes in the biological treatment of tuna cooking wastewater by an anaerobic sequencing batch reactor. Environ. Sci. Pollut. Res. 2015, 22, 18230–18238. [Google Scholar] [CrossRef] [PubMed]
- El-Bestawy, E.; El-Masry, M.H.; El-Adl, N.E. The potentiality of free Gram-negative bacteria for removing oil and grease from contaminated industrial effluents. World J. Microbiol. Biotechnol. 2005, 21, 815–822. [Google Scholar] [CrossRef]
- Garcia, J.L.; Uhia, I.; Galán, B. Catabolism and biotechnological applications of cholesterol degrading bacteria. Microb. Biotechnol. 2012, 5, 679–699. [Google Scholar] [CrossRef] [PubMed]
- Phong, N.T.; Duyen, N.T.; Diep, C.N. Isolation and characterization of lipid-degrading bacteria in wastewater of food processing plants and restaurants in Can Tho city, Vietnam. Am. J. Life Sci. 2014, 2, 382–388. [Google Scholar] [CrossRef]
- Hassan, S.W.M.; Latif, H.H.A.E.; Ali, S.M. Production of Cold-Active Lipase by Free and Immobilized Marine Bacillus cereus HSS: Application in Wastewater Treatment. Front. Microbiol. 2018, 9, 2377. [Google Scholar] [CrossRef]
- Sandaruwani, A.; Kumarasinghe, C.; Samarakoon, D.; Ariyadasa, T.U.; Gunawardena, S.H.P. Investigation of the Efficiency of Dairy Wastewater Treatment Using Lipid-Degrading Bacterial Strains. In Proceedings of the Moratuwa Engineering Research Conference (MERCon), Moratuwa, Sri Lanka, 30 May–1 June 2018; pp. 362–366. [Google Scholar] [CrossRef]
- Kurhekar, J.V. Isolation and Characterization of Lipase Producing Bacteria from Restaurant Waste Water. World J. Pharm. Res. 2018, 6, 685–693. [Google Scholar]
- Novotny, L.; Dvorska, L.; Lorencova, A.; Beran, V.; Pavlik, I. Fish: A potential source of bacterial pathogens for human beings. Vet. Med. Czech 2004, 49, 343–358. [Google Scholar] [CrossRef]
- Fazio, F.; Saoca, C.; Costa, G.; Zumbo, A.; Piccione, G.; Parrino, V. Flow cytometry and automatic blood cell analysis in striped bass Morone saxatilis (Walbaum, 1792): A new hematological approach. Aquaculture 2019, 513, 734398. [Google Scholar] [CrossRef]
- Parrino, V.; Cappello, T.; Costa, G.; Cannavà, C.; Sanfilippo, M.; Fazio, F.; Fasulo, S. Comparative study of haematology of two teleost fish (Mugil cephalus and Carassius auratus) from different environments and feeding habits. Eur. Zool. J. 2018, 85, 194–200. [Google Scholar] [CrossRef]
- Parrino, V.; Costa, G.; Cannavà, C.; Fazio, E.; Bonsignore, M.; Saoca, C.; Piccione, G.; Fazio, F. Flow cytometry and micro-Raman spectroscopy: Identification of hemocyte populations in the mussel Mytilus galloprovincialis (Bivalvia: Mytilidae) from Faro Lake and Tyrrhenian Sea (Sicily, Italy). Fish Shellfish Immunol. 2018, 87, 1–8. [Google Scholar] [CrossRef]
- Inagaki, F.; Nunoura, T.; Nakagawa, S.; Teske, A.; Lever, M.; Lauer, A.; Suzuki, M.; Takai, K.; Delwiche, M.; Colwell, F.S.; et al. Biogeographical distribution and diversity of microbes in methane hydrate-bearing deep marine sediments on the Pacific Ocean Margin. Proc. Natl. Acad. Sci. USA 2006, 103, 2815–2820. [Google Scholar] [CrossRef]
- Dyksma, S.; Bischof, K.; Fuchs, B.M.; Hoffmann, K.; Meier, D.; Meyerdierks, A.; Pjevac, P.; Probandt, D.; Richter, M.; Stepanauskas, R.; et al. Ubiquitous Gammaproteobacteria dominate dark carbon fixation in coastal sediments. ISME J. 2016, 10, 1939–1953. [Google Scholar] [CrossRef]
- Mussmann, M.; Pjevac, P.; Krüger, K.; Dyksma, S. Genomic repertoire of the Woeseiaceae/JTB255, cosmopolitan and abundant core members of microbial communities in marine sediments. ISME J. 2017, 11, 1276. [Google Scholar] [CrossRef]
- Zinger, L.; Amaral-Zettler, L.A.; Fuhrman, J.A.; Horner-Devine, M.C.; Huse, S.M.; Welch, D.B.M.; Martiny, J.B.H.; Sogin, M.; Boetius, A.; Ramette, A. Global Patterns of Bacterial Beta-Diversity in Seafloor and Seawater Ecosystems. PLoS ONE 2011, 6, e24570. [Google Scholar] [CrossRef]
- Gobet, A.; Boer, S.I.; Huse, S.M.; Van Beusekom, J.E.E.; Quince, C.; Sogin, M.L.; Boetius, A.; Ramette, A. Diversity and dynamics of rare and of resident bacterial populations in coastal sands. ISME J. 2012, 6, 542–553. [Google Scholar] [CrossRef]
- Pjevac, P.; Kamyshny, A., Jr.; Dyksma, S.; Mussmann, M. Microbial consumption of zero-valence sulfur in marine benthic habitats. Environ. Microbiol. 2014, 16, 3416–3430. [Google Scholar] [CrossRef]
- Zheng, B.; Wang, L.; Liu, L. Bacterial community structure and its regulating factors in the intertidal sediment along the Liaodong Bay of Bohai Sea, China. Microbiol. Res. 2014, 169, 585–592. [Google Scholar] [CrossRef]
- Probandt, D.; Knittel, K.; Tegetmeyer, H.E.; Ahmerkamp, S.; Holtappels, M.; Amann, R. Permeability shapes bacterial communities in sublittoral surface sediments. Environ. Microbiol. 2017, 19, 1584–1599. [Google Scholar] [CrossRef]
- Drury, B.; Rosy-Marshall, E.; Kelly, J.J. Wastewater Treatment Effluent Reduces the Abundance and Diversity of Benthic Bacterial Communities in Urban and Suburban Rivers. Appl. Environ. Microbiol. 2013, 79, 1897–1905. [Google Scholar] [CrossRef]
- Korlević, M.; Žućko, J.; Najdek, M.; Blažina, M.; Pustijanac, E.; Zeljko, T.V.; Gačeša, R.; Baranašić, D.; Starčević, A.; Diminić, J.; et al. Bacterial diversity of polluted surface sediments in the northern Adriatic Sea. Syst. Appl. Microbiol. 2015, 38, 189–197. [Google Scholar] [CrossRef]
- Micromeritics SediGraph 5100 Particle Size Analysis System Operator’ Manual; Micromeritics Instrument Corporation: Norcross, GA, USA, 2002; Available online: https://www.micromeritics.com/pdf/products/sedigraph_5100_brochure.pdf (accessed on 4 December 2019).
- Folk, R.L. The distinction between grain size and mineral composition in sedimentary rock nomenclature. J. Geol. 1954, 62, 344–356. [Google Scholar] [CrossRef]
- Gangi, A.F. Permeability of unconsolidated sands and porous rocks. J. Geophys. Res. Solid 1985, 90, 3099–3104. [Google Scholar] [CrossRef]
- WHO. Guidelines for Health Related Monitoring of Coastal Recreational and Shellfish Areas. Part I: General Guidelines. In Long-term Programme for Pollution Monitoring and Research in the Mediterranean Sea (MED/POL Phase II); WHO: Geneva, Switzerland, 1994. [Google Scholar]
- Condalab, EVA Broth (Ethyl, Violet, Azide, Litsky) Cat. 1230, Conda Manual. Available online: https://www.chemie-brunschwig.ch/ (accessed on 11 November 2019).
- Epstein, S.S.; Rossel, J. Enumeration of sandy sediment bacteria: Search for optimal protocol. Mar. Ecol. Prog. Ser. 1995, 117, 289–298. [Google Scholar] [CrossRef]
- Bligh, E.G.; Dyer, W.J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 1959, 37, 910–917. [Google Scholar] [CrossRef]
- Isobe, K.O.; Tarao, M.T.; Zakaria, M.; Chiem, N.H.; Minh, L.Y.; Takada, H. Quantitative application of fecal sterols using gas chromatography-mass spectrometry to investigate fecal pollution in tropical waters: Western Malaysia and Mekong Delta, Vietnam. Environ. Sci. Technol. 2002, 36, 4497–4507. [Google Scholar] [CrossRef]
- Szűcs, S.; Sárváry, A.; Cain, T.; Ádány, R. Method validation for the simultaneous determination of fecal sterols in surface waters by gas chromatography—Mass spectrometry. J. Chromatogr. Sci. 2006, 44, 70–76. [Google Scholar] [CrossRef][Green Version]
- Massana, R.; Murray, A.E.; Preston, C.M.; Delong, E.F. Vertical distribution and phylogenetic characterization of marine planktonic archaea in the Santa Barbara Channel. Appl. Environ. Microbiol. 1997, 63, 50–56. [Google Scholar]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microb. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, 590–596. [Google Scholar] [CrossRef]
- Ionescu, D.; Christian, S.; Polerecky, L.; Munwes, Y.Y.; Lott, C.; Häusler, S.; Bižić-Ionescu, M.; Quast, C.; Peplies, J.; Glöckner, F.O.; et al. Microbial and Chemical Characterization of Underwater Fresh Water Springs in the Dead Sea. PLoS ONE 2012, 7, e38319. [Google Scholar] [CrossRef] [PubMed]
- Pruesse, E.; Peplies, J.; Glöckner, F.O. SINA: Accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 2012, 28, 1823–1829. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Ivančić, I.; Paliaga, P.; Pfannkuchen, M.; Đakovac, T.; Najdek, M.; Steiner, P.; Korlević, M.; Markovski, M.; Baričević, A.; Smodlaka Tanković, M.; et al. Seasonal variations in extracellular enzymatic activity in marine snow-associated microbial communities and their impact on the surrounding water. FEMS Microbiol. Ecol. 2018, 94, 12. [Google Scholar] [CrossRef]
- Fuks, D.; Ivančić, I.; Najdek, M.; Njire, J.; Godrijan, J.; Maric Pfannkuchen, D.; Silovic, T.; Paliaga, P.; Blazina, M.; Precali, R.; et al. Changes in the planktonic community structure related to trophic conditions: The case study of the northern Adriatic Sea. J. Mar. Syst. 2012, 96, 95–102. [Google Scholar] [CrossRef][Green Version]
- Coyne, M.S.; Howell, J.M. The Fecal Coliform/Fecal Streptococci Ratio (FC/FS) And Water Quality in the Bluegrass Region of Kentucky. Soil Sci. 1994, 15, 1–4. [Google Scholar]
- Collado, L.; Figueras, M.J. Taxonomy, epidemiology and clinical relevance of the genus Arcobacter. Clin. Microbiol. Rev. 2011, 24, 174–192. [Google Scholar] [CrossRef]
- Paliaga, P.; Felja, I.; Ušić, U.; Ivančić, I.; Najdek, M. Accumulation and persistence of sewage and fish cannery pollution in coastal sediments (northern Adriatic Sea). J. Soils Sediments 2017. [Google Scholar] [CrossRef]
- Freier, T.A.; Beitz, D.C.; Li, L.; Hartman, P.A. Characterization of Eubacterium coprostanoligenes sp. nov., a cholesterol-reducing anaerobe. Int. J. Syst. Bacteriol. 1994, 44, 137–142. [Google Scholar] [CrossRef]
- Lye, H.S.; Rahmat-Ali, G.R.; Liong, M.T. Mechanisms of cholesterol removal by lactobacilli under conditions that mimic the human gastrointestinal tract. Int. Dairy J. 2010, 20, 169–175. [Google Scholar] [CrossRef]
- Omil, F.; Mendez, R.; Lema, J.M. Anaerobic treatment of saline wastewaters under high sulphide and ammonia content. Bioresour. Technol. 1995, 54, 269–278. [Google Scholar] [CrossRef]
- Murray, J.; Burt, J.R. The Composition of Fish; FAO Repository; FAO in Partnership with Support Unit for International Fisheries and Aquatic Research, SIFAR, Ministry of Technology, Torry Research Station: Rome, Italy, 2001. [Google Scholar]
- Gevertz, D.; Telang, A.J.; Voordouw, G.; Jenneman, G.E. Isolation and Characterization of Strains CVO and FWKO B, Two Novel Nitrate-Reducing, Sulfide-Oxidizing Bacteria Isolated from Oil Field Brine. Appl. Environ. Microbiol. 2000, 66, 2491–2501. [Google Scholar] [CrossRef] [PubMed]
- Lavik, G.; Stührmann, T.; Brüchert, V.; Van der Plas, A.; Mohrholz, V.; Lam, P.; Mussmann, M.; Fuchs, B.M.; Amann, R.; Lass, U.; et al. Detoxification of sulphidic African shelf waters by blooming chemolithotrophs. Nature 2009, 457, 581–585. [Google Scholar] [CrossRef]
- Inagaki, F.; Takai, K.; Kobayashi, H.; Nealson, K.H.; Horikoshi, K. Sulfurimonas autotrophica gen.nov., sp.nov., a novel sulfur-oxidizing epsilonproteobacterium isolated from hydrothermal sediments in the Mid-Okinawa Trough. Int. J. Syst. Evol. Microbiol. 2003, 53, 1801–1805. [Google Scholar] [CrossRef]
- Liang, B.; Wang, L.Y.; Mbadinga, S.M.; Liu, J.F.; Yang, S.Z.; Gu, J.D.; Mu, B.Z. Anaerolineaceae and Methanosaeta turned to be the dominant microorganisms in alkanes-dependent methanogenic culture after long-term of incubation. AMB Express 2015, 5, 37. [Google Scholar] [CrossRef]
- Böer, S.I.; Arnosti, C.; van Beusekom, J.E.E.; Boetius, A. Temporal variations in microbial activities and carbon turnover in subtidal sandy sediments. Biogeosciences 2009, 6, 1149–1165. [Google Scholar] [CrossRef]
| Station. | Sediment Type | dg (µm) | k (m2) |
|---|---|---|---|
| M0 | gS | 816.4 | 4.83 × 10−10 |
| M1 | (g)mS | 103.9 | 7.83 × 10−12 |
| M2 | (g)sM | 48.3 | 1.69 × 10−12 |
| VB | (g)sM | 26.7 | 5.16 × 10−13 |
| K | gS | 682.4 | 3.38 × 10−10 |
| Sample | Station | Core Section (cm) | COP | CHL | β-SIT |
|---|---|---|---|---|---|
| Effluent | FCD | 1.86 | 1230.20 | 247.83 | |
| Sediment | M0 | 0 | 7.58 | 29.92 | 2.14 |
| 5 | 5.53 | 27.86 | 1.49 | ||
| M1 | 0 | 0.68 | 2.37 | 0.47 | |
| 5 | 0.28 | 0.49 | 2.65 | ||
| M2 | 0 | 0.31 | 1.17 | 0.48 | |
| 5 | 0.09 | 0.41 | 0.25 | ||
| VB | 0 | 0.10 | 0.68 | 0.25 | |
| 5 | 0.05 | 0.26 | 0.33 | ||
| K | 0 | 0.09 | 0.32 | 0.22 | |
| 5 | nd | 0.32 | nd |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Paliaga, P.; Felja, I.; Budiša, A.; Ivančić, I. The Impact of a Fish Cannery Wastewater Discharge on the Bacterial Community Structure and Sanitary Conditions of Marine Coastal Sediments. Water 2019, 11, 2566. https://doi.org/10.3390/w11122566
Paliaga P, Felja I, Budiša A, Ivančić I. The Impact of a Fish Cannery Wastewater Discharge on the Bacterial Community Structure and Sanitary Conditions of Marine Coastal Sediments. Water. 2019; 11(12):2566. https://doi.org/10.3390/w11122566
Chicago/Turabian StylePaliaga, Paolo, Igor Felja, Andrea Budiša, and Ingrid Ivančić. 2019. "The Impact of a Fish Cannery Wastewater Discharge on the Bacterial Community Structure and Sanitary Conditions of Marine Coastal Sediments" Water 11, no. 12: 2566. https://doi.org/10.3390/w11122566
APA StylePaliaga, P., Felja, I., Budiša, A., & Ivančić, I. (2019). The Impact of a Fish Cannery Wastewater Discharge on the Bacterial Community Structure and Sanitary Conditions of Marine Coastal Sediments. Water, 11(12), 2566. https://doi.org/10.3390/w11122566




