Genome-Wide Identification of NBS-Encoding Resistance Genes in Sunflower (Helianthus annuus L.)
Abstract
1. Introduction
2. Materials and Methods
2.1. Retrieval and Identification of Sunflower NBS-Encoding Genes
2.2. Phylogenetic Tree Construction
2.3. Chromosomal Locations, Clustering and Gene Structure
2.4. Ka/Ks and Syntenic Analysis
2.5. Gene Homology and Expression Analysis
3. Results
3.1. Diversity of the NBS-Encoding Genes in Sunflower
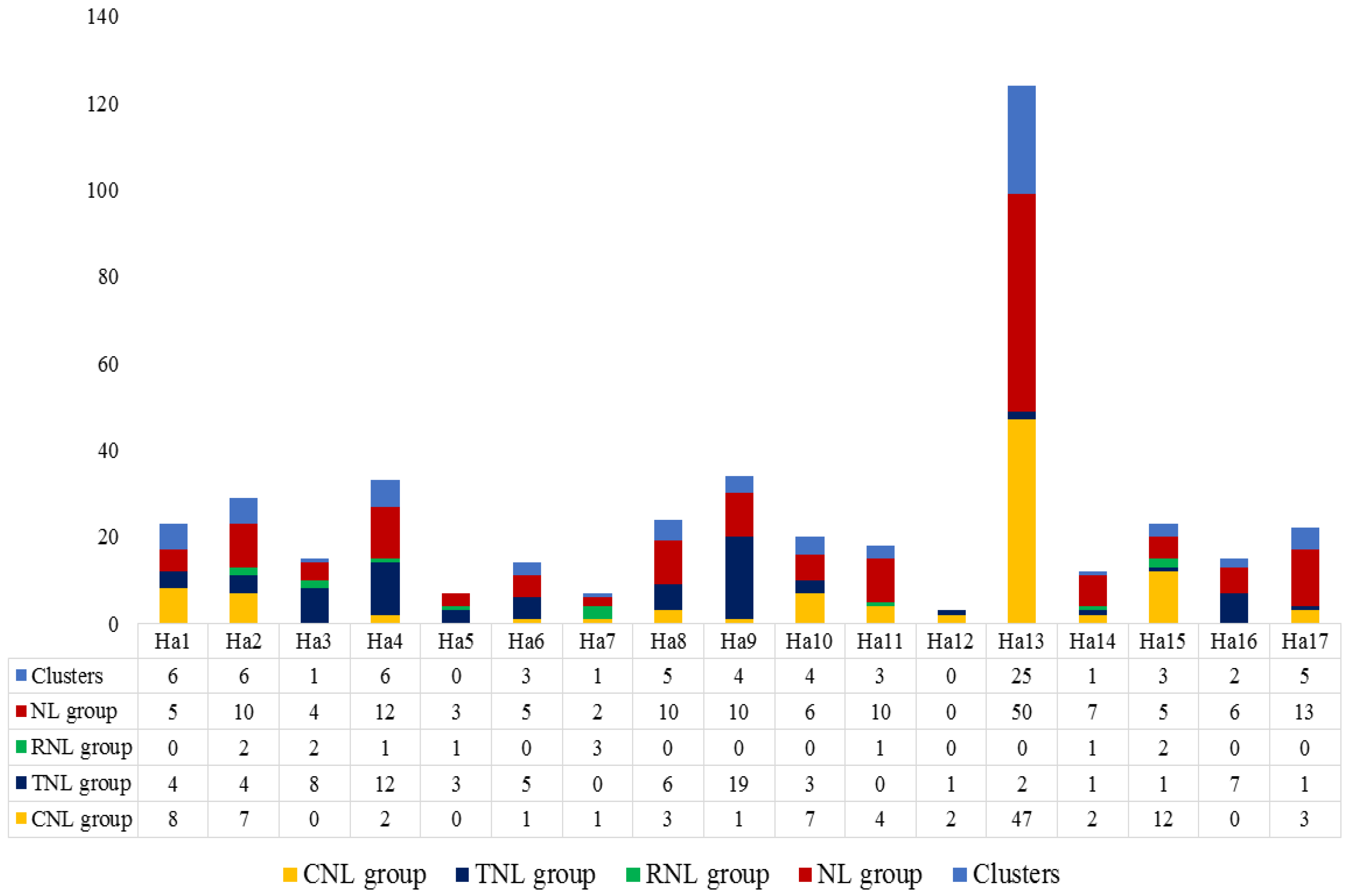
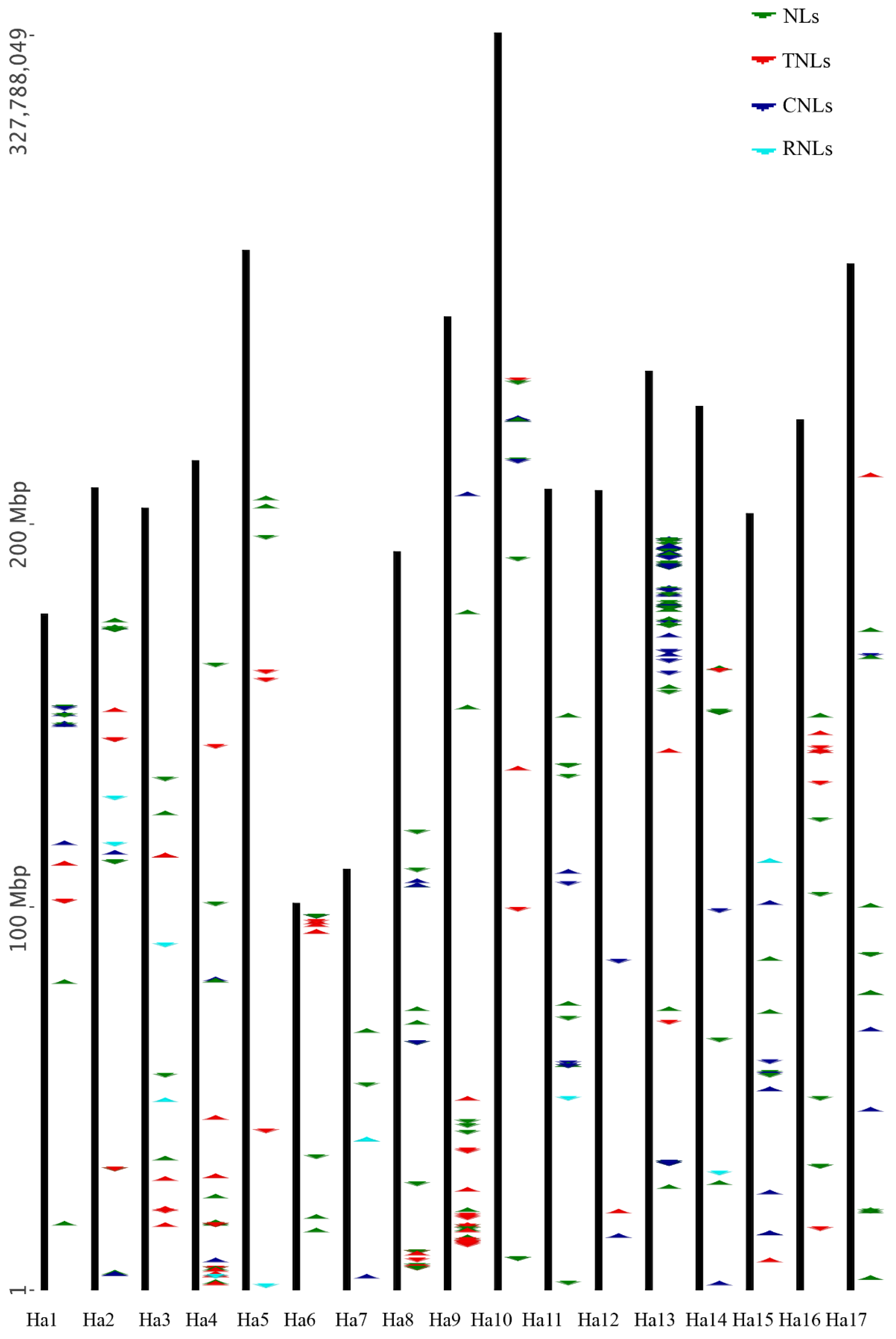
3.2. Gene Location, Clustering, Ka/Ks Values and Structural Variation
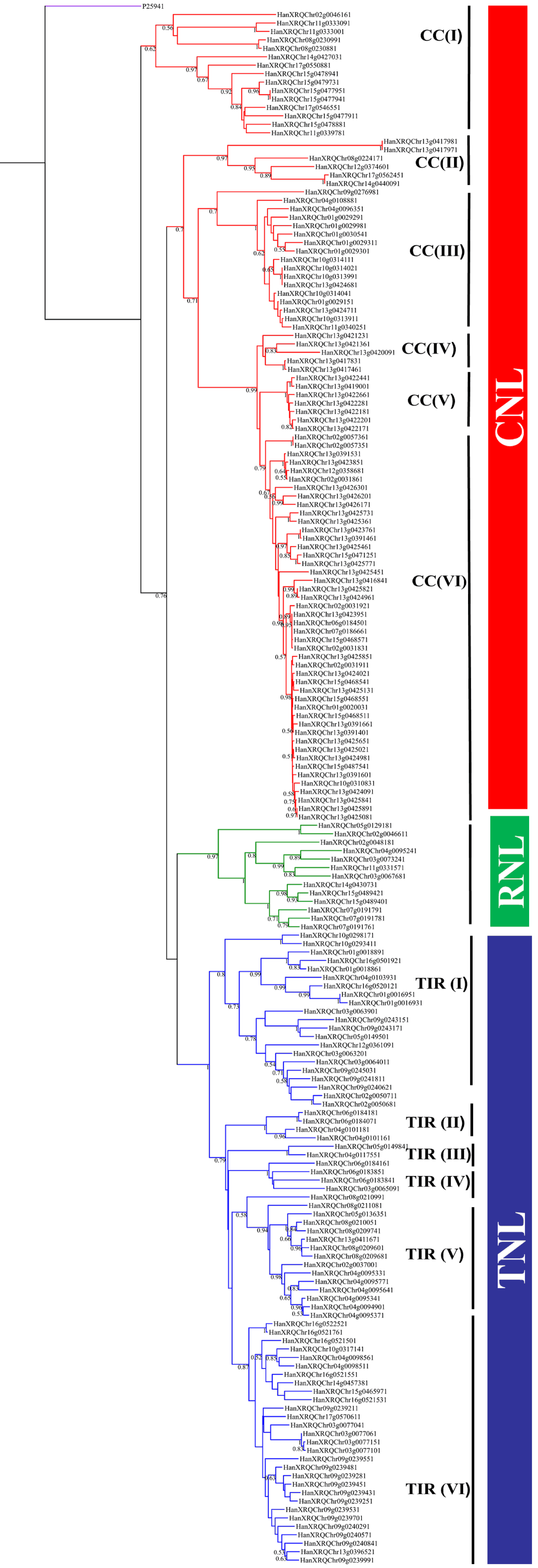
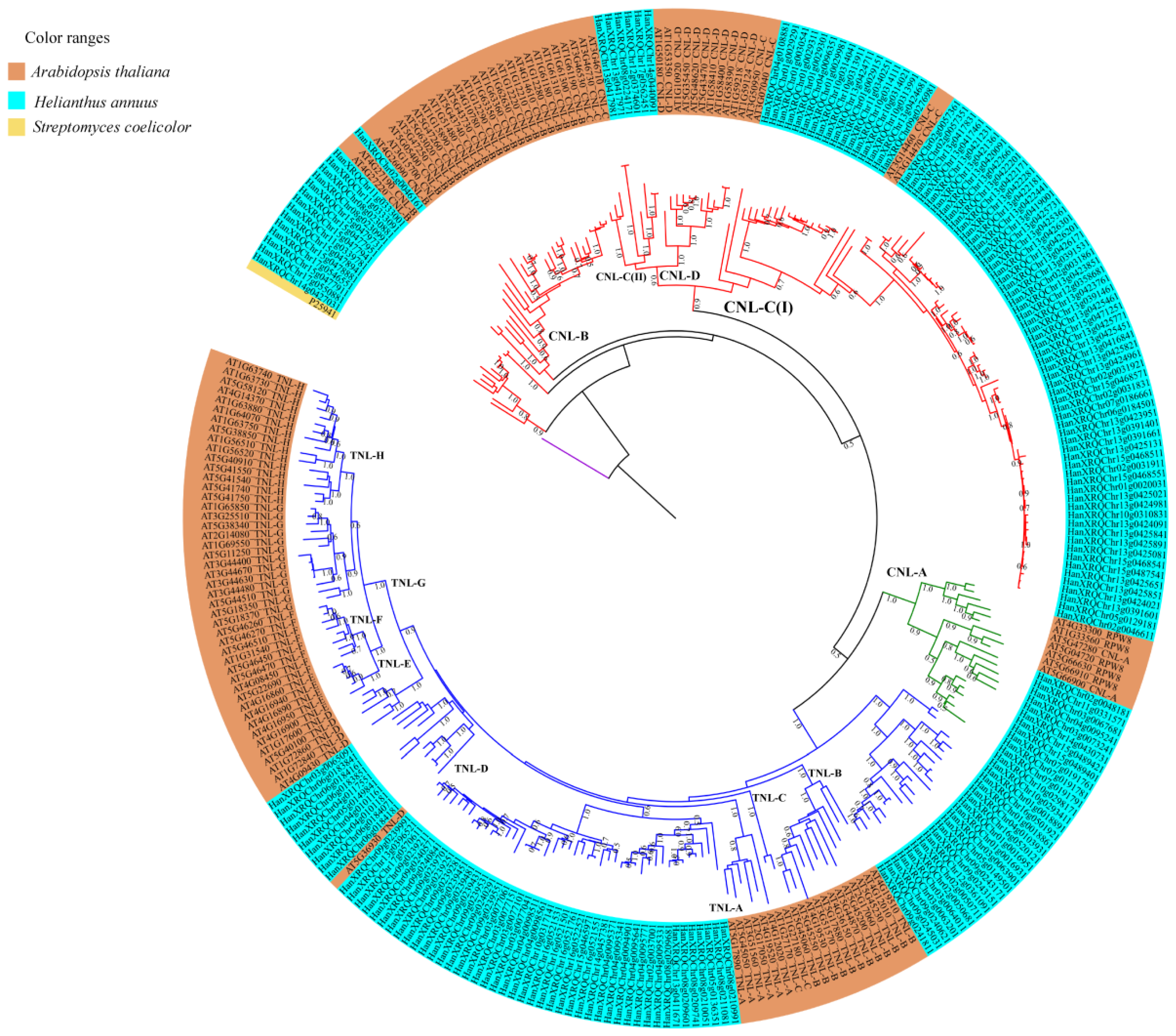
3.3. Phylogenetic and Syntenic Analysis
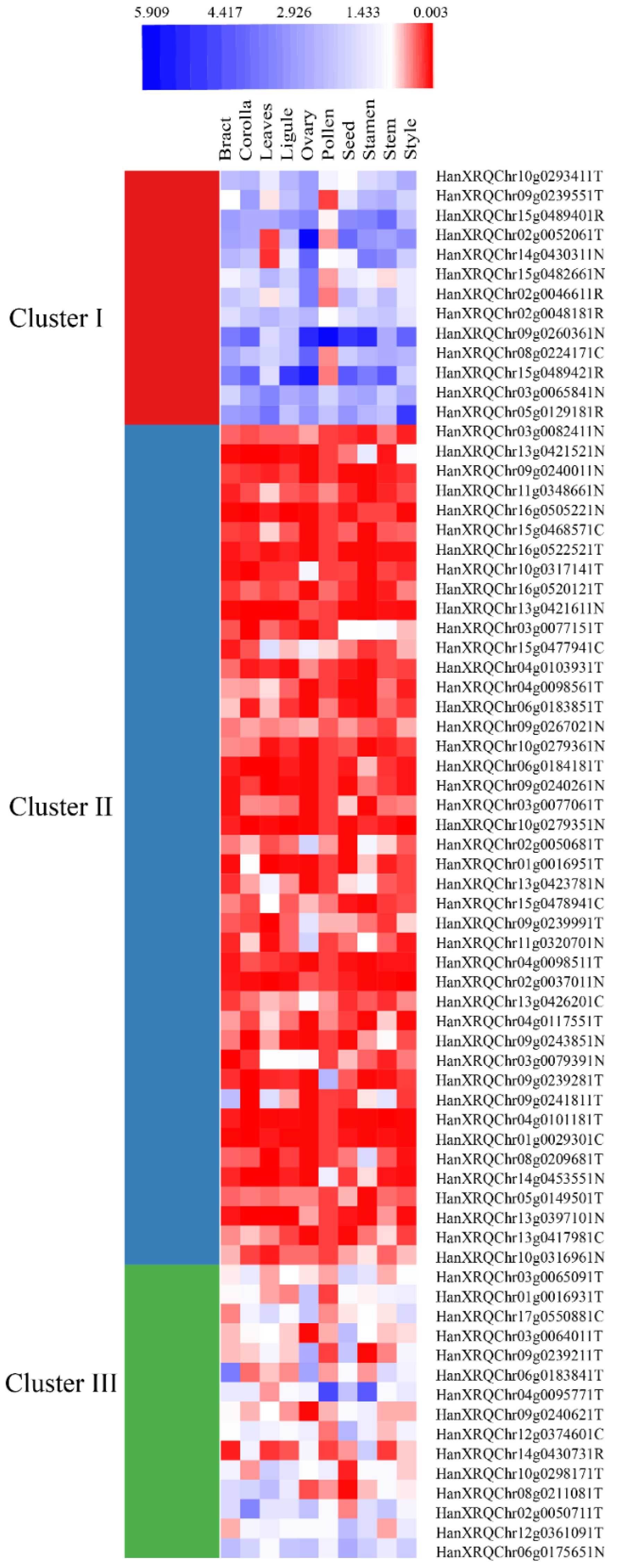
3.4. Homologs and Expression Analysis
4. Discussion
4.1. Diversity of NBS-Encoding Genes
4.2. Gene Location, Clustering, Ka/Ks Values and Structural Variation
4.3. Phylogenetic Relationships, Homology, Synteny and Expression Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Baluška, F.; Mancuso, S. Signaling in Plants, Signaling and Communication in Plants; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar]
- Wang, J.; Pan, C.; Wang, Y.; Ye, L.; Wu, J.; Chen, L.; Zou, T.; Lu, G. Genome-wide identification of MAPK, MAPKK, and MAPKKK gene families and transcriptional profiling analysis during development and stress response in cucumber. BMC Genom. 2015, 16, 386. [Google Scholar] [CrossRef] [PubMed]
- Andersen, E.; Ali, S.; Byamukama, E.; Yen, Y.; Nepal, M. Disease resistance mechanisms in plants. Genes 2018, 9, 339. [Google Scholar] [CrossRef] [PubMed]
- Sekhwal, M.K.; Li, P.; Lam, I.; Wang, X.; Cloutier, S.; You, F.M. Disease resistance gene analogs (RGAs) in Plants. Int. J. Mol. Sci. 2015, 16, 19248–19290. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Tehrim, S.; Zhang, F.; Tong, C.; Huang, J.; Cheng, X.; Dong, C.; Zhou, Y.; Qin, R.; Hua, W.; et al. Genome-wide comparative analysis of NBS-encoding genes between Brassica species and Arabidopsis thaliana. BMC Genom. 2014, 15, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Zipfel, C. Pattern-recognition receptors in plant innate immunity. Curr. Opin. Immunol. 2008, 20, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Andolfo, G.; Ercolano, M.R. Plant innate immunity multicomponent model. Front. Plant Sci. 2015, 6, 987. [Google Scholar] [CrossRef] [PubMed]
- Kuang, H.; Woo, S.-S.; Meyers, B.C.; Nevo, E.; Michelmore, R.W. Multiple genetic processes result in heterogeneous rates of evolution within the major cluster disease resistance genes in lettuce. Plant Cell 2004, 16, 2870–2894. [Google Scholar] [CrossRef] [PubMed]
- Vergne, E.; Grand, X.; Ballini, E.; Chalvon, V.; Saindrenan, P.; Tharreau, D.; Notteghem, J.-L.; Morel, J.-B. Preformed expression of defense is a hallmark of partial resistance to rice blast fungal pathogen Magnaporthe oryzae. BMC Plant Biol. 2010, 10, 206. [Google Scholar] [CrossRef] [PubMed]
- Osuna-Cruz, C.M.; Paytuvi-Gallart, A.; Di Donato, A.; Sundesha, V.; Andolfo, G.; Aiese Cigliano, R.; Sanseverino, W.; Ercolano, M.R. PRGdb 3.0: A comprehensive platform for prediction and analysis of plant disease resistance genes. Nucleic Acids Res. 2017, 46, D1197–D1201. [Google Scholar] [CrossRef] [PubMed]
- Gururani, M.A.; Venkatesh, J.; Upadhyaya, C.P.; Nookaraju, A.; Pandey, S.K.; Park, S.W. Plant disease resistance genes: Current status and future directions. Physiol. Mol. Plant Pathol. 2012, 78, 51–65. [Google Scholar] [CrossRef]
- Shao, Z.-Q.; Xue, J.-Y.; Wu, P.; Zhang, Y.-M.; Wu, Y.; Hang, Y.-Y.; Wang, B.; Chen, J.-Q. Large-scale analyses of angiosperm nucleotide-binding site-leucine-rich repeat (NBS-LRR) genes reveal three anciently diverged classes with distinct evolutionary patterns. Plant Physiol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Meyers, B.C.; Kozik, A.; Griego, A.; Kuang, H.; Michelmore, R.W. Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 2003, 15, 809–834. [Google Scholar] [CrossRef] [PubMed]
- Shao, Z.-Q.; Zhang, Y.-M.; Hang, Y.-Y.; Xue, J.-Y.; Zhou, G.-C.; Wu, P.; Wu, X.-Y.; Wu, X.-Z.; Wang, Q.; Wang, B. Long-term evolution of nucleotide-binding site-leucine-rich repeat genes: Understanding gained from and beyond the legume family. Plant Physiol. 2014, 166, 217–234. [Google Scholar] [CrossRef] [PubMed]
- Zheng, F.; Wu, H.; Zhang, R.; Li, S.; He, W.; Wong, F.-L.; Li, G.; Zhao, S.; Lam, H.-M. Molecular phylogeny and dynamic evolution of disease resistance genes in the legume family. BMC Genom. 2016, 17, 402. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.A.; Yeom, S.I. Plant NB-LRR proteins: Tightly regulated sensors in a complex manner. Brief Funct. Genom. 2015, 14, 233–242. [Google Scholar] [CrossRef] [PubMed]
- Michelmore, R.W.; Christopoulou, M.; Caldwell, K.S. Impacts of resistance gene genetics, function, and evolution on a durable future. Annu. Rev. Phytopathol. 2013, 51, 291–319. [Google Scholar] [CrossRef] [PubMed]
- Die, J.V.; Román, B.; Qi, X.; Rowland, L.J. Genome-scale examination of NBS-encoding genes in blueberry. Sci. Rep. 2018, 8, 3429. [Google Scholar] [CrossRef] [PubMed]
- Sharma, R.; Rawat, V.; Suresh, C. Genome-wide identification and tissue-specific expression analysis of nucleotide binding site-leucine rich repeat gene family in Cicer arietinum (kabuli chickpea). Genom. Data 2017, 14, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.J.; Kim, K.H.; Shim, S.; Yoon, M.Y.; Sun, S.; Kim, M.Y.; Van, K.; Lee, S.-H. Genome-wide mapping of NBS-LRR genes and their association with disease resistance in soybean. BMC Plant Biol. 2012, 12, 139. [Google Scholar] [CrossRef] [PubMed]
- Nepal, M.P.; Benson, B.V. CNL Disease resistance genes in soybean and their evolutionary divergence. Evol. Bioinform. Online 2015, 11, 49–63. [Google Scholar] [CrossRef] [PubMed]
- Nepal, M.P.; Andersen, E.J.; Neupane, S.; Benson, B.V. Comparative genomics of non-TNL Disease resistance genes from six plant species. Genes 2017, 8, 249. [Google Scholar] [CrossRef] [PubMed]
- Neupane, S.; Ma, Q.; Mathew, F.M.; Varenhorst, A.J.; Andersen, E.J.; Nepal, M.P. Evolutionary divergence of TNL disease-resistant proteins in soybean (Glycine max) and common bean (Phaseolus vulgaris). Biochem. Genet. 2018, 56, 397–442. [Google Scholar] [CrossRef] [PubMed]
- Monosi, B.; Wisser, R.J.; Pennill, L.; Hulbert, S.H. Full-genome analysis of resistance gene homologues in rice. Theor. Appl. Genet. 2004, 109, 1434–1477. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Wang, Y.; Chen, J.Q.; Araki, H.; Jing, Z.; Jiang, K.; Shen, J.; Tian, D. Genome-wide identification of NBS genes in japonica rice reveals significant expansion of divergent non-TIR NBS-LRR genes. Mol. Genet. Genom. 2004, 271, 402–415. [Google Scholar]
- Ameline-Torregrosa, C.; Wang, B.B.; O’Bleness, M.S.; Deshpande, S.; Zhu, H.; Roe, B.; Young, N.D.; Cannon, S.B. Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula. Plant Physiol. 2008, 146, 5–21. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Zhang, X.; Yue, J.-X.; Tian, D.; Chen, J.-Q. Recent duplications dominate NBS-encoding gene expansion in two woody species. Mol. Genet. Genom. 2008, 280, 187–198. [Google Scholar] [CrossRef] [PubMed]
- Lozano, R.; Ponce, O.; Ramirez, M.; Mostajo, N.; Orjeda, G. Genome-wide identification and mapping of NBS-encoding resistance genes in Solanum tuberosum group phureja. PLoS ONE 2012, 7, e34775. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.M.; Shao, Z.Q.; Wang, Q.; Hang, Y.Y.; Xue, J.Y.; Wang, B.; Chen, J.Q. Uncovering the dynamic evolution of nucleotide-binding site-leucine-rich repeat (NBS-LRR) genes in Brassicaceae. J. Integr. Plant Biol. 2016, 58, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Andersen, E.J.; Ali, S.; Reese, R.N.; Yen, Y.; Neupane, S.; Nepal, M.P. Diversity and evolution of disease resistance genes in Barley (Hordeum vulgare L.). Evol. Bioinform. Online 2016, 12, 99. [Google Scholar] [CrossRef] [PubMed]
- Andersen, E.J.; Nepal, M.P. Genetic diversity of disease resistance genes in foxtail millet (Setaria italica L.). Plant Gene 2017, 10, 8–16. [Google Scholar] [CrossRef]
- Wan, H.; Yuan, W.; Bo, K.; Shen, J.; Pang, X.; Chen, J. Genome-wide analysis of NBS-encoding disease resistance genes in Cucumis sativus and phylogenetic study of NBS-encoding genes in Cucurbitaceae crops. BMC Genom. 2013, 14, 109. [Google Scholar] [CrossRef] [PubMed]
- Xiang, L.; Liu, J.; Wu, C.; Deng, Y.; Cai, C.; Zhang, X.; Cai, Y. Genome-wide comparative analysis of NBS-encoding genes in four Gossypium species. BMC Genom. 2017, 18, 292. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Quan, X.; Jia, G.; Xiao, J.; Cloutier, S.; You, F.M. RGAugury: A pipeline for genome-wide prediction of resistance gene analogs (RGAs) in plants. BMC Genom. 2016, 17, 852. [Google Scholar] [CrossRef] [PubMed]
- Kane, N.C.; Rieseberg, L.H. Selective sweeps reveal candidate genes for adaptation to drought and salt tolerance in common sunflower, Helianthus annuus. Genetics 2007, 175, 1823–1834. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Martínez, J.; Melero-Vara, J.; Muñoz-Ruz, J.; Ruso, J.; Domínguez, J. Selection of wild and cultivated sunflower for resistance to a new broomrape race that overcomes resistance of the Or5 gene. Crop Sci. 2000, 40, 550–555. [Google Scholar] [CrossRef]
- Seiler, G. Wild annual Helianthus anomalus and H. deserticola for improving oil content and quality in sunflower. Ind. Crops Prod. 2007, 25, 95–100. [Google Scholar] [CrossRef]
- Markell, S.; Harveson, R.; Block, C.; Gulya, T. Sunflower Disease Diagnostic Series; PP1727-19; North Dakota State University Extension Service Publisher: Fargo, ND, USA, 2005. [Google Scholar]
- Plocik, A.; Layden, J.; Kesseli, R. Comparative analysis of NBS domain sequences of NBS-LRR disease resistance genes from sunflower, lettuce, and chicory. Mol. Phylogenet. Evol. 2004, 31, 153–163. [Google Scholar] [CrossRef]
- Hewezi, T.; Mouzeyar, S.; Thion, L.; Rickauer, M.; Alibert, G.; Nicolas, P.; Kallerhoff, J. Antisense expression of a NBS-LRR sequence in sunflower (Helianthus annuus L.) and tobacco (Nicotiana tabacum L.): Evidence for a dual role in plant development and fungal resistance. Transgenic Res. 2006, 15, 165–180. [Google Scholar] [CrossRef] [PubMed]
- Radwan, O.; Gandhi, S.; Heesacker, A.; Whitaker, B.; Taylor, C.; Plocik, A.; Kesseli, R.; Kozik, A.; Michelmore, R.W.; Knapp, S.J. Genetic diversity and genomic distribution of homologs encoding NBS-LRR disease resistance proteins in sunflower. Mol. Genet. Genom. 2008, 280, 111–125. [Google Scholar] [CrossRef] [PubMed]
- Radwan, O.; Mouzeyar, S.; Nicolas, P.; Bouzidi, M. Induction of a sunflower CC-NBS-LRR resistance gene analogue during incompatible interaction with Plasmopara halstedii. J. Exp. Bot. 2004, 56, 567–575. [Google Scholar] [CrossRef] [PubMed]
- Badouin, H.; Gouzy, J.; Grassa, C.J.; Murat, F.; Staton, S.E.; Cottret, L.; Lelandais-Brière, C.; Owens, G.L.; Carrère, S.; Mayjonade, B. The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 2017, 546, 148. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Clements, J.; Arndt, W.; Miller, B.L.; Wheeler, T.J.; Schreiber, F.; Bateman, A.; Eddy, S.R. HMMER web server: 2015 update. Nucleic Acids Res. 2015, 43, W30–W38. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J. Pfam: The protein families database. Nucleic Acids Res. 2013, 42, D222–D231. [Google Scholar] [CrossRef] [PubMed]
- Delorenzi, M.; Speed, T. An HMM model for coiled-coil domains and a comparison with PSSM-based predictions. Bioinformatics 2002, 18, 617–625. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in bipolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar] [PubMed]
- Emanuelsson, O.; Brunak, S.; von Heijne, G.; Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nat. Protoc. 2007, 2, 953–971. [Google Scholar] [CrossRef] [PubMed]
- Ba, A.N.N.; Pogoutse, A.; Provart, N.; Moses, A.M. NLStradamus: A simple Hidden Markov Model for nuclear localization signal prediction. BMC Bioinform. 2009, 10, 202. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016, 44, W242–W245. [Google Scholar] [CrossRef] [PubMed]
- Jupe, F.; Pritchard, L.; Etherington, G.J.; MacKenzie, K.; Cock, P.J.; Wright, F.; Sharma, S.K.; Bolser, D.; Bryan, G.J.; Jones, J.D. Identification and localisation of the NB-LRR gene family within the potato genome. BMC Genom. 2012, 13, 75. [Google Scholar] [CrossRef] [PubMed]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef] [PubMed]
- Soderlund, C.; Bomhoff, M.; Nelson, W.M. SyMAP v3. 4: A turnkey synteny system with application to plant genomes. Nucleic Acids Res. 2011, 39, e68. [Google Scholar] [CrossRef] [PubMed]
- Howe, E.A.; Sinha, R.; Schlauch, D.; Quackenbush, J. RNA-Seq analysis in MeV. Bioinformatics 2011, 27, 3209–3210. [Google Scholar] [CrossRef] [PubMed]
- Gedil, M.A.; Slabaugh, M.B.; Berry, S.; Johnson, R.; Michelmore, R.; Miller, J.; Gulya, T.; Knapp, S.J. Candidate disease resistance genes in sunflower cloned using conserved nucleotide-binding site motifs: Genetic mapping and linkage to the downy mildew resistance gene Pl1. Genome 2001, 44, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Ming, R.; Hou, S.; Feng, Y.; Yu, Q.; Dionne-Laporte, A.; Saw, J.H.; Senin, P.; Wang, W.; Ly, B.V.; Lewis, K.L. The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus). Nature 2008, 452, 991. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Zhu, J.; Wang, L.; Wang, S. Genome-wide association study identifies NBS-LRR-encoding genes related with anthracnose and common bacterial blight in the common Bean. Front. Plant Sci. 2017, 8, 1398. [Google Scholar] [CrossRef] [PubMed]
- Lozano, R.; Hamblin, M.T.; Prochnik, S.; Jannink, J.L. Identification and distribution of the NBS-LRR gene family in the Cassava genome. BMC Genom. 2015, 16, 360. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.-L.; Fitz, J.; Schneeberger, K.; Ossowski, S.; Cao, J.; Weigel, D. Genome-wide comparison of nucleotide-binding site-leucine-rich repeat-encoding genes in Arabidopsis. Plant Physiol. 2011, 157, 757–769. [Google Scholar] [CrossRef] [PubMed]
- Mun, J.-H.; Yu, H.-J.; Park, S.; Park, B.-S. Genome-wide identification of NBS-encoding resistance genes in Brassica rapa. Mol. Genet. Genom. 2009, 282, 617–631. [Google Scholar] [CrossRef] [PubMed]
- Christie, N.; Tobias, P.A.; Naidoo, S.; Külheim, C. The Eucalyptus grandis NBS-LRR gene family: Physical clustering and expression hotspots. Front. Plant Sci. 2016, 6, 1238. [Google Scholar] [CrossRef] [PubMed]
- Hao, W.; Collier, S.M.; Moffett, P.; Chai, J. Structural basis for the interaction between the potato virus X resistance protein (Rx) and its cofactor Ran GTPase-activating protein 2 (RanGAP2). J. Biol. Chem. 2013, 288, 35868–35876. [Google Scholar] [CrossRef] [PubMed]
- Kohler, A.; Rinaldi, C.; Duplessis, S.; Baucher, M.; Geelen, D.; Duchaussoy, F.; Meyers, B.C.; Boerjan, W.; Martin, F. Genome-wide identification of NBS resistance genes in Populus trichocarpa. Plant Mol. Biol. 2008, 66, 619–636. [Google Scholar] [CrossRef] [PubMed]
- Schumann, N.; Navarro-Quezada, A.; Ullrich, K.; Kuhl, C.; Quint, M. Molecular evolution and selection patterns of plant F-box proteins with C-terminal kelch repeats. Plant Physiol. 2011, 155, 835–850. [Google Scholar] [CrossRef] [PubMed]
- Afzal, A.J.; Wood, A.J.; Lightfoot, D.A. Plant receptor-like serine threonine kinases: Roles in signaling and plant defense. Mol. Plant-Microbe Interact. 2008, 21, 507–517. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Gómez, L.; Boller, T. Flagellin perception: A paradigm for innate immunity. Trends Plant Sci. 2002, 7, 251–256. [Google Scholar] [CrossRef]
- Zipfel, C.; Kunze, G.; Chinchilla, D.; Caniard, A.; Jones, J.D.; Boller, T.; Felix, G. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 2006, 125, 749–760. [Google Scholar] [CrossRef] [PubMed]
- Macho, A.P.; Zipfel, C. Plant PRRs and the activation of innate immune signaling. Mol. Cell 2014, 54, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Song, W.-Y.; Wang, G.-L.; Chen, L.-L.; Kim, H.-S.; Pi, L.-Y.; Holsten, T.; Gardner, J.; Wang, B.; Zhai, W.-X.; Zhu, L.-H. A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 1995, 270, 1804–1806. [Google Scholar] [CrossRef] [PubMed]
- Godiard, L.; Sauviac, L.; Torii, K.U.; Grenon, O.; Mangin, B.; Grimsley, N.H.; Marco, Y. ERECTA, an LRR receptor-like kinase protein controlling development pleiotropically affects resistance to bacterial wilt. Plant J. 2003, 36, 353–365. [Google Scholar] [CrossRef] [PubMed]
- Jeong, S.; Trotochaud, A.E.; Clark, S.E. The Arabidopsis CLAVATA2 gene encodes a receptor-like protein required for the stability of the CLAVATA1 receptor-like kinase. Plant Cell 1999, 11, 1925–1933. [Google Scholar] [CrossRef] [PubMed]
- Kruijt, M.; Kip, D.J.; Joosten, M.H.; Brandwagt, B.F.; de Wit, P.J. The Cf-4 and Cf-9 resistance genes against Cladosporium fulvum are conserved in wild tomato species. Mol. Plant-Microbe Interact. 2005, 18, 1011–1021. [Google Scholar] [CrossRef] [PubMed]
- Friedman, A.R.; Baker, B.J. The evolution of resistance genes in multi-protein plant resistance systems. Curr. Opin. Genet. Dev. 2007, 17, 493–499. [Google Scholar] [CrossRef] [PubMed]
- Qian, L.-H.; Zhou, G.-C.; Sun, X.-Q.; Lei, Z.; Zhang, Y.-M.; Xue, J.-Y.; Hang, Y.-Y. Distinct patterns of gene gain and loss: Diverse evolutionary modes of NBS-encoding genes in three Solanaceae crop species. G3 Genes Genomes Genet. 2017, 7, 1577–1585. [Google Scholar] [CrossRef] [PubMed]
- Leister, D. Tandem and segmental gene duplication and recombination in the evolution of plant disease resistance genes. Trends Genet. 2004, 20, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, D.; Ganfornina, M.D.; Gutiérrez, G.; Marín, A. Exon-intron structure and evolution of the Lipocalin gene family. Mol. Biol. Evol. 2003, 20, 775–783. [Google Scholar] [CrossRef] [PubMed]
- Zeng, L.; Zhang, Q.; Sun, R.; Kong, H.; Zhang, N.; Ma, H. Resolution of deep angiosperm phylogeny using conserved nuclear genes and estimates of early divergence times. Nat. Commun. 2014, 5, 4956. [Google Scholar] [CrossRef] [PubMed]
- Bonardi, V.; Tang, S.; Stallmann, A.; Roberts, M.; Cherkis, K.; Dangl, J.L. Expanded functions for a family of plant intracellular immune receptors beyond specific recognition of pathogen effectors. Proc. Natl. Acad. Sci. USA 2011, 108, 16463–16468. [Google Scholar] [CrossRef] [PubMed]
- Peart, J.R.; Mestre, P.; Lu, R.; Malcuit, I.; Baulcombe, D.C. NRG1, a CC-NB-LRR protein, together with N, a TIR-NB-LRR protein, mediates resistance against tobacco mosaic virus. Curr. Biol. 2005, 15, 968–973. [Google Scholar] [CrossRef] [PubMed]
- Collier, S.M.; Hamel, L.-P.; Moffett, P. Cell death mediated by the N-terminal domains of a unique and highly conserved class of NB-LRR protein. Mol. Plant-Microbe Interact. 2011, 24, 918–931. [Google Scholar] [CrossRef] [PubMed]
- Paal, J.; Henselewski, H.; Muth, J.; Meksem, K.; Menéndez, C.M.; Salamini, F.; Ballvora, A.; Gebhardt, C. Molecular cloning of the potato Gro1-4 gene conferring resistance to pathotype Ro1 of the root cyst nematode Globodera rostochiensis, based on a candidate gene approach. Plant J. 2004, 38, 285–297. [Google Scholar] [CrossRef] [PubMed]
- Vidal, S.; Cabrera, H.; Andersson, R.A.; Fredriksson, A.; Valkonen, J.P. Potato gene Y-1 is an N gene homolog that confers cell death upon infection with potato virus Y. Mol. Plant-Microbe Interact. 2002, 15, 717–727. [Google Scholar] [CrossRef] [PubMed]
- Levy, M.; Edelbaum, O.; Sela, I. Tobacco mosaic virus regulates the expression of its own resistance gene N. Plant Physiol. 2004, 135, 2392–2397. [Google Scholar] [CrossRef] [PubMed]
- Katagiri, F.; Thilmony, R.; He, S.Y. The Arabidopsis thaliana-Pseudomonas syringae interaction. Arabidopsis Book 2002, 1, e0039. [Google Scholar] [CrossRef] [PubMed]
- Pitrat, M.; Maestro, C.; Ferriere, C.; Ricard, M.; Alvarez, J. Resistance to Aphis gossypii in Spanish melon (Cucumis melo). Cucurbit. Genet. Coop. Rep. 1988, 11, 50–51. [Google Scholar]
- Liu, X.; Lin, F.; Wang, L.; Pan, Q. The in silico map-based cloning of Pi36, a rice coiled-coil–nucleotide-binding site–leucine-rich repeat gene that confers race-specific resistance to the blast fungus. Genetics 2007, 176, 2541–2549. [Google Scholar] [CrossRef] [PubMed]
- Sallam, A.H.; Tyagi, P.; Brown-Guedira, G.; Muehlbauer, G.J.; Hulse, A.; Steffenson, B.J. Genome-wide association mapping of stem rust resistance in Hordeum vulgare subsp. spontaneum. G3 Genes Genomes Genet. 2017, 7, 3491–3507. [Google Scholar] [CrossRef] [PubMed]
- Ron, M.; Avni, A. The receptor for the fungal elicitor ethylene-inducing xylanase is a member of a resistance-like gene family in tomato. Plant Cell 2004, 16, 1604–1615. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Liu, J.; Chang, C.; Zhang, L.; Maekawa, T.; Wang, Q.; Xiao, W.; Liu, Y.; Chai, J.; Takken, F.L. Structure-function analysis of barley NLR immune receptor MLA10 reveals its cell compartment specific activity in cell death and disease resistance. PLoS Pathog. 2012, 8, e1002752. [Google Scholar] [CrossRef] [PubMed]
- Casey, L.W.; Lavrencic, P.; Bentham, A.R.; Cesari, S.; Ericsson, D.J.; Croll, T.; Turk, D.; Anderson, P.A.; Mark, A.E.; Dodds, P.N. The CC domain structure from the wheat stem rust resistance protein Sr33 challenges paradigms for dimerization in plant NLR proteins. Proc. Natl. Acad. Sci. USA 2016, 113, 12856–12861. [Google Scholar] [CrossRef] [PubMed]
- Hammond-Kosack, K.E.; Jones, J.D. Plant disease resistance genes. Annu. Rev. Plant Biol. 1997, 48, 575–607. [Google Scholar] [CrossRef] [PubMed]
- Frazier, T.P.; Palmer, N.A.; Xie, F.; Tobias, C.M.; Donze-Reiner, T.J.; Bombarely, A.; Childs, K.L.; Shu, S.; Jenkins, J.W.; Schmutz, J. Identification, characterization, and gene expression analysis of nucleotide binding site (NB)-type resistance gene homologues in switchgrass. BMC Genom. 2016, 17, 892. [Google Scholar] [CrossRef] [PubMed]

| Protein Letter Code | Number of Proteins | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ha * | At a | Gm b,c | Mt a | Bo a | Br a | Tc a | Pt a | Vv a | Ca d | Cs e | Pv f,c | Lj f | Cc f | Gs f | Ga g | |
| CNL | 90 | 17 | 95 | 152 | 6 | 19 | 82 | 120 | 203 | 19 | 17 | 31 | 11 | 37 | 47 | 80 |
| CN | 5 | 8 | - | 25 | 5 | 15 | 46 | 14 | 26 | 33 | 1 | 40 | 26 | 41 | 62 | 44 |
| CNNL | 4 | - | 5 | - | - | 2 | - | - | - | 1 | - | - | - | - | - | - |
| CNN | 1 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| RNL | 10 | 2 | 6 | - | 1 | 4 | - | - | - | 2 | 2 | - | - | - | - | 3 |
| RN | 1 | 3 | - | - | 2 | 1 | - | - | - | 2 | - | - | - | - | - | - |
| RCNL | 2 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| TNL | 52 | 79 | 126 | 118 | 40 | 93 | 8 | 78 | 97 | 6 | 11 | 81 | 16 | 47 | 49 | 5 |
| TN | 21 | 17 | 22 | 38 | 29 | 23 | 4 | 10 | 14 | 7 | 2 | 11 | 53 | 36 | 76 | 2 |
| TNNL | 0 | 1 | - | - | 1 | 4 | - | - | - | - | - | - | - | - | - | - |
| TTNL | 1 | - | - | - | - | - | - | - | - | 1 | - | - | - | - | - | - |
| TNLTNL | 1 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| CTNL | 1 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| CTN | 1 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| N | 29 | 26 | 4 | 328 | 53 | 29 | 53 | 62 | 36 | 14 | 1 | 59 | 82 | 136 | 213 | 59 |
| NL | 125 | 20 | 73 | - | 24 | 27 | 104 | 132 | 159 | 12 | 23 | 20 | 18 | 56 | 58 | 53 |
| NN | 2 | - | - | - | 3 | 2 | - | - | - | 1 | - | - | - | - | - | - |
| NNL | 6 | - | - | - | - | 3 | - | - | - | - | - | - | - | - | - | - |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Neupane, S.; Andersen, E.J.; Neupane, A.; Nepal, M.P. Genome-Wide Identification of NBS-Encoding Resistance Genes in Sunflower (Helianthus annuus L.). Genes 2018, 9, 384. https://doi.org/10.3390/genes9080384
Neupane S, Andersen EJ, Neupane A, Nepal MP. Genome-Wide Identification of NBS-Encoding Resistance Genes in Sunflower (Helianthus annuus L.). Genes. 2018; 9(8):384. https://doi.org/10.3390/genes9080384
Chicago/Turabian StyleNeupane, Surendra, Ethan J. Andersen, Achal Neupane, and Madhav P. Nepal. 2018. "Genome-Wide Identification of NBS-Encoding Resistance Genes in Sunflower (Helianthus annuus L.)" Genes 9, no. 8: 384. https://doi.org/10.3390/genes9080384
APA StyleNeupane, S., Andersen, E. J., Neupane, A., & Nepal, M. P. (2018). Genome-Wide Identification of NBS-Encoding Resistance Genes in Sunflower (Helianthus annuus L.). Genes, 9(8), 384. https://doi.org/10.3390/genes9080384






