The 5′-3′ Exoribonuclease XRN4 Regulates Auxin Response via the Degradation of Auxin Receptor Transcripts
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material, Growth Conditions, and Treatments
2.2. Histochemical GUS Assays
2.3. RNA Decay Assay
2.4. RNA Preparation and RNA Analysis
3. Results
3.1. XRN4 Defective Plants Exhibit Pleiotropic Developmental Defects
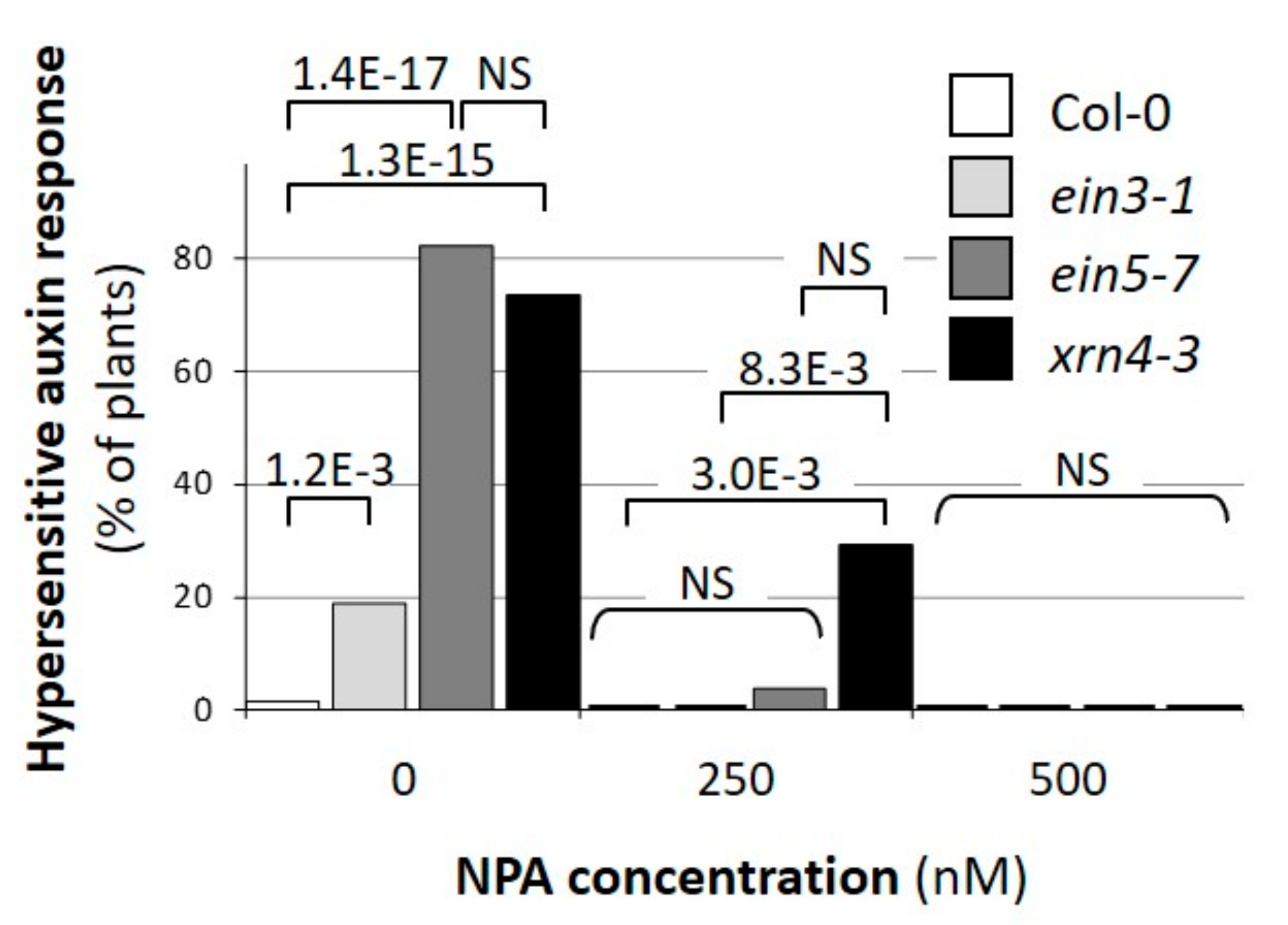
3.2. Defects in XRN4 Result in Hypersensitive Physiological Responses to Auxin
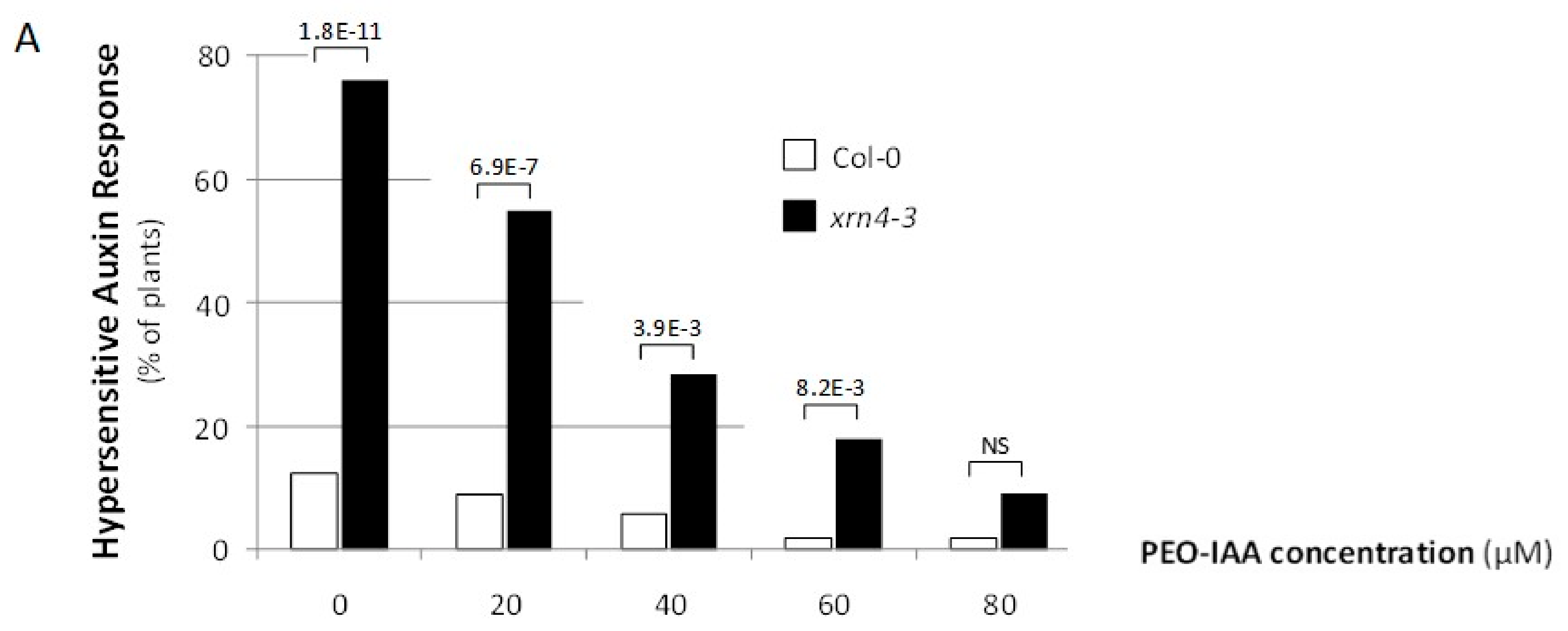
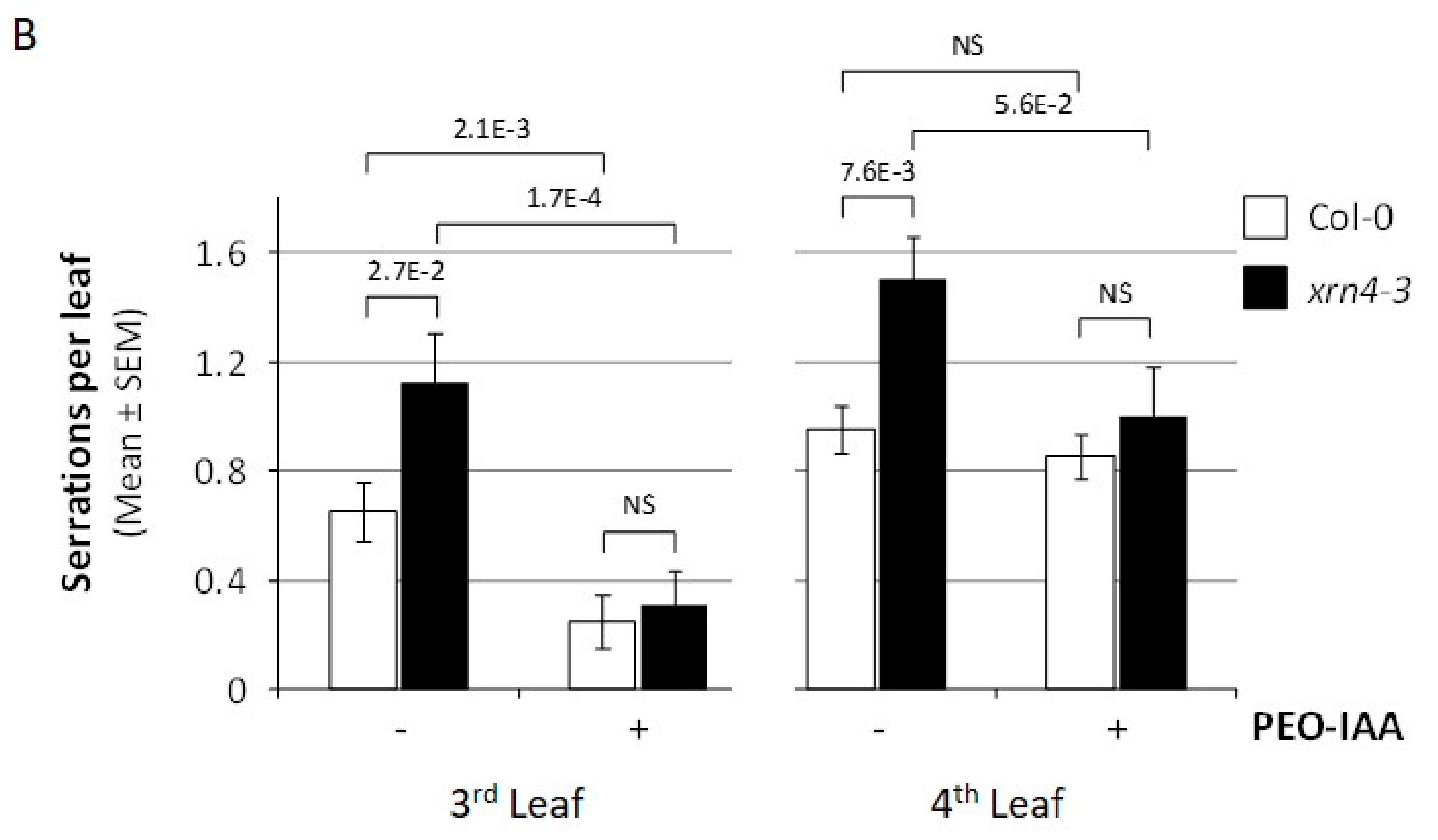
3.3. Chemical Inhibition of TAAR Function Suppresses Specific xrn4 Phenotypes
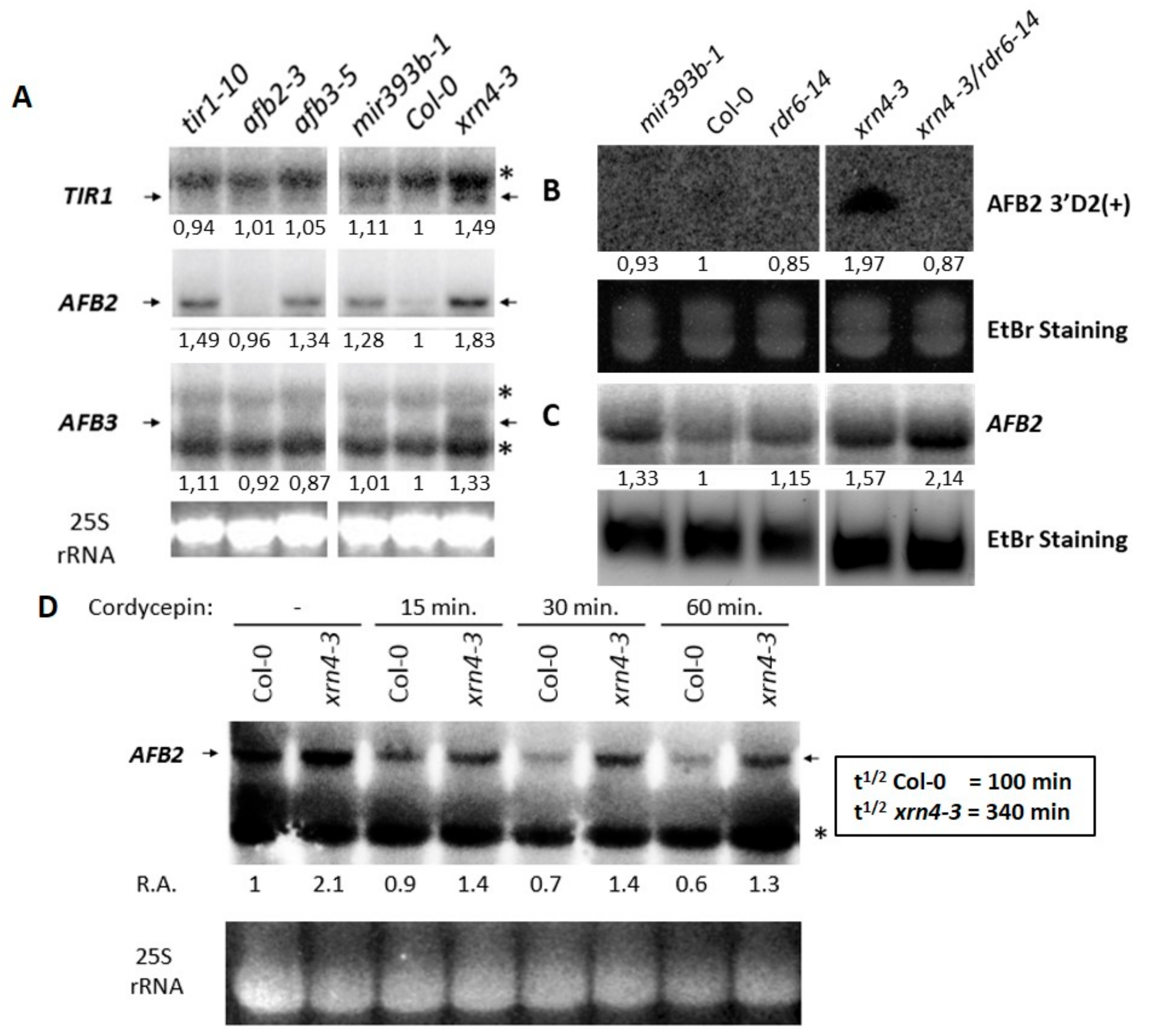
3.4. XRN4 Regulates the Turn-Over of TAAR Transcripts
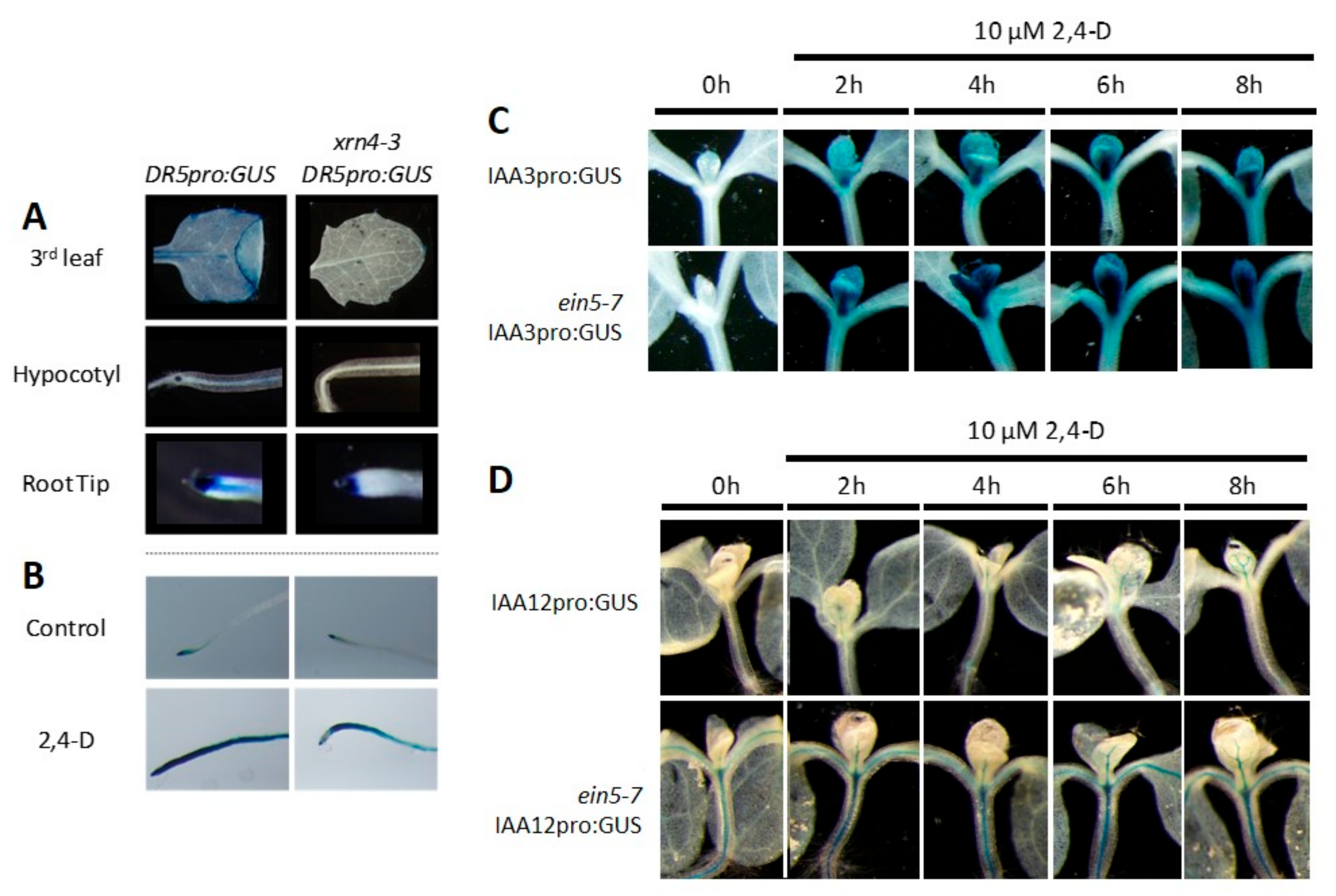
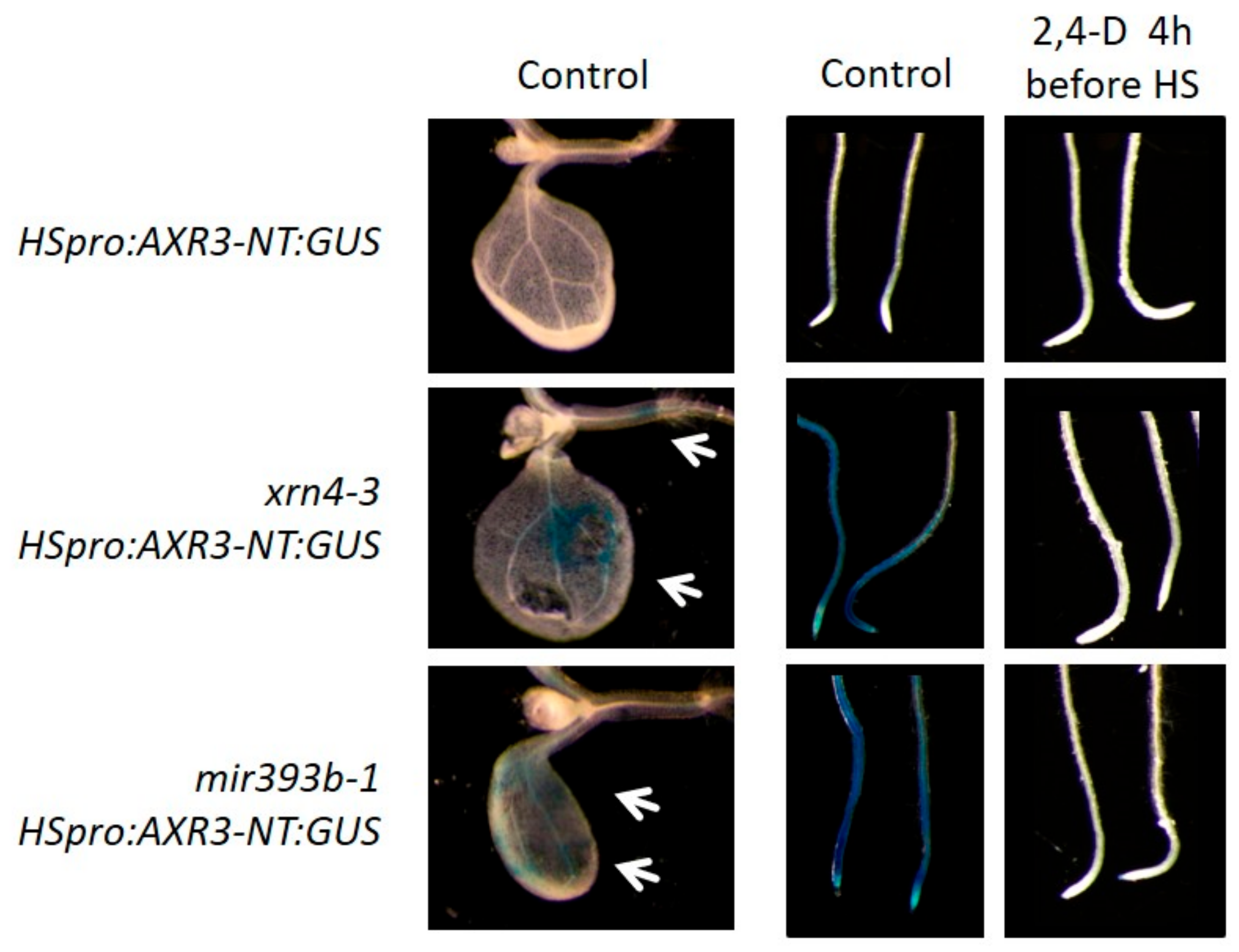
3.5. xrn4 Plants Exhibit Hypersensitive Auxin Signaling Responses
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mockaitis, K.; Estelle, M. Auxin receptors and plant development: A new signaling paradigm. Annu. Rev. Cell Dev. Biol. 2008, 24, 55–80. [Google Scholar] [CrossRef] [PubMed]
- Vanneste, S.; Friml, J. Auxin: A trigger for change in plant development. Cell 2009, 136, 1005–1016. [Google Scholar] [CrossRef] [PubMed]
- Dharmasiri, N.; Dharmasiri, S.; Estelle, M. The F-box protein TIR1 is an auxin receptor. Nature 2005, 435, 441–445. [Google Scholar] [CrossRef] [PubMed]
- Dharmasiri, N.; Dharmasiri, S.; Weijers, D.; Lechner, E.; Yamada, M.; Hobbie, L.; Ehrismann, J.S.; Jürgens, G.; Estelle, M. Plant development is regulated by a family of auxin receptor F box proteins. Dev. Cell 2005, 9, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Kepinski, S.; Leyser, O. The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 2005, 435, 446–451. [Google Scholar] [CrossRef] [PubMed]
- Gray, W.M.; Kepinski, S.; Rouse, D.; Leyser, O.; Estelle, M. Auxin regulates SCF(TIR1)-dependent degradation of AUX/IAA proteins. Nature 2001, 414, 271–276. [Google Scholar] [CrossRef] [PubMed]
- Lechner, E.; Achard, P.; Vansiri, A.; Potuschak, T.; Genschik, P. F-box proteins everywhere. Curr. Opin. Plant Biol. 2006, 9, 631–638. [Google Scholar] [CrossRef]
- Calderón Villalobos, L.I.; Lee, S.; De Oliveira, C.; Ivetac, A.; Brandt, W.; Armitage, L.; Sheard, L.B.; Tan, X.; Parry, G.; Mao, H.; et al. A combinatorial TIR1/AFB-Aux/IAA co-receptor system for differential sensing of auxin. Nat. Chem. Biol. 2012, 8, 477–485. [Google Scholar] [CrossRef]
- Ulmasov, T.; Hagen Gand Guilfoyle, T.J. Activation and repression of transcription by auxin-response factors. Proc. Natl. Acad. Sci. USA 1999, 96, 5844–5849. [Google Scholar] [CrossRef]
- Ljung, K. Auxin metabolism and homeostasis during plant development. Development 2013, 140, 943–950. [Google Scholar] [CrossRef]
- Rosquete, M.R.; Barbez, E.; Kleine-Vehn, J. Cellular auxin homeostasis: Gatekeeping is housekeeping. Mol. Plant 2012, 5, 772–786. [Google Scholar] [CrossRef] [PubMed]
- Navarro, L.; Dunoyer, P.; Jay, F.; Arnold, B.; Dharmasiri, N.; Estelle, M.; Voinnet, O.; Jones, J.D. A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 2006, 312, 436–439. [Google Scholar] [CrossRef] [PubMed]
- Vidal, E.A.; Araus, V.; Lu, C.; Parry, G.; Green, P.J.; Coruzzi, G.M.; Gutiérrez, R.A. Nitrate-responsive miR393/AFB3 regulatory module controls root system architecture in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2010, 107, 4477–4482. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.H.; Bao, M.L.; Sun, Y.Z.; Yang, Y.J.; Xu, X.H.; Wang, J.H.; Han, N.; Bian, H.W.; Zhu, M.Y. Regulation of auxin response by miR393-targeted transport inhibitor response protein 1 is involved in normal development in Arabidopsis. Plant Mol. Biol. 2011, 77, 619–629. [Google Scholar] [CrossRef] [PubMed]
- Si-Ammour, A.; Windels, D.; Arn-Bouldoires, E.; Kutter, C.; Ailhas, J.; Meins, F., Jr.; Vazquez, F. miR393 and secondary siRNAs regulate expression of the TIR1/AFB2 auxin receptor clade and auxin-related development of Arabidopsis leaves. Plant Physiol. 2011, 157, 683–691. [Google Scholar] [CrossRef] [PubMed]
- Windels, D.; Bielewicz, D.; Ebneter, M.; Jarmolowski, A.; Szweykowska-Kulinska, Z.; Vazquez, F. miR393 is required for production of proper auxin signalling output. PLoS ONE 2014, 9, e95972. [Google Scholar] [CrossRef] [PubMed]
- Iglesias, M.J.; Terrile, M.C.; Windels, D.; Lombardo, M.C.; Bartoli, C.G.; Vazquez, F.; Estelle, M.; Casalongué, C. miR393 regulation of auxin signaling and redox-related components during acclimation to salinity in Arabidopsis. PLoS ONE 2014, 9, e107678. [Google Scholar] [CrossRef]
- Windels, D.; Vazquez, F. miR393: Integrator of environmental cues in auxin signaling? Plant Signal. Behav. 2011, 6, 1672–1675. [Google Scholar] [CrossRef]
- Belostotsky, D.A. State of decay: An update on plant mRNA turnover. Curr. Top. Microbiol. Immunol. 2008, 326, 179–199. [Google Scholar]
- Houseley, J.; Tollervey, D. The many pathways of RNA degradation. Cell 2009, 136, 763–776. [Google Scholar] [CrossRef]
- Zakrzewska-Placzek, M.; Souret, F.F.; Sobczyk, G.J.; Green, P.J.; Kufel, J. Arabidopsis thaliana XRN2 is required for primary cleavage in the pre-ribosomal RNA. Nucleic Acids Res. 2010, 38, 4487–4502. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, Y.; Schmitz, R.J.; Nery, J.R.; Schultz, M.D.; Okubo-Kurihara, E.; Morosawa, T.; Tanaka, M.; Toyoda, T.; Seki, M.; Ecker, J.R. Surveillance of 3′ noncoding transcripts requires FIERY1 and XRN3 in Arabidopsis. G3 (Bethesda) 2012, 2, 487–498. [Google Scholar] [CrossRef] [PubMed]
- Kastenmayer, J.P.; Green, P.J. Novel features of the XRN-family in Arabidopsis: Evidence that AtXRN4, one of several orthologs of nuclear Xrn2p/Rat1p, functions in the cytoplasm. Proc. Natl. Acad. Sci. USA 2000, 97, 13985–13990. [Google Scholar] [CrossRef] [PubMed]
- Kastenmayer, J.P.; Johnson, M.A.; Green, P.J. Analysis of XRN orthologs by complementation of yeast mutants and localization of XRN-GFP fusion proteins. Methods Enzymol. 2001, 342, 269–282. [Google Scholar] [PubMed]
- Rymarquis, L.A.; Souret, F.F.; Green, P.J. Evidence that XRN4, an Arabidopsis homolog of exoribonuclease XRN1, preferentially impacts transcripts with certain sequences or in particular functional categories. RNA 2011, 17, 501–511. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, V.K.; Jones, C.I.; Newbury, S.F.; Green, P.J. XRN 5′→3′ exoribonucleases: Structure, mechanisms and functions. Biochim. Biophys. Acta 2013, 1829, 590–603. [Google Scholar] [CrossRef] [PubMed]
- Souret, F.F.; Kastenmayer, J.P.; Green, P.J. AtXRN4 degrades mRNA in Arabidopsis and its substrates include selected miRNA targets. Mol Cell. 2004, 15, 173–183. [Google Scholar] [CrossRef]
- Wang, K.L.; Li, H.; Ecker, J.R. Ethylene biosynthesis and signaling networks. Plant Cell 2002, 14, S131–S151. [Google Scholar] [CrossRef]
- Olmedo, G.; Guo, H.; Gregory, B.D.; Nourizadeh, S.D.; Aguilar-Henonin, L.; Li, H.; An, F.; Guzman, P.; Ecker, J.R. ETHYLENE-INSENSITIVE5 encodes a 5′→3′ exoribonuclease required for regulation of the EIN3-targeting F-box proteins EBF1/2. Proc. Natl. Acad. Sci. USA 2006, 103, 13286–13293. [Google Scholar] [CrossRef]
- Potuschak, T.; Vansiri, A.; Binder, B.M.; Lechner, E.; Vierstra, R.D.; Genschik, P. The exoribonuclease XRN4 is a component of the ethylene response pathway in Arabidopsis. Plant Cell 2006, 18, 3047–3057. [Google Scholar] [CrossRef]
- Vandenbussche, F.; Vaseva, I.; Vissenberg, K.; Van Der Straeten, D. Ethylene in vegetative development: A tale with a riddle. New Phytol. 2012, 194, 895–909. [Google Scholar] [CrossRef] [PubMed]
- Merret, R.; Descombin, J.; Juan, Y.T.; Favory, J.J.; Carpentier, M.C.; Chaparro, C.; Charng, Y.Y.; Deragon, J.M.; Bousquet-Antonelli, C. XRN4 and LARP1 are required for a heat-triggered mRNA decay pathway involved in plant acclimation and survival during thermal stress. Cell Rep. 2013, 5, 1279–1293. [Google Scholar] [CrossRef] [PubMed]
- Vazquez, F.; Blevins, T.; Ailhas, J.; Boller, T.; Meins, F., Jr. Evolution of Arabidopsis MIR genes generates novel microRNA classes. Nucleic Acids Res. 2008, 36, 6429–6438. [Google Scholar] [CrossRef]
- Gutierrez, R.A.; Ewing, R.M.; Cherry, J.M.; Green, P.J. Identification of unstable transcripts in Arabidopsis by cDNA microarray analysis: Rapid decay is associated with a group of touch- and specific clock-controlled genes. Proc. Natl. Acad. Sci. USA 2002, 99, 11513–11518. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, K.; Neve, J.; Hirose, M.; Kuboki, A.; Shimada, Y.; Kepinski, S.; Nozaki, H. Rational design of an auxin antagonist of the SCF(TIR1) auxin receptor complex. ACS Chem. Biol. 2012, 7, 590–598. [Google Scholar] [CrossRef] [PubMed]
- Gregory, B.D.; O’Malley, R.C.; Lister, R.; Urich, M.A.; Tonti-Filippini, J.; Chen, H.; Millar, A.H.; Ecker, J.R. A link between RNA metabolism and silencing affecting Arabidopsis development. Dev Cell 2008, 14, 854–866. [Google Scholar] [CrossRef] [PubMed]
- Martínez de Alba, A.E.; Moreno, A.B.; Gabriel, M.; Mallory, A.C.; Christ, A.; Bounon, R.; Balzergue, S.; Aubourg, S.; Gautheret, D.; Crespi, M.D.; et al. In plants, decapping prevents RDR6-dependent production of small interfering RNAs from endogenous mRNAs. Nucleic Acids Res. 2015, 43, 2902–2913. [Google Scholar] [CrossRef]
- Braun, J.E.; Truffault, V.; Boland, A.; Huntzinger, E.; Chang, C.T.; Haas, G.; Weichenrieder, O.; Coles, M.; Izaurralde, E. A direct interaction between DCP1 and XRN1 couples mRNA decapping to 5′ exonucleolytic degradation. Nat. Struct. Mol. Biol. 2012, 19, 1324–1331. [Google Scholar] [CrossRef]
- Benková, E.; Michniewicz, M.; Sauer, M.; Teichmann, T.; Seifertová, D.; Jürgens, G.; Friml, J. Local, efflux-dependent auxin gradients as a common module for plant organ formation. Cell 2003, 115, 591–602. [Google Scholar] [CrossRef]
- Bilsborough, G.D.; Runions, A.; Barkoulas, M.; Jenkins, H.W.; Hasson, A.; Galinha, C.; Laufs, P.; Hay, A.; Prusinkiewicz, P.; Tsiantis, M. Model for the regulation of Arabidopsis thaliana leaf margin development. Proc. Natl. Acad. Sci. USA 2011, 108, 3424–3429. [Google Scholar] [CrossRef]
- Hay, A.; Barkoulas, M.; Tsiantis, M. ASYMMETRIC LEAVES1 and auxin activities converge to repress BREVIPEDICELLUS expression and promote leaf development in Arabidopsis. Development 2006, 133, 3955–3961. [Google Scholar] [CrossRef] [PubMed]
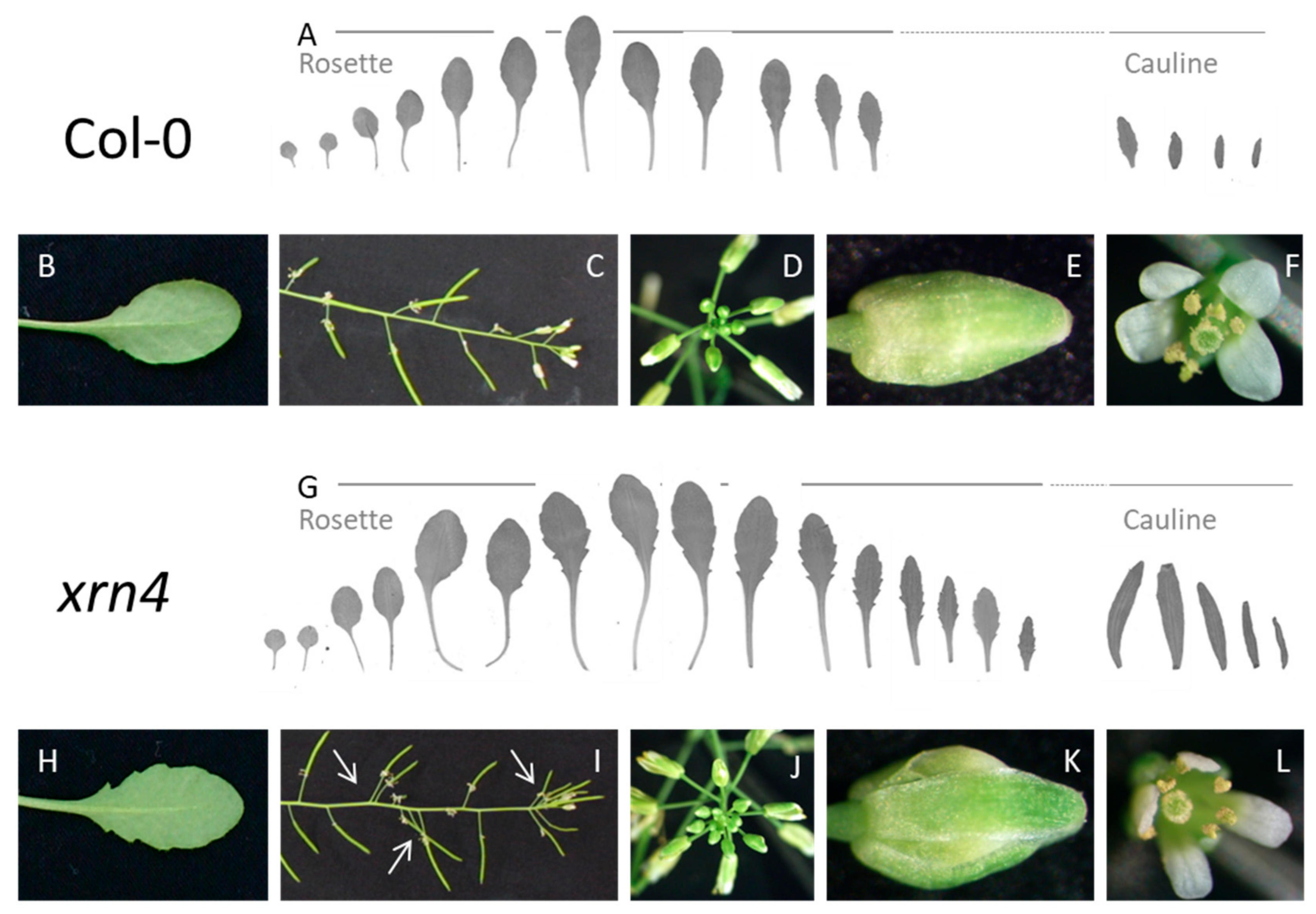

| Col-0 | xrn4 | p-Value | |
|---|---|---|---|
| 14 days-old | |||
| Primary Root Length (mm) | 51.3 ± 1.3 | 61.6 ± 1.4 | 4.0 × 10−6 |
| N. Lateral Roots | 19.9 ± 1.2 | 12.8 ± 0.7 | 1.6 × 10−5 |
| 28 days-old | |||
| Fresh Weight (mg) | 105 ± 10 | 325 ± 29 | 4.4 × 10−7 |
| N. of Serrations on 5th Leaf | 0.8 ± 0.2 | 2.0 ± 0.2 | 3.1 × 10−4 |
| N. of Serrations on 7th Leaf | 1.3 ± 0.3 | 3.5 ± 0.2 | 2.5 × 10−6 |
| 34 days-old | |||
| Number of Leaves | 11.9 ± 0.2 | 17.4 ± 1.0 | 1.2 × 10−5 |
| 45 days-old | |||
| N. of duplicated siliques | 0.0 ± 0.0 | 3.3 ± 0.6 | 3.1 × 10−4 |
| N. of aborted siliques | 0.1 ± 0.1 | 0.7 ± 0.2 | 3.4 × 10−2 |
| N. of Silique insertions | 25.0 ± 1.3 | 29.3 ± 0.7 | 2.9 × 10−3 |
| Internode length (mm) | 7.9 ± 0.1 | 6.3 ± 0.2 | 2.7 × 10−6 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Windels, D.; Bucher, E. The 5′-3′ Exoribonuclease XRN4 Regulates Auxin Response via the Degradation of Auxin Receptor Transcripts. Genes 2018, 9, 638. https://doi.org/10.3390/genes9120638
Windels D, Bucher E. The 5′-3′ Exoribonuclease XRN4 Regulates Auxin Response via the Degradation of Auxin Receptor Transcripts. Genes. 2018; 9(12):638. https://doi.org/10.3390/genes9120638
Chicago/Turabian StyleWindels, David, and Etienne Bucher. 2018. "The 5′-3′ Exoribonuclease XRN4 Regulates Auxin Response via the Degradation of Auxin Receptor Transcripts" Genes 9, no. 12: 638. https://doi.org/10.3390/genes9120638
APA StyleWindels, D., & Bucher, E. (2018). The 5′-3′ Exoribonuclease XRN4 Regulates Auxin Response via the Degradation of Auxin Receptor Transcripts. Genes, 9(12), 638. https://doi.org/10.3390/genes9120638






