Inherited Variation in Vitamin D Genes and Type 1 Diabetes Predisposition
Abstract
:1. Introduction
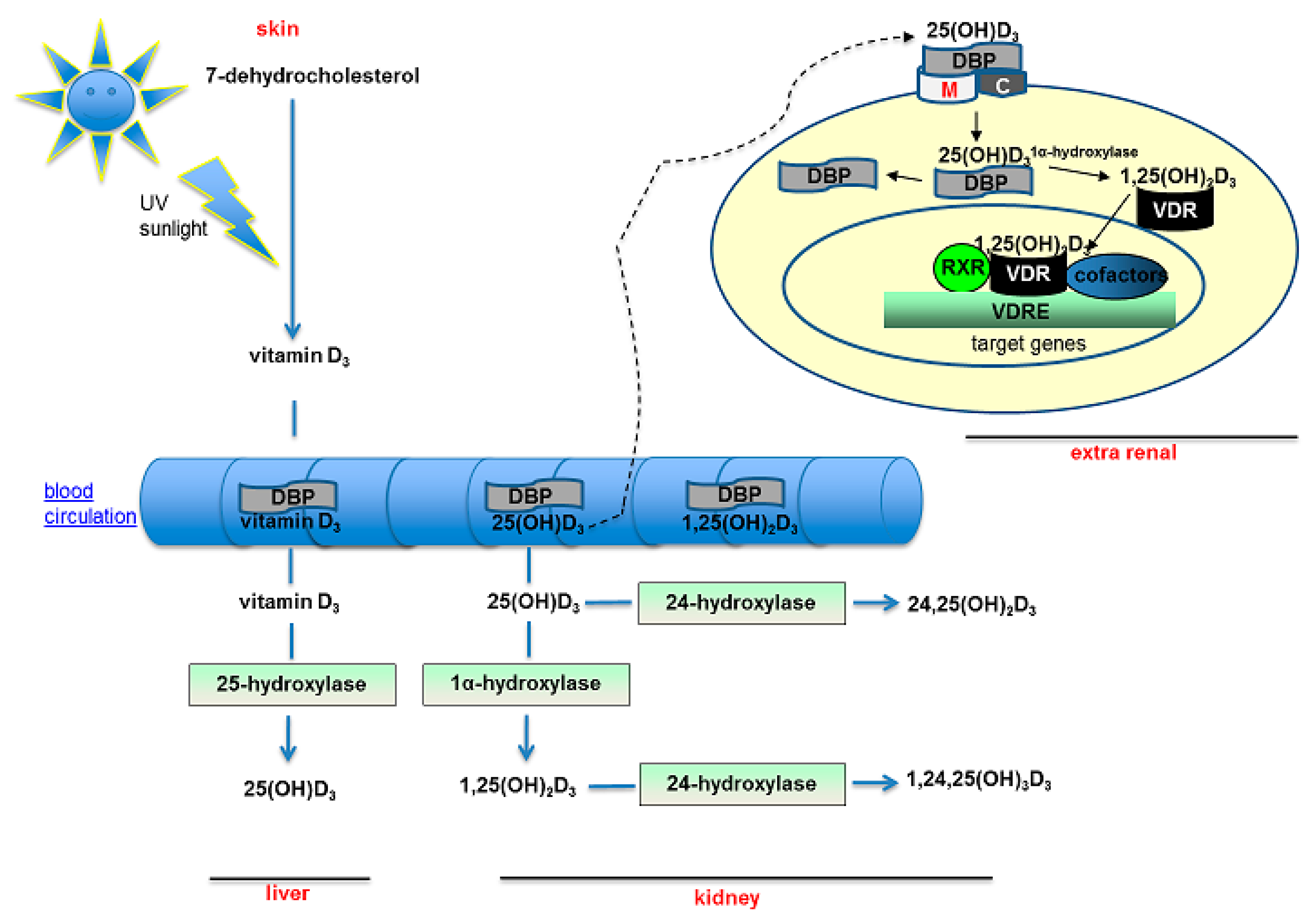
2. Vitamin D Pathways
2.1. Vitamin D Receptor Gene
2.2. Vitamin D Receptor and Meta-Analysis, Diabetes Complications and Monogenetic Vitamin D Disorders
2.3. Other Vitamin D Metabolism Components
2.4. Major Susceptibility to Type 1 Diabetes by HLA and Other Immune Genes: Vitamin D
2.5. Vitamin D Intervention in Type 1 Diabetes and Pharmacogenomics
3. Conclusions
Acknowledgments
Conflicts of Interest
References
- Rewers, M.; Ludvigsson, J. Environmental risk factors for type 1 diabetes. Lancet 2016, 387, 2340–2348. [Google Scholar] [CrossRef]
- Mathieu, C. Vitamin D and diabetes: Where do we stand? Diabetes Res. Clin. Pract. 2015, 108, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Riachy, R.; Vandewalle, B.; Belaich, S.; Kerr-Conte, J.; Gmyr, V.; Zerimech, F.; d’Herbomez, M.; Lefebvre, J.; Pattou, F. Beneficial effect of 1,25 dihydroxyvitamin D3 on cytokine-treated human pancreatic islets. J. Endocrinol. 2001, 169, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Van Belle, T.L.; Vanherwegen, A.S.; Feyaerts, D.; De Clercq, P.; Verstuyf, A.; Korf, H.; Gysemans, C.; Mathieu, C. 1,25-dihydroxyvitamin D3 and its analog TX527 promote a stable regulatory T cell phenotype in T cells from type 1 diabetes patients. PLoS ONE 2014, 9, e109194. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, G.B.; Gysemans, C.A.; Demengeot, J.; da Cunha, J.P.; Vanherwegen, A.S.; Overbergh, L.; Van Belle, T.L.; Pauwels, F.; Verstuyf, A.; Korf, H.; et al. 1,25-dihydroxyvitamin D3 promotes tolerogenic dendritic cells with functional migratory properties in nod mice. J. Immunol. 2014, 192, 4210–4220. [Google Scholar] [CrossRef] [PubMed]
- Holick, M.F.; Chen, T.C.; Lu, Z.; Sauter, E. Vitamin D and skin physiology: A d-lightful story. J. Bone Miner. Res. 2007, 22, V28–V33. [Google Scholar] [CrossRef] [PubMed]
- Fraser, D.R.; Kodicek, E. Unique biosynthesis by kidney of a biological active vitamin D metabolite. Nature 1970, 228, 764–766. [Google Scholar] [CrossRef] [PubMed]
- Kliewer, S.A.; Umesono, K.; Noonan, D.J.; Heyman, R.A.; Evans, R.M. Convergence of 9-cis retinoic acid and peroxisome proliferator signalling pathways through heterodimer formation of their receptors. Nature 1992, 358, 771–774. [Google Scholar] [CrossRef] [PubMed]
- Carlberg, C.; Campbell, M.J. Vitamin D receptor signaling mechanisms: Integrated actions of a well-defined transcription factor. Steroids 2013, 78, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Rowling, M.J.; Kemmis, C.M.; Taffany, D.A.; Welsh, J. Megalin-mediated endocytosis of vitamin D binding protein correlates with 25-hydroxycholecalciferol actions in human mammary cells. J. Nutr. 2006, 136, 2754–2759. [Google Scholar] [PubMed]
- Veldman, C.M.; Cantorna, M.T.; DeLuca, H.F. Expression of 1,25-dihydroxyvitamin D3 receptor in the immune system. Arch. Biochem. Biophys. 2000, 374, 334–338. [Google Scholar] [CrossRef] [PubMed]
- Verstuyf, A.; Carmeliet, G.; Bouillon, R.; Mathieu, C. Vitamin D: A pleiotropic hormone. Kidney Int. 2010, 78, 140–145. [Google Scholar] [CrossRef] [PubMed]
- Zehnder, D.; Bland, R.; Williams, M.C.; McNinch, R.W.; Howie, A.J.; Stewart, P.M.; Hewison, M. Extrarenal expression of 25-hydroxyvitamin D3-1 alpha-hydroxylase. J. Clin. Endocrinol. Metab. 2001, 86, 888–894. [Google Scholar] [PubMed]
- Pike, J.W.; Meyer, M.B. The vitamin d receptor: New paradigms for the regulation of gene expression by 1,25-dihydroxyvitamin D3. Endocrinol. Metab. Clin. N. Am. 2010, 39, 255–269. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, K.; Kesterson, R.A.; Yamamoto, H.; Taketani, Y.; Nishiwaki, E.; Tatsumi, S.; Inoue, Y.; Morita, K.; Takeda, E.; Pike, J.W. Structural organization of the human vitamin d receptor chromosomal gene and its promoter. Mol. Endocrinol. 1997, 11, 1165–1179. [Google Scholar] [CrossRef] [PubMed]
- Morrison, N.A.; Yeoman, R.; Kelly, P.J.; Eisman, J.A. Contribution of trans-acting factor alleles to normal physiological variability: Vitamin D receptor gene polymorphism and circulating osteocalcin. Proc. Natl. Acad. Sci. USA 1992, 89, 6665–6669. [Google Scholar] [CrossRef] [PubMed]
- Arai, H.; Miyamoto, K.; Taketani, Y.; Yamamoto, H.; Iemori, Y.; Morita, K.; Tonai, T.; Nishisho, T.; Mori, S.; Takeda, E. A vitamin D receptor gene polymorphism in the translation initiation codon: Effect on protein activity and relation to bone mineral density in Japanese women. J. Bone Miner. Res. 1997, 12, 915–921. [Google Scholar] [CrossRef] [PubMed]
- Jurutka, P.W.; Remus, L.S.; Whitfield, G.K.; Thompson, P.D.; Hsieh, J.C.; Zitzer, H.; Tavakkoli, P.; Galligan, M.A.; Dang, H.T.; Haussler, C.A.; et al. The polymorphic n terminus in human vitamin d receptor isoforms influences transcriptional activity by modulating interaction with transcription factor iib. Mol. Endocrinol. 2000, 14, 401–420. [Google Scholar] [CrossRef] [PubMed]
- McDermott, M.F.; Ramachandran, A.; Ogunkolade, B.W.; Aganna, E.; Curtis, D.; Boucher, B.J.; Snehalatha, C.; Hitman, G.A. Allelic variation in the vitamin D receptor influences susceptibility to iddm in Indian Asians. Diabetologia 1997, 40, 971–975. [Google Scholar] [CrossRef] [PubMed]
- Hauache, O.M.; Lazaretti-Castro, M.; Andreoni, S.; Gimeno, S.G.; Brandao, C.; Ramalho, A.C.; Kasamatsu, T.S.; Kunii, I.; Hayashi, L.F.; Dib, S.A.; et al. Vitamin D receptor gene polymorphism: Correlation with bone mineral density in a Brazilian population with insulin-dependent diabetes mellitus. Osteoporos. Int. 1998, 8, 204–210. [Google Scholar] [CrossRef] [PubMed]
- Pani, M.A.; Knapp, M.; Donner, H.; Braun, J.; Baur, M.P.; Usadel, K.H.; Badenhoop, K. Vitamin D receptor allele combinations influence genetic susceptibility to type 1 diabetes in Germans. Diabetes 2000, 49, 504–507. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.J.; Lei, H.H.; Yeh, J.I.; Chiu, K.C.; Lee, K.C.; Chen, M.C.; Tai, T.Y.; Chuang, L.M. Vitamin d receptor gene polymorphisms influence susceptibility to type 1 diabetes mellitus in the Taiwanese population. Clin. Endocrinol. 2000, 52, 575–580. [Google Scholar] [CrossRef]
- Ban, Y.; Taniyama, M.; Yanagawa, T.; Yamada, S.; Maruyama, T.; Kasuga, A.; Ban, Y. Vitamin D receptor initiation codon polymorphism influences genetic susceptibility to type 1 diabetes mellitus in the Japanese population. BMC Med. Genet. 2001. [Google Scholar] [CrossRef]
- Guja, C.; Marshall, S.; Welsh, K.; Merriman, M.; Smith, A.; Todd, J.A.; Ionescu-Tirgoviste, C. The study of ctla-4 and vitamin D receptor polymorphisms in the Romanian type 1 diabetes population. J. Cell. Mol. Med. 2002, 6, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Yokota, I.; Satomura, S.; Kitamura, S.; Taki, Y.; Naito, E.; Ito, M.; Nisisho, K.; Kuroda, Y. Association between vitamin D receptor genotype and age of onset in juvenile Japanese patients with type 1 diabetes. Diabetes Care 2002, 25, 1244. [Google Scholar] [CrossRef] [PubMed]
- Gyorffy, B.; Vasarhelyi, B.; Krikovszky, D.; Madacsy, L.; Tordai, A.; Tulassay, T.; Szabo, A. Gender-specific association of vitamin D receptor polymorphism combinations with type 1 diabetes mellitus. Eur. J. Endocrinol. 2002, 147, 803–808. [Google Scholar] [CrossRef] [PubMed]
- Fassbender, W.J.; Goertz, B.; Weismuller, K.; Steinhauer, B.; Stracke, H.; Auch, D.; Linn, T.; Bretzel, R.G. VDR gene polymorphisms are overrepresented in german patients with type 1 diabetes compared to healthy controls without effect on biochemical parameters of bone metabolism. Horm. Metab. Res. 2002, 34, 330–337. [Google Scholar] [CrossRef] [PubMed]
- Taverna, M.J.; Sola, A.; Guyot-Argenton, C.; Pacher, N.; Bruzzo, F.; Slama, G.; Reach, G.; Selam, J.L. Taq I polymorphism of the vitamin D receptor and risk of severe diabetic retinopathy. Diabetologia 2002, 45, 436–442. [Google Scholar] [CrossRef] [PubMed]
- Skrabic, V.; Zemunik, T.; Situm, M.; Terzic, J. Vitamin D receptor polymorphism and susceptibility to type 1 diabetes in the Dalmatian population. Diabetes Res. Clin. Pract. 2003, 59, 31–35. [Google Scholar] [CrossRef]
- Motohashi, Y.; Yamada, S.; Yanagawa, T.; Maruyama, T.; Suzuki, R.; Niino, M.; Fukazawa, T.; Kasuga, A.; Hirose, H.; Matsubara, K.; et al. Vitamin D receptor gene polymorphism affects onset pattern of type 1 diabetes. J. Clin. Endocrinol. Metab. 2003, 88, 3137–3140. [Google Scholar] [CrossRef] [PubMed]
- Turpeinen, H.; Hermann, R.; Vaara, S.; Laine, A.P.; Simell, O.; Knip, M.; Veijola, R.; Ilonen, J. Vitamin D receptor polymorphisms: No association with type 1 diabetes in the Finnish population. Eur. J. Endocrinol. 2003, 149, 591–596. [Google Scholar] [CrossRef] [PubMed]
- Nejentsev, S.; Cooper, J.D.; Godfrey, L.; Howson, J.M.; Rance, H.; Nutland, S.; Walker, N.M.; Guja, C.; Ionescu-Tirgoviste, C.; Savage, D.A.; et al. Analysis of the vitamin D receptor gene sequence variants in type 1 diabetes. Diabetes 2004, 53, 2709–2712. [Google Scholar] [CrossRef] [PubMed]
- Bianco, M.G.; Minicucci, L.; Calevo, M.G.; Lorini, R. Vitamin D receptor polymorphisms: Are they really associated with type 1 diabetes? Eur. J. Endocrinol. 2004, 151, 641–642. [Google Scholar] [CrossRef] [PubMed]
- Audi, L.; Marti, G.; Esteban, C.; Oyarzabal, M.; Chueca, M.; Gussinye, M.; Yeste, D.; Fernandez-Cancio, M.; Andaluz, P.; Carrascosa, A. VDR gene polymorphism at exon 2 start codon (FOKI) may have influenced type 1 diabetes mellitus susceptibility in two Spanish populations. Diabetic Med. 2004, 21, 393–394. [Google Scholar] [CrossRef] [PubMed]
- Zemunik, T.; Škrabić, V.; Boraska, V.; Diklić, D.; Terzić, I.M.; Čapkun, V.; Peruzović, M.; Terzić, J. Foki polymorphism, vitamin D receptor, and interleukin-1 receptor haplotypes are associated with type 1 diabetes in the Dalmatian population. J. Mol. Diagn. 2005, 7, 600–604. [Google Scholar] [CrossRef]
- San-Pedro, J.I.; Bilbao, J.R.; Perez de Nanclares, G.; Vitoria, J.C.; Martul, P.; Castano, L. Heterogeneity of vitamin D receptor gene association with celiac disease and type 1 diabetes mellitus. Autoimmunity 2005, 38, 439–444. [Google Scholar] [CrossRef] [PubMed]
- Taverna, M.J.; Selam, J.L.; Slama, G. Association between a protein polymorphism in the start codon of the vitamin D receptor gene and severe diabetic retinopathy in c-peptide-negative type 1 diabetes. J. Clin. Endocrinol. Metab. 2005, 90, 4803–4808. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Lopez, E.; Jansen, T.; Ivaskevicius, V.; Kahles, H.; Klepzig, C.; Oldenburg, J.; Badenhoop, K. Protection from type 1 diabetes by vitamin D receptor haplotypes. Ann. N. Y. Acad. Sci. 2006, 1079, 327–334. [Google Scholar] [CrossRef] [PubMed]
- Kanazawa, Y.; Motohashi, Y.; Yamada, S.; Oikawa, Y.; Shigihara, T.; Okubo, Y.; Maruyama, T.; Shimada, A. Frequency of ctla-4 gene ct60 polymorphism may not be affected by vitamin D receptor gene BSM I polymorphism or HLA DR9 in autoimmune-related type 1 diabetes in the Japanese. Ann. N. Y. Acad. Sci. 2006, 1079, 251–256. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.H.; Liu, Z.L.; Wang, H.; Sun, Q.; Li, W.H.; Yang, G.H.; Liu, Q.Y. Effects of vitamin D receptor gene polymorphisms on susceptibility to type 1 diabetes mellitus. Chin. Med. Sci. J. 2006, 21, 95–98. [Google Scholar] [PubMed]
- Capoluongo, E.; Pitocco, D.; Concolino, P.; Santonocito, C.; Di Stasio, E.; d’Onofrio, G.; Manto, A.; Giardina, B.; Ghirlanda, G.; Ameglio, F.; et al. Slight association between type 1 diabetes and “FF” VDR FOKI genotype in patients from the Italian Lazio region. Lack of association with diabetes complications. Clin. Biochem. 2006, 39, 888–892. [Google Scholar] [CrossRef] [PubMed]
- Mimbacas, A.; Trujillo, J.; Gascue, C.; Javiel, G.; Cardoso, H. Prevalence of vitamin D receptor gene polymorphism in a uruguayan population and its relation to type 1 diabetes mellitus. Genet. Mol. Res. 2007, 6, 534–542. [Google Scholar] [PubMed]
- Garcia, D.; Angel, B.; Carrasco, E.; Albala, C.; Santos, J.L.; Perez-Bravo, F. Vdr polymorphisms influence the immune response in type 1 diabetic children from Santiago, Chile. Diabetes Res. Clin. Pract. 2007, 77, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Shimada, A.; Kanazawa, Y.; Motohashi, Y.; Yamada, S.; Maruyama, T.; Ikegami, H.; Awata, T.; Kawasaki, E.; Kobayashi, T.; Nakanishi, K.; et al. Evidence for association between vitamin D receptor Bsmi polymorphism and type 1 diabetes in Japanese. J. Autoimmun. 2008, 30, 207–211. [Google Scholar] [CrossRef] [PubMed]
- Lemos, M.C.; Fagulha, A.; Coutinho, E.; Gomes, L.; Bastos, M.; Barros, L.; Carrilho, F.; Geraldes, E.; Regateiro, F.J.; Carvalheiro, M. Lack of association of vitamin D receptor gene polymorphisms with susceptibility to type 1 diabetes mellitus in the Portuguese population. Hum. Immunol. 2008, 69, 134–138. [Google Scholar] [CrossRef] [PubMed]
- Boraska, V.; Skrabic, V.; Zeggini, E.; Groves, C.J.; Buljubasic, M.; Peruzovic, M.; Zemunik, T. Family-based analysis of vitamin d receptor gene polymorphisms and type 1 diabetes in the population of South Croatia. J. Hum. Genet. 2008, 53, 210–214. [Google Scholar] [CrossRef] [PubMed]
- Lopez, T.; Garcia, D.; Angel, B.; Carrasco, E.; Codner, E.; Ugarte, F.; Perez-Bravo, F. Association between fok I vitamin D receptor (VDR) gene polymorphism and plasmatic concentrations of transforming growth factor-beta1 and interferon gamma in type 1 diabetes mellitus. Med. Clin. 2008, 130, 81–84. [Google Scholar] [CrossRef]
- Mory, D.B.; Rocco, E.R.; Miranda, W.L.; Kasamatsu, T.; Crispim, F.; Dib, S.A. Prevalence of vitamin D receptor gene polymorphisms Foki and Bsmi in Brazilian individuals with type 1 diabetes and their relation to beta-cell autoimmunity and to remaining beta-cell function. Hum. Immunol. 2009, 70, 447–451. [Google Scholar] [CrossRef] [PubMed]
- Panierakis, C.; Goulielmos, G.; Mamoulakis, D.; Maraki, S.; Papavasiliou, E.; Galanakis, E. Staphylococcus aureus nasal carriage might be associated with vitamin D receptor polymorphisms in type 1 diabetes. Int. J. Infect. Dis. 2009, 13, e437–e443. [Google Scholar] [CrossRef] [PubMed]
- Panierakis, C.; Goulielmos, G.; Mamoulakis, D.; Petraki, E.; Papavasiliou, E.; Galanakis, E. Vitamin D receptor gene polymorphisms and susceptibility to type 1 diabetes in Crete, Greece. Clin. Immunol. 2009, 133, 276–281. [Google Scholar] [CrossRef] [PubMed]
- Howson, J.M.; Walker, N.M.; Smyth, D.J.; Todd, J.A. Analysis of 19 genes for association with type I diabetes in the type I diabetes genetics consortium families. Genes Immun. 2009, 10, S74–S84. [Google Scholar] [CrossRef] [PubMed]
- Israni, N.; Goswami, R.; Kumar, A.; Rani, R. Interaction of vitamin D receptor with HLA DRB1 0301 in type 1 diabetes patients from North India. PLoS ONE 2009, 4, e8023. [Google Scholar] [CrossRef] [PubMed]
- Bucan, K.; Ivanisevic, M.; Zemunik, T.; Boraska, V.; Skrabic, V.; Vatavuk, Z.; Galetovic, D.; Znaor, L. Retinopathy and nephropathy in type 1 diabetic patients—Association with polymorphysms of vitamin D-receptor, tnf, neuro-D and IL-1 receptor 1 genes. Coll. Antropol. 2009, 33, 99–105. [Google Scholar] [PubMed]
- Kahles, H.; Morahan, G.; Todd, J.A.; Badenhoop, K.; Type, I.D.G.C. Association analyses of the vitamin D receptor gene in 1654 families with type I diabetes. Genes Immun. 2009, 10, S60–S63. [Google Scholar] [CrossRef] [PubMed]
- Kocabas, A.; Karaguzel, G.; Imir, N.; Yavuzer, U.; Akcurin, S. Effects of vitamin D receptor gene polymorphisms on susceptibility to disease and bone mineral density in turkish patients with type 1 diabetes mellitus. J. Pediatr. Endocrinol. Metab. 2010, 23, 1289–1297. [Google Scholar] [CrossRef] [PubMed]
- Martin, R.J.; McKnight, A.J.; Patterson, C.C.; Sadlier, D.M.; Maxwell, A.P.; Warren, U.K.G.S.G. A rare haplotype of the vitamin D receptor gene is protective against diabetic nephropathy. Nephrol. Dial. Transplant. 2010, 25, 497–503. [Google Scholar] [CrossRef] [PubMed]
- Cooper, J.D.; Smyth, D.J.; Walker, N.M.; Stevens, H.; Burren, O.S.; Wallace, C.; Greissl, C.; Ramos-Lopez, E.; Hypponen, E.; Dunger, D.B.; et al. Inherited variation in vitamin D genes is associated with predisposition to autoimmune disease type 1 diabetes. Diabetes 2011, 60, 1624–1631. [Google Scholar] [CrossRef] [PubMed]
- Gogas Yavuz, D.; Keskin, L.; Kiyici, S.; Sert, M.; Yazici, D.; Sahin, I.; Yuksel, M.; Deyneli, O.; Aydin, H.; Tuncel, E.; et al. Vitamin d receptor gene Bsmi, Foki, Apai, Taqi polymorphisms and bone mineral density in a group of turkish type 1 diabetic patients. Acta Diabetol. 2011, 48, 329–336. [Google Scholar] [CrossRef] [PubMed]
- Sahin, S.B.; Cetinkalp, S.; Erdogan, M.; Yilmaz, C.; Berdeli, A. Fas, fas ligand, and vitamin d receptor Foki gene polymorphisms in patients with type 1 diabetes mellitus in the Aegean Region of Turkey. Genet. Test Mol. Biomark. 2012, 16, 1179–1183. [Google Scholar] [CrossRef] [PubMed]
- Mohammadnejad, Z.; Ghanbari, M.; Ganjali, R.; Afshari, J.T.; Heydarpour, M.; Taghavi, S.M.; Fatemi, S.; Rafatpanah, H. Association between vitamin D receptor gene polymorphisms and type 1 diabetes mellitus in Iranian population. Mol. Biol. Rep. 2012, 39, 831–837. [Google Scholar] [CrossRef] [PubMed]
- Bonakdaran, S.; Abbaszadegan, M.R.; Dadkhah, E.; Khajeh-Dalouie, M. Vitamin D receptor gene polymorphisms in type 1 diabetes mellitus: A new pattern from khorasan province, islamic republic of Iran. East. Mediterr. Health J. 2012, 18, 614–619. [Google Scholar] [PubMed]
- Vedralova, M.; Kotrbova-Kozak, A.; Zeleznikova, V.; Zoubkova, H.; Rychlik, I.; Cerna, M. Polymorphisms in the vitamin D receptor gene and parathyroid hormone gene in the development and progression of diabetes mellitus and its chronic complications, diabetic nephropathy and non-diabetic renal disease. Kidney Blood Press. Res. 2012, 36, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Frederiksen, B.; Liu, E.; Romanos, J.; Steck, A.K.; Yin, X.; Kroehl, M.; Fingerlin, T.E.; Erlich, H.; Eisenbarth, G.S.; Rewers, M.; et al. Investigation of the vitamin d receptor gene (VDR) and its interaction with protein tyrosine phosphatase, non-receptor type 2 gene (PTPN2) on risk of islet autoimmunity and type 1 diabetes: The diabetes autoimmunity study in the young (daisy). J. Steroid Biochem. Mol. Biol. 2013, 133, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Hamed, E.O.; Abdel-Aal, A.M.; Din, A.K.; Atia, M.M. Vitamin D level and Fok-I vitamin D receptor gene polymorphism in Egyptian patients with type-1 diabetes. Egypt. J. Immunol. 2013, 20, 1–10. [Google Scholar] [PubMed]
- De Azevedo Silva, J.; Guimaraes, R.L.; Brandao, L.A.; Araujo, J.; Segat, L.; Crovella, S.; Sandrin-Garcia, P. Vitamin D receptor (VDR) gene polymorphisms and age onset in type 1 diabetes mellitus. Autoimmunity 2013, 46, 382–387. [Google Scholar] [CrossRef] [PubMed]
- Greer, R.M.; Portelli, S.L.; Hung, B.S.; Cleghorn, G.J.; McMahon, S.K.; Batch, J.A.; Conwell, L.S. Serum vitamin d levels are lower in Australian children and adolescents with type 1 diabetes than in children without diabetes. Pediatr. Diabetes 2013, 14, 31–41. [Google Scholar] [CrossRef] [PubMed]
- Thorsen, S.U.; Mortensen, H.B.; Carstensen, B.; Fenger, M.; Thuesen, B.H.; Husemoen, L.; Bergholdt, R.; Brorsson, C.; Pociot, F.; Linneberg, A.; et al. No association between type 1 diabetes and genetic variation in vitamin D metabolism genes: A danish study. Pediatr. Diabetes 2014, 15, 416–421. [Google Scholar] [CrossRef] [PubMed]
- Abd-Allah, S.H.; Pasha, H.F.; Hagrass, H.A.; Alghobashy, A.A. Vitamin d status and vitamin D receptor gene polymorphisms and susceptibility to type 1 diabetes in Egyptian children. Gene 2014, 536, 430–434. [Google Scholar] [CrossRef] [PubMed]
- Kamel, M.M.; Fouad, S.A.; Salaheldin, O.; El-Razek Ael, R.; El-Fatah, A.I. Impact of vitamin D receptor gene polymorphisms in pathogenesis of type-1 diabetes mellitus. Int. J. Clin. Exp. Med. 2014, 7, 5505–5510. [Google Scholar] [PubMed]
- Cheon, C.K.; Nam, H.K.; Lee, K.H.; Kim, S.Y.; Song, J.S.; Kim, C. Vitamin D receptor gene polymorphisms and type 1 diabetes mellitus in a Korean population. Pediatr. Int. 2015, 57, 870–874. [Google Scholar] [CrossRef] [PubMed]
- Miettinen, M.E.; Smart, M.C.; Kinnunen, L.; Mathews, C.; Harjutsalo, V.; Surcel, H.M.; Lamberg-Allardt, C.; Tuomilehto, J.; Hitman, G.A. Maternal VDR variants rather than 25-hydroxyvitamin d concentration during early pregnancy are associated with type 1 diabetes in the offspring. Diabetologia 2015, 58, 2278–2283. [Google Scholar] [CrossRef] [PubMed]
- Kahles, H.; Fain, P.R.; Baker, P.; Eisenbarth, G.; Badenhoop, K. Genetics of autoimmune thyroiditis in type 1 diabetes reveals a novel association with dpb1*0201: Data from the type 1 diabetes genetics consortium. Diabetes Care 2015, 38 (Suppl. 2), S21–S28. [Google Scholar] [CrossRef] [PubMed]
- Mory, D.B.; Gabbay, M.A.; Rocco, E.R.; Kasamatsu, T.; Crispim, F.; Miranda, W.L.; Dib, S.A. High frequency of vitamin D receptor gene polymorphism foki in Brazilian type 1 diabetes mellitus patients with clinical autoimmune thyroid disease. Diabetol. Metab. Syndr. 2016. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Wei, H.; Chen, X.; Zhao, Z.; Du, H.; Song, W.; Yang, Y.; Zhang, M.; Lu, W.; Pei, Z.; et al. Erbb3-rs2292239 as primary type 1 diabetes association locus among non-hla genes in chinese. Meta Gene 2016, 9, 120–123. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.W.; Magnuson, V.L.; Schiller, J.J.; Wang, X.; Wu, Y.; Ghosh, S. Meta-analysis of vitamin d receptor polymorphisms and type 1 diabetes: A huge review of genetic association studies. Am. J. Epidemiol. 2006, 164, 711–724. [Google Scholar] [CrossRef] [PubMed]
- Ponsonby, A.L.; Pezic, A.; Ellis, J.; Morley, R.; Cameron, F.; Carlin, J.; Dwyer, T. Variation in associations between allelic variants of the vitamin D receptor gene and onset of type 1 diabetes mellitus by ambient winter ultraviolet radiation levels: A meta-regression analysis. Am. J. Epidemiol. 2008, 168, 358–365. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Li, W.; Liu, J.; Wu, W.; Ouyang, H.; Zhang, Q.; Wang, Y.; Liu, L.; Yang, R.; Liu, X.; et al. Polymorphisms in the vitamin D receptor gene and type 1 diabetes mellitus risk: An update by meta-analysis. Mol. Cell. Endocrinol. 2012, 355, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Xi, B.; Reilly, K.H.; Liu, M.; Fu, M. Quantitative assessment of the associations between four polymorphisms (Foki, Apai, Bsmi, Taqi) of vitamin d receptor gene and risk of diabetes mellitus. Mol. Biol. Rep. 2012, 39, 9405–9414. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Zhang, Q.; Xu, N.; Xu, K.; Wang, J.; He, W.; Yang, T. Associations between two polymorphisms (Foki and Bsmi) of vitamin D receptor gene and type 1 diabetes mellitus in Asian population: A meta-analysis. PLoS ONE 2014, 9, e89325. [Google Scholar] [CrossRef] [PubMed]
- Qin, W.H.; Wang, H.X.; Qiu, J.L.; Huang, X.B.; Huang, Y.; Wu, N.R.; Liang, H.S. A meta-analysis of association of vitamin D receptor Bsmi gene polymorphism with the risk of type 1 diabetes mellitus. J. Recept. Signal Trans. Res. 2014, 34, 372–377. [Google Scholar] [CrossRef] [PubMed]
- Tizaoui, K.; Kaabachi, W.; Hamzaoui, A.; Hamzaoui, K. Contribution of VDR polymorphisms to type 1 diabetes susceptibility: Systematic review of case-control studies and meta-analysis. J. Steroid Biochem. Mol. Biol. 2014, 143, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Liu, L.; Chen, X.; He, W.; Yu, X. Associations study of vitamin d receptor gene polymorphisms with diabetic microvascular complications: A meta-analysis. Gene 2014, 546, 6–10. [Google Scholar] [CrossRef] [PubMed]
- Sahin, O.A.; Goksen, D.; Ozpinar, A.; Serdar, M.; Onay, H. Association of vitamin D receptor polymorphisms and type 1 diabetes susceptibility in children: A meta-analysis. Endocr. Connect. 2017, 6, 159–171. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, M.; d’Alesio, A.; Pascussi, J.M.; Kumar, R.; Griffin, M.D.; Dong, X.; Guillozo, H.; Rizk-Rabin, M.; Sinding, C.; Bougneres, P.; et al. Vitamin d-resistant rickets and type 1 diabetes in a child with compound heterozygous mutations of the vitamin d receptor (l263r and r391s): Dissociated responses of the cyp-24 and rel-b promoters to 1,25-dihydroxyvitamin d3. J. Bone Miner. Res. 2006, 21, 886–894. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, S.; Mondal, R.; Banerjee, I.; Sabui, T. Type II vitamin D-dependent rickets with diabetic ketoacidosis. J. Pediatr. Endocrinol. Metab. 2013, 26, 941–943. [Google Scholar] [CrossRef] [PubMed]
- Fang, C.; Li, H.; Li, X.; Xiao, W.; Huang, Y.; Cai, W.; Yang, Y.; Hu, J. De novo mutation of phex in a type 1 diabetes patient. J. Pediatr. Endocrinol. Metab. 2016, 29, 621–626. [Google Scholar] [CrossRef] [PubMed]
- Tiosano, D.; Wildbaum, G.; Gepstein, V.; Verbitsky, O.; Weisman, Y.; Karin, N.; Eztioni, A. The role of vitamin D receptor in innate and adaptive immunity: A study in hereditary vitamin D-resistant rickets patients. J. Clin. Endocrinol. Metab. 2013, 98, 1685–1693. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Lopez, E.; Bruck, P.; Jansen, T.; Herwig, J.; Badenhoop, K. Cyp2r1 (vitamin d 25-hydroxylase) gene is associated with susceptibility to type 1 diabetes and vitamin D levels in Germans. Diabetes Metab. Res. Rev. 2007, 23, 631–636. [Google Scholar] [CrossRef] [PubMed]
- Frederiksen, B.N.; Kroehl, M.; Fingerlin, T.E.; Wong, R.; Steck, A.K.; Rewers, M.; Norris, J.M. Association between vitamin D metabolism gene polymorphisms and risk of islet autoimmunity and progression to type 1 diabetes: The diabetes autoimmunity study in the young (daisy). J. Clin. Endocrinol. Metab. 2013, 98, E1845–E1851. [Google Scholar] [CrossRef] [PubMed]
- Lopez, E.R.; Zwermann, O.; Segni, M.; Meyer, G.; Reincke, M.; Seissler, J.; Herwig, J.; Usadel, K.H.; Badenhoop, K. A promoter polymorphism of the cyp27b1 gene is associated with addison’s disease, hashimoto’s thyroiditis, graves’ disease and type 1 diabetes mellitus in Germans. Eur. J. Endocrinol. 2004, 151, 193–197. [Google Scholar] [CrossRef] [PubMed]
- Bailey, R.; Cooper, J.D.; Zeitels, L.; Smyth, D.J.; Yang, J.H.; Walker, N.M.; Hypponen, E.; Dunger, D.B.; Ramos-Lopez, E.; Badenhoop, K.; et al. Association of the vitamin D metabolism gene CYP27B1 with type 1 diabetes. Diabetes 2007, 56, 2616–2621. [Google Scholar] [CrossRef] [PubMed]
- Fichna, M.; Zurawek, M.; Januszkiewicz-Lewandowska, D.; Fichna, P.; Nowak, J. Ptpn22, pdcd1 and cyp27b1 polymorphisms and susceptibility to type 1 diabetes in polish patients. Int. J. Immunogenet. 2010, 37, 367–372. [Google Scholar] [CrossRef] [PubMed]
- Hussein, A.G.; Mohamed, R.H.; Alghobashy, A.A. Synergism of cyp2r1 and cyp27b1 polymorphisms and susceptibility to type 1 diabetes in Egyptian children. Cell. Immunol. 2012, 279, 42–45. [Google Scholar] [CrossRef] [PubMed]
- Pani, M.A.; Donner, H.; Herwig, J.; Usadel, K.H.; Badenhoop, K. Vitamin d binding protein alleles and susceptibility for type 1 diabetes in germans. Autoimmunity 1999, 31, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Klupa, T.; Malecki, M.; Hanna, L.; Sieradzka, J.; Frey, J.; Warram, J.H.; Sieradzki, J.; Krolewski, A.S. Amino acid variants of the vitamin D-binding protein and risk of diabetes in white americans of european origin. Eur. J. Endocrinol. 1999, 141, 490–493. [Google Scholar] [CrossRef] [PubMed]
- Ongagna, J.C.; Kaltenbacher, M.C.; Sapin, R.; Pinget, M.; Belcourt, A. The hla-dqb alleles and amino acid variants of the vitamin D-binding protein in diabetic patients in alsace. Clin. Biochem. 2001, 34, 59–63. [Google Scholar] [CrossRef]
- Ongagna, J.C.; Pinget, M.; Belcourt, A. Vitamin d-binding protein gene polymorphism association with IA-2 autoantibodies in type 1 diabetes. Clin. Biochem. 2005, 38, 415–419. [Google Scholar] [CrossRef] [PubMed]
- Blanton, D.; Han, Z.; Bierschenk, L.; Linga-Reddy, M.V.; Wang, H.; Clare-Salzler, M.; Haller, M.; Schatz, D.; Myhr, C.; She, J.X.; et al. Reduced serum vitamin D-binding protein levels are associated with type 1 diabetes. Diabetes 2011, 60, 2566–2570. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Lopez, E.; Lange, B.; Penna-Martinez, M.; Bruck, P.; Swiech, K.; Mauf, S.; Kahles, H.; Badenhoop, K. The role of cubilin gene polymorphisms and their influence on 25(OH)D3 and 1,25(OH)2D3 plasma levels in type 1 diabetes patients. J. Steroid Biochem. Mol. Biol. 2010, 121, 442–444. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.B.; Levine, M.A.; Bell, N.H.; Mangelsdorf, D.J.; Russell, D.W. Genetic evidence that the human cyp2r1 enzyme is a key vitamin d 25-hydroxylase. Proc. Natl. Acad. Sci. USA 2004, 101, 7711–7715. [Google Scholar] [CrossRef] [PubMed]
- Thacher, T.D.; Levine, M.A. Cyp2r1 mutations causing vitamin d-deficiency rickets. J. Steroid Biochem. Mol. Biol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.B.; Motola, D.L.; Mangelsdorf, D.J.; Russell, D.W. De-orphanization of cytochrome p450 2r1: A microsomal vitamin d 25-hydroxilase. J. Biol. Chem. 2003, 278, 38084–38093. [Google Scholar] [CrossRef] [PubMed]
- Miller, W.L. Genetic disorders of vitamin d biosynthesis and degradation. J. Steroid Biochem. Mol. Biol. 2017, 165, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Yu, O.B.; Arnold, L.A. Calcitroic acid—A review. ACS Chem. Biol. 2016, 11, 2665–2672. [Google Scholar] [CrossRef] [PubMed]
- Schlingmann, K.P.; Kaufmann, M.; Weber, S.; Irwin, A.; Goos, C.; John, U.; Misselwitz, J.; Klaus, G.; Kuwertz-Broking, E.; Fehrenbach, H.; et al. Mutations in cyp24a1 and idiopathic infantile hypercalcemia. N. Engl. J. Med. 2011, 365, 410–421. [Google Scholar] [CrossRef] [PubMed]
- Chun, R.F.; Lauridsen, A.L.; Suon, L.; Zella, L.A.; Pike, J.W.; Modlin, R.L.; Martineau, A.R.; Wilkinson, R.J.; Adams, J.; Hewison, M. Vitamin D-binding protein directs monocyte responses to 25-hydroxy- and 1,25-dihydroxy vitamin D. J. Clin. Endocrinol. Metab. 2010, 95, 3368–3376. [Google Scholar] [CrossRef] [PubMed]
- Onengut-Gumuscu, S.; Chen, W.M.; Burren, O.; Cooper, N.J.; Quinlan, A.R.; Mychaleckyj, J.C.; Farber, E.; Bonnie, J.K.; Szpak, M.; Schofield, E.; et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat. Genet. 2015, 47, 381–386. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.K.; van den Berg, P.R.; Long, M.D.; Vreugdenhil, A.; Grieshober, L.; Ochs-Balcom, H.M.; Wang, J.; Delcambre, S.; Heikkinen, S.; Carlberg, C.; et al. Integration of VDR genome wide binding and gwas genetic variation data reveals co-occurrence of VDR and NF-kappab binding that is linked to immune phenotypes. BMC Genom. 2017, 18, 132. [Google Scholar] [CrossRef] [PubMed]
- Ramagopalan, S.V.; Heger, A.; Berlanga, A.J.; Maugeri, N.J.; Lincoln, M.R.; Burrell, A.; Handunnetthi, L.; Handel, A.E.; Disanto, G.; Orton, S.M.; et al. A chip-seq defined genome-wide map of vitamin D receptor binding: Associations with disease and evolution. Genome Res. 2010, 20, 1352–1360. [Google Scholar] [CrossRef] [PubMed]
- Eerligh, P.; Koeleman, B.P.; Dudbridge, F.; Jan Bruining, G.; Roep, B.O.; Giphart, M.J. Functional genetic polymorphisms in cytokines and metabolic genes as additional genetic markers for susceptibility to develop type 1 diabetes. Genes Immun. 2004, 5, 36–40. [Google Scholar] [CrossRef] [PubMed]
- Holick, M.F. Vitamin D: A D-lightful solution for health. J. Investig. Med. 2011, 59, 872–880. [Google Scholar] [CrossRef] [PubMed]
- Muscogiuri, G.; Mitri, J.; Mathieu, C.; Badenhoop, K.; Tamer, G.; Orio, F.; Mezza, T.; Vieth, R.; Colao, A.; Pittas, A. Mechanisms in endocrinology: Vitamin D as a potential contributor in endocrine health and disease. Eur. J. Endocrinol. 2014, 171, R101–R110. [Google Scholar] [CrossRef] [PubMed]
- Anonymous. Vitamin D supplement in early childhood and risk for type I (insulin-dependent) diabetes mellitus. The eurodiab substudy 2 study group. Diabetologia 1999, 42, 51–54. [Google Scholar]
- Hypponen, E.; Laara, E.; Reunanen, A.; Jarvelin, M.R.; Virtanen, S.M. Intake of vitamin D and risk of type 1 diabetes: A birth-cohort study. Lancet 2001, 358, 1500–1503. [Google Scholar] [CrossRef]
- Bogdanou, D.; Penna-Martinez, M.; Filmann, N.; Chung, T.L.; Moran-Auth, Y.; Wehrle, J.; Cappel, C.; Huenecke, S.; Herrmann, E.; Koehl, U.; et al. T-lymphocyte and glycemic status after vitamin D treatment in type 1 diabetes: A randomized controlled trial with sequential crossover. Diabetes Metab. Res. Rev. 2017, 33, e2865. [Google Scholar] [CrossRef] [PubMed]
- Moran-Auth, Y.; Penna-Martinez, M.; Badenhoop, K. Vdr foki polymorphism is associated with a reduced t-helper cell population under vitamin D stimulation in type 1 diabetes patients. J. Steroid Biochem. Mol. Biol. 2015, 148, 184–186. [Google Scholar] [CrossRef] [PubMed]
- Mauf, S.; Penna-Martinez, M.; Jentzsch, T.; Ackermann, H.; Henrich, D.; Radeke, H.H.; Bruck, P.; Badenhoop, K.; Ramos-Lopez, E. Immunomodulatory effects of 25-hydroxyvitamin D3 on monocytic cell differentiation and influence of vitamin D3 polymorphisms in type 1 diabetes. J. Steroid Biochem. Mol. Biol. 2015, 147, 17–23. [Google Scholar] [CrossRef] [PubMed]
| Acronym | Full Name | Protein Function | Chr | Position | SNP | locus | Gene Function | Amino Acid Change |
|---|---|---|---|---|---|---|---|---|
| VDR | vitamin D receptor | transcription factor transcriptional control of vitamin D target genes | 12 | 12q13.11 | rs7975232 | intron 8 | no | |
| rs10735810 | exon 2 | missense | Met → Thr | |||||
| rs1544410 | intron 8 | no | ||||||
| rs731236 | exon 9 | synonymous | Ile → Ile | |||||
| CYP2R1 | vitamin D 25-hydroxylase | transforming photo- synthesized and dietary vitamin D into 25(OH)D3 | 11 | 11p15.2 | rs10741657 | 5′ near gene | 2 kb mRNA transcript | |
| rs12794714 | exon 1 | synonymous | Ser → Ser | |||||
| CYP27B1 | 25(OH)D 1-α-hydroxylase | conversion of 25(OH)D3 to 1,25(OH)2D3 | 12 | 12q14.1 | rs10877012 | 5′ near gene | promoter (−1260) | |
| rs4646536 | intron 6 | (+2838) | no | |||||
| DBP or GC | vitamin D binding protein or group-specific component | transport of vitamin D metabolites | 4 | 4q11.13 | rs4588 | exon 11 | missense | Thr → Lys |
| rs7041 | missense | Asp → Glu | ||||||
| CUBN | cubilin | endocytotic receptors capable to internalize vitamin D into the cells | 10 | 10p12.33-p13 | rs3740165 | synonymous | Pro → Pro |
| VDR Gene | Susceptibility to T1D SNPs | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Reference | Author | Year | Population | Total | Case | Control | Comparison Groups | ||
| 1 | [19] | McDermott et al. | 1997 | Indian | 93 | rs1544410, bt, bAT | T1D families | ||
| 2 | [21] | Pani et al. | 2000 | German | 152 | At, Bt, Bat | T1D families | ||
| 3 | [22] | Chang et al. | 2000 | Chinese (Taiwan) | 405 | 157 | 248 | rs7975232, rs1544410 | T1D/control |
| 4 | [23] | Ban et al. | 2001 | Japanese | 360 | 110 | 250 | rs10735810 | T1D/control |
| 5 | [24] | Guja et al. | 2002 | Romanian | 204 | rs10735810, rs731236 | T1D families | ||
| 6 | [26] | Gyorffy et al. | 2002 | Hungarian | 210 | 107 | 103 | bau | T1D/control |
| 7 | [27] | Fassbender et al. | 2002 | German | 132 | 75 | 57 | rs731236 | T1D/control |
| 8 | [28] | Taverna et al. | 2002 | French | 200 | 101 | 99 | rs731236 | T1D with/without DR |
| 9 | [29] | Skrabic et al. | 2003 | Croatian (Dalmatian) | 266 | 134 | 132 | BBAAtt | T1D/control |
| 10 | [30] | Motohashi et al. | 2003 | Japanese | 425 | 203 | 222 | rs1544410 | T1D/control |
| 11 | [34] | Audi et al. | 2004 | Spanish (Barcelona) | 429 | 155 | 274 | rs1544410, rs10735810, bbFF | T1D/control |
| Spanish (Navarre) | 205 | 89 | 116 | rs1544410, rs10735810, bbff | T1D/control | ||||
| 12 | [35] | Zemunik et al. | 2005 | Croatian (Dalmatian) | 266 | 134 | 132 | rs10735810, FbATU | T1D/control |
| 13 | [36] | San Pedro et al. | 2005 | Spanish (Basque) | 159 | 71 | 88 | fBAt | T1D/control |
| 136 | 119 | T1D families | |||||||
| 14 | [37] | Taverna et al. | 2005 | French | 254 | 126 | 128 | rs10735810 | T1D with/without DR |
| 15 | [38] | Ramos-Lopez et al. | 2006 | German | 254 | rs9729, rs731236, rs7975232, rs757343 | T1D families | ||
| 16 | [40] | Xiao et al. | 2006 | Chinese | 54 | 82 | rs1544410 | T1D/control | |
| 17 | [41] | Capoluongo et al. | 2006 | Italian | 246 | 246 | rs10735810 | T1D/control | |
| 18 | [42] | Mimbacas et al. | 2007 | Uruguayan | 45 | rs10735810 | T1D families | ||
| 19 | [43] | Garcia et al. | 2007 | Chilean | 419 | 216 | 203 | BAT | T1D/control |
| 20 | [44] | Shimada et al. | 2008 | Japanese | 1373 | 774 | 599 | rs1544410 | T1D/control |
| 21 | [46] | Boraska et al. | 2008 | Croatian | 160 | rs757343, rs757343-rs1544410 | T1D families | ||
| 22 | [48] | Mory et al. | 2009 | Brazilian | 383 | 189 | 194 | rs1544410 | T1D/control |
| 23 | [49] | Panierakis et al. | 2009 | Greece | 93 | 29 | 64 | rs7975232, rs731236 | T1D with/without SAC |
| 24 | [50] | Panierakis et al. | 2009 | Greece | 196 | 100 | 96 | rs7975232, rs731236, rs1544410, rs10735810 | T1D/control |
| 25 | [52] | Israni et al. | 2009 | Indian | 424 | 233 | 191 | FBAt, fBAT | T1D/control |
| 26 | [53] | Bucan et al. | 2009 | Croatian | 120 | 66 | 54 | rs1544410 | T1D with/without DR |
| 27 | [55] | Kocabas et al. | 2010 | Turkish | 176 | 90 | 86 | rs10735810 | T1D/control |
| 28 | [56] | Martin et al. | 2010 | UK, Irish | 1329 | 655 | 674 | AGT | T1D with/without DN |
| 29 | [59] | Sahin et al. | 2012 | Turkish | 165 | 85 | 80 | rs10735810 | T1D/control |
| 30 | [60] | Mohammadnejad et al. | 2012 | Iranian | 187 | 87 | 100 | rs731236, tAbf, tabF, tAbF | T1D/control |
| 31 | [61] | Bonakdaran et al. | 2012 | Iranian | 114 | 69 | 45 | rs7975232, rs1544410, rs10735810 | T1D/control |
| 32 | [62] | Vedralová et al. | 2012 | Czech | 172 | 54 | 118 | rs10735810 | T1D/control |
| 250 | 132 | 118 | rs10735810, BBFFAATt | DN/control | |||||
| 33 | [63] | Frederiksen et al. | 2013 | North American | 38 | 84 | rs1544410 | T1D+IA/IA | |
| 34 | [65] | De Azevedo et al. | 2013 | Brazilian | 421 | 204 | 217 | rs1540339, rs4760648 | T1D/control |
| 35 | [68] | Abd-Allah et al. | 2014 | Egyptian | 240 | 120 | 120 | rs1544410, rs10735810 | T1D/control |
| 36 | [69] | Kamel et al. | 2014 | Egyptian | 102 | 74 | 28 | rs7975232, rs731236 | T1D/control |
| 37 | [70] | Cheon et al. | 2015 | Korean | 194 | 81 | 113 | rs731236, rs1544410 | T1D/control |
| 38 | [71] | Miettinen et al. | 2015 | Finnish | 2854 | rs731236, rs1544410 | T1D families | ||
| 39 | [73] | Mory et al. | 2016 | Brazilian | 180 | rs10735810 | T1D with/without Abs | ||
| VDR Gene and Meta-Analysis | Susceptibility to T1D SNPs | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Reference | Author | Year | Population | Total | Case | Control | Comparison Groups | ||
| 1 | [76] | Ponsonby et al. | 2008 | Asian, European, Latinos | 18,257 | 2549 | 15,708 | rs1544410, rs10735810 | T1D/control |
| 16 studies | |||||||||
| 2 | [77] | Zhang et al. | 2012 | Asian, European, Latinos | 11,591 | 5335 | 6256 | rs1544410 | T1D/control |
| 26 studies | |||||||||
| 3 | [78] | Wang et al. | 2012 | East Asian | 10,352 | 3854 | 6498 | rs1544410 | T1D/control |
| 25 studies | |||||||||
| 4 | [79] | Wang et al. | 2014 | East Asian, West Asian | 3959 | 1973 | 1986 | rs1544410, rs10735810 | T1D/control |
| 13 studies | |||||||||
| 5 | [81] | Tizaoui et al. | 2014 | Asian, European, Latinos | 8753 | 3332 | 5421 | BAT, bAT | T1D/control |
| 26 studies | |||||||||
| 6 | [82] | Liu et al. | 2014 | French, Polish, Croatian, Irish, Czech Iranian, Chinese | 2734 | 1394 | 1340 | rs10735810 | diabetic + DN/control |
| 8 studies | |||||||||
| 7 | [83] | Sahin et al. | 2017 | Asian, European, Latinos | 2070 | 1053 | 1017 | rs1544410, rs731236 | T1D/control |
| 8 studies | |||||||||
| Other Vitamin D System Components | Susceptibility to T1D SNPs | ||||||
|---|---|---|---|---|---|---|---|
| Author | Year | Population | Total | Case | Control | Comparison Groups | |
| CYP2R1 gene | |||||||
| Ramos-Lopez et al. [88] | 2007 | German | 203 | rs10741657 | T1D families | ||
| 578 | 284 | 294 | rs10741657 | T1D/control | |||
| Cooper et al. [57] | 2011 | British | 1933 | rs10741657, rs12794714 | T1D families | ||
| 18,955 | 8517 | 10,438 | rs10741657, rs12794714 | T1D/control | |||
| CYP27B1 gene | |||||||
| Ramos-Lopez et al. [90] | 2004 | German | 572 | 252 | 320 | rs10877012 | T1D/control |
| Bailey et al. [91] | 2007 | Great Britain, Northern Ireland, | 2774 | rs10877012, rs4646536 | T1D families | ||
| Finland, USA, Norway, Romania | |||||||
| Great Britain | 16,612 | 7854 | 8758 | rs10877012, rs4646536 | T1D/control | ||
| Cooper et al. [57] | 2011 | British | 1933 | rs10877012 | T1D families | ||
| 18,955 | 8517 | 10,438 | rs10877012 | T1D/control | |||
| Hussei et al. [93] | 2012 | Egyptian | 240 | 120 | 120 | rs10877012 | T1D/control |
| DBP (GC) gene | |||||||
| Ongagna et al. [96] | 2001 | Alsatian and North African origin | 95 | 43 | 52 | rs7041 | T1D/control |
| Ongagna et al. [97] | 2005 | 178 | 110 | 68 | rs7041 | T1D/control | |
| Cubilin gene | |||||||
| Ramos-Lopez et al. [99] | 2010 | German | 400 | 200 | 200 | rs3740165 | T1D/control |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Penna-Martinez, M.; Badenhoop, K. Inherited Variation in Vitamin D Genes and Type 1 Diabetes Predisposition. Genes 2017, 8, 125. https://doi.org/10.3390/genes8040125
Penna-Martinez M, Badenhoop K. Inherited Variation in Vitamin D Genes and Type 1 Diabetes Predisposition. Genes. 2017; 8(4):125. https://doi.org/10.3390/genes8040125
Chicago/Turabian StylePenna-Martinez, Marissa, and Klaus Badenhoop. 2017. "Inherited Variation in Vitamin D Genes and Type 1 Diabetes Predisposition" Genes 8, no. 4: 125. https://doi.org/10.3390/genes8040125
APA StylePenna-Martinez, M., & Badenhoop, K. (2017). Inherited Variation in Vitamin D Genes and Type 1 Diabetes Predisposition. Genes, 8(4), 125. https://doi.org/10.3390/genes8040125





