Replication Checkpoint: Tuning and Coordination of Replication Forks in S Phase
Abstract
:1. Introduction
| S. cerevisiae | S. pombe | H. sapiens | Function |
|---|---|---|---|
| Rad24-RFC | Rad17-RFC | RAD17-RFC | RFC-like complex, 9-1-1 clamp loader |
| Ddc1-Rad17-Mec3 | Rad9-Rad1-Hus1 | RAD9-RAD1-HUS1 | 9-1-1 complex, DNA damage checkpoint clamp, Mec1 activation |
| Dpb11 | Cut5/Rad4 | TOPBP1 | Mec1 ATR activation |
| Dna2 | Dna2 | DNA2 | Mec1 activation in S phase |
| Mre11-Rad50-Xrs2 | Mre11/Rad32-Rad50-Nbs1 | MRE11-RAD50-NBS1 | MRX/MRN complex, DSB resection, Tel1/ATM recruitment |
| Mec1-Ddc2 | Rad3-Rad26 | ATR-ATRIP | checkpoint signaling kinase |
| Tel1 | Tel1 | ATM | checkpoint signaling kinase |
| Mrc1 | Mrc1 | Claspin | fork-associated, checkpoint mediator |
| Rad9 | Crb2 | 53BP1, BRCA1 | checkpoint mediator |
| Sgs1 | Rqh1 | BLM, WRN | fork-associated, Rad53 activation |
| Rad53 | Cds1 | CHK2 | effector kinase |
| Chk1 | Chk1 | CHK1 | effector kinase |
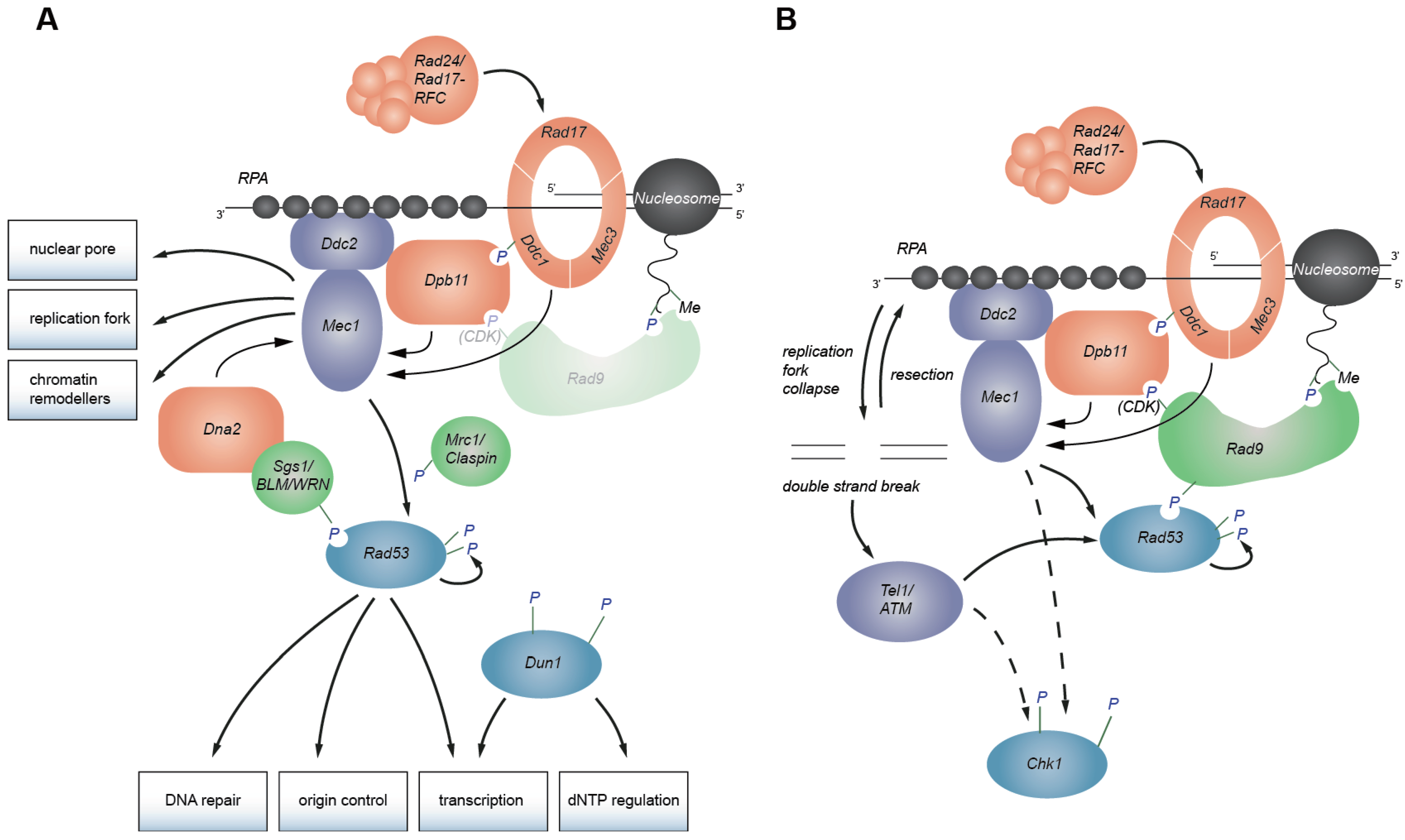
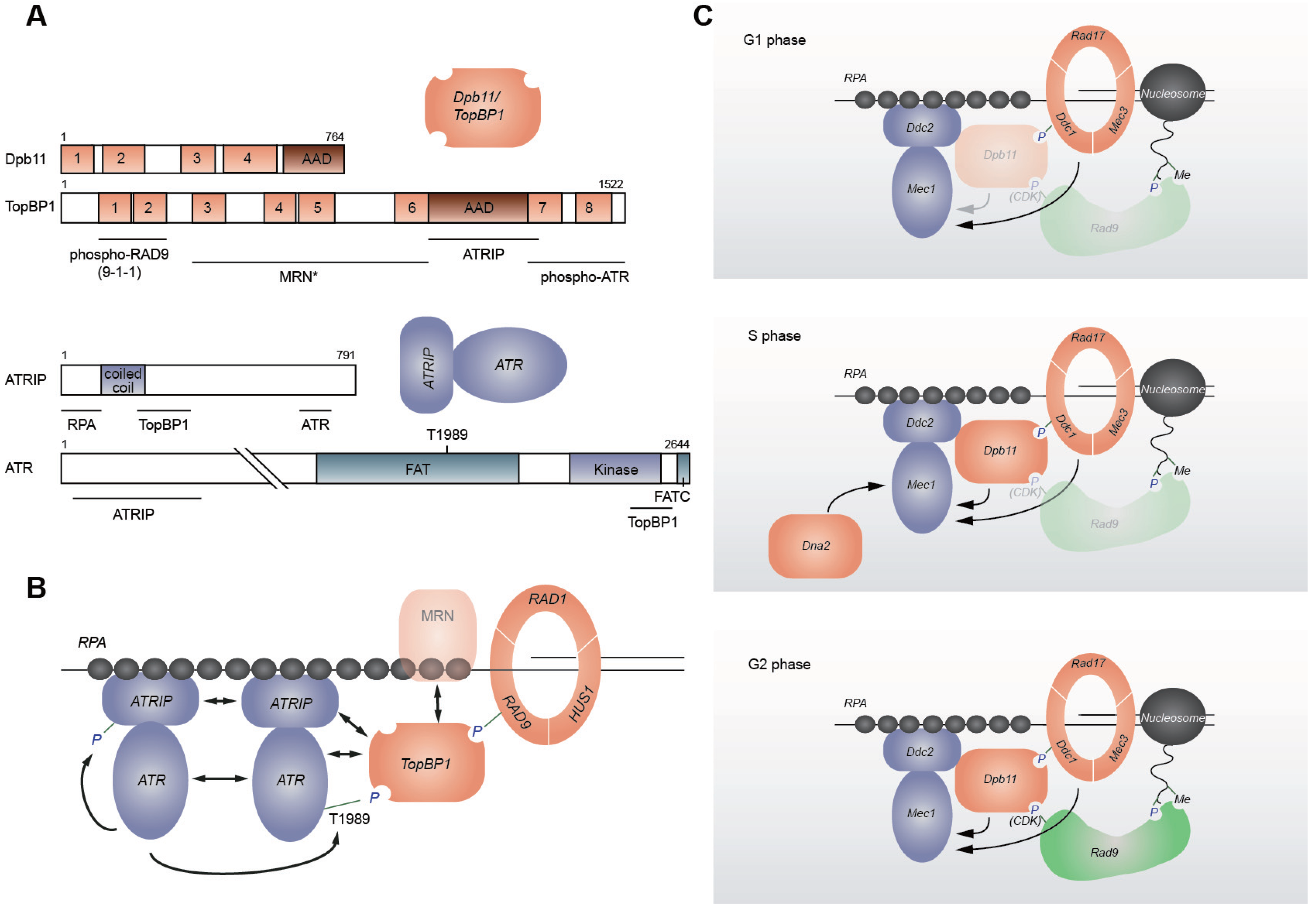
2. Replication Checkpoint Initiation
2.1. Lesions that Activate the Checkpoint
2.2. Drugs Used to Induce and Study Checkpoint Responses
| Treatment/Impediments | Mode of action | Result | Responders ( S.cerevisiae) |
|---|---|---|---|
| hydroxyurea (HU) | inhibits ribonucleotide reductase—dNTP pools become depleted | uncoupling of helicase and polymerase function; ssDNA is exposed | Mec1, Mrc1, Sgs1 |
| aphidicolin | inhibits DNA polymerases | uncoupling of helicase and polymerase function; ssDNA is exposed | Mec1, Mrc1 |
| methylmethanesulfonate (MMS) | alkylates DNA | uncoupling of helicase and polymerase function; ssDNA is exposed; in addition DNA repair takes place, that also leads to ssDNA; requires replication forks to induce checkpoint response | Mec1, Rad9 (Mrc1, Sgs1) |
| ultraviolet light/4-NQO | induces Thymidine dimerization | induces DNA repair, that leads to ssDNA | Mec1, Rad9 Mrc1 |
| crosslinking agents (cisplatin/nitrogen mustard) | causes DNA inter-strand crosslinks | both helicase and polymerase are blocked; in addition DNA repair takes place, that also leads to ssDNA | Mec1/Tel1, Rad9 |
| ionizing irradiation (IR)/bleomycin | causes single and double strand breaks | breaks are directly recognized by MRX-Tel1; resection leads to ssDNA | Mec1/Tel1; Rad9 |
| camptothecin (CPT) | inhibits Topoisomerase I, keeps it in a DNA-bound confirmation | both helicase and polymerase are blocked; double strand breaks are actively induced by DNA repair machinery | Mec1/Tel1, Rad9 |
| natural fork barriers (rDNA, t-RNA genes, transcription | slow down replisome progression | both helicase and polymerase are slowed down | - |
2.3. Mec1/ATR Activation
3. Activation of Effector Kinases
3.1. Rad53 Activation Is Mediated by Rad9 in Response to DNA Damage
3.2. Mrc1 Serves as a Mediator in the Replication Checkpoint
3.3. A Role for Sgs1 in Rad53 Activation
4. Targets of the Replication Checkpoint
4.1. Cell Cycle Regulation
4.2. Essential Function for Cell Viability—dNTP Pool Regulation and Fork Maintenance?
4.3. Replication Origin Control
4.4. Transcription Control
4.5. Coordinating DNA Repair
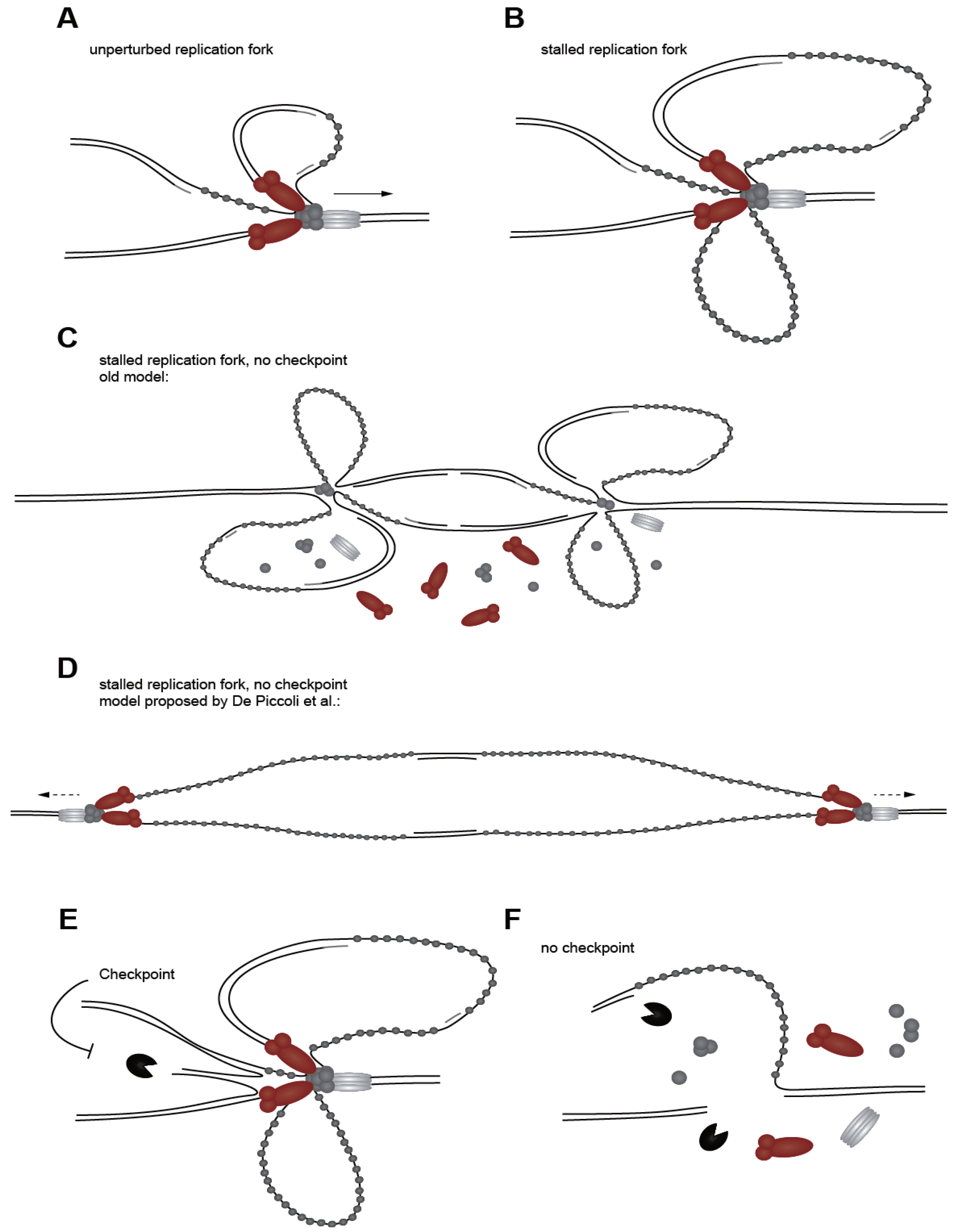
4.6. Replication Fork Stability
| Function | References | |
|---|---|---|
| S. cerevisiae (targeted by) | ||
| Rfa1 (Mec1/Tel1) | a subunit of RPA | [235,236,252] |
| Rfa2 (Mec1/Tel1) | a subunit of RPA | [236,242] |
| Pol1 (Rad53) | a subunit of DNA polα | [236] |
| Pol12 (Rad53) | a subunit of DNA polα | [24] |
| Pol31 (Mec1/Tel1) | a subunit of DNA polδ | [236] |
| Dpb4 (Mec1/Tel1) | a subunit of DNA polε and ISW2 | [235,236] |
| Mcm4(Mec1) | a subunit of MCM | [21] |
| Mcm6 (Mec1) | a subunit of MCM | [21] |
| Psf1 (Mec1) | a subunit of GINS | [224] |
| Mrc1 (Mec1, Rad53) | checkpoint mediator | [25,236] |
| Tof1 (Rad53) | fork protection complex | [235] |
| Ctf4 (Rad53) | polα interactor | [235] |
| Dbf4 (Rad53) | a subunit of DDK | [187,188,236,253] |
| Sgs1 (Mec1) | RecQ helicase | [103] |
| Higher eukaryotes | ||
| RPA1 (ATR/ATM) | a subunit of RPA | [237,241] |
| POLE (ATR/ATM) | a subunit of polε | [237] |
| POLEE4 (ATR/ATM) | a subunit of polε | [237] |
| POLL (ATR/ATM) | a subunit of polλ | [237] |
| MCM2 (ATR/ATM) | a subunit of MCM | [237,239,241] |
| MCM3 (ATR/ATM) | a subunit of MCM | [237] |
| MCM4 (ATR/ATM, CHK1) | a subunit of MCM | [237,239,250] |
| MCM5 (CHK1) | a subunit of MCM | [240] |
| MCM6 (ATR/ATM) | a subunit of MCM | [237,239] |
| MCM7 (ATR/ATM) | a subunit of MCM | [237] |
| POLDIP3 (ATR/ATM) | Polδ interacting factor | [237] |
| MCM10 (ATR/ATM) | replication initiation factor | [237] |
| HELB (ATR/ATM) | DNA helicase B | [237] |
| RFC1 (ATR/ATM) | clamp loader | [237,239,241] |
| RFC3 (ATR/ATM) | clamp loader | [237] |
| PSF2 (ATR/ATM) | a subunit of GINS | [237,241,249] |
| ORC3 (ATR/ATM) | a subunit of ORC | [237] |
| ORC6 (ATR/ATM) | a subunit of ORC | [237] |
| Higher eukaryotes | ||
| DBF4 (ATR/ATM) | a subunit of DDK | [237] |
| Claspin (ATR/ATM) | checkpoint mediator | [237,241] |
| CAF-1B (ATR/ATM) | histone assembly | [241] |
| CTF18 (ATR/ATM) | POLH interactor | [237,241] |
| TopBP1 (ATR/ATM, CHK1) | initiation and ATR activation | [237,240,241] |
| WDHD1 (ATR/ATM) | Polα interactor | [241] |
| BLM (ATR/CHK1) | RecQ helicase | [241,240] |
| FEN1 (CHK1) | 5' flap endonuclease | [240] |
| DNA Ligase 1 (CHK1) | DNA ligase | [239,240] |
| TIPIN (ATR/ATM) | fork protection complex | [237] |
| WRN (ATR/ATM) | RecQ helicase | [232,237] |
5. Checkpoint Recovery
5.1. Protein Phosphatases Down-Regulate the Checkpoint
5.2. Phosphatase-Independent Mechanisms of Checkpoint Down-Regulation
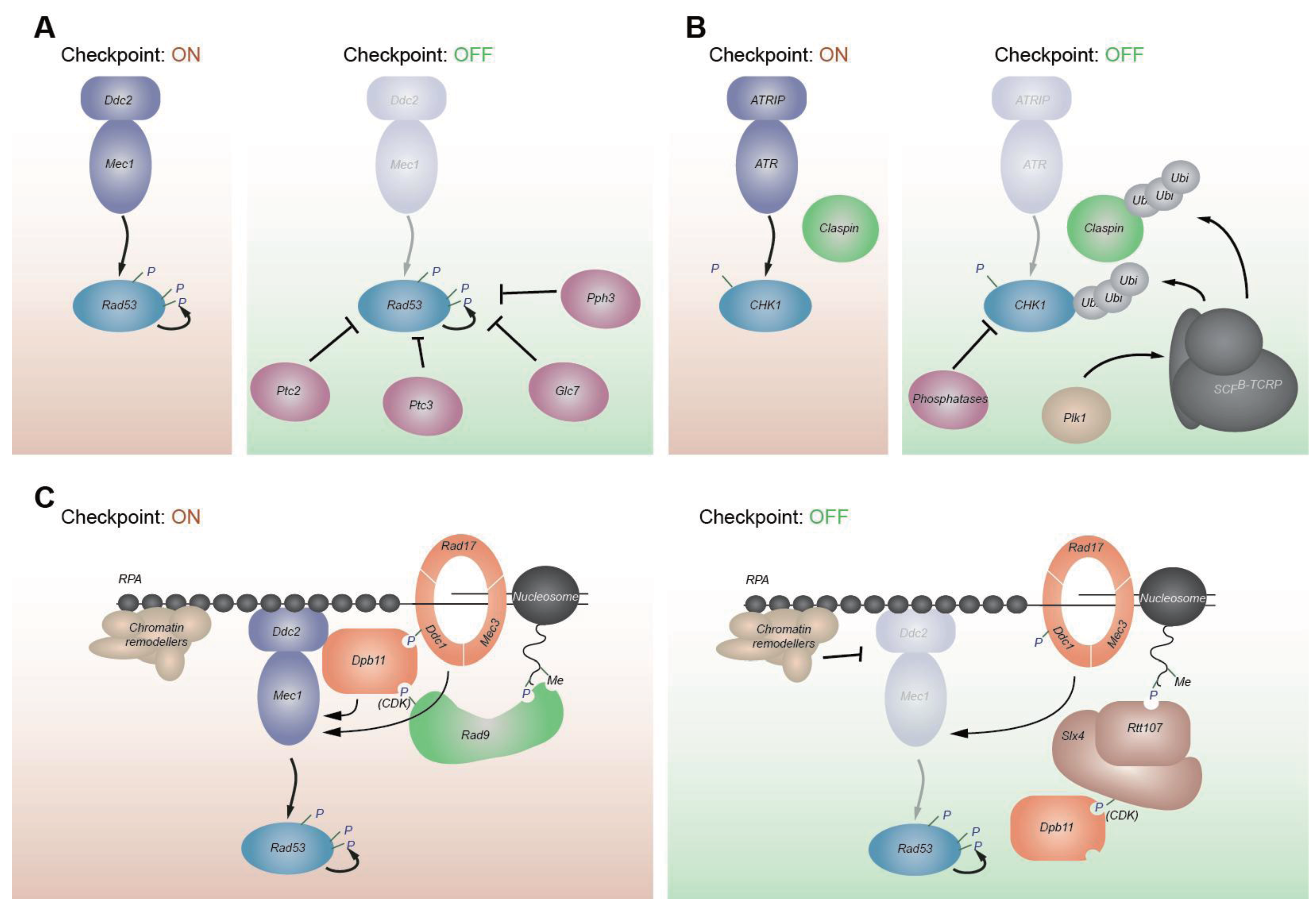
6. Coordination between ATRMec1 and ATMTel1
7. Perspectives
Acknowledgements
Conflicts of Interest
References and Notes
- Hartwell, L.H.; Weinert, T.A. Checkpoints: Controls that ensure the order of cell cycle events. Science 1989, 246, 629–634. [Google Scholar]
- Weinert, T.A.; Kiser, G.L.; Hartwell, L.H. Mitotic checkpoint genes in budding yeast and the dependence of mitosis on DNA replication and repair. Genes Dev. 1994, 8, 652–665. [Google Scholar] [CrossRef]
- Friedel, A.M.; Pike, B.L.; Gasser, S.M. ATR/Mec1: Coordinating fork stability and repair. Curr. Opin. Cell. Biol. 2009, 21, 237–244. [Google Scholar] [CrossRef]
- Allen, J.B.; Zhou, Z.; Siede, W.; Friedberg, E.C.; Elledge, S.J. The SAD1/RAD53 protein kinase controls multiple checkpoints and DNA damage-induced transcription in yeast. Genes Dev. 1994, 8, 2401–2415. [Google Scholar] [CrossRef]
- Harper, J.W.; Elledge, S.J. The DNA damage response: ten years after. Mol. Cell 2007, 28, 739–745. [Google Scholar] [CrossRef]
- Haering, C.H.; Nasmyth, K. Building and breaking bridges between sister chromatids. BioEssays 2003, 25, 1178–1191. [Google Scholar] [CrossRef]
- Halazonetis, T.D.; Gorgoulis, V.G.; Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 2008, 319, 1352–1355. [Google Scholar] [CrossRef]
- Aguilera, A.; Gomez-Gonzalez, B. Genome instability: A mechanistic view of its causes and consequences. Nat. Rev. Genet. 2008, 9, 204–217. [Google Scholar] [CrossRef]
- Elledge, S.J. Cell cycle checkpoints: Preventing an identity crisis. Science 1996, 274, 1664–1672. [Google Scholar] [CrossRef]
- Santocanale, C.; Diffley, J.F. A Mec1- and Rad53-dependent checkpoint controls late-firing origins of DNA replication. Nature 1998, 395, 615–618. [Google Scholar] [CrossRef]
- Shirahige, K.; Hori, Y.; Shiraishi, K.; Yamashita, M.; Takahashi, K.; Obuse, C.; Tsurimoto, T.; Yoshikawa, H. Regulation of DNA-replication origins during cell-cycle progression. Nature 1998, 395, 618–621. [Google Scholar] [CrossRef]
- Tercero, J.A.; Diffley, J.F. Regulation of DNA replication fork progression through damaged DNA by the Mec1/Rad53 checkpoint. Nature 2001, 412, 553–557. [Google Scholar] [CrossRef]
- Kim, S.T.; Lim, D.S.; Canman, C.E.; Kastan, M.B. Substrate specificities and identification of putative substrates of ATM kinase family members. J. Biol. Chem. 1999, 274, 37538–37543. [Google Scholar]
- Chan, D.W.; Ye, R.; Veillette, C.J.; Lees-Miller, S.P. DNA-dependent protein kinase phosphorylation sites in Ku 70/80 heterodimer. Biochemistry 1999, 38, 1819–1828. [Google Scholar] [CrossRef]
- Cortez, D.; Wang, Y.; Qin, J.; Elledge, S.J. Requirement of ATM-dependent phosphorylation of brca1 in the DNA damage response to double-strand breaks. Science 1999, 286, 1162–1166. [Google Scholar] [CrossRef]
- Sweeney, F.D.; Yang, F.; Chi, A.; Shabanowitz, J.; Hunt, D.F.; Durocher, D. Saccharomyces cerevisiae Rad9 acts as a Mec1 adaptor to allow Rad53 activation. Curr. Biol. 2005, 15, 1364–1375. [Google Scholar] [CrossRef]
- Traven, A.; Heierhorst, J. SQ/TQ cluster domains: concentrated ATM/ATR kinase phosphorylation site regions in DNA-damage-response proteins. BioEssays 2005, 27, 397–407. [Google Scholar] [CrossRef]
- Cimprich, K.A.; Cortez, D. ATR: An essential regulator of genome integrity. Nat. Rev. Mol. Cell. Biol. 2008, 9, 616–627. [Google Scholar] [CrossRef]
- Bosotti, R.; Isacchi, A.; Sonnhammer, E.L. FAT: A novel domain in PIK-related kinases. Trends Biochem. Sci. 2000, 25, 225–227. [Google Scholar] [CrossRef]
- Zhao, X.; Chabes, A.; Domkin, V.; Thelander, L.; Rothstein, R. The ribonucleotide reductase inhibitor Sml1 is a new target of the Mec1/Rad53 kinase cascade during growth and in response to DNA damage. EMBO J. 2001, 20, 3544–3553. [Google Scholar] [CrossRef]
- Randell, J.C.; Fan, A.; Chan, C.; Francis, L.I.; Heller, R.C.; Galani, K.; Bell, S.P. Mec1 is one of multiple kinases that prime the Mcm2-7 helicase for phosphorylation by Cdc7. Mol. Cell 2010, 40, 353–363. [Google Scholar] [CrossRef]
- Sanchez, Y.; Bachant, J.; Wang, H.; Hu, F.; Liu, D.; Tetzlaff, M.; Elledge, S.J. Control of the DNA damage checkpoint by chk1 and rad53 protein kinases through distinct mechanisms. Science 1999, 286, 1166–1171. [Google Scholar] [CrossRef]
- Sanchez, Y.; Desany, B.A.; Jones, W.J.; Liu, Q.; Wang, B.; Elledge, S.J. Regulation of RAD53 by the ATM-like kinases MEC1 and TEL1 in yeast cell cycle checkpoint pathways. Science 1996, 271, 357–360. [Google Scholar]
- Pellicioli, A.; Lucca, C.; Liberi, G.; Marini, F.; Lopes, M.; Plevani, P.; Romano, A.; di Fiore, P.P.; Foiani, M. Activation of Rad53 kinase in response to DNA damage and its effect in modulating phosphorylation of the lagging strand DNA polymerase. EMBO J. 1999, 18, 6561–6572. [Google Scholar] [CrossRef]
- Alcasabas, A.A.; Osborn, A.J.; Bachant, J.; Hu, F.; Werler, P.J.; Bousset, K.; Furuya, K.; Diffley, J.F.; Carr, A.M.; Elledge, S.J. Mrc1 transduces signals of DNA replication stress to activate Rad53. Nat. Cell Biol. 2001, 3, 958–965. [Google Scholar] [CrossRef]
- Gilbert, C.S.; Green, C.M.; Lowndes, N.F. Budding yeast Rad9 is an ATP-dependent Rad53 activating machine. Mol. Cell 2001, 8, 129–136. [Google Scholar] [CrossRef]
- Katou, Y.; Kanoh, Y.; Bando, M.; Noguchi, H.; Tanaka, H.; Ashikari, T.; Sugimoto, K.; Shirahige, K. S-phase checkpoint proteins Tof1 and Mrc1 form a stable replication-pausing complex. Nature 2003, 424, 1078–1083. [Google Scholar] [CrossRef]
- Gardner, R.; Putnam, C.W.; Weinert, T. RAD53, DUN1 and PDS1 define two parallel G2/M checkpoint pathways in budding yeast. EMBO J. 1999, 18, 3173–3185. [Google Scholar] [CrossRef]
- Downs, J.A.; Lowndes, N.F.; Jackson, S.P. A role for Saccharomyces cerevisiae histone H2A in DNA repair. Nature 2000, 408, 1001–1004. [Google Scholar] [CrossRef]
- Lindahl, T. Instability and decay of the primary structure of DNA. Nature 1993, 362, 709–715. [Google Scholar] [CrossRef]
- Cha, R.S.; Kleckner, N. ATR homolog Mec1 promotes fork progression, thus averting breaks in replication slow zones. Science 2002, 297, 602–606. [Google Scholar] [CrossRef]
- Cobb, J.A.; Schleker, T.; Rojas, V.; Bjergbaek, L.; Tercero, J.A.; Gasser, S.M. Replisome instability, fork collapse, and gross chromosomal rearrangements arise synergistically from Mec1 kinase and RecQ helicase mutations. Genes Dev. 2005, 19, 3055–3069. [Google Scholar] [CrossRef]
- Sogo, J.M.; Lopes, M.; Foiani, M. Fork reversal and ssDNA accumulation at stalled replication forks owing to checkpoint defects. Science 2002, 297, 599–602. [Google Scholar] [CrossRef]
- Tercero, J.A.; Longhese, M.P.; Diffley, J.F. A central role for DNA replication forks in checkpoint activation and response. Mol. Cell 2003, 11, 1323–1336. [Google Scholar] [CrossRef]
- Myung, K.; Datta, A.; Kolodner, R.D. Suppression of spontaneous chromosomal rearrangements by S phase checkpoint functions in Saccharomyces cerevisiae. Cell 2001, 104, 397–408. [Google Scholar] [CrossRef]
- Shimada, K.; Pasero, P.; Gasser, S.M. ORC and the intra-S-phase checkpoint: a threshold regulates Rad53p activation in S phase. Genes Dev. 2002, 16, 3236–3252. [Google Scholar] [CrossRef]
- MacDougall, C.A.; Byun, T.S.; van, C.; Yee, M.C.; Cimprich, K.A. The structural determinants of checkpoint activation. Genes Dev. 2007, 21, 898–903. [Google Scholar] [CrossRef]
- Huang, J.C.; Svoboda, D.L.; Reardon, J.T.; Sancar, A. Human nucleotide excision nuclease removes thymine dimers from DNA by incising the 22nd phosphodiester bond 5' and the 6th phosphodiester bond 3' to the photodimer. Proc. Nat. Acad. Sci. USA 1992, 89, 3664–3668. [Google Scholar] [CrossRef]
- Jazayeri, A.; Falck, J.; Lukas, C.; Bartek, J.; Smith, G.C.; Lukas, J.; Jackson, S.P. ATM- and cell cycle-dependent regulation of ATR in response to DNA double-strand breaks. Nat. Cell Biol. 2006, 8, 37–45. [Google Scholar] [CrossRef]
- Shiotani, B.; Zou, L. Single-stranded DNA orchestrates an ATM-to-ATR switch at DNA breaks. Mol. Cell 2009, 33, 547–558. [Google Scholar] [CrossRef]
- Costanzo, V.; Robertson, K.; Bibikova, M.; Kim, E.; Grieco, D.; Gottesman, M.; Carroll, D.; Gautier, J. Mre11 protein complex prevents double-strand break accumulation during chromosomal DNA replication. Mol. Cell 2001, 8, 137–147. [Google Scholar] [CrossRef]
- Myers, J.S.; Cortez, D. Rapid activation of ATR by ionizing radiation requires ATM and Mre11. J. Biol. Chem. 2006, 281, 9346–9350. [Google Scholar] [CrossRef]
- Alani, E.; Thresher, R.; Griffith, J.D.; Kolodner, R.D. Characterization of DNA-binding and strand-exchange stimulation properties of y-RPA, a yeast single-strand-DNA-binding protein. J. Mol. Biol. 1992, 227, 54–71. [Google Scholar] [CrossRef]
- Zou, L.; Elledge, S.J. Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science 2003, 300, 1542–1548. [Google Scholar] [CrossRef]
- Ball, H.L.; Myers, J.S.; Cortez, D. ATRIP binding to replication protein A-single-stranded DNA promotes ATR-ATRIP localization but is dispensable for Chk1 phosphorylation. Mol. Biol. Cell 2005, 16, 2372–2381. [Google Scholar] [CrossRef]
- Ball, H.L.; Ehrhardt, M.R.; Mordes, D.A.; Glick, G.G.; Chazin, W.J.; Cortez, D. Function of a conserved checkpoint recruitment domain in ATRIP proteins. Mol. Cell. Biol. 2007, 27, 3367–3377. [Google Scholar] [CrossRef]
- Kim, H.S.; Brill, S.J. Rfc4 interacts with Rpa1 and is required for both DNA replication and DNA damage checkpoints in Saccharomyces cerevisiae. Mol. Cell. Biol. 2001, 21, 3725–3737. [Google Scholar] [CrossRef]
- Dubrana, K.; van Attikum, H.; Hediger, F.; Gasser, S.M. The processing of double-strand breaks and binding of single-strand-binding proteins RPA and Rad51 modulate the formation of ATR-kinase foci in yeast. J. Cell Sci. 2007, 120, 4209–4220. [Google Scholar] [CrossRef]
- Nakada, D.; Hirano, Y.; Tanaka, Y.; Sugimoto, K. Role of the C terminus of Mec1 checkpoint kinase in its localization to sites of DNA damage. Mol. Biol. Cell 2005, 16, 5227–5235. [Google Scholar] [CrossRef]
- Majka, J.; Binz, S.K.; Wold, M.S.; Burgers, P.M. Replication protein A directs loading of the DNA damage checkpoint clamp to 5'-DNA junctions. J. Biol. Chem. 2006, 281, 27855–27861. [Google Scholar] [CrossRef]
- Delacroix, S.; Wagner, J.M.; Kobayashi, M.; Yamamoto, K.; Karnitz, L.M. The Rad9-Hus1-Rad1 (9-1-1) clamp activates checkpoint signaling via TopBP1. Genes Dev. 2007, 21, 1472–1477. [Google Scholar] [CrossRef]
- Lee, J.; Kumagai, A.; Dunphy, W.G. The Rad9-Hus1-Rad1 checkpoint clamp regulates interaction of TopBP1 with ATR. J. Biol. Chem. 2007, 282, 28036–28044. [Google Scholar] [CrossRef]
- Furuya, K.; Poitelea, M.; Guo, L.; Caspari, T.; Carr, A.M. Chk1 activation requires Rad9 S/TQ-site phosphorylation to promote association with C-terminal BRCT domains of Rad4TOPBP1. Genes Dev. 2004, 18, 1154–1164. [Google Scholar] [CrossRef]
- Mordes, D.A.; Glick, G.G.; Zhao, R.; Cortez, D. TopBP1 activates ATR through ATRIP and a PIKK regulatory domain. Genes Dev. 2008, 22, 1478–1489. [Google Scholar]
- Mordes, D.A.; Nam, E.A.; Cortez, D. Dpb11 activates the Mec1-Ddc2 complex. Proc. Nat. Acad. Sci. USA 2008, 105, 18730–18734. [Google Scholar]
- Puddu, F.; Granata, M.; di Nola, L.; Balestrini, A.; Piergiovanni, G.; Lazzaro, F.; Giannattasio, M.; Plevani, P.; Muzi-Falconi, M. Phosphorylation of the budding yeast 9-1-1 complex is required for Dpb11 function in the full activation of the UV-induced DNA damage checkpoint. Mol. Cell. Biol. 2008, 28, 4782–4793. [Google Scholar]
- Paciotti, V.; Lucchini, G.; Plevani, P.; Longhese, M.P. Mec1p is essential for phosphorylation of the yeast DNA damage checkpoint protein Ddc1p, which physically interacts with Mec3p. EMBO J. 1998, 17, 4199–4209. [Google Scholar] [CrossRef]
- Wang, Y.; Qin, J. MSH2 and ATR form a signaling module and regulate two branches of the damage response to DNA methylation. Proc. Nat. Acad. Sci. USA 2003, 100, 15387–15392. [Google Scholar]
- Liu, Y.; Fang, Y.; Shao, H.; Lindsey-Boltz, L.; Sancar, A.; Modrich, P. Interactions of human mismatch repair proteins MutSalpha and MutLalpha with proteins of the ATR-Chk1 pathway. J. Biol. Chem. 2010, 285, 5974–5982. [Google Scholar]
- Pabla, N.; Ma, Z.; McIlhatton, M.A.; Fishel, R.; Dong, Z. hMSH2 recruits ATR to DNA damage sites for activation during DNA damage-induced apoptosis. J. Biol. Chem. 2011, 286, 10411–10418. [Google Scholar] [CrossRef]
- Lambert, S.; Carr, A.M. Checkpoint responses to replication fork barriers. Biochimie 2005, 87, 591–602. [Google Scholar] [CrossRef]
- Azvolinsky, A.; Giresi, P.G.; Lieb, J.D.; Zakian, V.A. Highly transcribed RNA polymerase II genes are impediments to replication fork progression in Saccharomyces cerevisiae. Mol. Cell 2009, 34, 722–734. [Google Scholar] [CrossRef]
- Ray Chaudhuri, A.; Hashimoto, Y.; Herrador, R.; Neelsen, K.J.; Fachinetti, D.; Bermejo, R.; Cocito, A.; Costanzo, V.; Lopes, M. Topoisomerase I poisoning results in PARP-mediated replication fork reversal. Nat. Struct. Mol. Biol. 2012, 19, 417–423. [Google Scholar] [CrossRef]
- Huang, D.; Piening, B.D.; Paulovich, A.G. The preference for error-free or error-prone postreplication repair in Saccharomyces cerevisiae exposed to low-dose methyl methanesulfonate is cell cycle dependent. Mol. Cell. Biol. 2013, 33, 1515–1527. [Google Scholar] [CrossRef]
- Hishida, T.; Kubota, Y.; Carr, A.M.; Iwasaki, H. RAD6-RAD18-RAD5-pathway-dependent tolerance to chronic low-dose ultraviolet light. Nature 2009, 457, 612–615. [Google Scholar] [CrossRef]
- Byun, T.S.; Pacek, M.; Yee, M.C.; Walter, J.C.; Cimprich, K.A. Functional uncoupling of MCM helicase and DNA polymerase activities activates the ATR-dependent checkpoint. Genes Dev. 2005, 19, 1040–1052. [Google Scholar] [CrossRef]
- Eklund, H.; Uhlin, U.; Farnegardh, M.; Logan, D.T.; Nordlund, P. Structure and function of the radical enzyme ribonucleotide reductase. Prog. Biophys. Mol. Biol. 2001, 77, 177–268. [Google Scholar] [CrossRef]
- Poli, J.; Tsaponina, O.; Crabbe, L.; Keszthelyi, A.; Pantesco, V.; Chabes, A.; Lengronne, A.; Pasero, P. dNTP pools determine fork progression and origin usage under replication stress. EMBO J. 2012, 31, 883–894. [Google Scholar] [CrossRef]
- Ikegami, S.; Taguchi, T.; Ohashi, M.; Oguro, M.; Nagano, H.; Mano, Y. Aphidicolin prevents mitotic cell division by interfering with the activity of DNA polymerase-alpha. Nature 1978, 275, 458–460. [Google Scholar] [CrossRef]
- Pacek, M.; Walter, J.C. A requirement for MCM7 and Cdc45 in chromosome unwinding during eukaryotic DNA replication. EMBO J. 2004, 23, 3667–3676. [Google Scholar] [CrossRef]
- Lambert, S.; Mason, S.J.; Barber, L.J.; Hartley, J.A.; Pearce, J.A.; Carr, A.M.; McHugh, P.J. Schizosaccharomyces pombe checkpoint response to DNA interstrand cross-links. Mol. Cell. Biol. 2003, 23, 4728–4737. [Google Scholar] [CrossRef]
- Raschle, M.; Knipscheer, P.; Enoiu, M.; Angelov, T.; Sun, J.; Griffith, J.D.; Ellenberger, T.E.; Scharer, O.D.; Walter, J.C. Mechanism of replication-coupled DNA interstrand crosslink repair. Cell 2008, 134, 969–980. [Google Scholar] [CrossRef]
- Emili, A. MEC1-dependent phosphorylation of Rad9p in response to DNA damage. Mol. Cell 1998, 2, 183–189. [Google Scholar] [CrossRef]
- Liu, J.S.; Kuo, S.R.; Melendy, T. Comparison of checkpoint responses triggered by DNA polymerase inhibition versus DNA damaging agents. Mutat. Res. 2003, 532, 215–226. [Google Scholar] [CrossRef]
- Vazquez, M.V.; Rojas, V.; Tercero, J.A. Multiple pathways cooperate to facilitate DNA replication fork progression through alkylated DNA. DNA Repair 2008, 7, 1693–1704. [Google Scholar] [CrossRef]
- Lundin, C.; North, M.; Erixon, K.; Walters, K.; Jenssen, D.; Goldman, A.S.; Helleday, T. Methyl methanesulfonate (MMS) produces heat-labile DNA damage but no detectable in vivo DNA double-strand breaks. Nucl. Acid. Res. 2005, 33, 3799–3811. [Google Scholar]
- Povirk, L.F. DNA damage and mutagenesis by radiomimetic DNA-cleaving agents: Bleomycin, neocarzinostatin and other enediynes. Mutat. Res. 1996, 355, 71–89. [Google Scholar] [CrossRef]
- Neecke, H.; Lucchini, G.; Longhese, M.P. Cell cycle progression in the presence of irreparable DNA damage is controlled by a Mec1- and Rad53-dependent checkpoint in budding yeast. EMBO J. 1999, 18, 4485–4497. [Google Scholar] [CrossRef]
- Olson, E.; Nievera, C.J.; Lee, A.Y.; Chen, L.; Wu, X. The Mre11-Rad50-Nbs1 complex acts both upstream and downstream of ataxia telangiectasia mutated and Rad3-related protein (ATR) to regulate the S-phase checkpoint following UV treatment. J. Biol. Chem. 2007, 282, 22939–22952. [Google Scholar]
- Hsiang, Y.H.; Hertzberg, R.; Hecht, S.; Liu, L.F. Camptothecin induces protein-linked DNA breaks via mammalian DNA topoisomerase I. J. Biol. Chem. 1985, 260, 14873–14878. [Google Scholar]
- Hsiang, Y.H.; Lihou, M.G.; Liu, L.F. Arrest of replication forks by drug-stabilized topoisomerase I-DNA cleavable complexes as a mechanism of cell killing by camptothecin. Cancer Res. 1989, 49, 5077–5082. [Google Scholar]
- Redon, C.; Pilch, D.R.; Rogakou, E.P.; Orr, A.H.; Lowndes, N.F.; Bonner, W.M. Yeast histone 2A serine 129 is essential for the efficient repair of checkpoint-blind DNA damage. EMBO Rep. 2003, 4, 678–684. [Google Scholar] [CrossRef]
- Strumberg, D.; Pilon, A.A.; Smith, M.; Hickey, R.; Malkas, L.; Pommier, Y. Conversion of topoisomerase I cleavage complexes on the leading strand of ribosomal DNA into 5'-phosphorylated DNA double-strand breaks by replication runoff. Mol. Cell. Biol. 2000, 20, 3977–3987. [Google Scholar]
- Regairaz, M.; Zhang, Y.W.; Fu, H.; Agama, K.K.; Tata, N.; Agrawal, S.; Aladjem, M.I.; Pommier, Y. Mus81-mediated DNA cleavage resolves replication forks stalled by topoisomerase I-DNA complexes. J. Cell Biol. 2011, 195, 739–749. [Google Scholar] [CrossRef]
- Nielsen, I.; Bentsen, I.B.; Lisby, M.; Hansen, S.; Mundbjerg, K.; Andersen, A.H.; Bjergbaek, L. A Flp-nick system to study repair of a single protein-bound nick in vivo. Nat. Methods 2009, 6, 753–757. [Google Scholar] [CrossRef]
- Majka, J.; Niedziela-Majka, A.; Burgers, P.M. The checkpoint clamp activates Mec1 kinase during initiation of the DNA damage checkpoint. Mol. Cell 2006, 24, 891–901. [Google Scholar] [CrossRef]
- Navadgi-Patil, V.M.; Burgers, P.M. The unstructured C-terminal tail of the 9–1-1 clamp subunit Ddc1 activates Mec1/ATR via two distinct mechanisms. Mol. Cell 2009, 36, 743–753. [Google Scholar] [CrossRef]
- Bonilla, C.Y.; Melo, J.A.; Toczyski, D.P. Colocalization of sensors is sufficient to activate the DNA damage checkpoint in the absence of damage. Mol. Cell 2008, 30, 267–276. [Google Scholar] [CrossRef]
- Duursma, A.M.; Driscoll, R.; Elias, J.E.; Cimprich, K.A. A Role for the MRN Complex in ATR Activation via TOPBP1 Recruitment. Mol. Cell 2013, 50, 116–122. [Google Scholar] [CrossRef]
- Kumagai, A.; Lee, J.; Yoo, H.Y.; Dunphy, W.G. TopBP1 activates the ATR-ATRIP complex. Cell 2006, 124, 943–955. [Google Scholar] [CrossRef]
- Wang, H.; Elledge, S.J. Genetic and physical interactions between DPB11 and DDC1 in the yeast DNA damage response pathway. Genetics 2002, 160, 1295–1304. [Google Scholar]
- Navadgi-Patil, V.M.; Kumar, S.; Burgers, P.M. The unstructured C-terminal tail of yeast Dpb11 (human TopBP1) protein is dispensable for DNA replication and the S phase checkpoint but required for the G2/M checkpoint. J. Biol. Chem. 2011, 286, 40999–41007. [Google Scholar] [CrossRef]
- Pfander, B.; Diffley, J.F. Dpb11 coordinates Mec1 kinase activation with cell cycle-regulated Rad9 recruitment. EMBO J. 2011, 30, 4897–4907. [Google Scholar] [CrossRef]
- Navadgi-Patil, V.M.; Burgers, P.M. Cell-cycle-specific activators of the Mec1/ATR checkpoint kinase. Biochem. Soc. Trans. 2011, 39, 600–605. [Google Scholar] [CrossRef]
- Puddu, F.; Piergiovanni, G.; Plevani, P.; Muzi-Falconi, M. Sensing of replication stress and Mec1 activation act through two independent pathways involving the 9–1-1 complex and DNA polymerase epsilon. PLoS Genet. 2011, 7, e1002022. [Google Scholar] [CrossRef]
- Lin, S.J.; Wardlaw, C.P.; Morishita, T.; Miyabe, I.; Chahwan, C.; Caspari, T.; Schmidt, U.; Carr, A.M.; Garcia, V. The Rad4(TopBP1) ATR-activation domain functions in G1/S phase in a chromatin-dependent manner. PLoS Genet. 2012, 8, e1002801. [Google Scholar] [CrossRef]
- Bjergbaek, L.; Cobb, J.A.; Tsai-Pflugfelder, M.; Gasser, S.M. Mechanistically distinct roles for Sgs1p in checkpoint activation and replication fork maintenance. EMBO J. 2005, 24, 405–417. [Google Scholar] [CrossRef]
- Berens, T.J.; Toczyski, D.P. Colocalization of Mec1 and Mrc1 is sufficient for Rad53 phosphorylation in vivo. Mol. Biol. Cell 2012, 23, 1058–1067. [Google Scholar] [CrossRef]
- Kumar, S.; Burgers, P.M. Lagging strand maturation factor Dna2 is a component of the replication checkpoint initiation machinery. Genes Dev. 2013, 27, 313–321. [Google Scholar] [CrossRef]
- Cejka, P.; Cannavo, E.; Polaczek, P.; Masuda-Sasa, T.; Pokharel, S.; Campbell, J.L.; Kowalczykowski, S.C. DNA end resection by Dna2-Sgs1-RPA and its stimulation by Top3-Rmi1 and Mre11-Rad50-Xrs2. Nature 2010, 467, 112–116. [Google Scholar] [CrossRef]
- Frei, C.; Gasser, S.M. The yeast Sgs1p helicase acts upstream of Rad53p in the DNA replication checkpoint and colocalizes with Rad53p in S-phase-specific foci. Genes Dev. 2000, 14, 81–96. [Google Scholar]
- Fu, Y.; Zhu, Y.; Zhang, K.; Yeung, M.; Durocher, D.; Xiao, W. Rad6-Rad18 mediates a eukaryotic SOS response by ubiquitinating the 9-1-1 checkpoint clamp. Cell 2008, 133, 601–611. [Google Scholar] [CrossRef]
- Hegnauer, A.M.; Hustedt, N.; Shimada, K.; Pike, B.L.; Vogel, M.; Amsler, P.; Rubin, S.M.; van Leeuwen, F.; Guenole, A.; van Attikum, H.; et al. An N-terminal acidic region of Sgs1 interacts with Rpa70 and recruits Rad53 kinase to stalled forks. EMBO J. 2012, 31, 3768–3783. [Google Scholar] [CrossRef]
- Liu, S.; Shiotani, B.; Lahiri, M.; Marechal, A.; Tse, A.; Leung, C.C.; Glover, J.N.; Yang, X.H.; Zou, L. ATR autophosphorylation as a molecular switch for checkpoint activation. Mol. Cell 2011, 43, 192–202. [Google Scholar] [CrossRef]
- Nam, E.A.; Zhao, R.; Glick, G.G.; Bansbach, C.E.; Friedman, D.B.; Cortez, D. Thr-1989 phosphorylation is a marker of active ataxia telangiectasia-mutated and Rad3-related (ATR) kinase. J. Biol. Chem. 2011, 286, 28707–28714. [Google Scholar]
- Yue, M.; Singh, A.; Wang, Z.; Xu, Y.J. The phosphorylation network for efficient activation of the DNA replication checkpoint in fission yeast. J. Biol. Chem. 2011, 286, 22864–22874. [Google Scholar]
- Liu, S.; Ho, C.K.; Ouyang, J.; Zou, L. Nek1 kinase associates with ATR-ATRIP and primes ATR for efficient DNA damage signaling. Proc. Nat. Acad. Sci. USA 2013, 110, 2175–2180. [Google Scholar]
- Bakkenist, C.J.; Kastan, M.B. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 2003, 421, 499–506. [Google Scholar] [CrossRef]
- Ball, H.L.; Cortez, D. ATRIP oligomerization is required for ATR-dependent checkpoint signaling. J. Biol. Chem. 2005, 280, 31390–31396. [Google Scholar] [CrossRef]
- Itakura, E.; Sawada, I.; Matsuura, A. Dimerization of the ATRIP protein through the coiled-coil motif and its implication to the maintenance of stalled replication forks. Mol. Biol. Cell 2005, 16, 5551–5562. [Google Scholar] [CrossRef]
- Lindsay, H.D.; Griffiths, D.J.; Edwards, R.J.; Christensen, P.U.; Murray, J.M.; Osman, F.; Walworth, N.; Carr, A.M. S-phase-specific activation of Cds1 kinase defines a subpathway of the checkpoint response in Schizosaccharomyces pombe. Genes Dev. 1998, 12, 382–395. [Google Scholar] [CrossRef]
- Paciotti, V.; Clerici, M.; Scotti, M.; Lucchini, G.; Longhese, M.P. Characterization of mec1 kinase-deficient mutants and of new hypomorphic mec1 alleles impairing subsets of the DNA damage response pathway. Mol. Cell. Biol. 2001, 21, 3913–3925. [Google Scholar] [CrossRef]
- Lee, S.J.; Duong, J.K.; Stern, D.F. A Ddc2-Rad53 fusion protein can bypass the requirements for RAD9 and MRC1 in Rad53 activation. Mol. Biol. Cell 2004, 15, 5443–5455. [Google Scholar] [CrossRef]
- Kim, S.M.; Kumagai, A.; Lee, J.; Dunphy, W.G. Phosphorylation of Chk1 by ATM- and Rad3-related (ATR) in Xenopus egg extracts requires binding of ATRIP to ATR but not the stable DNA-binding or coiled-coil domains of ATRIP. J. Biol. Chem. 2005, 280, 38355–38364. [Google Scholar] [CrossRef]
- Durocher, D.; Jackson, S.P. The FHA domain. FEBS Lett. 2002, 513, 58–66. [Google Scholar] [CrossRef]
- Pike, B.L.; Yongkiettrakul, S.; Tsai, M.D.; Heierhorst, J. Diverse but overlapping functions of the two forkhead-associated (FHA) domains in Rad53 checkpoint kinase activation. J. Biol. Chem. 2003, 278, 30421–30424. [Google Scholar]
- Schwartz, M.F.; Lee, S.J.; Duong, J.K.; Eminaga, S.; Stern, D.F. FHA domain-mediated DNA checkpoint regulation of Rad53. Cell Cycle 2003, 2, 384–396. [Google Scholar]
- Lee, S.J.; Schwartz, M.F.; Duong, J.K.; Stern, D.F. Rad53 phosphorylation site clusters are important for Rad53 regulation and signaling. Mol. Cell. Biol. 2003, 23, 6300–6314. [Google Scholar] [CrossRef]
- Sun, Z.; Fay, D.S.; Marini, F.; Foiani, M.; Stern, D.F. Spk1/Rad53 is regulated by Mec1-dependent protein phosphorylation in DNA replication and damage checkpoint pathways. Genes Dev. 1996, 10, 395–406. [Google Scholar] [CrossRef]
- De la Torre-Ruiz, M.A.; Green, C.M.; Lowndes, N.F. RAD9 and RAD24 define two additive, interacting branches of the DNA damage checkpoint pathway in budding yeast normally required for Rad53 modification and activation. EMBO J. 1998, 17, 2687–2698. [Google Scholar] [CrossRef]
- Navas, T.A.; Sanchez, Y.; Elledge, S.J. RAD9 and DNA polymerase epsilon form parallel sensory branches for transducing the DNA damage checkpoint signal in Saccharomyces cerevisiae. Genes Dev. 1996, 10, 2632–2643. [Google Scholar] [CrossRef]
- Schleker, T.; Shimada, K.; Sack, R.; Pike, B.L.; Gasser, S.M. Cell cycle-dependent phosphorylation of Rad53 kinase by Cdc5 and Cdc28 modulates checkpoint adaptation. Cell Cycle 2010, 9, 350–363. [Google Scholar] [CrossRef]
- Weinert, T.A.; Hartwell, L.H. The RAD9 gene controls the cell cycle response to DNA damage in Saccharomyces cerevisiae. Science 1988, 241, 317–322. [Google Scholar]
- Grenon, M.; Costelloe, T.; Jimeno, S.; O'Shaughnessy, A.; Fitzgerald, J.; Zgheib, O.; Degerth, L.; Lowndes, N.F. Docking onto chromatin via the Saccharomyces cerevisiae Rad9 Tudor domain. Yeast 2007, 24, 105–119. [Google Scholar] [CrossRef]
- Van Leeuwen, F.; Gafken, P.R.; Gottschling, D.E. Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 2002, 109, 745–756. [Google Scholar] [CrossRef]
- Hammet, A.; Magill, C.; Heierhorst, J.; Jackson, S.P. Rad9 BRCT domain interaction with phosphorylated H2AX regulates the G1 checkpoint in budding yeast. EMBO Rep. 2007, 8, 851–857. [Google Scholar] [CrossRef]
- Van Attikum, H.; Fritsch, O.; Hohn, B.; Gasser, S.M. Recruitment of the INO80 complex by H2A phosphorylation links ATP-dependent chromatin remodeling with DNA double-strand break repair. Cell 2004, 119, 777–788. [Google Scholar] [CrossRef]
- Giannattasio, M.; Lazzaro, F.; Plevani, P.; Muzi-Falconi, M. The DNA damage checkpoint response requires histone H2B ubiquitination by Rad6-Bre1 and H3 methylation by Dot1. J. Biol. Chem. 2005, 280, 9879–9886. [Google Scholar] [CrossRef]
- Wysocki, R.; Javaheri, A.; Allard, S.; Sha, F.; Cote, J.; Kron, S.J. Role of Dot1-dependent histone H3 methylation in G1 and S phase DNA damage checkpoint functions of Rad9. Mol. Cell. Biol. 2005, 25, 8430–8443. [Google Scholar] [CrossRef]
- Schwartz, M.F.; Duong, J.K.; Sun, Z.; Morrow, J.S.; Pradhan, D.; Stern, D.F. Rad9 phosphorylation sites couple Rad53 to the Saccharomyces cerevisiae DNA damage checkpoint. Mol. Cell 2002, 9, 1055–1065. [Google Scholar] [CrossRef]
- Sun, Z.; Hsiao, J.; Fay, D.S.; Stern, D.F. Rad53 FHA domain associated with phosphorylated Rad9 in the DNA damage checkpoint. Science 1998, 281, 272–274. [Google Scholar] [CrossRef]
- Ma, J.L.; Lee, S.J.; Duong, J.K.; Stern, D.F. Activation of the checkpoint kinase Rad53 by the phosphatidyl inositol kinase-like kinase Mec1. J. Biol. Chem. 2006, 281, 3954–3963. [Google Scholar] [CrossRef]
- Tanaka, K.; Boddy, M.N.; Chen, X.B.; McGowan, C.H.; Russell, P. Threonine-11, phosphorylated by Rad3 and atm in vitro, is required for activation of fission yeast checkpoint kinase Cds1. Mol. Cell. Biol. 2001, 21, 3398–3404. [Google Scholar] [CrossRef]
- Matsuoka, S.; Rotman, G.; Ogawa, A.; Shiloh, Y.; Tamai, K.; Elledge, S.J. Ataxia telangiectasia-mutated phosphorylates Chk2 in vivo and in vitro. Proc. Nat. Acad. Sci. USA 2000, 97, 10389–10394. [Google Scholar]
- Usui, T.; Foster, S.S.; Petrini, J.H. Maintenance of the DNA-damage checkpoint requires DNA-damage-induced mediator protein oligomerization. Mol. Cell 2009, 33, 147–159. [Google Scholar] [CrossRef]
- Osborn, A.J.; Elledge, S.J. Mrc1 is a replication fork component whose phosphorylation in response to DNA replication stress activates Rad53. Genes Dev. 2003, 17, 1755–1767. [Google Scholar] [CrossRef]
- Tanaka, K.; Russell, P. Mrc1 channels the DNA replication arrest signal to checkpoint kinase Cds1. Nat. Cell Biol. 2001, 3, 966–972. [Google Scholar] [CrossRef]
- Chen, S.H.; Zhou, H. Reconstitution of Rad53 activation by Mec1 through adaptor protein Mrc1. J. Biol. Chem. 2009, 284, 18593–18604. [Google Scholar] [CrossRef]
- Kumagai, A.; Dunphy, W.G. Claspin, a novel protein required for the activation of Chk1 during a DNA replication checkpoint response in Xenopus egg extracts. Mol. Cell 2000, 6, 839–849. [Google Scholar] [CrossRef]
- Kumagai, A.; Kim, S.M.; Dunphy, W.G. Claspin and the activated form of ATR-ATRIP collaborate in the activation of Chk1. J. Biol. Chem. 2004, 279, 49599–49608. [Google Scholar] [CrossRef]
- Chini, C.C.; Chen, J. Human claspin is required for replication checkpoint control. J. Biol. Chem. 2003, 278, 30057–30062. [Google Scholar] [CrossRef]
- Bando, M.; Katou, Y.; Komata, M.; Tanaka, H.; Itoh, T.; Sutani, T.; Shirahige, K. Csm3, Tof1, and Mrc1 form a heterotrimeric mediator complex that associates with DNA replication forks. J. Biol. Chem. 2009, 284, 34355–34365. [Google Scholar] [CrossRef]
- Lou, H.; Komata, M.; Katou, Y.; Guan, Z.; Reis, C.C.; Budd, M.; Shirahige, K.; Campbell, J.L. Mrc1 and DNA polymerase epsilon function together in linking DNA replication and the S phase checkpoint. Mol. Cell 2008, 32, 106–117. [Google Scholar]
- Komata, M.; Bando, M.; Araki, H.; Shirahige, K. The direct binding of Mrc1, a checkpoint mediator, to Mcm6, a replication helicase, is essential for the replication checkpoint against methyl methanesulfonate-induced stress. Mol. Cell. Biol. 2009, 29, 5008–5019. [Google Scholar] [CrossRef]
- Szyjka, S.J.; Viggiani, C.J.; Aparicio, O.M. Mrc1 is required for normal progression of replication forks throughout chromatin in S. cerevisiae. Mol. Cell 2005, 19, 691–697. [Google Scholar] [CrossRef]
- Tourriere, H.; Versini, G.; Cordon-Preciado, V.; Alabert, C.; Pasero, P. Mrc1 and Tof1 promote replication fork progression and recovery independently of Rad53. Mol. Cell 2005, 19, 699–706. [Google Scholar] [CrossRef]
- Naylor, M.L.; Li, J.M.; Osborn, A.J.; Elledge, S.J. Mrc1 phosphorylation in response to DNA replication stress is required for Mec1 accumulation at the stalled fork. Proc. Nat. Acad. Sci. USA 2009, 106, 12765–12770. [Google Scholar]
- Cobb, J.A.; Bjergbaek, L.; Shimada, K.; Frei, C.; Gasser, S.M. DNA polymerase stabilization at stalled replication forks requires Mec1 and the RecQ helicase Sgs1. EMBO J. 2003, 22, 4325–4336. [Google Scholar] [CrossRef]
- Lopes, M.; Cotta-Ramusino, C.; Pellicioli, A.; Liberi, G.; Plevani, P.; Muzi-Falconi, M.; Newlon, C.S.; Foiani, M. The DNA replication checkpoint response stabilizes stalled replication forks. Nature 2001, 412, 557–561. [Google Scholar] [CrossRef]
- Lucca, C.; Vanoli, F.; Cotta-Ramusino, C.; Pellicioli, A.; Liberi, G.; Haber, J.; Foiani, M. Checkpoint-mediated control of replisome-fork association and signalling in response to replication pausing. Oncogene 2004, 23, 1206–1213. [Google Scholar] [CrossRef]
- Patro, B.S.; Frohlich, R.; Bohr, V.A.; Stevnsner, T. WRN helicase regulates the ATR-CHK1-induced S-phase checkpoint pathway in response to topoisomerase-I-DNA covalent complexes. J. Cell Sci. 2011, 124, 3967–3979. [Google Scholar] [CrossRef]
- Sanchez, Y.; Wong, C.; Thoma, R.S.; Richman, R.; Wu, Z.; Piwnica-Worms, H.; Elledge, S.J. Conservation of the Chk1 checkpoint pathway in mammals: linkage of DNA damage to Cdk regulation through Cdc25. Science 1997, 277, 1497–1501. [Google Scholar] [CrossRef]
- Rhind, N.; Russell, P. Tyrosine phosphorylation of cdc2 is required for the replication checkpoint in Schizosaccharomyces pombe. Mol. Cell. Biol. 1998, 18, 3782–3787. [Google Scholar]
- Karlsson-Rosenthal, C.; Millar, J.B. Cdc25: Mechanisms of checkpoint inhibition and recovery. Trends Cell Biol. 2006, 16, 285–292. [Google Scholar] [CrossRef]
- Sorensen, C.S.; Syljuasen, R.G. Safeguarding genome integrity: The checkpoint kinases ATR, CHK1 and WEE1 restrain CDK activity during normal DNA replication. Nucl. Acid. Res. 2012, 40, 477–486. [Google Scholar] [CrossRef]
- Peng, C.Y.; Graves, P.R.; Thoma, R.S.; Wu, Z.; Shaw, A.S.; Piwnica-Worms, H. Mitotic and G2 checkpoint control: Regulation of 14–3-3 protein binding by phosphorylation of Cdc25C on serine-216. Science 1997, 277, 1501–1505. [Google Scholar] [CrossRef]
- Busino, L.; Donzelli, M.; Chiesa, M.; Guardavaccaro, D.; Ganoth, D.; Dorrello, N.V.; Hershko, A.; Pagano, M.; Draetta, G.F. Degradation of Cdc25A by beta-TrCP during S phase and in response to DNA damage. Nature 2003, 426, 87–91. [Google Scholar] [CrossRef]
- Jin, J.; Shirogane, T.; Xu, L.; Nalepa, G.; Qin, J.; Elledge, S.J.; Harper, J.W. SCFbeta-TRCP links Chk1 signaling to degradation of the Cdc25A protein phosphatase. Genes Dev. 2003, 17, 3062–3074. [Google Scholar] [CrossRef]
- Sorger, P.K.; Murray, A.W. S-phase feedback control in budding yeast independent of tyrosine phosphorylation of p34cdc28. Nature 1992, 355, 365–368. [Google Scholar] [CrossRef]
- Paulovich, A.G.; Hartwell, L.H. A checkpoint regulates the rate of progression through S phase in S. cerevisiae in response to DNA damage. Cell 1995, 82, 841–847. [Google Scholar] [CrossRef]
- Clarke, D.J.; Segal, M.; Jensen, S.; Reed, S.I. Mec1p regulates Pds1p levels in S phase: Complex coordination of DNA replication and mitosis. Nat. Cell Biol. 2001, 3, 619–627. [Google Scholar] [CrossRef]
- Hu, F.; Wang, Y.; Liu, D.; Li, Y.; Qin, J.; Elledge, S.J. Regulation of the Bub2/Bfa1 GAP complex by Cdc5 and cell cycle checkpoints. Cell 2001, 107, 655–665. [Google Scholar] [CrossRef]
- Krishnan, V.; Nirantar, S.; Crasta, K.; Cheng, A.Y.; Surana, U. DNA replication checkpoint prevents precocious chromosome segregation by regulating spindle behavior. Mol. Cell 2004, 16, 687–700. [Google Scholar] [CrossRef]
- Desany, B.A.; Alcasabas, A.A.; Bachant, J.B.; Elledge, S.J. Recovery from DNA replicational stress is the essential function of the S-phase checkpoint pathway. Genes Dev. 1998, 12, 2956–2970. [Google Scholar] [CrossRef]
- Zhao, X.; Muller, E.G.; Rothstein, R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol. Cell 1998, 2, 329–340. [Google Scholar] [CrossRef]
- Brown, E.J.; Baltimore, D. ATR disruption leads to chromosomal fragmentation and early embryonic lethality. Genes Dev. 2000, 14, 397–402. [Google Scholar]
- De Klein, A.; Muijtjens, M.; van Os, R.; Verhoeven, Y.; Smit, B.; Carr, A.M.; Lehmann, A.R.; Hoeijmakers, J.H. Targeted disruption of the cell-cycle checkpoint gene ATR leads to early embryonic lethality in mice. Curr. Biol. 2000, 10, 479–482. [Google Scholar]
- Liu, Q.; Guntuku, S.; Cui, X.S.; Matsuoka, S.; Cortez, D.; Tamai, K.; Luo, G.; Carattini-Rivera, S.; DeMayo, F.; Bradley, A.; et al. Chk1 is an essential kinase that is regulated by Atr and required for the G(2)/M DNA damage checkpoint. Genes Dev. 2000, 14, 1448–1459. [Google Scholar]
- Takai, H.; Tominaga, K.; Motoyama, N.; Minamishima, Y.A.; Nagahama, H.; Tsukiyama, T.; Ikeda, K.; Nakayama, K.; Nakanishi, M.; Nakayama, K. Aberrant cell cycle checkpoint function and early embryonic death in Chk1(−/−) mice. Genes Dev. 2000, 14, 1439–1447. [Google Scholar]
- Bester, A.C.; Roniger, M.; Oren, Y.S.; Im, M.M.; Sarni, D.; Chaoat, M.; Bensimon, A.; Zamir, G.; Shewach, D.S.; Kerem, B. Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell 2011, 145, 435–446. [Google Scholar] [CrossRef]
- Chabes, A.; Georgieva, B.; Domkin, V.; Zhao, X.; Rothstein, R.; Thelander, L. Survival of DNA damage in yeast directly depends on increased dNTP levels allowed by relaxed feedback inhibition of ribonucleotide reductase. Cell 2003, 112, 391–401. [Google Scholar] [CrossRef]
- Nordlund, P.; Reichard, P. Ribonucleotide reductases. Annu. Rev. Biochem. 2006, 75, 681–706. [Google Scholar] [CrossRef]
- Zhao, X.; Rothstein, R. The Dun1 checkpoint kinase phosphorylates and regulates the ribonucleotide reductase inhibitor Sml1. Proc. Nat. Acad. Sci. USA 2002, 99, 3746–3751. [Google Scholar] [CrossRef]
- Lee, Y.D.; Wang, J.; Stubbe, J.; Elledge, S.J. Dif1 is a DNA-damage-regulated facilitator of nuclear import for ribonucleotide reductase. Mol. Cell 2008, 32, 70–80. [Google Scholar] [CrossRef]
- Huang, M.; Zhou, Z.; Elledge, S.J. The DNA replication and damage checkpoint pathways induce transcription by inhibition of the Crt1 repressor. Cell 1998, 94, 595–605. [Google Scholar] [CrossRef]
- Manfrini, N.; Gobbini, E.; Baldo, V.; Trovesi, C.; Lucchini, G.; Longhese, M.P. G(1)/S and G(2)/M cyclin-dependent kinase activities commit cells to death in the absence of the S-phase checkpoint. Mol. Cell. Biol. 2012, 32, 4971–4985. [Google Scholar] [CrossRef]
- Casper, A.M.; Nghiem, P.; Arlt, M.F.; Glover, T.W. ATR regulates fragile site stability. Cell 2002, 111, 779–789. [Google Scholar] [CrossRef]
- Tanaka, S.; Araki, H. Regulation of the initiation step of DNA replication by cyclin-dependent kinases. Chromosoma 2010, 119, 565–574. [Google Scholar] [CrossRef]
- Zegerman, P.; Diffley, J.F. DNA replication as a target of the DNA damage checkpoint. DNA Repair 2009, 8, 1077–1088. [Google Scholar] [CrossRef]
- Watase, G.; Takisawa, H.; Kanemaki, M.T. Mcm10 plays a role in functioning of the eukaryotic replicative DNA helicase, Cdc45-Mcm-GINS. Curr. Biol. 2012, 22, 343–349. [Google Scholar] [CrossRef]
- Van Deursen, F.; Sengupta, S.; de Piccoli, G.; Sanchez-Diaz, A.; Labib, K. Mcm10 associates with the loaded DNA helicase at replication origins and defines a novel step in its activation. EMBO J. 2012, 31, 2195–2206. [Google Scholar] [CrossRef]
- Kanke, M.; Kodama, Y.; Takahashi, T.S.; Nakagawa, T.; Masukata, H. Mcm10 plays an essential role in origin DNA unwinding after loading of the CMG components. EMBO J. 2012, 31, 2182–2194. [Google Scholar] [CrossRef]
- Gilbert, D.M.; Takebayashi, S.I.; Ryba, T.; Lu, J.; Pope, B.D.; Wilson, K.A.; Hiratani, I. Space and time in the nucleus: Developmental control of replication timing and chromosome architecture. Cold Spring Harbor Symp. 2010, 75, 143–153. [Google Scholar] [CrossRef]
- Barberis, M.; Spiesser, T.W.; Klipp, E. Replication origins and timing of temporal replication in budding yeast: How to solve the conundrum? Curr. Genomics 2010, 11, 199–211. [Google Scholar] [CrossRef]
- Tanaka, S.; Nakato, R.; Katou, Y.; Shirahige, K.; Araki, H. Origin association of Sld3, Sld7, and Cdc45 proteins is a key step for determination of origin-firing timing. Curr. Biol. 2011, 21, 2055–2063. [Google Scholar] [CrossRef]
- Mantiero, D.; Mackenzie, A.; Donaldson, A.; Zegerman, P. Limiting replication initiation factors execute the temporal programme of origin firing in budding yeast. EMBO J. 2011, 30, 4805–4814. [Google Scholar] [CrossRef]
- Zegerman, P.; Diffley, J.F. Checkpoint-dependent inhibition of DNA replication initiation by Sld3 and Dbf4 phosphorylation. Nature 2010, 467, 474–478. [Google Scholar] [CrossRef]
- Lopez-Mosqueda, J.; Maas, N.L.; Jonsson, Z.O.; Defazio-Eli, L.G.; Wohlschlegel, J.; Toczyski, D.P. Damage-induced phosphorylation of Sld3 is important to block late origin firing. Nature 2010, 467, 479–483. [Google Scholar] [CrossRef]
- Karnani, N.; Dutta, A. The effect of the intra-S-phase checkpoint on origins of replication in human cells. Genes Dev. 2011, 25, 621–633. [Google Scholar] [CrossRef]
- Kim, J.M.; Kakusho, N.; Yamada, M.; Kanoh, Y.; Takemoto, N.; Masai, H. Cdc7 kinase mediates Claspin phosphorylation in DNA replication checkpoint. Oncogene 2008, 27, 3475–3482. [Google Scholar] [CrossRef]
- Tenca, P.; Brotherton, D.; Montagnoli, A.; Rainoldi, S.; Albanese, C.; Santocanale, C. Cdc7 is an active kinase in human cancer cells undergoing replication stress. J. Biol. Chem. 2007, 282, 208–215. [Google Scholar]
- Matsumoto, S.; Shimmoto, M.; Kakusho, N.; Yokoyama, M.; Kanoh, Y.; Hayano, M.; Russell, P.; Masai, H. Hsk1 kinase and Cdc45 regulate replication stress-induced checkpoint responses in fission yeast. Cell Cycle 2010, 9, 4627–4637. [Google Scholar] [CrossRef]
- Furuya, K.; Miyabe, I.; Tsutsui, Y.; Paderi, F.; Kakusho, N.; Masai, H.; Niki, H.; Carr, A.M. DDK phosphorylates checkpoint clamp component Rad9 and promotes its release from damaged chromatin. Mol. Cell 2010, 40, 606–618. [Google Scholar] [CrossRef]
- Tsuji, T.; Lau, E.; Chiang, G.G.; Jiang, W. The role of Dbf4/Drf1-dependent kinase Cdc7 in DNA-damage checkpoint control. Mol. Cell 2008, 32, 862–869. [Google Scholar] [CrossRef]
- Jelinsky, S.A.; Samson, L.D. Global response of Saccharomyces cerevisiae to an alkylating agent. Proc. Nat. Acad. Sci. USA 1999, 96, 1486–1491. [Google Scholar] [CrossRef]
- Gasch, A.P.; Huang, M.; Metzner, S.; Botstein, D.; Elledge, S.J.; Brown, P.O. Genomic expression responses to DNA-damaging agents and the regulatory role of the yeast ATR homolog Mec1p. Mol. Biol. Cell 2001, 12, 2987–3003. [Google Scholar] [CrossRef]
- Benton, M.G.; Somasundaram, S.; Glasner, J.D.; Palecek, S.P. Analyzing the dose-dependence of the Saccharomyces cerevisiae global transcriptional response to methyl methanesulfonate and ionizing radiation. BMC Genomics 2006, 7, 305. [Google Scholar]
- Travesa, A.; Kuo, D.; de Bruin, R.A.; Kalashnikova, T.I.; Guaderrama, M.; Thai, K.; Aslanian, A.; Smolka, M.B.; Yates, J.R., 3rd; Ideker, T.; et al. DNA replication stress differentially regulates G1/S genes via Rad53-dependent inactivation of Nrm1. EMBO J. 2012, 31, 1811–1822. [Google Scholar] [CrossRef]
- Bastos de Oliveira, F.M.; Harris, M.R.; Brazauskas, P.; de Bruin, R.A.; Smolka, M.B. Linking DNA replication checkpoint to MBF cell-cycle transcription reveals a distinct class of G1/S genes. EMBO J. 2012, 31, 1798–1810. [Google Scholar] [CrossRef]
- Iyer, V.R.; Horak, C.E.; Scafe, C.S.; Botstein, D.; Snyder, M.; Brown, P.O. Genomic binding sites of the yeast cell-cycle transcription factors SBF and MBF. Nature 2001, 409, 533–538. [Google Scholar] [CrossRef]
- Bean, J.M.; Siggia, E.D.; Cross, F.R. High functional overlap between MluI cell-cycle box binding factor and Swi4/6 cell-cycle box binding factor in the G1/S transcriptional program in Saccharomyces cerevisiae. Genetics 2005, 171, 49–61. [Google Scholar] [CrossRef]
- De Bruin, R.A.; Kalashnikova, T.I.; Chahwan, C.; McDonald, W.H.; Wohlschlegel, J.; Yates, J., 3rd; Russell, P.; Wittenberg, C. Constraining G1-specific transcription to late G1 phase: The MBF-associated corepressor Nrm1 acts via negative feedback. Mol. Cell 2006, 23, 483–496. [Google Scholar] [CrossRef]
- De Bruin, R.A.; Kalashnikova, T.I.; Aslanian, A.; Wohlschlegel, J.; Chahwan, C.; Yates, J.R., 3rd; Russell, P.; Wittenberg, C. DNA replication checkpoint promotes G1-S transcription by inactivating the MBF repressor Nrm1. Proc. Nat. Acad. Sci. USA 2008, 105, 11230–11235. [Google Scholar] [CrossRef]
- Alabert, C.; Bianco, J.N.; Pasero, P. Differential regulation of homologous recombination at DNA breaks and replication forks by the Mrc1 branch of the S-phase checkpoint. EMBO J. 2009, 28, 1131–1141. [Google Scholar] [CrossRef]
- Barlow, J.H.; Rothstein, R. Rad52 recruitment is DNA replication independent and regulated by Cdc28 and the Mec1 kinase. EMBO J. 2009, 28, 1121–1130. [Google Scholar] [CrossRef]
- Meister, P.; Taddei, A.; Vernis, L.; Poidevin, M.; Gasser, S.M.; Baldacci, G. Temporal separation of replication and recombination requires the intra-S checkpoint. J. Cell Biol. 2005, 168, 537–544. [Google Scholar] [CrossRef]
- Sirbu, B.M.; Lachmayer, S.J.; Wulfing, V.; Marten, L.M.; Clarkson, K.E.; Lee, L.W.; Gheorghiu, L.; Zou, L.; Powell, S.N.; Dahm-Daphi, J.; et al. ATR-p53 restricts homologous recombination in response to replicative stress but does not limit DNA interstrand crosslink repair in lung cancer cells. PLoS One 2011, 6, e23053. [Google Scholar] [CrossRef]
- Sorensen, C.S.; Hansen, L.T.; Dziegielewski, J.; Syljuasen, R.G.; Lundin, C.; Bartek, J.; Helleday, T. The cell-cycle checkpoint kinase Chk1 is required for mammalian homologous recombination repair. Nat. Cell Biol. 2005, 7, 195–201. [Google Scholar] [CrossRef]
- Huang, M.; Miao, Z.H.; Zhu, H.; Cai, Y.J.; Lu, W.; Ding, J. Chk1 and Chk2 are differentially involved in homologous recombination repair and cell cycle arrest in response to DNA double-strand breaks induced by camptothecins. Mol. Cancer Ther. 2008, 7, 1440–1449. [Google Scholar] [CrossRef]
- Helt, C.E.; Wang, W.; Keng, P.C.; Bambara, R.A. Evidence that DNA damage detection machinery participates in DNA repair. Cell Cycle 2005, 4, 529–532. [Google Scholar] [CrossRef]
- Paulovich, A.G.; Armour, C.D.; Hartwell, L.H. The Saccharomyces cerevisiae RAD9, RAD17, RAD24 and MEC3 genes are required for tolerating irreparable, ultraviolet-induced DNA damage. Genetics 1998, 150, 75–93. [Google Scholar]
- Kai, M.; Wang, T.S. Checkpoint activation regulates mutagenic translesion synthesis. Genes Dev. 2003, 17, 64–76. [Google Scholar] [CrossRef]
- Karras, G.I.; Fumasoni, M.; Sienski, G.; Vanoli, F.; Branzei, D.; Jentsch, S. Noncanonical role of the 9-1-1 clamp in the error-free DNA damage tolerance pathway. Mol. Cell 2013, 49, 536–546. [Google Scholar] [CrossRef]
- Aylon, Y.; Kupiec, M. The checkpoint protein Rad24 of Saccharomyces cerevisiae is involved in processing double-strand break ends and in recombination partner choice. Mol. Cell. Biol. 2003, 23, 6585–6596. [Google Scholar] [CrossRef]
- Toueille, M.; El-Andaloussi, N.; Frouin, I.; Freire, R.; Funk, D.; Shevelev, I.; Friedrich-Heineken, E.; Villani, G.; Hottiger, M.O.; Hubscher, U. The human Rad9/Rad1/Hus1 damage sensor clamp interacts with DNA polymerase beta and increases its DNA substrate utilisation efficiency: Implications for DNA repair. Nucl. Acid. Res. 2004, 32, 3316–3324. [Google Scholar] [CrossRef]
- Wang, W.; Brandt, P.; Rossi, M.L.; Lindsey-Boltz, L.; Podust, V.; Fanning, E.; Sancar, A.; Bambara, R.A. The human Rad9-Rad1-Hus1 checkpoint complex stimulates flap endonuclease 1. Proc. Nat. Acad. Sci. USA 2004, 101, 16762–16767. [Google Scholar] [CrossRef]
- Balakrishnan, L.; Brandt, P.D.; Lindsey-Boltz, L.A.; Sancar, A.; Bambara, R.A. Long patch base excision repair proceeds via coordinated stimulation of the multienzyme DNA repair complex. J. Biol. Chem. 2009, 284, 15158–15172. [Google Scholar]
- Gembka, A.; Toueille, M.; Smirnova, E.; Poltz, R.; Ferrari, E.; Villani, G.; Hubscher, U. The checkpoint clamp, Rad9-Rad1-Hus1 complex, preferentially stimulates the activity of apurinic/apyrimidinic endonuclease 1 and DNA polymerase beta in long patch base excision repair. Nucl. Acid. Res. 2007, 35, 2596–2608. [Google Scholar] [CrossRef]
- Guan, X.; Madabushi, A.; Chang, D.Y.; Fitzgerald, M.E.; Shi, G.; Drohat, A.C.; Lu, A.L. The human checkpoint sensor Rad9-Rad1-Hus1 interacts with and stimulates DNA repair enzyme TDG glycosylase. Nucl. Acid. Res. 2007, 35, 6207–6218. [Google Scholar] [CrossRef]
- Chang, D.Y.; Lu, A.L. Interaction of checkpoint proteins Hus1/Rad1/Rad9 with DNA base excision repair enzyme MutY homolog in fission yeast, Schizosaccharomyces pombe. J. Biol. Chem. 2005, 280, 408–417. [Google Scholar]
- Kai, M.; Furuya, K.; Paderi, F.; Carr, A.M.; Wang, T.S. Rad3-dependent phosphorylation of the checkpoint clamp regulates repair-pathway choice. Nat. Cell Biol. 2007, 9, 691–697. [Google Scholar] [CrossRef]
- Inagaki, A.; Sleddens-Linkels, E.; van Cappellen, W.A.; Hibbert, R.G.; Sixma, T.K.; Hoeijmakers, J.H.; Grootegoed, J.A.; Baarends, W.M. Human RAD18 interacts with ubiquitylated chromatin components and facilitates RAD9 recruitment to DNA double strand breaks. PLoS One 2011, 6, e23155. [Google Scholar] [CrossRef]
- Segurado, M.; Diffley, J.F. Separate roles for the DNA damage checkpoint protein kinases in stabilizing DNA replication forks. Genes Dev. 2008, 22, 1816–1827. [Google Scholar] [CrossRef]
- De Piccoli, G.; Katou, Y.; Itoh, T.; Nakato, R.; Shirahige, K.; Labib, K. Replisome stability at defective DNA replication forks is independent of S phase checkpoint kinases. Mol. Cell 2012, 45, 696–704. [Google Scholar] [CrossRef]
- Crabbe, L.; Thomas, A.; Pantesco, V.; de Vos, J.; Pasero, P.; Lengronne, A. Analysis of replication profiles reveals key role of RFC-Ctf18 in yeast replication stress response. Nat. Struct. Mol. Biol. 2010, 17, 1391–1397. [Google Scholar] [CrossRef]
- Morin, I.; Ngo, H.P.; Greenall, A.; Zubko, M.K.; Morrice, N.; Lydall, D. Checkpoint-dependent phosphorylation of Exo1 modulates the DNA damage response. EMBO J. 2008, 27, 2400–2410. [Google Scholar] [CrossRef]
- Cotta-Ramusino, C.; Fachinetti, D.; Lucca, C.; Doksani, Y.; Lopes, M.; Sogo, J.; Foiani, M. Exo1 processes stalled replication forks and counteracts fork reversal in checkpoint-defective cells. Mol. Cell 2005, 17, 153–159. [Google Scholar] [CrossRef]
- El-Shemerly, M.; Hess, D.; Pyakurel, A.K.; Moselhy, S.; Ferrari, S. ATR-dependent pathways control hEXO1 stability in response to stalled forks. Nucl. Acid. Res. 2008, 36, 511–519. [Google Scholar]
- Hu, J.; Sun, L.; Shen, F.; Chen, Y.; Hua, Y.; Liu, Y.; Zhang, M.; Hu, Y.; Wang, Q.; Xu, W.; et al. The intra-S phase checkpoint targets Dna2 to prevent stalled replication forks from reversing. Cell 2012, 149, 1221–1232. [Google Scholar] [CrossRef]
- Kai, M.; Boddy, M.N.; Russell, P.; Wang, T.S. Replication checkpoint kinase Cds1 regulates Mus81 to preserve genome integrity during replication stress. Genes Dev. 2005, 19, 919–932. [Google Scholar] [CrossRef]
- Davies, S.L.; North, P.S.; Dart, A.; Lakin, N.D.; Hickson, I.D. Phosphorylation of the Bloom’s syndrome helicase and its role in recovery from S-phase arrest. Mol. Cell. Biol. 2004, 24, 1279–1291. [Google Scholar] [CrossRef]
- Pichierri, P.; Rosselli, F.; Franchitto, A. Werner’s syndrome protein is phosphorylated in an ATR/ATM-dependent manner following replication arrest and DNA damage induced during the S phase of the cell cycle. Oncogene 2003, 22, 1491–1500. [Google Scholar] [CrossRef]
- Davies, S.L.; North, P.S.; Hickson, I.D. Role for BLM in replication-fork restart and suppression of origin firing after replicative stress. Nat. Struct. Mol. Biol. 2007, 14, 677–679. [Google Scholar] [CrossRef]
- Ammazzalorso, F.; Pirzio, L.M.; Bignami, M.; Franchitto, A.; Pichierri, P. ATR and ATM differently regulate WRN to prevent DSBs at stalled replication forks and promote replication fork recovery. EMBO J. 2010, 29, 3156–3169. [Google Scholar] [CrossRef]
- Smolka, M.B.; Albuquerque, C.P.; Chen, S.H.; Zhou, H. Proteome-wide identification of in vivo targets of DNA damage checkpoint kinases. Proc. Nat. Acad. Sci. USA 2007, 104, 10364–10369. [Google Scholar]
- Chen, S.H.; Albuquerque, C.P.; Liang, J.; Suhandynata, R.T.; Zhou, H. A proteome-wide analysis of kinase-substrate network in the DNA damage response. J. Biol. Chem. 2010, 285, 12803–12812. [Google Scholar]
- Matsuoka, S.; Ballif, B.A.; Smogorzewska, A.; McDonald, E.R., 3rd; Hurov, K.E.; Luo, J.; Bakalarski, C.E.; Zhao, Z.; Solimini, N.; Lerenthal, Y.; et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 2007, 316, 1160–1166. [Google Scholar] [CrossRef]
- Mu, J.J.; Wang, Y.; Luo, H.; Leng, M.; Zhang, J.; Yang, T.; Besusso, D.; Jung, S.Y.; Qin, J. A proteomic analysis of ataxia telangiectasia-mutated (ATM)/ATM-Rad3-related (ATR) substrates identifies the ubiquitin-proteasome system as a regulator for DNA damage checkpoints. J. Biol. Chem. 2007, 282, 17330–17334. [Google Scholar]
- Bennetzen, M.V.; Larsen, D.H.; Bunkenborg, J.; Bartek, J.; Lukas, J.; Andersen, J.S. Site-specific phosphorylation dynamics of the nuclear proteome during the DNA damage response. Mol. Cell. Proteomics 2010, 9, 1314–1323. [Google Scholar] [CrossRef]
- Blasius, M.; Forment, J.V.; Thakkar, N.; Wagner, S.A.; Choudhary, C.; Jackson, S.P. A phospho-proteomic screen identifies substrates of the checkpoint kinase Chk1. Genome Biol. 2011, 12, R78. [Google Scholar] [CrossRef]
- Stokes, M.P.; Rush, J.; Macneill, J.; Ren, J.M.; Sprott, K.; Nardone, J.; Yang, V.; Beausoleil, S.A.; Gygi, S.P.; Livingstone, M.; et al. Profiling of UV-induced ATM/ATR signaling pathways. Proc. Nat. Acad. Sci. USA 2007, 104, 19855–19860. [Google Scholar] [CrossRef]
- Brush, G.S.; Morrow, D.M.; Hieter, P.; Kelly, T.J. The ATM homologue MEC1 is required for phosphorylation of replication protein A in yeast. Proc. Nat. Acad. Sci. USA 1996, 93, 15075–15080. [Google Scholar] [CrossRef]
- Liu, S.; Opiyo, S.O.; Manthey, K.; Glanzer, J.G.; Ashley, A.K.; Amerin, C.; Troksa, K.; Shrivastav, M.; Nickoloff, J.A.; Oakley, G.G. Distinct roles for DNA-PK, ATM and ATR in RPA phosphorylation and checkpoint activation in response to replication stress. Nucl. Acid. Res. 2012, 40, 10780–10794. [Google Scholar] [CrossRef]
- Michael, W.M.; Ott, R.; Fanning, E.; Newport, J. Activation of the DNA replication checkpoint through RNA synthesis by primase. Science 2000, 289, 2133–2137. [Google Scholar] [CrossRef]
- Cortez, D.; Glick, G.; Elledge, S.J. Minichromosome maintenance proteins are direct targets of the ATM and ATR checkpoint kinases. Proc. Nat. Acad. Sci. USA 2004, 101, 10078–10083. [Google Scholar] [CrossRef]
- Yoo, H.Y.; Shevchenko, A.; Shevchenko, A.; Dunphy, W.G. Mcm2 is a direct substrate of ATM and ATR during DNA damage and DNA replication checkpoint responses. J. Biol. Chem. 2004, 279, 53353–53364. [Google Scholar] [CrossRef]
- Trenz, K.; Errico, A.; Costanzo, V. Plx1 is required for chromosomal DNA replication under stressful conditions. EMBO J. 2008, 27, 876–885. [Google Scholar] [CrossRef]
- Shi, Y.; Dodson, G.E.; Mukhopadhyay, P.S.; Shanware, N.P.; Trinh, A.T.; Tibbetts, R.S. Identification of carboxyl-terminal MCM3 phosphorylation sites using polyreactive phosphospecific antibodies. J. Biol. Chem. 2007, 282, 9236–9243. [Google Scholar] [CrossRef]
- Ilves, I.; Tamberg, N.; Botchan, M.R. Checkpoint kinase 2 (Chk2) inhibits the activity of the Cdc45/MCM2–7/GINS (CMG) replicative helicase complex. Proc. Nat. Acad. Sci. USA 2012, 109, 13163–13170. [Google Scholar] [CrossRef]
- Ishimi, Y.; Komamura-Kohno, Y.; Kwon, H.J.; Yamada, K.; Nakanishi, M. Identification of MCM4 as a target of the DNA replication block checkpoint system. J. Biol. Chem. 2003, 278, 24644–24650. [Google Scholar]
- Bailis, J.M.; Luche, D.D.; Hunter, T.; Forsburg, S.L. Minichromosome maintenance proteins interact with checkpoint and recombination proteins to promote s-phase genome stability. Mol. Cell. Biol 2008, 28, 1724–1738. [Google Scholar] [CrossRef]
- Brush, G.S.; Kelly, T.J. Phosphorylation of the replication protein A large subunit in the Saccharomyces cerevisiae checkpoint response. Nucl. Acid. Res. 2000, 28, 3725–3732. [Google Scholar] [CrossRef]
- Weinreich, M.; Stillman, B. Cdc7p-Dbf4p kinase binds to chromatin during S phase and is regulated by both the APC and the RAD53 checkpoint pathway. EMBO J. 1999, 18, 5334–5346. [Google Scholar] [CrossRef]
- Morrison, A.J.; Kim, J.A.; Person, M.D.; Highland, J.; Xiao, J.; Wehr, T.S.; Hensley, S.; Bao, Y.; Shen, J.; Collins, S.R.; et al. Mec1/Tel1 phosphorylation of the INO80 chromatin remodeling complex influences DNA damage checkpoint responses. Cell 2007, 130, 499–511. [Google Scholar] [CrossRef]
- Papamichos-Chronakis, M.; Peterson, C.L. The Ino80 chromatin-remodeling enzyme regulates replisome function and stability. Nat. Struct. Mol. Biol. 2008, 15, 338–345. [Google Scholar] [CrossRef]
- Shimada, K.; Oma, Y.; Schleker, T.; Kugou, K.; Ohta, K.; Harata, M.; Gasser, S.M. Ino80 chromatin remodeling complex promotes recovery of stalled replication forks. Curr. Biol. 2008, 18, 566–575. [Google Scholar] [CrossRef]
- Vincent, J.A.; Kwong, T.J.; Tsukiyama, T. ATP-dependent chromatin remodeling shapes the DNA replication landscape. Nat. Struct. Mol. Biol. 2008, 15, 477–484. [Google Scholar] [CrossRef]
- Falbo, K.B.; Alabert, C.; Katou, Y.; Wu, S.; Han, J.; Wehr, T.; Xiao, J.; He, X.; Zhang, Z.; Shi, Y.; et al. Involvement of a chromatin remodeling complex in damage tolerance during DNA replication. Nat. Struct. Mol. Biol. 2009, 16, 1167–1172. [Google Scholar] [CrossRef]
- Shen, X.; Mizuguchi, G.; Hamiche, A.; Wu, C. A chromatin remodelling complex involved in transcription and DNA processing. Nature 2000, 406, 541–544. [Google Scholar] [CrossRef]
- Au, T.J.; Rodriguez, J.; Vincent, J.A.; Tsukiyama, T. ATP-dependent chromatin remodeling factors tune S phase checkpoint activity. Mol. Cell. Biol. 2011, 31, 4454–4463. [Google Scholar] [CrossRef]
- Bermejo, R.; Capra, T.; Jossen, R.; Colosio, A.; Frattini, C.; Carotenuto, W.; Cocito, A.; Doksani, Y.; Klein, H.; Gomez-Gonzalez, B.; et al. The replication checkpoint protects fork stability by releasing transcribed genes from nuclear pores. Cell 2011, 146, 233–246. [Google Scholar] [CrossRef]
- Palancade, B.; Liu, X.; Garcia-Rubio, M.; Aguilera, A.; Zhao, X.; Doye, V. Nucleoporins prevent DNA damage accumulation by modulating Ulp1-dependent sumoylation processes. Mol. Biol. Cell 2007, 18, 2912–2923. [Google Scholar] [CrossRef]
- Cheung, H.C.; San Lucas, F.A.; Hicks, S.; Chang, K.; Bertuch, A.A.; Ribes-Zamora, A. An S/T-Q cluster domain census unveils new putative targets under Tel1/Mec1 control. BMC Genomics 2012, 13, 664. [Google Scholar] [CrossRef]
- Clemenson, C.; Marsolier-Kergoat, M.C. DNA damage checkpoint inactivation: adaptation and recovery. DNA Repair 2009, 8, 1101–1109. [Google Scholar] [CrossRef]
- Heideker, J.; Lis, E.T.; Romesberg, F.E. Phosphatases, DNA damage checkpoints and checkpoint deactivation. Cell. Cycle 2007, 6, 3058–3064. [Google Scholar] [CrossRef]
- Leroy, C.; Lee, S.E.; Vaze, M.B.; Ochsenbein, F.; Guerois, R.; Haber, J.E.; Marsolier-Kergoat, M.C. PP2C phosphatases Ptc2 and Ptc3 are required for DNA checkpoint inactivation after a double-strand break. Mol. Cell 2003, 11, 827–835. [Google Scholar] [CrossRef]
- Keogh, M.C.; Kim, J.A.; Downey, M.; Fillingham, J.; Chowdhury, D.; Harrison, J.C.; Onishi, M.; Datta, N.; Galicia, S.; Emili, A.; et al. A phosphatase complex that dephosphorylates gammaH2AX regulates DNA damage checkpoint recovery. Nature 2006, 439, 497–501. [Google Scholar] [CrossRef]
- Guillemain, G.; Ma, E.; Mauger, S.; Miron, S.; Thai, R.; Guerois, R.; Ochsenbein, F.; Marsolier-Kergoat, M.C. Mechanisms of checkpoint kinase Rad53 inactivation after a double-strand break in Saccharomyces cerevisiae. Mol. Cell. Biol. 2007, 27, 3378–3389. [Google Scholar] [CrossRef]
- O'Neill, B.M.; Szyjka, S.J.; Lis, E.T.; Bailey, A.O.; Yates, J.R., 3rd; Aparicio, O.M.; Romesberg, F.E. Pph3-Psy2 is a phosphatase complex required for Rad53 dephosphorylation and replication fork restart during recovery from DNA damage. Proc. Nat. Acad. Sci. USA 2007, 104, 9290–9295. [Google Scholar] [CrossRef]
- Szyjka, S.J.; Aparicio, J.G.; Viggiani, C.J.; Knott, S.; Xu, W.; Tavare, S.; Aparicio, O.M. Rad53 regulates replication fork restart after DNA damage in Saccharomyces cerevisiae. Genes Dev. 2008, 22, 1906–1920. [Google Scholar] [CrossRef]
- Bazzi, M.; Mantiero, D.; Trovesi, C.; Lucchini, G.; Longhese, M.P. Dephosphorylation of gamma H2A by Glc7/protein phosphatase 1 promotes recovery from inhibition of DNA replication. Mol. Cell. Biol. 2010, 30, 131–145. [Google Scholar] [CrossRef]
- den Elzen, N.R.; O'Connell, M.J. Recovery from DNA damage checkpoint arrest by PP1-mediated inhibition of Chk1. EMBO J. 2004, 23, 908–918. [Google Scholar] [CrossRef]
- Freeman, A.K.; Monteiro, A.N. Phosphatases in the cellular response to DNA damage. Cell Commun. Signal. 2010, 8, 27. [Google Scholar] [CrossRef]
- Shimada, M.; Nakanishi, M. Response to DNA damage: Why do we need to focus on protein phosphatases? Front. Oncol. 2013, 3, 8. [Google Scholar]
- Goodarzi, A.A.; Jonnalagadda, J.C.; Douglas, P.; Young, D.; Ye, R.; Moorhead, G.B.; Lees-Miller, S.P.; Khanna, K.K. Autophosphorylation of ataxia-telangiectasia mutated is regulated by protein phosphatase 2A. EMBO J. 2004, 23, 4451–4461. [Google Scholar] [CrossRef]
- Shreeram, S.; Demidov, O.N.; Hee, W.K.; Yamaguchi, H.; Onishi, N.; Kek, C.; Timofeev, O.N.; Dudgeon, C.; Fornace, A.J.; Anderson, C.W.; et al. Wip1 phosphatase modulates ATM-dependent signaling pathways. Mol. Cell 2006, 23, 757–764. [Google Scholar] [CrossRef]
- Lu, X.; Nannenga, B.; Donehower, L.A. PPM1D dephosphorylates Chk1 and p53 and abrogates cell cycle checkpoints. Genes Dev. 2005, 19, 1162–1174. [Google Scholar] [CrossRef]
- Leung-Pineda, V.; Ryan, C.E.; Piwnica-Worms, H. Phosphorylation of Chk1 by ATR is antagonized by a Chk1-regulated protein phosphatase 2A circuit. Mol. Cell. Biol. 2006, 26, 7529–7538. [Google Scholar] [CrossRef]
- Liang, X.; Reed, E.; Yu, J.J. Protein phosphatase 2A interacts with Chk2 and regulates phosphorylation at Thr-68 after cisplatin treatment of human ovarian cancer cells. Int. J. Mol. Med. 2006, 17, 703–708. [Google Scholar] [Green Version]
- Fujimoto, H.; Onishi, N.; Kato, N.; Takekawa, M.; Xu, X.Z.; Kosugi, A.; Kondo, T.; Imamura, M.; Oishi, I.; Yoda, A.; et al. Regulation of the antioncogenic Chk2 kinase by the oncogenic Wip1 phosphatase. Cell. Death Differ. 2006, 13, 1170–1180. [Google Scholar] [CrossRef]
- Oliva-Trastoy, M.; Berthonaud, V.; Chevalier, A.; Ducrot, C.; Marsolier-Kergoat, M.C.; Mann, C.; Leteurtre, F. The Wip1 phosphatase (PPM1D) antagonizes activation of the Chk2 tumour suppressor kinase. Oncogene 2007, 26, 1449–1458. [Google Scholar] [CrossRef]
- Macurek, L.; Benada, J.; Mullers, E.; Halim, V.A.; Krejcikova, K.; Burdova, K.; Pechackova, S.; Hodny, Z.; Lindqvist, A.; Medema, R.H.; et al. Downregulation of Wip1 phosphatase modulates the cellular threshold of DNA damage signaling in mitosis. Cell Cycle 2013, 12, 251–262. [Google Scholar] [CrossRef]
- Zhang, Y.W.; Brognard, J.; Coughlin, C.; You, Z.; Dolled-Filhart, M.; Aslanian, A.; Manning, G.; Abraham, R.T.; Hunter, T. The F box protein Fbx6 regulates Chk1 stability and cellular sensitivity to replication stress. Mol. Cell 2009, 35, 442–453. [Google Scholar] [CrossRef]
- Leung-Pineda, V.; Huh, J.; Piwnica-Worms, H. DDB1 targets Chk1 to the Cul4 E3 ligase complex in normal cycling cells and in cells experiencing replication stress. Cancer Res. 2009, 69, 2630–2637. [Google Scholar] [CrossRef]
- Mamely, I.; van Vugt, M.A.; Smits, V.A.; Semple, J.I.; Lemmens, B.; Perrakis, A.; Medema, R.H.; Freire, R. Polo-like kinase-1 controls proteasome-dependent degradation of Claspin during checkpoint recovery. Curr. Biol. 2006, 16, 1950–1955. [Google Scholar] [CrossRef]
- Peschiaroli, A.; Dorrello, N.V.; Guardavaccaro, D.; Venere, M.; Halazonetis, T.; Sherman, N.E.; Pagano, M. SCFbetaTrCP-mediated degradation of Claspin regulates recovery from the DNA replication checkpoint response. Mol. Cell 2006, 23, 319–329. [Google Scholar] [CrossRef]
- Mailand, N.; Bekker-Jensen, S.; Bartek, J.; Lukas, J. Destruction of Claspin by SCFbetaTrCP restrains Chk1 activation and facilitates recovery from genotoxic stress. Mol. Cell 2006, 23, 307–318. [Google Scholar] [CrossRef]
- Faustrup, H.; Bekker-Jensen, S.; Bartek, J.; Lukas, J.; Mailand, N. USP7 counteracts SCFbetaTrCP- but not APCCdh1-mediated proteolysis of Claspin. J. Cell. Biol. 2009, 184, 13–19. [Google Scholar] [CrossRef]
- Macurek, L.; Lindqvist, A.; Lim, D.; Lampson, M.A.; Klompmaker, R.; Freire, R.; Clouin, C.; Taylor, S.S.; Yaffe, M.B.; Medema, R.H. Polo-like kinase-1 is activated by aurora A to promote checkpoint recovery. Nature 2008, 455, 119–123. [Google Scholar] [CrossRef]
- Peng, A.; Wang, L.; Fisher, L.A. Greatwall and Polo-like kinase 1 coordinate to promote checkpoint recovery. J. Biol. Chem. 2011, 286, 28996–29004. [Google Scholar] [CrossRef]
- Toczyski, D.P.; Galgoczy, D.J.; Hartwell, L.H. CDC5 and CKII control adaptation to the yeast DNA damage checkpoint. Cell 1997, 90, 1097–1106. [Google Scholar] [CrossRef]
- Vidanes, G.M.; Sweeney, F.D.; Galicia, S.; Cheung, S.; Doyle, J.P.; Durocher, D.; Toczyski, D.P. CDC5 inhibits the hyperphosphorylation of the checkpoint kinase Rad53, leading to checkpoint adaptation. PLoS Biol. 2010, 8, e1000286. [Google Scholar] [CrossRef]
- Donnianni, R.A.; Ferrari, M.; Lazzaro, F.; Clerici, M.; Tamilselvan Nachimuthu, B.; Plevani, P.; Muzi-Falconi, M.; Pellicioli, A. Elevated levels of the polo kinase Cdc5 override the Mec1/ATR checkpoint in budding yeast by acting at different steps of the signaling pathway. PLoS Genet. 2010, 6, e1000763. [Google Scholar] [CrossRef]
- Song, B.; Liu, X.S.; Davis, K.; Liu, X. Plk1 phosphorylation of Orc2 promotes DNA replication under conditions of stress. Mol. Cell. Biol. 2011, 31, 4844–4856. [Google Scholar] [CrossRef]
- Veaute, X.; Jeusset, J.; Soustelle, C.; Kowalczykowski, S.C.; Le Cam, E.; Fabre, F. The Srs2 helicase prevents recombination by disrupting Rad51 nucleoprotein filaments. Nature 2003, 423, 309–312. [Google Scholar] [CrossRef]
- Krejci, L.; van Komen, S.; Li, Y.; Villemain, J.; Reddy, M.S.; Klein, H.; Ellenberger, T.; Sung, P. DNA helicase Srs2 disrupts the Rad51 presynaptic filament. Nature 2003, 423, 305–309. [Google Scholar] [CrossRef]
- Vaze, M.B.; Pellicioli, A.; Lee, S.E.; Ira, G.; Liberi, G.; Arbel-Eden, A.; Foiani, M.; Haber, J.E. Recovery from checkpoint-mediated arrest after repair of a double-strand break requires Srs2 helicase. Mol. Cell 2002, 10, 373–385. [Google Scholar] [CrossRef]
- Jain, S.; Sugawara, N.; Lydeard, J.; Vaze, M.; Tanguy Le Gac, N.; Haber, J.E. A recombination execution checkpoint regulates the choice of homologous recombination pathway during DNA double-strand break repair. Genes Dev. 2009, 23, 291–303. [Google Scholar] [CrossRef]
- Yeung, M.; Durocher, D. Srs2 enables checkpoint recovery by promoting disassembly of DNA damage foci from chromatin. DNA Repair 2011, 10, 1213–1222. [Google Scholar]
- McKee, A.H.; Kleckner, N. A general method for identifying recessive diploid-specific mutations in Saccharomyces cerevisiae, its application to the isolation of mutants blocked at intermediate stages of meiotic prophase and characterization of a new gene SAE2. Genetics 1997, 146, 797–816. [Google Scholar]
- Prinz, S.; Amon, A.; Klein, F. Isolation of COM1, a new gene required to complete meiotic double-strand break-induced recombination in Saccharomyces cerevisiae. Genetics 1997, 146, 781–795. [Google Scholar]
- Rattray, A.J.; McGill, C.B.; Shafer, B.K.; Strathern, J.N. Fidelity of mitotic double-strand-break repair in Saccharomyces cerevisiae: a role for SAE2/COM1. Genetics 2001, 158, 109–122. [Google Scholar]
- Clerici, M.; Mantiero, D.; Lucchini, G.; Longhese, M.P. The Saccharomyces cerevisiae Sae2 protein promotes resection and bridging of double strand break ends. J. Biol. Chem. 2005, 280, 38631–38638. [Google Scholar] [CrossRef]
- Clerici, M.; Mantiero, D.; Lucchini, G.; Longhese, M.P. The Saccharomyces cerevisiae Sae2 protein negatively regulates DNA damage checkpoint signalling. EMBO Rep. 2006, 7, 212–218. [Google Scholar] [CrossRef]
- Kim, H.S.; Vijayakumar, S.; Reger, M.; Harrison, J.C.; Haber, J.E.; Weil, C.; Petrini, J.H. Functional interactions between Sae2 and the Mre11 complex. Genetics 2008, 178, 711–723. [Google Scholar] [CrossRef]
- Guenole, A.; Srivas, R.; Vreeken, K.; Wang, Z.Z.; Wang, S.; Krogan, N.J.; Ideker, T.; van Attikum, H. Dissection of DNA damage responses using multiconditional genetic interaction maps. Mol. Cell 2013, 49, 346–358. [Google Scholar] [CrossRef]
- Javaheri, A.; Wysocki, R.; Jobin-Robitaille, O.; Altaf, M.; Cote, J.; Kron, S.J. Yeast G1 DNA damage checkpoint regulation by H2A phosphorylation is independent of chromatin remodeling. Proc. Nat. Acad. Sci USA 2006, 103, 13771–13776. [Google Scholar]
- Ohouo, P.Y.; Bastos de Oliveira, F.M.; Almeida, B.S.; Smolka, M.B. DNA damage signaling recruits the Rtt107-Slx4 scaffolds via Dpb11 to mediate replication stress response. Mol. Cell 2010, 39, 300–306. [Google Scholar] [CrossRef]
- Li, X.; Liu, K.; Li, F.; Wang, J.; Huang, H.; Wu, J.; Shi, Y. Structure of C-terminal tandem BRCT repeats of Rtt107 protein reveals critical role in interaction with phosphorylated histone H2A during DNA damage repair. J. Biol. Chem. 2012, 287, 9137–9146. [Google Scholar] [CrossRef]
- Ohouo, P.Y.; Bastos de Oliveira, F.M.; Liu, Y.; Ma, C.J.; Smolka, M.B. DNA-repair scaffolds dampen checkpoint signalling by counteracting the adaptor Rad9. Nature 2013, 493, 120–124. [Google Scholar]
- Mimitou, E.P.; Symington, L.S. DNA end resection—Unraveling the tail. DNA Repair 2011, 10, 344–348. [Google Scholar] [CrossRef]
- Costelloe, T.; Louge, R.; Tomimatsu, N.; Mukherjee, B.; Martini, E.; Khadaroo, B.; Dubois, K.; Wiegant, W.W.; Thierry, A.; Burma, S.; et al. The yeast Fun30 and human SMARCAD1 chromatin remodellers promote DNA end resection. Nature 2012, 489, 581–584. [Google Scholar] [CrossRef]
- Chen, X.; Cui, D.; Papusha, A.; Zhang, X.; Chu, C.D.; Tang, J.; Chen, K.; Pan, X.; Ira, G. The Fun30 nucleosome remodeller promotes resection of DNA double-strand break ends. Nature 2012, 489, 576–580. [Google Scholar] [CrossRef]
- Eapen, V.V.; Sugawara, N.; Tsabar, M.; Wu, W.H.; Haber, J.E. The Saccharomyces cerevisiae chromatin remodeler Fun30 regulates DNA end resection and checkpoint deactivation. Mol. Cell. Biol. 2012, 32, 4727–4740. [Google Scholar] [CrossRef]
- Peterson, S.E.; Li, Y.; Chait, B.T.; Gottesman, M.E.; Baer, R.; Gautier, J. Cdk1 uncouples CtIP-dependent resection and Rad51 filament formation during M-phase double-strand break repair. J. Cell. Biol. 2011, 194, 705–720. [Google Scholar] [CrossRef]
- Hill, R.; Lee, P.W. The DNA-dependent protein kinase (DNA-PK): More than just a case of making ends meet? Cell Cycle 2010, 9, 3460–3469. [Google Scholar] [CrossRef]
- Trenz, K.; Smith, E.; Smith, S.; Costanzo, V. ATM and ATR promote Mre11 dependent restart of collapsed replication forks and prevent accumulation of DNA breaks. EMBO J. 2006, 25, 1764–1774. [Google Scholar] [CrossRef]
- Doksani, Y.; Bermejo, R.; Fiorani, S.; Haber, J.E.; Foiani, M. Replicon dynamics, dormant origin firing, and terminal fork integrity after double-strand break formation. Cell 2009, 137, 247–258. [Google Scholar] [CrossRef]
- Stiff, T.; Walker, S.A.; Cerosaletti, K.; Goodarzi, A.A.; Petermann, E.; Concannon, P.; O'Driscoll, M.; Jeggo, P.A. ATR-dependent phosphorylation and activation of ATM in response to UV treatment or replication fork stalling. EMBO J. 2006, 25, 5775–5782. [Google Scholar] [CrossRef]
- Baldo, V.; Testoni, V.; Lucchini, G.; Longhese, M.P. Dominant TEL1-hy mutations compensate for Mec1 lack of functions in the DNA damage response. Mol. Cell. Biol. 2008, 28, 358–375. [Google Scholar] [CrossRef]
- D'Amours, D.; Jackson, S.P. The yeast Xrs2 complex functions in S phase checkpoint regulation. Genes Dev. 2001, 15, 2238–2249. [Google Scholar] [CrossRef]
- Lee, J.; Dunphy, W.G. The Mre11-Rad50-Nbs1 (MRN) complex has a specific role in the activation of Chk1 in response to stalled replication forks. Mol. Biol. Cell 2013, 24, 1343–1353. [Google Scholar] [CrossRef]
- Tittel-Elmer, M.; Alabert, C.; Pasero, P.; Cobb, J.A. The MRX complex stabilizes the replisome independently of the S phase checkpoint during replication stress. EMBO J. 2009, 28, 1142–1156. [Google Scholar] [CrossRef]
- Tittel-Elmer, M.; Lengronne, A.; Davidson, M.B.; Bacal, J.; Francois, P.; Hohl, M.; Petrini, J.H.; Pasero, P.; Cobb, J.A. Cohesin association to replication sites depends on rad50 and promotes fork restart. Mol. Cell 2012, 48, 98–108. [Google Scholar] [CrossRef]
- Watrin, E.; Peters, J.M. The cohesin complex is required for the DNA damage-induced G2/M checkpoint in mammalian cells. EMBO J. 2009, 28, 2625–2635. [Google Scholar] [CrossRef]
- Subramanian, L.; Nakamura, T.M. A kinase-independent role for the Rad3(ATR)-Rad26(ATRIP) complex in recruitment of Tel1(ATM) to telomeres in fission yeast. PLoS Genet. 2010, 6, e1000839. [Google Scholar] [CrossRef]
- Gorgoulis, V.G.; Vassiliou, L.V.; Karakaidos, P.; Zacharatos, P.; Kotsinas, A.; Liloglou, T.; Venere, M.; Ditullio, R.A., Jr.; Kastrinakis, N.G.; Levy, B.; et al. Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 2005, 434, 907–913. [Google Scholar] [CrossRef]
- Bartkova, J.; Horejsi, Z.; Koed, K.; Kramer, A.; Tort, F.; Zieger, K.; Guldberg, P.; Sehested, M.; Nesland, J.M.; Lukas, C.; et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 2005, 434, 864–870. [Google Scholar] [CrossRef]
- Bartkova, J.; Rezaei, N.; Liontos, M.; Karakaidos, P.; Kletsas, D.; Issaeva, N.; Vassiliou, L.V.; Kolettas, E.; Niforou, K.; Zoumpourlis, V.C.; et al. Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature 2006, 444, 633–637. [Google Scholar] [CrossRef]
- Di Micco, R.; Fumagalli, M.; Cicalese, A.; Piccinin, S.; Gasparini, P.; Luise, C.; Schurra, C.; Garre, M.; Nuciforo, P.G.; Bensimon, A.; et al. Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature 2006, 444, 638–642. [Google Scholar] [CrossRef]
- Kastan, M.B.; Bartek, J. Cell-cycle checkpoints and cancer. Nature 2004, 432, 316–323. [Google Scholar] [CrossRef]
- Sidi, S.; Sanda, T.; Kennedy, R.D.; Hagen, A.T.; Jette, C.A.; Hoffmans, R.; Pascual, J.; Imamura, S.; Kishi, S.; Amatruda, J.F.; et al. Chk1 suppresses a caspase-2 apoptotic response to DNA damage that bypasses p53, Bcl-2, and caspase-3. Cell 2008, 133, 864–877. [Google Scholar] [CrossRef]
- Myers, K.; Gagou, M.E.; Zuazua-Villar, P.; Rodriguez, R.; Meuth, M. ATR and Chk1 suppress a caspase-3-dependent apoptotic response following DNA replication stress. PLoS Genet. 2009, 5, e1000324. [Google Scholar] [CrossRef]
- Murga, M.; Bunting, S.; Montana, M.F.; Soria, R.; Mulero, F.; Canamero, M.; Lee, Y.; McKinnon, P.J.; Nussenzweig, A.; Fernandez-Capetillo, O. A mouse model of ATR-Seckel shows embryonic replicative stress and accelerated aging. Nat. Genet. 2009, 41, 891–898. [Google Scholar] [CrossRef]
- Toledo, L.I.; Murga, M.; Zur, R.; Soria, R.; Rodriguez, A.; Martinez, S.; Oyarzabal, J.; Pastor, J.; Bischoff, J.R.; Fernandez-Capetillo, O. A cell-based screen identifies ATR inhibitors with synthetic lethal properties for cancer-associated mutations. Nat. Struct. Mol. Biol. 2011, 18, 721–727. [Google Scholar] [CrossRef]
- Koniaras, K.; Cuddihy, A.R.; Christopoulos, H.; Hogg, A.; O'Connell, M.J. Inhibition of Chk1-dependent G2 DNA damage checkpoint radiosensitizes p53 mutant human cells. Oncogene 2001, 20, 7453–7463. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Hustedt, N.; Gasser, S.M.; Shimada, K. Replication Checkpoint: Tuning and Coordination of Replication Forks in S Phase. Genes 2013, 4, 388-434. https://doi.org/10.3390/genes4030388
Hustedt N, Gasser SM, Shimada K. Replication Checkpoint: Tuning and Coordination of Replication Forks in S Phase. Genes. 2013; 4(3):388-434. https://doi.org/10.3390/genes4030388
Chicago/Turabian StyleHustedt, Nicole, Susan M. Gasser, and Kenji Shimada. 2013. "Replication Checkpoint: Tuning and Coordination of Replication Forks in S Phase" Genes 4, no. 3: 388-434. https://doi.org/10.3390/genes4030388
APA StyleHustedt, N., Gasser, S. M., & Shimada, K. (2013). Replication Checkpoint: Tuning and Coordination of Replication Forks in S Phase. Genes, 4(3), 388-434. https://doi.org/10.3390/genes4030388




