A Recipe to Evolve Complex Life Chemically on Earth
Highlights
- tRNA evolved in pre-life according to the three 31 nucleotide minihelix evolution theorem.
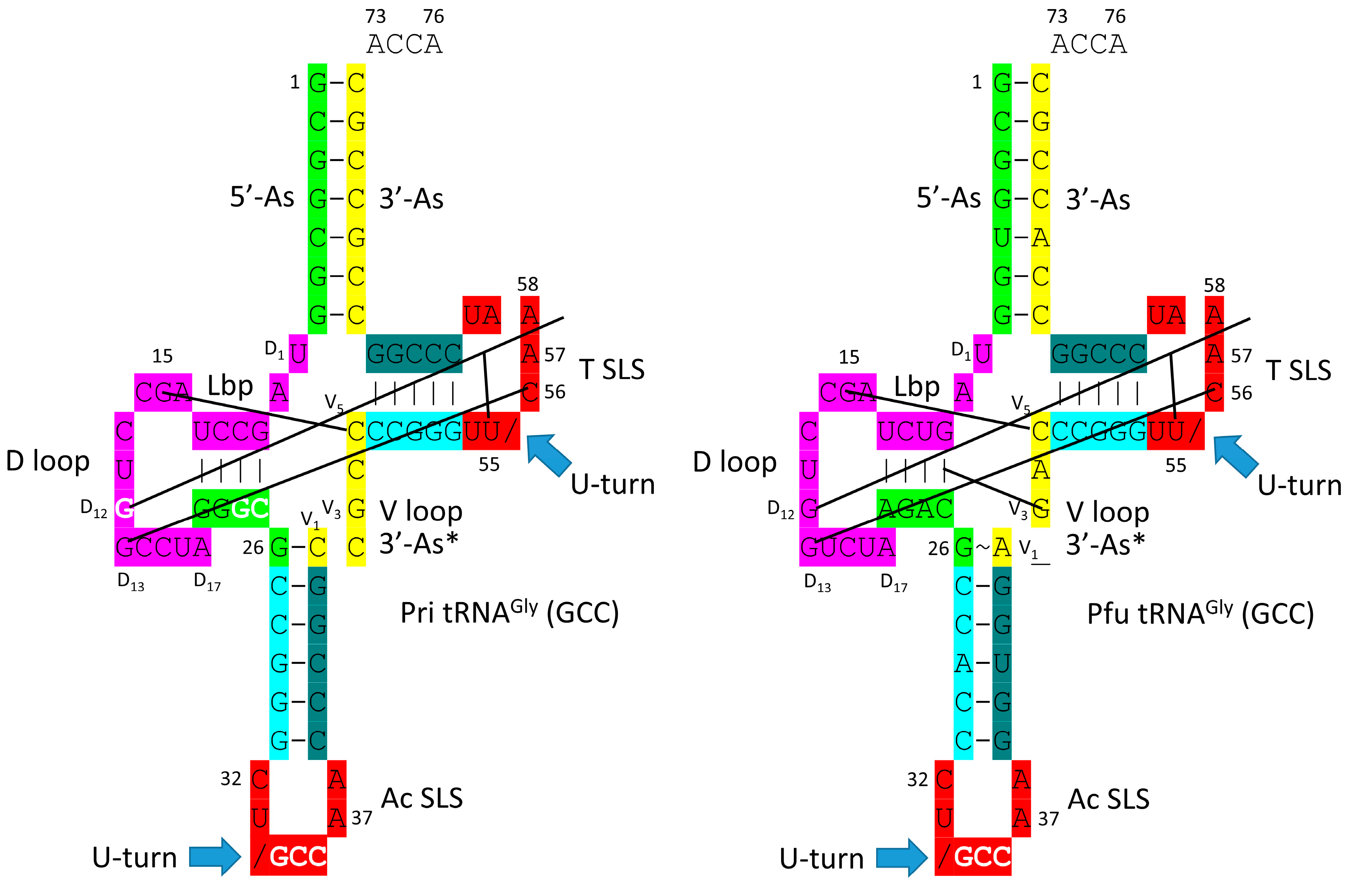
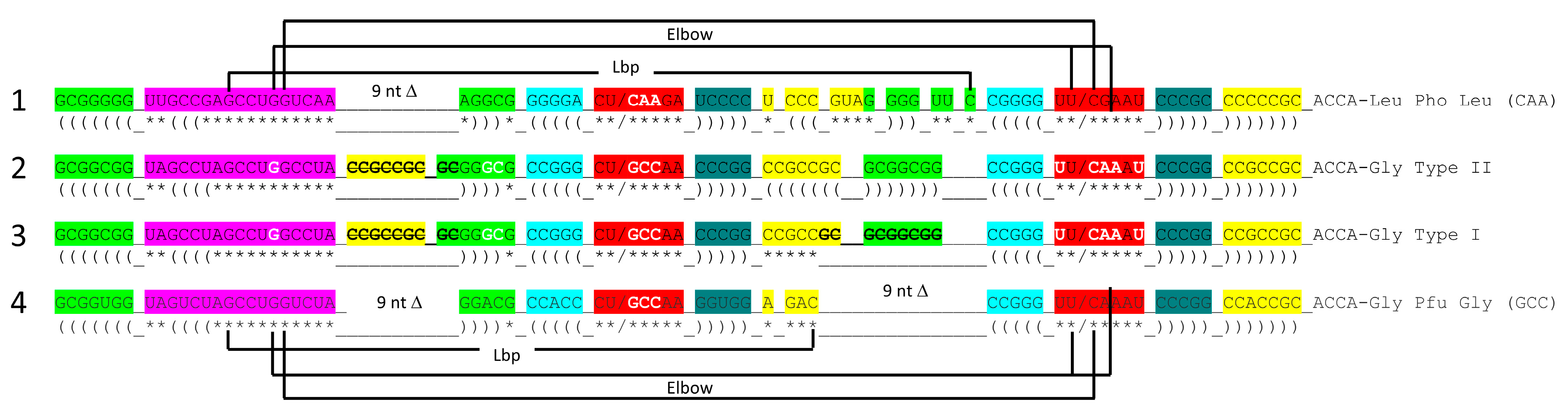
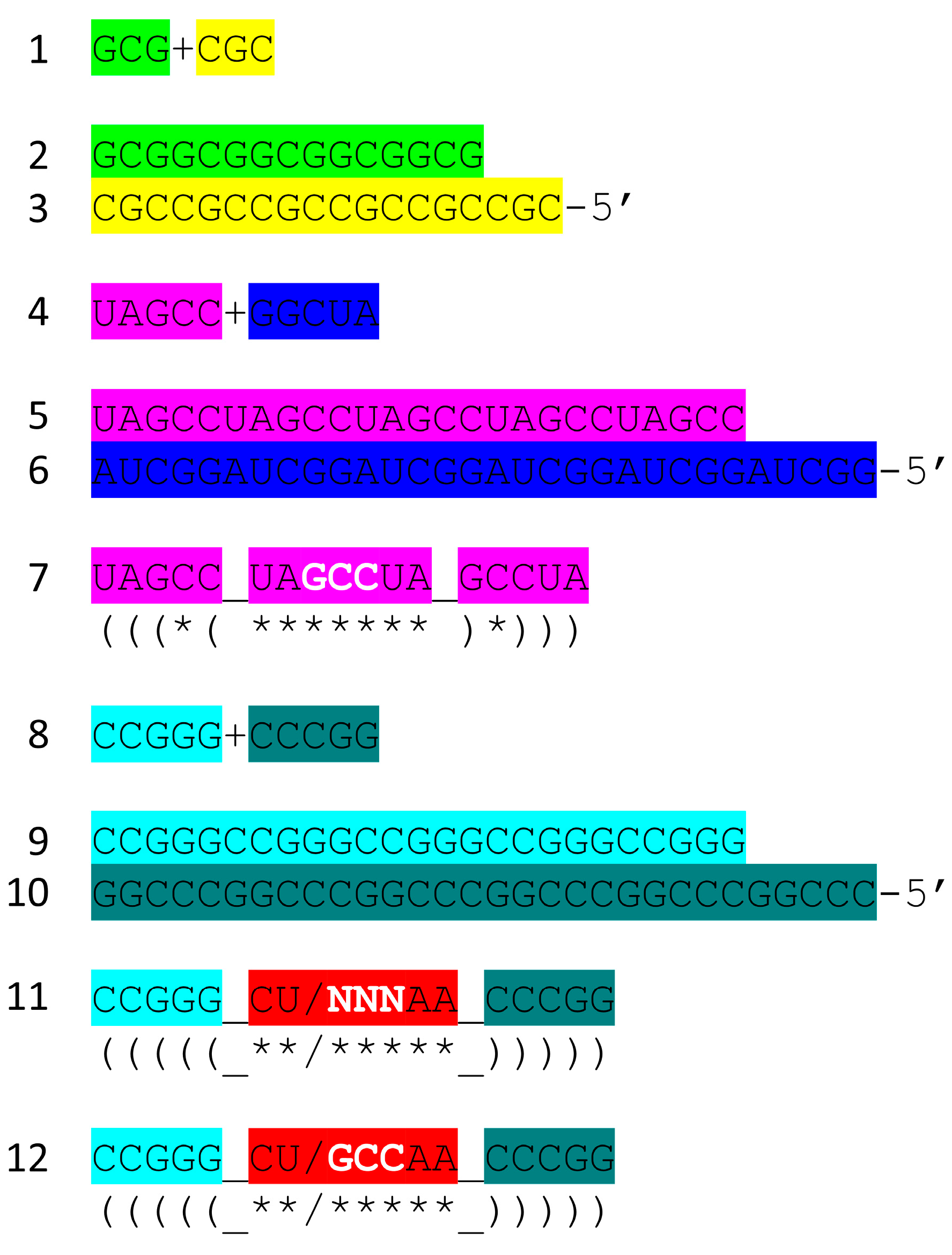
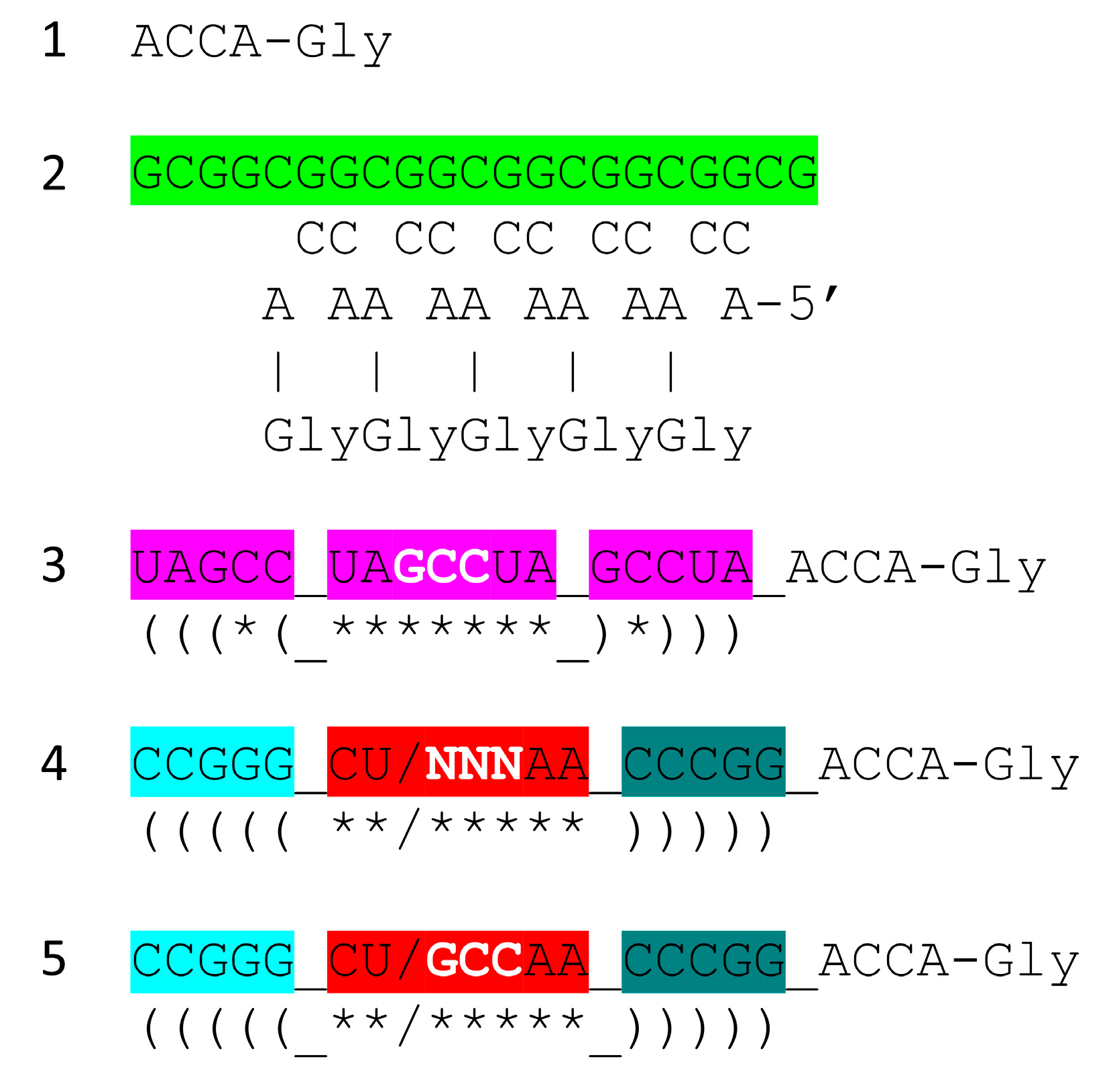
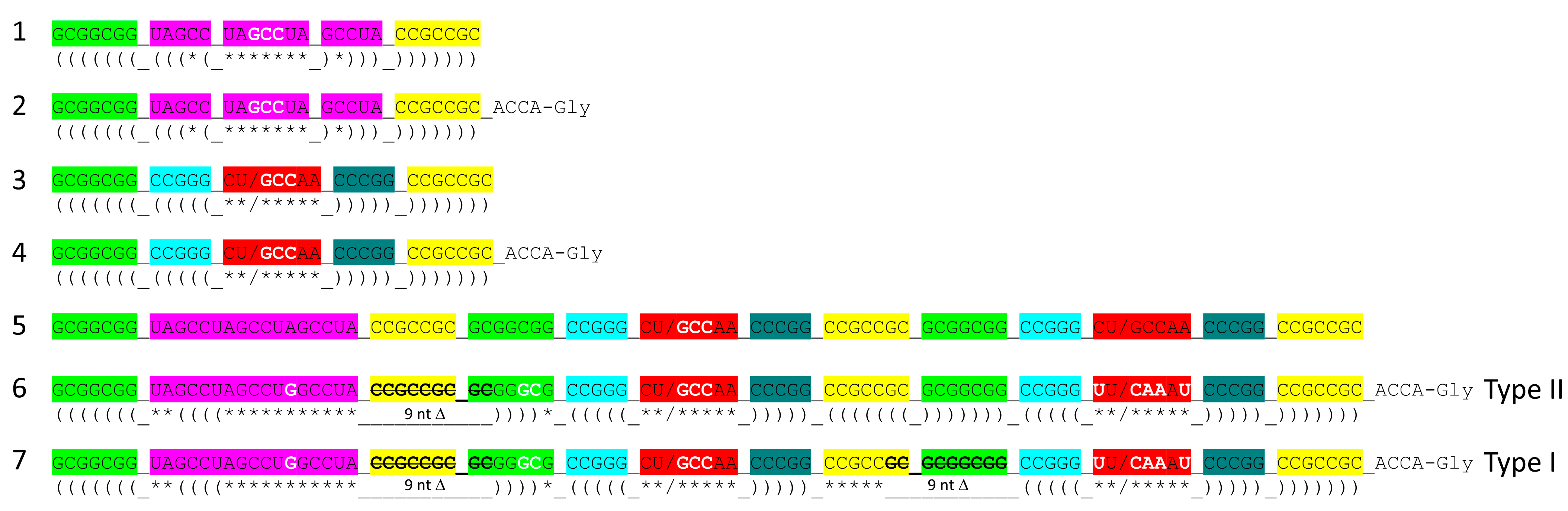
- tRNA evolved from RNA repeats (GCG, CGC and UAGCC) and inverted repeats (~CCGGG_CU/BNNAA_CCCGG) (_ separates stems and loop; / indicates the U-turn) and 3’-ACCA-Gly.
- The component parts of tRNA can be generated in a laboratory and assembled using pre-life reactants.
- Complex life such as microbial life and LUCA is only possible with a genetic code, and tRNA evolution is the core story in the evolution of this code.
Abstract
1. Introduction
2. Materials and Methods
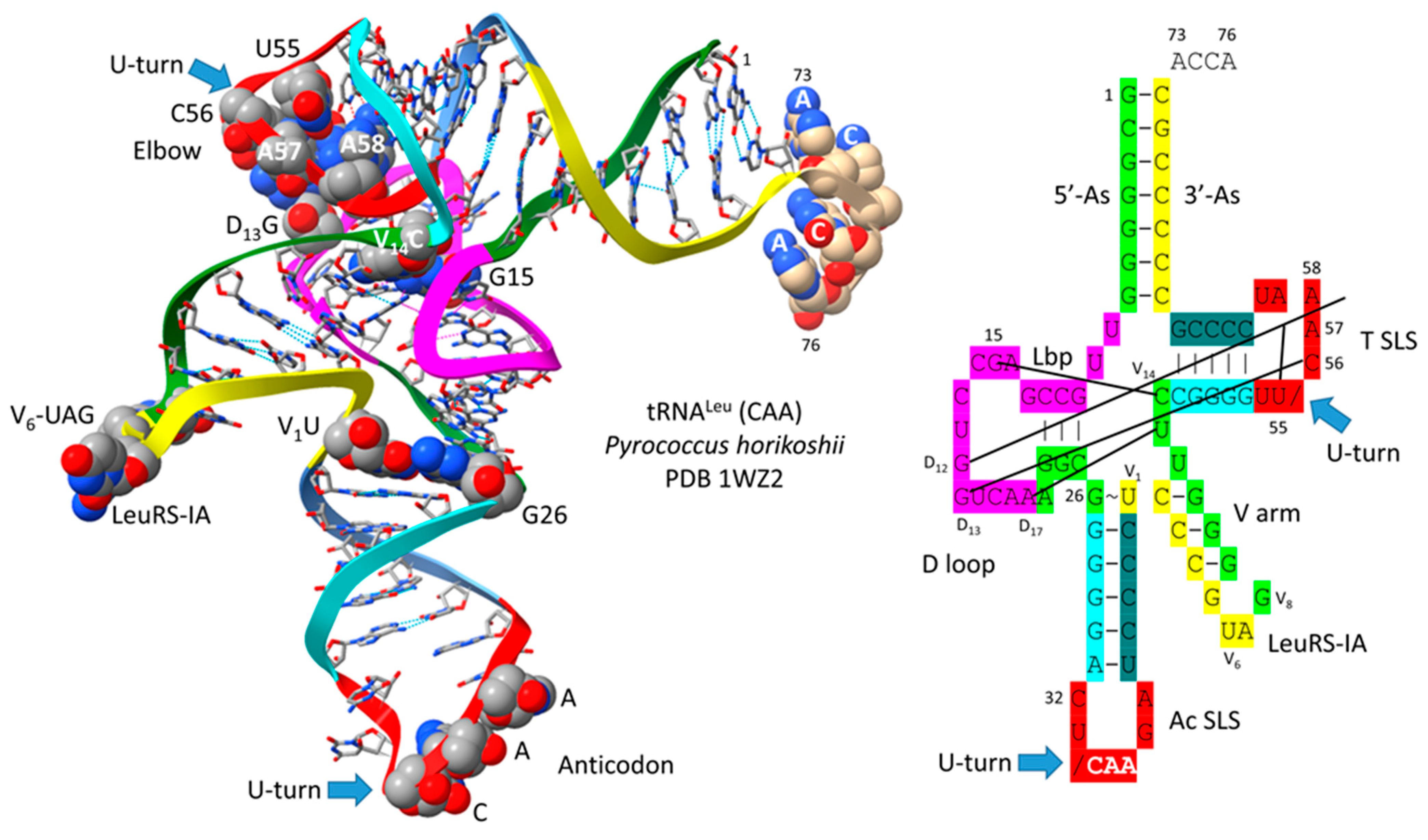
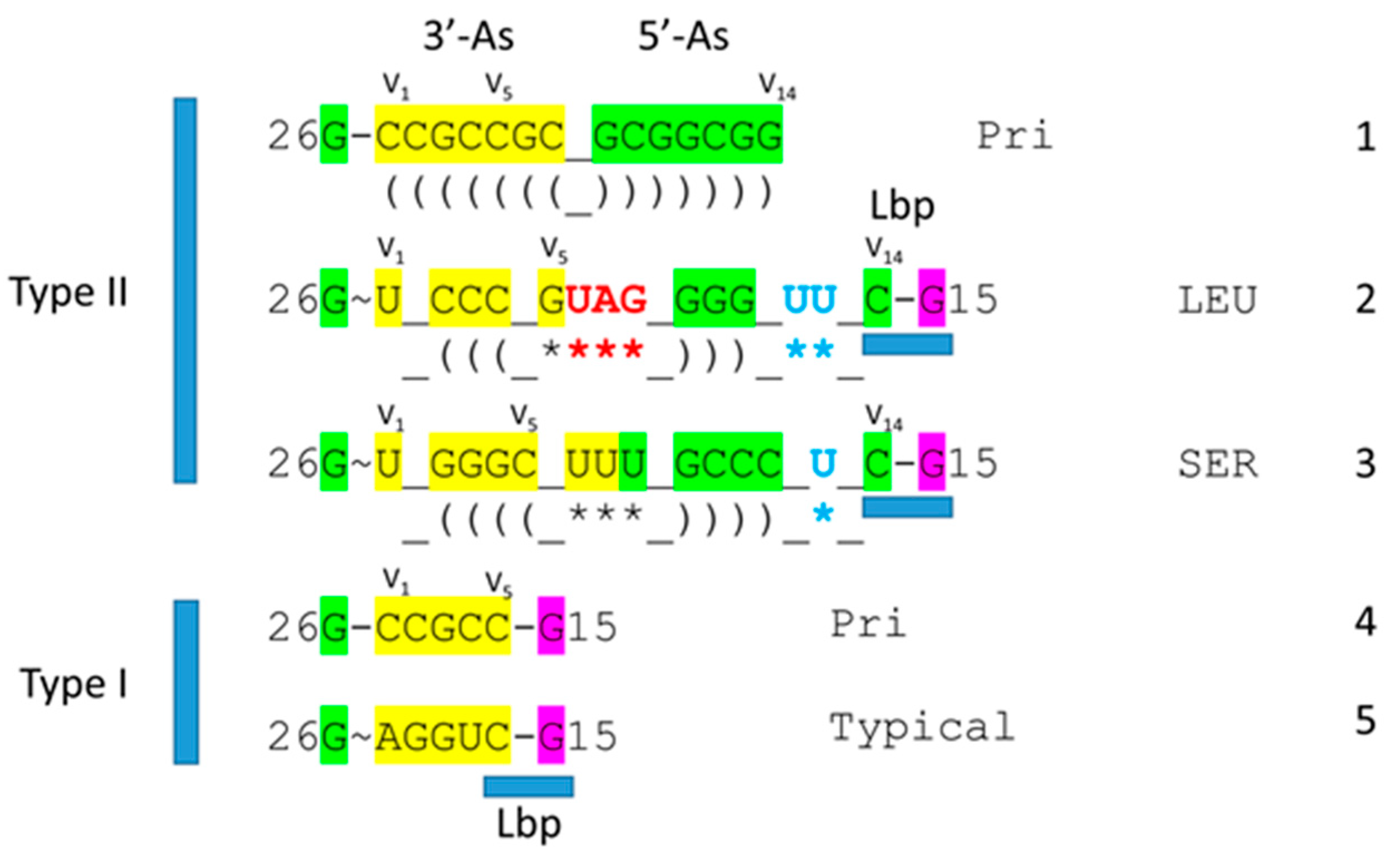
3. Type II tRNA
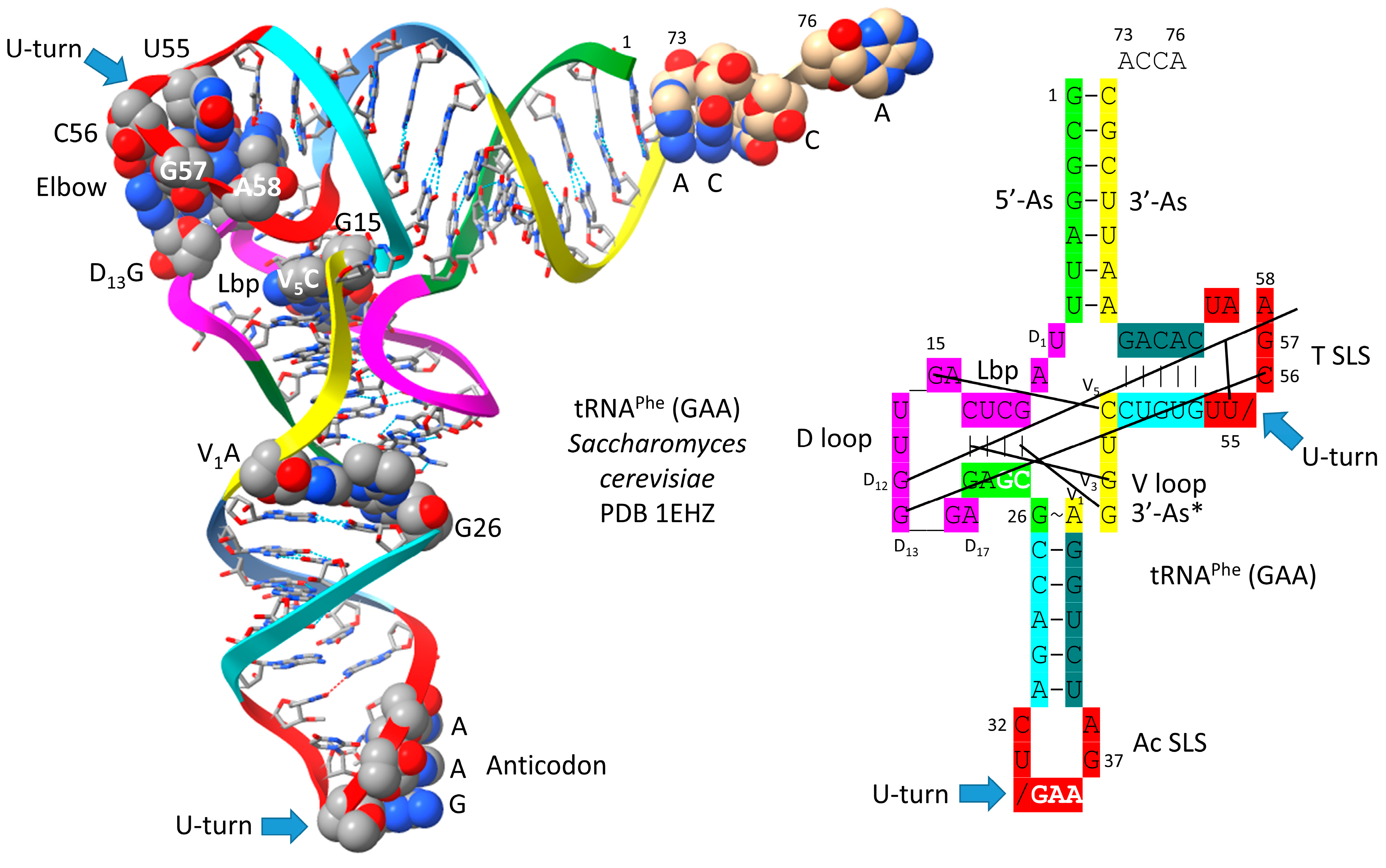
4. Type I tRNA
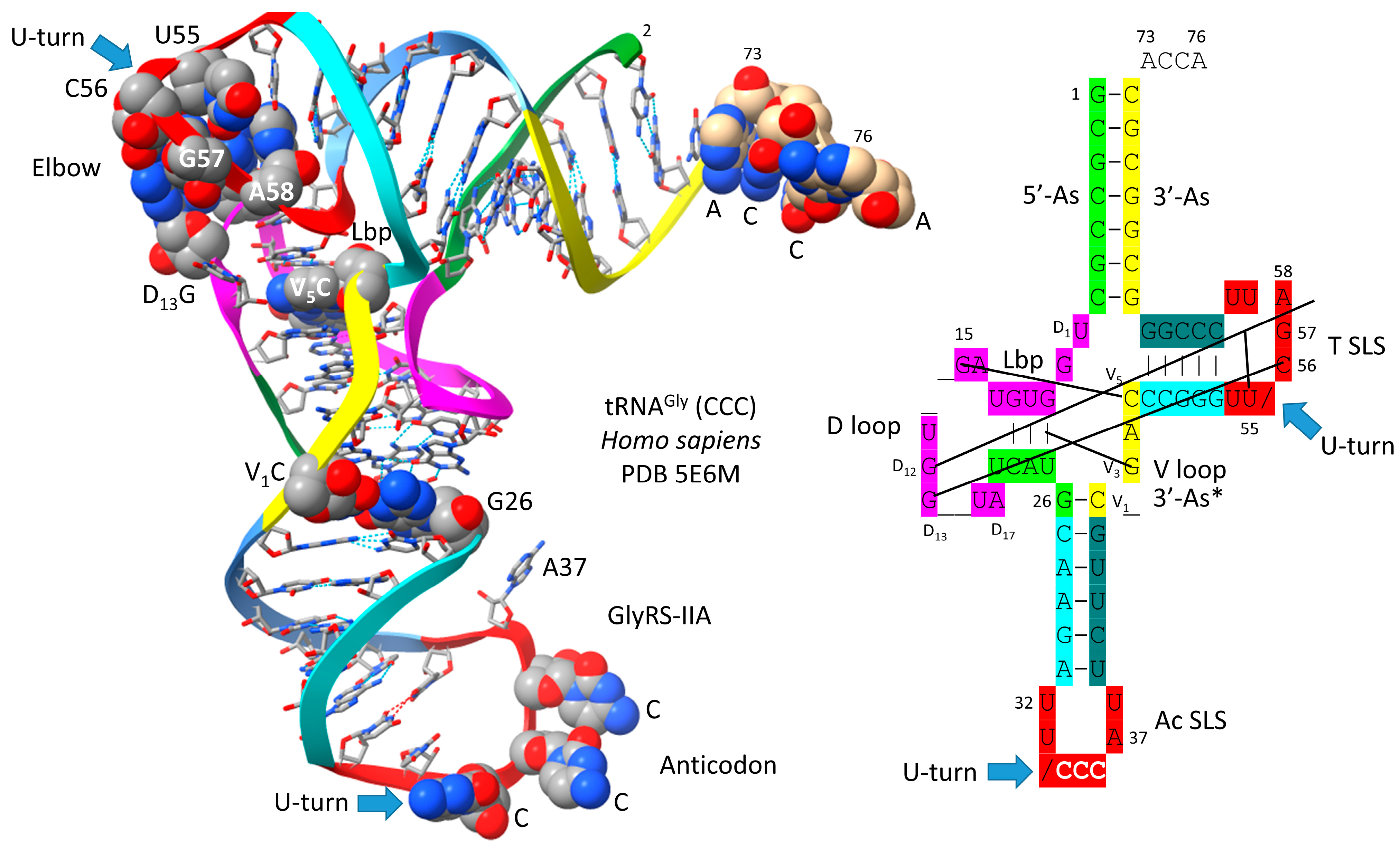
5. Was the First tRNA a tRNAGly?
6. Breaking tRNAPri into Its Component Parts
7. Chemical Evolution of the First Translation Systems
8. Evolution of Third- and Fourth-Generation Polyglycine Synthesis Systems
9. Alternate Genetic Adapters
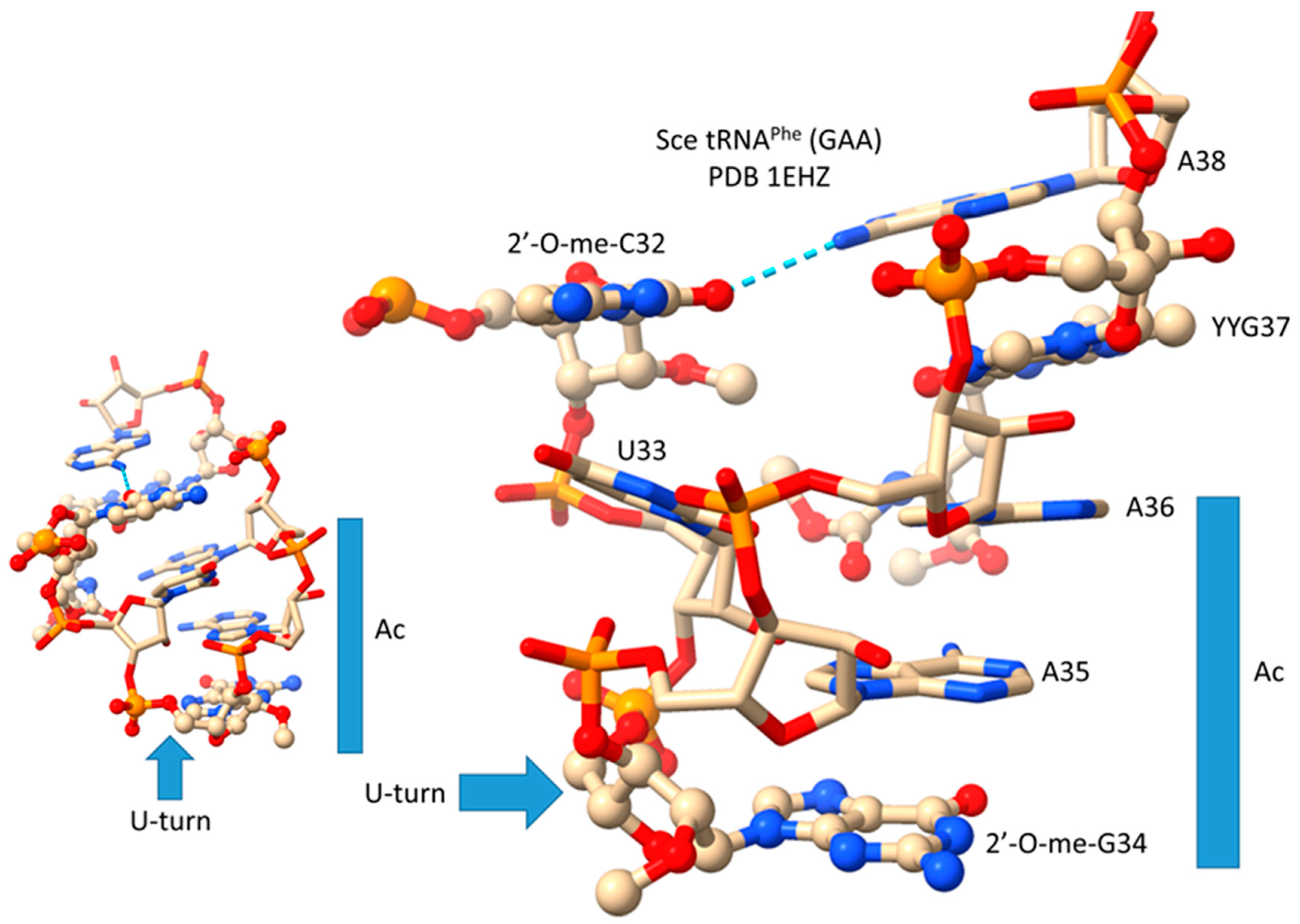
10. The Anticodon Loop as Essential Intellectual Property to Evolve Life on Earth
11. Determinants on tRNA for Cognate Aminoacyl-tRNA Synthetase Recognition
12. Dirty Polyglycine and Emulsification at the Origin of Life
13. Ribozymes and RNA in Pre-Life
14. The Three 31 nt Minihelix tRNA Evolution Theorem
15. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AARS | Aminoacyl-tRNA synthetase (ligase) |
| LUCA | Last Universal Common (Cellular) Ancestor |
| Elp3 | Elongator Protein 3 |
| Indel | Insertions and Deletions |
| Ac | Anticodon |
| As | Acceptor stem |
| SLS | Stem–loop–stem |
References
- Lei, L.; Burton, Z.F. Chemical Evolution of Life on Earth. Genes 2025, 16, 220. [Google Scholar] [CrossRef]
- Lei, L.; Burton, Z.F. Origin of Type II tRNA Variable Loops, Aminoacyl-tRNA Synthetase Allostery from Distal Determinants, and Diversification of Life. DNA 2024, 4, 252–275. [Google Scholar] [CrossRef]
- Lei, L.; Burton, Z.F. The 3 31 Nucleotide Minihelix tRNA Evolution Theorem and the Origin of Life. Life 2023, 13, 2224. [Google Scholar] [CrossRef]
- Lei, L.; Burton, Z.F. Evolution of the genetic code. Transcription 2021, 12, 28–53. [Google Scholar] [CrossRef]
- Chan, P.P.; Lowe, T.M. GtRNAdb 2.0: An expanded database of transfer RNA genes identified in complete and draft genomes. Nucleic Acids Res. 2016, 44, D184–D189. [Google Scholar] [CrossRef]
- Abe, T.; Inokuchi, H.; Yamada, Y.; Muto, A.; Iwasaki, Y.; Ikemura, T. tRNADB-CE: tRNA gene database well-timed in the era of big sequence data. Front. Genet. 2014, 5, 114. [Google Scholar] [CrossRef]
- Abe, T.; Ikemura, T.; Sugahara, J.; Kanai, A.; Ohara, Y.; Uehara, H.; Kinouchi, M.; Kanaya, S.; Yamada, Y.; Muto, A.; et al. tRNADB-CE 2011: tRNA gene database curated manually by experts. Nucleic Acids Res. 2011, 39, D210–D213. [Google Scholar] [CrossRef]
- Abe, T.; Ikemura, T.; Ohara, Y.; Uehara, H.; Kinouchi, M.; Kanaya, S.; Yamada, Y.; Muto, A.; Inokuchi, H. tRNADB-CE: tRNA gene database curated manually by experts. Nucleic Acids Res. 2009, 37, D163–D168. [Google Scholar] [CrossRef] [PubMed]
- Burley, S.K.; Piehl, D.W.; Vallat, B.; Zardecki, C. RCSB Protein Data Bank: Supporting research and education worldwide through explorations of experimentally determined and computationally predicted atomic level 3D biostructures. IUCrJ 2024, 11, 279–286. [Google Scholar] [CrossRef]
- Bittrich, S.; Bhikadiya, C.; Bi, C.; Chao, H.; Duarte, J.M.; Dutta, S.; Fayazi, M.; Henry, J.; Khokhriakov, I.; Lowe, R.; et al. RCSB Protein Data Bank: Efficient Searching and Simultaneous Access to One Million Computed Structure Models Alongside the PDB Structures Enabled by Architectural Advances. J. Mol. Biol. 2023, 435, 167994. [Google Scholar] [CrossRef]
- Burley, S.K.; Bhikadiya, C.; Bi, C.; Bittrich, S.; Chao, H.; Chen, L.; Craig, P.A.; Crichlow, G.V.; Dalenberg, K.; Duarte, J.M.; et al. RCSB Protein Data Bank (RCSB.org): Delivery of experimentally-determined PDB structures alongside one million computed structure models of proteins from artificial intelligence/machine learning. Nucleic Acids Res. 2023, 51, D488–D508. [Google Scholar] [CrossRef]
- Meng, E.C.; Goddard, T.D.; Pettersen, E.F.; Couch, G.S.; Pearson, Z.J.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 2023, 32, e4792. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Goddard, T.D.; Huang, C.C.; Meng, E.C.; Pettersen, E.F.; Couch, G.S.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 2018, 27, 14–25. [Google Scholar] [CrossRef]
- Fukunaga, R.; Yokoyama, S. Aminoacylation complex structures of leucyl-tRNA synthetase and tRNALeu reveal two modes of discriminator-base recognition. Nat. Struct. Mol. Biol. 2005, 12, 915–922. [Google Scholar] [CrossRef]
- Fukunaga, R.; Yokoyama, S. Crystal structure of leucyl-tRNA synthetase from the archaeon Pyrococcus horikoshii reveals a novel editing domain orientation. J. Mol. Biol. 2005, 346, 57–71. [Google Scholar] [CrossRef]
- Weiss, M.C.; Preiner, M.; Xavier, J.C.; Zimorski, V.; Martin, W.F. The last universal common ancestor between ancient Earth chemistry and the onset of genetics. PLoS Genet. 2018, 14, e1007518. [Google Scholar] [CrossRef]
- Weiss, M.C.; Sousa, F.L.; Mrnjavac, N.; Neukirchen, S.; Roettger, M.; Nelson-Sathi, S.; Martin, W.F. The physiology and habitat of the last universal common ancestor. Nat. Microbiol. 2016, 1, 16116. [Google Scholar] [CrossRef]
- Shi, H.; Moore, P.B. The crystal structure of yeast phenylalanine tRNA at 1.93 A resolution: A classic structure revisited. RNA 2000, 6, 1091–1105. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Deng, X.; Chen, L.; Xie, W. Crystal Structure of the Wild-Type Human GlyRS Bound with tRNA(Gly) in a Productive Conformation. J. Mol. Biol. 2016, 428, 3603–3614. [Google Scholar] [CrossRef]
- Pak, D.; Du, N.; Kim, Y.; Sun, Y.; Burton, Z.F. Rooted tRNAomes and evolution of the genetic code. Transcription 2018, 9, 137–151. [Google Scholar] [CrossRef]
- Paul, N.; Joyce, G.F. A self-replicating ligase ribozyme. Proc. Natl. Acad. Sci. USA 2002, 99, 12733–12740. [Google Scholar] [CrossRef] [PubMed]
- Kakoti, A.; Joyce, G.F. RNA Polymerase Ribozyme That Recognizes the Template-Primer Complex through Tertiary Interactions. Biochemistry 2023, 62, 1916–1928. [Google Scholar] [CrossRef] [PubMed]
- Tjhung, K.F.; Shokhirev, M.N.; Horning, D.P.; Joyce, G.F. An RNA polymerase ribozyme that synthesizes its own ancestor. Proc. Natl. Acad. Sci. USA 2020, 117, 2906–2913. [Google Scholar] [CrossRef]
- Samanta, B.; Horning, D.P.; Joyce, G.F. 3’-End labeling of nucleic acids by a polymerase ribozyme. Nucleic Acids Res. 2018, 46, e103. [Google Scholar] [CrossRef]
- Horning, D.P.; Joyce, G.F. Amplification of RNA by an RNA polymerase ribozyme. Proc. Natl. Acad. Sci. USA 2016, 113, 9786–9791. [Google Scholar] [CrossRef]
- Muller, F.; Escobar, L.; Xu, F.; Wegrzyn, E.; Nainyte, M.; Amatov, T.; Chan, C.Y.; Pichler, A.; Carell, T. A prebiotically plausible scenario of an RNA-peptide world. Nature 2022, 605, 279–284. [Google Scholar] [CrossRef]
- Burton, Z.F. The 3-Minihelix tRNA Evolution Theorem. J. Mol. Evol. 2020, 88, 234–242. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Burton, Z.F. “Superwobbling” and tRNA-34 Wobble and tRNA-37 Anticodon Loop Modifications in Evolution and Devolution of the Genetic Code. Life 2022, 12, 252. [Google Scholar] [CrossRef]
- Alkatib, S.; Scharff, L.B.; Rogalski, M.; Fleischmann, T.T.; Matthes, A.; Seeger, S.; Schottler, M.A.; Ruf, S.; Bock, R. The contributions of wobbling and superwobbling to the reading of the genetic code. PLoS Genet. 2012, 8, e1003076. [Google Scholar] [CrossRef]
- Rogalski, M.; Karcher, D.; Bock, R. Superwobbling facilitates translation with reduced tRNA sets. Nat. Struct. Mol. Biol. 2008, 15, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Tang, G.Q.; Hu, H.; Douglas, J.; Carter, C.W., Jr. Primordial aminoacyl-tRNA synthetases preferred minihelices to full-length tRNA. Nucleic Acids Res. 2024, 52, 7096–7111. [Google Scholar] [CrossRef]
- Martinez-Rodriguez, L.; Erdogan, O.; Jimenez-Rodriguez, M.; Gonzalez-Rivera, K.; Williams, T.; Li, L.; Weinreb, V.; Collier, M.; Chandrasekaran, S.N.; Ambroggio, X.; et al. Functional Class I and II Amino Acid-activating Enzymes Can Be Coded by Opposite Strands of the Same Gene. J. Biol. Chem. 2015, 290, 19710–19725. [Google Scholar] [CrossRef]
- Carter, C.W., Jr.; Li, L.; Weinreb, V.; Collier, M.; Gonzalez-Rivera, K.; Jimenez-Rodriguez, M.; Erdogan, O.; Kuhlman, B.; Ambroggio, X.; Williams, T.; et al. The Rodin-Ohno hypothesis that two enzyme superfamilies descended from one ancestral gene: An unlikely scenario for the origins of translation that will not be dismissed. Biol. Direct. 2014, 9, 11. [Google Scholar] [CrossRef]
- Stephen, P.; Ye, S.; Zhou, M.; Song, J.; Zhang, R.; Wang, E.D.; Giege, R.; Lin, S.X. Structure of Escherichia coli Arginyl-tRNA Synthetase in Complex with tRNA(Arg): Pivotal Role of the D-loop. J. Mol. Biol. 2018, 430, 1590–1606. [Google Scholar] [CrossRef]
- Giege, R.; Eriani, G. The tRNA identity landscape for aminoacylation and beyond. Nucleic Acids Res. 2023, 51, 1528–1570. [Google Scholar] [CrossRef]
- Tawfik, D.S.; Gruic-Sovulj, I. How evolution shapes enzyme selectivity—lessons from aminoacyl-tRNA synthetases and other amino acid utilizing enzymes. FEBS J. 2020, 287, 1284–1305. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Seligmann, H. Evolution of small and large ribosomal RNAs from accretion of tRNA subelements. Biosystems 2022, 222, 104796. [Google Scholar] [CrossRef]
- Demongeot, J.; Seligmann, H. Codon assignment evolvability in theoretical minimal RNA rings. Gene 2021, 769, 145208. [Google Scholar] [CrossRef] [PubMed]
- Di Giulio, M. An RNA Ring was Not the Progenitor of the tRNA Molecule. J. Mol. Evol. 2020, 88, 228–233. [Google Scholar] [CrossRef]
- Demongeot, J.; Seligmann, H. Theoretical minimal RNA rings mimick molecular evolution before tRNA-mediated translation: Codon-amino acid affinities increase from early to late RNA rings. C. R. Biol. 2020, 343, 111–122. [Google Scholar] [CrossRef]
- Demongeot, J.; Seligmann, H. RNA Rings Strengthen Hairpin Accretion Hypotheses for tRNA Evolution: A Reply to Commentaries by Z.F. Burton and M. Di Giulio. J. Mol. Evol. 2020, 88, 243–252. [Google Scholar] [CrossRef]
- Demongeot, J.; Seligmann, H. The primordial tRNA acceptor stem code from theoretical minimal RNA ring clusters. BMC Genet. 2020, 21, 7. [Google Scholar] [CrossRef]
- Demongeot, J.; Seligmann, H. The Uroboros Theory of Life’s Origin: 22-Nucleotide Theoretical Minimal RNA Rings Reflect Evolution of Genetic Code and tRNA-rRNA Translation Machineries. Acta Biotheor. 2019, 67, 273–297. [Google Scholar] [CrossRef] [PubMed]
- Di Giulio, M. A polyphyletic model for the origin of tRNAs has more support than a monophyletic model. J. Theor. Biol. 2013, 318, 124–128. [Google Scholar] [CrossRef] [PubMed]
- Di Giulio, M. The ‘recently’ split transfer RNA genes may be close to merging the two halves of the tRNA rather than having just separated them. J. Theor. Biol. 2012, 310, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Di Giulio, M. The origin of the tRNA molecule: Independent data favor a specific model of its evolution. Biochimie 2012, 94, 1464–1466. [Google Scholar] [CrossRef]
- Branciamore, S.; Di Giulio, M. The presence in tRNA molecule sequences of the double hairpin, an evolutionary stage through which the origin of this molecule is thought to have passed. J. Mol. Evol. 2011, 72, 352–363. [Google Scholar] [CrossRef]
- Di Giulio, M. Transfer RNA genes in pieces are an ancestral character. EMBO Rep. 2008, 9, 820, Reply in EMBO Rep. 2008, 9, 820–821. [Google Scholar] [CrossRef]
- Widmann, J.; Di Giulio, M.; Yarus, M.; Knight, R. tRNA creation by hairpin duplication. J. Mol. Evol. 2005, 61, 524–530. [Google Scholar] [CrossRef]
- Di Giulio, M. The origin of the tRNA molecule: Implications for the origin of protein synthesis. J. Theor. Biol. 2004, 226, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Caetano-Anolles, G. Ancestral Insertions and Expansions of rRNA do not Support an Origin of the Ribosome in Its Peptidyl Transferase Center. J. Mol. Evol. 2015, 80, 162–165. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Kowiatek, B.; Opron, K.; Burton, Z.F. Type-II tRNAs and Evolution of Translation Systems and the Genetic Code. Int. J. Mol. Sci. 2018, 19, 3275. [Google Scholar] [CrossRef]
- Pak, D.; Root-Bernstein, R.; Burton, Z.F. tRNA structure and evolution and standardization to the three nucleotide genetic code. Transcription 2017, 8, 205–219. [Google Scholar] [CrossRef]
- Petrov, A.S.; Gulen, B.; Norris, A.M.; Kovacs, N.A.; Bernier, C.R.; Lanier, K.A.; Fox, G.E.; Harvey, S.C.; Wartell, R.M.; Hud, N.V.; et al. History of the ribosome and the origin of translation. Proc. Natl. Acad. Sci. USA 2015, 112, 15396–15401. [Google Scholar] [CrossRef] [PubMed]
- Ikehara, K. Why Were [GADV]-amino Acids and GNC Codons Selected and How Was GNC Primeval Genetic Code Established? Genes 2023, 14, 375. [Google Scholar] [CrossRef]
- Ikehara, K. [GADV]-protein world hypothesis on the origin of life. Orig. Life Evol. Biosph. 2014, 44, 299–302. [Google Scholar] [CrossRef]
- Wehbi, S.; Wheeler, A.; Morel, B.; Manepalli, N.; Minh, B.Q.; Lauretta, D.S.; Masel, J. Order of amino acid recruitment into the genetic code resolved by last universal common ancestor’s protein domains. Proc. Natl. Acad. Sci. USA 2024, 121, e2410311121. [Google Scholar] [CrossRef]
- Ikehara, K. Evolutionary Steps in the Emergence of Life Deduced from the Bottom-Up Approach and GADV Hypothesis (Top-Down Approach). Life 2016, 6, 6. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lei, L.; Burton, Z.F. A Recipe to Evolve Complex Life Chemically on Earth. Genes 2025, 16, 1136. https://doi.org/10.3390/genes16101136
Lei L, Burton ZF. A Recipe to Evolve Complex Life Chemically on Earth. Genes. 2025; 16(10):1136. https://doi.org/10.3390/genes16101136
Chicago/Turabian StyleLei, Lei, and Zachary Frome Burton. 2025. "A Recipe to Evolve Complex Life Chemically on Earth" Genes 16, no. 10: 1136. https://doi.org/10.3390/genes16101136
APA StyleLei, L., & Burton, Z. F. (2025). A Recipe to Evolve Complex Life Chemically on Earth. Genes, 16(10), 1136. https://doi.org/10.3390/genes16101136







