Identification and Functional Annotation of Genes Related to Bone Stability in Laying Hens Using Random Forests
Abstract
1. Introduction
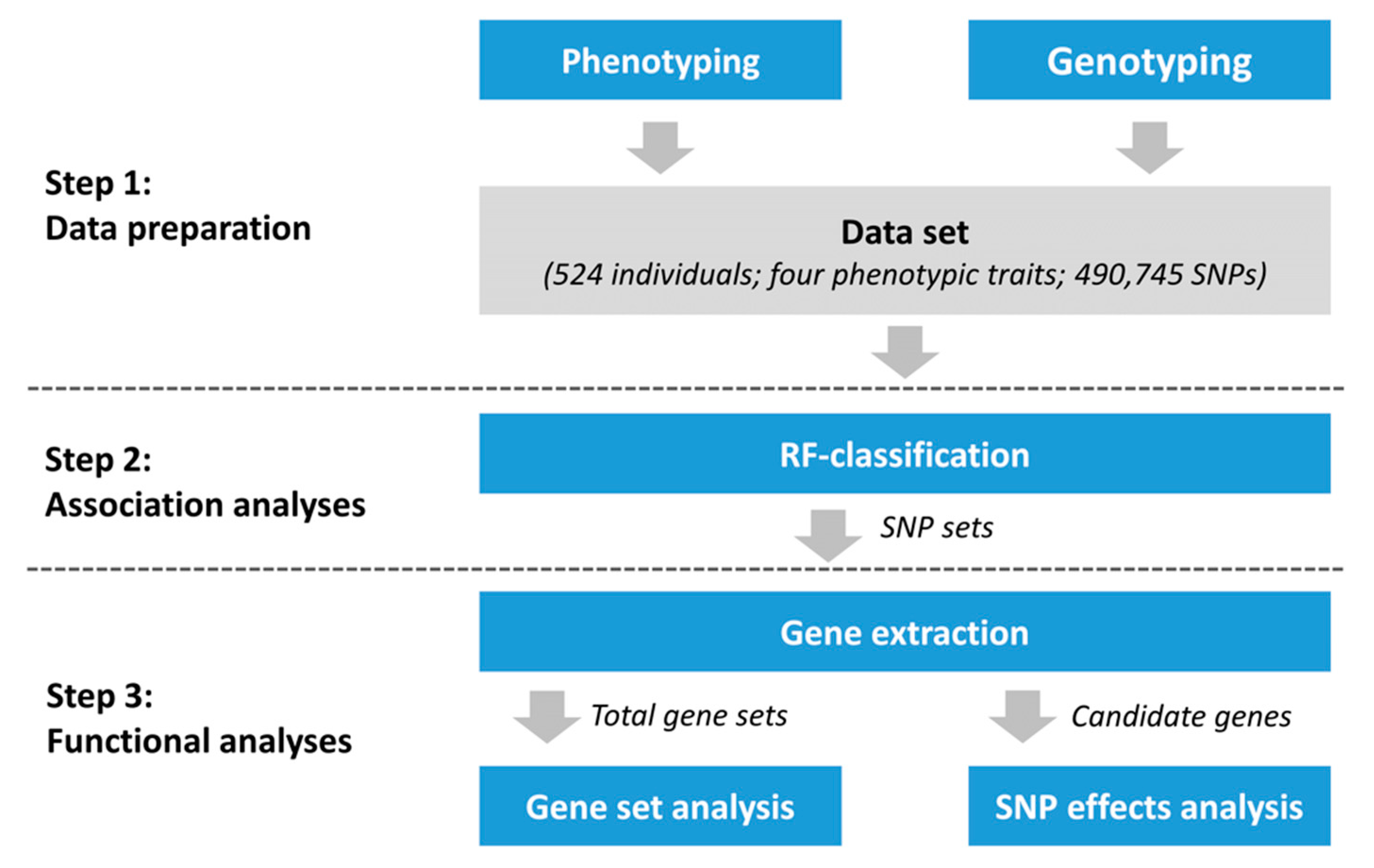
2. Materials and Methods
2.1. Population and Experimental Setup
2.2. Genotyping
2.3. Random Forests Classification
2.4. Functional Analyses
2.4.1. Gene Extraction
2.4.2. SNP Effects Analysis
2.4.3. Gene Set Analysis
| SNP | Trait | Location | GGA 1 | Position 2 | Genotypes | N Individuals | Genotype Frequencies | EA/OA 3 | EA Frequency | Candidate Gene | Reference 4 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| AX-75268181 | Tib_BMD | intragenic | 1 | 139,001,157 | CC/CT/TT | 392/96/36 | 0.75/0.18/0.07 | T/C | 0.16 | MCF2L | [54] |
| AX-76044166 | Tib_BBS | intragenic | 2 | 15,440,861 | AA/AG/GG | 421/63/40 | 0.80/0.12/0.08 | G/A | 0.14 | MPP7 | [55] |
| AX-80813610 | Tib_BMD | downstream | 2 | 23,056,581 | CC/CG/GG | 339/113/72 | 0.65/0.22/0.13 | G/C | 0.25 | CALCR | [56] |
| AX-76099065 | Tib_BMD | intragenic | 2 | 46,101,680 | GG/GA/AA | 392/77/55 | 0.75/0.15/0.10 | A/G | 0.18 | SFRP4 | [57] |
| AX-76601713 | Tib_BBS | intragenic | 3 | 10,617,925 | AA/AG/GG | 265/102/157 | 0.51/0.19/0.30 | G/A | 0.40 | ACTR2 | [15] |
| AX-77276717 | Tib_BBS | intragenic | 3 | 19,498,104 | GG/GA/AA | 322/145/57 | 0.61/0.28/0.11 | A/G | 0.25 | TGFB2 | [58] |
| AX-76491534 | Tib_BBS | intragenic | 3 | 49,027,160 | AA/AG/GG | 432/62/30 | 0.82/0.12/0.06 | G/A | 0.12 | CCDC170 | [59] |
| AX-76772658 | Tib_BBS/Hum_BBS | intragenic | 5 | 11,438,677 | TT/TC/CC | 219/199/109 | 0.41/0.38/0.21 | C/T | 0.40 | SOX6 | [60] |
| AX-77113061 | Tib_BMD | upstream | 8 | 5,889,886 | GG/AG/AA | 202/156/166 | 0.38/0.30/0.32 | A/G | 0.47 | TMCO1 | [61] |
| AX-77091655 | Hum_BBS/Hum_BMD | upstream | 8 | 24,931,025 | CC/CA/AA | 286/139/99 | 0.54/0.27/0.19 | A/C | 0.32 | PODN | [15] |
| AX-75597497 | Hum_BBS | downstream | 10 | 19,108,829 | AA/AG/GG | 376/124/24 | 0.72/0.24/0.04 | G/A | 0.16 | SMAD6 | [62] |
| AX-75677174 | Tib_BMD | intragenic | 11 | 10,044,055 | CC/CT/TT | 377/107/40 | 0.72/020/0.08 | T/C | 0.18 | GPATCH1 | [55] |
| AX-75711229 | Tib_BBS | intragenic | 12 | 3,804,145 | GG/AG/AA | 459/58/7 | 0.88/0.11/0.01 | A/G | 0.07 | ASPN | [63] |
| AX-75913642 | Tib_BBS | upstream | 18 | 8,793,585 | GG/AG/AA | 451/61/12 | 0.86/0.12/0.02 | A/G | 0.08 | SOX9 | [64] |
| AX-76351785 | Hum_BBS | intragenic | 27 | 3,497,444 | CC/CT/TT | 316/138/70 | 0.61/0.26/0.13 | T/C | 0.26 | WNT9B | [65] |
| AX-76351898 | Hum_BMD | downstream | 27 | 3,518,924 | GG/GA/AA | 483/31/10 | 0.92/0.06/0.02 | A/G | 0.05 | WNT3 | [55] |
| AX-76351899 | Hum_BMD | downstream | 27 | 3,519,091 | TT/TC/CC | 483/31/10 | 0.92/0.06/0.02 | C/T | 0.05 | WNT3 | [55] |
3. Results
3.1. Identified Single Nucleotide Polymorphisms
3.2. Functional Analyses
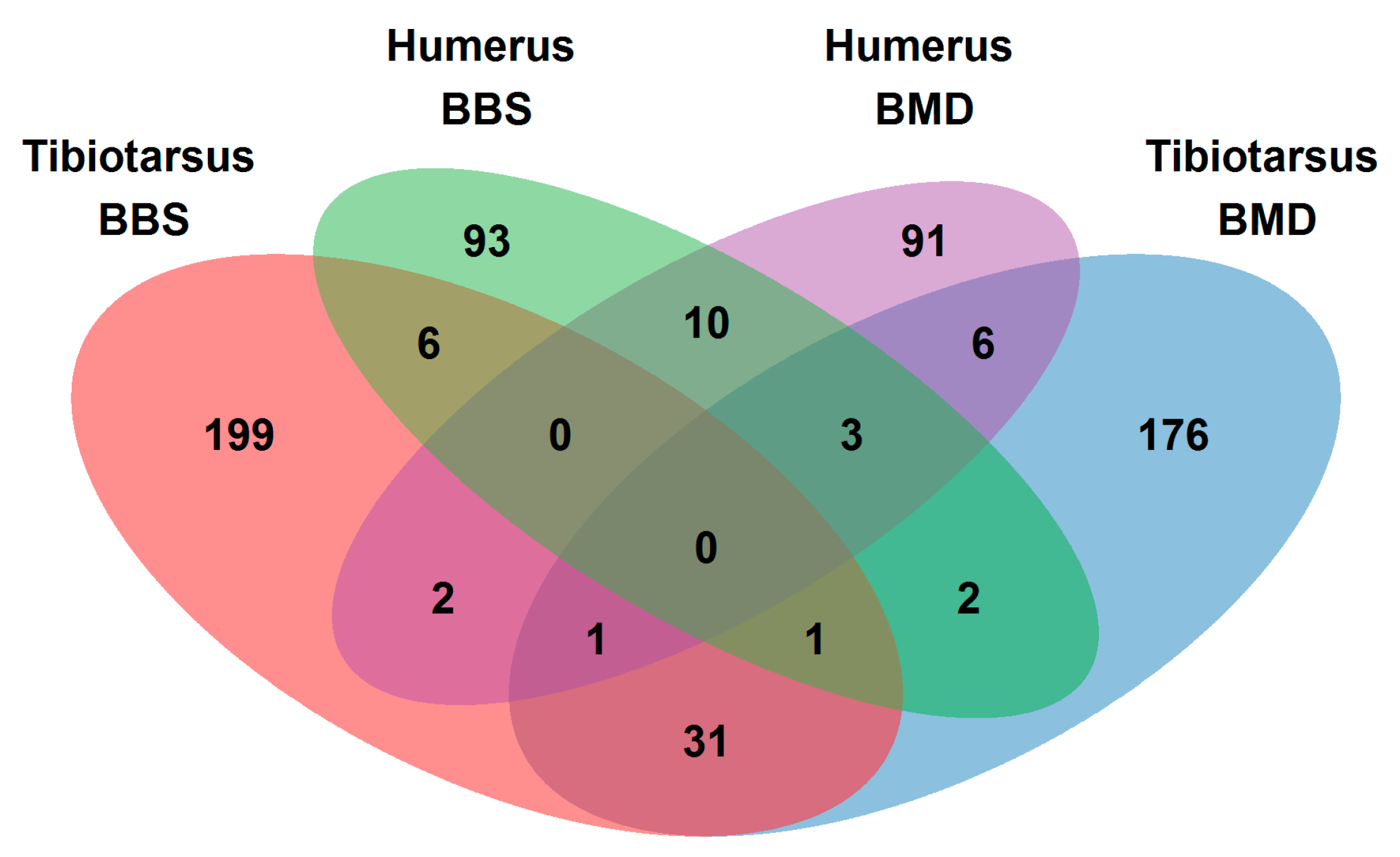
3.2.1. Extracted Gene Sets
3.2.2. SNP Effects Analysis
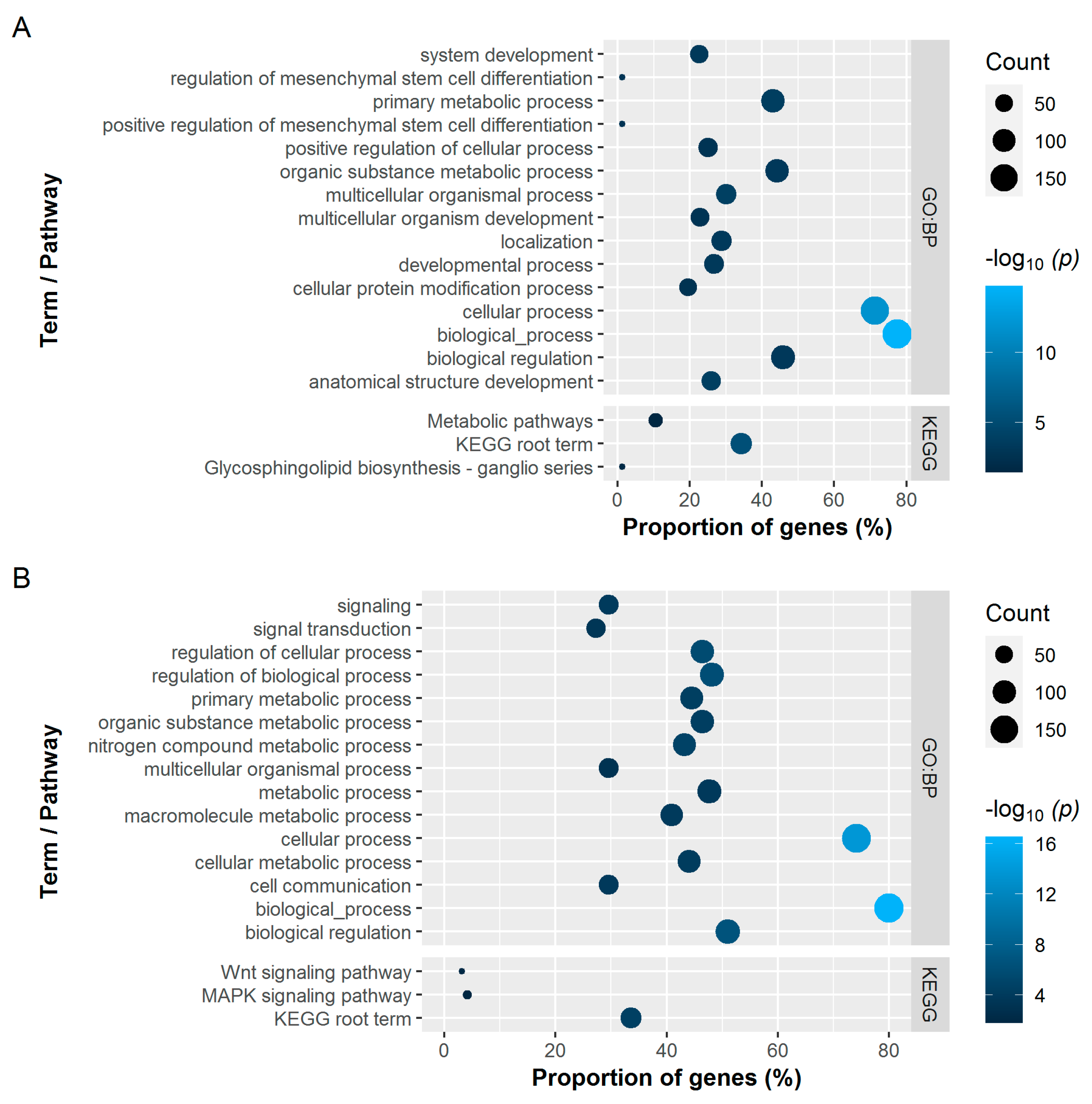
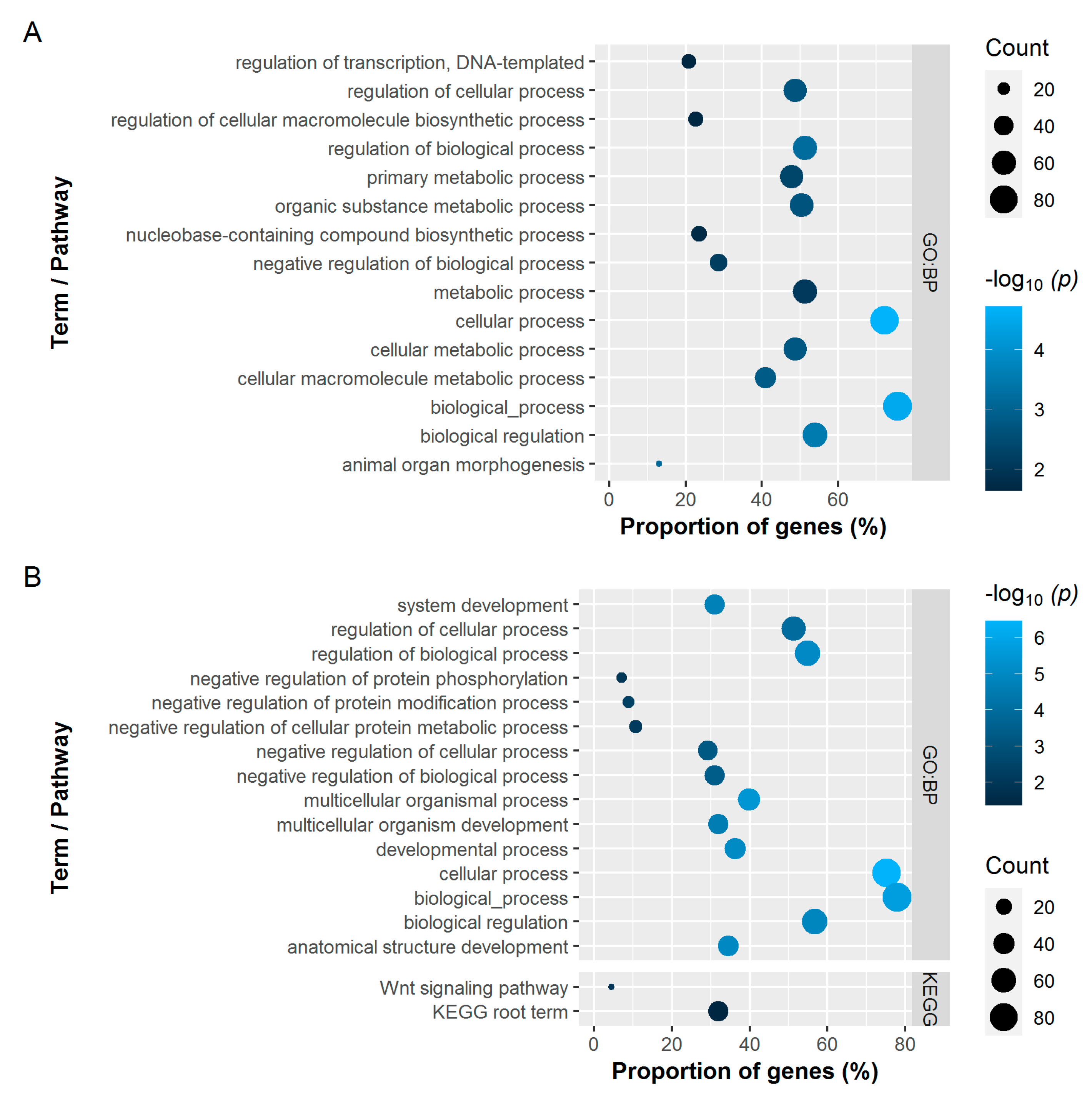
3.2.3. Gene Set Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sandilands, V. The laying hen and bone fractures. Vet. Rec. 2011, 169, 411. [Google Scholar] [CrossRef]
- Petrik, M.T.; Guerin, M.T.; Widowski, T.M. On-farm comparison of keel fracture prevalence and other welfare indicators in conventional cage and floor-housed laying hens in Ontario, Canada. Poult. Sci. 2015, 94, 579–585. [Google Scholar] [CrossRef]
- Fleming, R.H.; McCormack, H.A.; McTeir, L.; Whitehead, C.C. Relationships between genetic, environmental and nutritional factors influencing osteoporosis in laying hens. Br. Poult. Sci. 2006, 47, 742–755. [Google Scholar] [CrossRef]
- Cransberg, P.H.; Parkinson, G.B.; Wilson, S.; Thorp, B.H. Sequential studies of skeletal calcium reserves and structural bone volume in a commercial layer flock. Br. Poult. Sci. 2001, 42, 260–265. [Google Scholar] [CrossRef]
- McCoy, M.A.; Reilly, G.A.C.; Kilpatrick, D.J. Density and breaking strength of bones of mortalities among caged layers. Res. Vet. Sci. 1996, 60, 185–186. [Google Scholar] [CrossRef]
- Nasr, M.A.F.; Nicol, C.J.; Murrell, J.C. Do laying hens with keel bone fractures experience pain? PLoS ONE 2012, 7, e42420. [Google Scholar] [CrossRef]
- Webster, A.B. Welfare implications of avian osteoporosis. Poult. Sci. 2004, 83, 184–192. [Google Scholar] [CrossRef]
- Nasr, M.A.F.; Murrell, J.; Nicol, C.J. The effect of keel fractures on egg production, feed and water consumption in individual laying hens. Br. Poult. Sci. 2013, 54, 165–170. [Google Scholar] [CrossRef]
- Eusemann, B.K.; Patt, A.; Schrader, L.; Weigend, S.; Thöne-Reineke, C.; Petow, S. The role of egg production in the etiology of keel bone damage in laying hens. Front. Vet. Sci. 2020, 7, 81. [Google Scholar] [CrossRef]
- Habig, C.; Baulain, U.; Henning, M.; Scholz, A.; Sharifi, A.; Janisch, S.; Simianer, H.; Weigend, S. How bone stability in laying hens is affected by phylogenetic background and performance level. Eur. Poult. Sci. 2017, 81, 1–17. [Google Scholar]
- Hocking, P.M.; Bain, M.; Channing, C.E.; Fleming, R.; Wilson, S. Genetic variation for egg production, egg quality and bone strength in selected and traditional breeds of laying fowl. Br. Poult. Sci. 2003, 44, 365–373. [Google Scholar] [CrossRef]
- Gebhardt-Henrich, S.G.; Fröhlich, E.K. Early onset of laying and bumblefoot favor keel bone fractures. Animals 2015, 5, 1192–1206. [Google Scholar] [CrossRef]
- Dunn, I.C.; De Koning, D.-J.; McCormack, H.A.; Fleming, R.H.; Wilson, P.W.; Andersson, B.; Schmutz, M.; Benavides, C.; Dominguez-Gasca, N.; Sanchez-Rodriguez, E.; et al. No evidence that selection for egg production persistency causes loss of bone quality in laying hens. Genet. Sel. Evol. 2021, 53, 11. [Google Scholar] [CrossRef]
- Jansen, S.; Baulain, U.; Habig, C.; Weigend, A.; Halle, I.; Scholz, A.M.; Simianer, H.; Sharifi, A.R.; Weigend, S. Relationship between Bone Stability and Egg Production in Genetically Divergent Chicken Layer Lines. Animals 2020, 10, 850. [Google Scholar] [CrossRef]
- Raymond, B.; Johansson, A.M.; McCormack, H.A.; Fleming, R.H.; Schmutz, M.; Dunn, I.C.; De Koning, D.J. Genome-wide association study for bone strength in laying hens. J. Anim. Sci. 2018, 96, 2525–2535. [Google Scholar] [CrossRef]
- Bishop, S.C.; Fleming, R.H.; McCormack, H.A.; Flock, D.K.; Whitehead, C.C. Inheritance of bone characteristics affecting osteoporosis in laying hens. Br. Poult. Sci. 2000, 41, 33–40. [Google Scholar] [CrossRef]
- Rubin, C.J.; Brändström, H.; Wright, D.; Kerje, S.; Gunnarsson, U.; Schutz, K.; Fredriksson, R.; Jensen, P.; Andersson, L.; Ohlsson, C.; et al. Quantitative trait loci for BMD and bone strength in an intercross between domestic and wildtype chickens. J. Bone Miner. Res. 2007, 22, 375–384. [Google Scholar] [CrossRef]
- Zhou, H.; Deeb, N.; Evock-Clover, C.M.; Mitchell, A.D.; Ashwell, C.M.; Lamont, S.J. Genome-Wide Linkage Analysis to Identify Chromosomal Regions Affecting Phenotypic Traits in the Chicken. III. Skeletal Integrity. Poult. Sci. 2007, 86, 255–266. [Google Scholar] [CrossRef]
- Bennett, A.K.; Hester, P.Y.; Spurlock, D.M. Relationships of a transforming growth factor-beta2 single nucleotide polymorphism and messenger ribonucleic acid abundance with bone and production traits in chickens. Poult. Sci. 2007, 86, 829–834. [Google Scholar] [CrossRef]
- Johnsson, M.; Gustafson, I.; Rubin, C.-J.; Sahlqvist, A.-S.; Jonsson, K.B.; Kerje, S.; Ekwall, O.; Kämpe, O.; Andersson, L.; Jensen, P.; et al. A Sexual Ornament in Chickens Is Affected by Pleiotropic Alleles at HAO1 and BMP2, Selected during Domestication. PLoS Genetics 2012, 8, e1002914. [Google Scholar] [CrossRef]
- Schreiweis, M.A.; Hester, P.Y.; Moody, D.E. Identification of quantitative trait loci associated with bone traits and body weight in an F2 resource population of chickens. Genet. Sel. Evol. 2005, 37, 677. [Google Scholar] [CrossRef]
- Podisi, B.K.; Knott, S.A.; Dunn, I.C.; Burt, D.W.; Hocking, P.M. Bone mineral density QTL at sexual maturity and end of lay. Br. Poult. Sci. 2012, 53, 763–769. [Google Scholar] [CrossRef]
- Dunn, I.C.; Fleming, R.H.; McCormack, H.A.; Morrice, D.; Burt, D.W.; Preisinger, R.; Whitehead, C.C. A QTL for osteoporosis detected in an F2 population derived from White Leghorn chicken lines divergently selected for bone index. Anim. Genet. 2007, 38, 45–49. [Google Scholar] [CrossRef]
- De Koning, D.-J.; Dominguez-Gasca, N.; Fleming, R.H.; Gill, A.; Kurian, D.; Law, A.; McCormack, H.A.; Morrice, D.; Sanchez-Rodriguez, E.; Rodriguez-Navarro, A.B.; et al. An eQTL in the cystathionine beta synthase gene is linked to osteoporosis in laying hens. Genet. Sel. Evol. 2020, 52, 13. [Google Scholar] [CrossRef]
- Guo, J.; Sun, C.; Qu, L.; Shen, M.; Dou, T.; Ma, M.; Wang, K.; Yang, N. Genetic architecture of bone quality variation in layer chickens revealed by a genome-wide association study. Sci. Rep. 2017, 7, 45317. [Google Scholar] [CrossRef]
- Rothammer, S.; Bernau, M.; Kremer-Rücker, P.V.; Medugorac, I.; Scholz, A.M. Genome-wide QTL mapping results for regional DXA body composition and bone mineral density traits in pigs. Arch. Anim. Breed. 2017, 60, 51–59. [Google Scholar] [CrossRef]
- Ramzan, F.; Klees, S.; Schmitt, A.O.; Cavero, D.; Gültas, M. Identification of Age-Specific and Common Key Regulatory Mechanisms Governing Eggshell Strength in Chicken Using Random Forests. Genes 2020, 11, 464. [Google Scholar] [CrossRef]
- Romagnoni, A.; Jégou, S.; Van Steen, K.; Wainrib, G.; Hugot, J.-P.; Peyrin-Biroulet, L.; Chamaillard, M.; Colombel, J.-F.; Cottone, M.; D’Amato, M.; et al. Comparative performances of machine learning methods for classifying Crohn Disease patients using genome-wide genotyping data. Sci. Rep. 2019, 9, 10351. [Google Scholar] [CrossRef]
- Brieuc, M.S.O.; Waters, C.D.; Drinan, D.P.; Naish, K.A. A practical introduction to Random Forest for genetic association studies in ecology and evolution. Mol. Ecol. Resour. 2018, 18, 755–766. [Google Scholar] [CrossRef]
- Brieuc, M.S.O.; Ono, K.; Drinan, D.P.; Naish, K.A. Integration of Random Forest with population-based outlier analyses provides insight on the genomic basis and evolution of run timing in Chinook salmon (Oncorhynchus tshawytscha). Mol. Ecol. 2015, 24, 2729–2746. [Google Scholar] [CrossRef]
- Nguyen, T.-T.; Huang, J.Z.; Wu, Q.; Nguyen, T.T.; Li, M.J. Genome-wide association data classification and SNPs selection using two-stage quality-based Random Forests. BMC Genom. 2015, 16, S5. [Google Scholar] [CrossRef]
- Lieboldt, M.-A.; Halle, I.; Frahm, J.; Schrader, L.; Baulain, U.; Henning, M.; Preisinger, R.; Dänicke, S.; Weigend, S. Phylogenic versus selection effects on growth development, egg laying and egg quality in purebred laying hens. Eur. Poult. Sci. 2015, 79, 1–16. [Google Scholar]
- Dudde, A.; Weigend, S.; Krause, E.T.; Jansen, S.; Habig, C.; Schrader, L. Chickens in motion: Effects of egg production level and pen size on the motor abilities and bone stability of laying hens (Gallus gallus forma domestica). Appl. Anim. Behav. Sci. 2020, 227, 104998. [Google Scholar] [CrossRef]
- Kranis, A.; Gheyas, A.A.; Boschiero, C.; Turner, F.; Yu, L.; Smith, S.; Talbot, R.; Pirani, A.; Brew, F.; Kaiser, P.; et al. Development of a high density 600K SNP genotyping array for chicken. BMC Genom. 2013, 14, 59. [Google Scholar] [CrossRef]
- SNP & Variation Suite ™ (Version 8.9); Golden, Helix, Inc.: Bozeman, MT, USA; Available online: https://www.goldenhelix.com (accessed on 6 May 2021).
- Browning, B.L.; Zhou, Y.; Browning, S.R. A One-Penny Imputed Genome from Next-Generation Reference Panels. Am. J. Hum. Genet. 2018, 103, 338–348. [Google Scholar] [CrossRef]
- National Center for Biotechnology Information (NCBI). Genome Assembly GRCg6a. Available online: https://www.ncbi.nlm.nih.gov/assembly/GCF_000002315.6 (accessed on 17 June 2020).
- Ramzan, F.; Gültas, M.; Bertram, H.; Cavero, D.; Schmitt, A.O. Combining Random Forests and a Signal Detection Method Leads to the Robust Detection of Genotype-Phenotype Associations. Genes 2020, 11, 892. [Google Scholar] [CrossRef]
- Kursa, M.B.; Rudnicki, W.R. Feature selection with the Boruta package. J. Stat. Softw. 2010, 36, 1–13. [Google Scholar] [CrossRef]
- Python Software Foundation. Python Language Reference, Version 3.8.9. Available online: https://docs.python.org/3.8/reference/index.html (accessed on 6 May 2021).
- Homola, D. Python Implementations of the Boruta All-Relevant Feature Selection Method. Available online: https://github.com/scikit-learn-contrib/boruta_py (accessed on 14 October 2020).
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Yates, A.D.; Achuthan, P.; Akanni, W.; Allen, J.; Allen, J.; Alvarez-Jarreta, J.; Amode, M.R.; Armean, I.M.; Azov, A.G.; Bennett, R.; et al. Ensembl 2020. Nucleic Acids Res. 2020, 48, 682–688. [Google Scholar] [CrossRef] [PubMed]
- Kinsella, R.J.; Kähäri, A.; Haider, S.; Zamora, J.; Proctor, G.; Spudich, G.; Almeida-King, J.; Staines, D.; Derwent, P.; Kerhornou, A.; et al. Ensembl BioMarts: A hub for data retrieval across taxonomic space. Database 2011, 2011, bar030. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.R.; Hem, V.; Katz, K.S.; Ovetsky, M.; Wallin, C.; Ermolaeva, O.; Tolstoy, I.; Tatusova, T.; Pruitt, K.D.; Maglott, D.R.; et al. Gene: A gene-centered information resource at NCBI. Nucleic Acids Res. 2015, 43, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Wiedemann, I.; Maehlmeyer, A.; Jansen, S.; Sharifi, A.R.; Knorr, C. SNP g.1007A>G within the porcine DNAL4 gene affects sperm motility traits and percentage of midpiece abnormalities. Reprod. Domest. Anim. 2018, 53, 401–413. [Google Scholar] [CrossRef] [PubMed]
- Bates, D.; Mächler, M.; Bolker, B.; Walker, S. Fitting Linear Mixed-Effects Models Using lme4. J. Stat. Softw. 2015, 67, 1–48. [Google Scholar] [CrossRef]
- Van Lenth, R. Emmeans: Estimated Marginal Means, aka Least-Squares Means. R Package Version 1.5.3. Available online: https://CRAN.R-project.org/package=emmeans (accessed on 8 January 2021).
- Raudvere, U.; Kolberg, L.; Kuzmin, I.; Arak, T.; Adler, P.; Peterson, H.; Vilo, J. g:Profiler: A web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 2019, 47, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Furumichi, M.; Sato, Y.; Ishiguro-Watanabe, M.; Tanabe, M. KEGG: Integrating viruses and cellular organisms. Nucleic Acids Res. 2021, 49, D545–D551. [Google Scholar] [CrossRef]
- Reimand, J.; Kull, M.; Peterson, H.; Hansen, J.; Vilo, J. g:Profiler--a web-based toolset for functional profiling of gene lists from large-scale experiments. Nucleic Acids Res. 2007, 35, 193–200. [Google Scholar] [CrossRef]
- Sayols, S. Rrvgo: A Bioconductor Package to Reduce and Visualize Gene Ontology Terms. Available online: https://ssayols.github.io/rrvgo (accessed on 7 February 2021).
- Day-Williams, A.G.; Southam, L.; Panoutsopoulou, K.; Rayner, N.W.; Esko, T.; Estrada, K.; Helgadottir, H.T.; Hofman, A.; Ingvarsson, T.; Jonsson, H.; et al. A variant in MCF2L is associated with osteoarthritis. Am. J. Hum. Genet. 2011, 89, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Estrada, K.; Styrkarsdottir, U.; Evangelou, E.; Hsu, Y.-H.; Duncan, E.L.; Ntzani, E.E.; Oei, L.; Albagha, O.M.E.; Amin, N.; Kemp, J.P.; et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat. Genet. 2012, 44, 491–501. [Google Scholar] [CrossRef]
- Zhang, H.; Tao, X.; Wu, J. Association of calcitonin receptor gene polymorphism with bone mineral density in postmenopausal Chinese women: A meta-analysis. Arch. Gynecol. Obstet. 2015, 291, 165–172. [Google Scholar] [CrossRef]
- Kiper, P.O.S.; Saito, H.; Gori, F.; Unger, S.; Hesse, E.; Yamana, K.; Kiviranta, R.; Solban, N.; Liu, J.; Brommage, R.; et al. Cortical-Bone Fragility--Insights from sFRP4 Deficiency in Pyle’s Disease. N. Engl. J. Med. 2016, 374, 2553–2562. [Google Scholar] [CrossRef]
- Li, H.; Deeb, N.; Zhou, H.; Mitchell, A.D.; Ashwell, C.M.; Lamont, S.J. Chicken quantitative trait loci for growth and body composition associated with transforming growth factor-beta genes. Poult. Sci. 2003, 82, 347–356. [Google Scholar] [CrossRef] [PubMed]
- Mullin, B.H.; Walsh, J.P.; Zheng, H.-F.; Brown, S.J.; Surdulescu, G.L.; Curtis, C.; Breen, G.; Dudbridge, F.; Richards, J.B.; Spector, T.D.; et al. Genome-wide association study using family-based cohorts identifies the WLS and CCDC170/ESR1 loci as associated with bone mineral density. BMC Genom. 2016, 17, 136. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.L.; Guo, Y.; Liu, Y.J.; Shen, H.; Liu, Y.Z.; Lei, S.F.; Li, J.; Tian, Q.; Deng, H.W. Genetic variants in the SOX6 gene are associated with bone mineral density in both Caucasian and Chinese populations. Osteoporos. Int. 2012, 23, 781–787. [Google Scholar] [CrossRef]
- Li, J.; Liu, C.; Li, Y.; Zheng, Q.; Xu, Y.; Liu, B.; Sun, W.; Li, Y.; Ji, S.; Liu, M.; et al. TMCO1-mediated Ca(2+) leak underlies osteoblast functions via CaMKII signaling. Nat. Commun. 2019, 10, 1589. [Google Scholar] [CrossRef]
- Borton, A.J.; Frederick, J.P.; Datto, M.B.; Wang, X.F.; Weinstein, R.S. The loss of Smad3 results in a lower rate of bone formation and osteopenia through dysregulation of osteoblast differentiation and apoptosis. J. Bone Miner. Res. 2001, 16, 1754–1764. [Google Scholar] [CrossRef]
- Mishra, A.; Awasthi, S.; Raj, S.; Mishra, P.; Srivastava, R.N. Identifying the role of ASPN and COMP genes in knee osteoarthritis development. J. Orthop. Surg. Res. 2019, 14, 337. [Google Scholar] [CrossRef]
- Szwedowski, D.; Szczepanek, J.; Paczesny, Ł.; Pękała, P.; Zabrzyński, J.; Kruczyński, J. Genetics in Cartilage Lesions: Basic Science and Therapy Approaches. Int. J. Mol. Sci. 2020, 21, 5430. [Google Scholar] [CrossRef]
- Karasik, D.; Rivadeneira, F.; Johnson, M.L. The genetics of bone mass and susceptibility to bone diseases. Nat. Rev. Rheumatol. 2016, 12, 323–334. [Google Scholar] [CrossRef]
- Johnsson, M.; Jonsson, K.B.; Andersson, L.; Jensen, P.; Wright, D. Genetic Regulation of Bone Metabolism in the Chicken: Similarities and Differences to Mammalian Systems. PLoS Genetics 2015, 11, e1005250. [Google Scholar] [CrossRef]
- Xiao, S.-M.; Kung, A.W.C.; Gao, Y.; Lau, K.-S.; Ma, A.; Zhang, Z.-L.; Liu, J.-M.; Xia, W.; He, J.-W.; Zhao, L.; et al. Post-genome wide association studies and functional analyses identify association of MPP7 gene variants with site-specific bone mineral density. Hum. Mol. Genet. 2012, 21, 1648–1657. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Guo, J.; Kanwal, Z.; Wu, M.; Lv, X.; Ibrahim, N.A.; Li, P.; Buabeid, M.A.; Arafa, E.-S.A.; Sun, Q. Calcitonin and Bone Physiology: In Vitro, In Vivo, and Clinical Investigations. J. Endocrinol. 2020, 2020, 3236828. [Google Scholar] [CrossRef]
- Zofková, I.; Zajícková, K.; Hill, M.; Krepelová, A. Does polymorphism C1377T of the calcitonin receptor gene determine bone mineral density in postmenopausal women? Exp. Clin. Endocrinol. Diabetes 2003, 111, 447–449. [Google Scholar] [CrossRef] [PubMed]
- Schinke, T.; Liese, S.; Priemel, M.; Haberland, M.; Schilling, A.F.; Catala-Lehnen, P.; Blicharski, D.; Rueger, J.M.; Gagel, R.F.; Emeson, R.B.; et al. Decreased Bone Formation and Osteopenia in Mice Lacking α-Calcitonin Gene-Related Peptide. J. Bone Miner. Res. 2004, 19, 2049–2056. [Google Scholar] [CrossRef] [PubMed]
- Goetz, S.C.; Anderson, K.V. The primary cilium: A signalling centre during vertebrate development. Nat. Rev. Genet. 2010, 11, 331–344. [Google Scholar] [CrossRef]
- Ali, I.H.A.; Brazil, D.P. Bone morphogenetic proteins and their antagonists: Current and emerging clinical uses. Br. J. Pharmacol. 2014, 171, 3620–3632. [Google Scholar] [CrossRef] [PubMed]
- Hata, A.; Lagna, G.; Massagué, J.; Hemmati-Brivanlou, A. Smad6 inhibits BMP/Smad1 signaling by specifically competing with the Smad4 tumor suppressor. Genes Dev. 1998, 12, 186–197. [Google Scholar] [CrossRef]
- Hidalgo-Bravo, A.; Parra-Torres, A.Y.; Casas-Avila, L.; Jimenez-Ortega, R.F.; Ramírez-Salazar, E.G.; Patiño, N.; Rivera-Paredez, B.; Salmerón, J.; Valdés-Flores, M.; Velázquez-Cruz, R. Association of RMND1/CCDC170–ESR1 single nucleotide polymorphisms with hip fracture and osteoporosis in postmenopausal women. Climacteric 2019, 22, 97–104. [Google Scholar] [CrossRef]
- Rivadeneira, F.; Styrkársdottir, U.; Estrada, K.; Halldórsson, B.V.; Hsu, Y.-H.; Richards, J.B.; Zillikens, M.C.; Kavvoura, F.K.; Amin, N.; Aulchenko, Y.S.; et al. Twenty bone-mineral-density loci identified by large-scale meta-analysis of genome-wide association studies. Nat. Genetic. 2009, 41, 1199–1206. [Google Scholar]
- Smits, P.; Li, P.; Mandel, J.; Zhang, Z.; Deng, J.M.; Behringer, R.R.; de Crombrugghe, B.; Lefebvre, V. The transcription factors L-Sox5 and Sox6 are essential for cartilage formation. Dev. Cell 2001, 1, 277–290. [Google Scholar] [CrossRef]
- Sammels, E.; Parys, J.B.; Missiaen, L.; De Smedt, H.; Bultynck, G. Intracellular Ca2+ storage in health and disease: A dynamic equilibrium. Cell Calcium 2010, 47, 297–314. [Google Scholar] [CrossRef]
- Shimizu-Hirota, R.; Sasamura, H.; Kuroda, M.; Kobayashi, E.; Saruta, T. Functional characterization of podocan, a member of a new class in the small leucine-rich repeat protein family. FEBS Lett. 2004, 563, 69–74. [Google Scholar] [CrossRef]
- Viguet-Carrin, S.; Garnero, P.; Delmas, P.D. The role of collagen in bone strength. Osteoporos. Int. 2006, 17, 319–336. [Google Scholar] [CrossRef] [PubMed]
- Sparke, A.J.; Sims, T.J.; Avery, N.C.; Bailey, A.J.; Fleming, R.H.; Whitehead, C.C. Differences in composition of avian bone collagen following genetic selection for resistance to osteoporosis. Br. Poult. Sci. 2002, 43, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Bultink, I.E.; Lems, W.F. Osteoarthritis and osteoporosis: What is the overlap? Curr. Rheumatol. Rep. 2013, 15, 328. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, M.C.; Vullings, J.; van de Loo, F.A.J. Osteoporosis and osteoarthritis are two sides of the same coin paid for obesity. Nutrition 2020, 70, 110486. [Google Scholar] [CrossRef] [PubMed]
- Cong, Y.; Ru, J.Y.; Bao, N.R.; Guo, T.; Zhao, J.N. A single nucleotide polymorphism in the TGF-β1 gene (rs1982073 C>T) may contribute to increased risks of bone fracture, osteoporosis, and osteoarthritis: A meta-analysis. Clin. Rheumatol. 2016, 35, 973–985. [Google Scholar] [CrossRef] [PubMed]
- Shepherd, C.; Skelton, A.J.; Rushton, M.D.; Reynard, L.N.; Loughlin, J. Expression analysis of the osteoarthritis genetic susceptibility locus mapping to an intron of the MCF2L gene and marked by the polymorphism rs11842874. BMC Med. Genet. 2015, 16, 108. [Google Scholar] [CrossRef]
- Mao, J.H.; Sui, Y.X.; Ao, S.; Wang, Y.; Liu, Y.; Leng, H. miR-140-3p exhibits repressive functions on preosteoblast viability and differentiation by downregulating MCF2L in osteoporosis. In Vitro Cell. Dev. Biol. Anim. 2020, 56, 49–58. [Google Scholar] [CrossRef] [PubMed]
- González-Huerta, N.C.; Borgonio-Cuadra, V.M.; Zenteno, J.C.; Cortés-González, S.; Duarte-Salazar, C.; Miranda-Duarte, A. D14 repeat polymorphism of the asporin gene is associated with primary osteoarthritis of the knee in a Mexican Mestizo population. Int. J. Rheum. Dis. 2017, 20, 1935–1941. [Google Scholar] [CrossRef]
- Sobhan, M.R.; Mehdinejad, M.; Jamaladini, M.H.; Mazaheri, M.; Zare-Shehneh, M.; Neamatzadeh, H. Association between aspartic acid repeat polymorphism of the asporin gene and risk of knee osteoarthritis: A systematic review and meta-analysis. Acta Orthop. Traumatol. Turc. 2017, 51, 409–415. [Google Scholar] [CrossRef]
- Hsu, Y.H.; Kiel, D.P. Clinical review: Genome-wide association studies of skeletal phenotypes: What we have learned and where we are headed. J. Clin. Endocrinol. Metab. 2012, 97, 1958–1977. [Google Scholar] [CrossRef] [PubMed]
- Juhász, T.; Matta, C.; Somogyi, C.; Katona, É.; Takács, R.; Soha, R.F.; Szabó, I.A.; Cserháti, C.; Sződy, R.; Karácsonyi, Z.; et al. Mechanical loading stimulates chondrogenesis via the PKA/CREB-Sox9 and PP2A pathways in chicken micromass cultures. Cell. Signal. 2014, 26, 468–482. [Google Scholar] [CrossRef] [PubMed]
- Kawano, Y.; Kypta, R. Secreted antagonists of the Wnt signalling pathway. J. Cell Sci. 2003, 116, 2627–2634. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.L. LRP5 and bone mass regulation: Where are we now? BoneKEy Rep. 2012, 1, 1. [Google Scholar] [CrossRef]
- Zhang, Y.; Pizzute, T.; Pei, M. A review of crosstalk between MAPK and Wnt signals and its impact on cartilage regeneration. Cell Tissue Res. 2014, 358, 633–649. [Google Scholar] [CrossRef]
- Gu, H.; Huang, Z.; Chen, G.; Zhou, K.; Zhang, Y.; Chen, J.; Xu, J.; Yin, X. Network and pathway-based analyses of genes associated with osteoporosis. Medicine 2020, 99, e19120. [Google Scholar] [CrossRef] [PubMed]
- Yi, G.; Shen, M.; Yuan, J.; Sun, C.; Duan, Z.; Qu, L.; Dou, T.; Ma, M.; Lu, J.; Guo, J.; et al. Genome-wide association study dissects genetic architecture underlying longitudinal egg weights in chickens. BMC Genom. 2015, 16, 746. [Google Scholar] [CrossRef]
| SNP | Trait 1 | Candidate Gene | Generation | Layer Line | SNP Genotype | Allele Substitution Effect 2 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| F-Statistics | p-Value | F-Statistics | p-Value | F-Statistics | p-Value | Beta (SE 3) | Standardised Beta 4 (SE) | t-Value | p-Value | |||
| AX-76044166 | Tib_BBS | MPP7 | 80.92 | <0.0001 | 46.34 | <0.0001 | 4.05 | 0.0448 | 8.22 (4.09) | 0.10 (0.05) | 2.01 | 0.0448 |
| AX-76601713 | Tib_BBS | ACTR2 | 86.02 | <0.0001 | 106.86 | <0.0001 | 13.33 | 0.0003 | −10.19 (2.79) | −0.18 (0.05) | −3.65 | 0.0003 |
| AX-77276717 | Tib_BBS | TGFB2 | 81.07 | <0.0001 | 102.16 | <0.0001 | 3.32 | 0.0696 | 4.67 (2.57) | 0.06 (0.04) | 1.82 | 0.0696 |
| AX-76491534 | Tib_BBS | CCDC170 | 91.49 | <0.0001 | 84.86 | <0.0001 | 12.58 | 0.0004 | −15.63 (4.41) | −0.17 (0.05) | −3.55 | 0.0004 |
| AX-76772658 | Tib_BBS | SOX6 | 81.50 | <0.0001 | 117.84 | <0.0001 | 10.71 | 0.0012 | 7.63 (2.33) | 0.12 (0.04) | 3.27 | 0.0012 |
| AX-75711229 | Tib_BBS | ASPN | 79.24 | <0.0001 | 84.23 | <0.0001 | 2.08 | 0.1503 | 6.66 (4.62) | 0.05 (0.04) | 1.44 | 0.1503 |
| AX-75913642 | Tib_BBS | SOX9 | 83.08 | <0.0001 | 111.94 | <0.0001 | 9.67 | 0.0019 | −12.87 (4.14) | −0.11 (0.04) | −3.11 | 0.0019 |
| AX-76772658 | Hum_BBS | SOX6 | 36.26 | <0.0001 | 52.59 | <0.0001 | 5.67 | 0.0177 | −5.32 (2.23) | −0.10 (0.04) | −2.38 | 0.0177 |
| AX-77091655 | Hum_BBS | PODN | 39.91 | <0.0001 | 41.64 | <0.0001 | 8.35 | 0.0041 | 6.69 (2.31) | 0.13 (0.04) | 2.89 | 0.0041 |
| AX-75597497 | Hum_BBS | SMAD6 | 36.38 | <0.0001 | 53.40 | <0.0001 | 4.62 | 0.0321 | −7.13 (3.32) | −0.10 (0.05) | −2.15 | 0.0321 |
| AX-76351785 | Hum_BBS | WNT9B | 37.27 | <0.0001 | 67.22 | <0.0001 | 21.57 | <0.0001 | 11.51 (2.48) | 0.21 (0.04) | 4.64 | <0.0001 |
| AX-75268181 | Tib_BMD | MCF2L | 4.30 | 0.0401 | 106.46 | <0.0001 | 13.53 | 0.0003 | −0.015 (0.004) | −0.15 (0.05) | −3.67 | 0.0003 |
| AX-80813610 | Tib_BMD | CALCR | 4.24 | 0.0415 | 56.10 | <0.0001 | 4.86 | 0.0298 | 0.008 (0.004) | 0.10 (0.05) | 2.21 | 0.028 |
| AX-76099065 | Tib_BMD | SFRP4 | 4.31 | 0.0400 | 65.23 | <0.0001 | 8.55 | 0.0036 | −0.016 (0.006) | −0.18 (0.06) | −2.92 | 0.0036 |
| AX-77113061 | Tib_BMD | TMCO1 | 4.45 | 0.0369 | 99.26 | <0.0001 | 5.27 | 0.0221 | 0.008 (0.003) | 0.11 (0.05) | 2.30 | 0.0221 |
| AX-75677174 | Tib_BMD | GPATCH1 | 4.27 | 0.0406 | 61.13 | <0.0001 | 10.84 | 0.0011 | 0.013 (0.004) | 0.13(0.04) | 3.29 | 0.0011 |
| AX-77091655 | Hum_BMD | PODN | 20.70 | <0.0001 | 51.56 | <0.0001 | 11.53 | 0.0008 | 0.007 (0.002) | 0.14 (0.04) | 3.39 | 0.0008 |
| AX-76351898 | Hum_BMD | WNT3 | 19.82 | <0.0001 | 77.58 | <0.0001 | 13.81 | 0.0002 | 0.016 (0.004) | 0.15 (0.04) | 3.72 | 0.0002 |
| AX-76351899 | Hum_BMD | WNT3 | 19.82 | <0.0001 | 77.58 | <0.0001 | 13.81 | 0.0002 | 0.016 (0.004) | 0.15 (0.04) | 3.72 | 0.0002 |
| SNP | Trait 1 | Candidate Gene | Genotypic Values | Homozygous Additive Allele Effect 5 | Dominance Effect 5 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| AA 2,3 (SE 4) | AB 2,3 (SE) | BB 2,3 (SE) | Estimate (SE) | t-Value | p-Value | Estimate (SE) | t-Value | p-Value | |||
| AX-76044166 | Tib_BBS | MPP7 | 155.33 (2.26) ab | 145.80 (5.85) b | 172.76 (7.25) a | −8.71 (4.05) | −2.15 | 0.0320 | −18.20 (5.45) | −3.35 | 0.0009 |
| AX-76601713 | Tib_BBS | ACTR2 | 162.77 (3.08) a | 156.79 (3.81) a | 143.10 (3.79) b | 9.83 (2.82) | 3.49 | 0.0005 | 3.86 (4.03) | 0.96 | 0.3392 |
| AX-77276717 | Tib_BBS | TGFB2 | 153.42 (2.25) a | 157.05 (3.06) a | 163.72 (5.10) a | −5.15 (2.83) | −1.82 | 0.0694 | −1.52 (3.73) | −0.41 | 0.6843 |
| AX-76491534 | Tib_BBS | CCDC170 | 159.13 (2.19) a | 144.04 (6.28) ab | 127.83 (8.09) b | 15.70 (4.42) | 3.54 | 0.0004 | 0.56 (5.88) | 0.096 | 0.9239 |
| AX-76772658 | Tib_BBS | SOX6 | 149.06 (2.65) b | 158.53 (2.58) a | 163.29 (3.90) a | −7.11 (2.43) | −2.93 | 0.0035 | 2.36 (3.13) | 0.75 | 0.4520 |
| AX-75711229 | Tib_BBS | ASPN | 155.14 (1.94) b | 154.78 (5.29) b | 188.53 (13.11) a | −16.70 (6.62) | −2.52 | 0.0120 | −17.10 (8.02) | −2.13 | 0.0340 |
| AX-75913642 | Tib_BBS | SOX9 | 157.50 (1.93) a | 148.13 (4.83) ab | 124.13 (10.37) b | 16.70 (5.31) | 3.14 | 0.0018 | 7.40 (6.44) | 1.15 | 0.2506 |
| AX-76772658 | Hum_BBS | SOX6 | 127.04 (2.51) a | 116.24 (2.46) b | 119.38 (3.71) ab | 3.83 (2.31) | 1.66 | 0.0984 | −6.96 (3.02) | −2.31 | 0.0215 |
| AX-77091655 | Hum_BBS | PODN | 118.01 (2.31) b | 120.73 (3.04) b | 132.21 (3.73) a | −7.10 (2.31) | −3.07 | 0.0023 | −4.38 (3.44) | −1.27 | 0.2043 |
| AX-75597497 | Hum_BBS | SMAD6 | 122.16 (2.08) a | 123.48 (3.64) a | 98.16 (6.97) b | 12.0 (3.71) | 3.23 | 0.0013 | 13.30 (4.61) | 2.88 | 0.0040 |
| AX-76351785 | Hum_BBS | WNT9B | 115.73 (2.19) c | 124.86 (3.05) b | 139.61 (4.34) a | −11.90 (2.54) | −4.70 | <0.0001 | −2.81 (3.49) | −0.80 | 0.4215 |
| AX-75268181 | Tib_BMD | MCF2L | 0.263 (0.003) a | 0.253 (0.005) a | 0.228 (0.008) b | 0.017 (0.004) | 3.92 | 0.0001 | 0.008 (0.006) | 1.35 | 0.1768 |
| AX-80813610 | Tib_BMD | CALCR | 0.256 (0.003) a | 0.258 (0.005) a | 0.273 (0.006) a | −0.009 (0.004) | −2.24 | 0.0257 | −0.007 (0.005) | −1.27 | 0.2051 |
| AX-76099065 | Tib_BMD | SFRP4 | 0.261 (0.003) ab | 0.265 (0.008) a | 0.235 (0.009) b | 0.013 (0.006) | 2.32 | 0.0206 | 0.018 (0.006) | 2.71 | 0.0071 |
| AX-77113061 | Tib_BMD | TMCO1 | 0.246 (0.005) a | 0.267 (0.004) a | 0.266 (0.004) a | −0.01 (0.004) | −2.82 | 0.0050 | 0.011 (0.004) | 2.51 | 0.0125 |
| AX-75677174 | Tib_BMD | GPATCH1 | 0.254 (0.003) b | 0.269 (0.005) a | 0.278 (0.007) a | −0.012 (0.004) | −3.05 | 0.0024 | 0.004 (0.005) | 0.56 | 0.5739 |
| AX-77091655 | Hum_BMD | PODN | 0.164 (0.002) b | 0.167 (0.003) b | 0.178 (0.003) a | −0.007 (0.002) | −3.53 | 0.0005 | −0.004 (0.003) | −1.25 | 0.2117 |
| AX-76351898 | Hum_BMD | WNT3 | 0.166 (0.002) b | 0.176 (0.006) b | 0.206 (0.010) a | −0.02 (0.005) | −3.84 | 0.0001 | −0.009 (0.007) | −1.29 | 0.1991 |
| AX-76351899 | Hum_BMD | WNT3 | 0.166 (0.002) b | 0.176 (0.006) b | 0.206 (0.010) a | −0.02 (0.005) | −3.84 | 0.0001 | −0.009 (0.007) | −1.29 | 0.1991 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jansen, S.; Baulain, U.; Habig, C.; Ramzan, F.; Schauer, J.; Schmitt, A.O.; Scholz, A.M.; Sharifi, A.R.; Weigend, A.; Weigend, S. Identification and Functional Annotation of Genes Related to Bone Stability in Laying Hens Using Random Forests. Genes 2021, 12, 702. https://doi.org/10.3390/genes12050702
Jansen S, Baulain U, Habig C, Ramzan F, Schauer J, Schmitt AO, Scholz AM, Sharifi AR, Weigend A, Weigend S. Identification and Functional Annotation of Genes Related to Bone Stability in Laying Hens Using Random Forests. Genes. 2021; 12(5):702. https://doi.org/10.3390/genes12050702
Chicago/Turabian StyleJansen, Simon, Ulrich Baulain, Christin Habig, Faisal Ramzan, Jens Schauer, Armin Otto Schmitt, Armin Manfred Scholz, Ahmad Reza Sharifi, Annett Weigend, and Steffen Weigend. 2021. "Identification and Functional Annotation of Genes Related to Bone Stability in Laying Hens Using Random Forests" Genes 12, no. 5: 702. https://doi.org/10.3390/genes12050702
APA StyleJansen, S., Baulain, U., Habig, C., Ramzan, F., Schauer, J., Schmitt, A. O., Scholz, A. M., Sharifi, A. R., Weigend, A., & Weigend, S. (2021). Identification and Functional Annotation of Genes Related to Bone Stability in Laying Hens Using Random Forests. Genes, 12(5), 702. https://doi.org/10.3390/genes12050702








