The Maze Pathway of Coevolution: A Critical Review over the Leishmania and Its Endosymbiotic History
Abstract
1. Introduction
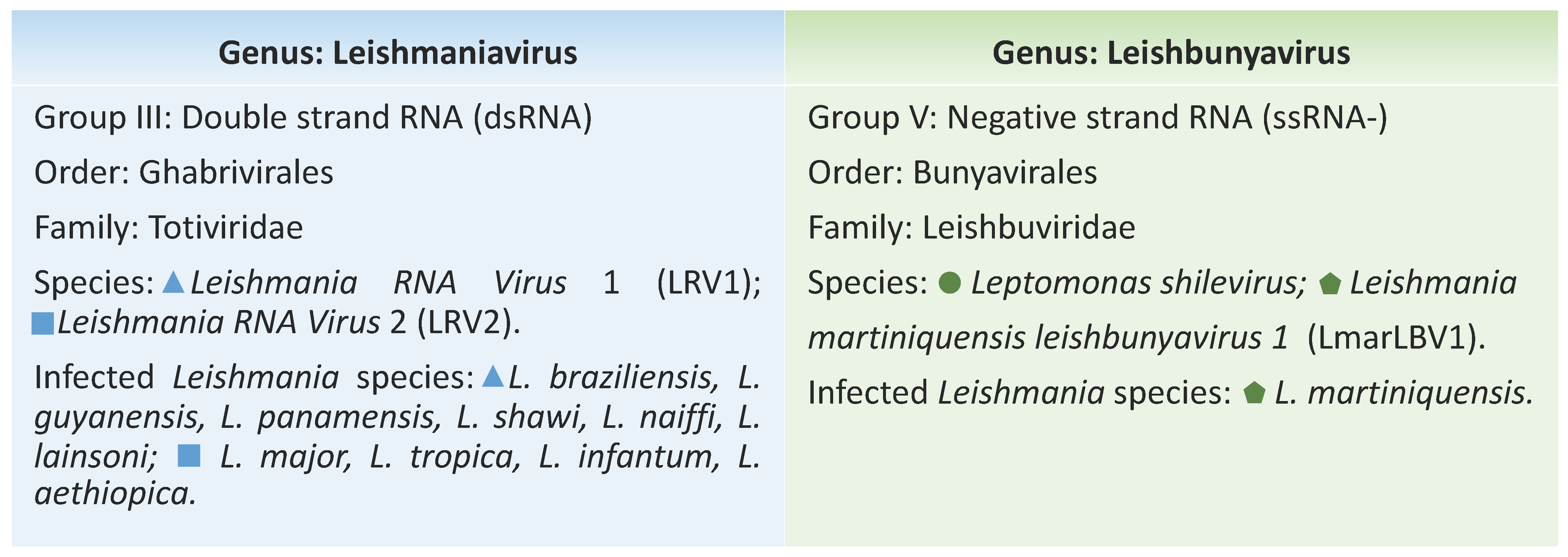
2. The Leishmania viruses
3. Exploiting Characteristics of Leishmania-Infecting Viruses
4. A Brief History of the Detection and Dispersion of LRV1, LRV2 and LBV
5. LRV and LBV Modulating Leishmania spp. Phenotypes
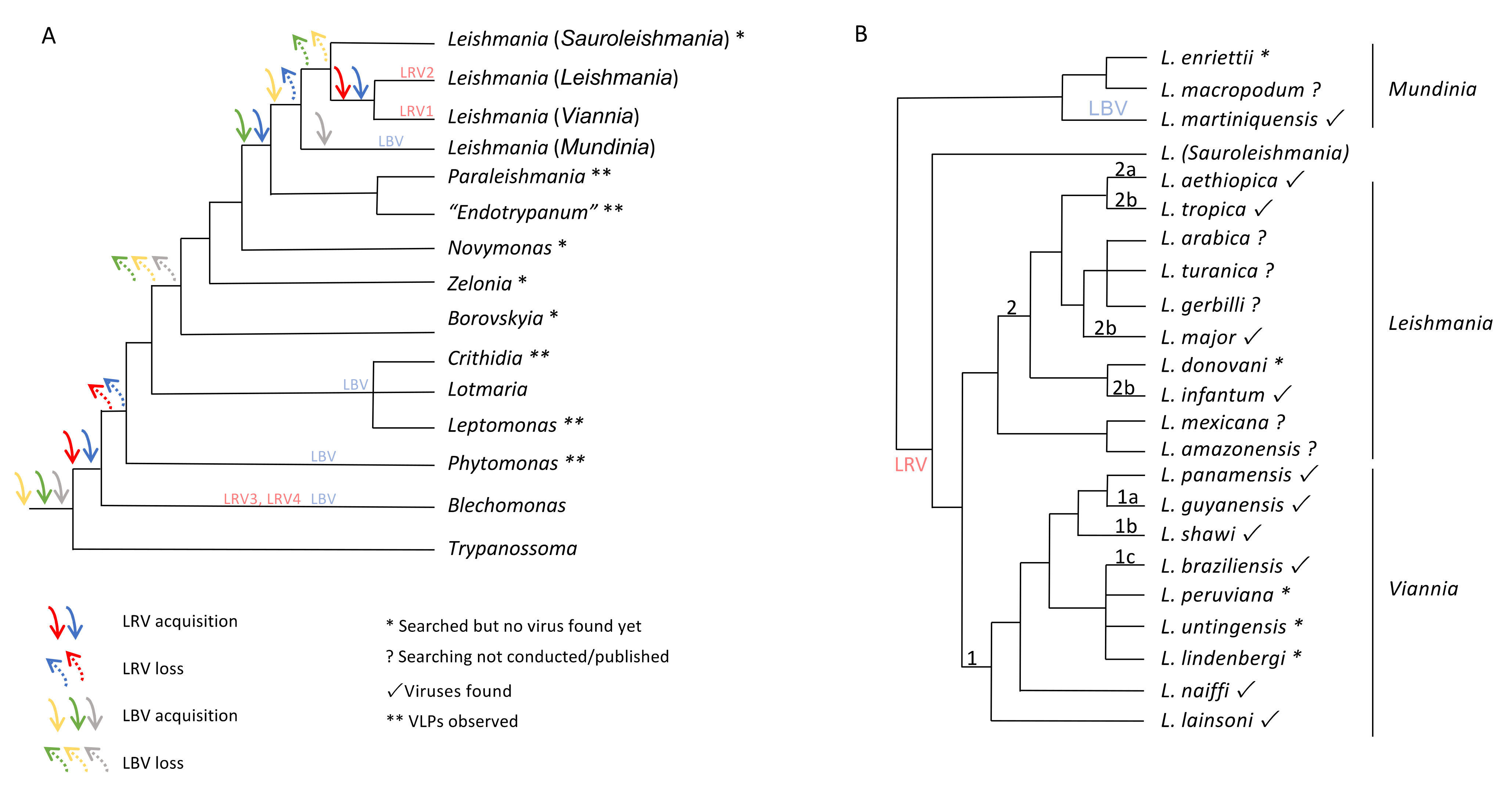
6. The Maze Pathway of Coevolution of Leishmania spp. and Its Viruses
7. Concluding Remarks
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Steverding, D. The history of leishmaniasis. Parasites Vectors 2017, 10, 1–10. [Google Scholar] [CrossRef]
- Kostygov, A.Y.; Yurchenko, V. Revised classification of the subfamily Leishmaniinae (Trypanosomatidae). Folia Parasitol. 2017, 64, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Espinosa, O.A.; Serrano, M.G.; Camargo, E.P.; Teixeira, M.M.G.; Shaw, J.J. An appraisal of the taxonomy and nomenclature of trypanosomatids presently classified as Leishmania and Endotrypanum. Parasitology 2018, 145, 430–442. [Google Scholar] [CrossRef]
- Serafim, T.D.; Coutinho-Abreu, I.V.; Oliveira, F.; Meneses, C.; Kamhawi, S.; Valenzuela, J.G. Sequential blood meals promote Leishmania replication and reverse metacyclogenesis augmenting vector infectivity. Nat. Microbiol. 2018, 3, 548–555. [Google Scholar] [CrossRef]
- Cupolillo, E.; Medina-Acosta, E.; Noyes, H.; Momen, H.; Grimaldi, G. A revised classification for Leishmania and Endotrypanum. Parasitol. Today 2000, 16, 142–144. [Google Scholar] [CrossRef]
- Jirků, M.; Yurchenko, V.Y.; Lukeš, J.; Maslov, D.A. New species of insect trypanosomatids from costa rica and the proposal for a new subfamily within the Trypanosomatidae. J. Eukaryot. Microbiol. 2012, 59, 537–547. [Google Scholar] [CrossRef] [PubMed]
- Croft, S.L.; Molyneux, D.H. Studies on the ultrastructure, virus-like particles and infectivity of Leishmania hertigi. Ann. Trop. Med. Parasitol. 1979, 73, 213–226. [Google Scholar] [CrossRef]
- Tarr, P.I.; Aline, R.F.; Smiley, B.L.; Scholler, J.; Keithly, J.; Stuart, K. LR1: A candidate RNA virus of Leishmania. Proc. Natl. Acad. Sci. USA 1988, 85, 9572–9575. [Google Scholar] [CrossRef]
- Widmer, G.; Comeau, A.M.; Furlong, D.B.; Wirth, D.F.; Patterson, J.L. Characterization of a RNA virus from the parasite Leishmania. Proc. Natl. Acad. Sci. USA 1989, 86, 5979–5982. [Google Scholar] [CrossRef]
- Weeks, R.S.; Patterson, J.L.; Stuart, K.; Widmer, G. Transcribing and replicating particles in a double-stranded RNA virus from Leishmania. Mol. Biochem. Parasitol. 1992, 52, 207–213. [Google Scholar] [CrossRef]
- Cadd, T.L.; MacBeth, K.; Furlong, D.; Patterson, J.L. Mutational analysis of the capsid protein of Leishmania RNA virus LRV1-4. J. Virol. 1994, 68, 7738–7745. [Google Scholar] [CrossRef] [PubMed]
- Ives, A.; Ronet, C.; Prevel, F.; Ruzzante, G.; Fuertes-Marraco, S.; Schutz, F.; Zangger, H.; Revaz-Breton, M.; Lye, L.-F.; Hickerson, S.M.; et al. Leishmania RNA virus controls the severity of mucocutaneous leishmaniasis. Science 2011, 331, 775–778. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.H.; Swartzwelder, J.C. Virus-like Particles in an Entamoeba histolytica Trophozoite. J. Parasitol. 1960, 46, 523. [Google Scholar] [CrossRef]
- Walker, P.J.; Siddell, S.G.; Lefkowitz, E.J.; Mushegian, A.R.; Dempsey, D.M.; Dutilh, B.E.; Harrach, B.; Harrison, R.L.; Hendrickson, R.C.; Junglen, S.; et al. Changes to virus taxonomy and the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses (2019). Arch. Virol. 2019, 164, 2417–2429. [Google Scholar] [CrossRef] [PubMed]
- Grybchuk, D.; Akopyants, N.S.; Kostygov, A.Y.; Konovalovas, A.; Lye, L.F.; Dobson, D.E.; Zangger, H.; Fasel, N.; Butenko, A.; Frolov, A.O.; et al. Viral discovery and diversity in trypanosomatid protozoa with a focus on relatives of the human parasite Leishmania. Proc. Natl. Acad. Sci. USA 2018, 115, E506–E515. [Google Scholar] [CrossRef]
- Grybchuk, D.; MacEdo, D.H.; Kleschenko, Y.; Kraeva, N.; Lukashev, A.N.; Bates, P.A.; Kulich, P.; Leštinová, T.; Volf, P.; Kostygov, A.Y.; et al. The first Non-LRV RNA virus in leishmania. Viruses 2020, 12. [Google Scholar] [CrossRef]
- Butenko, A.; Kostygov, A.Y.; Sádlová, J.; Kleschenko, Y.; Bečvář, T.; Podešvová, L.; MacEdo, D.H.; Žihala, D.; Lukeš, J.; Bates, P.A.; et al. Comparative genomics of Leishmania (Mundinia). Bmc Genom. 2019, 20, 1–12. [Google Scholar] [CrossRef]
- Zangger, H.; Hailu, A.; Desponds, C.; Lye, L.-F.; Akopyants, N.S.; Dobson, D.E.; Ronet, C.; Ghalib, H.; Beverley, S.M.; Fasel, N. Leishmania aethiopica field isolates bearing an endosymbiontic dsRNA virus induce pro-inflammatory cytokine response. PLoS Negl. Trop. Dis. 2014, 8, e2836. [Google Scholar] [CrossRef]
- Salinas, G.; Zamora, M.; Stuart, K.; Saravia, N. Leishmania RNA viruses in Leishmania of the Viannia subgenus. Am. J. Trop. Med. Hyg. 1996, 54, 425–429. [Google Scholar] [CrossRef]
- Cantanhêde, L.M.; da Silva Júnior, C.F.; Ito, M.M.; Felipin, K.P.; Nicolete, R.; Salcedo, J.M.V.; Porrozzi, R.; Cupolillo, E.; de Godoi Mattos Ferreira, R. Further Evidence of an Association between the Presence of Leishmania RNA Virus 1 and the Mucosal Manifestations in Tegumentary Leishmaniasis Patients. Plos Negl. Trop. Dis. 2015, 9, e0004079. [Google Scholar] [CrossRef]
- Scheffter, S.M.; Ro, Y.T.; Chung, I.K.; Patterson, J.L. The complete sequence of Leishmania RNA virus LRV2-1, a virus of an Old World parasite strain. Virology 1995, 212, 84–90. [Google Scholar] [CrossRef]
- Hajjaran, H.; Mahdi, M.; Mohebali, M.; Samimi-Rad, K.; Ataei-Pirkooh, A.; Kazemi-Rad, E.; Naddaf, S.R.; Raoofian, R. Detection and molecular identification of leishmania RNA virus (LRV) in Iranian Leishmania species. Arch. Virol. 2016, 161, 3385–3390. [Google Scholar] [CrossRef]
- Stuart, K.D.; Weeks, R.; Guilbride, L.; Myler, P.J. Molecular organization of Leishmania RNA virus 1. Proc. Natl. Acad. Sci. USA 1992, 89, 8596–8600. [Google Scholar] [CrossRef]
- Widmer, G.; Patterson, J.L. Genomic structure and RNA polymerase activity in Leishmania virus. J. Virol. 1991, 65, 4211–4215. [Google Scholar] [CrossRef]
- Ghabrial, S.A. University of K. Totivirus. In Encyclopedia of Virology; Elsevier Ltd.: Lexington, KY, USA, 2008; pp. 163–174. [Google Scholar]
- Allen, A.; Islamovic, E.; Kaur, J.; Gold, S.; Shah, D.; Smith, T.J. Transgenic maize plants expressing the Totivirus antifungal protein, KP4, are highly resistant to corn smut. Plant Biotechnol. J. 2011, 9, 857–864. [Google Scholar] [CrossRef] [PubMed]
- Provenzano, D.; Khoshnan, A.; Alderete, J.F. Involvement of dsRNA virus in the protein composition and growth kinetics of host Trichomonas vaginalis. Arch. Virol. 1997, 142, 939–952. [Google Scholar] [CrossRef] [PubMed]
- Ro, Y.; Patterson, J.L. Identification of the Minimal Essential RNA Sequences Responsible for Site-Specific Targeting of the Leishmania RNA Virus 1-4 Capsid Endoribonuclease Identification of the Minimal Essential RNA Sequences Responsible for Site-Specific Targeting of the Leis. J. Virol. 2000. [Google Scholar] [CrossRef] [PubMed]
- MacBeth, K.J.; Ro, Y.T.; Gehrke, L.; Patterson, J.L. Cleavage site mapping and substrate-specificity of Leishmaniavirus 2-1 capsid endoribonuclease activity. J. Biochem. 1997, 122, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Saiz, M.; Ro, Y.T.; Wirth, D.F.; Patterson, J.L. Host cell proteins bind specifically to the capsid-cleaved 5′ end of Leishmaniavirus RNA. J. Biochem. 1999, 126, 538–544. [Google Scholar] [CrossRef] [PubMed]
- Patterson, J.L. Viruses of protozoan parasites. Exp. Parasitol. 1990, 70, 111–113. [Google Scholar] [CrossRef]
- Scheffter, S.; Widmer, G.; Patterson, J.L. Complete sequence of Leishmania RNA virus 1-4 and identification of conserved sequences. Virology 1994, 199, 479–483. [Google Scholar] [CrossRef]
- Lefkowitz, E.J.; Dempsey, D.M.; Hendrickson, R.C.; Orton, R.J.; Siddell, S.G.; Smith, D.B. Virus taxonomy: The database of the International Committee on Taxonomy of Viruses (ICTV). Nucleic Acids Res. 2018, 46, D708–D717. [Google Scholar] [CrossRef]
- Wichgers Schreur, P.J.; Kormelink, R.; Kortekaas, J. Genome packaging of the Bunyavirales. Curr. Opin. Virol. 2018, 33, 151–155. [Google Scholar] [CrossRef]
- Sun, Y.; Li, J.; Gao, G.F.; Tien, P.; Liu, W. Bunyavirales ribonucleoproteins: The viral replication and transcription machinery. Crit. Rev. Microbiol. 2018, 44, 522–540. [Google Scholar] [CrossRef] [PubMed]
- Elliott, R.M. Molecular biology of the Bunyaviridae. J. Gen. Virol. 1990, 71, 501–522. [Google Scholar] [CrossRef] [PubMed]
- Akopyants, N.S.; Lye, L.F.; Dobson, D.E.; Lukeš, J.; Beverley, S.M. A novel bunyavirus-like virus of trypanosomatid protist parasites. Genome Announc. 2016, 4, 4–5. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Carthew, R.W.; Sontheimer, E.J. Origins and Mechanisms of miRNAs and siRNAs. Cell 2009, 136, 642–655. [Google Scholar] [CrossRef] [PubMed]
- Gitlin, L.; Andino, R. Nucleic Acid-Based Immune System: The Antiviral Potential of Mammalian RNA Silencing. J. Virol. 2003, 77, 7159–7165. [Google Scholar] [CrossRef]
- Maga, J.A.; Widmer, G.; LeBowitz, J.H. Leishmania RNA virus 1-mediated cap-independent translation. Mol. Cell. Biol. 1995, 15, 4884–4889. [Google Scholar] [CrossRef]
- Guilbride, L.; Myler, P.J.; Stuart, K. Distribution and sequence divergence of LRV1 viruses among different Leishmania species. Mol. Biochem. Parasitol. 1992, 54, 101–104. [Google Scholar] [CrossRef]
- Widmer, G.; Dooley, S. Phylogenetic analysis of Leishmania RNA virus and leishmania suggests ancient virus-parasite association. Nucleic Acids Res. 1995, 23, 2300–2304. [Google Scholar] [CrossRef] [PubMed]
- Pereira, L.D.O.R.; Maretti-Mira, A.C.; Rodrigues, K.M.; Lima, R.B.; de Oliveira-Neto, M.P.; Cupolillo, E.; Pirmez, C.; de Oliveira, M.P. Severity of tegumentary leishmaniasis is not exclusively associated with Leishmania RNA virus 1 infection in Brazil. Memórias Do Inst. Oswaldo Cruz 2013, 108, 665–667. [Google Scholar] [CrossRef]
- Macedo, D.H.; Menezes-Neto, A.; Rugani, J.M.; Rocha, A.C.; Silva, S.O.; Melo, M.N.; Lye, L.F.; Beverley, S.M.; Gontijo, C.M.; Soares, R.P. Low frequency of LRV1 in Leishmania braziliensis strains isolated from typical and atypical lesions in the State of Minas Gerais, Brazil. Mol. Biochem. Parasitol. 2016, 210, 50–54. [Google Scholar] [CrossRef] [PubMed]
- Cantanhêde, L.M.; Fernandes, F.G.; Eduardo Melim Ferreira, G.; Porrozzi, R.; De Godoi Mattos Ferreira, R.; Cupolillo, E. New insights into the genetic diversity of Leishmania RNA Virus 1 and its species-specific relationship with Leishmania parasites. PLoS ONE 2018, 13, 1–16. [Google Scholar] [CrossRef]
- Vieira-Gonçalves, R.; Fagundes-Silva, G.A.; Heringer, J.F.; Fantinatti, M.; Da-Cruz, A.M.; Oliveira-Neto, M.P.; Guerra, J.A.O.; Gomes-Silva, A. First report of treatment failure in a patient with cutaneous leishmaniasis infected by Leishmania (Viannia) naiffi carrying Leishmania RNA virus: A fortuitous combination? Rev. Da Soc. Bras. De Med. Trop. 2019, 52, 10–12. [Google Scholar] [CrossRef]
- Kariyawasam, R.; Mukkala, A.N.; Lau, R.; Valencia, B.M.; Llanos-Cuentas, A.; Boggild, A.K. Virulence factor RNA transcript expression in the Leishmania Viannia subgenus: Influence of species, isolate source, and Leishmania RNA virus-1. Trop. Med. Health 2019, 47, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kleschenko, Y.; Grybchuk, D.; Matveeva, N.S.; Macedo, D.H.; Ponirovsky, E.N.; Lukashev, A.N.; Yurchenko, V. Molecular Characterization of Leishmania RNA virus 2 in Leishmaniamajor from Uzbekistan. Genes 2019, 10, 830. [Google Scholar] [CrossRef] [PubMed]
- Saberi, R.; Fakhar, M.; Hajjaran, H.; Ataei-Pirkooh, A.; Mohebali, M.; Taghipour, N.; Ziaei Hezarjaribi, H.; Moghadam, Y.; Bagheri, A. Presence and diversity of Leishmania RNA virus in an old zoonotic cutaneous leishmaniasis focus, northeastern Iran: Haplotype and phylogenetic based approach. Int. J. Infect. Dis. 2020, 101, 6–13. [Google Scholar] [CrossRef]
- Nalçacı, M.; Karakuş, M.; Yılmaz, B.; Demir, S.; Özbilgin, A.; Özbel, Y.; Töz, S. Detection of Leishmania RNA virus 2 in Leishmania species from Turkey. Trans. R. Soc. Trop. Med. Hyg. 2019, 113, 410–417. [Google Scholar] [CrossRef]
- Lukeš, J.; Butenko, A.; Hashimi, H.; Maslov, D.A.; Votýpka, J.; Yurchenko, V. Trypanosomatids Are Much More than Just Trypanosomes: Clues from the Expanded Family Tree. Trends Parasitol. 2018, 34, 466–480. [Google Scholar] [CrossRef]
- Harkins, K.M.; Schwartz, R.S.; Cartwright, R.A.; Stone, A.C. Phylogenomic reconstruction supports supercontinent origins for Leishmania. Infect. Genet. Evol. 2016, 38, 101–109. [Google Scholar] [CrossRef]
- Ogg, M.M.; Carrion, R.; Botelho, A.C.D.C.; Mayrink, W.; Correa-Oliveira, R.; Patterson, J.L. Short report: Quantification of leishmaniavirus RNA in clinical samples and its possible role in pathogenesis. Am. J. Trop. Med. Hyg. 2003, 69, 309–313. [Google Scholar] [CrossRef] [PubMed]
- Hartley, M.-A.; Ronet, C.; Zangger, H.; Beverley, S.M.; Fasel, N. Leishmania RNA virus: When the host pays the toll. Front. Cell. Infect. Microbiol. 2012, 2, 99. [Google Scholar] [CrossRef] [PubMed]
- Bourreau, E.; Ginouves, M.; Prévot, G.; Hartley, M.-A.; Gangneux, J.-P.; Robert-Gangneux, F.; Dufour, J.; Sainte-Marie, D.; Bertolotti, A.; Pratlong, F.; et al. Leishmania-RNA virus presence in L. guyanensis parasites increases the risk of first-line treatment failure and symptomatic relapse. J. Infect. Dis. 2015; 1–28. [Google Scholar] [CrossRef]
- Adaui, V.; Lye, L.; Akopyants, N.S.; Zimic, M.; Llanos-cuentas, A.; Garcia, L.; Maes, I.; Doncker, S. De Association of the Endobiont Double-Stranded RNA Virus LRV1 With Treatment Failure for Human Leishmaniasis Caused by Leishmania braziliensis in Peru and Bolivia. J. Infect. Dis. 2016, 213. [Google Scholar] [CrossRef]
- Abtahi, M.; Eslami, G.; Cavallero, S.; Vakili, M.; Hosseini, S.S.; Ahmadian, S.; Boozhmehrani, M.J.; Khamesipour, A. Relationship of Leishmania RNA Virus (LRV) and treatment failure in clinical isolates of Leishmania major. BMC Res. Notes 2020, 13, 1–6. [Google Scholar] [CrossRef]
- Ginouvès, M.; Couppié, P.; Simon, S.; Bourreau, E.; Rogier, S.; Brousse, P.; Travers, P.; Pommier de Santi, V.; Demar, M.; Briolant, S.; et al. Leishmania virus genetic diversity is not related to leishmaniasis treatment failure. Clin. Microbiol. Infect. 2020. [Google Scholar] [CrossRef]
- Choisy, M.; Hide, M.; Bañuls, A.-L.; Guégan, J.-F. Rocking the curve. Trends Microbiol. 2004, 12, 534–536. [Google Scholar] [CrossRef] [PubMed]
- Mendes, B.P.; Da Silva, I.A.; Damata, J.P.; Castro-Gomes, T.; Vieira, L.Q.; Ribeiro-Dias, F.; Horta, M.F. Metacyclogenesis of Leishmania (Viannia) guyanensis: A comprehensive study of the main transformation features in axenic culture and purification of metacyclic promastigotes by negative selection with Bauhinia purpurea lectin. Parasitology 2019, 716–727. [Google Scholar] [CrossRef] [PubMed]
- Zangger, H.; Ronet, C.; Desponds, C.; Kuhlmann, F.M.; Robinson, J.; Hartley, M.-A.; Prevel, F.; Castiglioni, P.; Pratlong, F.; Bastien, P.; et al. Detection of Leishmania RNA virus in Leishmania parasites. PLoS Negl. Trop. Dis. 2013, 7, e2006. [Google Scholar] [CrossRef]
- Tirera, S.; Ginouves, M.; Donato, D.; Caballero, I.S.; Bouchier, C.; Lavergne, A.; Bourreau, E.; Mosnier, E.; Vantilcke, V.; Couppié, P.; et al. Unraveling the genetic diversity and phylogeny of Leishmania RNA virus 1 strains of infected Leishmania isolates circulating in French Guiana. PLoS Negl. Trop. Dis. 2017, 11, 1–20. [Google Scholar] [CrossRef]
- Silvester, E.; Young, J.; Ivens, A.; Matthews, K.R. Europe PMC Funders Group Interspecies quorum-sensing in co-infections can manipulate trypanosome transmission potential. Interspecies 2018, 2, 1471–1479. [Google Scholar] [CrossRef]
- Schuh, C.M.A.P.; Aguayo, S.; Zavala, G.; Khoury, M. Exosome-like vesicles in Apis mellifera bee pollen, honey and royal jelly contribute to their antibacterial and pro-regenerative activity. J. Exp. Biol. 2019, 222. [Google Scholar] [CrossRef] [PubMed]
- Atayde, V.D.; da Silva Lira Filho, A.; Chaparro, V.; Zimmermann, A.; Martel, C.; Jaramillo, M.; Olivier, M. Exploitation of the Leishmania exosomal pathway by Leishmania RNA virus 1. Nat. Microbiol. 2019, 4, 714–723. [Google Scholar] [CrossRef] [PubMed]
- Atayde, V.D.; Aslan, H.; Townsend, S.; Hassani, K.; Kamhawi, S.; Olivier, M. Exosome Secretion by the Parasitic Protozoan Leishmania within the Sand Fly Midgut. Cell Rep. 2015, 13, 957–967. [Google Scholar] [CrossRef] [PubMed]
- Castiglioni, P.; Hartley, M.-A.; Rossi, M.; Prevel, F.; Desponds, C.; Utzschneider, D.T.; Eren, R.-O.; Zangger, H.; Brunner, L.; Collin, N.; et al. Exacerbated Leishmaniasis Caused by a Viral Endosymbiont can be Prevented by Immunization with Its Viral Capsid. PLoS Negl. Trop. Dis. 2017, 11, e0005240. [Google Scholar] [CrossRef] [PubMed]
- Ronet, C.; Beverley, S.M.; Fasel, N. Muco-cutaneous leishmaniasis in the New World: The ultimate subversion. Virulence 2011, 2, 547–552. [Google Scholar] [CrossRef]
- de Carvalho, R.V.H.; Lima-Junior, D.S.; da Silva, M.V.G.; Dilucca, M.; Rodrigues, T.S.; Horta, C.V.; Silva, A.L.N.; da Silva, P.F.; Frantz, F.G.; Lorenzon, L.B.; et al. Leishmania RNA virus exacerbates Leishmaniasis by subverting innate immunity via TLR3-mediated NLRP3 inflammasome inhibition. Nat. Commun. 2019, 10. [Google Scholar] [CrossRef]
- Baneth, G.; Zivotofsky, D.; Nachum-Biala, Y.; Yasur-Landau, D.; Botero, A.M. Mucocutaneous Leishmania tropica infection in a dog from a human cutaneous leishmaniasis focus. Parasites Vectors 2014, 7, 1–5. [Google Scholar] [CrossRef]
- Shirian, S.; Oryan, A.; Hatam, G.R.; Daneshbod, Y. Mixed mucosal leishmaniasis infection caused by Leishmania tropica and leishmania major. J. Clin. Microbiol. 2012, 50, 3805–3808. [Google Scholar] [CrossRef]
- Morsy, T.A.; Khalil, N.M.; Salama, M.M.; Hamdi, K.N.; al Shamrany, Y.A.; Abdalla, K.F. Mucosal leishmaniasis caused by Leishmania tropica in Saudi Arabia. J. Egypt. Soc. Parasitol. 1995, 25, 73–79. [Google Scholar] [PubMed]
- Strazzulla, A.; Cocuzza, S.; Pinzone, M.R.; Postorino, M.C.; Cosentino, S.; Serra, A.; Cacopardo, B.; Nunnari, G. Mucosal leishmaniasis: An underestimated presentation of a neglected disease. Biomed Res. Int. 2013, 2013. [Google Scholar] [CrossRef] [PubMed]
- Schönian, G.; Akuffo, H.; Lewin, S.; Maasho, K.; Nylén, S.; Pratlong, F.; Eisenberger, C.L.; Schnur, L.F.; Presber, W. Genetic variability within the species Leishmania aethiopica does not correlate with clinical variations of cutaneous leishmaniasis. Mol. Biochem. Parasitol. 2000, 106, 239–248. [Google Scholar] [CrossRef]
- Dalzoto, P.R.; Glienke-Blanco, C.; Kava-Cordeiro, V.; Ribeiro, J.Z.; Kitajima, E.W.; Azevedo, J.L. Horizontal transfer and hypovirulence associated with double-stranded RNA in Beauveria bassiana. Mycol. Res. 2006, 110, 1475–1481. [Google Scholar] [CrossRef]
- Armstrong, T.C.; Keenan, M.C.; Widmer, G.; Patterson, J.L. Successful transient introduction of Leishmania RNA virus into a virally infected and an uninfected strain of Leishmania. Proc. Natl. Acad. Sci. USA 1993, 90, 1736–1740. [Google Scholar] [CrossRef]
- Ro, Y.; Scheffter, S.M.; Patterson, J.L. Specific in vitro cleavage of a Leishmania virus capsid-RNA-dependent RNA polymerase polyprotein by a host cysteine-like protease. J. Virol. 1997, 71, 8983–8990. [Google Scholar] [CrossRef]
- Maslov, D.A.; Votýpka, J.; Yurchenko, V.; Lukeš, J. Diversity and phylogeny of insect trypanosomatids: All that is hidden shall be revealed. Trends Parasitol. 2013, 29, 43–52. [Google Scholar] [CrossRef]
- Moriconi, M.; Rugna, G.; Calzolari, M.; Bellini, R.; Albieri, A.; Angelini, P.; Cagarelli, R.; Landini, M.P.; Charrel, R.N.; Varani, S. Phlebotomine sand fly–borne pathogens in the Mediterranean Basin: Human leishmaniasis and phlebovirus infections. Plos Negl. Trop. Dis. 2017, 11, 1–19. [Google Scholar] [CrossRef]
- da Silva, L.A.; de Sousa, C.d.S.; da Graça, G.C.; Porrozzi, R.; Cupolillo, E. Sequence analysis and PCR-RFLP profiling of the hsp70 gene as a valuable tool for identifying Leishmania species associated with human leishmaniasis in Brazil. Infect. Genet. Evol. 2010, 10, 77–83. [Google Scholar] [CrossRef]
- Boité, M.C.; Mauricio, I.L.; Miles, M.A.; Cupolillo, E. New Insights on Taxonomy, Phylogeny and Population Genetics of Leishmania (Viannia) Parasites Based on Multilocus Sequence Analysis. PLoS Negl. Trop. Dis. 2012, 6. [Google Scholar] [CrossRef]
- Cupolillo, E.; Grimaldi, G.; Momen, H. A General Classification of New World Leishmania Using Numerical Zymotaxonomy. Am. J. Trop. Med. Hyg. 1994, 50, 296–311. [Google Scholar] [CrossRef] [PubMed]
- Romero, G.A.S.; Ishikawa, E.; Cupolillo, E.; Toaldo, C.B.; Guerra, M.V.; Vinitius de Farias Guerra, M.; Gomes Paes, M.; de Oliveira Macêdo, M.V.; Shaw, J.J. Identification of antigenically distinct populations of Leishmania (Viannia) guyanensis from Manaus, Brazil, using monoclonal antibodies. Acta Trop. 2002, 82, 25–29. [Google Scholar] [CrossRef]
- Grimaldi, G.; Momen, H.; Naiff, R.D.; McMahon-Pratt, D.; Barrett, T.V. Characterization and classification of leishmanial parasites from humans, wild mammals, and sand flies in the Amazon region of Brazil. Am. J. Trop. Med. Hyg. 1991, 44, 645–661. [Google Scholar] [CrossRef] [PubMed]
- Oddone, R.; Schweynoch, C.; Schönian, G.; De Sousa, C.D.S.; Cupolillo, E.; Espinosa, D.; Arevalo, J.; Noyes, H.; Mauricio, I.; Kuhls, K. Development of a multilocus microsatellite typing approach for discriminating strains of Leishmania (Viannia) species. J. Clin. Microbiol. 2009, 47, 2818–2825. [Google Scholar] [CrossRef]
- Kuhls, K.; Cupolillo, E.; Silva, S.O.; Schweynoch, C.; Boité, M.C.; Mello, M.N.; Mauricio, I.; Miles, M.; Wirth, T.; Schönian, G. Population Structure and Evidence for Both Clonality and Recombination among Brazilian Strains of the Subgenus Leishmania (Viannia). PLoS Negl. Trop. Dis. 2013, 7. [Google Scholar] [CrossRef] [PubMed]
- Gomes, R.F.; Macedo, A.M.; Pena, S.D.J.; Melo, M.N. Leishmania (Viannia) braziliensis: Genetic Relationships between Strains Isolated from Different Areas of Brazil as Revealed by DNA Fingerprinting and RAPD. Exp. Parasitol. 1995, 80, 681–687. [Google Scholar] [CrossRef]
- Rougeron, V.; De Meeûs, T.; Hide, M.; Waleckx, E.; Bermudez, H.; Arevalo, J.; Llanos-Cuentas, A.; Dujardin, J.C.; De Doncker, S.; Le Ray, D.; et al. Extreme inbreeding in Leishmania braziliensis. Proc. Natl. Acad. Sci. USA 2009, 106, 10224–10229. [Google Scholar] [CrossRef]
- Odiwuor, S.; Veland, N.; Maes, I.; Arévalo, J.; Dujardin, J.C.; Van der Auwera, G. Evolution of the Leishmania braziliensis species complex from amplified fragment length polymorphisms, and clinical implications. Infect. Genet. Evol. 2012, 12, 1994–2002. [Google Scholar] [CrossRef]
- Gomez, P.; Ashby, B.; Buckling, A. Population mixing promotes arms race host-parasite coevolution. Proc. R. Soc. B Biol. Sci. 2014, 282, 20142297. [Google Scholar] [CrossRef]
- Gandon, S.; Buckling, A.; Decaestecker, E.; Day, T. Host-parasite coevolution and patterns of adaptation across time and space. J. Evol. Biol. 2008, 21, 1861–1866. [Google Scholar] [CrossRef]
- Sasaki, A. Host-parasite coevolution in a multilocus gene-for-gene system. Proc. R. Soc. B Biol. Sci. 2000, 267, 2183–2188. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cantanhêde, L.M.; Mata-Somarribas, C.; Chourabi, K.; Pereira da Silva, G.; Dias das Chagas, B.; de Oliveira R. Pereira, L.; Côrtes Boité, M.; Cupolillo, E. The Maze Pathway of Coevolution: A Critical Review over the Leishmania and Its Endosymbiotic History. Genes 2021, 12, 657. https://doi.org/10.3390/genes12050657
Cantanhêde LM, Mata-Somarribas C, Chourabi K, Pereira da Silva G, Dias das Chagas B, de Oliveira R. Pereira L, Côrtes Boité M, Cupolillo E. The Maze Pathway of Coevolution: A Critical Review over the Leishmania and Its Endosymbiotic History. Genes. 2021; 12(5):657. https://doi.org/10.3390/genes12050657
Chicago/Turabian StyleCantanhêde, Lilian Motta, Carlos Mata-Somarribas, Khaled Chourabi, Gabriela Pereira da Silva, Bruna Dias das Chagas, Luiza de Oliveira R. Pereira, Mariana Côrtes Boité, and Elisa Cupolillo. 2021. "The Maze Pathway of Coevolution: A Critical Review over the Leishmania and Its Endosymbiotic History" Genes 12, no. 5: 657. https://doi.org/10.3390/genes12050657
APA StyleCantanhêde, L. M., Mata-Somarribas, C., Chourabi, K., Pereira da Silva, G., Dias das Chagas, B., de Oliveira R. Pereira, L., Côrtes Boité, M., & Cupolillo, E. (2021). The Maze Pathway of Coevolution: A Critical Review over the Leishmania and Its Endosymbiotic History. Genes, 12(5), 657. https://doi.org/10.3390/genes12050657





