Hybridogenesis in the Water Frogs from Western Russian Territory: Intrapopulation Variation in Genome Elimination
Abstract
1. Introduction
2. Materials and Methods
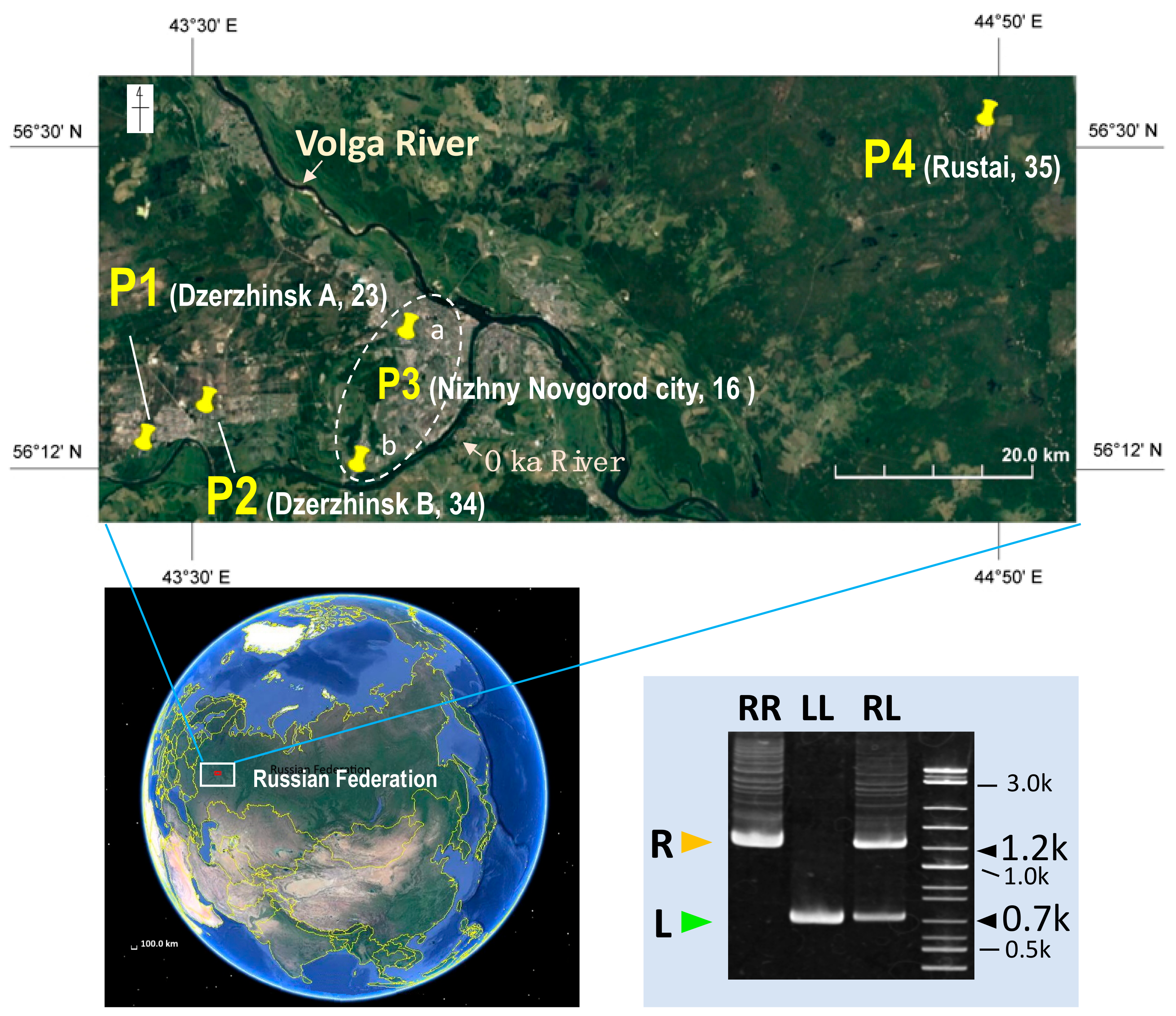
2.1. Collecting Frogs and Taxonomic Identification
2.2. Artificial Crossing and Microscopic Observation of Gonads
2.3. Cytochrome b and Serum albumin Gene Analyses
3. Results
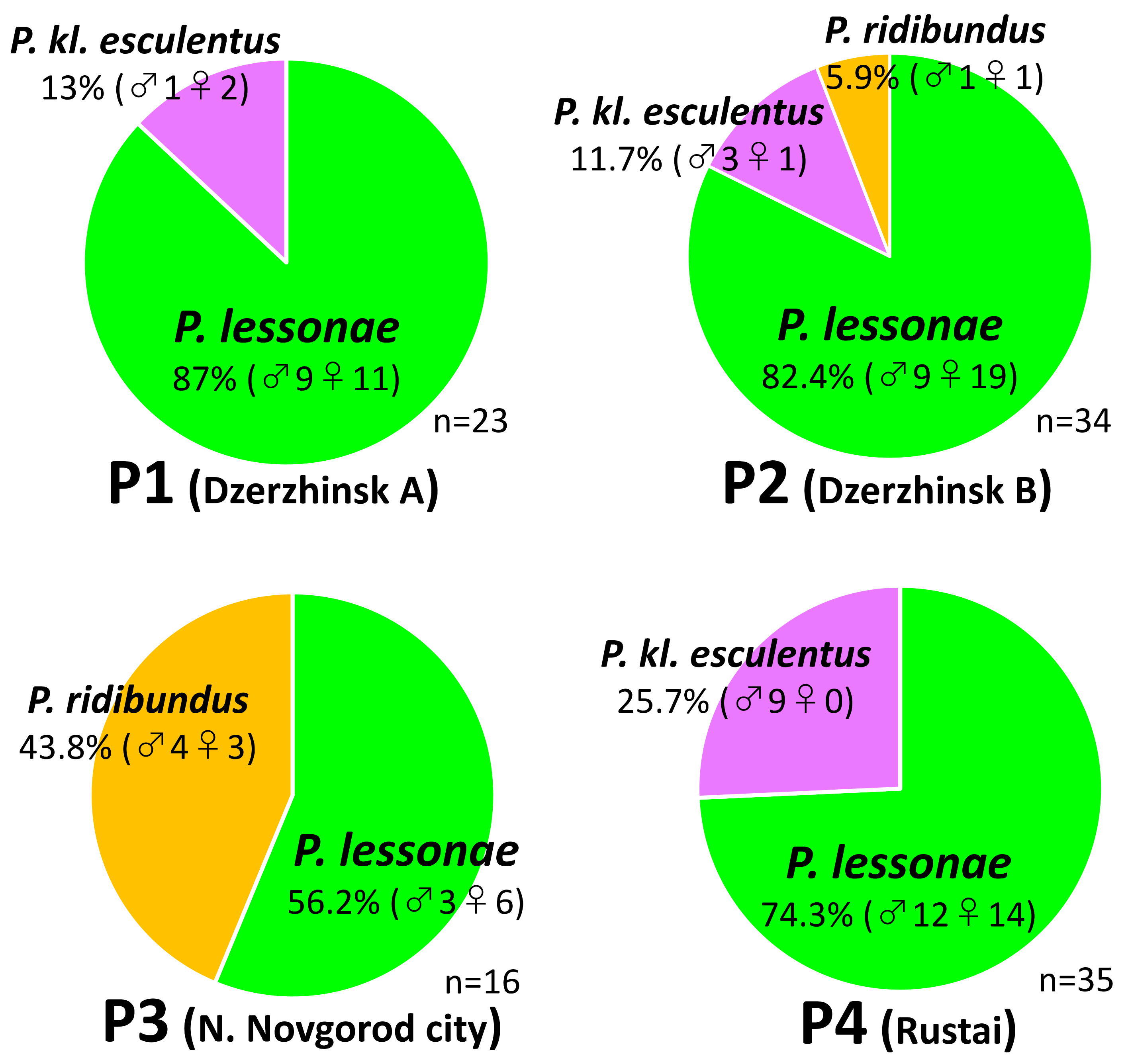
3.1. Taxonomic Composition
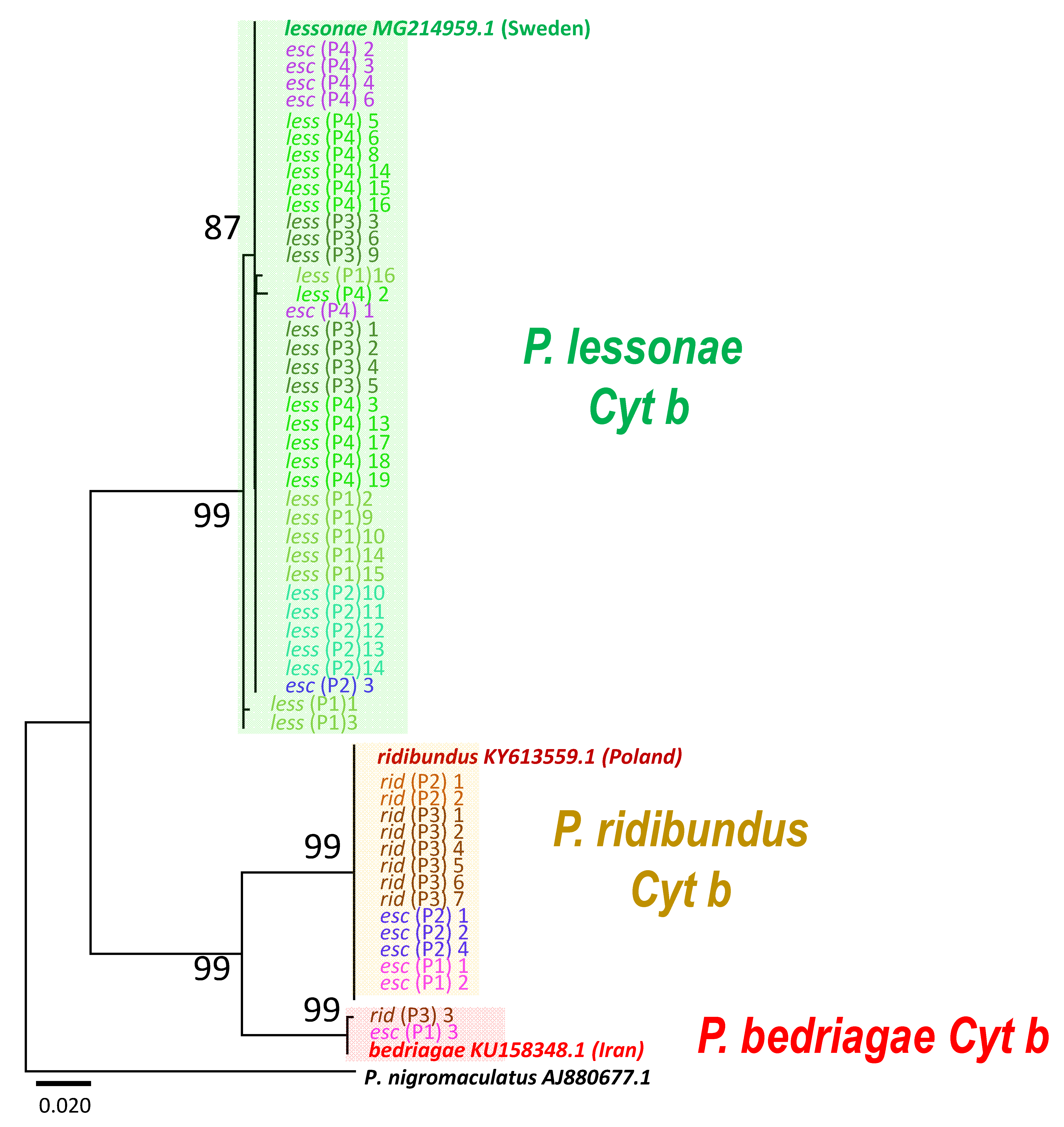
3.2. Mitochondrial Origins
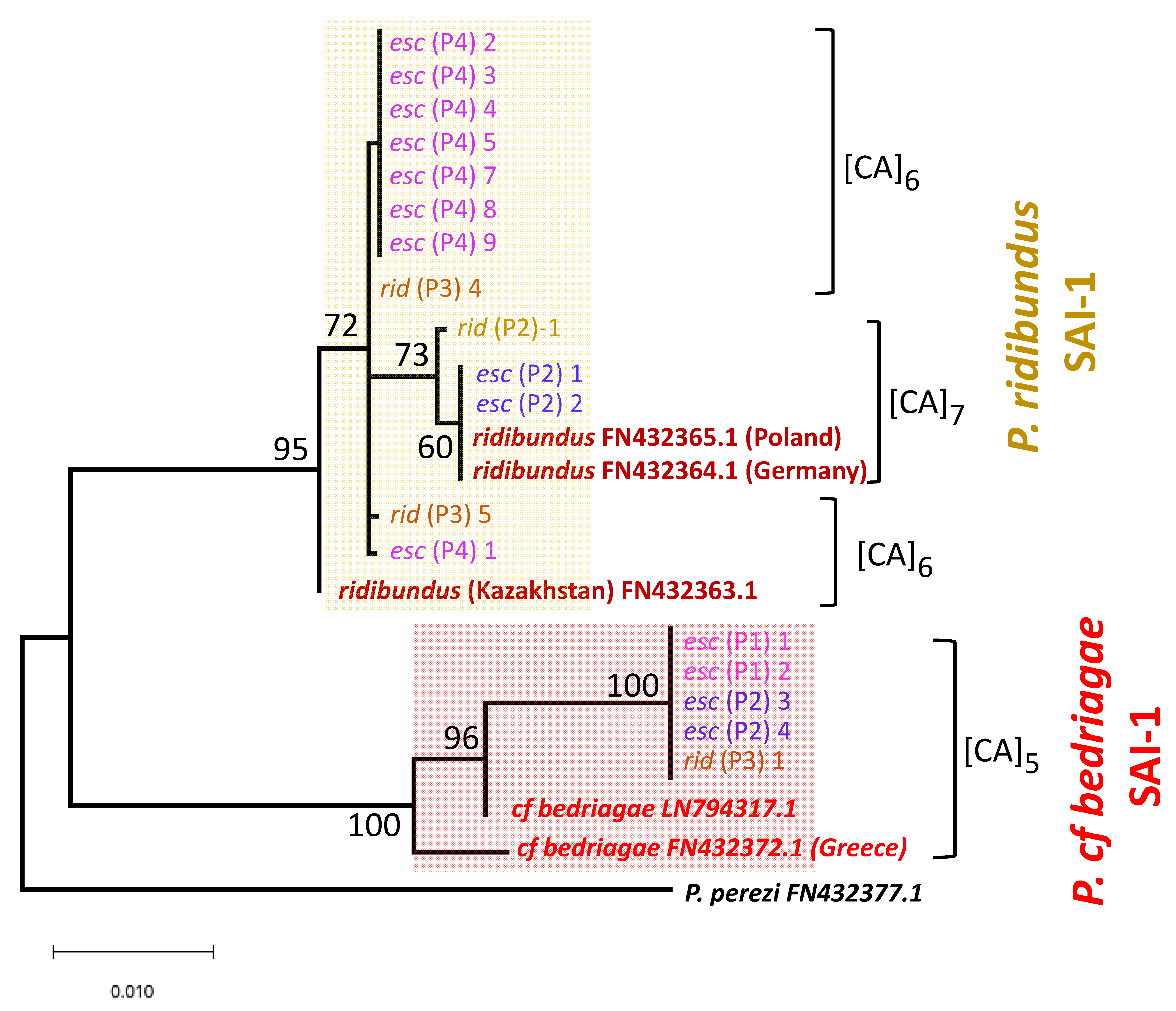
3.3. Nuclear Genome of P. bedriagae
3.4. Genome Elimination in P. kl. esculentus
3.5. Genome Endoduplication in P. kl. esculentus
3.6. Genome Elimination in the F1 Generation
3.7. Hybridogenesis in Spermatogenic Cells
4. Discussion
4.1. Hybridogenesis in P. kl. esculentus
4.2. Basic L-E System in Western Russian Territoty
4.3. Introgression from Marsh and Levant Frogs
4.4. Variation in Genome Elimination
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Mayr, E. Systematics and the Origin of Species; Columbia University Press: New York, NY, USA, 1942. [Google Scholar]
- Burt, A.; Trivers, R. Genes in Conflict: The Biology of Selfish Genetic Elements; Harvard University Press: Cambridge, MA, USA, 2006. [Google Scholar]
- Schultz, R.J. Reproductive Mechanism of Unisexual and Bisexual Strains of the Viviparous Fish Poeciliopsis. Evolution 1961, 15, 302–325. [Google Scholar] [CrossRef]
- Schultz, R.J. Hybridization Experiments with an All-Female Fish of the Genus Poeciliopsis. Biol. Bull. 1966, 130, 415–429. [Google Scholar] [CrossRef]
- Schultz, R.J. Hybridization, unisexuality, and polyploidy in the teleost Poeciliopsis (Poeciliidae) and other vertebrates. Am. Nat. 1969, 103, 605–619. [Google Scholar] [CrossRef]
- Berger, L. Embryonal and larval development of F1 generation of green frogs different combinations. Acta Zool. Cracov. 1967, 12, 123–160. [Google Scholar]
- Berger, L. Morphology of the F1 generation of various crosses within Rana esculentus complex. Acta Zool. Cracov. 1968, 13, 301–324. [Google Scholar]
- Berger, L. Some characteristics of the crosses within Rana esculenta complex in post larval development. Ann. Zoologici Warszawa 1969, 2, 374–416. [Google Scholar]
- Graf, J.-D.; Polls-Pelaz, M. Evolutionary genetics of Rana esculenta complex. In Evolution and Ecology of Unisexual Vertebrates; Dawley, R.M., Bogart, J.P., Eds.; The New York State Museum Bulletin: Albany, NY, USA, 1989; pp. 298–302. [Google Scholar]
- Holsbeek, G.; Jooris, R. Potential impact of genome exclusion by alien species in the hybridogenetic water frogs (Pelophylax esculentus complex). Biol. Invasions 2010, 12, 1–13. [Google Scholar] [CrossRef]
- Tunner, H. Die klonale struktur einre Wassersroschpopulation. Z. Zool. Syst. Evol. Forsch. 1974, 12, 309–314. [Google Scholar] [CrossRef]
- Uzzell, T.; Berger, L. Electrophoretic phenotypes of Rana ridibunda, Rana lessonae, and their hybridogenetic associate, Rana esculenta. Proc. Acad. Nat. Sci. Phila. 1975, 127, 13–24. [Google Scholar]
- Graf, J.-D.; Müller, W.P. Experimental gynogenesis provides evidence of hybridogenetic reproduction in the Rana esculenta complex. Experientia 1979, 35, 1574–1576. [Google Scholar] [CrossRef] [PubMed]
- Uzzell, T.; Hotz, H.; Berger, L. Genome exclusion in gametogenesis by an interspecific Rana hybrid: Evidence from electrophoresis of individual oocytes. J. Exp. Zool. 1980, 214, 251–259. [Google Scholar] [CrossRef]
- Berger, L.; Günther, R. Inheritance patterns of water frog males from the environments of Nature Reserve Steckby, Germany. Zool. Pol. 1991, 37, 87–100. [Google Scholar]
- Krizmanić, I.I.; Ivanović, A.T. Population systems of the Pelophylax esculentus complex in the southern part of its range. J. Vertebr. Biol. 2010, 59, 215–222. [Google Scholar]
- Doležálková, M.; Sember, A.; Marec, F.; Ráb, P.; Plötner, J.; Choleva, L. Is premeiotic genome elimination an exclusive mechanism for hemiclonal reproduction in hybrid males of the genus Pelophylax? BMC Genet. 2016, 17, 100. [Google Scholar] [CrossRef] [PubMed]
- Dedukh, D.; Litvinchuk, J.; Svinin, A.; Litvinchuk, S.; Rosanov, J.; Krasikova, A. Variation in hybridogenetic hybrid emergence between populations of water frogs from the Pelophylax esculentus complex. PLoS ONE 2019. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, A.Y.; Korzikov, V.A.; Alekseev, S.K.; Ermakov, O.A. Molecular genetic characteristics of marsh frogs Pelophylax ridibundus s.l. from the upper Poochye. In Proceedings of the Modern Problems of Zoology, Ecology and Nature Protection: Materials of Readings and Scientific Conference Dedicated to the Memory of Prof. Andrey Grigorievich Bannikov, and the 100th anniversary of his birth (Sovremenniye problemy zoologii, ekologii I ohrany prirody; materialy chtenii I nauchnoi konferencii, posvyashchennoi pamyati professor Andreya Grigorievicha Bannikova I 100-letiyu so dnya ego rozhdeniya), Moscow, Russia, 24 April 2015; pp. 228–232. [Google Scholar]
- Svinin, A.O.; Ivanov, A.Y.; Zaks, M.M.; Litvinchuk, S.N.; Borkin, L.Y.; Rozanov, Y.M.; Ermakov, O.A. Distribution of “western” and “Eastern” forms of the marsh frog, Pelophylax ridibundus, and their participation in the formation of semi-clonal hybrids of P. esculentus in the Republic of Mari El. Mod. Herpetol. 2015, 15, 120–129. [Google Scholar]
- Zamaletdinov, R.I.; Pavlov, A.V.; Zaks, M.M.; Ivanov, A.Y.; Ermakov, O.A. Molecular genetic characteristics of frogs Pelophylax esculentus complex on the eastern periphery of the range (Volga region, Republic of Tatarstan). Bull. Tomsk State Univ. Biol. 2015, 3, 54–66. [Google Scholar] [CrossRef] [PubMed]
- Plötner, J.; Köhler, F.; Uzzell, T.; Beerl, P.; Schreibe, R.; Guex, G.-D.; Hotzae, H. Evolution of serum albumin intron-1 is shaped by a 5′ truncated non-long terminal repeat retrotransposon in western Palearctic water frogs (Neobatrachia). Mol. Phylogenet. Evol. 2009, 53, 784–791. [Google Scholar] [CrossRef] [PubMed]
- Berger, L.; Rybacki, M.; Hotz, H. Artificial fertilization of water frogs. Amphib. Reptil. 1994, 15, 408–413. [Google Scholar]
- Ohtani, H.; Miura, I.; Kondo, Y.; Uchibori, M. Amphidiploidy recovers the viability of hybrids between the European and Far Eastern water frogs. J. Exp. Zool. 1997, 279, 113–117. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Reyer, H.-U.; Niederer, B.; Hettyey, A. Variation in fertilisation abilities between hemiclonal hybrid and sexual parental males of sympatric water frogs (Rana lessonae, R. esculenta, R. ridibunda). Behav. Ecol. Sociobiol. 2003, 54, 274–284. [Google Scholar] [CrossRef][Green Version]
- Ogielska, M.; Bartmanska, J. Development of testes and differentiation of germ cells in water frogs of the Rana esculenta-complex (Amphibia, Anura). Amphib. Reptil. 1999, 20, 251–263. [Google Scholar] [CrossRef]
- Tunner, H.G.; Heppich-Tunner, S. A new population system of water frogs discovered in Hungary. In Proceedings of the Sixth Ordinary General Meeting of the Societas Europaea Herpetologica; Korsós, Z., Kiss, I., Eds.; pp. 453–460.
- Dubey, S.; Maddalena, T.; Bonny, L.; Jeffries, D.L.; Dufresnes, C. Population genomics of an exceptional hybridogenetic system of Pelophylax water frogs. BMC Evolutionary Bio. 2019, 19, 164. [Google Scholar] [CrossRef] [PubMed]
- Pruvost, N.B.M.; Hoffmann, A.; Reyer, H.-U. Gamete production patterns, ploidy, and population genetics reveal evolutionary significant units in hybrid water frogs (Pelophylax esculentus). Ecol. Evol. 2013, 3, 2933–2946. [Google Scholar] [CrossRef]
- Dedukh, D.; Litvinchuk, S.; Rosanov, J.; Shabanov, D.; Krasikova, A. Mutual maintenance of di- and triploid Pelophylax esculentus hybrids in R-E systems: Results from artificial crossings experiments. BMC Evol. Biol. 2017, 17. [Google Scholar] [CrossRef]
- Suriadna, N.M.; Mykytynets, G.I.; Pupiņš, M.; Gasso, V.Y. Population systems of Eurasian water frogs (Pelophylax) in the south of Ukraine. Biosyst. Divers. 2020, 28, 154–162. [Google Scholar] [CrossRef]
- Uzzell, T.; Günther, R.; Berger, L. Rana ridibunda and Rana esculenta: A leaky hybridogenetic system (Amphibia Salientia). Proc. Acad. Nat. Sci. Phila. 1976, 128, 147–171. [Google Scholar]
- Tunner, H.G.; Heppich, S. Premeiotic genome exclusion during oogenesis in the common edible frog, Rana esculenta. Die Naturwissenschaften 1981, 68, 207–208. [Google Scholar] [CrossRef] [PubMed]
- Heppich, S.; Tunner, H.G.; Greilhuber, J. Premeiotic Chromosome doubling after genome elimination during spermatogenesisof the species hybrid Rana esculenta. Theor. Appl. Genet. 1982, 61, 101–104. [Google Scholar] [CrossRef] [PubMed]
- Spolsky, C.; Uzzel, T. Evolutionary History of the Hybridogenetic Hybrid Frog Rana esculenta as Deduced from mtDNA Analyses. Mol. Biol. Evol. 1986, 3, 44–56. [Google Scholar]
- Chmielewska, M.; Dedukh, D.; Haczkiewicz, K.; Rozenblut-Kościsty, B.; Kaźmierczak, M.; Kolenda, K.; Serwa, E.; Pietras-Lebioda, A.; Krasikova, A.; Ogielska, M. The programmed DNA elimination and formation of micronuclei in germ line cells of the natural hybridogenetic water frog Pelophylax esculentus. Sci. Rep. 2018, 8, 7870. [Google Scholar] [CrossRef] [PubMed]
- Leuenberger, J.; Gander, A.; Benedikt, R.; Schmidt, B.R.; Perrin, N. Are invasive marsh frogs (Pelophylax ridibundus) replacing the native P. lessonae/P. esculentus hybridogenetic complex in Western Europe? Genetic evidence from a field study. Conserv. Genet. 2014, 15, 869–878. [Google Scholar] [CrossRef]
- Plötner, J. Die westpaläarktischen Wasserfrösche: Von Märtyrern der Wissenschaft zur biologischen Sensation; Laurenti-Verlag: Bielefeld, Germany, 2005; p. 160. [Google Scholar]
- Guerrini, F.; Bucci, S.; Ragghianti, M.; Mancino, G.; Hotz, H.; Uzzell, T.; Berger, L. Genomes of two water frog species resist germ line exclusion in interspecies hybrids. J. Exp. Zool. 1997, 279, 163–176. [Google Scholar] [CrossRef]
- Ragghianti, M.; Bucci, S.; Marracci, S.; Casola, C.; Mancino, G.; Hotz, H.; Guex, G.-D.; Plötner, J.; Uzzell, T. Gametogenesis of intergroup hybrids of hemiclonal frogs. Genet. Res. 2007, 89, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Gibeaux, R.; Acker, R.; Kitaoka, M.; Georgiou, G.; van Kruijsbergen, I.; Ford, B.; Marcotte, E.M.; Nomura, D.K.; Kwon, T.; Veenstra, G.J.C.; et al. Paternal chromosome loss and metabolic crisis contribute to hybrid inviability in Xenopus. Nature 2018, 553, 337–341. [Google Scholar] [CrossRef]
- Quilodrán, C.S.; Currat, M.; Montoya-Burgos, J.I. Effect of hybridization with genome exclusion on extinction risk. Conserv. Biol. 2018, 32, 1139–1149. [Google Scholar] [CrossRef] [PubMed]
- Ågren, J.A.; Clark, A.G. Selfish genetic elements. PLoS Genet. 2018, 14, e1007700. [Google Scholar] [CrossRef] [PubMed]
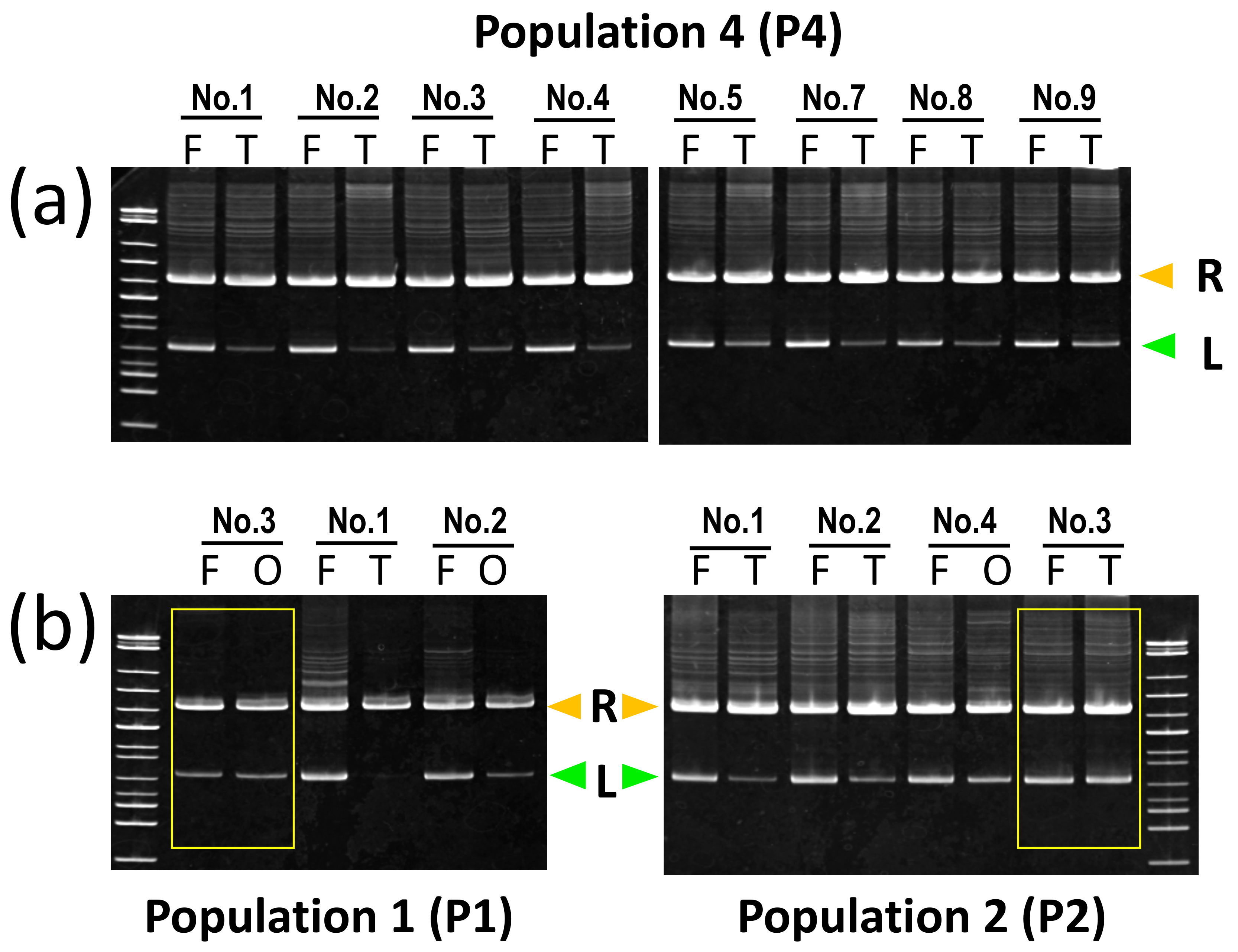
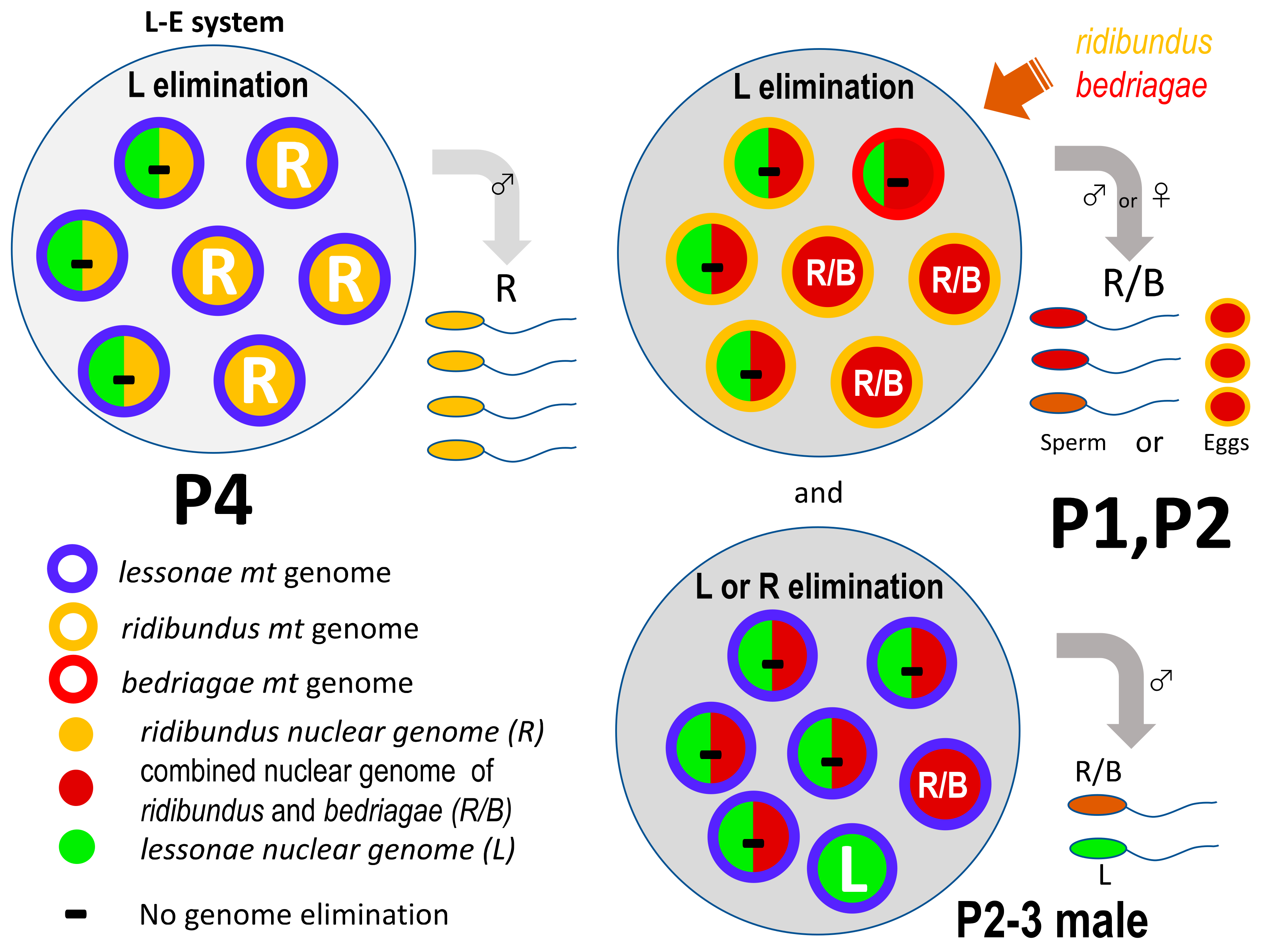
| Species | Population | Individual Number | Sex | Cyt b (1) | Serum A Intron 1 (SAI-1) (2) | Reduced SAI-1 Allele in Gonad (3) | SAI-1 Allele of Gamete | Fertility (4) |
|---|---|---|---|---|---|---|---|---|
| P. esculentus | P1 | No.1 | male | R | B | L | NE | |
| No.2 | female | R | B | L | NE | |||
| No.3 | female | B | BR | No change | NE | |||
| P2 | No.1 | male | R | R | L | R | 42.8 | |
| No.2 | male | R | R | L | NE | 0 | ||
| No.3 | male | L | B | No change | B or L | 0–2.3 | ||
| No.4 | female | R | B | L | B | 8.5 | ||
| P4 | No.1 | male | L | R | L | R | 66.7 | |
| No.2 | male | L | R | L | NE | |||
| No.3 | male | L | R | L | R | 37.5 | ||
| No.4 | male | L | R | L | R | 63 | ||
| No.5 | male | L | R | L | R | 11.3 | ||
| No.6 | male | L | NE | NE | NE | |||
| No.7 | male | L | R | L | R | 44.4 | ||
| No.8 | male | L | R | L | R | 36.4–50.9 | ||
| No.9 | male | L | R | L | R | 1.5–33.3 | ||
| P. ridibundus | P2 | No.1 | male | R | RR | |||
| No.2 | female | R | RB | |||||
| P3 | No.1 | male | R | BB | ||||
| No.2 | male | R | RR | 30.4–75.1 | ||||
| No.3 | male | B | RB | |||||
| No.4 | male | R | RR | |||||
| No.5 | female | R | RR | |||||
| No.6 | female | R | RB | 75.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Miura, I.; Vershinin, V.; Vershinina, S.; Lebedinskii, A.; Trofimov, A.; Sitnikov, I.; Ito, M. Hybridogenesis in the Water Frogs from Western Russian Territory: Intrapopulation Variation in Genome Elimination. Genes 2021, 12, 244. https://doi.org/10.3390/genes12020244
Miura I, Vershinin V, Vershinina S, Lebedinskii A, Trofimov A, Sitnikov I, Ito M. Hybridogenesis in the Water Frogs from Western Russian Territory: Intrapopulation Variation in Genome Elimination. Genes. 2021; 12(2):244. https://doi.org/10.3390/genes12020244
Chicago/Turabian StyleMiura, Ikuo, Vladimir Vershinin, Svetlana Vershinina, Andrei Lebedinskii, Alexander Trofimov, Ivan Sitnikov, and Michihiko Ito. 2021. "Hybridogenesis in the Water Frogs from Western Russian Territory: Intrapopulation Variation in Genome Elimination" Genes 12, no. 2: 244. https://doi.org/10.3390/genes12020244
APA StyleMiura, I., Vershinin, V., Vershinina, S., Lebedinskii, A., Trofimov, A., Sitnikov, I., & Ito, M. (2021). Hybridogenesis in the Water Frogs from Western Russian Territory: Intrapopulation Variation in Genome Elimination. Genes, 12(2), 244. https://doi.org/10.3390/genes12020244









