Exploration of Alternative Splicing (AS) Events in MDV-Infected Chicken Spleens
Abstract
1. Introduction
2. Materials and Methods
2.1. Biological Samples
2.2. Alternative Splicing Analysis
2.3. Function Enrichment Analysis
2.4. Analysis of SNP and Indel
2.5. Validation of AS Events
3. Results
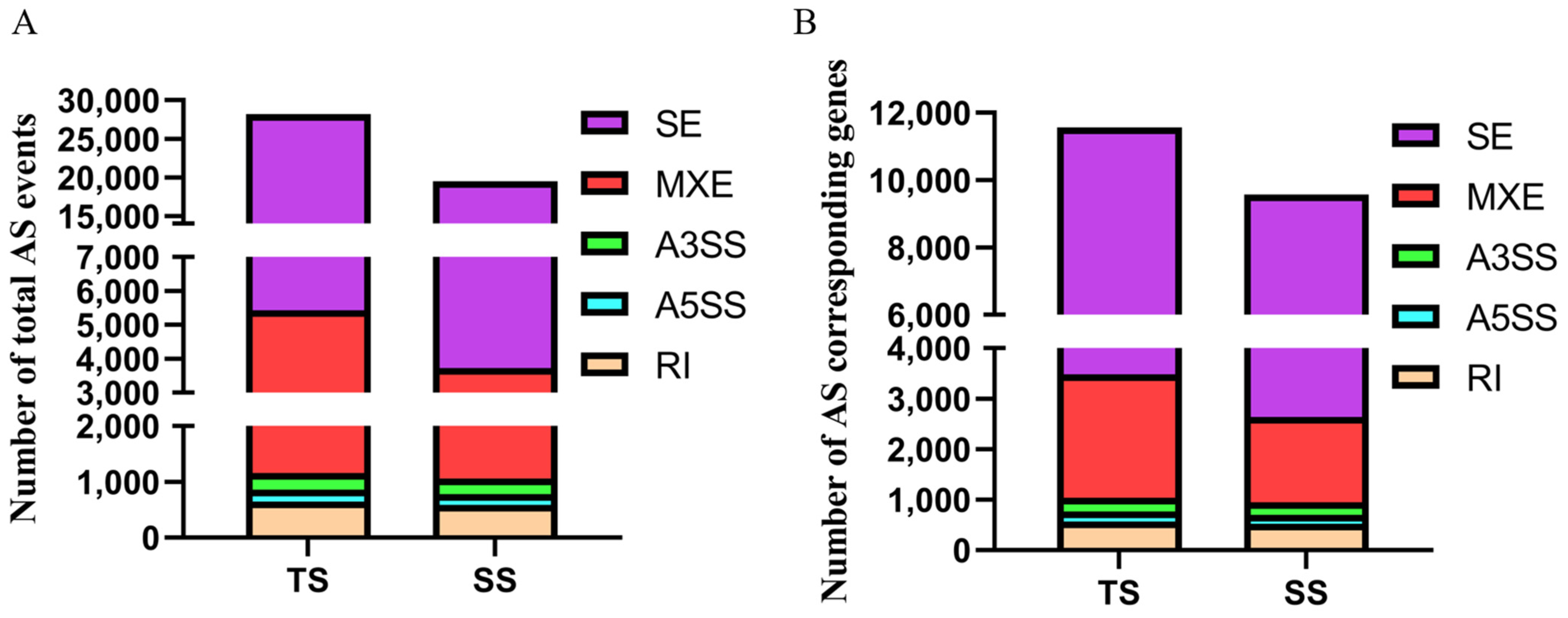
3.1. Identification of Alternative Splicing Events
3.2. Validation and Expression Analysis of AS Events
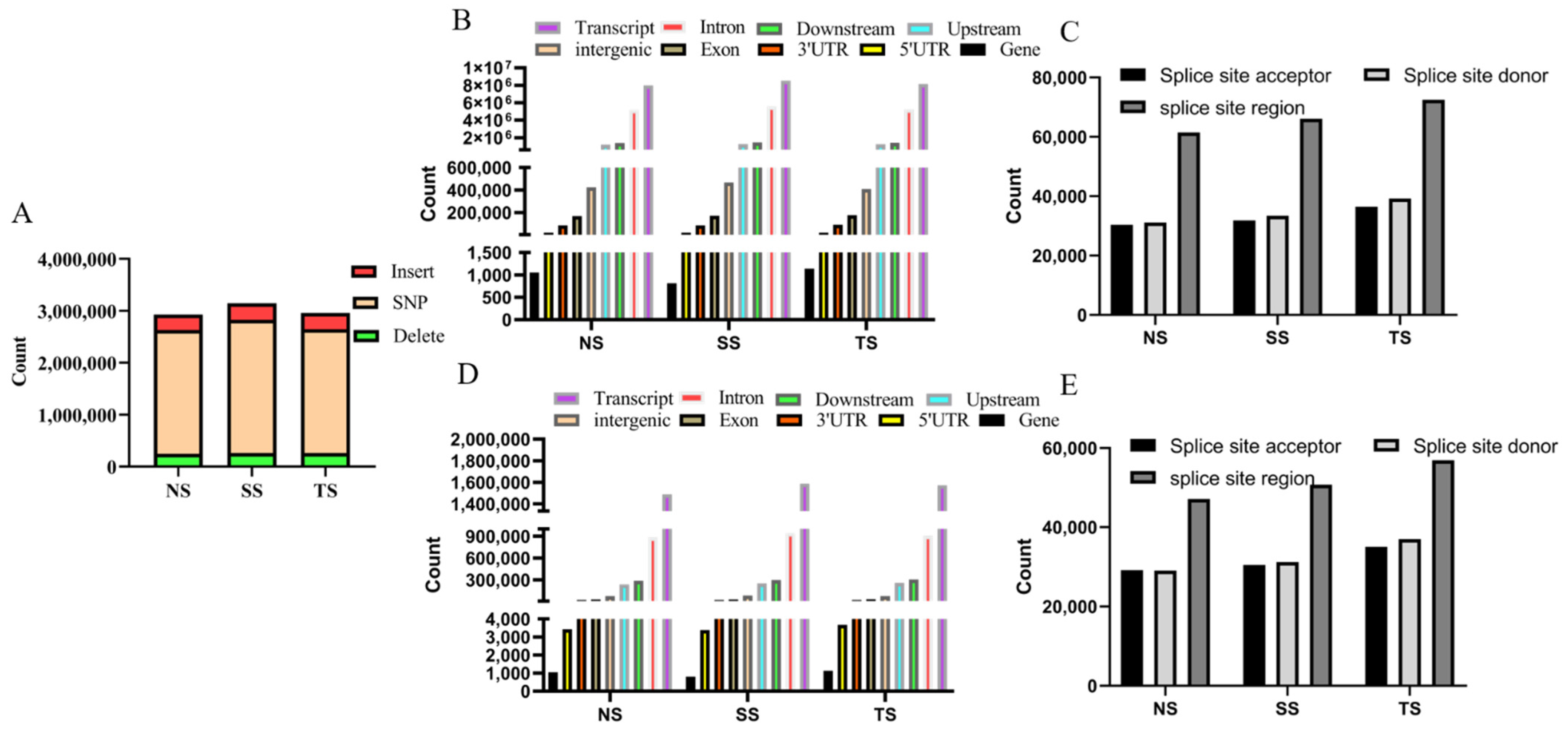
3.3. Analysis of SNP and Indel
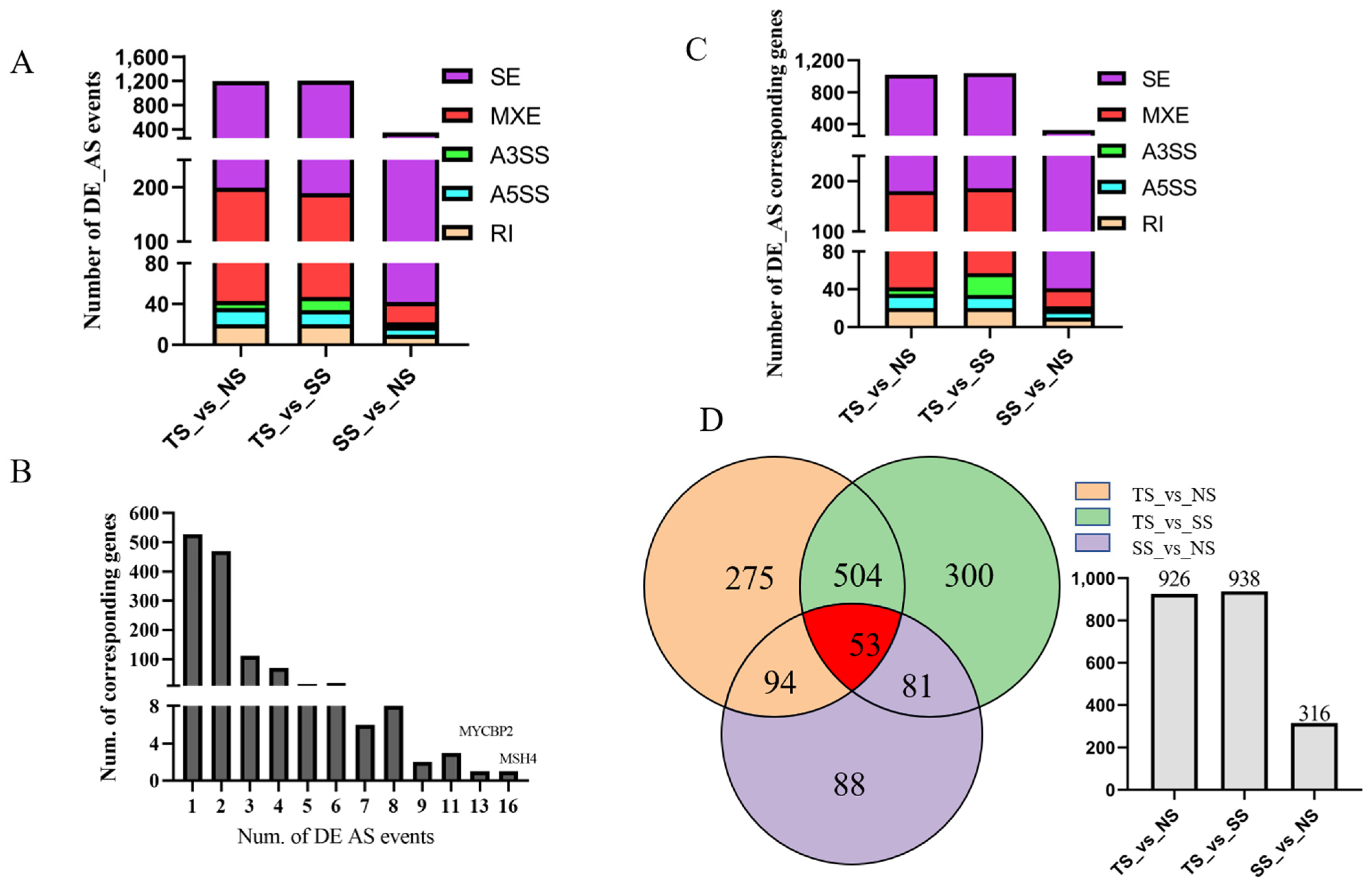
3.4. Identification of Differentially Alternative Splicing (DE AS) Event and Corresponded Genes
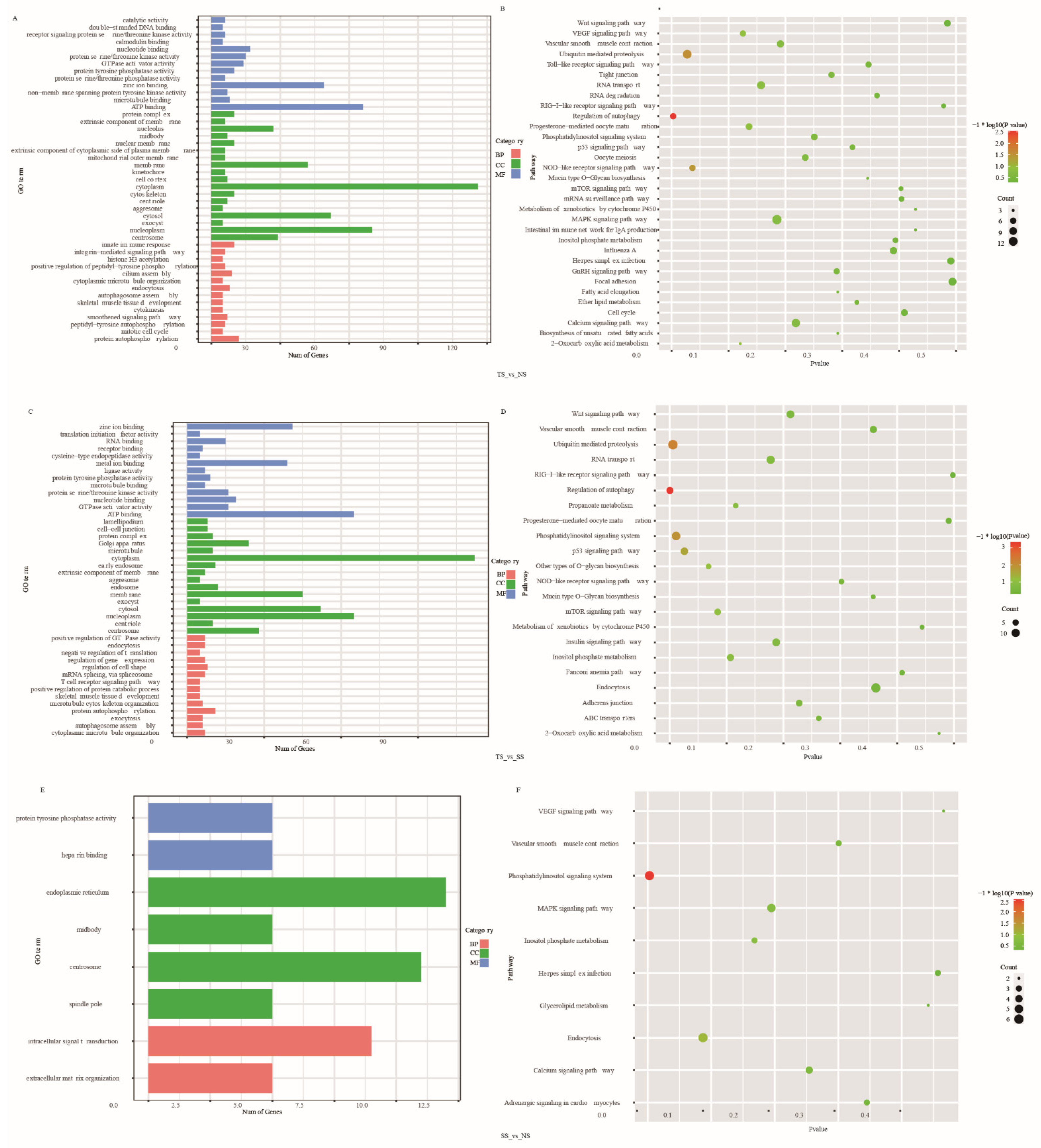
3.5. Function Enrichment Analysis of DE AS Corresponding Genes
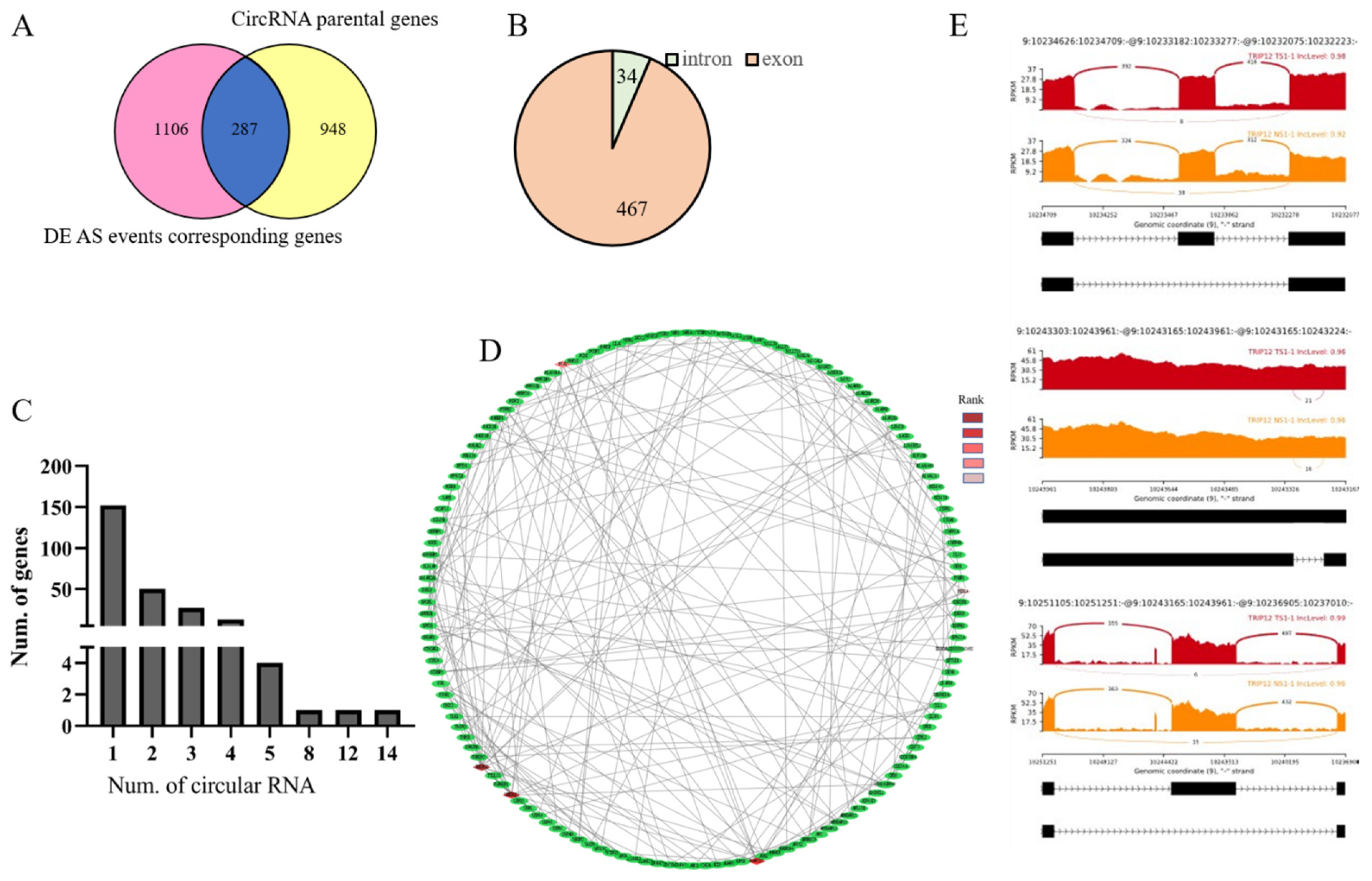
3.6. Identification of Genes between DE AS Corresponding Genes and CircRNA Parental Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gennart, I.; Coupeau, D.; Pejakovic, S.; Laurent, S.; Rasschaert, D.; Muylkens, B. Marek’s disease: Genetic regulation of gallid herpesvirus 2 infection and latency. Vet. J. 2015, 205, 339–348. [Google Scholar] [CrossRef] [PubMed]
- McPherson, M.; Delany, M. Virus and host genomic, molecular, and cellular interactions during Marek’s disease pathogenesis and oncogenesis. Poult. Sci. 2016, 95, 419–425. [Google Scholar] [CrossRef]
- Boodhoo, N.; Gurung, A.; Sharif, S.; Behboudi, S. Marek’s disease in chickens: A review with focus on immunology. Veter. Res. 2016, 47, 119. [Google Scholar] [CrossRef] [PubMed]
- Nair, V. Evolution of Marek’s disease—A paradigm for incessant race between the pathogen and the host. Vet. J. 2005, 170, 175–183. [Google Scholar] [CrossRef] [PubMed]
- Arbuckle, J.H.; Medveczky, M.M.; Luka, J.; Hadley, S.H.; Luegmayr, A.; Ablashi, D.; Lund, T.C.; Tolar, J.; De Meirleir, K.; Montoya, J.G.; et al. The latent human herpesvirus-6A genome specifically integrates in telomeres of human chromosomes in vivo and in vitro. Proc. Natl. Acad. Sci. USA 2010, 107, 5563–5568. [Google Scholar] [CrossRef] [PubMed]
- Delecluse, H.J.; Bartnizke, S.; Hammerschmidt, W.; Bullerdiek, J.; Bornkamm, G.W. Episomal and Integrated Copies of Epstein-Barr Virus__Coexist in Burkitt Lymphoma Cell Lines. J. Virol. 1993, 67, 1292–1299. [Google Scholar] [CrossRef]
- Delecluse, H.J.; Hammerschmidt, W. Status of Mareks-Disease Virus In Established Lymphoma Cell-Lines—Herpesvirus Integration Is Common. J. Virol. 1993, 67, 82–92. [Google Scholar] [CrossRef]
- Delecluse, H.J.; Schuller, S.; Hammerschmidt, W. Latent Mareks-Disease Virus Can Be Activated From Its Chromosomally Integrated State In Herpesvirus-Transformed Lymphoma-Cells. EMBO J. 1993, 12, 3277–3286. [Google Scholar] [CrossRef]
- Osterrieder, N.; Kamil, J.P.; Schumacher, D.; Tischer, B.K.; Trapp, S. Marek’s disease virus: From miasma to model. Nat. Rev. Microbiol. 2006, 4, 283–294. [Google Scholar] [CrossRef] [PubMed]
- Daibata, M.; Taguchi, T.; Taguchi, H.; Miyoshi, I. Integration of human herpesvirus 6 in a Burkitt’s lymphoma cell line. Br. J. Haematol. 1998, 102, 1307–1313. [Google Scholar] [CrossRef] [PubMed]
- Kaschka-Dierich, C.; Adams, A.; Lindahl, T.; Bornkamm, G.W.; Bjursell, G.; Klein, G.; Giovanella, B.C.; Singh, S. Intracellular forms of Epstein–Barr virus DNA in human tumour cells in vivo. Nat. Cell Biol. 1976, 260, 302–306. [Google Scholar] [CrossRef] [PubMed]
- Benetatos, L.; Hatzimichael, E.; Londin, E.; Vartholomatos, G.; Loher, P.; Rigoutsos, I.; Briasoulis, E. The microRNAs within the DLK1-DIO3 genomic region: Involvement in disease pathogenesis. Cell. Mol. Life Sci. 2012, 70, 795–814. [Google Scholar] [CrossRef]
- Mao, S.; Zhang, S.; Zhou, Z.; Shi, X.; Huang, T.; Feng, W.; Yao, C.; Gu, X.; Yu, B. Alternative RNA splicing associated with axon regeneration after rat peripheral nerve injury. Exp. Neurol. 2018, 308, 80–89. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.-S.; Zhu, T.; Li, L.; Xie, P.; Li, X.; Zhou, H.-H.; Liu, Z.-Q. LncRNAs and CircRNAs from the same gene: Masterpieces of RNA splicing. Cancer Lett. 2018, 415, 49–57. [Google Scholar] [CrossRef]
- Stamm, S.; Ben-Ari, S.; Rafalska, I.; Tang, Y.; Zhang, Z.; Toiber, D.; Thanaraj, T.A.; Soreq, H. Function of alternative splicing. Gene 2005, 344, 1–20. [Google Scholar] [CrossRef]
- Wang, R.; Zhang, S.; Chen, X.; Li, N.; Li, J.; Jia, R.; Pan, Y.; Liang, H. CircNT5E Acts as a Sponge of miR-422a to Promote Glioblastoma Tumorigenesis. Cancer Res. 2018, 78, 4812–4825. [Google Scholar] [CrossRef]
- Hansen, T.B.; Wiklund, E.D.; Bramsen, J.B.; Villadsen, S.B.; Statham, A.L.; Clark, S.J.; Kjems, J. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 2011, 30, 4414–4422. [Google Scholar] [CrossRef]
- Xu, S.; Zhou, L.; Ponnusamy, M.; Zhang, L.; Dong, Y.; Zhang, Y.; Wang, Q.; Liu, J.; Wang, K. A comprehensive review of circRNA: From purification and identification to disease marker potential. Peer J. 2018, 6, e5503. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S.; Lu, C.; Luber, B.; Wang, H.; Chen, Y.; Zhu, Y.; Silberstein, J.L.; Taylor, M.N.; Maughan, B.L.; Denmeade, S.R.; et al. Clinical Significance of Androgen Receptor Splice Variant-7 mRNA Detection in Circulating Tumor Cells of Men With Metastatic Castration-Resistant Prostate Cancer Treated With First- and Second-Line Abiraterone and Enzalutamide. J. Clin. Oncol. 2017, 35, 2149–2156. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Ji, M.; Lv, Y.; Feng, Q.; Zheng, P.; Mao, Y.; Xu, Y.; He, G.; Xu, J. A signature predicting relapse based on integrated analysis on relapse-associated alternative mRNA splicing in I–III rectal cancer. Genomics 2020, 112, 3274–3283. [Google Scholar] [CrossRef] [PubMed]
- Nandagopalan, S.R.; Agatheeswaran, S.; Vadlamudi, Y.; Biswas, S.; Biswas, G.; Pattnayak, N.C.; Chakraborty, S. PTBP2 exon 10 inclusion is associated with the progression of CML and it is BCR-ABL1 dependent. Int. J. Biochem. Cell Biol. 2019, 109, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Tominaga, K.; Pereira-Smith, O.M. The genomic organization, promoter position and expression profile of the mouse MRG15 gene. Gene 2002, 294, 215–224. [Google Scholar] [CrossRef]
- Tajul-Arifin, K.; Teasdale, R.; Ravasi, T.; Hume, D.; RIKEN GER Group; Members, G.; Mattick, J. Identification and Analysis of Chromodomain-Containing Proteins Encoded in the Mouse Transcriptome. Genome Res. 2003, 13, 1416–1429. [Google Scholar] [CrossRef] [PubMed]
- Lois, S.; Blanco, N.; Martínez-Balbás, M.; de la Cruz, X. The functional modulation of epigenetic regulators by alternative splicing. BMC Genom. 2007, 8, 252. [Google Scholar] [CrossRef] [PubMed]
- Peterson, C.L.; Laniel, M.A. Histones and histone modifications. Curr. Biol. 2004, 14, 546–551. [Google Scholar] [CrossRef]
- Brown, A.C.; Nair, V.; Allday, M.J. Epigenetic regulation of the latency-associated region of Marek’s disease virus in tumor-derived T-cell lines and primary lymphoma. J. Virol. 2012, 86, 1683–1695. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.C.; Huang, L.; Wei, X.X.; Zhao, Z.Y.; Li, Y.; Wei, P. The relationship between the alternative exon 7 splice variant of the BF gene and MHC-related Marek’s disease resistance in chickens. Scand. J. Immunol. 2014, 80, 323–326. [Google Scholar] [CrossRef] [PubMed]
- Jarosinski, K.W.; Schat, K.A. Multiple alternative splicing to exons II and III of viral interleukin-8 (vIL-8) in the Marek’s disease virus genome: The importance of vIL-8 exon I. Virus Genes 2007, 34, 9–22. [Google Scholar] [CrossRef] [PubMed]
- Kaya, M.; Preeyanon, L.; Dodgson, J.B.; Cheng, H.H. Validation of Alternative Transcript Splicing in Chicken Lines that Differ in Genetic Resistance to Marek’s Disease. Anim. Biotechnol. 2016, 27, 238–244. [Google Scholar] [CrossRef] [PubMed]
- Galeano, C.H.; Fernández, A.C.; Gómez, M.; Blair, M.W. Single strand conformation polymorphism based SNP and Indel markers for genetic mapping and synteny analysis of common bean (Phaseolus vulgaris L.). BMC Genom. 2009, 10, 629. [Google Scholar] [CrossRef]
- Schmidt, C.; Wolff, M.; von Ahsen, N.; Lange, K.; Friede, T.; Zerr, I. CR1 is potentially associated with rate of decline in sporadic Alzheimer’s disease. J. Clin. Neurosci. 2014, 21, 1705–1708. [Google Scholar] [CrossRef] [PubMed]
- Stone, L. Prostate cancer: In a SNP: KLK6 mutations in aggressive disease. Nat. Rev. Urol. 2017, 14, 326–327. [Google Scholar]
- Vernerová, K.; Tothova, L.; Mikova, A.; Vodrazka, P.; Simek, B.; Hanusová, L.; Citek, J. BSE-associated polymorphisms in the prion protein gene: An investigation. J. Anim. Breed. Genet. 2014, 131, 403–408. [Google Scholar] [CrossRef] [PubMed]
- Peletto, S.; Bertolini, S.; Maniaci, M.G.; Colussi, S.; Modesto, P.; Biolatti, C.; Bertuzzi, S.; Caramelli, M.; Maurella, C.; Acutis, P.L. Association of an indel polymorphism in the 3′UTR of the caprine SPRN gene with scrapie positivity in the central nervous system. J. Gen. Virol. 2012, 93, 1620–1623. [Google Scholar] [CrossRef] [PubMed]
- Li, D.F.; Lian, L.; Qu, L.J.; Chen, Y.M.; Liu, W.B.; Chen, S.R.; Zheng, J.X.; Xu, G.Y.; Yang, N. A genome-wide SNP scan reveals two loci associated with the chicken resistance to Marek’s disease. Anim. Genet. 2013, 44, 217–222. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Kung, H.; Fulton, J.; Morgan, R.; Cheng, H. Growth hormone interacts with the Marek’s disease virus SORF2 protein and is associated with disease resistance in chicken. Proc. Nat. Acad. Sci. USA 2001, 98, 9203–9208. [Google Scholar] [CrossRef] [PubMed]
- Kuhnlein, U.; Ni, L.; Weigend, S.; Gavora, J.; Fairfull, W.; Zadworny, D. DNA polymorphisms in the chicken growth hormone gene: Response to selection for disease resistance and association with egg production. Anim. Genet. 1997, 28, 116–123. [Google Scholar] [CrossRef]
- Ju, Z.; Wang, C.; Wang, X.; Yang, C.; Sun, Y.; Jiang, Q.; Wang, F.; Li, M.; Zhong, J.; Huang, J. Role of an SNP in Alternative Splicing of Bovine NCF4 and Mastitis Susceptibility. PLoS ONE 2015, 10, e0143705. [Google Scholar] [CrossRef] [PubMed]
- Lian, L.; Qu, L.J.; Sun, H.Y.; Chen, Y.M.; Lamont, S.J.; Liu, C.J.; Yang, N. Gene expression analysis of host spleen responses to Marek’s disease virus infection at late tumor transformation phase. Poult. Sci. 2012, 91, 2130–2138. [Google Scholar] [CrossRef]
- Shen, S.; Park, J.W.; Lu, Z.-X.; Lin, L.; Henry, M.; Wu, Y.N.; Zhou, Q.; Xing, Y. rMATS: Robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc. Natl. Acad. Sci. USA 2014, 111, E5593–E5601. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [PubMed]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef] [PubMed]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Jarosinski, K.W.; Tischer, B.K.; Trapp, S.; Osterrieder, N. Marek’s disease virus: Lytic replication, oncogenesis and control. Expert Rev. Vaccines 2006, 5, 761–772. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.C.; Smith, L.P.; Kgosana, L.; Baigent, S.J.; Nair, V.; Allday, M.J. Homodimerization of the Meq viral oncoprotein is necessary for induction of T-cell lymphoma by Marek’s disease virus. J. Virol. 2009, 83, 11142–11151. [Google Scholar] [CrossRef]
- Artandi, S.E. Telomerase flies the coop: The telomerase RNA component as a viral-encoded oncogene. J. Exp. Med. 2006, 203, 1143–1145. [Google Scholar] [CrossRef][Green Version]
- Engel, A.T.; Selvaraj, R.K.; Kamil, J.P.; Osterrieder, N.; Kaufer, B.B. Marek’s disease viral interleukin-8 promotes lymphoma formation through targeted recruitment of B cells and CD4+ CD25+ T cells. J. Virol. 2012, 86, 8536–8545. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.B.; Sondermeijer, P.J.A.; Velicer, L.F. Identification of a unique mareks-disease virus gene which encodes a 38-kilodalton phosphoprotein and is expressed in both lytically infected-cells and latently infected lymphoblastoid tumor-cells. J. Virol. 1992, 66, 85–94. [Google Scholar] [CrossRef] [PubMed]
- Prigge, J.T.; Majerciak, V.; Hunt, H.D.; Dienglewicz, R.L.; Parcells, M.S. Construction and characterization of Marek’s disease viruses having green fluorescent protein expression tied directly or indirectly to phosphoprotein 38 expression. Avian. Dis. 2004, 48, 471–487. [Google Scholar] [CrossRef]
- Zhao, Y.; Yao, Y.; Xu, H.; Lambeth, L.; Smith, L.P.; Kgosana, L.; Wang, X.; Nair, V. A functional MicroRNA-155 ortholog encoded by the oncogenic Marek’s disease virus. J. Virol. 2009, 83, 489–492. [Google Scholar] [CrossRef] [PubMed]
- Jarosinski, K.W.; Hunt, H.D.; Osterrieder, N. Down-regulation of MHC class I by the Marek’s disease virus (MDV) UL49.5 gene product mildly affects virulence in a haplotype-specific fashion. Virology 2010, 405, 457–463. [Google Scholar] [CrossRef]
- Ding, K.; Yu, Z.H.; Yu, C.; Jia, Y.Y.; He, L.; Liao, C.S.; Li, J.; Zhang, C.J.; Li, Y.J.; Wu, T.C.; et al. Effect of gga-miR-155 on cell proliferation, apoptosis and invasion of Marek’s disease virus (MDV) transformed cell line MSB1 by targeting RORA. BMC Vet. Res. 2020, 16, 23. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Lian, L.; Zhang, D.X.; Qu, L.J.; Yang, N. Gga-miR-26a targets NEK6 and suppresses Marek’s disease lymphoma cell proliferation. Poul. Sci. 2014, 93, 1097–1105. [Google Scholar] [CrossRef]
- Yan, Y.; Yang, N.; Cheng, H.H.; Song, J.; Qu, L. Genome-wide identification of copy number variations between two chicken lines that differ in genetic resistance to Marek’s disease. BMC Genom. 2015, 16, 843. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; He, Y.; Ding, Y.; Sun, G.; Carrillo, J.A.; Li, Y.; Ghaly, M.M.; Ma, L.; Zhang, H.; Liu, G.E.; et al. Characterization of Copy Number Variation’s Potential Role in Marek’s Disease. Int. J. Mol. Sci. 2017, 18, 1020. [Google Scholar] [CrossRef]
- Luo, J.; Yu, Y.; Chang, S.; Tian, F.; Zhang, H.; Song, J. DNA Methylation Fluctuation Induced by Virus Infection Differs between MD-resistant and -susceptible Chickens. Front. Genet. 2012, 3, 20. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; De Vecchi, G.; Caleca, L.; Foglia, C.; Ripamonti, C.B.; Ficarazzi, F.; Barile, M.; Varesco, L.; Peissel, B.; Manoukian, S.; et al. Comparative In Vitro and In Silico Analyses of Variants in Splicing Regions of BRCA1 and BRCA2 Genes and Characterization of Novel Pathogenic Mutations. PLoS ONE 2013, 8, e57173. [Google Scholar] [CrossRef]
- Pan, S.-S.; Han, Y.; Farabaugh, P.; Xia, H. Implication of alternative splicing for expression of a variant NAD(P)H:quinone oxidoreductase-1 with a single nucleotide polymorphism at 465C>T. Pharmacogenetics 2002, 12, 479–488. [Google Scholar] [CrossRef]
- Santucci-Darmanin, S.; Walpita, D.; Lespinasse, F.; Desnuelle, C.; Ashley, T.; Paquis-Flucklinger, V. MSH4 acts in conjunction with MLH1 during mammalian meiosis. FASEB J. 2000, 14, 1539–1547. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Tian, D.; Wang, S.; Ruan, Y.; Qiu, B.; Zhang, L.; Lu, B. Expressions of genes related to genome stability and DNA repair in nasopharyngeal carcinoma clustering families. Chin-German J. Clin. Oncol. 2009, 8, 713–718. [Google Scholar] [CrossRef]
- Tang, D.D.; Xu, C.; Geng, H.; Gao, Y.; Cheng, H.R.; Ni, X.Q.; He, X.J.; Gao, Y.X. A novel homozygous mutation in the meiotic gene MSH4 leading to male infertility due to non-obstructive azoospermia. Am. J. Transl. Res. 2020, 12, 8185–8191. [Google Scholar] [PubMed]
- Her, C.; Wu, X.; Griswold, M.D.; Zhou, F. Human MutS homologue MSH4 physically interacts with von Hippel-Lindau tumor suppressor-binding protein 1. Cancer Res. 2003, 63, 865–872. [Google Scholar] [PubMed]
- Liang, J.; Zhou, W.; Sakre, N.; DeVecchio, J.; Ferrandon, S.; Ting, A.H.; Bao, S.; Bissett, I.; Church, J.; Kalady, M.F. Epigenetically regulated miR-1247 functions as a novel tumour suppressor via MYCBP2 in methylator colon cancers. Br. J. Cancer 2018, 119, 1267–1277. [Google Scholar] [CrossRef] [PubMed]
- Cole, M.D.; McMahon, S.B. The Myc oncoprotein: A critical evaluation of transactivation and target gene regulation. Oncogene 1999, 18, 2916–2924. [Google Scholar] [CrossRef]
- Ge, Z.; Guo, X.; Li, J.; Hartman, M.; Kawasawa, Y.I.; Dovat, S.; Song, C. Clinical significance of high c-MYC and low MYCBP2 expression and their association with Ikaros dysfunction in adult acute lymphoblastic leukemia. Oncotarget 2015, 6, 42300–42311. [Google Scholar] [CrossRef] [PubMed]
- Calvayrac, O.; Pradines, A.; Favre, G. RHOB expression controls the activity of serine/threonine protein phosphatase PP2A to modulate mesenchymal phenotype and invasion in non-small cell lung cancers. Small GTPases 2016, 9, 339–344. [Google Scholar] [CrossRef] [PubMed]
- Sahin, F.; Celik, H.A.; Aydin, H.; Oktem, G.; Omay, S.B.; Saydam, G. The interaction between taxoids and serine/threonine protein phosphatase activities during taxan-induced apoptosis of HL 60 leukemic cells. Hematology 2008, 13, 215–223. [Google Scholar] [CrossRef]
- Bayer, A.L.; Heidkamp, M.C.; Patel, N.; Porter, M.; Engman, S.; Samarel, A.M. Alterations in protein kinase C isoenzyme expression and autophosphorylation during the progression of pressure overload-induced left ventricular hypertrophy. Mol. Cell. Biochem. 2003, 242, 145–152. [Google Scholar] [CrossRef]
- Ralph, A.B.; Edward, A.D. Handbook of Cell Signaling, 2nd ed.; Academic Press: Cambridge, MA, USA, 2009. [Google Scholar]
- Leroy, E.; Boyer, R.; Auburger, G.; Leube, B.; Ulm, G.; Mezey, E.; Harta, G.; Brownstein, M.J.; Jonnalagada, S.; Chernova, T.; et al. The ubiquitin pathway in Parkinson’s disease. Nature 1998, 395, 451–452. [Google Scholar] [CrossRef] [PubMed]
- Guo, G.; Gui, Y.; Gao, S.; Tang, A.; Hu, X.; Huang, Y.; Jia, W.; Li, Z.; He, M.; Sun, L.; et al. Frequent mutations of genes encoding ubiquitin-mediated proteolysis pathway components in clear cell renal cell carcinoma. Nat. Genet. 2011, 44, 17–19. [Google Scholar] [CrossRef] [PubMed]
- Broz, P.; Dixit, V.M. Inflammasomes: Mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 2016, 16, 407–420. [Google Scholar] [CrossRef]
- Wan, X.; Jin, X.; Xie, M.; Liu, J.; Gontcharov, A.A.; Wang, H.; Lv, R.; Liu, D.; Wang, Q.; Li, Y. Characterization of a polysaccharide from Sanghuangporus vaninii and its antitumor regulation via activation of the p53 signaling pathway in breast cancer MCF-7 cells. Int. J. Biol. Macromol. 2020, 163, 865–877. [Google Scholar] [CrossRef]
- Guo, L.; Kang, J.S.; Kang, N.J.; Choi, Y.W. S-petasin induces apoptosis and inhibits cell migration through activation of p53 pathway signaling in melanoma B16F10 cells and A375 cells. Arch. Biochem. Biophys. 2020, 692, 108519. [Google Scholar] [CrossRef]
- Li, H.; Zhu, J.; He, M.; Luo, Q.; Liu, F.; Chen, R. Marek’s Disease Virus Activates the PI3K/Akt Pathway Through Interaction of Its Protein Meq With the P85 Subunit of PI3K to Promote Viral Replication. Front. Microbiol. 2018, 9, 2547. [Google Scholar] [CrossRef] [PubMed]
- Roberti, M.C.; La Starza, R.; Surace, C.; Sirleto, P.; Pinto, R.M.; Pierini, V.; Crescenzi, B.; Mecucci, C.; Angioni, A. RABGAP1L gene rearrangement resulting from a der(Y)t(Y;1)(q12;q25) in acute myeloid leukemia arising in a child with Klinefelter syndrome. Virchows Arch. 2009, 454, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Kowarz, E.; Burmeister, T.; Nigro, L.L.; Jansen, M.W.J.C.; Delabesse, E.; Klingebiel, T.; Dingermann, T.; Meyer, C.; Marschalek, R.; Dingermann, T. Complex MLL rearrangements in t(4;11) leukemia patients with absent AF4 · MLL fusion allele. Leukemia 2007, 21, 1232–1238. [Google Scholar] [CrossRef] [PubMed]
- Anderson, D.; Neri, J.I.C.F.; Souza, C.R.M.; Valverde, J.G.; De Araújo, J.M.G.; Nascimento, M.D.S.B.; Branco, R.C.C.; Arrais, N.M.R.; Lassmann, T.; Blackwell, J.M.; et al. Zika Virus Changes Methylation of Genes Involved in Immune Response and Neural Development in Brazilian Babies Born With Congenital Microcephaly. J. Infect. Dis. 2021, 223, 435–440. [Google Scholar] [CrossRef] [PubMed]
- Marat, A.L.; McPherson, P.S. The Connecdenn Family, Rab35 Guanine Nucleotide Exchange Factors Interfacing with the Clathrin Machinery. J. Biol. Chem. 2010, 285, 10627–10637. [Google Scholar] [CrossRef]
- Chen, Z.-J.; Zhao, H.; He, L.; Shi, Y.; Qin, Y.; Shi, Y.; Li, Z.; You, L.; Zhao, J.; Liu, J.; et al. Genome-wide association study identifies susceptibility loci for polycystic ovary syndrome on chromosome 2p16.3, 2p21 and 9q33.3. Nat. Genet. 2010, 43, 55–59. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.-N.; Zhang, Y.-T.; Liu, Q.; Shen, S.-M.; Zou, X.; Cao, Y.-X.; Wang, W.-J.; Yi, L.; Gao, Q.; Yang, W.-D.; et al. Association analysis between the tag single nucleotide polymorphisms of DENND1A and the risk of polycystic ovary syndrome in Chinese Han women. BMC Med. Genet. 2020, 21, 10–14. [Google Scholar] [CrossRef]
- McAllister, J.M.; Modi, B.; Miller, B.A.; Biegler, J.; Bruggeman, R.; Legro, R.; Strauss, J.F. Overexpression of a DENND1A isoform produces a polycystic ovary syndrome theca phenotype. Proc. Nat. Acad. Sci. USA 2014, 111, E1519–E1527. [Google Scholar] [CrossRef]
- Wang, Z.; Li, T.; Zhang, W.; You, L.; Zhao, Y.; Xia, M.; Zhao, H.; Chen, Z.-J. Variants in DENND1A and LHCGR are associated with endometrioid adenocarcinoma. Gynecol. Oncol. 2012, 127, 403–405. [Google Scholar] [CrossRef]
- Larrieu, D.; Brunet, M.; Vargas, C.; Hanoun, N.; Ligat, L.; Dagnon, L.; Lulka, H.; Pommier, R.; Selves, J.; Jády, B.E.; et al. The E3 ubiquitin ligase TRIP12 participates in cell cycle progression and chromosome stability. Sci. Rep. 2020, 10, 789. [Google Scholar] [CrossRef]
- Liu, X.; Yang, X.; Li, Y.; Zhao, S.; Li, C.; Ma, P.; Mao, B. Trip12 is an E3 ubiquitin ligase for USP7/HAUSP involved in the DNA damage response. FEBS Lett. 2016, 590, 4213–4222. [Google Scholar] [CrossRef] [PubMed]
- Hagiwara, N.; Yeh, M.; Liu, A. Sox6 is required for normal fiber type differentiation of fetal skeletal muscle in mice. Dev. Dyn. 2007, 236, 2062–2076. [Google Scholar] [CrossRef] [PubMed]
- Gao, P.; Jin, Z.; Cheng, Y.; Cao, X. RNA-Seq analysis identifies aberrant RNA splicing of TRIP12 in acute myeloid leukemia patients at remission. Tumor. Biol. 2014, 35, 9585–9590. [Google Scholar] [CrossRef] [PubMed]
- Jacobsen, A.; Bosch, L.J.W.; Kemp, S.M.-D.; Carvalho, B.; Sillars-Hardebol, A.H.; Dobson, R.J.; De Rinaldis, E.; Meijer, G.A.; Abeln, S.; Heringa, J.; et al. Aurora kinase A (AURKA) interaction with Wnt and Ras-MAPK signalling pathways in colorectal cancer. Sci. Rep. 2018, 8, 7522. [Google Scholar] [CrossRef] [PubMed]
- Anaya, J. OncoLnc: Linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. PeerJ Comput. Sci. 2016, 2, e67. [Google Scholar] [CrossRef]
- Hou, L.; Li, Y.; Wang, Y.; Xu, D.; Cui, H.; Xu, X.; Cong, Y.; Yu, C. UBE2D1 RNA Expression Was an Independent Unfavorable Prognostic Indicator in Lung Adenocarcinoma, but Not in Lung Squamous Cell Carcinoma. Dis. Markers 2018, 2018, 4108919. [Google Scholar] [CrossRef] [PubMed]
- Tokumoto, M.; Fujiwara, Y.; Shimada, A.; Hasegawa, T.; Seko, Y.; Nagase, H.; Satoh, M. Cadmium toxicity is caused by accumulation of p53 through the down-regulation of Ube2d family genes in vitro and in vivo. J. Toxicol. Sci. 2011, 36, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Saville, M.K.; Sparks, A.; Xirodimas, D.P.; Wardrop, J.; Stevenson, L.F.; Bourdon, J.-C.; Woods, Y.L.; Lane, D.P. Regulation of p53 by the Ubiquitin-conjugating Enzymes UbcH5B/C in Vivo. J. Biol. Chem. 2004, 279, 42169–42181. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Bi, F.; Yuan, J.; Yang, F.; Sun, S. Gain of UBE2D1 facilitates hepatocellular carcinoma progression and is associated with DNA damage caused by continuous IL-6. J. Exp. Clin. Cancer Res. 2018, 37, 290. [Google Scholar] [CrossRef] [PubMed]
- Alsina, D.; Lytovchenko, O.; Schab, A.; Atanassov, I.; Schober, F.; Jiang, M.; Koolmeister, C.; Wedell, A.; Taylor, R.W.; Wredenberg, A.; et al. FBXL 4 deficiency increases mitochondrial removal by autophagy. EMBO Mol. Med. 2020, 12, 11659. [Google Scholar] [CrossRef]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol. Cell 2017, 66, 22–37.e9. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef] [PubMed]
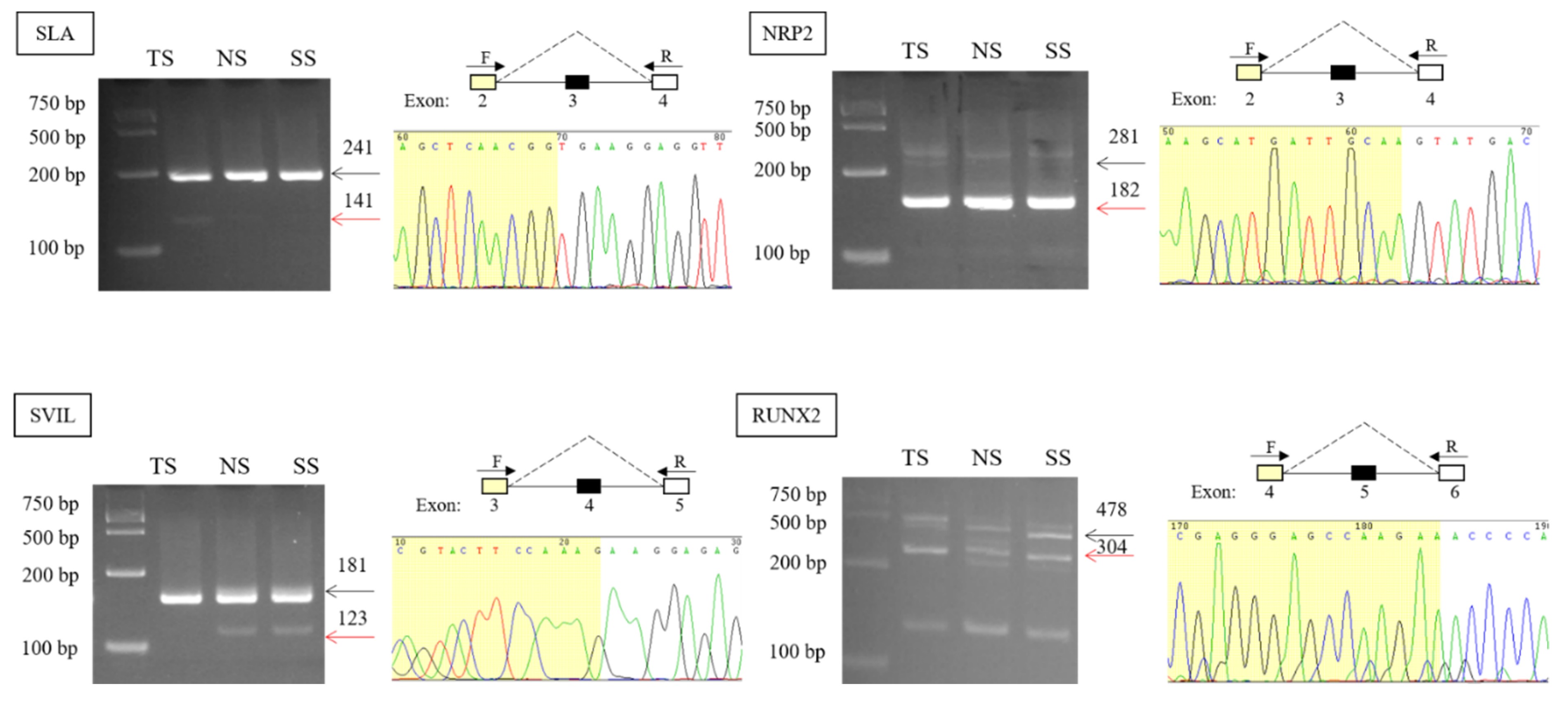
| Gene_Name | IJC_TS | SJC_TS | IJC_NS | SJC_NS | IJC_SS | SJC_SS |
|---|---|---|---|---|---|---|
| SLA | 1730, 701, 912, 599, 833, 980, 1362 | 55, 43, 83, 19, 16, 28, 22 | 1379, 1025, 940, 1399, 1600 | 23, 9, 8, 17, 14 | 10, 229, 697, 271, 717, 700 | 14, 11, 68, 22, 6 |
| NRP2 | 0, 58, 79, 22, 26, 15, 8 | 126, 272, 326, 146, 102, 57, 128 | 4, 1, 7, 0, 11 | 127, 129, 113, 204, 140 | 17, 11, 8, 20, 14 | 172, 182, 148, 184, 156 |
| SVIL | 658, 398, 738, 318, 133, 188, 482 | 20, 4, 4, 13, 18, 14, 36 | 327, 271, 220, 366, 296 | 43, 53, 60, 39, 33 | 220, 228, 289, 371, 427 | 29, 24, 28, 42, 34 |
| RUNX2 | 45, 130, 38, 33, 53, 63, 66, 29 | 6, 47, 24, 3, 4, 6, 1 | 4, 7, 3, 5, 5 | 3, 0, 1, 1, 1 | 11, 13, 4, 7, 1 | 5, 0, 0, 2, 0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, L.; Zheng, G.; Yuan, Y.; Wang, Z.; Liu, C.; Zhang, H.; Lian, L. Exploration of Alternative Splicing (AS) Events in MDV-Infected Chicken Spleens. Genes 2021, 12, 1857. https://doi.org/10.3390/genes12121857
Wang L, Zheng G, Yuan Y, Wang Z, Liu C, Zhang H, Lian L. Exploration of Alternative Splicing (AS) Events in MDV-Infected Chicken Spleens. Genes. 2021; 12(12):1857. https://doi.org/10.3390/genes12121857
Chicago/Turabian StyleWang, Lulu, Gang Zheng, Yiming Yuan, Ziyi Wang, Changjun Liu, Hao Zhang, and Ling Lian. 2021. "Exploration of Alternative Splicing (AS) Events in MDV-Infected Chicken Spleens" Genes 12, no. 12: 1857. https://doi.org/10.3390/genes12121857
APA StyleWang, L., Zheng, G., Yuan, Y., Wang, Z., Liu, C., Zhang, H., & Lian, L. (2021). Exploration of Alternative Splicing (AS) Events in MDV-Infected Chicken Spleens. Genes, 12(12), 1857. https://doi.org/10.3390/genes12121857







