A Genome-Wide Identification Study Reveals That HmoCYP76AD1, HmoDODAα1 and HmocDOPA5GT Involved in Betalain Biosynthesis in Hylocereus
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Gene Identification and Phylogenetic Analyses
2.3. Chromosomal Distribution Analysis
2.4. Relative Expression Analyses by qRT-PCR
2.5. Full-Length cDNA Amplification and Plasmid Construction
2.6. Subcellular Localization Assay
2.7. VIGS of Candidate Genes in H. monacanthus
2.8. Genetic Transformation of A. thaliana
2.9. Transient Expression Assay in N. benthamiana
2.10. Recombinant Expression in Yeast
2.11. Statistical Analysis
3. Results
3.1. Identification and Classification of Candidate Genes Involved in Betalain Biosynthesis
3.2. Genomic Distribution among Pitaya Chromosomes
3.3. Gene Characteristics and Structures
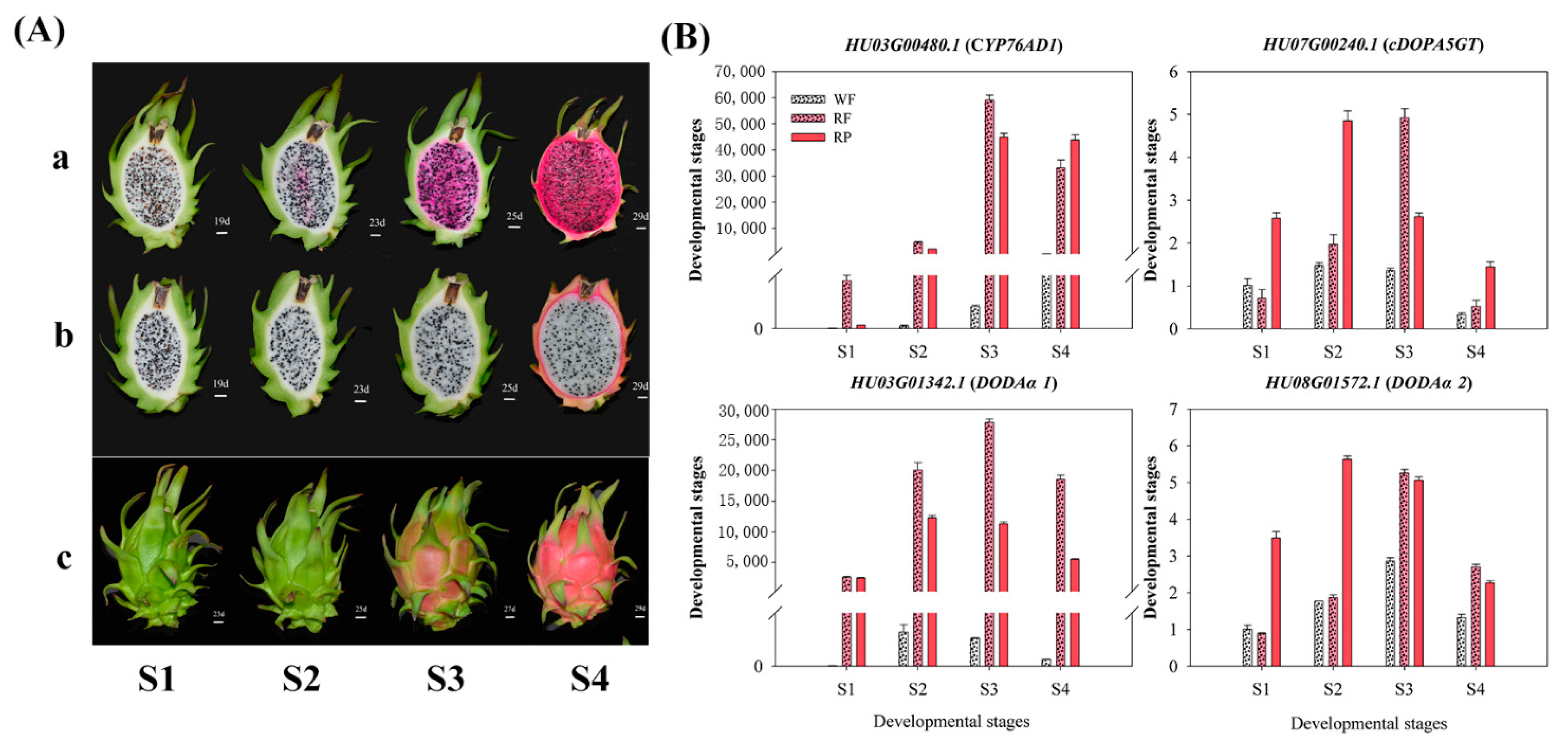
3.4. Expression Patterns of Candidate Genes Involved in Betalain Biosynthesis
3.5. Subcellular Localizations
3.6. VIGS Analyses of Candidate Genes
3.7. Overexpression of HmoDODAα1 and HmoDODAα2 in Arabidopsis and Yeast
3.8. Co-Expression of HmoCYP76AD1, HmoDODAα1/α2 and HmoB5GT1/HmocDOPA5GT in N. benthamiana
3.9. Recombinant Expression of HmoCYP76AD1, HmoDODAα1 and HmocDOPA5GT in Yeast
4. Discussion
4.1. Identification of Genes Involved in Betalain Biosynthesis
4.2. HmoCYP76AD1 Catalyzes Tyrosine to L-DOPA and to Cyclo-DOPA
4.3. HmoDODAα1 Exhibits a High Degree of L-DOPA 4,5-Dioxygenase Activity
4.4. Reconstruction of Heterogenous Production of Betalain Biosynthesis Pathway
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Tenore, G.C.; Novellino, E.; Basile, A. Nutraceutical potential and antioxidant benefits of red pitaya (Hylocereus polyrhizus) extracts. J. Funct. Foods 2012, 4, 129–136. [Google Scholar] [CrossRef]
- Khan, M.I. Plant betalains: Safety, antioxidant activity, clinical efficacy, and bioavailability. Compr. Rev. Food Sci. Food Saf. 2016, 15, 316–330. [Google Scholar] [CrossRef] [Green Version]
- Ibrahim, S.R.M.; Mohamed, G.A.; Khedr, A.I.M.; Zayed, M.F.; El-Kholy, A.A.S. Genus Hylocereus: Beneficial phytochemicals, nutritional importance, and biological relevancea review. J. Food Biochem. 2018, 42, e12491. [Google Scholar] [CrossRef]
- Belhadj Slimen, I.; Najar, T.; Abderrabba, M. Chemical and antioxidant properties of betalains. J. Agric. Food Chem. 2017, 65, 675–689. [Google Scholar] [CrossRef] [PubMed]
- Ibdah, M.; Krins, A.; Seidlitz, H.K.; Heller, W.; Strack, D.; Vogt, T. Spectral dependence of flavonol and betacyanin accumulation in Mesembryanthemum crystallinum under enhanced ultraviolet radiation. Plant Cell Environ. 2002, 25, 1145–1154. [Google Scholar] [CrossRef]
- Savitha, B.C.; Thimmaraju, R.; Bhagyalakshmi, N.; Ravishankar, G.A. Different biotic and abiotic elicitors influence betalain production in hairy root cultures of Beta vulgaris in shake-flask and bioreactor. Process Biochem. 2006, 41, 50–60. [Google Scholar] [CrossRef]
- D’Andrea, R.M.; Andreo, C.S.; Lara, M.V. Deciphering the mechanisms involved in Portulaca oleracea (C4) response to drought: Metabolic changes including crassulacean acid-like metabolism induction and reversal upon re-watering. Physiol. Plant. 2014, 152, 414–430. [Google Scholar] [CrossRef]
- Jain, G.; Gould, K.S. Functional significance of betalain biosynthesis in leaves of Disphyma australe under salinity stress. Environ. Exp. Bot. 2015, 109, 131–140. [Google Scholar] [CrossRef]
- Jain, G.; Schwinn, K.E.; Gould, K.S. Betalain induction by L-DOPA application confers photoprotection to saline-exposed leaves of Disphyma australe. New Phytol. 2015, 207, 1075–1083. [Google Scholar] [CrossRef]
- Sarker, U.; Oba, S. Salinity stress enhances color parameters, bioactive leaf pigments, vitamins, polyphenols, flavonoids and antioxidant activity in selected Amaranthus leafy vegetables. J. Sci. Food Agric. 2019, 99, 2275–2284. [Google Scholar] [CrossRef]
- Strack, D.; Vogt, T.; Schliemann, W. Recent advances in betalain research. Phytochemistry 2003, 62, 247–269. [Google Scholar] [CrossRef]
- Gandia-Herrero, F.; Garcia-Carmona, F. Biosynthesis of betalains: Yellow and violet plant pigments. Trends Plant Sci. 2013, 18, 334–343. [Google Scholar] [CrossRef] [PubMed]
- Hatlestad, G.J.; Sunnadeniya, R.M.; Akhavan, N.A.; Gonzalez, A.; Goldman, I.L.; McGrath, J.M.; Lloyd, A.M. The beet R locus encodes a new cytochrome P450 required for red betalain production. Nat. Genet. 2012, 44, 130–816. [Google Scholar] [CrossRef] [PubMed]
- Brockington, S.F.; Yang, Y.; Gandia-Herrero, F.; Covshoff, S.; Hibberd, J.M.; Sage, R.F.; Wong, G.K.S.; Moore, M.J.; Smith, S.A. Lineage-specific gene radiations underlie the evolution of novel betalain pigmentation in Caryophyllales. New Phytol. 2015, 207, 1170–1180. [Google Scholar] [CrossRef]
- Suzuki, M.; Miyahara, T.; Tokumoto, H.; Hakamatsuka, T.; Goda, Y.; Ozeki, Y.; Sasaki, N. Transposon-mediated mutation of CYP76AD3 affects betalain synthesis and produces variegated flowers in four o’clock (Mirabilis jalapa). J. Plant Physiol. 2014, 171, 1586–1590. [Google Scholar] [CrossRef] [PubMed]
- Imamura, T.; Takagi, H.; Miyazato, A.; Ohki, S.; Mizukoshi, H.; Mori, M. Isolation and characterization of the betalain biosynthesis gene involved in hypocotyl pigmentation of the allotetraploid Chenopodium quinoa. Biochem. Bioph. Res. Commun. 2018, 496, 280–286. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.C.; Chiu, Y.C.; Tsao, N.W.; Chou, Y.L.; Tan, C.M.; Chiang, Y.H.; Liao, P.C.; Lee, Y.C.; Hsieh, L.C.; Wang, S.Y.; et al. Elucidation of the core betalain biosynthesis pathway in Amaranthus tricolor. Sci. Rep. 2021, 11, 6086. [Google Scholar] [CrossRef]
- Polturak, G.; Breitel, D.; Grossman, N.; Sarrion-Perdigones, A.; Weithorn, E.; Pliner, M.; Orzaez, D.; Granell, A.; Rogachev, I.; Aharoni, A. Elucidation of the first committed step in betalain biosynthesis enables the heterologous engineering of betalain pigments in plants. New Phytol. 2016, 210, 269–283. [Google Scholar] [CrossRef] [Green Version]
- Sunnadeniya, R.; Bean, A.; Brown, M.; Akhavan, N.; Hatlestad, G.; Gonzalez, A.; Symonds, V.V.; Lloyd, A. Tyrosine hydroxylation in betalain pigment biosynthesis is performed by cytochrome P450 enzymes in beets (Beta vulgaris). PLoS ONE 2016, 11, e0149417. [Google Scholar] [CrossRef] [Green Version]
- Bean, A.; Sunnadeniya, R.; Akhavan, N.; Campbell, A.; Brown, M.; Lloyd, A. Gain-of-function mutations in beet DODA2 identify key residues for betalain pigment evolution. New Phytol. 2018, 219, 287–296. [Google Scholar] [CrossRef] [Green Version]
- Christinet, L.; Burdet, F.; Zaiko, M.; Hinz, U.; Zryd, J.P. Characterization and functional identification of a novel plant 4,5-extradiol dioxygenase involved in betalain pigment biosynthesis in Portulaca grandiflora. Plant Physiol. 2004, 134, 265–274. [Google Scholar] [CrossRef] [Green Version]
- Sheehan, H.; Feng, T.; Walker-Hale, N.; Lopez-Nieves, S.; Pucker, B.; Guo, R.; Yim, W.C.; Badgami, R.; Timoneda, A.; Zhao, L.; et al. Evolution of l-DOPA 4,5-dioxygenase activity allows for recurrent specialisation to betalain pigmentation in Caryophyllales. New Phytol. 2020, 227, 914–929. [Google Scholar] [CrossRef] [Green Version]
- Sasaki, N.; Abe, Y.; Goda, Y.; Adachi, T.; Kasahara, K.; Ozeki, Y. Detection of dopa 4,5-dioxygenase (DOD) activity using recombinant protein prepared from Escherichia coli cells harboring cDNA encoding DOD from Mirabilis jalapa. Plant Cell Physiol. 2009, 50, 1012–1016. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chung, H.; Schwinn, K.E.; Ngo, H.M.; Lewis, D.H.; Massey, B.; Calcottt, K.E.; Crowhurst, R.; Joyce, D.C.; Gould, K.S.; Davies, K.M.; et al. Characterisation of betalain biosynthesis in Parakeelya flowers identifies the key biosynthetic gene DOD as belonging to an expanded LigB gene family that is conserved in betalain-producing species. Front. Plant Sci. 2015, 6, 499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Casique-Arroyo, G.; Martinez-Gallardo, N.; Gonzalez De La Vara, L.; Delano-Frier, J.P. Betacyanin biosynthetic genes and enzymes are differentially induced by (a)biotic stress in Amaranthus hypochondriacus. PLoS ONE 2014, 9, e99012. [Google Scholar] [CrossRef]
- Heuer, S.; Strack, D. Synthesis of betanin from betanidin and UDP-glucose by a protein preparation from cell suspension cultures of Dorotheanthus bellidiformis (Burm. f.) N. E. Br. Planta 1992, 186, 626–628. [Google Scholar] [CrossRef] [PubMed]
- Heuer, S.; Vogt, T.; Boehm, H.; Strack, D. Partial purification and characterization of UDP-glucose: Betanidin 5-O- and 6-O-glucosyltransferases from cell suspension cultures of Dorotheanthus bellidiformis (Burm. f.) N.E.Br. Planta 1996, 199, 244–250. [Google Scholar] [CrossRef]
- Vogt, T.; Grimm, R.; Strack, D. Cloning and expression of a cDNA encoding betanidin 5-O-glucosyltransferase, a betanidin- and flavonoid-specific enzyme with high homology to inducible glucosyltransferases from the Solanaceae. Plant J. 1999, 19, 509–519. [Google Scholar] [CrossRef] [Green Version]
- Vogt, T. Substrate specificity and sequence analysis define a polyphyletic origin of betanidin 5- and 6-O-glucosyltransferase from Dorotheanthus bellidiformis. Planta 2002, 214, 492–495. [Google Scholar] [CrossRef]
- Sasaki, N.; Adachi, T.; Koda, T.; Ozeki, Y. Detection of UDP-glucose: Cyclo-DOPA 5-O-glucosyltransferase activity in four o’clocks (Mirabilis jalapa L.). FEBS Lett. 2004, 568, 159–162. [Google Scholar] [CrossRef] [Green Version]
- Owen, N.A.; Fahy, K.F.; Griffiths, H. Crassulacean acid metabolism (CAM) offers sustainable bioenergy production and resilience to climate change. GCB Bioenergy 2016, 8, 737–749. [Google Scholar] [CrossRef] [Green Version]
- Lina, A.; Eloisa, L. How do young cacti (seeds and seedlings) from tropical xeric environments cope with extended drought periods? J. Arid Environ. 2018, 154, 1–7. [Google Scholar] [CrossRef]
- Tel-Zur, N.; Abbo, S.; Bar-Zvi, D.; Mizrahi, Y. Genetic relationships among Hylocereus and Selenicereus vine cacti (Cactaceae): Evidence from hybridization and cytological studies. Ann. Bot. 2004, 94, 527–534. [Google Scholar] [CrossRef] [Green Version]
- Ortiz-Hernández, Y.D.; Carrillo-Salazar, J.A. Pitahaya (Hylocereus spp.): A short review. Comun. Sci. 2012, 3, 220–237. [Google Scholar]
- Hua, Q.Z.; Chen, C.J.; Chen, Z.; Chen, P.K.; Ma, Y.W.; Wu, J.Y.; Zheng, J.; Hu, G.B.; Zhao, J.T.; Qin, Y.H. Transcriptomic analysis reveals key genes related to betalain biosynthesis in pulp coloration of Hylocereus polyrhizus. Front. Plant Sci. 2016, 6, 1179. [Google Scholar] [CrossRef] [Green Version]
- Hua, Q.Z.; Zhou, Q.J.; Gan, S.S.; Wu, J.Y.; Chen, C.B.; Li, J.Q.; Ye, Y.X.; Zhao, J.T.; Hu, G.B.; Qin, Y.H. Proteomic analysis of Hylocereus polyrhizus reveals metabolic pathway changes. Int. J. Mol. Sci. 2016, 17, 1606. [Google Scholar] [CrossRef]
- Wu, Y.W.; Xu, J.; Han, X.M.; Qiao, G.; Yang, K.; Wen, Z.; Wen, X.P. Comparative transcriptome analysis combining SMRT- and illumina-based RNA-seq identifies potential candidate genes involved in betalain biosynthesis in pitaya fruit. Int. J. Mol. Sci. 2020, 21, 3288. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.X.; Gao, H.M.; Ming, J.H.; Ding, Z.L.; Lin, X.E.; Zhan, R.L. Combined transcriptome and metabolome analysis of pitaya fruit unveiled the mechanisms underlying peel and pulp color formation. BMC Genom. 2020, 21, 734. [Google Scholar] [CrossRef]
- Xie, F.F.; Hua, Q.Z.; Chen, C.B.; Zhang, L.L.; Zhang, Z.K.; Chen, J.Y.; Zhang, R.; Zhao, J.S.; Hu, G.B.; Zhao, J.T.; et al. Transcriptomics-based identification and characterization of glucosyltransferases involved in betalain biosynthesis in Hylocereus megalanthus. Plant Physiol. Biochem. 2020, 152, 112–124. [Google Scholar] [CrossRef]
- Cheng, M.N.; Huang, Z.J.; Hua, Q.Z.; Shan, W.; Kuang, J.F.; Lu, W.J.; Qin, Y.H.; Chen, J.Y. The WRKY transcription factor HPWRKY44 regulates CytP450-like1 expression in red pitaya fruit (Hylocereus polyrhizus). Hortic. Res. 2017, 4, 17039. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.L.; Chen, C.B.; Xie, F.F.; Hua, Q.Z.; Zhang, Z.K.; Zhang, R.; Chen, J.Y.; Zhao, J.T.; Hu, G.B.; Qin, Y.H. A novel WRKY transcription factor HmoWRKY40 associated with betalain biosynthesis in pitaya (Hylocereus monacanthus) through regulating HmoCYP76AD1. Int. J. Mol. Sci. 2021, 22, 2171. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.F.; Meinhardt, L.W.; Goenaga, R.; Zhang, D.P.; Yin, Y.B. The chromosome-level genome of dragon fruit reveals whole-genome duplication and chromosomal co-localization of betacyanin biosynthetic genes. Hortic. Res. 2021, 8, 63. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.Y.; Xie, F.F.; Cui, Y.Z.; Chen, C.B.; Lu, W.J.; Hu, X.L.; Hua, Q.Z.; Zhao, J.; Wu, Z.J.; Gao, D.; et al. A chromosome-scale genome sequence of pitaya (Hylocereus undatus) provides novel insights into the genome evolution and regulation of betalain biosynthesis. Hortic. Res. 2021, 8, 164. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using realtime quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Chen, C.B.; Wu, J.Y.; Hua, Q.Z.; Tel-Zur, N.; Xie, F.F.; Zhang, Z.K.; Chen, J.Y.; Zhang, R.; Hu, G.B.; Zhao, J.T.; et al. Identification of reliable reference genes for quantitative real-time PCR normalization in pitaya. Plant Methods 2019, 15, 70. [Google Scholar] [CrossRef] [Green Version]
- Hong, S.M.; Bahn, S.C.; Lyu, A.; Jung, H.S.; Ahn, J.H. Identification and testing of superior reference genes for a starting pool of transcript normalization in Arabidopsis. Plant Cell Physiol. 2010, 51, 1694–1706. [Google Scholar] [CrossRef] [Green Version]
- Jailani, A.A.K.; Solanki, V.; Roy, A.; Sivasudha, T.; Mandal, B. A CGMMV genome-replicon vector with partial sequences of coat protein gene efficiently expresses GFP in Nicotiana benthamiana. Virus Res. 2017, 233, 77–85. [Google Scholar] [CrossRef]
- Nelson, B.K.; Cai, X.; Nebenführ, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 2007, 51, 1126–1136. [Google Scholar] [CrossRef]
- Zhang, X.R.; Henriques, R.; Lin, S.S.; Niu, Q.W.; Chua, N.H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 2006, 1, 641–646. [Google Scholar] [CrossRef]
- Timoneda, A.; Sheehan, H.; Feng, T.; Lopez-Nieves, S.; Maeda, H.A.; Brockington, S. Redirecting primary metabolism to boost production of tyrosine-derived specialised metabolites in planta. Sci. Rep. 2018, 8, 17256. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.; Yu, Z.H.; Xiao, X.G. Cytosolic and nuclear co-localization of betalain biosynthetic enzymes in tobacco suggests that betalains are synthesized in the cytoplasm and/or nucleus of betalainic plant cells. Front. Plant Sci. 2017, 8, 831. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stintzing, F.C.; Schieber, A.; Carle, R. Identification of betalains from yellow beet (Beta vulgaris L.) and cactus pear [Opuntia ficus-indica (L.) Mill.] By high-performance liquid chromatography-electrospray ionization mass spectrometry. J. Agric. Food Chem. 2002, 50, 2302–2307. [Google Scholar] [CrossRef] [PubMed]
- Suh, D.H.; Lee, S.; Heo, D.Y.; Kim, Y.; Cho, S.K.; Lee, S.; Lee, C.H. Metabolite profiling of red and white pitayas (Hylocereus polyrhizus and Hylocereus undatus) for comparing betalain biosynthesis and antioxidant activity. J. Agric. Food Chem. 2014, 62, 8764–8771. [Google Scholar] [CrossRef] [PubMed]








Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hua, Q.; Chen, C.; Xie, F.; Zhang, Z.; Zhang, R.; Zhao, J.; Hu, G.; Qin, Y. A Genome-Wide Identification Study Reveals That HmoCYP76AD1, HmoDODAα1 and HmocDOPA5GT Involved in Betalain Biosynthesis in Hylocereus. Genes 2021, 12, 1858. https://doi.org/10.3390/genes12121858
Hua Q, Chen C, Xie F, Zhang Z, Zhang R, Zhao J, Hu G, Qin Y. A Genome-Wide Identification Study Reveals That HmoCYP76AD1, HmoDODAα1 and HmocDOPA5GT Involved in Betalain Biosynthesis in Hylocereus. Genes. 2021; 12(12):1858. https://doi.org/10.3390/genes12121858
Chicago/Turabian StyleHua, Qingzhu, Canbin Chen, Fangfang Xie, Zhike Zhang, Rong Zhang, Jietang Zhao, Guibing Hu, and Yonghua Qin. 2021. "A Genome-Wide Identification Study Reveals That HmoCYP76AD1, HmoDODAα1 and HmocDOPA5GT Involved in Betalain Biosynthesis in Hylocereus" Genes 12, no. 12: 1858. https://doi.org/10.3390/genes12121858
APA StyleHua, Q., Chen, C., Xie, F., Zhang, Z., Zhang, R., Zhao, J., Hu, G., & Qin, Y. (2021). A Genome-Wide Identification Study Reveals That HmoCYP76AD1, HmoDODAα1 and HmocDOPA5GT Involved in Betalain Biosynthesis in Hylocereus. Genes, 12(12), 1858. https://doi.org/10.3390/genes12121858






