Flavonoids as Potential Drugs for VPS13-Dependent Rare Neurodegenerative Diseases
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains, Media, and Growth Conditions
2.2. Plasmids
2.3. Drug Screening Assay
2.4. Screen of an In-House Library of Natural Compounds and Their Derivatives
3. Results
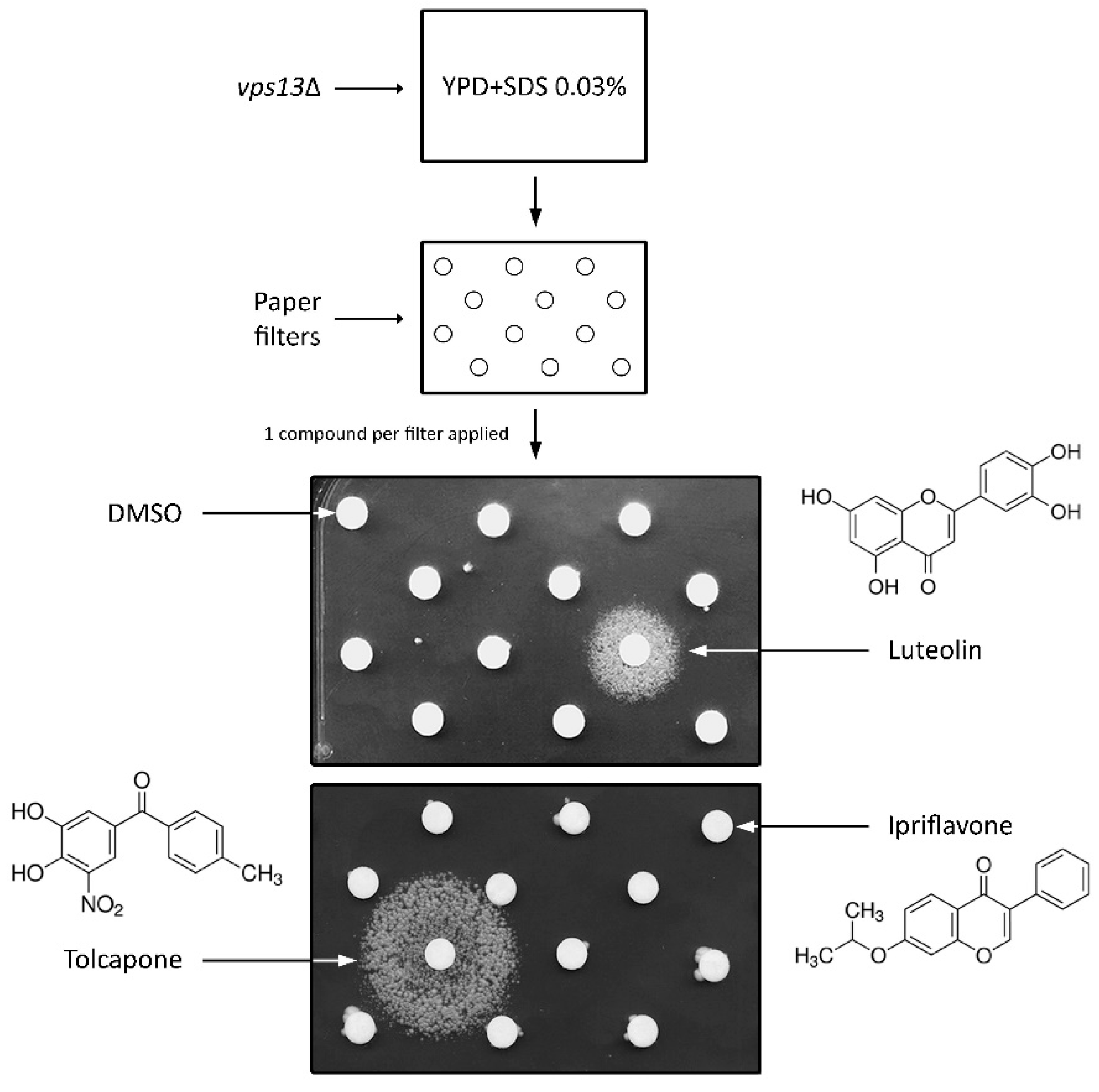
3.1. Drug Library Screening Assay Using vps13∆ Strain and SDS Hypersensitivity Phenotype Revealed Luteolin and Tolcapone as Active Compounds
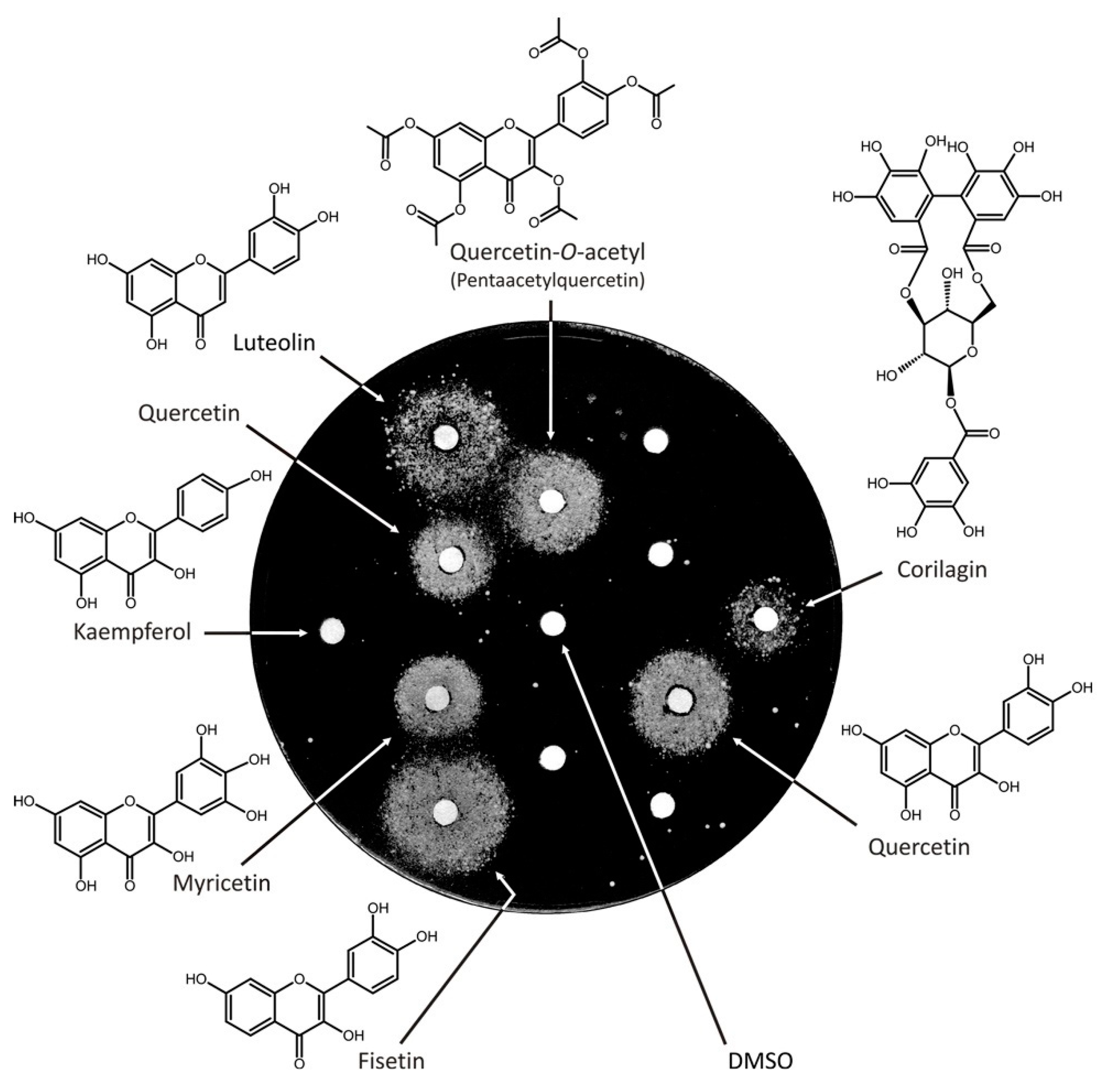
3.2. Screen of an In-House Library of Natural Compounds and Their Derivatives Revealed Six Compounds from Two Structural Classes, Flavonoids, and Tannins
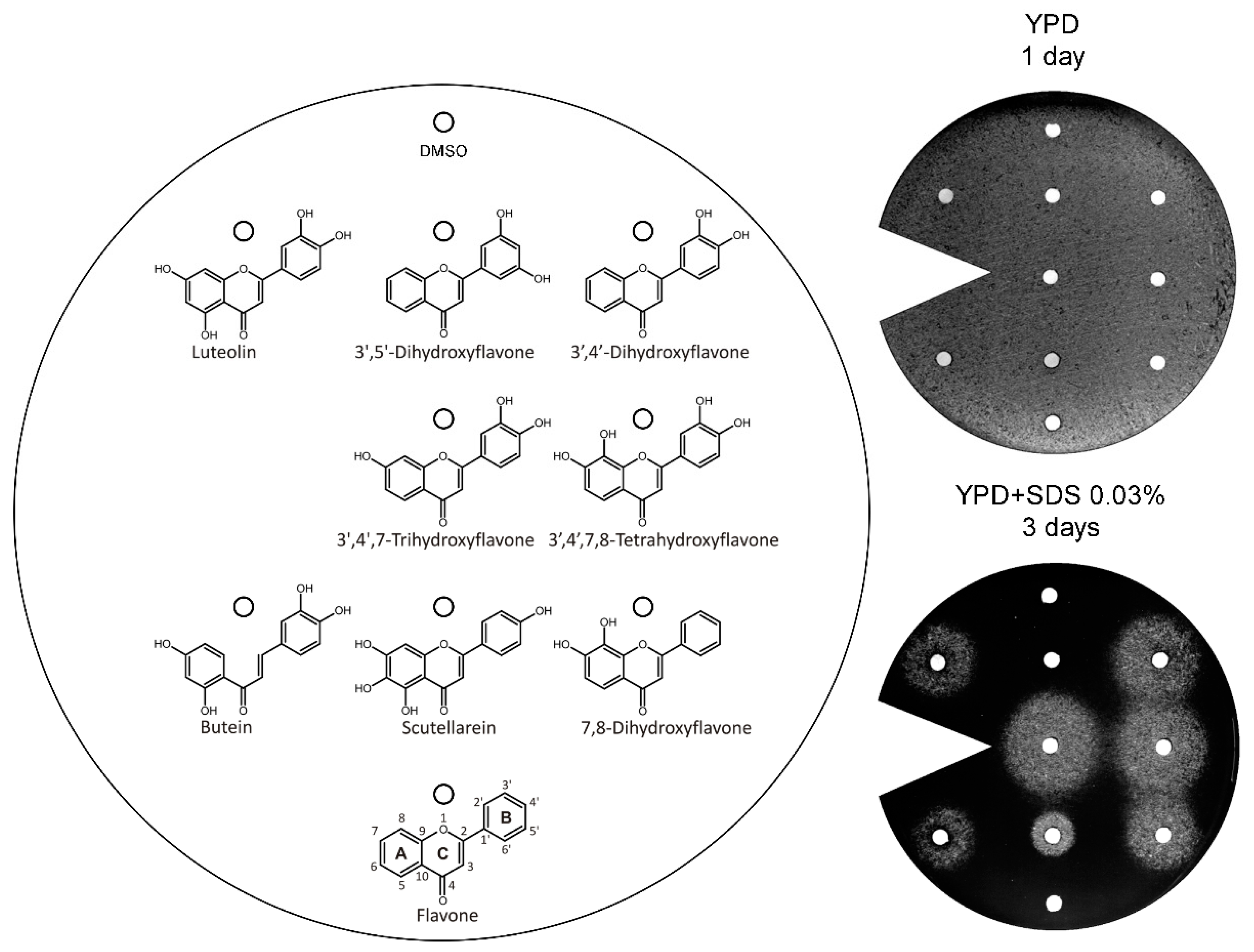
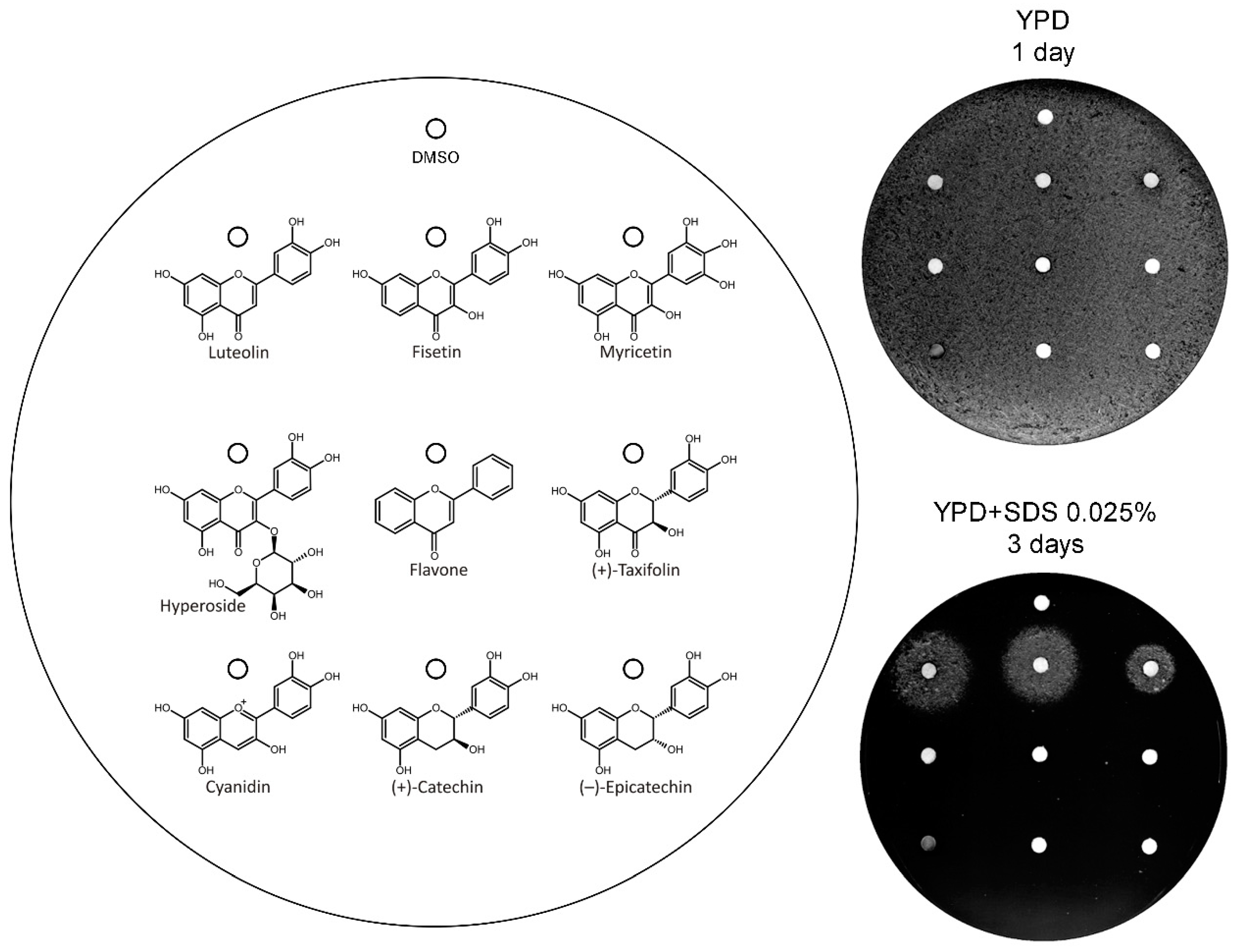
3.3. Analysis of the Structural Determinants of Flavonoids Activity as vps13∆ Suppressors
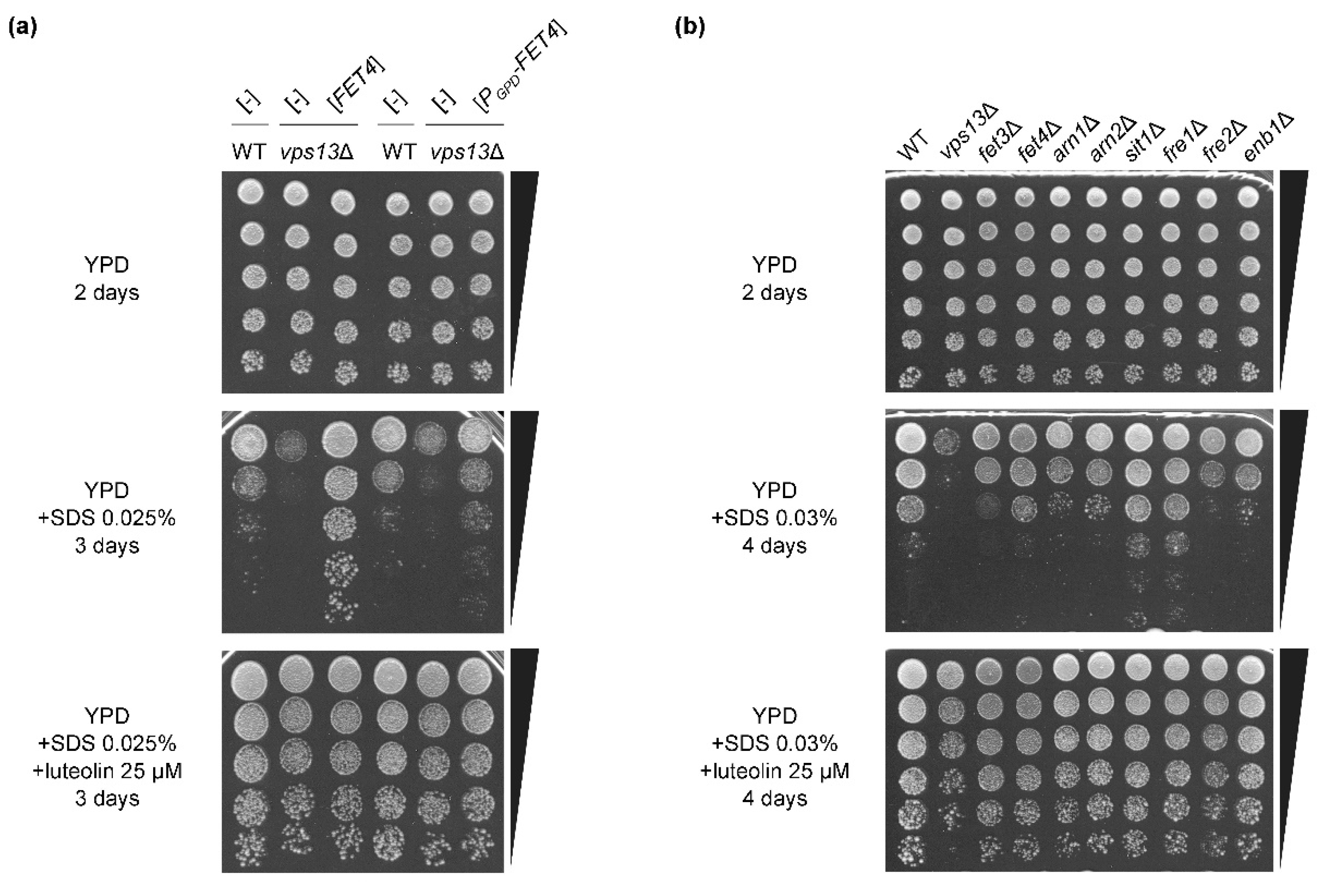
3.4. The FET4 Gene Encoding Iron Transporter Fet4 Suppresses vps13Δ and Mutations Affecting Iron Transport Result in SDS Hypersensitivity
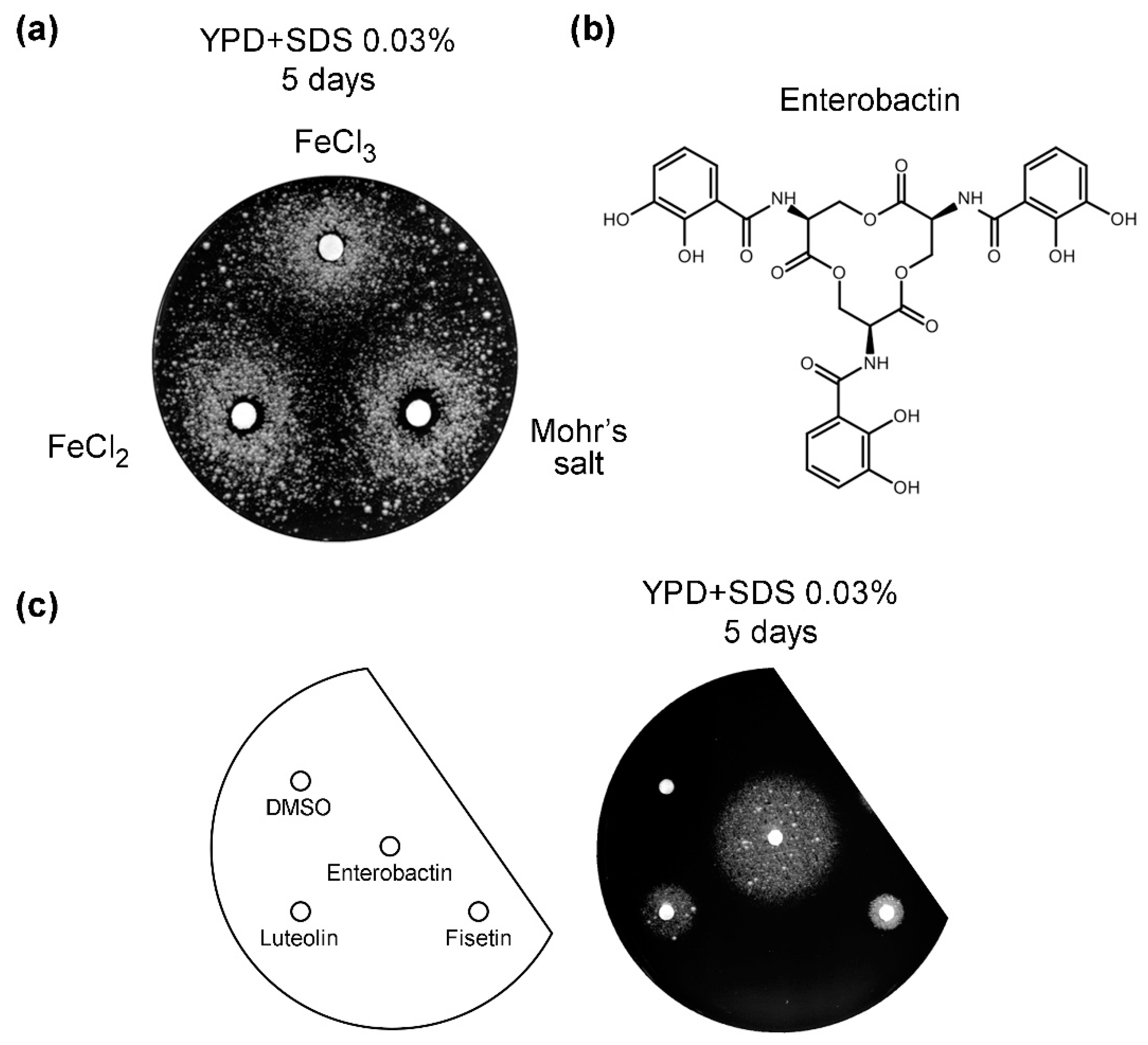
3.5. Iron Salts and Enterobactin Are Chemical Suppressors of vps13Δ
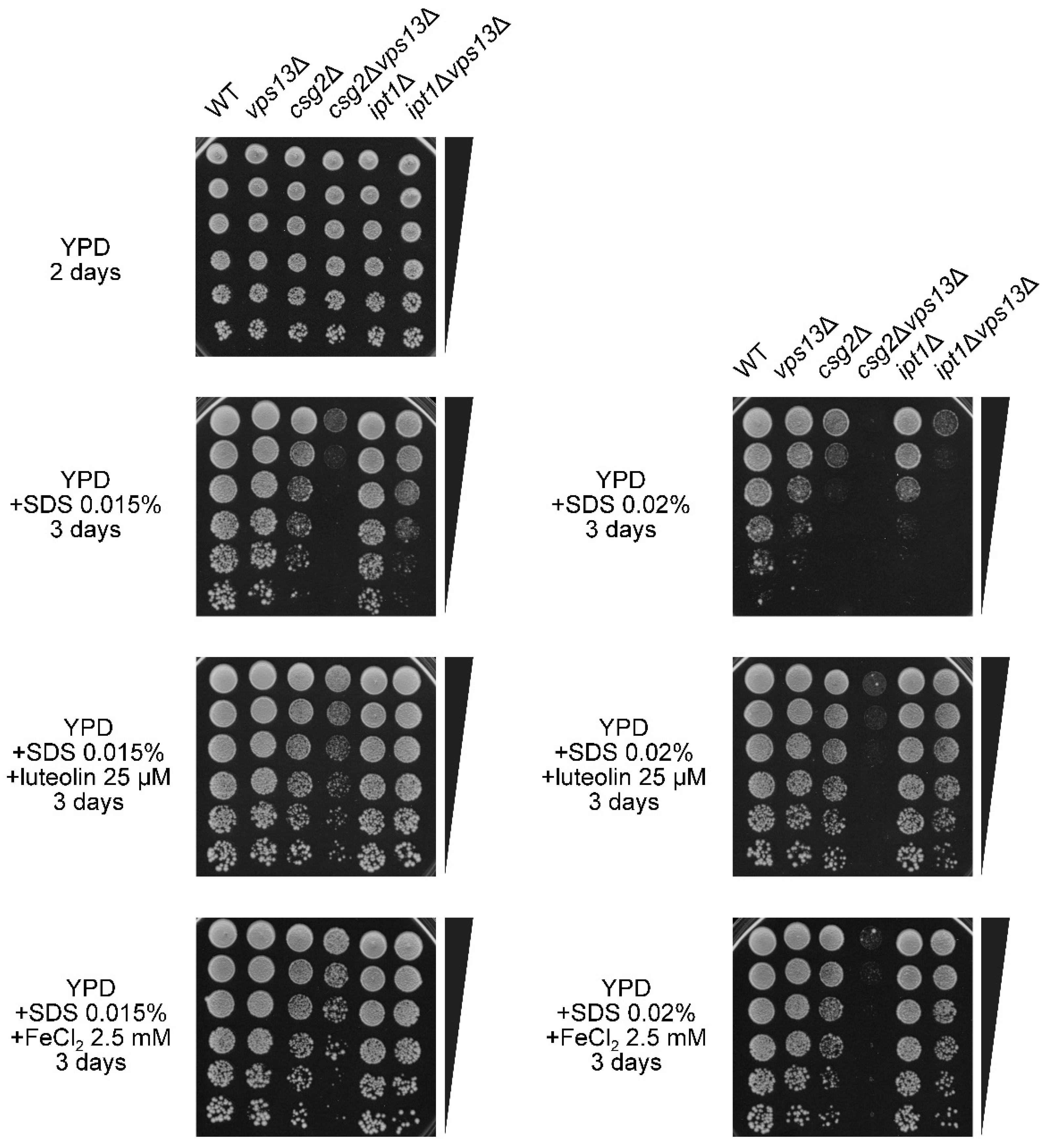
3.6. Characterization of the Possible Mechanism of Flavonoids Action
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Shamsi, T.N.; Athar, T.; Parveen, R.; Fatima, S. A review on protein misfolding, aggregation and strategies to prevent related ailments. Int. J. Biol. Macromol. 2017, 105, 993–1000. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.A.; Rizzo, V.; Puthanveettil, S.V. Pathologies of axonal transport in neurodegenerative diseases. Transl. Neurosci. 2012, 3, 355–372. [Google Scholar] [CrossRef]
- Belaidi, A.A.; Bush, A.I. Iron neurochemistry in Alzheimer’s disease and Parkinson’s disease: Targets for therapeutics. J. Neurochem. 2016, 139, 179–197. [Google Scholar] [CrossRef]
- Hroudová, J.; Singh, N.; Fišar, Z. Mitochondrial dysfunctions in neurodegenerative diseases: Relevance to Alzheimer’s disease. Biomed. Res. Int. 2014, 2014, 175062. [Google Scholar] [CrossRef] [PubMed]
- Jung, H.H.; Danek, A.; Walker, R.H. Neuroacanthocytosis Syndromes. Orphanet J. Rare Dis. 2011, 6, 68. [Google Scholar] [CrossRef]
- Rampoldi, L.; Dobson-Stone, C.; Rubio, J.P.; Danek, A.; Chalmers, R.M.; Wood, N.W.; Verellen, C.; Ferrer, X.; Malandrini, A.; Fabrizi, G.M.; et al. A conserved sorting-associated protein is mutant in chorea-acanthocytosis. Nat. Genet. 2001, 28, 119–120. [Google Scholar] [CrossRef]
- Ueno, S.I.; Maruki, Y.; Nakamura, M.; Tomemori, Y.; Kamae, K.; Tanabe, H.; Yamashita, Y.; Matsuda, S.; Kaneko, S.; Sano, A. The gene encoding a newly discovered protein, chorein, is mutated in chorea-acanthocytosis. Nat. Genet. 2001, 28, 121–122. [Google Scholar] [CrossRef] [PubMed]
- Velayos-Baeza, A.; Vettori, A.; Copley, R.R.; Dobson-Stone, C.; Monaco, A.P. Analysis of the human VPS13 gene family. Genomics 2004, 84, 536–549. [Google Scholar] [CrossRef]
- Rzepnikowska, W.; Flis, K.; Muñoz-Braceras, S.; Menezes, R.; Escalante, R.; Zoladek, T. Yeast and other lower eukaryotic organisms for studies of Vps13 proteins in health and disease. Traffic 2017, 18, 711–719. [Google Scholar] [CrossRef]
- Kolehmainen, J.; Black, G.C.M.; Saarinen, A.; Chandler, K.; Clayton-Smith, J.; Träskelin, A.-L.; Perveen, R.; Kivitie-Kallio, S.; Norio, R.; Warburg, M.; et al. Cohen Syndrome Is Caused by Mutations in a Novel Gene, COH1, Encoding a Transmembrane Protein with a Presumed Role in Vesicle-Mediated Sorting and Intracellular Protein Transport. Am. J. Hum. Genet. 2003, 72, 1359–1369. [Google Scholar] [CrossRef]
- Lesage, S.; Drouet, V.; Majounie, E.; Deramecourt, V.; Jacoupy, M.; Nicolas, A.; Cormier-Dequaire, F.; Hassoun, S.M.; Pujol, C.; Ciura, S.; et al. Loss of VPS13C Function in Autosomal-Recessive Parkinsonism Causes Mitochondrial Dysfunction and Increases PINK1/Parkin-Dependent Mitophagy. Am. J. Hum. Genet. 2016, 98, 500–513. [Google Scholar] [CrossRef]
- Gauthier, J.; Meijer, I.A.; Lessel, D.; Mencacci, N.E.; Krainc, D.; Hempel, M.; Tsiakas, K.; Prokisch, H.; Rossignol, E.; Helm, M.H.; et al. Recessive mutations in VPS13D cause childhood onset movement disorders. Ann. Neurol. 2018, 83, 1089–1095. [Google Scholar] [CrossRef] [PubMed]
- Koh, K.; Ishiura, H.; Shimazaki, H.; Tsutsumiuchi, M.; Ichinose, Y.; Nan, H.; Hamada, S.; Ohtsuka, T.; Tsuji, S.; Takiyama, Y. VPS13D-related disorders presenting as a pure and complicated form of hereditary spastic paraplegia. Mol. Genet. Genomic Med. 2020, 8, e1108. [Google Scholar] [CrossRef] [PubMed]
- Seong, E.; Insolera, R.; Dulovic, M.; Kamsteeg, E.J.; Trinh, J.; Brüggemann, N.; Sandford, E.; Li, S.; Ozel, A.B.; Li, J.Z.; et al. Mutations in VPS13D lead to a new recessive ataxia with spasticity and mitochondrial defects. Ann. Neurol. 2018, 83, 1075–1088. [Google Scholar] [CrossRef] [PubMed]
- Peikert, K.; Danek, A.; Hermann, A. Current state of knowledge in Chorea-Acanthocytosis as core Neuroacanthocytosis syndrome. Eur. J. Med. Genet. 2018, 61, 699–705. [Google Scholar] [CrossRef]
- Rodrigues, J.M.; Fernandes, H.D.; Caruthers, C.; Braddock, S.R.; Knutsen, A.P. Cohen Syndrome: Review of the Literature. Cureus 2018, 10, e3330. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, A.V.; Vilaça, R.; Santos, C.N.; Costa, V.; Menezes, R. Exploring the power of yeast to model aging and age-related neurodegenerative disorders. Biogerontology 2017, 18, 3–34. [Google Scholar] [CrossRef]
- Menezes, R.; Tenreiro, S.; Macedo, D.; Santos, C.N.; Outeiro, T.F. From the baker to the bedside: Yeast models of parkinson’s disease. Microb. Cell 2015, 2, 262–279. [Google Scholar] [CrossRef]
- Winderickx, J.; Delay, C.; De Vos, A.; Klinger, H.; Pellens, K.; Vanhelmont, T.; Van Leuven, F.; Zabrocki, P. Protein folding diseases and neurodegeneration: Lessons learned from yeast. Biochim. Biophys. Acta-Mol. Cell Res. 2008, 1783, 1381–1395. [Google Scholar] [CrossRef]
- Dudnik, A.; Almeida, A.F.; Andrade, R.; Avila, B.; Bañados, P.; Barbay, D.; Bassard, J.E.; Benkoulouche, M.; Bott, M.; Braga, A.; et al. BacHBerry: BACterial Hosts for production of Bioactive phenolics from bERRY fruits. Phytochem. Rev. 2018, 17, 291–326. [Google Scholar] [CrossRef]
- Chung, C.Y.; Khurana, V.; Auluck, P.K.; Tardiff, D.F.; Mazzulli, J.R.; Soldner, F.; Baru, V.; Lou, Y.; Freyzon, Y.; Cho, S.; et al. Identification and rescue of α-synuclein toxicity in Parkinson patient-derived neurons. Science 2013, 342, 983–987. [Google Scholar] [CrossRef]
- Dajas, F.; Rivera, F.; Blasina, F.; Arredondo, F.; Echeverry, C.; Lafon, L.; Morquio, A.; Heizen, H. Cell culture protection and in vivo neuroprotective capacity of flavonoids. Neurotox. Res. 2003, 5, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Griffioen, G.; Duhamel, H.; Van Damme, N.; Pellens, K.; Zabrocki, P.; Pannecouque, C.; Van Leuven, F.; Winderickx, J.; Wera, S. A yeast-based model of α-synucleinopathy identifies compounds with therapeutic potential. Biochim. Biophys. Acta-Mol. Basis Dis. 2006, 1762, 312–318. [Google Scholar] [CrossRef] [PubMed]
- Rencus-Lazar, S.; DeRowe, Y.; Adsi, H.; Gazit, E.; Laor, D. Yeast models for the study of amyloid-associated disorders and development of future therapy. Front. Mol. Biosci. 2019, 6, 15. [Google Scholar] [CrossRef] [PubMed]
- Tardiff, D.F.; Jui, N.T.; Khurana, V.; Tambe, M.A.; Thompson, M.L.; Chung, C.Y.; Kamadurai, H.B.; Kim, H.T.; Lancaster, A.K.; Caldwell, K.A.; et al. Yeast Reveal a “Druggable” Rsp5/Nedd4 Network that Ameliorates-Synuclein Toxicity in Neurons. Science 2013, 342, 979–983. [Google Scholar] [CrossRef]
- Kumar, S.; Pandey, A.K. Chemistry and biological activities of flavonoids: An overview. ScientificWorldJournal 2013, 2013, 162750. [Google Scholar] [CrossRef] [PubMed]
- Panche, A.N.; Diwan, A.D.; Chandra, S.R. Flavonoids: An overview. J. Nutr. Sci. 2016, 5, e47. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.y.; Li, Q.; Bi, K.s. Bioactive flavonoids in medicinal plants: Structure, activity and biological fate. Asian J. Pharm. Sci. 2018, 13, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Naeimi, A.F.; Alizadeh, M. Antioxidant properties of the flavonoid fisetin: An updated review of in vivo and in vitro studies. Trends Food Sci. Technol. 2017, 70, 34–44. [Google Scholar] [CrossRef]
- Rodríguez-García, C.; Sánchez-Quesada, C.; Gaforio, J.J. Dietary Flavonoids as Cancer Chemopreventive Agents: An Updated Review of Human Studies. Antioxidants 2019, 8, 137. [Google Scholar] [CrossRef]
- De Andrade Teles, R.B.; Diniz, T.C.; Costa Pinto, T.C.; de Oliveira Júnior, R.G.; Gama e Silva, M.; de Lavor, É.M.; Fernandes, A.W.C.; de Oliveira, A.P.; de Almeida Ribeiro, F.P.R.; da Silva, A.A.M.; et al. Flavonoids as Therapeutic Agents in Alzheimer’s and Parkinson’s Diseases: A Systematic Review of Preclinical Evidences. Oxid. Med. Cell. Longev. 2018, 2018, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Nabavi, S.F.; Braidy, N.; Gortzi, O.; Sobarzo-Sanchez, E.; Daglia, M.; Skalicka-Woźniak, K.; Nabavi, S.M. Luteolin as an anti-inflammatory and neuroprotective agent: A brief review. Brain Res. Bull. 2015, 119, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Figueira, I.; Menezes, R.; Macedo, D.; Costa, I.; Dos Santos, C.N. Polyphenols Beyond Barriers: A Glimpse into the Brain. Curr. Neuropharmacol. 2017, 15, 562–594. [Google Scholar] [CrossRef] [PubMed]
- Kaniak-Golik, A.; Skoneczna, A. Mitochondria-nucleus network for genome stability. Free Radic. Biol. Med. 2015, 82, 73–104. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Pastor, M.T.; Perea-García, A.; Puig, S. Mechanisms of iron sensing and regulation in the yeast Saccharomyces cerevisiae. World J. Microbiol. Biotechnol. 2017, 33, 75. [Google Scholar] [CrossRef] [PubMed]
- Pimentel, C.; Vicente, C.; Menezes, R.A.; Caetano, S.; Carreto, L.; Rodrigues-Pousada, C. The Role of the Yap5 Transcription Factor in Remodeling Gene Expression in Response to Fe Bioavailability. PLoS ONE 2012, 7, e37434. [Google Scholar] [CrossRef] [PubMed]
- Cyert, M.S.; Philpott, C.C. Regulation of Cation Balance in Saccharomyces cerevisiae. Genetics 2013, 193, 677–713. [Google Scholar] [CrossRef]
- Dix, D.R.; Bridgham, J.T.; Broderius, M.A.; Byersdorfer, C.A.; Eide, D.J. The FET4 gene encodes the low affinity Fe(II) transport protein of Saccharomyces cerevisiae. J. Biol. Chem. 1994, 269, 26092–26099. [Google Scholar]
- Hassett, R.; Dix, D.R.; Eide, D.J.; Kosman, D.J. The Fe(II) permease Fet4p functions as a low affinity copper transporter and supports normal copper trafficking in Saccharomyces cerevisiae. Biochem. J. 2000, 351 Pt 2, 477–484. [Google Scholar] [CrossRef]
- Waters, B.M.; Eide, D.J. Combinatorial control of yeast FET4 gene expression by iron, zinc, and oxygen. J. Biol. Chem. 2002, 277, 33749–33757. [Google Scholar] [CrossRef] [PubMed]
- Kolakowski, D.; Kaminska, J.; Zoladek, T. The binding of the APT1 domains to phosphoinositides is regulated by metal ions in vitro. Biochim. Biophys. Acta-Biomembr. 2020, 1862, 183349. [Google Scholar] [CrossRef] [PubMed]
- Rzepnikowska, W.; Flis, K.; Kaminska, J.; Grynberg, M.; Urbanek, A.; Ayscough, K.R.; Zoladek, T. Amino acid substitution equivalent to human choreaacanthocytosis I2771R in yeast Vps13 protein affects its binding to phosphatidylinositol 3-phosphate. Hum. Mol. Genet. 2017, 26, 1497–1510. [Google Scholar] [CrossRef] [PubMed]
- Soczewka, P.; Kolakowski, D.; Smaczynska-de Rooij, I.; Rzepnikowska, W.; Ayscough, K.R.; Kaminska, J.; Zoladek, T. Yeast-model-based study identified myosin- and calcium-dependent calmodulin signalling as a potential target for drug intervention in chorea-acanthocytosis. Dis. Model. Mech. 2019, 12, dmm036830. [Google Scholar] [CrossRef] [PubMed]
- Kabir, M.A.; Kaminska, J.; Segel, G.B.; Bethlendy, G.; Lin, P.; Della Seta, F.; Blegen, C.; Swiderek, K.M.; Zoładek, T.; Arndt, K.T.; et al. Physiological effects of unassembled chaperonin Cct subunits in the yeast Saccharomyces cerevisiae. Yeast 2005, 22, 219–239. [Google Scholar] [CrossRef]
- Gietz, R.D.; Sugino, A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 1988, 14, 527–534. [Google Scholar] [CrossRef]
- Sikorski, R.S.; Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 1989, 122, 19–27. [Google Scholar]
- Bach, S.; Talarek, N.; Andrieu, T.; Vierfond, J.M.; Mettey, Y.; Galons, H.; Dormont, D.; Meijer, L.; Cullin, C.; Blondel, M. Isolation of drugs active against mammalian prions using a yeast-based screening assay. Nat. Biotechnol. 2003, 21, 1075–1081. [Google Scholar] [CrossRef] [PubMed]
- Couplan, E.; Aiyar, R.S.; Kucharczyk, R.; Kabala, A.; Ezkurdia, N.; Gagneur, J.; St Onge, R.P.; Salin, B.; Soubigou, F.; Le Cann, M.; et al. A yeast-based assay identifies drugs active against human mitochondrial disorders. Proc. Natl. Acad. Sci. USA 2011, 108, 11989–11994. [Google Scholar] [CrossRef]
- Almeida, A.F.; Santos, C.N.; Ventura, M.R. Synthesis of New Sulfated and Glucuronated Metabolites of Dietary Phenolic Compounds Identified in Human Biological Samples. J. Agric. Food Chem. 2017, 65, 6460–6466. [Google Scholar] [CrossRef]
- Heins-Marroquin, U.; Jung, P.P.; Cordero-Maldonado, M.L.; Crawford, A.D.; Linster, C.L. Phenotypic assays in yeast and zebrafish reveal drugs that rescue ATP13A2 deficiency. Brain Commun. 2019, 1, fcz019. [Google Scholar] [CrossRef]
- Antonini, A.; Abbruzzese, G.; Barone, P.; Bonuccelli, U.; Lopiano, L.; Onofrj, M.; Zappia, M.; Quattrone, A. COMT inhibition with tolcapone in the treatment algorithm of patients with Parkinson’s disease (PD): Relevance for motor and non-motor features. Neuropsychiatr. Dis. Treat. 2008, 4, 1–9. [Google Scholar] [CrossRef]
- Bors, W.; Heller, W.; Michel, C.; Saran, M. Flavonoids as antioxidants: Determination of radical-scavenging efficiencies. Methods Enzymol. 1990, 186, 343–355. [Google Scholar] [PubMed]
- Fernandez, M.T.; Mira, M.L.; Florêncio, M.H.; Jennings, K.R. Iron and copper chelation by flavonoids: An electrospray mass spectrometry study. J. Inorg. Biochem. 2002, 92, 105–111. [Google Scholar] [CrossRef]
- Rice-Evans, C.A.; Miller, N.J.; Paganga, G. Structure-antioxidant activity relationships of flavonoids and phenolic acids. Free Radic. Biol. Med. 1996, 20, 933–956. [Google Scholar] [CrossRef]
- Khokhar, S.; Owusu Apenten, R.K. Iron binding characteristics of phenolic compounds: Some tentative structure-activity relations. Food Chem. 2003, 81, 133–140. [Google Scholar] [CrossRef]
- Martins, T.S.; Pereira, C.; Canadell, D.; Vilaça, R.; Teixeira, V.; Moradas-Ferreira, P.; de Nadal, E.; Posas, F.; Costa, V. The Hog1p kinase regulates Aft1p transcription factor to control iron accumulation. Biochim. Biophys. Acta-Mol. Cell Biol. Lipids 2018, 1863, 61–70. [Google Scholar] [CrossRef]
- Heymann, P.; Ernst, J.F.; Winkelmann, G. A gene of the major facilitator superfamily encodes a transporter for enterobactin (Enb1p) in Saccharomyces cerevisiae. BioMetals 2000, 13, 65–72. [Google Scholar] [CrossRef]
- López-García, B.; Gandía, M.; Muñoz, A.; Carmona, L.; Marcos, J.F. A genomic approach highlights common and diverse effects and determinants of susceptibility on the yeast Saccharomyces cerevisiae exposed to distinct antimicrobial peptides. BMC Microbiol. 2010, 10, 289. [Google Scholar] [CrossRef]
- Zhang, M.; Liu, Z.; Yu, Q.; Mao, J.; Zhang, B.; Xing, L.; Li, M. Deletion of genes encoding fatty acid desaturases leads to alterations in stress sensitivity in Pichia pastoris. FEMS Yeast Res. 2015, 15, fov020. [Google Scholar] [CrossRef] [PubMed]
- Dickson, R.C.; Nagiec, E.E.; Wells, G.B.; Nagiec, M.M.; Lester, R.L. Synthesis of mannose-(inositol-P)2-ceramide, the major sphingolipid in Saccharomyces cerevisiae, requires the IPT1 (YDR072c) gene. J. Biol. Chem. 1997, 272, 29620–29625. [Google Scholar] [CrossRef]
- Beeler, T.J.; Fu, D.; Rivera, J.; Monaghan, E.; Gable, K.; Dunn, T.M. SUR1 (CSG1/BCL21), a gene necessary for growth of Saccharomyces cerevisiae in the presence of high Ca2+ concentrations at 37 °C, is required for mannosylation of inositolphosphorylceramide. Mol. Gen. Genet. MGG 1997, 255, 570–579. [Google Scholar] [CrossRef] [PubMed]
- Uemura, S.; Kihara, A.; Inokuchi, J.I.; Igarashi, Y. Csg1p and Newly Identified Csh1p Function in Mannosylinositol Phosphorylceramide Synthesis by Interacting with Csg2p. J. Biol. Chem. 2003, 278, 45049–45055. [Google Scholar] [CrossRef] [PubMed]
- Di Meo, F.; Valentino, A.; Petillo, O.; Peluso, G.; Filosa, S.; Crispi, S. Bioactive Polyphenols and Neuromodulation: Molecular Mechanisms in Neurodegeneration. Int. J. Mol. Sci. 2020, 21, 2564. [Google Scholar] [CrossRef] [PubMed]
- Casamayor, A.; Torrance, P.D.; Kobayashi, T.; Thorner, J.; Alessi, D.R. Functional counterparts of mammalian protein kinases PDK1 and SGK in budding yeast. Curr. Biol. 1999, 9, 186–197. [Google Scholar] [CrossRef]
- Hechtberger, P.; Zinser, E.; Saf, R.; Hummel, K.; Paltauf, F.; Daum, G. Characterization, quantification and subcellular localization of inositol-containing sphingolipids of the yeast, Saccharomyces cerevisiae. Eur. J. Biochem. 1994, 225, 641–649. [Google Scholar] [CrossRef] [PubMed]
- Redding, K.; Brickner, J.H.; Marschall, L.G.; Nichols, J.W.; Fuller, R.S. Allele-specific suppression of a defective trans-Golgi network (TGN) localization signal in Kex2p identifies three genes involved in localization of TGN transmembrane proteins. Mol. Cell. Biol. 1996, 16, 6208–6217. [Google Scholar] [CrossRef] [PubMed]
- Brickner, J.H.; Fuller, R.S. SOI1 encodes a novel, conserved protein that promotes TGN-endosomal cycling of Kex2p and other membrane proteins by modulating the function of two TGN localization signals. J. Cell Biol. 1997, 139, 23–36. [Google Scholar] [CrossRef]
- Fröhlich, F.; Petit, C.; Kory, N.; Christiano, R.; Hannibal-Bach, H.K.; Graham, M.; Liu, X.; Ejsing, C.S.; Farese, R.V.; Walther, T.C. The GARP complex is required for cellular sphingolipid homeostasis. Elife 2015, 4, e08712. [Google Scholar]
- Sato, K.; Noda, Y.; Yoda, K. Kei1: A novel subunit of inositolphosphorylceramide synthase, essential for its enzyme activity and golgi localization. Mol. Biol. Cell 2009, 20, 4444–4457. [Google Scholar] [CrossRef]
- Park, J.S.; Neiman, A.M. VPS13 regulates membrane morphogenesis during sporulation in Saccharomyces cerevisiae. J. Cell Sci. 2012, 125, 3004–3011. [Google Scholar] [CrossRef]
- Breslow, D.K.; Collins, S.R.; Bodenmiller, B.; Aebersold, R.; Simons, K.; Shevchenko, A.; Ejsing, C.S.; Weissman, J.S. Orm family proteins mediate sphingolipid homeostasis. Nature 2010, 463, 1048–1053. [Google Scholar] [CrossRef] [PubMed]
- Walther, T.C. Keeping sphingolipid levels nORMal. Proc. Natl. Acad. Sci. USA 2010, 107, 5701–5702. [Google Scholar] [CrossRef] [PubMed]
- Brice, S.E.; Alford, C.W.; Cowart, L.A. Modulation of sphingolipid metabolism by the phosphatidylinositol-4-phosphate phosphatase Sac1p through regulation of phosphatidylinositol in Saccharomyces cerevisiae. J. Biol. Chem. 2009, 284, 7588–7596. [Google Scholar] [CrossRef] [PubMed]
- Davis, D.; Kannan, M.; Wattenberg, B. Orm/ORMDL proteins: Gate guardians and master regulators. Adv. Biol. Regul. 2018, 70, 3–18. [Google Scholar] [CrossRef]
- Muir, A.; Ramachandran, S.; Roelants, F.M.; Timmons, G.; Thorner, J. TORC2-dependent protein kinase Ypk1 phosphorylates ceramide synthase to stimulate synthesis of complex sphingolipids. Elife 2014, 3, e03779. [Google Scholar] [CrossRef]
- Roelants, F.M.; Torrance, P.D.; Thorner, J. Differential roles of PDK1- and PDK2- phosphorylation sites in the yeast AGC kinases Ypk1, Pkc1 and Sch9. Microbiology 2004, 150, 3289–3304. [Google Scholar] [CrossRef]
- Lee, Y.J.; Huang, X.; Kropat, J.; Henras, A.; Merchant, S.S.; Dickson, R.C.; Chanfreau, G.F. Sphingolipid signaling mediates iron toxicity. Cell Metab. 2012, 16, 90–96. [Google Scholar] [CrossRef]
- Xue, Y.; Schmollinger, S.; Attar, N.; Campos, O.A.; Vogelauer, M.; Carey, M.F.; Merchant, S.S.; Kurdistani, S.K. Endoplasmic reticulum–mitochondria junction is required for iron homeostasis. J. Biol. Chem. 2017, 292, 13197–13204. [Google Scholar] [CrossRef]
- Abdel Hadi, L.; Di Vito, C.; Marfia, G.; Ferraretto, A.; Tringali, C.; Viani, P.; Riboni, L. Sphingosine Kinase 2 and Ceramide Transport as Key Targets of the Natural Flavonoid Luteolin to Induce Apoptosis in Colon Cancer Cells. PLoS ONE 2015, 10, e0143384. [Google Scholar] [CrossRef] [PubMed]
- Goins, L.; Spassieva, S. Sphingoid bases and their involvement in neurodegenerative diseases. Adv. Biol. Regul. 2018, 70, 65–73. [Google Scholar] [CrossRef]
- Feinstein, M.; Flusser, H.; Tally, L.S.; Ben-Zeev, B.; Lev, D.; Agamy, O.; Cohen, I.; Kadir, R.; Sivan, S.; Leshinsky-Silver, E.; et al. VPS53 mutations cause progressive cerebello-cerebral atrophy type 2 (PCCA2). J. Med. Genet. 2014, 51, 303–308. [Google Scholar] [CrossRef] [PubMed]
- Park, J.S.; Halegoua, S.; Kishida, S.; Neiman, A.M. A conserved function in phosphatidylinositol metabolism for mammalian Vps13 family proteins. PLoS ONE 2015, 10, e0124836. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Soczewka, P.; Flis, K.; Tribouillard-Tanvier, D.; di Rago, J.-P.; Santos, C.N.; Menezes, R.; Kaminska, J.; Zoladek, T. Flavonoids as Potential Drugs for VPS13-Dependent Rare Neurodegenerative Diseases. Genes 2020, 11, 828. https://doi.org/10.3390/genes11070828
Soczewka P, Flis K, Tribouillard-Tanvier D, di Rago J-P, Santos CN, Menezes R, Kaminska J, Zoladek T. Flavonoids as Potential Drugs for VPS13-Dependent Rare Neurodegenerative Diseases. Genes. 2020; 11(7):828. https://doi.org/10.3390/genes11070828
Chicago/Turabian StyleSoczewka, Piotr, Krzysztof Flis, Déborah Tribouillard-Tanvier, Jean-Paul di Rago, Cláudia N. Santos, Regina Menezes, Joanna Kaminska, and Teresa Zoladek. 2020. "Flavonoids as Potential Drugs for VPS13-Dependent Rare Neurodegenerative Diseases" Genes 11, no. 7: 828. https://doi.org/10.3390/genes11070828
APA StyleSoczewka, P., Flis, K., Tribouillard-Tanvier, D., di Rago, J.-P., Santos, C. N., Menezes, R., Kaminska, J., & Zoladek, T. (2020). Flavonoids as Potential Drugs for VPS13-Dependent Rare Neurodegenerative Diseases. Genes, 11(7), 828. https://doi.org/10.3390/genes11070828





