Molecular Cytogenetic Characterization of the Sicilian Endemic Pond Turtle Emys trinacris and the Yellow-Bellied Slider Trachemys scripta scripta (Testudines, Emydidae)
Abstract
1. Introduction
2. Materials and Methods
2.1. Studied Material
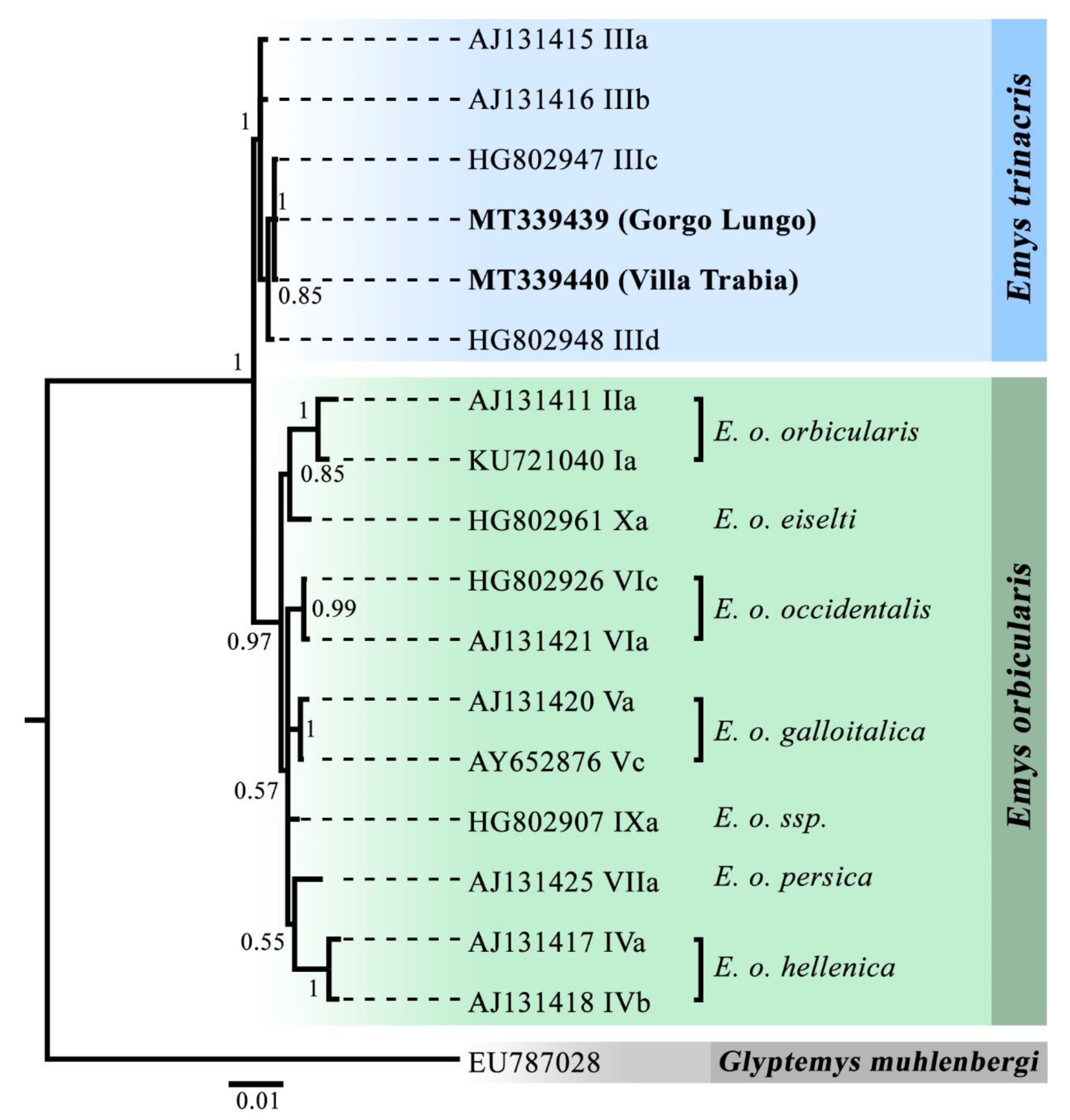
2.2. Species Identification
2.3. Cytogenetic Examination
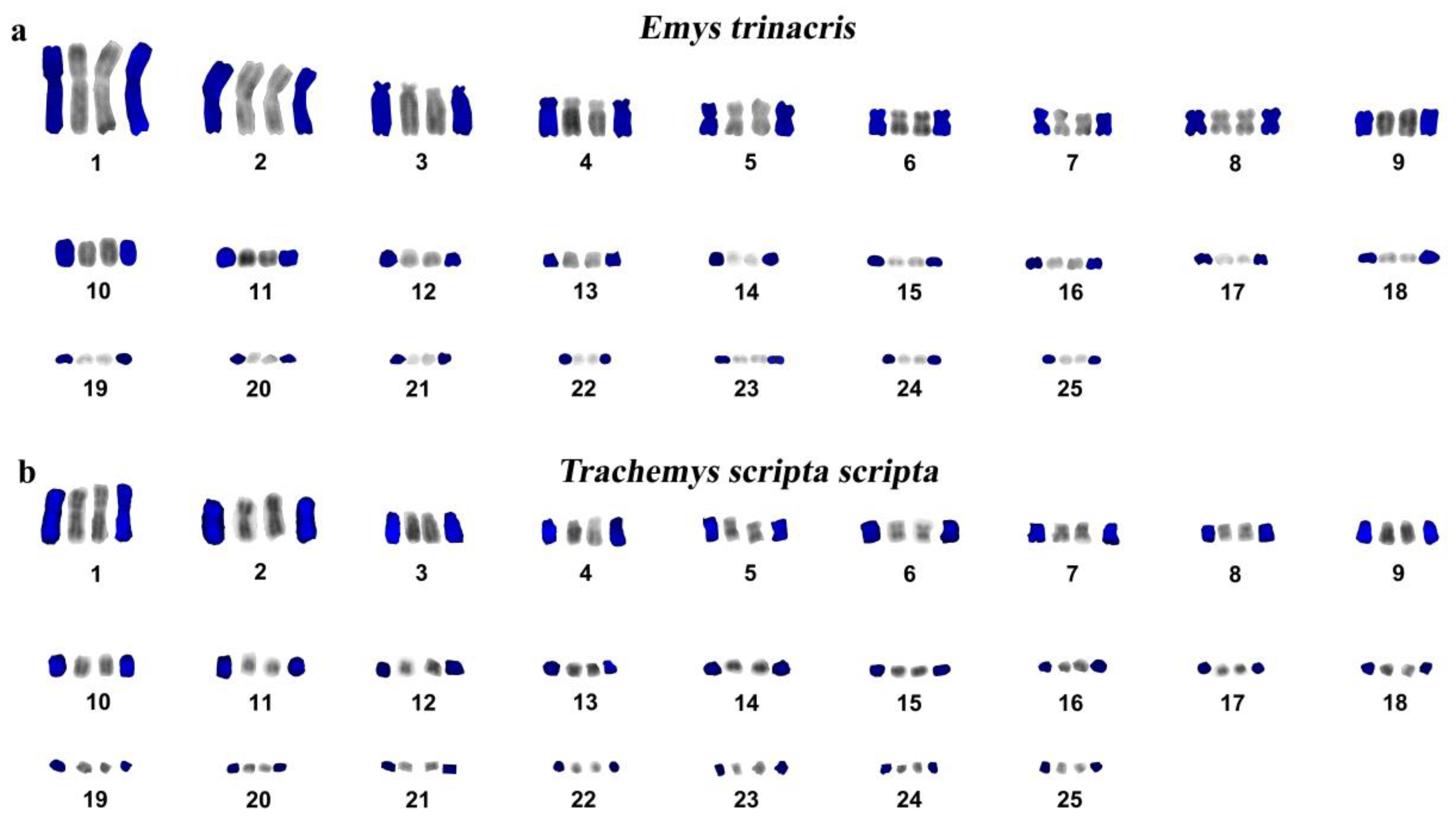
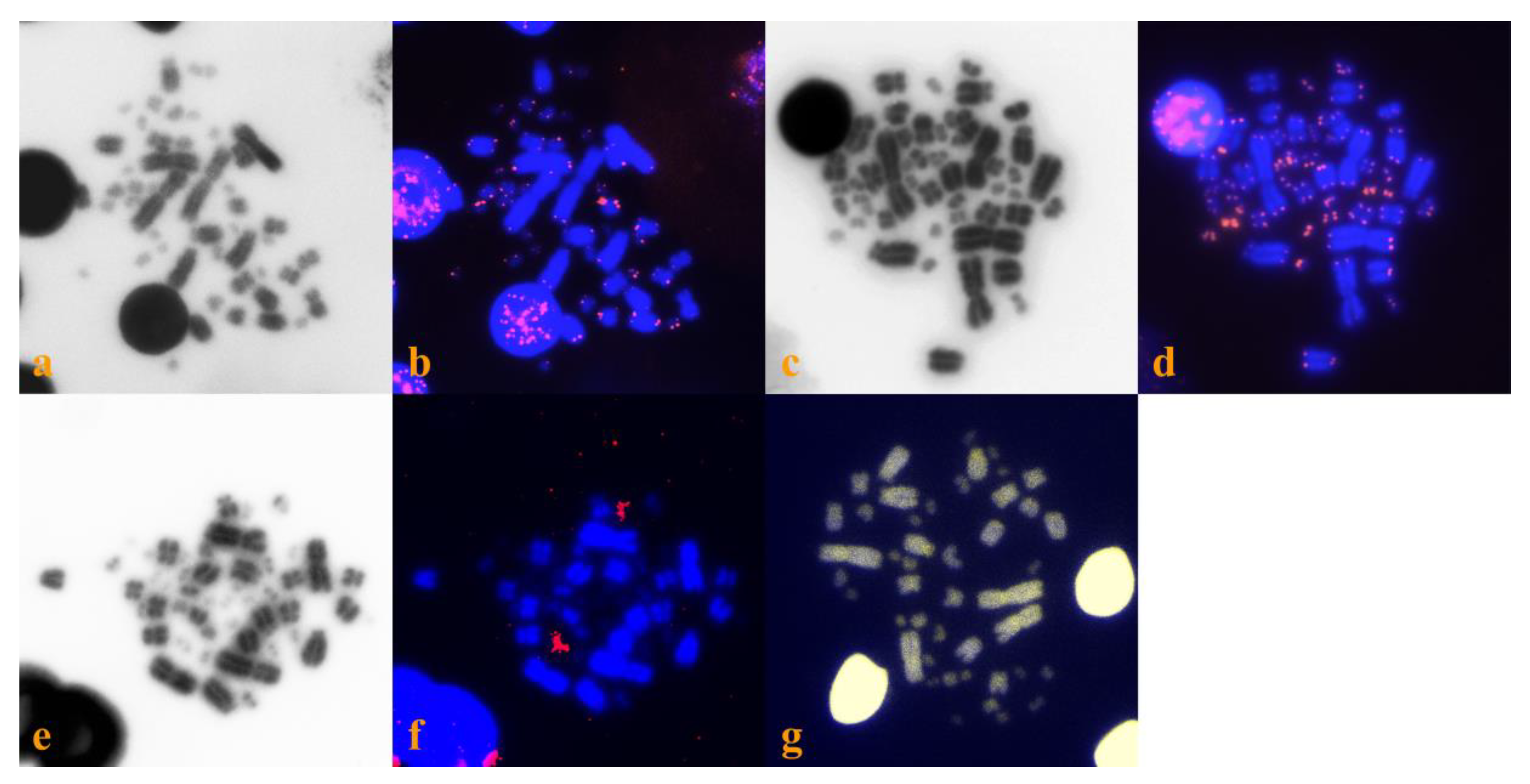
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Uetz, P.; Freed, P.; Hošek, J. The Reptile Database. Available online: http://www.reptile-database.org (accessed on 28 May 2020).
- Parham, J.F.; Papenfuss, T.J.; van Dijk, P.P.; Wilson, B.S.; Marte, C.; Rodriguez-Schettino, L.; Simison, W.B. Genetic introgression and hybridization in Antillean freshwater turtles (Trachemys) revealed by coalescent analyses of mitochondrial and cloned nuclear markers. Mol. Phylogenet. Evol. 2013, 67, 176–187. [Google Scholar] [CrossRef] [PubMed]
- Parham, J.F.; Papenfuss, T.J.; Buskirk, J.R.; Parra-Olea, G.; Chen, J.Y.; Simison, W.B. Trachemys ornata or not ornata: Reassessment of a taxonomic revision for Mexican Trachemys. Proc. Calif. Acad. Sci. 2015, 62, 359–367. [Google Scholar]
- Rogner, M. European Pond Turtle: Emys Orbicularis; Chelonian Library, Edition Chimaira: Frankfurt, Germany, 2009. [Google Scholar]
- Gaffney, E.S.; Meylan, P.A. A Phylogeny of Turtles in: The Phylogeny and Classification of Tetrapods; Benton, M.J., Ed.; Clarendon: Oxford, UK, 1988; pp. 157–219. [Google Scholar]
- Marrone, F.; Sacco, F.; Arizza, V.; Arculeo, M. Amendment of the type locality of the endemic Sicilian pond turtle Emys trinacris, Fritz et al. 2005, with some notes on the highest altitude reached by the species (Testudines, Emydidae). Acta Herpetol. 2016, 11, 63–65. [Google Scholar]
- Vamberger, M.; Fritz, U. Big data can cause big mistakes: Using the Societas Europaea Herpetologica atlas by Sillero et al. (2014), the distribution of Emys orbicularis will be misunderstood. Biologia 2018, 73, 281–283. [Google Scholar] [CrossRef]
- Feldman, C.R.; Parham, J.F. Molecular phylogenetics of emydine turtles: Taxonomic revision and the evolution of shell kinesis. Mol. Phylogenet. Evol. 2002, 22, 388–398. [Google Scholar] [CrossRef]
- Fritz, U.; Schmidt, C.; Ernst, C.H. Competing generic concepts for Blanding’s, Pacific and European pond turtles (Emydoidea, Actinemys and Emys)—Which is best? Zootaxa 2011, 2791, 41–53. [Google Scholar] [CrossRef]
- Seidel, M.E.; Ernst, C.H. A systematic review of the turtle family Emydidae. Vertebr. Zool. 2017, 67, 1–122. [Google Scholar]
- D’Angelo, S.; Pennisi, M.G.; Lo Valvo, M.; Fritz, U. Variation of Sicilian pond turtles, Emys trinacris—What makes a species cryptic? Amphib-Reptil. 2006, 27, 513–529. [Google Scholar] [CrossRef]
- Arizza, V.; Russo, D.; Marrone, F.; Sacco, F.; Arculeo, M. Morphological characterization of the blood cells in the endangered endemic pond turtle, Emys trinacris (Testudines: Emydidae). Ital. J. Zool. 2014, 81, 344–353. [Google Scholar] [CrossRef]
- Fritz, U.; Fattizzo, T.; Guicking, D.; Tripepi, S.; Pennisi, M.G.; Lenk, P.; Joger, U.; Wink, M. A new cryptic species of pond turtle from southern Italy, the hottest spot in the range of the genus Emys. Zool. Scr. 2005, 34, 351–371. [Google Scholar] [CrossRef]
- Stuckas, H.; Velo-Antón, G.; Fahd, S.; Kalboussi, M.; Rouag, R.; Arculeo, M.; Marrone, F.; Sacco, F.; Vamberger, M.; Fritz, U. Where are you from, stranger? The enigmatic biogeography of North African pond turtles (Emys orbicularis). Org. Divers. Evol. 2014, 14, 295–306. [Google Scholar] [CrossRef]
- Vamberger, M.; Stuckas, H.; Sacco, F.; D’Angelo, S.; Arculeo, M.; Cheylan, M.; Corti, C.; Lo Valvo, M.; Marrone, F.; Wink, M.; et al. Differences in gene flow in a twofold secondary contact zone of pond turtles in southern Italy (Testudines: Emydidae: Emys orbicularis galloitalica, E. o. hellenica, E. trinacris). Zool. Scr. 2015, 44, 233–249. [Google Scholar] [CrossRef]
- Marrone, F.; Sacco, F.; Kehlmaier, C.; Arizza, V.; Arculeo, M. Some like it cold: The glossiphoniid parasites of the Sicilian endemic pond turtle Emys trinacris (Testudines, Emydidae), an example of ‘parasite inertia’? J. Zool. Syst. Evol. Res. 2016, 54, 60–66. [Google Scholar] [CrossRef]
- Olmo, E.; Signorino, G.G. Chromorep: A Reptile Chromosomes Database. Available online: http://chromorep.univpm.it (accessed on 14 April 2020).
- Bickham, J.W.; Baker, R.J. Chromosome homology and evolution of emydid turtles. Chromosoma 1976, 54, 201–219. [Google Scholar] [CrossRef] [PubMed]
- Bickham, J.W.; Carr, J.L. Taxonomy and phylogeny of the higher categories of Cryptodiran turtles based on a cladistics analysis of chromosomal data. Copeia 1983, 4, 918–932. [Google Scholar] [CrossRef]
- Cleiton, F.; Giuliano-Caetano, L. Cytogenetic characterization of two turtle species: Trachemys dorbigni and Trachemys scripta elegans. Caryologia 2008, 61, 253–257. [Google Scholar] [CrossRef]
- Iannucci, A.; Svartman, M.; Bellavita, M.; Chelazzi, G.; Stanyon, R.; Ciofi, C. Insights into emydid turtle cytogenetics: The european pond turtle as a model species. Cytogenet. Genome Res. 2019, 157, 166–171. [Google Scholar] [CrossRef]
- Montiel, E.E.; Badenhorst, D.; Lee, L.S.; Literman, R.; Trifonov, V.; Valenzuela, N. Cytogenetic insights into the evolution of chromosomes and sex determination reveal striking homology of turtle sex chromosomes to amphibian autosomes. Cytogenet. Genome Res. 2016, 148, 292–304. [Google Scholar] [CrossRef]
- Montiel, E.E.; Badenhorst, D.; Tamplin, J.; Burke, R.; Valenzuela, N. Discovery of youngest sex chromosomes reveals first case of convergent co-option of ancestral autosomes in turtles. Chromosoma 2017, 126, 105–113. [Google Scholar] [CrossRef]
- Valenzuela, N.; Badenhorst, D.; Montiel, E.E.; Literman, R. Molecular cytogenetic search for cryptic sex chromosomes in painted turtles Chrysemys picta. Cytogenet. Genome Res. 2014, 144, 39–46. [Google Scholar] [CrossRef]
- Martinez, P.A.; Boeris, J.M.; Sánchez, J.; Pastori, M.C.; Bolzán, A.D.; Ledesma, M.A. Karyotypic characterization of Trachemys dorbigni (Testudines: Emydidae) and Chelonoidis (Geochelone) donosobarrosi (Testudines: Testudinidae), two species of cryptodiran turtles from Argentina. Genetica 2009, 137, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Kasai, F.; O’Brien, P.C.M.; Martin, S.; Ferguson-Smith, M.A. Extensive homology of chicken macrochromosomes in the karyotypes of Trachemys scripta elegans and Crocodylus niloticus revealed by chromosome painting despite long divergence times. Cytogenet. Genome Res. 2012, 136, 303–307. [Google Scholar] [CrossRef] [PubMed]
- Badenhorst, D.; Hillier, L.W.; Literman, R.; Montiel, E.E.; Radhakrishnan, S.; Shen, Y.; Minx, P.; Janes, D.E.; Warren, W.C.; Edwards, S.V.; et al. Physical mapping and refinement of the painted turtle genome (Chrysemys picta) inform amniote genome evolution and challenge turtle-bird chromosomal conservation. Genome Biol. Evol. 2015, 7, 2038–2050. [Google Scholar] [CrossRef][Green Version]
- Picone, B.; Dumas, F.; Stanyon, R.; Lannino, A.; Bigoni, F.; Privitera, O.; Sineo, L. Exploring evolution in 298 Ceboidea (Platyrrhini, Primates) by Williams-Beuren Probe (HSA 7q11.23) chromosome mapping. Folia Primatol. 2008, 79, 417–427. [Google Scholar] [CrossRef] [PubMed]
- Dumas, F.; Houck, M.; Bigoni, F.; Perelman, P.; Romanenko, S.; Stanyon, R. Chromosome painting of the.pygmy tree shrew shows that no derived cytogenetic traits link primates and scandentia. Cytogenet. Genome Res. 2012, 136, 175–179. [Google Scholar] [CrossRef] [PubMed]
- Dumas, F.; Stanyon, R.; Sineo, L.; Stone, G.; Bigoni, F. Phylogenomics of species from four genera of New World monkeys by flow sorting and reciprocal chromosome painting. BMC Evol. Boil. 2007, 7, S11. [Google Scholar] [CrossRef]
- Dumas, F.; Sineo, L. Chromosomal dynamics in platyrrhinae by mapping BACs probes. J. Biol. Res 2012, LXXXV, 299–301. [Google Scholar]
- Dumas, F.; Sineo, L. Chromosomal dynamics in Cercopithecini studied by Williams-Beuren probe mapping. Caryologia 2010, 63, 435–442. [Google Scholar]
- Dumas, F.; Sineo, L. The evolution of human synteny 4 by mapping sub-chromosomal specific probes in Primates. Caryologia 2014, 67, 281–291. [Google Scholar] [CrossRef]
- Scardino, R.; Milioto, V.; Proskuryakova, A.A.; Serdyukova, N.A.; Perelman, P.L.; Dumas, F. Evolution of the human chromosome 13 synteny: Evolutionary rearrangements, plasticity, human disease genes and cancer breakpoints. Genes 2020, 11, 383. [Google Scholar] [CrossRef] [PubMed]
- Dumas, F.; Sineo, L.; Ishida, T. Taxonomic identification of Aotus (Platyrrhinae) through cytogenetics | Identificazione tassonomica di Aotus (Platyrrhinae) mediante la citogenetica. J. Biol. Res. 2015, 88, 65–66. [Google Scholar]
- Dumas, F.; Mazzoleni, S. Neotropical primate evolution and phylogenetic reconstruction using chromosomal data. Eur. Zoöl. J. 2017, 84, 1–18. [Google Scholar] [CrossRef][Green Version]
- Rovatsos, M.; Kratochvíl, L.; Altmanová, M.; Johnson Pokorná, M. Interstitial telomeric motifs in squamate reptiles: When the exceptions outnumber the rule. PLoS ONE 2015, 10, e0134985. [Google Scholar] [CrossRef] [PubMed]
- Matsubara, K.; Uno, Y.; Srikulnath, K.; Matsuda, Y.; Miller, E.; Olsson, M. No interstitial telomeres on autosomes but remarkable amplification of telomeric repeats on the W sex chromosome in the sand lizard (Lacerta agilis). J. Hered. 2015, 106, 753–757. [Google Scholar] [CrossRef] [PubMed]
- Dumas, F.; Cuttaia, H.; Sineo, L. Chromosomal distribution of interstitial telomeric sequences in nine neotropical primates (Platyrrhini): Possible implications in evolution and phylogeny. J. Zoöl. Syst. Evol. Res. 2016, 54, 226–236. [Google Scholar] [CrossRef]
- Mazzoleni, S.; Schillaci, O.; Sineo, L.; Dumas, F. Distribution of interstitial telomeric sequences in primates and the pygmy tree shrew (Scandentia). Cytogenet. Genome Res. 2017, 151, 141–150. [Google Scholar] [CrossRef]
- Mazzoleni, S.; Rovatsos, M.; Schillaci, O.; Dumas, F. Evolutionary insight on localization of 18S, 28S rDNA genes on homologous chromosomes in Primates genomes. Comp. Cytogenet. 2018, 12, 27–40. [Google Scholar] [CrossRef]
- Milioto, V.; Vlah, S.; Mazzoleni, S.; Rovatsos, M.; Dumas, F. Chromosomal localization of 18S-28S rDNA and (TTAGGG)n sequences in two south african dormice of the genus Graphiurus (Rodentia: Gliridae). Cytogenet. Genome Res. 2019, 158, 145–151. [Google Scholar] [CrossRef]
- Rovatsos, M.; Altmanová, M.; Augstenová, B.; Mazzoleni, S.; Velenský, P.; Kratochvíl, L. ZZ/ZW sex determination with multiple neo-sex chromosomes is common in madagascan chameleons of the genus Furcifer (Reptilia: Chamaeleonidae). Genes 2019, 10, 1020. [Google Scholar] [CrossRef]
- Mazzoleni, S.; Augstenová, B.; Clemente, L.; Auer, M.; Fritz, U.; Praschag, P.; Protiva, T.; Velenský, P.; Kratochvíl, L.; Rovatsos, M. Sex is determined by XX/XY sex chromosomes in Australasian side-necked turtles (Testudines: Chelidae). Sci. Rep. 2020, 10, 1–11. [Google Scholar] [CrossRef]
- Meyne, J.; Ratliff, R.L.; Moyzis, R.K. Conservation of the human telomere sequence (TTAGGG)n among vertebrates. Proc. Natl. Acad. Sci. USA 1989, 86, 7049–7053. [Google Scholar] [CrossRef] [PubMed]
- Meyne, J.; Baker, R.J.; Hobart, H.H.; Hsu, T.C.; Ryder, O.A.; Ward, O.G.; Wiley, J.E.; Wurster-Hill, D.H.; Yates, T.L.; Moyzis, R.K. Distribution of non-telomeric sites of the (TTAGGG)n telomeric sequence in vertebrate chromosomes. Chromosoma 1990, 99, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Herrera, A.; Nergadze, S.G.; Santagostino, M.; Giulotto, E. Telomeric repeats far from the ends: Mechanisms of origin and role in evolution. Cytogenet. Genome Res 2008, 122, 219–228. [Google Scholar] [CrossRef] [PubMed]
- Ficetola, G.F.; Scali, S. Invasive amphibians and reptiles in Italy. In Atti VIII Congresso Nazionale Societas Herpetologica Italica; Ianieri Edizioni: Pescara, Italy, 2010; pp. 335–340. [Google Scholar]
- Redrobe, S.; MacDonald, J. Sample collection and clinical pathology of reptiles. Vet. Clin. N. Am. Exot. Anim. Pract. 1999, 2, 709–730. [Google Scholar] [CrossRef]
- Naselli-Flores, L.; Marrone, F. Different invasibility of permanent and temporary waterbodies in a semiarid Mediterranean Island. Inland Waters 2019, 9, 41–421. [Google Scholar] [CrossRef]
- Mazzoleni, S.; Augstenová, B.; Clemente, L.; Auer, M.; Fritz, U.; Praschag, P.; Protiva, T.; Velenský, P.; Kratochvíl, L.; Rovatsos, M. Turtles of the genera Geoemyda and Pangshura (Testudines: Geoemydidae) lack differentiated sex chromosomes: The end of a 40-year error cascade for Pangshura. PeerJ 2019, 7, e6241. [Google Scholar] [CrossRef]
- Belaiba, E.; Marrone, F.; Vecchioni, L.; Bahri-Sfar, L.; Arculeo, M. An exhaustive phylogeny of the combtooth blenny genus Salaria (Pisces, Blenniidae) shows introgressive hybridization and lack of reciprocal mtDNA monophyly between the marine species Salaria basilisca and Salaria pavo. Mol. Phylogenet. Evol. 2019, 135, 210–221. [Google Scholar] [CrossRef]
- Kasai, F.; O’Brien, P.C.M.; Ferguson-Smith, M.A. Reassessment of genome size in turtle and crocodile based on chromosome measurement by flow karyotyping: Close similarity to chicken. Biol Lett. 2012, 8, 631–635. [Google Scholar] [CrossRef]
- Olmo, E. Trends in the evolution of reptilian chromosomes. Integr. Comp. Biol. 2008, 48, 486–493. [Google Scholar] [CrossRef]
- Bista, B.; Valenzuela, N. Turtle insights into the evolution of the reptilian karyotype and the genomic architecture of sex determination. Genes 2020, 11, 416. [Google Scholar] [CrossRef]
- Vamberger, M.; Ihlow, F.; Asztalos, M.; Dawson, J.E.; Jasinski, S.E.; Praschag, P.; Fritz, U. So different, yet so alike: North American slider turtles (Trachemys scripta). Vertebr. Zool. 2020, 70, 87–96. [Google Scholar]
| Latin Name | Code | Samples | Specimens |
|---|---|---|---|
| Emys trinacris | ETR | blood | male collected in a natural pond (Gorgo Lungo, WGS84 geographical coordinates: 37.901131 N, 13.408438 E; altitude: 890 m a.s.l.) |
| blood | male collected in an ornamental basin of a public garden within the town of Palermo (Villa Trabia, WGS84 geographical coordinates: 38.129757 N, 13.347749 E; altitude: 20 m a.s.l.). | ||
| Trachemys scripta scripta | TSS | blood | female collected at the Botanical Garden of the University of Palermo (Italy) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Scardino, R.; Mazzoleni, S.; Rovatsos, M.; Vecchioni, L.; Dumas, F. Molecular Cytogenetic Characterization of the Sicilian Endemic Pond Turtle Emys trinacris and the Yellow-Bellied Slider Trachemys scripta scripta (Testudines, Emydidae). Genes 2020, 11, 702. https://doi.org/10.3390/genes11060702
Scardino R, Mazzoleni S, Rovatsos M, Vecchioni L, Dumas F. Molecular Cytogenetic Characterization of the Sicilian Endemic Pond Turtle Emys trinacris and the Yellow-Bellied Slider Trachemys scripta scripta (Testudines, Emydidae). Genes. 2020; 11(6):702. https://doi.org/10.3390/genes11060702
Chicago/Turabian StyleScardino, Rita, Sofia Mazzoleni, Michail Rovatsos, Luca Vecchioni, and Francesca Dumas. 2020. "Molecular Cytogenetic Characterization of the Sicilian Endemic Pond Turtle Emys trinacris and the Yellow-Bellied Slider Trachemys scripta scripta (Testudines, Emydidae)" Genes 11, no. 6: 702. https://doi.org/10.3390/genes11060702
APA StyleScardino, R., Mazzoleni, S., Rovatsos, M., Vecchioni, L., & Dumas, F. (2020). Molecular Cytogenetic Characterization of the Sicilian Endemic Pond Turtle Emys trinacris and the Yellow-Bellied Slider Trachemys scripta scripta (Testudines, Emydidae). Genes, 11(6), 702. https://doi.org/10.3390/genes11060702






