Hybridization Facilitates Adaptive Evolution in Two Major Fungal Pathogens
Abstract
1. Introduction
2. Hybridization in Cryptococcus Species Complex
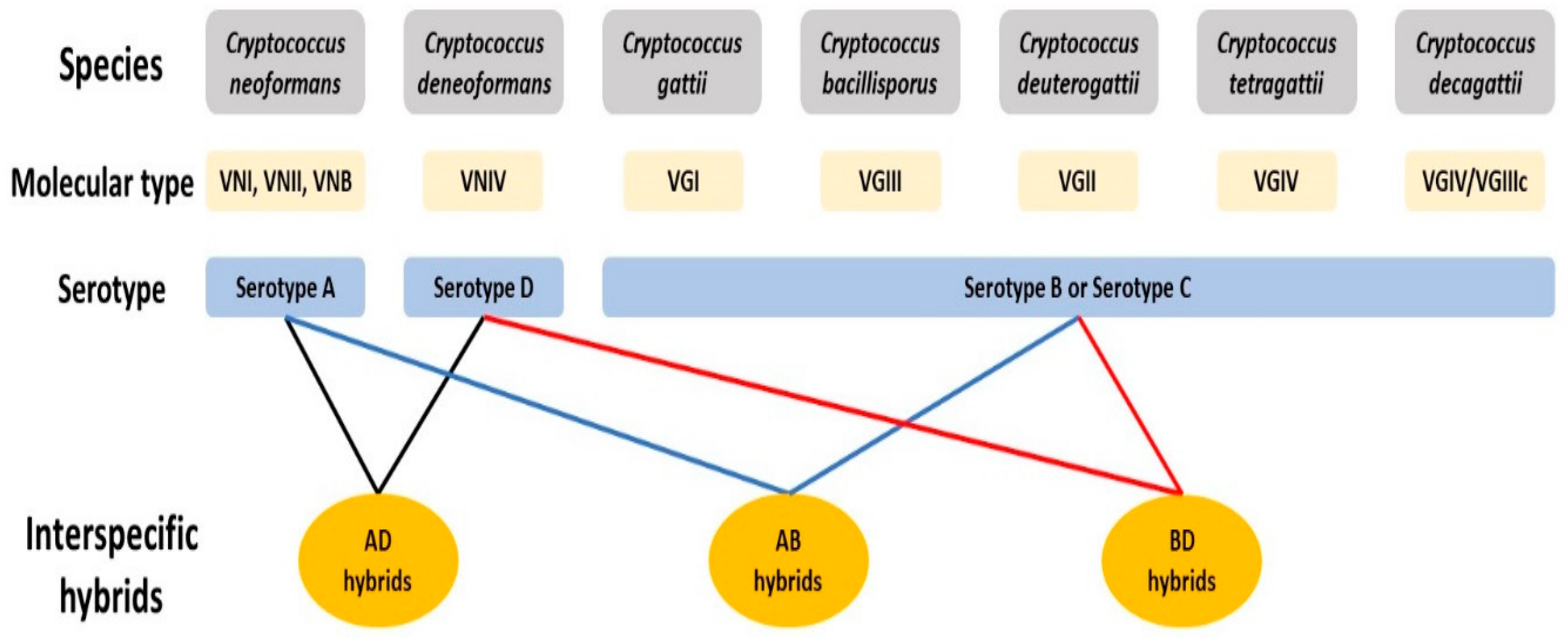
2.1. Cryptococcus Species Complex
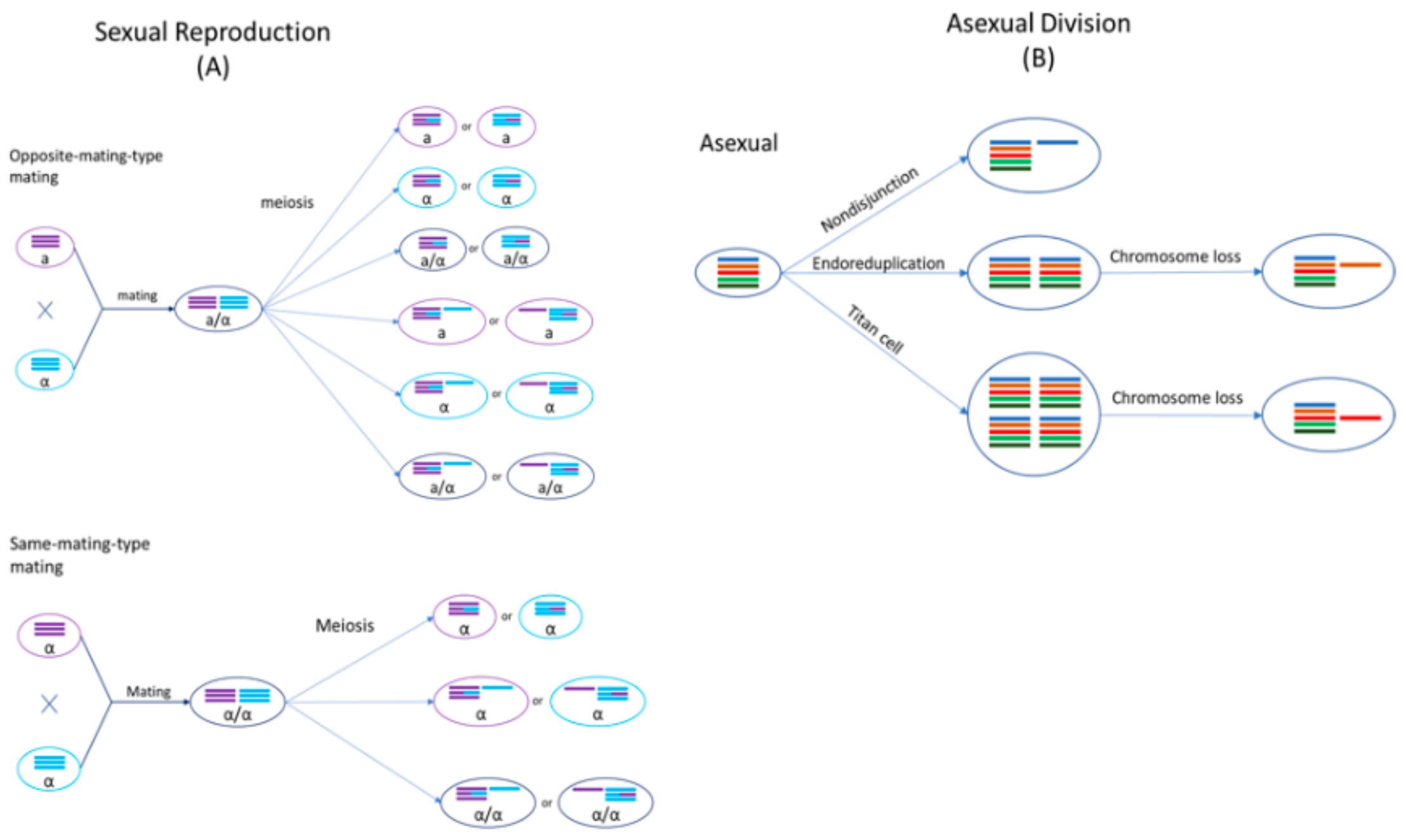
2.2. Sexual Cycle of Cryptococcus
2.3. Hybrids in the Cryptococcus Species Complex
3. Outcomes of Hybridization between Cryptococcal Lineages
3.1. Hybrid Inviability
3.2. Hybrid Sterility
3.3. Phenotypic Diversity and Hybrid Vigor
4. Genetics of Cryptococcal Hybrids
4.1. Aneuploidy in Cryptococcal Hybrids
4.2. Loss of Heterozygosity
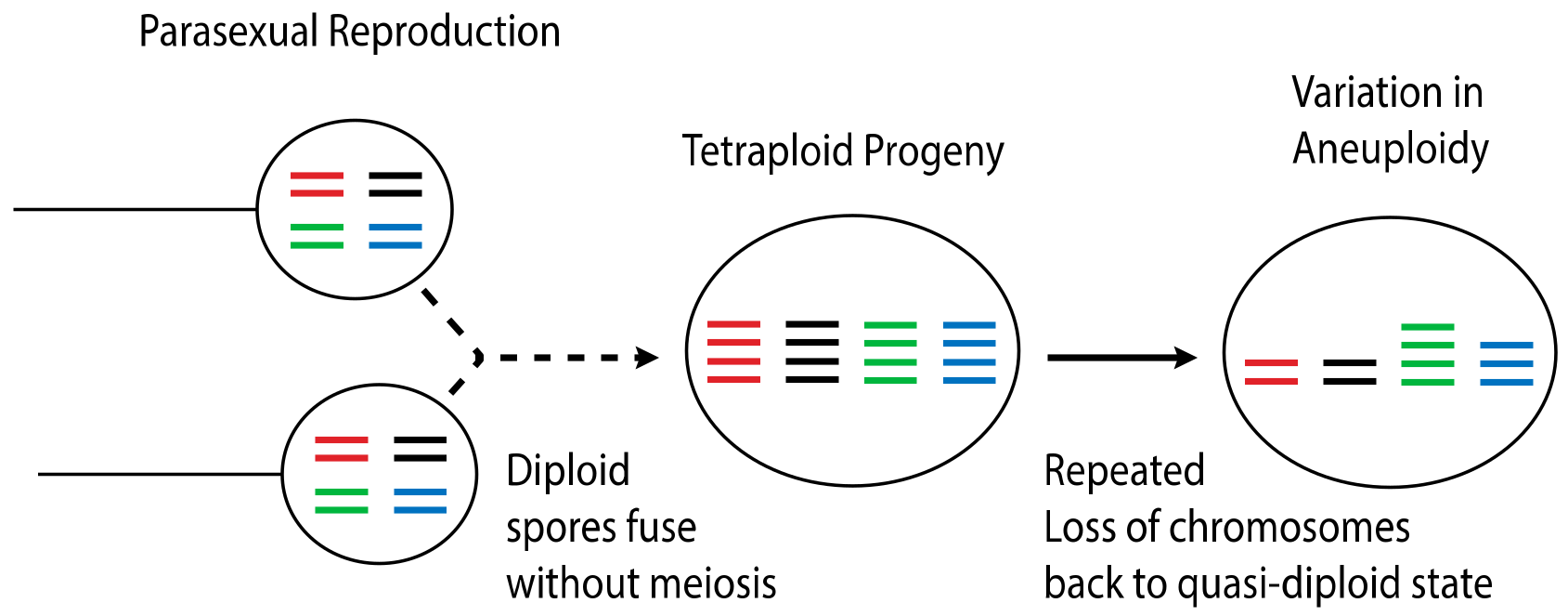
4.3. Dynamic Ploidy Changes in Cryptococcus
4.4. Cryptococcus as a Model System for Fungal Hybridization
5. Hybridization in an Aquatic Chytrid Fungus Associated with Amphibian Declines
5.1. The Amphibian Chytrid Batrachochytrium Dendrobatidis
5.2. Hybrids in Batrachochytrium Dendrobatidis
5.3. Outcomes of Hybridization in Batrachochytrium Dendrobatidis
5.4. Aneuploidy in Batrachochytrium Dendrobatidis
5.5. LOH in Batrachochytrium Dendrobatidis
6. Conclusions and Perspectives
Author Contributions
Conflicts of Interest
References
- Barton, N.H. The role of hybridization in evolution. Mol. Ecol. 2001, 10, 551–568. [Google Scholar] [CrossRef]
- Barton, N.H.; Hewitt, G.M. Analysis of hybrid zones. Annu. Rev. Ecol. Syst. 1985, 16, 113–148. [Google Scholar] [CrossRef]
- Rieseberg, L.H. Hybrid origins of plant species. Annu. Rev. Ecol. Syst. 1997, 28, 359–389. [Google Scholar] [CrossRef]
- Arnold, M.L.; Hodges, S.A. Are natural hybrids fit or unfit relative to their parents? Trends Ecol. Evol. (Amsterdam) 1995, 10, 67–71. [Google Scholar] [CrossRef]
- Mallet, J. Hybrid speciation. Nature 2007, 446, 279–283. [Google Scholar] [CrossRef]
- Ramsey, J.; Schemske, D.W. Neopolyploidy in Flowering Plants. Annu. Rev. Ecol. Syst. 2002, 33, 589–639. [Google Scholar] [CrossRef]
- Harrison, R.G.; Larson, E.L. Hybridization, introgression, and the nature of species boundaries. J. Hered. 2014, 105 (Suppl. 1), 795–809. [Google Scholar] [CrossRef]
- Dunn, B.; Richter, C.; Kvitek, D.J.; Pugh, T.; Sherlock, G. Analysis of the Saccharomyces cerevisiae pan-genome reveals a pool of copy number variants distributed in diverse yeast strains from differing industrial environments. Genome Res. 2012, 22, 908–924. [Google Scholar] [CrossRef]
- Schardl, C.L.; Craven, K.D. Interspecific hybridization in plant-associated fungi and oomycetes: A review. Mol. Ecol. 2003, 12, 2861–2873. [Google Scholar] [CrossRef]
- Stukenbrock, E.H. The role of hybridization in the evolution and emergence of new fungal plant pathogens. Phytopathology 2016, 106, 104–112. [Google Scholar] [CrossRef]
- Nieuwenhuis, B.P.S.; James, T.Y. The frequency of sex in fungi. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2016, 371, 20150540. [Google Scholar] [CrossRef] [PubMed]
- Haag, C.R.; Roze, D. Genetic load in sexual and asexual diploids: Segregation, dominance and genetic drift. Genetics 2007, 176, 1663–1678. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rajasingham, R.; Smith, R.M.; Park, B.J.; Jarvis, J.N.; Govender, N.P.; Chiller, T.M.; Denning, D.W.; Loyse, A.; Boulware, D.R. Global burden of disease of HIV-associated cryptococcal meningitis: An updated analysis. Lancet Infect. Dis. 2017, 17, 873–881. [Google Scholar] [CrossRef]
- Cogliati, M. Global Molecular Epidemiology of Cryptococcus neoformans and Cryptococcus gattii: An Atlas of the Molecular Types. Scientifica 2013, 2013, 1–23. [Google Scholar] [CrossRef]
- Hagen, F.; Khayhan, K.; Theelen, B.; Kolecka, A.; Polacheck, I.; Sionov, E.; Falk, R.; Parnmen, S.; Lumbsch, H.T.; Boekhout, T. Recognition of seven species in the Cryptococcus gattii/Cryptococcus neoformans species complex. Fungal Genet. Biol. 2015, 78, 16–48. [Google Scholar] [CrossRef]
- Hagen, F.; Lumbsch, H.T.; Arsic Arsenijevic, V.; Badali, H.; Bertout, S.; Billmyre, R.B.; Bragulat, M.R.; Cabañes, F.J.; Carbia, M.; Chakrabarti, A.; et al. Importance of resolving fungal nomenclature: The case of multiple pathogenic species in the Cryptococcus genus. mSphere 2017, 2, e00238-17. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Bennett, J.E.; Wickes, B.L.; Meyer, W.; Cuomo, C.A.; Wollenburg, K.R.; Bicanic, T.A.; Castañeda, E.; Chang, Y.C.; Chen, J.; et al. The case for adopting the “species complex” nomenclature for the etiologic agents of cryptococcosis. mSphere 2017, 2, e00357-16. [Google Scholar] [CrossRef]
- Dromer, F.; Gueho, E.; Ronin, O.; Dupont, B. Serotyping of Cryptococcus neoformans by using a monoclonal antibody specific for capsular polysaccharide. J. Clin. Microbiol. 1993, 31, 359–363. [Google Scholar] [CrossRef]
- Casadevall, A.; Freij, J.B.; Hann-Soden, C.; Taylor, J. Continental drift and speciation of the Cryptococcus neoformans and Cryptococcus gattii species complexes. mSphere 2017, 2, e00103-17. [Google Scholar] [CrossRef]
- Xu, J.; Vilgalys, R.; Mitchell, T.G. Multiple gene genealogies reveal recent dispersion and hybridization in the human pathogenic fungus Cryptococcus neoformans. Mol. Ecol. 2000, 9, 1471–1481. [Google Scholar] [CrossRef]
- Ngamskulrungroj, P.; Gilgado, F.; Faganello, J.; Litvintseva, A.P.; Leal, A.L.; Tsui, K.M.; Mitchell, T.G.; Vainstein, M.H.; Meyer, W. Genetic diversity of the Cryptococcus species complex suggests that Cryptococcus gattii deserves to have varieties. PLoS ONE 2009, 4, e5862. [Google Scholar] [CrossRef]
- Sharpton, T.J.; Neafsey, D.E.; Galagan, J.E.; Taylor, J.W. Mechanisms of intron gain and loss in Cryptococcus. Genome Biol. 2008, 9, R24. [Google Scholar] [CrossRef] [PubMed]
- Kwon-Chung, K.J. A New Genus, Filobasidiella, the Perfect State of Cryptococcus neoformans. Mycologia 1975, 67, 1197–1200. [Google Scholar] [CrossRef] [PubMed]
- Kwon-Chung, K.J. A new species of Filobasidiella, the sexual state of Cryptococcus neoformans B and C serotypes. Mycologia 1976, 68, 943–946. [Google Scholar] [CrossRef]
- Lin, X.; Heitman, J. Chlamydospore formation during hyphal growth in Cryptococcus neoformans. Eukaryot. Cell 2005, 4, 1746–1754. [Google Scholar] [CrossRef]
- Lin, X. Cryptococcus neoformans: Morphogenesis, infection, and evolution. Infect. Genet. Evol. 2009, 9, 401–416. [Google Scholar] [CrossRef]
- Lin, X.; Hull, C.M.; Heitman, J. Sexual reproduction between partners of the same mating type in Cryptococcus neoformans. Nature 2005, 434, 1017–1021. [Google Scholar] [CrossRef]
- Zhao, Y.; Lin, J.; Fan, Y.; Lin, X. Life Cycle of Cryptococcus neoformans. Annu. Rev. Microbiol. 2019, 73, 17–42. [Google Scholar] [CrossRef]
- Hull, C.M.; Heitman, J. Genetics of Cryptococcus neoformans. Annu. Rev. Genet. 2002, 36, 557–615. [Google Scholar] [CrossRef]
- Wang, P.; Perfect, J.R.; Heitman, J. The G-protein beta subunit GPB1 is required for mating and haploid fruiting in Cryptococcus neoformans. Mol. Cell. Biol. 2000, 20, 352–362. [Google Scholar] [CrossRef] [PubMed]
- Wickes, B.L.; Mayorga, M.E.; Edman, U.; Edman, J.C. Dimorphism and haploid fruiting in Cryptococcus neoformans: Association with the alpha-mating type. Proc. Natl. Acad. Sci. USA 1996, 93, 7327–7331. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Huang, J.C.; Mitchell, T.G.; Heitman, J. Virulence attributes and hyphal growth of C. neoformans are quantitative traits and the MATα allele enhances filamentation. PLoS Genet. 2006, 2, e187. [Google Scholar] [CrossRef] [PubMed]
- Kwon-Chung, K.J.; Bennett, J.E. Distribution of alpha and alpha mating types of Cryptococcus neoformans among natural and clinical isolates. Am. J. Epidemiol. 1978, 108, 337–340. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Li, X.; Xu, J. Geographic distribution of mating type alleles of Cryptococcus neoformans in four areas of the United States. J. Clin. Microbiol. 2002, 40, 965–972. [Google Scholar] [CrossRef]
- Fu, C.; Heitman, J. PRM1 and KAR5 function in cell-cell fusion and karyogamy to drive distinct bisexual and unisexual cycles in the Cryptococcus pathogenic species complex. PLoS Genet. 2017, 13, e1007113. [Google Scholar] [CrossRef]
- Lin, X.; Heitman, J. The biology of the Cryptococcus neoformans species complex. Annu. Rev. Microbiol. 2006, 60, 69–105. [Google Scholar] [CrossRef]
- Lin, X.; Litvintseva, A.P.; Nielsen, K.; Patel, S.; Floyd, A.; Mitchell, T.G.; Heitman, J. αADα hybrids of Cryptococcus neoformans: Evidence of same-sex mating in nature and hybrid fitness. PLoS Genet. 2007, 3, e186. [Google Scholar] [CrossRef]
- Rhodes, J.; Desjardins, C.A.; Sykes, S.M.; Beale, M.A.; Vanhove, M.; Sakthikumar, S.; Chen, Y.; Gujja, S.; Saif, S.; Chowdhary, A.; et al. Tracing genetic exchange and biogeography of Cryptococcus neoformans var. grubii at the global population level. Genetics 2017, 207, 327–346. [Google Scholar] [CrossRef]
- Ni, M.; Feretzaki, M.; Li, W.; Floyd-Averette, A.; Mieczkowski, P.; Dietrich, F.S.; Heitman, J. Unisexual and heterosexual meiotic reproduction generate aneuploidy and phenotypic diversity de novo in the yeast Cryptococcus neoformans. PLoS Biol. 2013, 11, e1001653. [Google Scholar] [CrossRef]
- You, M.; Xu, J. The effects of environmental and genetic factors on the germination of basidiospores in the Cryptococcus gattii species complex. Sci. Rep. 2018, 8, 1–15. [Google Scholar] [CrossRef]
- Bovers, M.; Hagen, F.; Kuramae, E.E.; Diaz, M.R.; Spanjaard, L.; Dromer, F.; Hoogveld, H.L.; Boekhout, T. Unique hybrids between the fungal pathogens Cryptococcus neoformans and Cryptococcus gattii. FEMS Yeast Res. 2006, 6, 599–607. [Google Scholar] [CrossRef] [PubMed]
- Bovers, M.; Hagen, F.; Kuramae, E.E.; Hoogveld, H.L.; Dromer, F.; St-Germain, G.; Boekhout, T. AIDS patient death caused by novel Cryptococcus neoformans × C. gattii hybrid. Emerg. Infect. Dis. 2008, 14, 1105–1108. [Google Scholar] [CrossRef] [PubMed]
- Aminnejad, M.; Diaz, M.; Arabatzis, M.; Castañeda, E.; Lazera, M.; Velegraki, A.; Marriott, D.; Sorrell, T.C.; Meyer, W. Identification of novel hybrids between Cryptococcus neoformans var. grubii VNI and Cryptococcus gattii VGII. Mycopathologia 2012, 173, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Smith, I.M.; Stephan, C.; Hogardt, M.; Klawe, C.; Tintelnot, K.; Rickerts, V. Cryptococcosis due to Cryptococcus gattii in Germany from 2004 to 2013. Int. J. Med. Microbiol. 2015, 305, 719–723. [Google Scholar] [CrossRef] [PubMed]
- Hagen, F.; Hare Jensen, R.; Meis, J.F.; Arendrup, M.C. Molecular epidemiology and in vitro antifungal susceptibility testing of 108 clinical Cryptococcus neoformans sensu lato and Cryptococcus gattii sensu lato isolates from Denmark. Mycoses 2016, 59, 576–584. [Google Scholar] [CrossRef]
- Rhodes, J.; Desjardins, C.A.; Sykes, S.M.; Beale, M.A.; Vanhove, M.; Sakthikumar, S.; Chen, Y.; Gujja, S.; Saif, S.; Chowdhary, A.; et al. Population genomics of Cryptococcus neoformans var. grubii reveals new biogeographic relationships and finely maps hybridization. BioRxiv 2017, 132894. [Google Scholar] [CrossRef]
- Bennett, J.E.; Kwon-Chung, K.J.; Howard, D.H. Epidemiologic differences among serotypes of Cryptococcus neoformans. Am. J. Epidemiol. 1977, 105, 582–586. [Google Scholar] [CrossRef]
- Maduro, A.P.; Mansinho, K.; Teles, F.; Silva, I.; Meyer, W.; Martins, M.L.; Inácio, J. Insights on the genotype distribution among Cryptococcus neoformans and C. gattii Portuguese clinical isolates. Curr. Microbiol. 2014, 68, 199–203. [Google Scholar] [CrossRef]
- Cogliati, M.; Esposto, M.C.; Clarke, D.L.; Wickes, B.L.; Viviani, M.A. Origin of Cryptococcus neoformans var. neoformans diploid strains. J. Clin. Microbiol. 2001, 39, 3889–3894. [Google Scholar] [CrossRef]
- Viviani, M.A.; Antinori, S.; Cogliati, M.; Esposto, M.C.; Pinsi, G.; Casari, S.; Bergamasco, A.F.; Santis, M.D.; Ghirga, P.; Bonaccorso, C.; et al. European Confederation of Medical Mycology (ECMM) prospective survey of cryptococcosis: Report from Italy. Med. Mycol. 2002, 40, 507–517. [Google Scholar]
- Samarasinghe, H.; Xu, J. Hybrids and hybridization in the Cryptococcus neoformans and Cryptococcus gattii species complexes. Infect. Genet. Evol. 2018, 66, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Lengeler, K.B.; Cox, G.M.; Heitman, J. Serotype AD strains of Cryptococcus neoformans are diploid or aneuploid and are heterozygous at the mating-type locus. Infect. Immun. 2001, 69, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Xu, J. Genetic analyses of a hybrid cross between serotypes A and D strains of the human pathogenic fungus Cryptococcus neoformans. Genetics 2007, 177, 1475–1486. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Averette, A.F.; Desnos-Ollivier, M.; Ni, M.; Dromer, F.; Heitman, J. Genetic diversity and genomic plasticity of Cryptococcus neoformans AD hybrid strains. G3 (Bethesda) 2012, 2, 83–97. [Google Scholar] [CrossRef] [PubMed]
- Campbell, L.T.; Fraser, J.A.; Nichols, C.B.; Dietrich, F.S.; Carter, D.; Heitman, J. Clinical and environmental isolates of Cryptococcus gattii from Australia that retain sexual fecundity. Eukaryot. Cell 2005, 4, 1410–1419. [Google Scholar] [CrossRef]
- Kidd, S.E.; Hagen, F.; Tscharke, R.L.; Huynh, M.; Bartlett, K.H.; Fyfe, M.; Macdougall, L.; Boekhout, T.; Kwon-Chung, K.J.; Meyer, W. A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). Proc. Natl. Acad. Sci. USA 2004, 101, 17258–17263. [Google Scholar] [CrossRef] [PubMed]
- Fraser, J.A.; Subaran, R.L.; Nichols, C.B.; Heitman, J. Recapitulation of the sexual cycle of the primary fungal pathogen Cryptococcus neoformans var. gattii: Implications for an outbreak on Vancouver Island, Canada. Eukaryot. Cell 2003, 2, 1036–1045. [Google Scholar] [CrossRef]
- Voelz, K.; Ma, H.; Phadke, S.; Byrnes, E.J.; Zhu, P.; Mueller, O.; Farrer, R.A.; Henk, D.A.; Lewit, Y.; Hsueh, Y.-P.; et al. Transmission of hypervirulence traits via sexual reproduction within and between lineages of the human fungal pathogen Cryptococcus gattii. PLoS Genet. 2013, 9, e1003771. [Google Scholar] [CrossRef]
- Vogan, A.A.; Khankhet, J.; Xu, J. Evidence for mitotic recombination within the basidia of a hybrid cross of Cryptococcus neoformans. PLoS ONE 2013, 8, e62790. [Google Scholar] [CrossRef]
- Forsythe, A.; Vogan, A.; Xu, J. Genetic and environmental influences on the germination of basidiospores in the Cryptococcus neoformans species complex. Sci. Rep. 2016, 6, 1–12. [Google Scholar] [CrossRef]
- Tanaka, R.; Nishimura, K.; Miyaji, M. Ploidy of serotype AD strains of Cryptococcus neoformans. Nihon Ishinkin Gakkai Zasshi 1999, 40, 31–34. [Google Scholar] [CrossRef] [PubMed]
- Desnos-Ollivier, M.; Patel, S.; Raoux-Barbot, D.; Heitman, J.; Dromer, F.; French Cryptococcosis Study Group. Cryptococcosis serotypes impact outcome and provide evidence of Cryptococcus neoformans speciation. MBio 2015, 6, e00311. [Google Scholar] [CrossRef] [PubMed]
- Litvintseva, A.P.; Lin, X.; Templeton, I.; Heitman, J.; Mitchell, T.G. Many globally isolated AD hybrid strains of Cryptococcus neoformans originated in Africa. PLoS Pathog. 2007, 3, e114. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Nielsen, K.; Patel, S.; Heitman, J. Impact of mating type, serotype, and ploidy on the virulence of Cryptococcus neoformans. Infect. Immun. 2008, 76, 2923–2938. [Google Scholar] [CrossRef]
- Li, M.; Liao, Y.; Chen, M.; Pan, W.; Weng, L. Antifungal susceptibilities of Cryptococcus species complex isolates from AIDS and non-AIDS patients in Southeast China. Braz. J. Infect. Dis. 2012, 16, 175–179. [Google Scholar] [PubMed]
- Vogan, A.A.; Khankhet, J.; Samarasinghe, H.; Xu, J. Identification of QTLs associated with virulence related traits and drug resistance in Cryptococcus neoformans. G3 (Bethesda) 2016, 6, 2745–2759. [Google Scholar] [CrossRef]
- Sun, S.; Xu, J. Chromosomal rearrangements between serotype A and D strains in Cryptococcus neoformans. PLoS ONE 2009, 4, e5524. [Google Scholar] [CrossRef]
- Hu, G.; Liu, I.; Sham, A.; Stajich, J.E.; Dietrich, F.S.; Kronstad, J.W. Comparative hybridization reveals extensive genome variation in the AIDS-associated pathogen Cryptococcus neoformans. Genome Biol. 2008, 9, R41. [Google Scholar] [CrossRef]
- Kavanaugh, L.A.; Fraser, J.A.; Dietrich, F.S. Recent evolution of the human pathogen Cryptococcus neoformans by intervarietal transfer of a 14-gene fragment. Mol. Biol. Evol. 2006, 23, 1879–1890. [Google Scholar] [CrossRef]
- Vogan, A.A.; Xu, J. Evidence for genetic incompatibilities associated with post-zygotic reproductive isolation in the human fungal pathogen Cryptococcus neoformans. Genome 2014, 57, 335–344. [Google Scholar] [CrossRef]
- Cogliati, M.; Esposto, M.C.; Tortorano, A.M.; Viviani, M.A. Cryptococcus neoformans population includes hybrid strains homozygous at mating-type locus. FEMS Yeast Res. 2006, 6, 608–613. [Google Scholar] [CrossRef] [PubMed]
- Llorente, B.; Smith, C.E.; Symington, L.S. Break-induced replication: What is it and what is it for? Cell Cycle 2008, 7, 859–864. [Google Scholar] [CrossRef] [PubMed]
- Bennett, R.J.; Forche, A.; Berman, J. Rapid mechanisms for generating genome diversity: Whole ploidy shifts, aneuploidy, and loss of heterozygosity. Cold Spring Harb. Perspect. Med. 2014, 4, a019604. [Google Scholar] [CrossRef] [PubMed]
- Samarasinghe, H.; Vogan, A.; Pum, N.; Xu, J. Patterns of allele distribution in a hybrid population of the Cryptococcus neoformans species complex. Mycoses 2019. [CrossRef]
- Sionov, E.; Lee, H.; Chang, Y.C.; Kwon-Chung, K.J. Cryptococcus neoformans overcomes stress of azole drugs by formation of disomy in specific multiple chromosomes. PLoS Pathog. 2010, 6, e1000848. [Google Scholar] [CrossRef]
- Dong, K.; You, M.; Xu, J. Genetic changes in experimental populations of a hybrid in the Cryptococcus neoformans species complex. Pathogens 2020, 9, 3. [Google Scholar] [CrossRef]
- Lin, X.; Patel, S.; Litvintseva, A.P.; Floyd, A.; Mitchell, T.G.; Heitman, J. Diploids in the Cryptococcus neoformans serotype A population homozygous for the alpha mating type originate via unisexual mating. PLoS Pathog. 2009, 5, e1000283. [Google Scholar] [CrossRef]
- Hata, K.; Ohkusu, M.; Aoki, S.; Ito-Kuwa, S.; Pienthaweechai, K.; Takeo, K. Cells of different ploidy are often present together in Cryptococcus neoformans strains. Nihon Ishinkin Gakkai Zasshi 2000, 41, 161–167. [Google Scholar] [CrossRef][Green Version]
- Toffaletti, D.L.; Nielsen, K.; Dietrich, F.; Heitman, J.; Perfect, J.R. Cryptococcus neoformans mitochondrial genomes from serotype A and D strains do not influence virulence. Curr. Genet. 2004, 46, 193–204. [Google Scholar] [CrossRef]
- Skosireva, I.; James, T.Y.; Sun, S.; Xu, J. Mitochondrial inheritance in haploid x non-haploid crosses in Cryptococcus neoformans. Curr. Genet. 2010, 56, 163–176. [Google Scholar] [CrossRef][Green Version]
- Zaragoza, O.; Nielsen, K. Titan cells in Cryptococcus neoformans: Cells with a giant impact. Curr. Opin. Microbiol. 2013, 16, 409–413. [Google Scholar] [CrossRef]
- Hu, G.; Wang, J.; Choi, J.; Jung, W.H.; Liu, I.; Litvintseva, A.P.; Bicanic, T.; Aurora, R.; Mitchell, T.G.; Perfect, J.R.; et al. Variation in chromosome copy number influences the virulence of Cryptococcus neoformans and occurs in isolates from AIDS patients. BMC Genom. 2011, 12, 526. [Google Scholar] [CrossRef] [PubMed]
- Toffaletti, D.L.; Rude, T.H.; Johnston, S.A.; Durack, D.T.; Perfect, J.R. Gene transfer in Cryptococcus neoformans by use of biolistic delivery of DNA. J. Bacteriol. 1993, 175, 1405–1411. [Google Scholar] [CrossRef]
- Tanaka, R.; Taguchi, H.; Takeo, K.; Miyaji, M.; Nishimura, K. Determination of ploidy in Cryptococcus neoformans by flow cytometry. J. Med. Vet. Mycol. 1996, 34, 299–301. [Google Scholar] [CrossRef] [PubMed]
- Meyer, W.; Aanensen, D.M.; Boekhout, T.; Cogliati, M.; Diaz, M.R.; Esposto, M.C.; Fisher, M.; Gilgado, F.; Hagen, F.; Kaocharoen, S.; et al. Consensus multi-locus sequence typing scheme for Cryptococcus neoformans and Cryptococcus gattii. Med. Mycol. 2009, 47, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Sabiiti, W.; May, R.C.; Pursall, E.R. Experimental models of cryptococcosis. Int. J. Microbiol. 2012, 2012, 626745. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Lin, X. Multiple Applications of a transient CRISPR-Cas9 coupled with electroporation (TRACE) system in the Cryptococcus neoformans species complex. Genetics 2018, 208, 1357–1372. [Google Scholar] [CrossRef] [PubMed]
- Xie, G.Y.; Olson, D.H.; Blaustein, A.R. Projecting the global distribution of the emerging amphibian fungal pathogen, Batrachochytrium dendrobatidis, based on IPCC climate futures. PLoS ONE 2016, 11, e0160746. [Google Scholar] [CrossRef]
- Berger, L.; Speare, R.; Daszak, P.; Green, D.E.; Cunningham, A.A.; Goggin, C.L.; Slocombe, R.; Ragan, M.A.; Hyatt, A.D.; McDonald, K.R.; et al. Chytridiomycosis causes amphibian mortality associated with population declines in the rain forests of Australia and Central America. PNAS 1998, 95, 9031–9036. [Google Scholar] [CrossRef]
- Rachowicz, L.J.; Knapp, R.A.; Morgan, J.A.T.; Stice, M.J.; Vredenburg, V.T.; Parker, J.M.; Briggs, C.J. Emerging infectious disease as a proximate cause of amphibian mass mortality. Ecology 2006, 87, 1671–1683. [Google Scholar] [CrossRef]
- Olson, D.H.; Aanensen, D.M.; Ronnenberg, K.L.; Powell, C.I.; Walker, S.F.; Bielby, J.; Garner, T.W.J.; Weaver, G.; Bd Mapping Group; Fisher, M.C. Mapping the global emergence of Batrachochytrium dendrobatidis, the amphibian chytrid fungus. PLoS ONE 2013, 8, e56802. [Google Scholar] [CrossRef] [PubMed]
- O’Hanlon, S.J.; Rieux, A.; Farrer, R.A.; Rosa, G.M.; Waldman, B.; Bataille, A.; Kosch, T.A.; Murray, K.A.; Brankovics, B.; Fumagalli, M.; et al. Recent Asian origin of chytrid fungi causing global amphibian declines. Science 2018, 360, 621–627. [Google Scholar] [CrossRef] [PubMed]
- Farrer, R.A.; Weinert, L.A.; Bielby, J.; Garner, T.W.J.; Balloux, F.; Clare, F.; Bosch, J.; Cunningham, A.A.; Weldon, C.; du Preez, L.H.; et al. Multiple emergences of genetically diverse amphibian-infecting chytrids include a globalized hypervirulent recombinant lineage. PNAS 2011, 108, 18732–18736. [Google Scholar] [CrossRef] [PubMed]
- Schloegel, L.M.; Toledo, L.F.; Longcore, J.E.; Greenspan, S.E.; Vieira, C.A.; Lee, M.; Zhao, S.; Wangen, C.; Ferreira, C.M.; Hipolito, M.; et al. Novel, panzootic and hybrid genotypes of amphibian chytridiomycosis associated with the bullfrog trade. Mol. Ecol. 2012, 21, 5162–5177. [Google Scholar] [CrossRef] [PubMed]
- Byrne, A.Q.; Vredenburg, V.T.; Martel, A.; Pasmans, F.; Bell, R.C.; Blackburn, D.C.; Bletz, M.C.; Bosch, J.; Briggs, C.J.; Brown, R.M.; et al. Cryptic diversity of a widespread global pathogen reveals expanded threats to amphibian conservation. Proc. Natl. Acad. Sci. USA 2019, 116, 20382–20387. [Google Scholar] [CrossRef]
- James, T.Y.; Litvintseva, A.P.; Vilgalys, R.; Morgan, J.A.T.; Taylor, J.W.; Fisher, M.C.; Berger, L.; Weldon, C.; du Preez, L.; Longcore, J.E. Rapid global expansion of the fungal disease chytridiomycosis into declining and healthy amphibian populations. PLoS Pathog. 2009, 5, e1000458. [Google Scholar] [CrossRef]
- Rosenblum, E.B.; James, T.Y.; Zamudio, K.R.; Poorten, T.J.; Ilut, D.; Rodriguez, D.; Eastman, J.M.; Richards-Hrdlicka, K.; Joneson, S.; Jenkinson, T.S.; et al. Complex history of the amphibian-killing chytrid fungus revealed with genome resequencing data. Proc. Natl. Acad. Sci. USA 2013, 110, 9385–9390. [Google Scholar] [CrossRef]
- Jenkinson, T.S.; Betancourt Román, C.M.; Lambertini, C.; Valencia-Aguilar, A.; Rodriguez, D.; Nunes-de-Almeida, C.H.L.; Ruggeri, J.; Belasen, A.M.; da Silva Leite, D.; Zamudio, K.R.; et al. Amphibian-killing chytrid in Brazil comprises both locally endemic and globally expanding populations. Mol. Ecol. 2016, 25, 2978–2996. [Google Scholar] [CrossRef]
- Giraud, T.; Enjalbert, J.; Fournier, E.; Delmote, F.; Dutech, C. Population genetics of fungal diseases of plants. Parasite 2008, 15, 449–454. [Google Scholar] [CrossRef]
- Bennett, R.J. The parasexual lifestyle of Candida albicans. Curr. Opin. Microbiol. 2015, 28, 10–17. [Google Scholar] [CrossRef]
- Greenspan, S.E.; Lambertini, C.; Carvalho, T.; James, T.Y.; Toledo, L.F.; Haddad, C.F.B.; Becker, C.G. Hybrids of amphibian chytrid show high virulence in native hosts. Sci. Rep. 2018, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Becker, C.G.; Rodriguez, D.; Toledo, L.F.; Longo, A.V.; Lambertini, C.; Corrêa, D.T.; Leite, D.S.; Haddad, C.F.B.; Zamudio, K.R. Partitioning the net effect of host diversity on an emerging amphibian pathogen. Proc. Biol. Sci. 2014, 281, 20141796. [Google Scholar] [CrossRef] [PubMed]
- Farrer, R.A.; Henk, D.A.; Garner, T.W.J.; Balloux, F.; Woodhams, D.C.; Fisher, M.C. Chromosomal copy number variation, selection and uneven rates of recombination reveal cryptic genome diversity linked to pathogenicity. PLoS Genet. 2013, 9, e1003703. [Google Scholar] [CrossRef] [PubMed]
- Daum, J.M.; Davis, L.R.; Bigler, L.; Woodhams, D.C. Hybrid advantage in skin peptide immune defenses of water frogs (Pelophylax esculentus) at risk from emerging pathogens. Infect. Genet. Evol. 2012, 12, 1854–1864. [Google Scholar] [CrossRef]
- Refsnider, J.M.; Poorten, T.J.; Langhammer, P.F.; Burrowes, P.A.; Rosenblum, E.B. Genomic correlates of virulence attenuation in the deadly amphibian chytrid fungus, Batrachochytrium dendrobatidis. G3 (Bethesda) 2015, 5, 2291–2298. [Google Scholar] [CrossRef]
- Langhammer, P.F.; Lips, K.R.; Burrowes, P.A.; Tunstall, T.; Palmer, C.M.; Collins, J.P. A fungal pathogen of amphibians, Batrachochytrium dendrobatidis, attenuates in pathogenicity with in vitro passages. PLoS ONE 2013, 8, e77630. [Google Scholar] [CrossRef]
- James, T.Y.; Toledo, L.F.; Rödder, D.; da Silva Leite, D.; Belasen, A.M.; Betancourt-Román, C.M.; Jenkinson, T.S.; Soto-Azat, C.; Lambertini, C.; Longo, A.V.; et al. Disentangling host, pathogen, and environmental determinants of a recently emerged wildlife disease: Lessons from the first 15 years of amphibian chytridiomycosis research. Ecol. Evol. 2015, 5, 4079–4097. [Google Scholar] [CrossRef]
- Joneson, S.; Stajich, J.E.; Shiu, S.-H.; Rosenblum, E.B. Genomic transition to pathogenicity in chytrid fungi. PLoS Pathog. 2011, 7, e1002338. [Google Scholar] [CrossRef]
- Rosenblum, E.B.; Stajich, J.E.; Maddox, N.; Eisen, M.B. Global gene expression profiles for life stages of the deadly amphibian pathogen Batrachochytrium dendrobatidis. Proc. Natl. Acad. Sci. USA 2008, 105, 17034–17039. [Google Scholar] [CrossRef]
- Rosenblum, E.B.; Poorten, T.J.; Joneson, S.; Settles, M. Substrate-specific gene expression in Batrachochytrium dendrobatidis, the chytrid pathogen of amphibians. PLoS ONE 2012, 7, e49924. [Google Scholar] [CrossRef]
- Saikkonen, K.; Young, C.A.; Helander, M.; Schardl, C.L. Endophytic Epichloë species and their grass hosts: From evolution to applications. Plant Mol. Biol. 2016, 90, 665–675. [Google Scholar] [CrossRef]
- Inderbitzin, P.; Davis, R.M.; Bostock, R.M.; Subbarao, K.V. The ascomycete Verticillium longisporum is a hybrid and a plant pathogen with an expanded host range. PLoS ONE 2011, 6, e18260. [Google Scholar] [CrossRef] [PubMed]
- Fogelqvist, J.; Tzelepis, G.; Bejai, S.; Ilbäck, J.; Schwelm, A.; Dixelius, C. Analysis of the hybrid genomes of two field isolates of the soil-borne fungal species Verticillium longisporum. BMC Genom. 2018, 19, 14. [Google Scholar] [CrossRef] [PubMed]
- Marcet-Houben, M.; Gabaldón, T. Beyond the whole-genome duplication: Phylogenetic evidence for an ancient interspecies hybridization in the baker’s yeast lineage. PLoS Biol. 2015, 13, e1002220. [Google Scholar] [CrossRef] [PubMed]
- Fay, J.C.; Liu, P.; Ong, G.T.; Dunham, M.J.; Cromie, G.A.; Jeffery, E.W.; Ludlow, C.L.; Dudley, A.M. A polyploid admixed origin of beer yeasts derived from European and Asian wine populations. PLoS Biol. 2019, 17, e3000147. [Google Scholar] [CrossRef]
- Langdon, Q.K.; Peris, D.; Baker, E.P.; Opulente, D.A.; Nguyen, H.-V.; Bond, U.; Gonçalves, P.; Sampaio, J.P.; Libkind, D.; Hittinger, C.T. Fermentation innovation through complex hybridization of wild and domesticated yeasts. Nat. Ecol. Evol. 2019, 3, 1576–1586. [Google Scholar] [CrossRef] [PubMed]
- Gostinčar, C.; Stajich, J.E.; Zupančič, J.; Zalar, P.; Gunde-Cimerman, N. Genomic evidence for intraspecific hybridization in a clonal and extremely halotolerant yeast. BMC Genom. 2018, 19, 364. [Google Scholar] [CrossRef]
- Pryszcz, L.P.; Németh, T.; Saus, E.; Ksiezopolska, E.; Hegedűsová, E.; Nosek, J.; Wolfe, K.H.; Gacser, A.; Gabaldón, T. The genomic aftermath of hybridization in the opportunistic pathogen Candida metapsilosis. PLoS Genet. 2015, 11, e1005626. [Google Scholar] [CrossRef]
- Stukenbrock, E.H.; Christiansen, F.B.; Hansen, T.T.; Dutheil, J.Y.; Schierup, M.H. Fusion of two divergent fungal individuals led to the recent emergence of a unique widespread pathogen species. PNAS 2012, 109, 10954–10959. [Google Scholar] [CrossRef]
- Le Gac, M.; Hood, M.E.; Fournier, E.; Giraud, T. Phylogenetic evidence of host-specific cryptic species in the anther smut fungus. Evolution 2007, 61, 15–26. [Google Scholar] [CrossRef]
- Turner, E.; Jacobson, D.J.; Taylor, J.W. Genetic architecture of a reinforced, postmating, reproductive isolation barrier between Neurospora species indicates evolution via natural selection. PLoS Genet. 2011, 7, e1002204. [Google Scholar] [CrossRef] [PubMed]
- Büker, B.; Petit, E.; Begerow, D.; Hood, M.E. Experimental hybridization and backcrossing reveal forces of reproductive isolation in Microbotryum. BMC Evol. Biol. 2013, 13, 224. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Chacko, N.; Wang, L.; Pavuluri, Y. Generation of stable mutants and targeted gene deletion strains in Cryptococcus neoformans through electroporation. Med. Mycol. 2015, 53, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Olson, A.; Stenlid, J. Plant pathogens. Mitochondrial control of fungal hybrid virulence. Nature 2001, 411, 438. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Samarasinghe, H.; You, M.; Jenkinson, T.S.; Xu, J.; James, T.Y. Hybridization Facilitates Adaptive Evolution in Two Major Fungal Pathogens. Genes 2020, 11, 101. https://doi.org/10.3390/genes11010101
Samarasinghe H, You M, Jenkinson TS, Xu J, James TY. Hybridization Facilitates Adaptive Evolution in Two Major Fungal Pathogens. Genes. 2020; 11(1):101. https://doi.org/10.3390/genes11010101
Chicago/Turabian StyleSamarasinghe, Himeshi, Man You, Thomas S. Jenkinson, Jianping Xu, and Timothy Y. James. 2020. "Hybridization Facilitates Adaptive Evolution in Two Major Fungal Pathogens" Genes 11, no. 1: 101. https://doi.org/10.3390/genes11010101
APA StyleSamarasinghe, H., You, M., Jenkinson, T. S., Xu, J., & James, T. Y. (2020). Hybridization Facilitates Adaptive Evolution in Two Major Fungal Pathogens. Genes, 11(1), 101. https://doi.org/10.3390/genes11010101





