Reconstruction and Analysis of Gene Networks of Human Neurotransmitter Systems Reveal Genes with Contentious Manifestation for Anxiety, Depression, and Intellectual Disabilities
Abstract
:1. Introduction
2. Materials and Methods
2.1. Reconstruction of Transcriptional Regulation Networks
2.2. SNP Analysis
2.3. Comparison with Current Gene Panels
3. Results
3.1. Formation of the Initial List of Genes
- SLC6A4—serotonin transporter gene. Many studies have shown a relationship between 5-HTTLPR polymorphism and anxiety [2], depression, and bipolar disorder, antidepressant sensitivity [28], risk of depression, and suicide [29]. However, in a recent study, it has been shown to reverse the effects of this polymorphism in several Asian populations such as Japanese [12], Chinese [30], and Tuvinians [11].
- HLA-B (human leukocyte antigen) is one of the genes of the histocompatibility complex of class 1 in humans. Its allele, HLA-B *1502, plays an important role in the development of such carbamazepine-caused diseases as Stevens–Johnson syndrome and Lyell’s syndrome, but its effect is seen only in Chinese [31] and Malaysian populations [32]. In individuals of European origin, its influence was not found to be significant [33].
- ABCB1 (ATP-binding cassette) is a gene of the membrane protein P-glycoprotein, which may transport xenobiotic substances through the cell membrane. C3435T allele of this gene supposedly plays a role in the development of drug resistance to epilepsy in a number of populations: Japan [34], Taiwan [35], but does not show a significant role in Korea [36], Turkey [37], Great Britain [38,39], and Ireland [40].
- Apolipoprotein E (ApoE) is one of the most important apolipoproteins, involved in the metabolism of blood lipids on the one hand, and cholesterol metabolism in the brain on the other. The ε4 allele of this gene appears to be associated with an increased risk of temporal epilepsy after head injuries. Also, ApoE genotype is related to a level of intellectual disability, and presence of dementia in Down syndrome [41]. However, studies confirming this relationship were obtained only by researchers from the United States [42,43]. In other countries, no such relationship has been identified [44,45].
- BDNF—neurotrophin, a signaling protein that stimulates and supports the development of neurons. Peripheral BDNF level is associated with attention deficit and intellectual disability in preschool children [46]. In Japanese population, it is a trait of susceptibility to partial epilepsy [47], but in European populations, such dependence could not be detected [48,49].
- COMT is an enzyme that plays an important role in the degradation of catecholamines, including dopamine, adrenaline and norepinephrine and the role of rs4680 polymorphism in the etiology of idiopathic intellectual disability was revealed [50]. Additionally, the role of rs4680 as a risk factor for bipolar disorder was shown in Chinese population [51], as well as in Ashkenazi population [52], but its influence was considered insignificant in European populations [53].
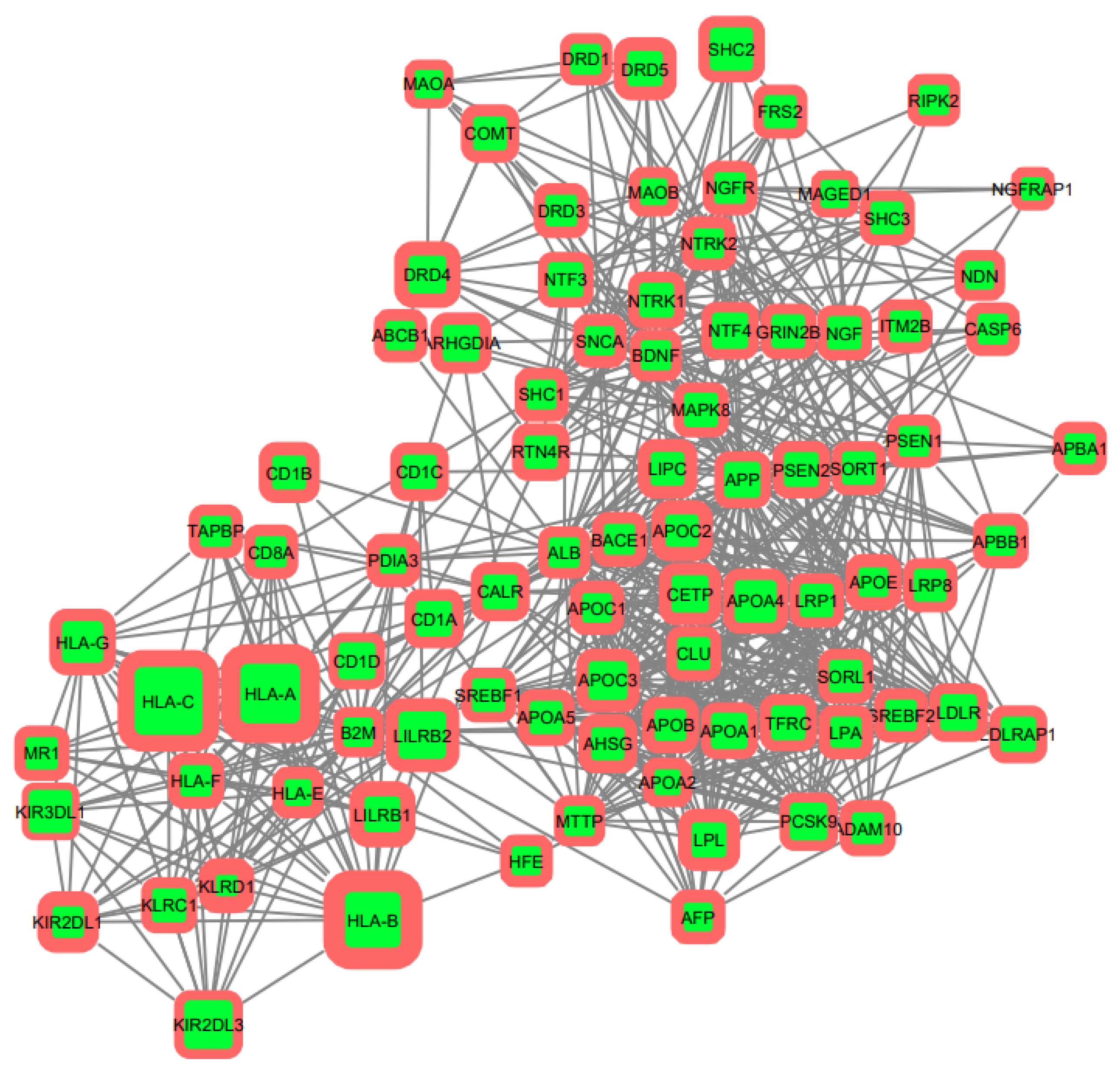
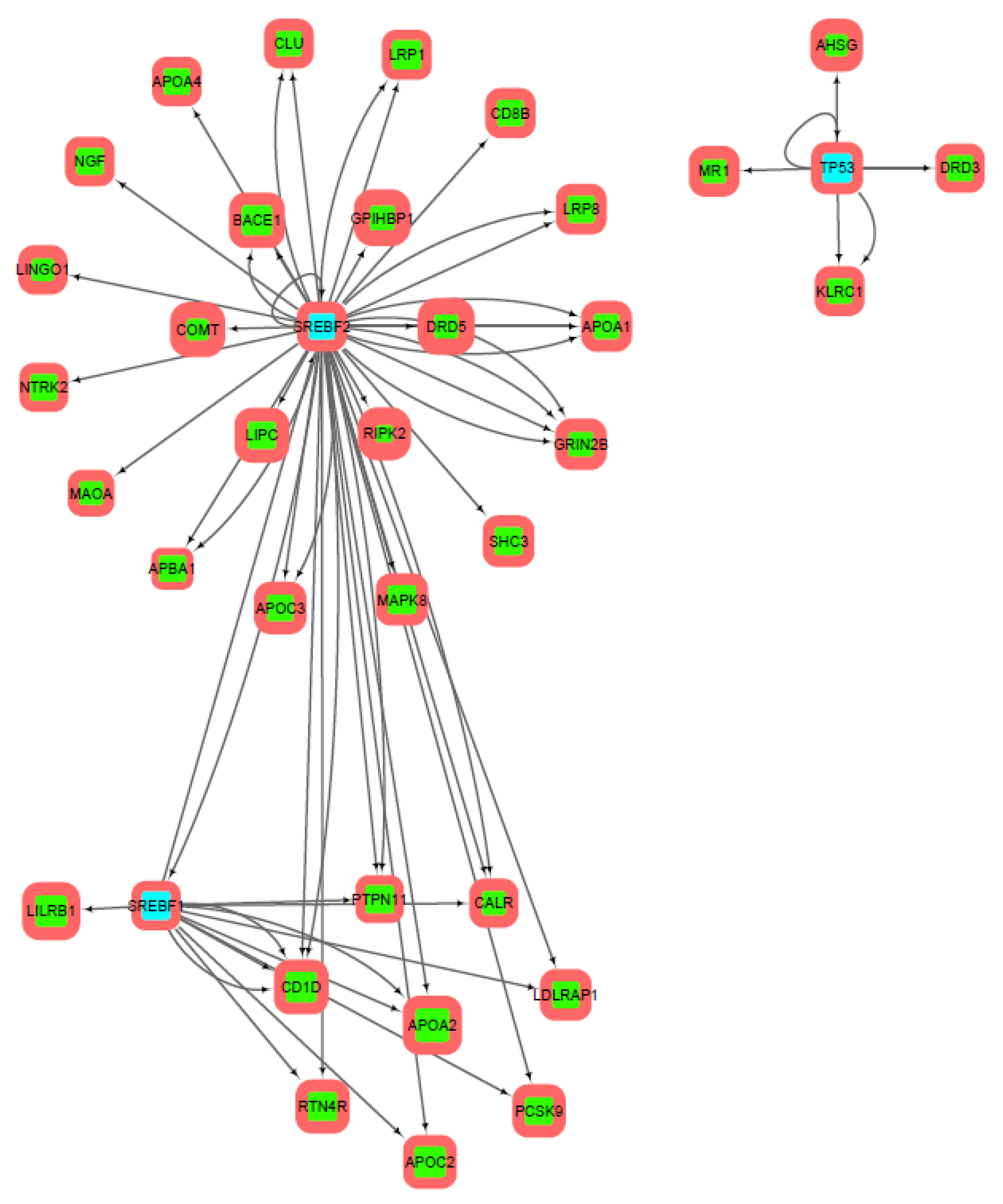
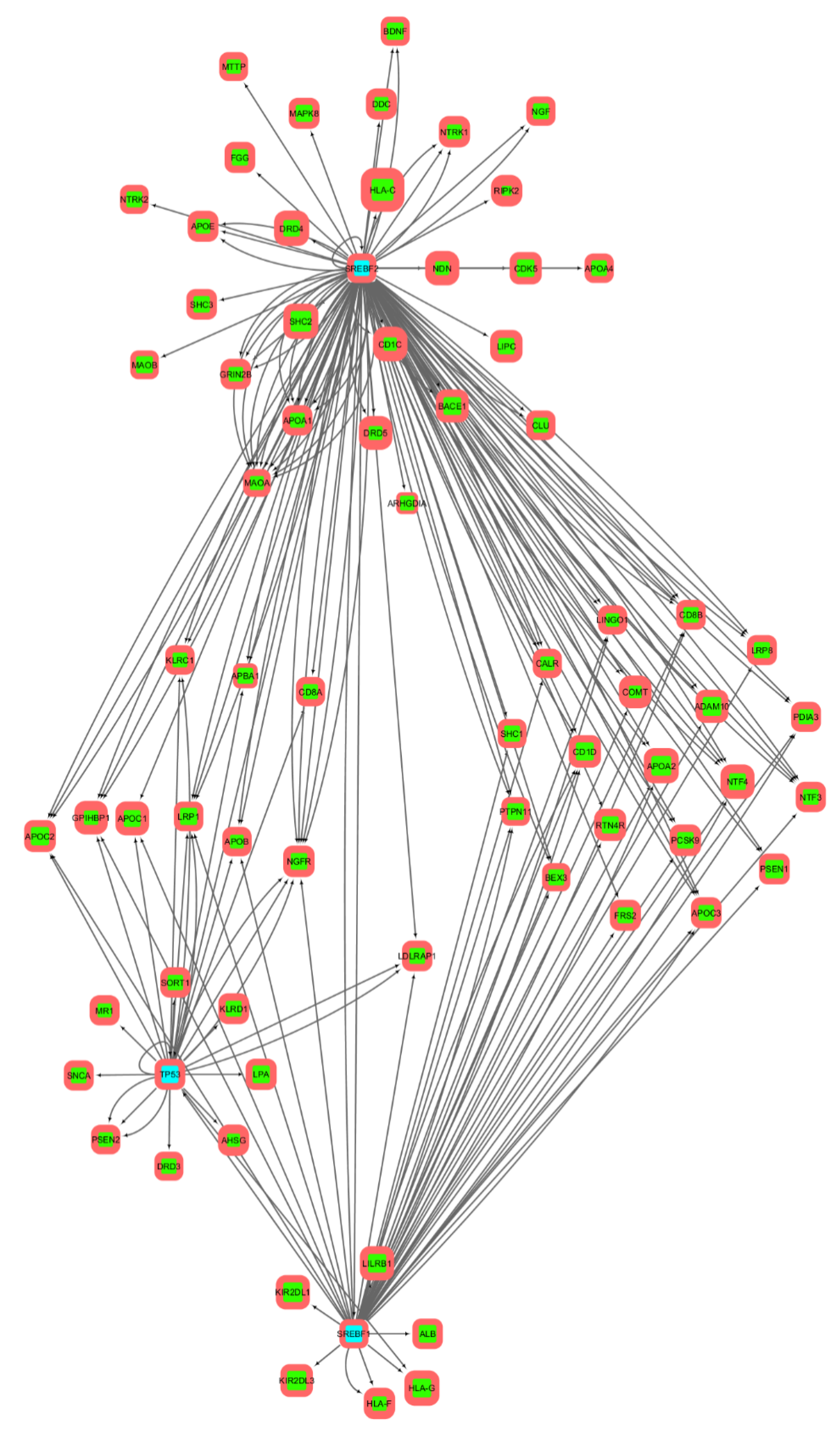
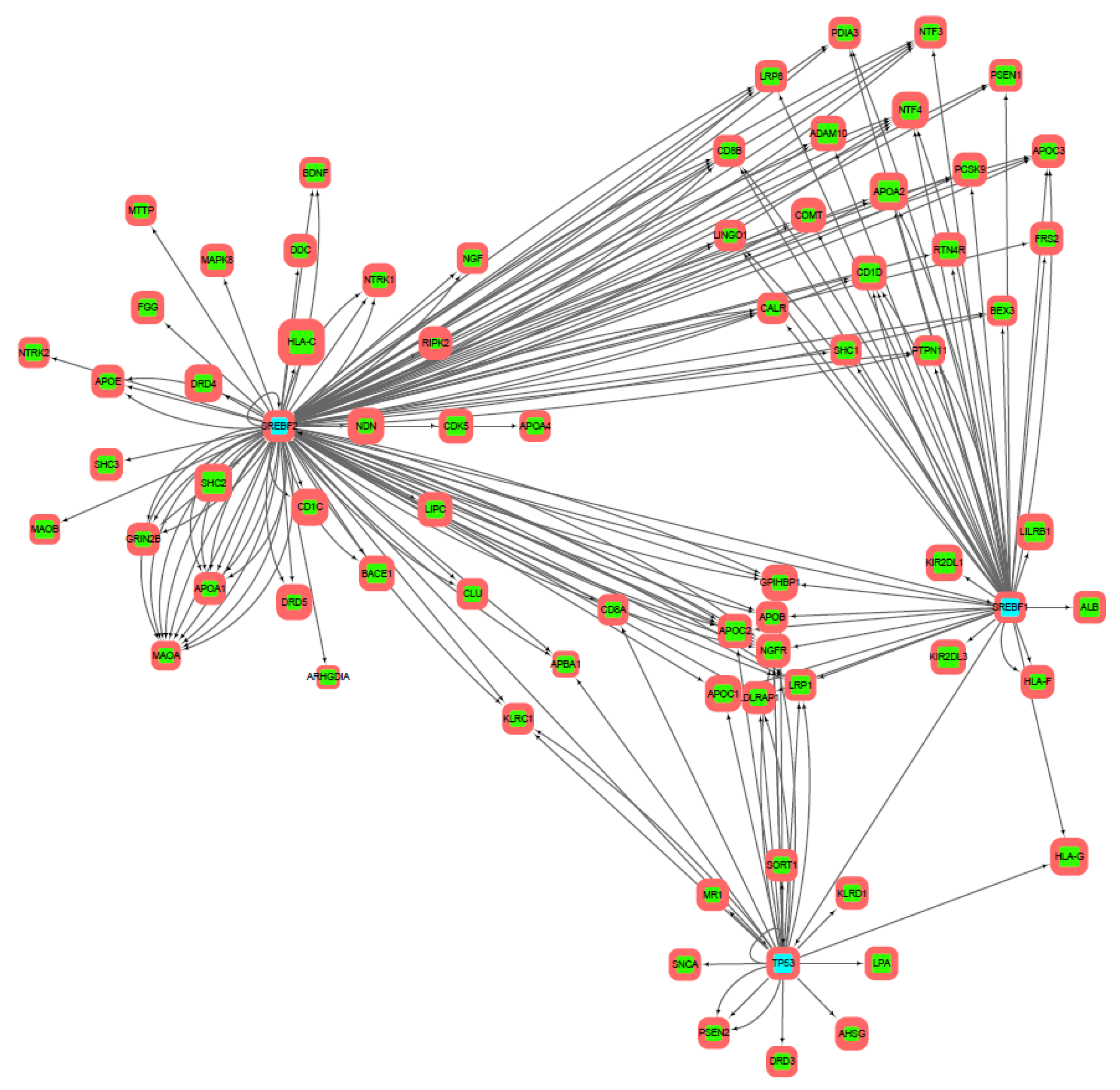
3.2. Gene Networks Reconstruction
3.3. Gene Sets Formation
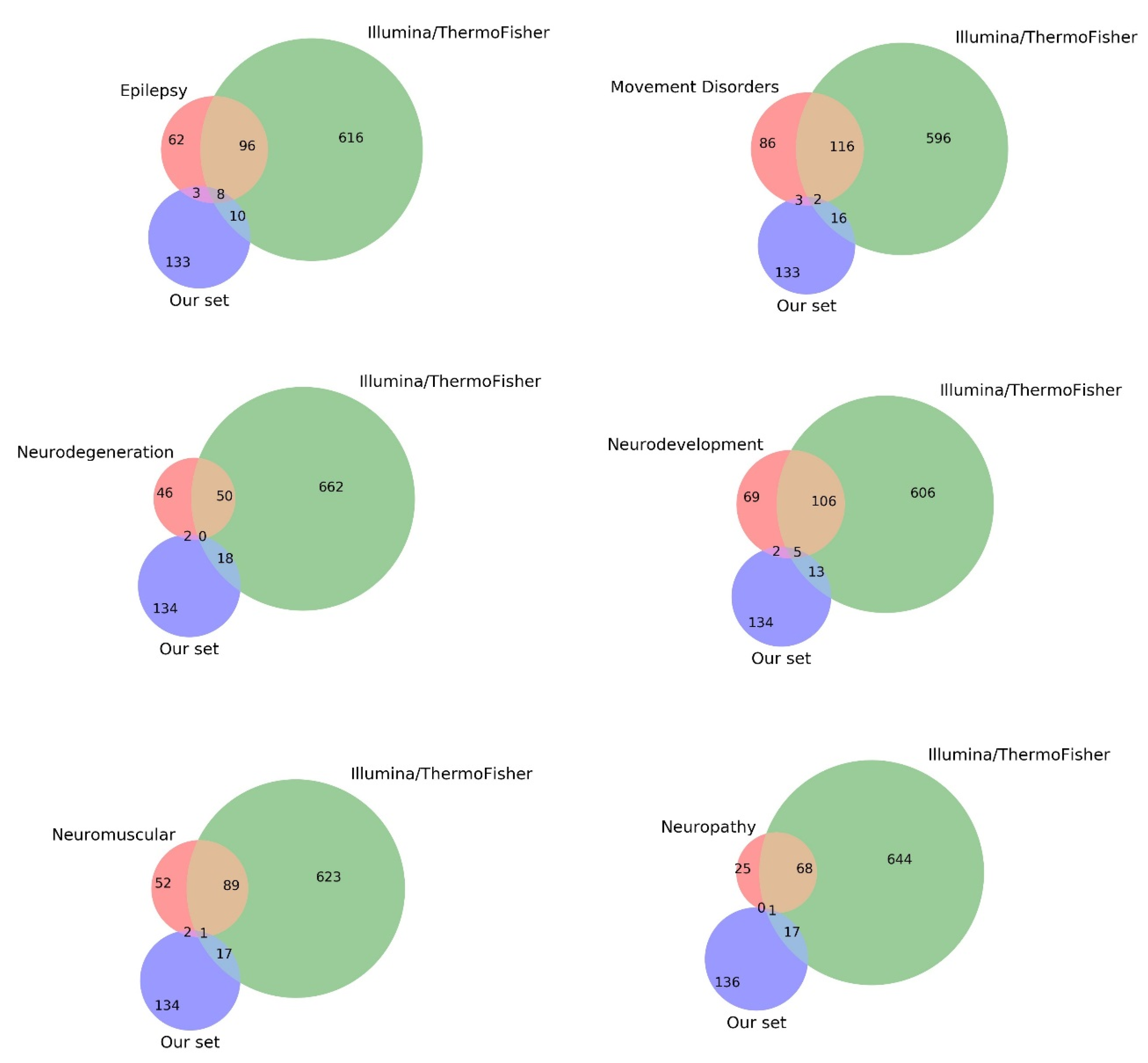
3.4. Comparison with Current Gene Panels
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Appendix A
References
- Reul, J.M.H.M.; Holsboer, F. Corticotropin-releasing factor receptors 1 and 2 in anxiety and depression. Curr. Opin. Pharmacol. 2002, 2, 23–33. [Google Scholar] [CrossRef]
- Benmansour, S.; Cecchi, M.; Morilak, D.A.; Gerhardt, G.A.; Javors, M.A.; Gould, G.G.; Frazer, A. Effects of chronic antidepressant treatments on serotonin transporter function, density, and mRNA level. J. Neurosci. 1999, 19, 10494–10501. [Google Scholar] [CrossRef]
- Chang, S.H.; Gao, L.; Li, Z.; Zhang, W.N.; Du, Y.; Wang, J. BDgene: A genetic database for bipolar disorder and its overlap with schizophrenia and major depressive disorder. Biol. Psychiatry 2013, 74, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Baudewijns, L.; Ronsse, E.; Verstraete, V.; Sabbe, B.; Morrens, M.; Bertelli, M.O. Problem behaviours and Major Depressive Disorder in adults with intellectual disability and autism. Psychiatry Res. 2018, 270, 769–774. [Google Scholar] [CrossRef] [PubMed]
- El Mrayyan, N.; Eberhard, J.; Ahlström, G. The occurrence of comorbidities with affective and anxiety disorders among older people with intellectual disability compared with the general population: A register study. BMC Psychiatry 2019, 19, 1–12. [Google Scholar] [CrossRef]
- Zuckerman, H.; Pan, Z.; Park, C.; Brietzke, E.; Musial, N.; Shariq, A.S.; Iacobucci, M.; Yim, S.J.; Lui, L.M.; Rong, C.; et al. Recognition and Treatment of Cognitive Dysfunction in Major Depressive Disorder. Front. Psychiatry 2018, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Whitney, D.G.; Shapiro, D.N.; Peterson, M.D.; Warschausky, S.A. Factors associated with depression and anxiety in children with intellectual disabilities. J. Intellect. Disabil. Res. 2019, 63, 408–417. [Google Scholar] [CrossRef]
- Scherer, N.; Verhey, I.; Kuper, H. Depression and anxiety in parents of children with intellectual and developmental disabilities: A systematic review and meta-analysis. PLoS ONE 2019, 14, e0219888. [Google Scholar] [CrossRef]
- Duncan, L.E.; Pouastri, A.R.; Smoller, J.W. Mind the gap: Why many geneticists and psychological scientists have discrepant views about gene–environment interaction (G× E) research. Am. Psychol. 2014, 69, 249–268. [Google Scholar] [CrossRef]
- Risch, N.; Herrell, R.; Lehner, T. Interaction between the Serotonin. JAMA 2009, 301, 2462–2472. [Google Scholar] [CrossRef]
- Savostyanov, A.N.; Naumenko, V.S.; Sinyakova, N.A.; L’vova, M.N.; Levin, E.A.; Zaleshin, M.S.; Kavay-ool, U.N.; Mordvinov, V.A.; Kolchanov, N.A.; Aftanas, L.I. Association of anxiety level with polymorphic variants of serotonin transporter gene in Russians and Tuvinians. Russ. J. Genet. Appl. Res. 2015, 5, 656–665. [Google Scholar] [CrossRef]
- Nomura, M.; Kaneko, M.; Okuma, Y.; Nomura, J.; Kusumi, I.; Koyama, T.; Nomura, Y. Involvement of serotonin transporter gene polymorphisms (5-HTT) in impulsive behavior in the Japanese population. PLoS ONE 2015, 10, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Wakasugi, M.; Kazama, J.J.; Narita, I.; Iseki, K.; Moriyama, T.; Yamagata, K.; Fujimoto, S.; Tsuruya, K.; Asahi, K.; Konta, T.; et al. Association between Combined Lifestyle Factors and Non-Restorative Sleep in Japan: A Cross-Sectional Study Based on a Japanese Health Database. PLoS ONE 2014. [Google Scholar] [CrossRef] [PubMed]
- Bosman, C.A.; Lansink, C.S.; Pennartz, C.M.A. Functions of gamma-band synchronization in cognition: From single circuits to functional diversity across cortical and subcortical systems. Eur. J. Neurosci. 2014, 39, 1982–1999. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.Y.; Iga, J.I.; Numata, S.; Umehara, H.; Nishi, A.; Kinoshita, M.; Inoshita, M.; Ohmori, T. Polymorphism in the promoter of the gene for the serotonin transporter affects the age of onset of major depressive disorder in the Japanese population. J. Affect. Disord. 2015, 183, 156–158. [Google Scholar] [CrossRef] [PubMed]
- Abrahams, B.S.; Arking, D.E.; Campbell, D.B.; Mefford, H.C.; Morrow, E.M.; Weiss, L.A.; Menashe, I.; Wadkins, T.; Banerjee-Basu, S.; Packer, A. SFARI Gene 2.0: A community-driven knowledgebase for the autism spectrum disorders (ASDs). Mol. Autism 2013, 4, 36. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez-Sacristán, A.; Grosdidier, S.; Valverde, O.; Torrens, M.; Bravo, À.; Piñero, J.; Sanz, F.; Furlong, L.I. PsyGeNET: A knowledge platform on psychiatric disorders and their genes. Bioinformatics 2015, 31, 3075–3077. [Google Scholar] [CrossRef]
- Tan, N.C.K.; Berkovic, S.F. The epilepsy genetic association database (epiGAD): Analysis of 165 genetic association studies, 1996–2008. Epilepsia 2010, 51, 686–689. [Google Scholar] [CrossRef]
- Guo, L.; Zhang, W.; Chang, S.; Zhang, L.; Ott, J.; Wang, J. MK4MDD: A Multi-Level Knowledge Base and Analysis Platform for Major Depressive Disorder. PLoS ONE 2012, 7, e46335. [Google Scholar] [CrossRef]
- Braschi, B.; Denny, P.; Gray, K.; Jones, T.; Seal, R.; Tweedie, S.; Yates, B.; Bruford, E. Genenames. Org: The HGNC and VGNC resources in 2019. Nucleic Acids Res. 2018, 47, D786–D792. [Google Scholar] [CrossRef]
- Sullivan, P.F.; Agrawal, A.; Bulik, C.M.; Andreassen, O.A.; Børglum, A.D.; Breen, G.; Cichon, S.; Edenberg, H.J.; Faraone, S.V.; Gelernter, J.; et al. Psychiatric genomics: An update and an Agenda. Am. J. Psychiatry 2017, 175, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Von Mering, C.; Jensen, L.J.; Snel, B.; Hooper, S.D.; Krupp, M.; Foglierini, M.; Jouffre, N.; Huynen, M.A.; Bork, P. STRING: Known and predicted protein-protein associations, integrated and transferred across organisms. Nucleic Acids Res. 2005, 33, 433–437. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.M.; Liu, T.; Liu, C.J.; Song, S.; Zhang, X.; Liu, W.; Jia, H.; Xue, Y.; Guo, A.Y. AnimalTFDB 2 0: A resource for expression, prediction and functional study of animal transcription factors. Nucleic Acids Res. 2015, 43, 76–81. [Google Scholar] [CrossRef] [PubMed]
- Kulakovskiy, I.V.; Vorontsov, I.E.; Yevshin, I.S.; Sharipov, R.N.; Fedorova, A.D.; Rumynskiy, E.I.; Medvedeva, Y.A.; Magana-Mora, A.; Bajic, V.B.; Papatsenko, D.A.; et al. HOCOMOCO: Towards a complete collection of transcription factor binding models for human and mouse via large-scale ChIP-Seq analysis. Nucleic Acids Res. 2017, 46, D252–D259. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Sudmant, P.H.; Rausch, T.; Gardner, E.J.; Handsaker, R.E.; Abyzov, A.; Huddleston, J.; Zhang, Y.; Ye, K.; Jun, G.; Fritz, M.H.Y.; et al. An integrated map of structural variation in 2,504 human genomes. Nature 2015, 526, 75–81. [Google Scholar] [CrossRef] [Green Version]
- Porcelli, S.; Fabbri, C.; Serretti, A. Meta-analysis of serotonin transporter gene promoter polymorphism (5-HTTLPR) association with antidepressant efficacy. Eur. Neuropsychopharmacol. 2012, 22, 239–258. [Google Scholar] [CrossRef]
- Kato, T. Molecular genetics of bipolar disorder and depression. Psychiatry Clin. Neurosci. 2007, 61, 3–19. [Google Scholar] [CrossRef]
- Guo, W.Y.; Zhang, Z.H.; Mu, J.L.; Liu, D.; Zhao, L.; Yao, Z.Y.; Song, J.G. Relationship between 5-HTTLPR polymorphism and post-stroke depression. Genet. Mol. Res. 2016, 15, 1–6. [Google Scholar] [CrossRef]
- Hung, S.I.; Chung, W.H.; Jee, S.H.; Chen, W.C.; Chang, Y.T.; Lee, W.R.; Hu, S.L.; Wu, M.T.; Chen, G.S.; Wong, T.W.; et al. Genetic susceptibility to carbamazepine-induced cutaneous adverse drug reactions. Pharmacogenet. Genomics 2006, 16, 297–306. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.C.; Too, C.L.; Murad, S.; Hussein, S.H. Association of HLA-B1502 allele with carbamazepine-induced toxic epidermal necrolysis and Stevens-Johnson syndrome in the multi-ethnic Malaysian population. Int. J. Dermatol. 2011, 50, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Alfirevic, A.; Jorgensen, A.L.; Williamson, P.R.; Chadwick, D.W.; Park, B.K.; Pirmohamed, M. HLA-B locus in Caucasian patients with carbamazepine hypersensitivity. Pharmacogenomics 2006, 7, 813–818. [Google Scholar] [CrossRef] [PubMed]
- Seo, T.; Ishitsu, T.; Ueda, N.; Nakada, N.; Yurube, K.; Ueda, K.; Nakagawa, K. ABCB1 polymorphisms influence the response to antiepileptic drugs in Japanese epilepsy patients. Pharmacogenomics 2006, 7, 551–561. [Google Scholar] [CrossRef] [PubMed]
- Hung, C.C.; Tai, J.J.; Lin, C.J.; Lee, M.J.; Liou, H.H. Complex haplotypic effects of the ABCB1 gene on epilepsy treatment response. Pharmacogenomics 2005, 6, 411–417. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.O.; Kim, M.K.; Woo, Y.J.; Lee, M.C.; Kim, J.H.; Park, K.W.; Kim, E.Y.; Roh, Y.I.; Kim, C.J. Single nucleotide polymorphisms in the multidrug resistance 1 gene in Korean epileptics. Seizure 2006, 15, 67–72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dericioglu, N.; Babaoglu, M.O.; Yasar, U.; Bal, I.B.; Bozkurt, A.; Saygi, S. Multidrug resistance in patients undergoing resective epilepsy surgery is not associated with C3435T polymorphism in the ABCB1 (MDR1) gene. Epilepsy Res. 2008, 80, 42–46. [Google Scholar] [CrossRef] [PubMed]
- Leschziner, G.D.; Andrew, T.; Leach, J.P.; Chadwick, D.; Coffey, A.J.; Balding, D.J.; Bentley, D.R.; Pirmohamed, M.; Johnson, M.R. Common ABCB1 polymorphisms are not associated with multidrug resistance in epilepsy using a gene-wide tagging approach. Pharmacogenet. Genomics 2007, 17, 217–220. [Google Scholar] [CrossRef] [PubMed]
- Sills, G.J.; Mohanraj, R.; Butler, E.; McCrindle, S.; Collier, L.; Wilson, E.A.; Brodie, M.J. Lack of association between the C3435T polymorphism in the human multidrug resistance (MDR1) gene and response to antiepileptic drug treatment. Epilepsia 2005, 46, 643–647. [Google Scholar] [CrossRef]
- Shahwan, A.; Murphy, K.; Doherty, C.; Cavalleri, G.L.; Muckian, C.; Dicker, P.; McCarthy, M.; Kinirons, P.; Goldstein, D.; Delanty, N. The controversial association of ABCB1 polymorphisms in refractory epilepsy: An analysis of multiple SNPs in an Irish population. Epilepsy Res. 2007, 73, 192–198. [Google Scholar] [CrossRef]
- Head, E.; Doran, E.; Nistor, M.; Hill, M.; Schmitt, F.A.; Haier, R.J.; Lott, I.T. Plasma amyloid-β as a function of age, level of intellectual disability, and presence of dementia in down syndrome. J. Alzheimer’s Dis. 2011, 23, 399–409. [Google Scholar] [CrossRef] [PubMed]
- Chapin, J.S.; Busch, R.M.; Janigro, D.; Dougherty, M.; Tilelli, C.Q.; Lineweaver, T.T.; Naugle, R.I.; Diaz-Arrastia, R.; Najm, I.M. APOE ε4 is associated with postictal confusion in patients with medically refractory temporal lobe epilepsy. Epilepsy Res. 2008, 81, 220–224. [Google Scholar] [CrossRef] [PubMed]
- Busch, R.M.; Lineweaver, T.T.; Naugle, R.I.; Kim, K.H.; Gong, Y.; Tilelli, C.Q.; Prayson, R.A.; Bingaman, W.; Najm, I.M.; Diaz-Arrastia, R.; et al. APOE ɛ4 is associated with reduced memory in long-standing intractable temporal lobe epilepsy. Neurology 2007, 68, 409–414. [Google Scholar] [CrossRef] [PubMed]
- Blümcke, I.; Brockhaus, A.; Scheiwe, C.; Rollbrocker, B.; Wolf, H.K.; Elger, C.E.; Wiestler, O.D. The apolipoprotein E ε4 allele is not associated with early onset temporal lobe epilepsy. Neuroreport 1997, 8, 1235–1237. [Google Scholar]
- Cavalleri, G.L.; Lynch, J.M.; Depondt, C.; Burley, M.W.; Wood, N.W.; Sisodiya, S.M.; Goldstein, D.B. Failure to replicate previously reported genetic associations with sporadic temporal lobe epilepsy: Where to from here? Brain 2005, 128, 1832–1840. [Google Scholar] [CrossRef] [PubMed]
- Yeom, C.W.; Park, Y.J.; Choi, S.W.; Bhang, S.Y. Association of peripheral BDNF level with cognition, attention and behavior in preschool children. Child. Adolesc. Psychiatry Ment. Health 2016, 10, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kanemoto, K.; Kawasaki, J.; Tarao, Y.; Kumaki, T.; Oshima, T.; Kaji, R.; Nishimura, M. Association of partial epilepsy with brain-derived neurotrophic factor (BDNF) gene polymorphisms. Epilepsy Res. 2003, 53, 255–258. [Google Scholar] [CrossRef]
- Louhivuori, V.; Arvio, M.; Soronen, P.; Oksanen, V.; Paunio, T.; Castrén, M.L. The Val66Met polymorphism in the BDNF gene is associated with epilepsy in fragile X syndrome. Epilepsy Res. 2009, 85, 114–117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lohoff, F.W.; Ferraro, T.N.; Dahl, J.P.; Hildebrandt, M.A.; Scattergood, T.M.; O’Connor, M.J.; Sperling, M.R.; Dlugos, D.J.; Berrettini, W.H.; Buono, R.J. Lack of association between variations in the brain-derived neurotrophic factor (BDNF) gene and temporal lobe epilepsy. Epilepsy Res. 2005, 66, 59–62. [Google Scholar] [CrossRef] [PubMed]
- Bhowmik, A.D.; Chaudhury, S.; Dutta, S.; Shaw, J.; Chatterjee, A.; Choudhury, A.; Saha, A.; Sadhukhan, D.; Kar, T.; Sinha, S.; et al. Role of functional dopaminergic gene polymorphisms in the etiology of idiopathic intellectual disability. Prog. Neuro-Psychopharmacology Biol. Psychiatry 2011, 35, 1714–1722. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Lindpaintner, K.; Che, R.; He, Z.; Wang, P.; Yang, P.; Feng, G.; He, L.; Shi, Y. The Val/Met functional polymorphism in COMT confers susceptibility to bipolar disorder: Evidence from an association study and a meta-analysis. J. Neural Transm. 2009, 116, 1193–1200. [Google Scholar] [CrossRef] [PubMed]
- Shifman, S.; Bronstein, M.; Sternfeld, M.; Pisanté, A.; Weizman, A.; Reznik, I.; Spivak, B.; Grisaru, N.; Karp, L.; Schiffer, R.; et al. COMT: A Common Susceptibility Gene in Bipolar Disorder and Schizophrenia. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2004, 64, 61–64. [Google Scholar] [CrossRef] [PubMed]
- Lachman, H.M.; Saoto, T.; Papolos, D.F.; Weinshilboum, R.M.; Szumlanski, C.L. Human catechol-O-methyltransferase pharmacogenetics: Description of a functional polymorphism and its potential application to neuropsychiatric disorders. Pharmacogenetics 1996, 6, 234–250. [Google Scholar] [CrossRef]
| Category | Term | Count | Set Content (%) | P Value |
|---|---|---|---|---|
| GAD_DISEASE | Schizophrenia | 59 | 3806 | 1.22 × 10−33 |
| GAD_DISEASE | several psychiatric disorders | 56 | 3612 | 1.51 × 10−53 |
| GAD_DISEASE | Tobacco Use Disorder | 53 | 3419 | 3.05 × 10−4 |
| GAD_DISEASE | Type 2 Diabetes| edema | rosiglitazone | 51 | 3290 | 1.293 × 10−7 |
| GAD_DISEASE | Autism | 46 | 2967 | 1.126 × 10−35 |
| GAD_DISEASE | Bipolar Disorder | 41 | 2645 | 2.60 × 10−27 |
| GAD_DISEASE | alcohol consumption | 35 | 2258 | 3.92 × 10−38 |
| GAD_DISEASE | Alzheimer’s disease | 27 | 1741 | 1.31 × 10−5 |
| GAD_DISEASE | ADHD | attention-deficit hyperactivity disorder | 24 | 1548 | 7.66 × 10−24 |
| GOTERM_BP_DIRECT | GO:0007268~chemical synaptic transmission | 38 | 2451 | 1.42 × 10−35 |
| GOTERM_BP_DIRECT | GO:0007165~signal transduction | 25 | 1612 | 9.60 × 10−5 |
| GOTERM_CC_DIRECT | GO:0005886~plasma membrane | 96 | 6193 | 7.52 × 10−26 |
| GOTERM_CC_DIRECT | GO:0016021~integral component of membrane | 81 | 5225 | 3.35 × 10−10 |
| GOTERM_CC_DIRECT | GO:0005887~integral component of plasma membrane | 63 | 4064 | 1.77 × 10−29 |
| GOTERM_CC_DIRECT | GO:0070062~extracellular exosome | 41 | 2645 | 4.12 × 10−4 |
| GOTERM_CC_DIRECT | GO:0016020~membrane | 37 | 2387 | 5.55 × 10−5 |
| GOTERM_CC_DIRECT | GO:0005576~extracellular region | 34 | 2193 | 1.15 × 10−6 |
| GOTERM_CC_DIRECT | GO:0030054~cell junction | 33 | 2129 | 1.23 × 10−20 |
| GOTERM_CC_DIRECT | GO:0005615~extracellular space | 28 | 1806 | 1.94 × 10−5 |
| KEGG_PATHWAY | hsa04080:Neuroactive ligand-receptor interaction | 51 | 3290 | 6.98 × 10−40 |
| KEGG_PATHWAY | hsa04024:cAMP signaling pathway | 26 | 1677 | 1.35 × 10−15 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ivanov, R.; Zamyatin, V.; Klimenko, A.; Matushkin, Y.; Savostyanov, A.; Lashin, S. Reconstruction and Analysis of Gene Networks of Human Neurotransmitter Systems Reveal Genes with Contentious Manifestation for Anxiety, Depression, and Intellectual Disabilities. Genes 2019, 10, 699. https://doi.org/10.3390/genes10090699
Ivanov R, Zamyatin V, Klimenko A, Matushkin Y, Savostyanov A, Lashin S. Reconstruction and Analysis of Gene Networks of Human Neurotransmitter Systems Reveal Genes with Contentious Manifestation for Anxiety, Depression, and Intellectual Disabilities. Genes. 2019; 10(9):699. https://doi.org/10.3390/genes10090699
Chicago/Turabian StyleIvanov, Roman, Vladimir Zamyatin, Alexandra Klimenko, Yury Matushkin, Alexander Savostyanov, and Sergey Lashin. 2019. "Reconstruction and Analysis of Gene Networks of Human Neurotransmitter Systems Reveal Genes with Contentious Manifestation for Anxiety, Depression, and Intellectual Disabilities" Genes 10, no. 9: 699. https://doi.org/10.3390/genes10090699
APA StyleIvanov, R., Zamyatin, V., Klimenko, A., Matushkin, Y., Savostyanov, A., & Lashin, S. (2019). Reconstruction and Analysis of Gene Networks of Human Neurotransmitter Systems Reveal Genes with Contentious Manifestation for Anxiety, Depression, and Intellectual Disabilities. Genes, 10(9), 699. https://doi.org/10.3390/genes10090699






