Meeting the Challenge of Eliminating Chronic Hepatitis B Infection
Abstract
:1. Background
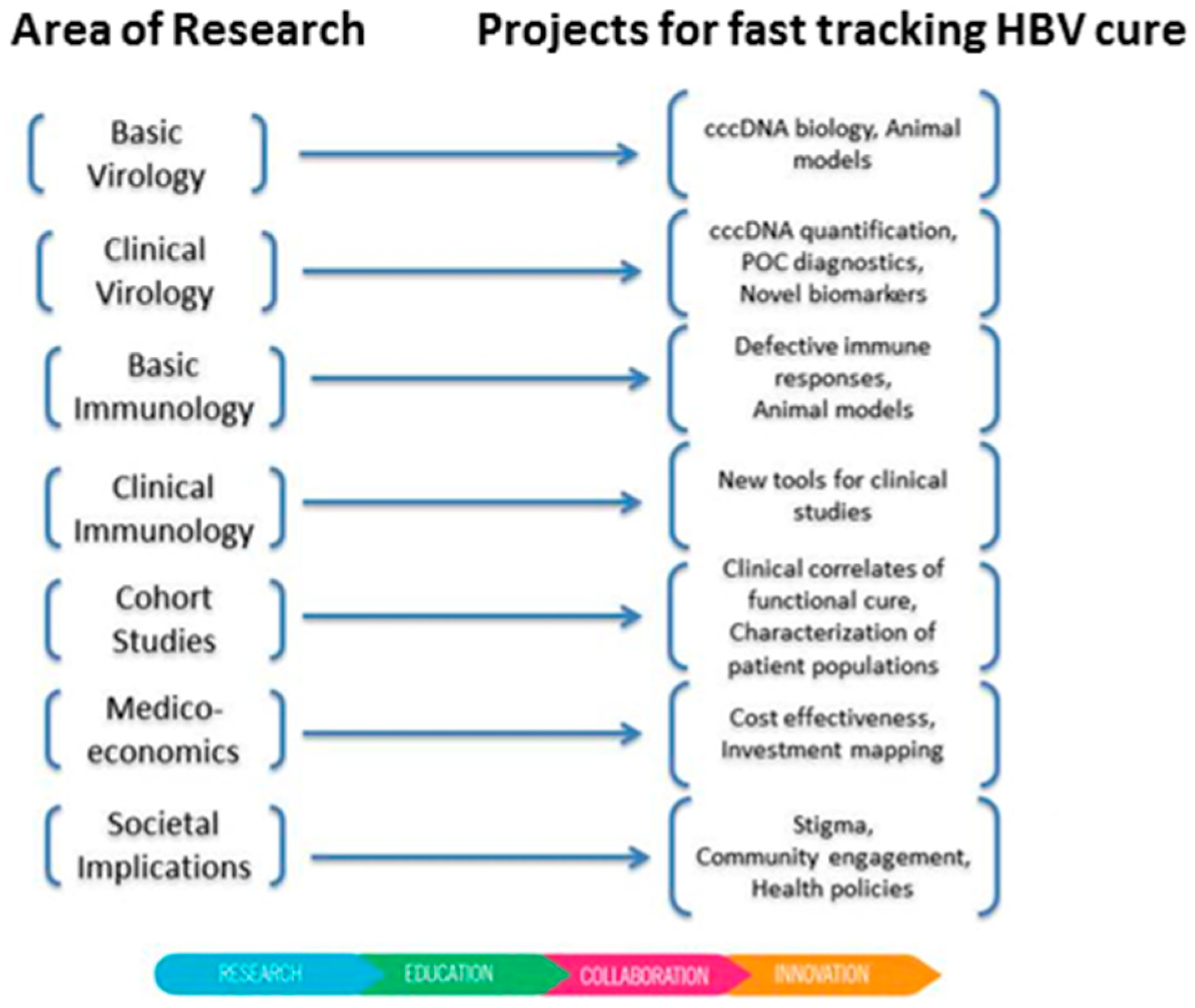
2. A New Global Initiative Promoting Hepatitis B Cure
3. Hepatitis B Virus Life Cycle
4. Barriers to Hepatitis B Virus Cure
5. Clinical Advances towards Hepatitis B Virus Cure
6. Further Challenges
7. Finding the Missing Millions
8. Conclusions
Funding
Conflicts of Interest
References
- World Health Organization. Hepatitis B Fact Sheet. Available online: http://www.who.int/mediacentre/factsheets/fs204/en/ (accessed on 10 December 2018).
- Revill, P.; Testoni, B.; Locarnini, S.; Zoulim, F. Global strategies are required to cure and eliminate HBV infection. Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 239–248. [Google Scholar] [CrossRef] [PubMed]
- Zeisel, M.B.; Lucifora, J.; Mason, W.S.; Sureau, C.; Beck, J.; Levrero, M.; Kann, M.; Knolle, P.A.; Benkirane, M.; Durantel, D.; et al. Towards an HBV cure: State-of-the-art and unresolved questions—Report of the ANRS workshop on HBV cure. Gut 2015, 64, 1314–1326. [Google Scholar] [CrossRef] [PubMed]
- Zoulim, F.; Testoni, B.; Lebosse, F. Kinetics of intrahepatic covalently closed circular DNA and serum hepatitis B surface antigen during antiviral therapy for chronic hepatitis B: Lessons from experimental and clinical studies. Clin. Gastroenterol. Hepatol. 2013, 11, 1011–1013. [Google Scholar] [CrossRef] [PubMed]
- Lok, A.S.; Zoulim, F.; Dusheiko, G.; Ghany, M.G. Hepatitis B cure: From discovery to regulatory approval. Hepatology 2017, 66, 1296–1313. [Google Scholar] [CrossRef] [PubMed]
- Kramvis, A. Genotypes and genetic variability of hepatitis B virus. Intervirology 2014, 57, 141–150. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.K.; Revill, P.A.; Ahn, S.H. HBV genotypes: Relevance to natural history, pathogenesis and treatment of chronic hepatitis B. Antivir. Ther. 2011, 16, 1169–1186. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization: Finding a Cure for Hepatitis B: Are We Close? Available online: https://protect-au.mimecast.com/s/HVSYCvl0E5unJwJWHQC44B?domain=who.int (accessed on 10 December 2018).
- Alter, H.; Block, T.M.; Brown, N.; Brownstein, A.; Brosgart, C.; Chang, K.M.; Chen, P.J.; Chisari, F.V.; Cohen, C.; El-Serag, H.; et al. A Research Agenda for Curing Chronic Hepatitis B Virus Infection. Hepatology 2017. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Zhong, G.; Xu, G.; He, W.; Jing, Z.; Gao, Z.; Huang, Y.; Qi, Y.; Peng, B.; Wang, H.; et al. Sodium taurocholate cotransporting polypeptide is a functional receptor for human hepatitis B and D virus. eLife 2012, 1, e00049. [Google Scholar] [CrossRef]
- Nassal, M. HBV cccDNA: Viral persistence reservoir and key obstacle for a cure of chronic hepatitis B. Gut 2015, 64, 1972–1984. [Google Scholar] [CrossRef]
- Newbold, J.E.; Xin, H.; Tencza, M.; Sherman, G.; Dean, J.; Bowden, S.; Locarnini, S. The covalently closed duplex form of the hepadnavirus genome exists in situ as a heterogeneous population of viral minichromosomes. J. Virol. 1995, 69, 3350–3357. [Google Scholar]
- Bock, C.T.; Schwinn, S.; Locarnini, S.; Fyfe, J.; Manns, M.P.; Trautwein, C.; Zentgraf, H. Structural organization of the hepatitis B virus minichromosome. J. Mol. Biol. 2001, 307, 183–196. [Google Scholar] [CrossRef] [PubMed]
- Zoulim, F.; Locarnini, S. Hepatitis B virus resistance to nucleos(t)ide analogues. Gastroenterology 2009, 137, 1593–1608. [Google Scholar] [CrossRef] [PubMed]
- Bayliss, J.; Yuen, L.; Rosenberg, G.; Wong, D.; Littlejohn, M.; Jackson, K.; Gaggar, A.; Kitrinos, K.M.; Subramanian, G.M.; Marcellin, P.; et al. Deep sequencing shows that HBV basal core promoter and precore variants reduce the likelihood of HBsAg loss following tenofovir disoproxil fumarate therapy in HBeAg-positive chronic hepatitis B. Gut 2016. [Google Scholar] [CrossRef] [PubMed]
- Soussan, P.; Pol, J.; Garreau, F.; Schneider, V.; Le Pendeven, C.; Nalpas, B.; Lacombe, K.; Bonnard, P.; Pol, S.; Kremsdorf, D. Expression of defective hepatitis B virus particles derived from singly spliced RNA is related to liver disease. J. Infect. Dis. 2008, 198, 218–225. [Google Scholar] [CrossRef] [PubMed]
- Soussan, P.; Garreau, F.; Zylberberg, H.; Ferray, C.; Brechot, C.; Kremsdorf, D. In vivo expression of a new hepatitis B virus protein encoded by a spliced RNA. J. Clin. Invest. 2000, 105, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Redelsperger, F.; Lekbaby, B.; Mandouri, Y.; Giang, E.; Duriez, M.; Desire, N.; Roque Afonso, A.M.; Brichler, S.; Dubreuil, P.; Dobrin, A.; et al. Production of hepatitis B defective particles is dependent on liver status. Virology 2012, 431, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Soussan, P.; Tuveri, R.; Nalpas, B.; Garreau, F.; Zavala, F.; Masson, A.; Pol, S.; Brechot, C.; Kremsdorf, D. The expression of hepatitis B spliced protein (HBSP) encoded by a spliced hepatitis B virus RNA is associated with viral replication and liver fibrosis. J. Hepatol. 2003, 38, 343–348. [Google Scholar] [CrossRef]
- Bayliss, J.; Lim, L.; Thompson, A.J.; Desmond, P.; Angus, P.; Locarnini, S.; Revill, P.A. Hepatitis B virus splicing is enhanced prior to development of hepatocellular carcinoma. J. Hepatol. 2013, 59, 1022–1028. [Google Scholar] [CrossRef]
- Pan, M.-H.; Hu, H.-H.; Mason, H.; Bayliss, J.; Littlejohn, M.; Lin, Y.-L.; Su, C.-Y.; Chiang, C.-T.; Chen, C.-J.; Locarnini, S.; et al. Hepatitis B splice variants are strongly associated with and are indeed predictive of hepatocellular carcinoma. J. Hepatol. 2018, 68, 474–475. [Google Scholar] [CrossRef]
- Wooddell, C.I.; Yuen, M.F.; Chan, H.L.; Gish, R.G.; Locarnini, S.A.; Chavez, D.; Ferrari, C.; Given, B.D.; Hamilton, J.; Kanner, S.B.; et al. RNAi-based treatment of chronically infected patients and chimpanzees reveals that integrated hepatitis B virus DNA is a source of HBsAg. Sci. Transl. Med. 2017, 9, eaan0241. [Google Scholar] [CrossRef]
- Thi, E.P.; Dhillon, A.P.; Ardzinski, A.; Bidirici-Ertekin, L.; Cobarrubias, K.D.; Cuconati, A.; Kondratowicz, A.S.; Kwak, K.; Li, A.H.L.; Miller, A.; et al. ARB-1740, a RNA Interference Therapeutic for Chronic Hepatitis B Infection. ACS Infect. Dis. 2018. [Google Scholar] [CrossRef] [PubMed]
- Klumpp, K.; Lam, A.M.; Lukacs, C.; Vogel, R.; Ren, S.; Espiritu, C.; Baydo, R.; Atkins, K.; Abendroth, J.; Liao, G.; et al. High-resolution crystal structure of a hepatitis B virus replication inhibitor bound to the viral core protein. Proc. Natl. Acad. Sci. USA 2015, 112, 15196–15201. [Google Scholar] [CrossRef] [PubMed]
- Klumpp, K.; Shimada, T.; Allweiss, L.; Volz, T.; Lutgehetmann, M.; Hartman, G.; Flores, O.A.; Lam, A.M.; Dandri, M. Efficacy of NVR 3-778, Alone and In Combination With Pegylated Interferon, vs. Entecavir In uPA/SCID Mice With Humanized Livers and HBV Infection. Gastroenterology 2018, 154, 652–662. [Google Scholar] [CrossRef]
- Lam, A.M.; Espiritu, C.; Vogel, R.; Ren, S.; Lau, V.; Kelly, M.; Kuduk, S.D.; Hartman, G.D.; Flores, O.A.; Klumpp, K. Preclinical Characterization of NVR 3-778, a First-in-Class Capsid Assembly Modulator against Hepatitis B Virus. Antimicrob. Agents Chemother. 2019, 63, e01734-18. [Google Scholar] [CrossRef] [PubMed]
- Bazinet, M.; Pantea, V.; Cebotarescu, V.; Cojuhari, L.; Jimbei, P.; Albrecht, J.; Schmid, P.; Le Gal, F.; Gordien, E.; Krawczyk, A.; et al. Safety and efficacy of REP 2139 and pegylated interferon alfa-2a for treatment-naive patients with chronic hepatitis B virus and hepatitis D virus co-infection (REP 301 and REP 301-LTF): A non-randomised, open-label, phase 2 trial. Lancet Gastroenterol. Hepatol. 2017, 2, 877–889. [Google Scholar] [CrossRef]
- Roehl, I.; Seiffert, S.; Brikh, C.; Quinet, J.; Jamard, C.; Dorfler, N.; Lockridge, J.A.; Cova, L.; Vaillant, A. Nucleic Acid Polymers with Accelerated Plasma and Tissue Clearance for Chronic Hepatitis B Therapy. Mol. Nucleic Acids 2017, 8, 1–12. [Google Scholar] [CrossRef]
- Agarwal, K.; Brunetto, M.; Seto, W.K.; Lim, Y.S.; Fung, S.; Marcellin, P.; Ahn, S.H.; Izumi, N.; Chuang, W.L.; Bae, H.; et al. 96weeks treatment of tenofovir alafenamide vs. tenofovir disoproxil fumarate for hepatitis B virus infection. J. Hepatol. 2018, 68, 672–681. [Google Scholar] [CrossRef]
- Cathcart, A.L.; Chan, H.L.; Bhardwaj, N.; Liu, Y.; Marcellin, P.; Pan, C.Q.; Shalimar; Buti, M.; Cox, S.; Parhy, B.; et al. No Resistance to Tenofovir Alafenamide Detected through 96 Weeks of Treatment in Patients with Chronic Hepatitis B Infection. Antimicrob. Agents Chemother. 2018, 62. [Google Scholar] [CrossRef]
- Petersen, J.; Dandri, M.; Mier, W.; Lutgehetmann, M.; Volz, T.; von Weizsacker, F.; Haberkorn, U.; Fischer, L.; Pollok, J.M.; Erbes, B.; et al. Prevention of hepatitis B virus infection in vivo by entry inhibitors derived from the large envelope protein. Nat. Biotechnol. 2008, 26, 335–341. [Google Scholar] [CrossRef]
- Long, K.R.; Lomonosova, E.; Li, Q.; Ponzar, N.L.; Villa, J.A.; Touchette, E.; Rapp, S.; Liley, R.M.; Murelli, R.P.; Grigoryan, A.; et al. Efficacy of hepatitis B virus ribonuclease H inhibitors, a new class of replication antagonists, in FRG human liver chimeric mice. Antivir. Res. 2018, 149, 41–47. [Google Scholar] [CrossRef]
- Schiwon, M.; Ehrke-Schulz, E.; Oswald, A.; Bergmann, T.; Michler, T.; Protzer, U.; Ehrhardt, A. One-Vector System for Multiplexed CRISPR/Cas9 against Hepatitis B Virus cccDNA Utilizing High-Capacity Adenoviral Vectors. Mol. Therapy. Nucleic Acids 2018, 12, 242–253. [Google Scholar] [CrossRef]
- Song, J.; Zhang, X.; Ge, Q.; Yuan, C.; Chu, L.; Liang, H.F.; Liao, Z.; Liu, Q.; Zhang, Z.; Zhang, B. CRISPR/Cas9-mediated knockout of HBsAg inhibits proliferation and tumorigenicity of HBV-positive hepatocellular carcinoma cells. J. Cell. Biochem. 2018, 119, 8419–8431. [Google Scholar] [CrossRef]
- Joung, J.K.; Sander, J.D. TALENs: A widely applicable technology for targeted genome editing. Nat. Reviews. Mol. Cell Biol. 2013, 14, 49–55. [Google Scholar] [CrossRef]
- Bloom, K.; Ely, A.; Mussolino, C.; Cathomen, T.; Arbuthnot, P. Inactivation of hepatitis B virus replication in cultured cells and in vivo with engineered transcription activator-like effector nucleases. Mol. Ther. 2013, 21, 1889–1897. [Google Scholar] [CrossRef]
- Wieland, S.F.; Chisari, F.V. Stealth and cunning: Hepatitis B and hepatitis C viruses. J. Virol. 2005, 79, 9369–9380. [Google Scholar] [CrossRef]
- Cheng, X.; Xia, Y.; Serti, E.; Block, P.D.; Chung, M.; Chayama, K.; Rehermann, B.; Liang, T.J. Hepatitis B virus evades innate immunity of hepatocytes but activates cytokine production by macrophages. Hepatology 2017, 66, 1779–1793. [Google Scholar] [CrossRef] [PubMed]
- Wieland, D.; Hofmann, M.; Thimme, R. Overcoming CD8+ T-Cell Exhaustion in Viral Hepatitis: Lessons from the Mouse Model and Clinical Perspectives. Dig. Dis. 2017, 35, 334–338. [Google Scholar] [CrossRef]
- Burton, A.R.; Pallett, L.J.; McCoy, L.E.; Suveizdyte, K.; Amin, O.E.; Swadling, L.; Alberts, E.; Davidson, B.R.; Kennedy, P.T.; Gill, U.S.; et al. Circulating and intrahepatic antiviral B cells are defective in hepatitis B. J. Clin. Invest. 2018, 128, 4588–4603. [Google Scholar] [CrossRef]
- Hepatitis B Foundation: Drug Watch. Available online: http://www.hepb.org/treatment-and-management/drug-watch (accessed on 22 January 2019).
- ClinicalTrials.gov: Home. Available online: https://clinicaltrials.gov (accessed on 22 January 2019).
- World Hepatitis Alliance. Available online: http://www.worldhepatitisalliance.org/ (accessed on 10 December 2018).
- CEVHAP. Available online: http://www.cevhap.org /index.php/en/ (accessed on 10 December 2018).
- El-Khoueiry, A.B.; Sangro, B.; Yau, T.; Crocenzi, T.S.; Kudo, M.; Hsu, C.; Kim, T.Y.; Choo, S.P.; Trojan, J.; Welling, T.H.R.; et al. Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): An open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet 2017, 389, 2492–2502. [Google Scholar] [CrossRef]
- Kramvis, A.; Kew, M.; Francois, G. Hepatitis B virus genotypes. Vaccine 2005, 23, 2409–2423. [Google Scholar] [CrossRef]
- Kramvis, A.; Kew, M.C. Relationship of genotypes of hepatitis B virus to mutations, disease progression and response to antiviral therapy. J. Viral Hepat. 2005, 12, 456–464. [Google Scholar] [CrossRef]
- Sozzi, V.; Walsh, R.; Littlejohn, M.; Colledge, D.; Jackson, K.; Warner, N.; Yuen, L.; Locarnini, S.A.; Revill, P.A. In Vitro Studies Show that Sequence Variability Contributes to Marked Variation in Hepatitis B Virus Replication, Protein Expression, and Function Observed across Genotypes. J. Virol. 2016, 90, 10054–10064. [Google Scholar] [CrossRef]
- Kew, M.C.; Kramvis, A.; Yu, M.C.; Arakawa, K.; Hodkinson, J. Increased hepatocarcinogenic potential of hepatitis B virus genotype A in Bantu-speaking sub-saharan Africans. J. Med. Virol. 2005, 75, 513–521. [Google Scholar] [CrossRef]
- Kramvis, A.; Kew, M.C. Epidemiology of hepatitis B virus in Africa, its genotypes and clinical associations of genotypes. Hepatol. Res. 2007, 37, S9–S19. [Google Scholar] [CrossRef] [PubMed]
- Kramvis, A.; Kew, M.C. Molecular characterization of subgenotype A1 (subgroup Aa) of hepatitis B virus. Hepatol. Res. 2007, 37, S27–S32. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.; Littlejohn, M.; Locarnini, S.A.; Whiting, S.; Hajkowicz, K.; Cowie, B.C.; Bowden, D.S.; Tong, S.Y.; Davis, J.S. Molecular epidemiology of hepatitis B in the Indigenous people of northern Australia. J. Gastroenterol. Hepatol. 2013, 28, 1234–1241. [Google Scholar] [CrossRef] [PubMed]
- Littlejohn, M.; Davies, J.; Yuen, L.; Edwards, R.; Sozzi, T.; Jackson, K.; Cowie, B.; Tong, S.; Davis, J.; Locarnini, S. Molecular virology of hepatitis B virus, sub-genotype C4 in northern Australian Indigenous populations. J. Med. Virol. 2014, 86, 695–706. [Google Scholar] [CrossRef]
- Shen, F.; Li, Y.; Wang, Y.; Sozzi, V.; Revill, P.A.; Liu, J.; Gao, L.; Yang, G.; Lu, M.; Sutter, K.; et al. Hepatitis B virus sensitivity to interferon-alpha in hepatocytes is more associated with cellular interferon response than with viral genotype. Hepatology 2018, 67, 1237–1252. [Google Scholar] [CrossRef]
- Marcellin, P.; Heathcote, E.J.; Buti, M.; Gane, E.; de Man, R.A.; Krastev, Z.; Germanidis, G.; Lee, S.S.; Flisiak, R.; Kaita, K.; et al. Tenofovir disoproxil fumarate versus adefovir dipivoxil for chronic hepatitis B. N. Engl. J. Med. 2008, 359, 2442–2455. [Google Scholar] [CrossRef]
- The World Alliance Hepatitis. Available online: http://www.worldhepatitisalliance.org/find-missing-millions (accessed on 10 December 2018).
- Polaris Observatory, C. Global prevalence, treatment, and prevention of hepatitis B virus infection in 2016: A modelling study. Lancet Gastroenterol. Hepatol. 2018, 3, 383–403. [Google Scholar]
- HIV, Viral Hepatitis and Sexually Transmissible Infections in Australia: Annual Surveillance Report 2018. Available online: https://kirby.unsw.edu.au/report/hiv-viral-hepatitis-and-sexually-transmissible-infections-australia-annual-surveillance (accessed on 10 December 2018).
- World Health Organization, Public Reports of WHO prequalified IVDs. Available online: https://www.who.int/diagnostics_laboratory/evaluations/pq-list/hiv-rdts/public_report/en/ (accessed on 10 December 2018).
- International HBV Meeting. Available online: www.hbvmeeting.org/ (accessed on 10 December 2018).
- World Health Organization. Available online: www.who.int (accessed on 10 December 2018).
| Approach | Name/Type | Company | Status |
|---|---|---|---|
| Silencing HBV RNAs | RNAi gene silencer (1.0) | Arrowhead Pharma | NCT03365947 (R) |
| HBV Locked Nucleic Acid (LNA) RO7062931 | Roche | NCT03038113 (R) NCT03505190 (R) | |
| SiRNA VIR-2218 | Alnylam and Vir Biotech | NCT03672188 (R) | |
| Liquid nano-particle (LNP) RNAi (ARB-1462) | Arbutus Biopharma | Phase 2 (IMPACT study) | |
| Antisense Molecules | IONIS-HBVRx (GSK3228836) (GSK33389404) | IONIS/GSK | NCT02981602 (R) NCT03020745 (R) |
| Entry inhibitor | Myrcludex B | Myr-pharma | NCT02888106 Recruiting hepatitis delta virus HDV/HBV coinfected patients |
| Capsid Inhibitors | GLS4 | HEC Pharma | NCT03638076 (R) |
| JNJ 56136379 | Janssen Sciences | NCT03439488 (R) NCT03361956 (R) | |
| ABI-H0731 | Assembly Biosciences | NCT03714152 (R) | |
| RO7049389 | Roche | NCT 02952924 (R) NCT 03570658 (R) NCT 03717064 (A) | |
| AB-506 | Arbutus Biopharma | Phase 1a studies completed | |
| HBsAg Inhibitors | REP 2139/2165 | Replicor, Canada | NCT02565719 (A) NCT02876419 (A) |
| TLR7 Antagonist | JNJ-64794964 (AL-034) | Janssen Sciences | NCT03285620 (R) |
| RO7020531 | Roche | NCT02956850 (R) NCT03530917 (R) | |
| TLR8 Antagonist | GS-9688 | Gilead, USA | NCT03615066 (R) |
| Innate Immune Activators | Inarigavir RIG-I agonist (also an HBV replication inhibitor). | Springbank Pharmaceuticals | NCT02751996 (R) |
| Immune Therapy | HBsAg monoclonal antibody | Green Cross | NCT03519113 (R) |
| Therapeutic DNA Vaccine | JNJ-64300535 | Janssen Sciences | NCT03463369 (R) |
| Undisclosed | RO7239958 | Roche | NCT03762681 Not yet recruiting |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Revill, P.A.; Penicaud, C.; Brechot, C.; Zoulim, F. Meeting the Challenge of Eliminating Chronic Hepatitis B Infection. Genes 2019, 10, 260. https://doi.org/10.3390/genes10040260
Revill PA, Penicaud C, Brechot C, Zoulim F. Meeting the Challenge of Eliminating Chronic Hepatitis B Infection. Genes. 2019; 10(4):260. https://doi.org/10.3390/genes10040260
Chicago/Turabian StyleRevill, Peter A., Capucine Penicaud, Christian Brechot, and Fabien Zoulim. 2019. "Meeting the Challenge of Eliminating Chronic Hepatitis B Infection" Genes 10, no. 4: 260. https://doi.org/10.3390/genes10040260
APA StyleRevill, P. A., Penicaud, C., Brechot, C., & Zoulim, F. (2019). Meeting the Challenge of Eliminating Chronic Hepatitis B Infection. Genes, 10(4), 260. https://doi.org/10.3390/genes10040260





