Assessing Global DNA Methylation Changes Associated with Plasticity in Seven Highly Inbred Lines of Snapdragon Plants (Antirrhinum majus)
Abstract
:1. Introduction
2. Material and Methods
2.1. Plant Material
2.2. Experimental Design
2.3. Phenotypic Measurements
2.4. Global DNA Methylation (%mC)
2.5. Statistical Analysis
3. Results
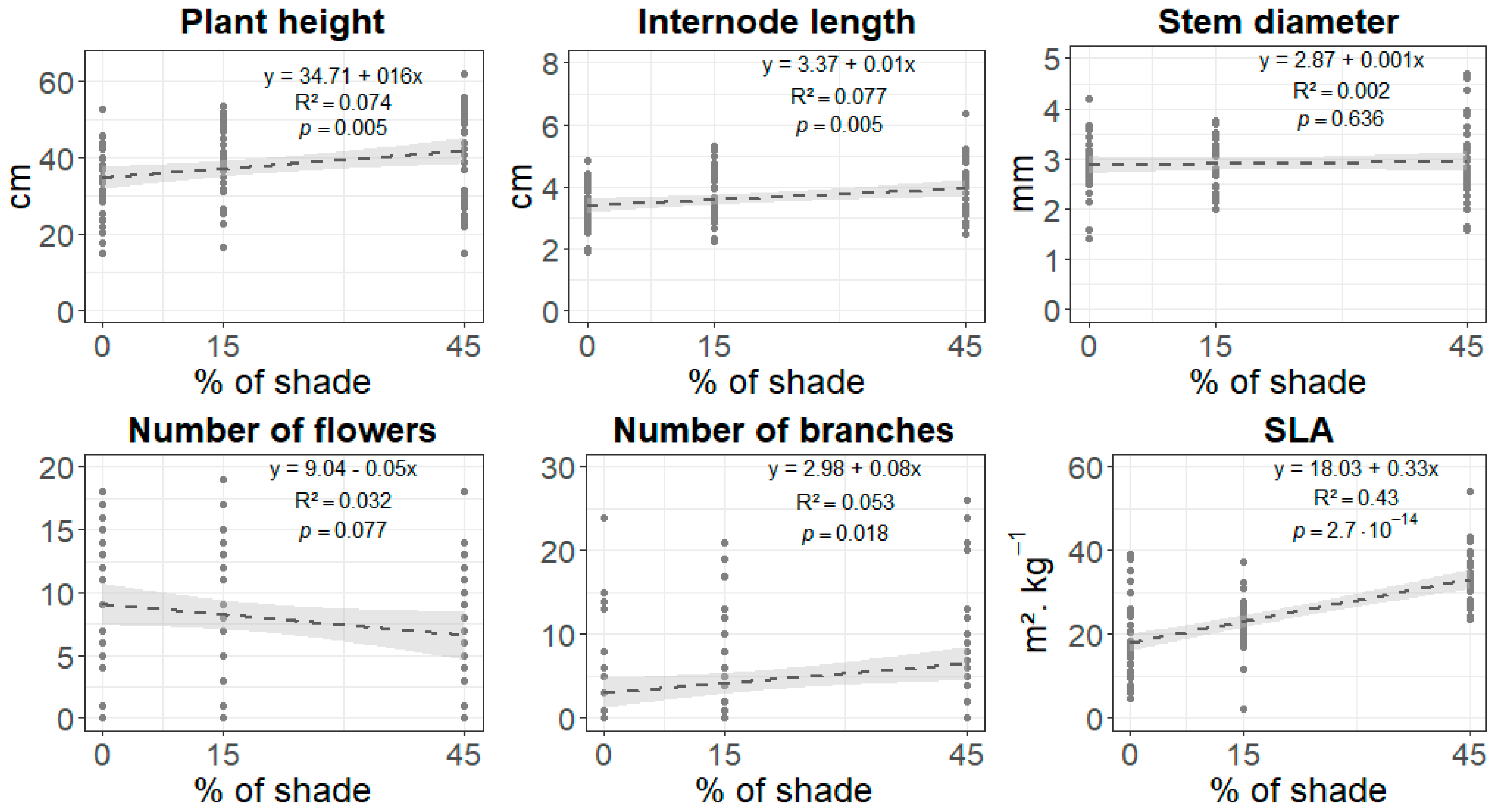
3.1. Phenotypic Response to Shade
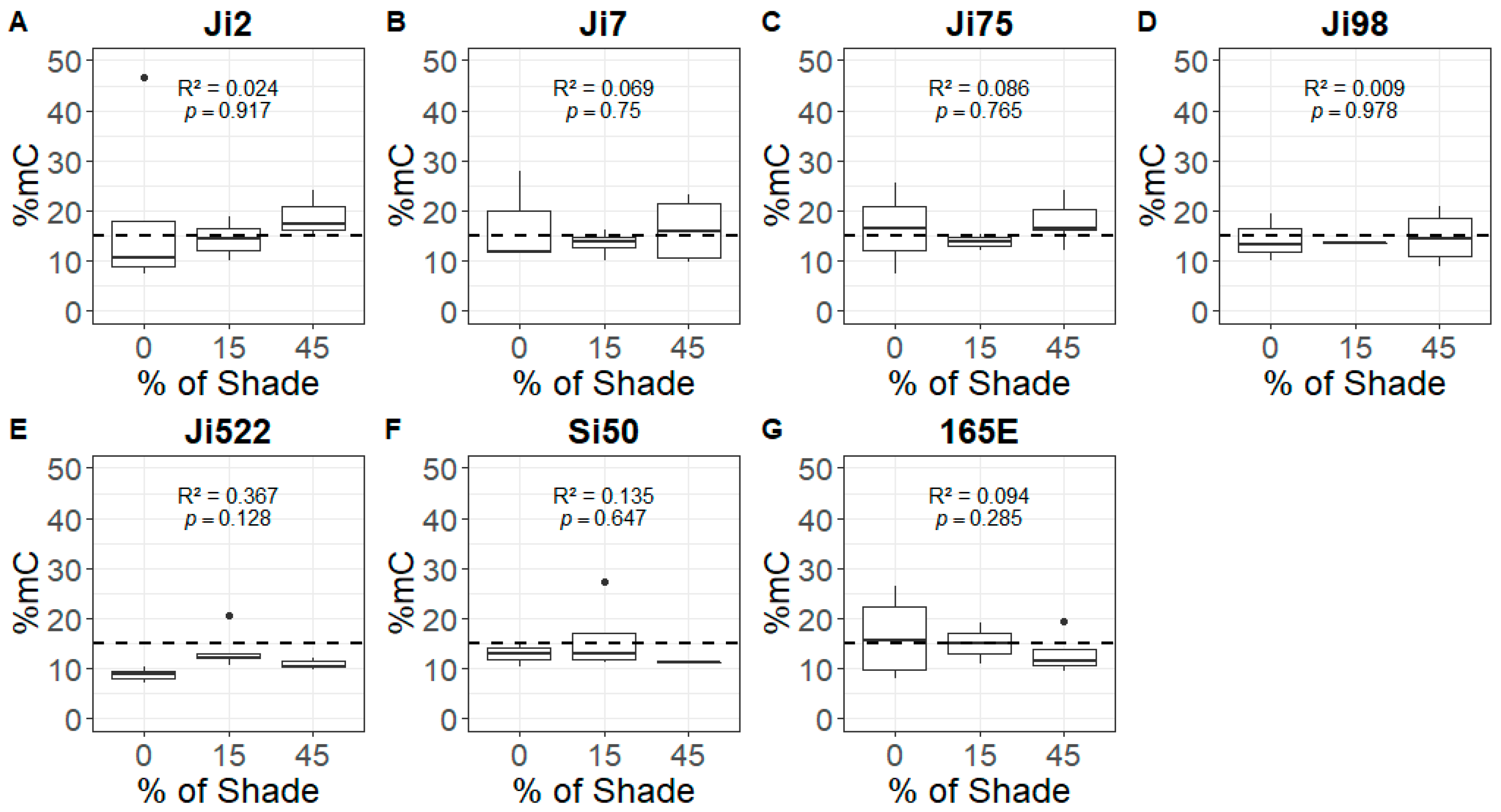
3.2. Relationship between Shade and Global DNA Methylation
3.3. Relationship between Traits and Global DNA Methylation
3.4. Relationship between the Slopes of the Reaction Norms and the Global DNA Methylation Changes
4. Discussion
4.1. Phenotypic Shade Avoidance Response
4.2. Global DNA Methylation
4.3. The Link between Phenotypic Plasticity and Epigenetic Changes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bradshaw, A.D. Evolutionary significance of phenotypic plasticity in plants. In Advances in Genetics; Caspari, E.W., Thoday, J.M., Eds.; Academic Press: New York, NY, USA, 1965; Volume 13, pp. 115–155. [Google Scholar]
- Pigliucci, M. Phenotypic Plasticity: Beyond Nature and Nurture; Johns Hopkins University Press: Baltimore, MD, USA, 2001. [Google Scholar]
- West-Eberhard, M.J. Phenotypic accommodation: Adaptive innovation due to developmental plasticity. J. Exp. Zoolog. B Mol. Dev. Evol. 2005, 304, 610–618. [Google Scholar] [CrossRef] [PubMed]
- Ghalambor, C.K.; McKay, J.K.; Carroll, S.P.; Reznick, D.N. Adaptive versus non-adaptive phenotypic plasticity and the potential for contemporary adaptation in new environments. Funct. Ecol. 2007, 21, 394–407. [Google Scholar] [CrossRef]
- Schmid, M.; Guillaume, F. The role of phenotypic plasticity on population differentiation. Heredity 2017, 119, 214–225. [Google Scholar] [CrossRef] [PubMed]
- Sultan, S.E. Plant developmental responses to the environment: Eco-devo insights. Curr. Opin. Plant Biol. 2010, 13, 96–101. [Google Scholar] [CrossRef] [PubMed]
- Baulcombe, D.C.; Dean, C. Epigenetic regulation in plant responses to the environment. Cold Spring Harb. Perspect. Biol. 2014, 6, a019471. [Google Scholar] [CrossRef]
- Geng, Y.; Gao, L.; Yang, J. Epigenetic flexibility underlying phenotypic plasticity. In Progress in Botany; Springer: Berlin, Germany, 2013; pp. 153–163. ISBN 978-3-642-30966-3. [Google Scholar]
- Nicotra, A.B.; Atkin, O.K.; Bonser, S.P.; Davidson, A.M.; Finnegan, E.J.; Mathesius, U.; Poot, P.; Purugganan, M.D.; Richards, C.L.; Valladares, F.; et al. Plant phenotypic plasticity in a changing climate. Trends Plant Sci. 2010, 15, 684–692. [Google Scholar] [CrossRef]
- Rey, O.; Danchin, E.; Mirouze, M.; Loot, C.; Blanchet, S. Adaptation to Global Change: A Transposable element-epigenetics perspective. Trends Ecol. Evol. 2016, 31, 514–526. [Google Scholar] [CrossRef] [PubMed]
- Richards, C.L.; Alonso, C.; Becker, C.; Bossdorf, O.; Bucher, E.; Colomé-Tatché, M.; Durka, W.; Engelhardt, J.; Gaspar, B.; Gogol-Döring, A.; et al. Ecological plant epigenetics: Evidence from model and non-model species, and the way forward. Ecol. Lett. 2017, 20, 1576–1590. [Google Scholar] [CrossRef]
- Meyer, P. Chapter One—Epigenetics—A historical perspective. In Advances in Botanical Research; Mirouze, M., Bucher, E., Gallusci, P., Eds.; Plant Epigenetics Coming of Age for Breeding Applications; Academic Press: New York, NY, USA, 2018; Volume 88, pp. 1–19. [Google Scholar]
- Waddington, C.H. The epigenotype. Endeavour 1942, 1, 18–20. [Google Scholar] [CrossRef]
- Eichten, S.R.; Schmitz, R.J.; Springer, N.M. Epigenetics: Beyond chromatin modifications and complex genetic regulation. Plant Physiol. 2014, 165, 933–947. [Google Scholar] [CrossRef]
- Lanciano, S.; Mirouze, M. Transposable elements: All mobile, all different, some stress responsive, some adaptive? Curr. Opin. Genet. Dev. 2018, 49, 106–114. [Google Scholar] [CrossRef]
- Maeji, H.; Nishimura, T. Chapter Two - Epigenetic mechanisms in plants. In Advances in Botanical Research; Plant Epigenetics Coming of Age for Breeding Applications; Mirouze, M., Bucher, E., Gallusci, P., Eds.; Academic Press: New York, NY, USA, 2018; Volume 88, pp. 21–47. [Google Scholar]
- Mirouze, M.; Paszkowski, J. Epigenetic contribution to stress adaptation in plants. Curr. Opin. Plant Biol. 2011, 14, 267–274. [Google Scholar] [CrossRef]
- Zentner, G.E.; Henikoff, S. Regulation of nucleosome dynamics by histone modifications. Nat. Struct. Mol. Biol. 2013, 20, 259–266. [Google Scholar] [CrossRef]
- Herrera, C.M.; Bazaga, P. Epigenetic correlates of plant phenotypic plasticity: DNA methylation differs between prickly and nonprickly leaves in heterophyllous Ilex aquifolium (Aquifoliaceae) trees. Bot. J. Linn. Soc. 2013, 171, 441–452. [Google Scholar] [CrossRef]
- Richards, C.L.; Bossdorf, O.; Pigliucci, M. What role does heritable epigenetic variation play in phenotypic evolution? BioScience 2010, 60, 232–237. [Google Scholar] [CrossRef]
- Chinnusamy, V.; Zhu, J.-K. Epigenetic regulation of stress responses in plants. Genome Stud. Mol. Genet. 2009, 12, 133–139. [Google Scholar] [CrossRef]
- Kinoshita, T.; Seki, M. Epigenetic memory for stress response and adaptation in plants. Plant Cell Physiol. 2014, 55, 1859–1863. [Google Scholar] [CrossRef]
- Liu, J.; Feng, L.; Li, J.; He, Z. Genetic and epigenetic control of plant heat responses. Front. Plant Sci. 2015, 6, 267. [Google Scholar] [CrossRef]
- Banerjee, A.; Wani, S.H.; Roychoudhury, A. Epigenetic control of plant cold responses. Front. Plant Sci. 2017, 8, 1643. [Google Scholar] [CrossRef]
- Espinas, N.A.; Saze, H.; Saijo, Y. Epigenetic control of defense signaling and priming in plants. Front. Plant Sci. 2016, 7, 1201. [Google Scholar] [CrossRef]
- Ballare, C.L.; Pierik, R. The shade-avoidance syndrome: Multiple signals and ecological consequences. Plant Cell Environ. 2017, 40, 2530–2543. [Google Scholar] [CrossRef]
- Pierik, R.; Testerink, C. The art of being flexible: How to escape from shade, salt, and drought. Plant Physiol. 2014, 166, 5–22. [Google Scholar] [CrossRef]
- Batschauer, A. Light perception in higher plants. Cell. Mol. Life Sci. 1999, 55, 153–166. [Google Scholar] [CrossRef] [PubMed]
- Pedmale, U.V.; Celaya, R.B.; Liscum, E. Phototropism: Mechanism and outcomes. Arab. Book 2010, 8, e0125. [Google Scholar] [CrossRef]
- Sessa, G.; Carabelli, M.; Possenti, M.; Morelli, G.; Ruberti, I. Multiple pathways in the control of the shade avoidance response. Plants Basel Switz. 2018, 7, 102. [Google Scholar] [CrossRef] [PubMed]
- Franklin, K.A.; Whitelam, G.C. Phytochromes and shade-avoidance responses in plants. Ann. Bot. 2005, 96, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Morelli, G.; Ruberti, I. Shade avoidance responses. Driving auxin along lateral routes. Plant Physiol. 2000, 122, 621–626. [Google Scholar] [CrossRef] [PubMed]
- Pierik, R.; Cuppens, M.L.; Voesenek, L.A.; Visser, E.J. Interactions between ethylene and gibberellins in phytochrome-mediated shade avoidance responses in tobacco. Plant Physiol. 2004, 136, 2928–2936. [Google Scholar] [CrossRef]
- Ruberti, I.; Sessa, G.; Ciolfi, A.; Possenti, M.; Carabelli, M.; Morelli, G. Plant adaptation to dynamically changing environment: The shade avoidance response. Biotechnol. Adv. 2012, 30, 1047–1058. [Google Scholar] [CrossRef]
- Bell, D.L.; Galloway, L.F. Plasticity to neighbour shade: Fitness consequences and allometry. Funct. Ecol. 2007, 21, 1146–1153. [Google Scholar] [CrossRef]
- Van Kleunen, M.; Fischer, M. Adaptive evolution of plastic foraging responses in a clonal plant. Ecology 2001, 82, 3309–3319. [Google Scholar] [CrossRef]
- Schmitt, J.; Stinchcombe, J.R.; Heschel, M.S.; Huber, H. The adaptive evolution of plasticity: Phytochrome-mediated shade avoidance responses. Integr. Comp. Biol. 2003, 43, 459–469. [Google Scholar] [CrossRef]
- Haddad, N.M.; Brudvig, L.A.; Clobert, J.; Davies, K.F.; Gonzalez, A.; Holt, R.D.; Lovejoy, T.E.; Sexton, J.O.; Austin, M.P.; Collins, C.D.; et al. Habitat fragmentation and its lasting impact on Earth’s ecosystems. Sci. Adv. 2015, 1, e1500052. [Google Scholar] [CrossRef]
- Poyatos, R.; Latron, J.; Llorens, P. Land use and land cover change after agricultural abandonment. Mt. Res. Dev. 2003, 23, 362–368. [Google Scholar] [CrossRef]
- Pujol, B.; Mühlen, G.; Garwood, N.; Horoszowski, Y.; Douzery, E.J.P.; McKey, D. Evolution under domestication: Contrasting functional morphology of seedlings in domesticated cassava and its closest wild relatives. New Phytol. 2005, 166, 305–318. [Google Scholar] [CrossRef] [PubMed]
- Sessa, G.; Carabelli, M.; Sassi, M.; Ciolfi, A.; Possenti, M.; Mittempergher, F.; Becker, J.; Morelli, G.; Ruberti, I. A dynamic balance between gene activation and repression regulates the shade avoidance response in Arabidopsis. Genes Dev. 2005, 19, 2811–2815. [Google Scholar] [CrossRef]
- Donohue, K.; Schmitt, J. The genetic architecture of plasticity to density in Impatiens capensis. Evolution 1999, 53, 1377–1386. [Google Scholar] [CrossRef]
- Galloway, L.F.; Etterson, J.R. Transgenerational plasticity is adaptive in the wild. Science 2007, 318, 1134. [Google Scholar] [CrossRef] [PubMed]
- Huber, H.; Kane, N.C.; Heschel, M.S.; von Wettberg, E.J.; Banta, J.; Leuck, A.-M.; Schmitt, J. Frequency and microenvironmental pattern of selection on plastic shade-avoidance traits in a natural population of Impatiens capensis. Am. Nat. 2004, 163, 548–563. [Google Scholar] [CrossRef]
- Peng, M.; Li, Z.; Zhou, N.; Ma, M.; Jiang, Y.; Dong, A.; Shen, W.-H.; Li, L. Linking PHYTOCHROME-INTERACTING FACTOR to histone modification in plant shade avoidance. Plant Physiol. 2018, 176, 1341–1351. [Google Scholar] [CrossRef]
- Tatra, G.S.; Miranda, J.; Chinnappa, C.C.; Reid, D.M. Effect of light quality and 5-azacytidine on genomic methylation and stem elongation in two ecotypes of Stellaria longipes. Physiol. Plant. 2000, 109, 313–321. [Google Scholar] [CrossRef]
- Schwarz-Sommer, Z.; Davies, B.; Hudson, A. An everlasting pioneer: The story of Antirrhinum research. Nat. Rev. Genet. 2003, 4, 655. [Google Scholar] [CrossRef]
- Khattak, A.M.; Pearson, S. Light quality and temperature effects on Antirrhinum growth and development. J. Zhejiang Univ. Sci. B 2005, 6, 119–124. [Google Scholar] [CrossRef]
- Munir, M.; Jamil, M.; Baloch, J.; Khattak, K.R. Impact of light intensity on flowering time and plant quality of Antirrhinum majus L. cultivar Chimes White. J. Zhejiang Univ. Sci. A 2004, 5, 400–405. [Google Scholar] [CrossRef]
- Khimoun, A.; Burrus, M.; Andalo, C.; Liu, Z.-L.; Vicédo-Cazettes, C.; Thébaud, C.; Pujol, B. Locally asymmetric introgressions between subspecies suggest circular range expansion at the Antirrhinum majus global scale. J. Evol. Biol. 2011, 24, 1433–1441. [Google Scholar] [CrossRef] [PubMed]
- Pujol, B.; Archambeau, J.; Bontemps, A.; Lascoste, M.; Marin, S.; Meunier, A. Mountain landscape connectivity and subspecies appurtenance shape genetic differentiation in natural plant populations of the snapdragon (Antirrhinum majus L.). Bot. Lett. 2017, 164, 111–119. [Google Scholar] [CrossRef]
- Tavares, H.; Whibley, A.; Field, D.L.; Bradley, D.; Couchman, M.; Copsey, L.; Elleouet, J.; Burrus, M.; Andalo, C.; Li, M.; et al. Selection and gene flow shape genomic islands that control floral guides. Proc. Natl. Acad. Sci. USA 2018, 115, 11006–11011. [Google Scholar] [CrossRef]
- Khimoun, A.; Cornuault, J.; Burrus, M.; Pujol, B.; Thebaud, C.; Andalo, C. Ecology predicts parapatric distributions in two closely related Antirrhinum majus subspecies. Evolut. Ecol. 2013, 27, 51–64. [Google Scholar] [CrossRef]
- Yong, W.-S.; Hsu, F.-M.; Chen, P.-Y. Profiling genome-wide DNA methylation. Epigenetics Chromatin 2016, 9, 26. [Google Scholar] [CrossRef]
- Li, M.; Zhang, D.; Gao, Q.; Luo, Y.; Zhang, H.; Ma, B.; Chen, C.; Whibley, A.; Zhang, Y.; Cao, Y.; et al. Genome structure and evolution of Antirrhinum majus L. Nat. Plants 2019, 5, 174. [Google Scholar] [CrossRef]
- Van Gurp, T.P.; Wagemaker, N.C.A.M.; Wouters, B.; Vergeer, P.; Ouborg, J.N.J.; Verhoeven, K.J.F. epiGBS: Reference-free reduced representation bisulfite sequencing. Nat. Methods 2016, 13, 322–324. [Google Scholar] [CrossRef]
- Trucchi, E.; Mazzarella, A.B.; Gilfillan, G.D.; Lorenzo, M.T.; Schonswetter, P.; Paun, O. BsRADseq: Screening DNA methylation in natural populations of non-model species. Mol. Ecol. 2016, 25, 1697–1713. [Google Scholar] [CrossRef] [PubMed]
- Schield, D.R.; Walsh, M.R.; Card, D.C.; Andrew, A.L.; Adams, R.H.; Castoe, T.A. EpiRADseq: Scalable analysis of genomewide patterns of methylation using next-generation sequencing. Methods Ecol. Evol. 2016, 7, 60–69. [Google Scholar] [CrossRef]
- Weber, M.; Davies, J.J.; Wittig, D.; Oakeley, E.J.; Haase, M.; Lam, W.L.; Schübeler, D. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat. Genet. 2005, 37, 853. [Google Scholar] [CrossRef]
- Alonso, C.; Perez, R.; Bazaga, P.; Herrera, C.M. Global DNA cytosine methylation as an evolving trait: Phylogenetic signal and correlated evolution with genome size in angiosperms. Front. Genet. 2015, 6, 4. [Google Scholar] [CrossRef]
- Alonso, C.; Balao, F.; Bazaga, P.; Pérez, R. Epigenetic contribution to successful polyploidizations: Variation in global cytosine methylation along an extensive ploidy series in Dianthus broteri (Caryophyllaceae). New Phytol. 2016, 212, 571–576. [Google Scholar] [CrossRef] [PubMed]
- Lambé, P.; Mutambel, H.S.N.; Fouché, J.-G.; Deltour, R.; Foidart, J.-M.; Gaspar, T. DNA methylation as a key process in regulation of organogenic totipotency and plant neoplastic progression? Vitro Cell. Dev. Biol. Plant 1997, 33, 155–162. [Google Scholar] [CrossRef]
- Sow, M.D.; Segura, V.; Chamaillard, S.; Jorge, V.; Delaunay, A.; Lafon-Placette, C.; Fichot, R.; Faivre-Rampant, P.; Villar, M.; Brignolas, F.; et al. Narrow-sense heritability and PST estimates of DNA methylation in three Populus nigra L. populations under contrasting water availability. Tree Genet. Genomes 2018, 14, 78. [Google Scholar] [CrossRef]
- Teyssier, C.; Maury, S.; Beaufour, M.; Grondin, C.; Delaunay, A.; Le Metté, C.; Ader, K.; Cadene, M.; Label, P.; Lelu-Walter, M.-A. In search of markers for somatic embryo maturation in hybrid larch (Larix × eurolepis): Global DNA methylation and proteomic analyses. Physiol. Plant. 2014, 150, 271–291. [Google Scholar] [CrossRef] [PubMed]
- Trap-Gentil, M.-V.; Hébrard, C.; Lafon-Placette, C.; Delaunay, A.; Hagège, D.; Joseph, C.; Brignolas, F.; Lefebvre, M.; Barnes, S.; Maury, S. Time course and amplitude of DNA methylation in the shoot apical meristem are critical points for bolting induction in sugar beet and bolting tolerance between genotypes. J. Exp. Bot. 2011, 62, 2585–2597. [Google Scholar] [CrossRef] [PubMed]
- Kurdyukov, S.; Bullock, M. DNA Methylation Analysis: Choosing the right method. Biology 2016, 5, 3. [Google Scholar] [CrossRef] [PubMed]
- Pujol, B.; Salager, J.-L.; Beltran, M.; Bousquet, S.; McKey, D. Photosynthesis and leaf structure in domesticated cassava (Euphorbiaceae) and a close wild relative: Have leaf photosynthetic parameters evolved under domestication? Biotropica 2008, 40, 305–312. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671. [Google Scholar] [CrossRef] [PubMed]
- Causevic, A.; Delaunay, A.; Ounnar, S.; Righezza, M.; Delmotte, F.; Brignolas, F.; Hagege, D.; Maury, S. DNA methylating and demethylating treatments modify phenotype and cell wall differentiation state in sugarbeet cell lines. Plant Physiol. Biochem. PPB 2005, 43, 681–691. [Google Scholar] [CrossRef] [PubMed]
- Gourcilleau, D.; Bogeat-Triboulot, M.-B.; Le Thiec, D.; Lafon-Placette, C.; Delaunay, A.; El-Soud, W.A.; Brignolas, F.; Maury, S. DNA methylation and histone acetylation: Genotypic variations in hybrid poplars, impact of water deficit and relationships with productivity. Ann. For. Sci. 2010, 67, 208. [Google Scholar] [CrossRef]
- Zhu, R.; Shevchenko, O.; Ma, C.; Maury, S.; Freitag, M.; Strauss, S.H. Poplars with a PtDDM1-RNAi transgene have reduced DNA methylation and show aberrant post-dormancy morphology. Planta 2013, 237, 1483–1493. [Google Scholar] [CrossRef] [PubMed]
- Gentil, M.-V.; Maury, S. Characterization of epigenetic biomarkers using new molecular approaches. In Genomics-Assisted Crop Improvement; Springer: Dordrecht, The Netherlands, 2007; Volume 1, pp. 351–370. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; ISBN 3-900051-07-0. [Google Scholar]
- Bates, D.; Mächler, M.; Bolker, B.; Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 2015, 1, 1–48. [Google Scholar]
- Ada, N.-L.; Farkash, L.; Hamburger, D.; Ovadia, R.; Forrer, I.; Kagan, S.; Michal, O.-S. Light-scattering shade net increases branching and flowering in ornamental pot plants. J. Hortic. Sci. Biotechnol. 2008, 83, 9–14. [Google Scholar] [CrossRef]
- Park, Y.; Runkle, E.S. Far-red radiation promotes growth of seedlings by increasing leaf expansion and whole-plant net assimilation. Environ. Exp. Bot. 2017, 136, 41–49. [Google Scholar] [CrossRef]
- Greally, J.M. Population epigenetics. Curr. Opin. Syst. Biol. 2017, 1, 84–89. [Google Scholar] [CrossRef]
- Richards, E.J. Population epigenetics. Curr. Opin. Genet. Dev. 2008, 18, 221–226. [Google Scholar] [CrossRef]
- Karan, R.; DeLeon, T.; Biradar, H.; Subudhi, P.K. Salt stress induced variation in DNA methylation pattern and its influence on gene expression in contrasting rice genotypes. PLoS ONE 2012, 7, e40203. [Google Scholar] [CrossRef]
- Kumar, S.; Beena, A.S.; Awana, M.; Singh, A. Salt-induced tissue-specific cytosine methylation downregulates expression of HKT genes in contrasting wheat (Triticum aestivum L.) genotypes. DNA Cell Biol. 2017, 36, 283–294. [Google Scholar] [CrossRef]
- Le Gac, A.-L.; Lafon-Placette, C.; Chauveau, D.; Segura, V.; Delaunay, A.; Fichot, R.; Marron, N.; Le Jan, I.; Berthelot, A.; Bodineau, G.; et al. Winter-dormant shoot apical meristem in poplar trees shows environmental epigenetic memory. J. Exp. Bot. 2018, 69, 4821–4837. [Google Scholar] [CrossRef]
- Danchin, E.; Pocheville, A.; Rey, O.; Pujol, B.; Blanchet, S. Epigenetically facilitated mutational assimilation: Epigenetics as a hub within the inclusive evolutionary synthesis. Biol. Rev. 2019, 94, 259–282. [Google Scholar] [CrossRef]
- Pfeifer, G.P. DNA methylation and mutation. In eLS.; American Cancer Society: Atlanta, GA, USA, 2017; pp. 1–5. ISBN 978-0-470-01590-2. [Google Scholar]
- Zhang, M.; Xu, C.; von Wettstein, D.; Liu, B. Tissue-specific differences in cytosine methylation and their association with differential gene expression in Sorghum. Plant Physiol. 2011, 156, 1955–1966. [Google Scholar] [CrossRef]
- Kawakatsu, T.; Huang, S.-S.C.; Jupe, F.; Sasaki, E.; Schmitz, R.J.; Urich, M.A.; Castanon, R.; Nery, J.R.; Barragan, C.; He, Y.; et al. Epigenomic diversity in a global collection of Arabidopsis thaliana accessions. Cell 2016, 166, 492–505. [Google Scholar] [CrossRef]
- Schmid, M.W.; Heichinger, C.; Schmid, D.C.; Guthörl, D.; Gagliardini, V.; Bruggmann, R.; Aluri, S.; Aquino, C.; Schmid, B.; Turnbull, L.A.; et al. Contribution of epigenetic variation to adaptation in Arabidopsis. Nat. Commun. 2018, 9, 4446. [Google Scholar] [CrossRef]
- Pujol, B.; Blanchet, S.; Charmantier, A.; Danchin, E.; Facon, B.; Marrot, P.; Roux, F.; Scotti, I.; Teplitsky, C.; Thomson, C.E.; et al. The missing response to selection in the wild. Trends Ecol. Evol. 2018, 33, 337–346. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gourcilleau, D.; Mousset, M.; Latutrie, M.; Marin, S.; Delaunay, A.; Maury, S.; Pujol, B. Assessing Global DNA Methylation Changes Associated with Plasticity in Seven Highly Inbred Lines of Snapdragon Plants (Antirrhinum majus). Genes 2019, 10, 256. https://doi.org/10.3390/genes10040256
Gourcilleau D, Mousset M, Latutrie M, Marin S, Delaunay A, Maury S, Pujol B. Assessing Global DNA Methylation Changes Associated with Plasticity in Seven Highly Inbred Lines of Snapdragon Plants (Antirrhinum majus). Genes. 2019; 10(4):256. https://doi.org/10.3390/genes10040256
Chicago/Turabian StyleGourcilleau, Delphine, Mathilde Mousset, Mathieu Latutrie, Sara Marin, Alain Delaunay, Stéphane Maury, and Benoît Pujol. 2019. "Assessing Global DNA Methylation Changes Associated with Plasticity in Seven Highly Inbred Lines of Snapdragon Plants (Antirrhinum majus)" Genes 10, no. 4: 256. https://doi.org/10.3390/genes10040256
APA StyleGourcilleau, D., Mousset, M., Latutrie, M., Marin, S., Delaunay, A., Maury, S., & Pujol, B. (2019). Assessing Global DNA Methylation Changes Associated with Plasticity in Seven Highly Inbred Lines of Snapdragon Plants (Antirrhinum majus). Genes, 10(4), 256. https://doi.org/10.3390/genes10040256





