Does the Presence of Transposable Elements Impact the Epigenetic Environment of Human Duplicated Genes?
Abstract
:1. Introduction
2. Material and Methods
2.1. Duplicated Genes
2.2. Epigenetic Modification and Expression Data
2.3. Transposable Elements Neighborhood
2.4. Gene Classification
2.5. Statistical Tests
3. RESULTS
3.1. Duplicated Genes in the Human Genome Are Mainly Located on Different Chromosomes, Represent Old Events and Display Similar TE Environment
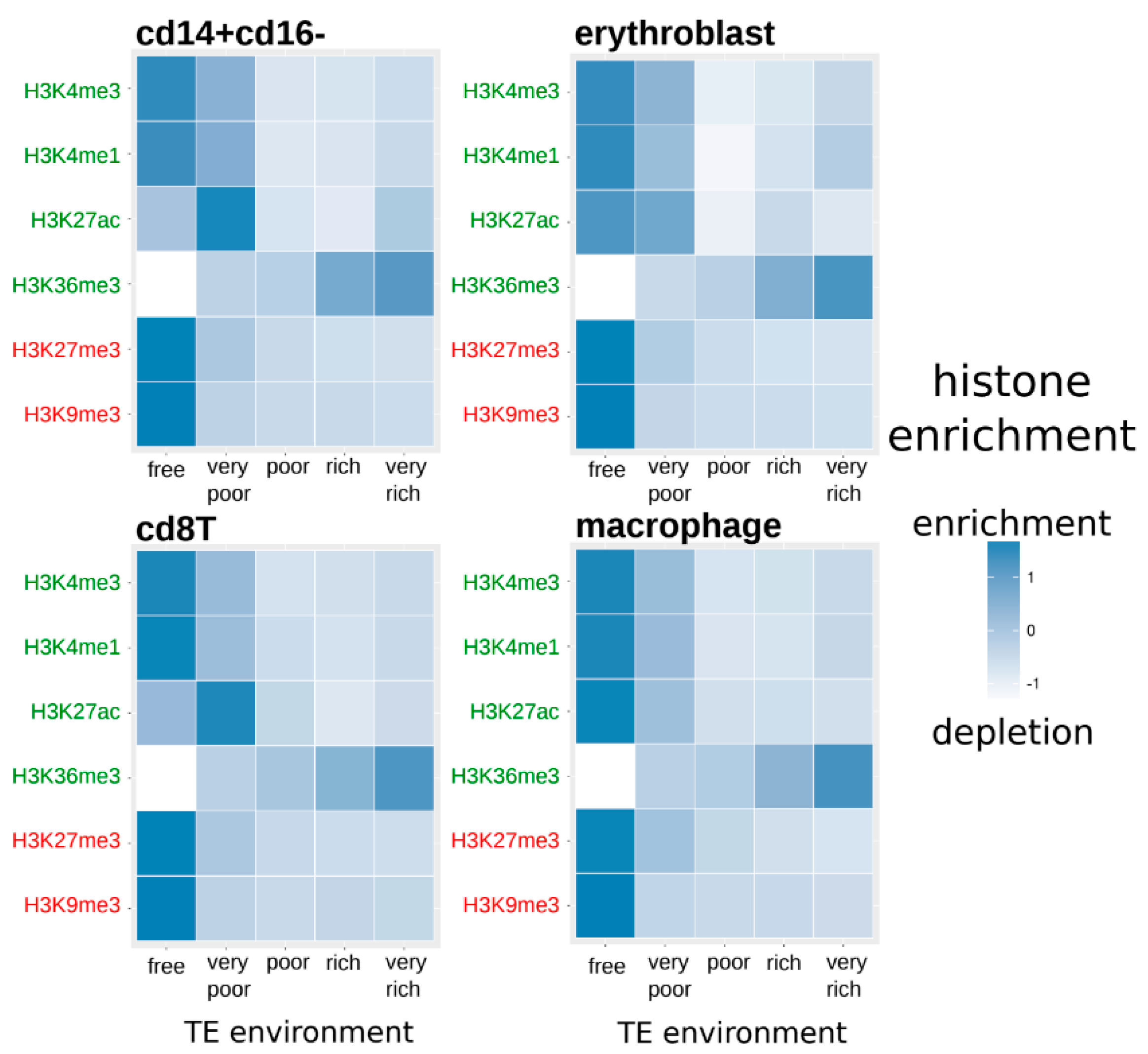
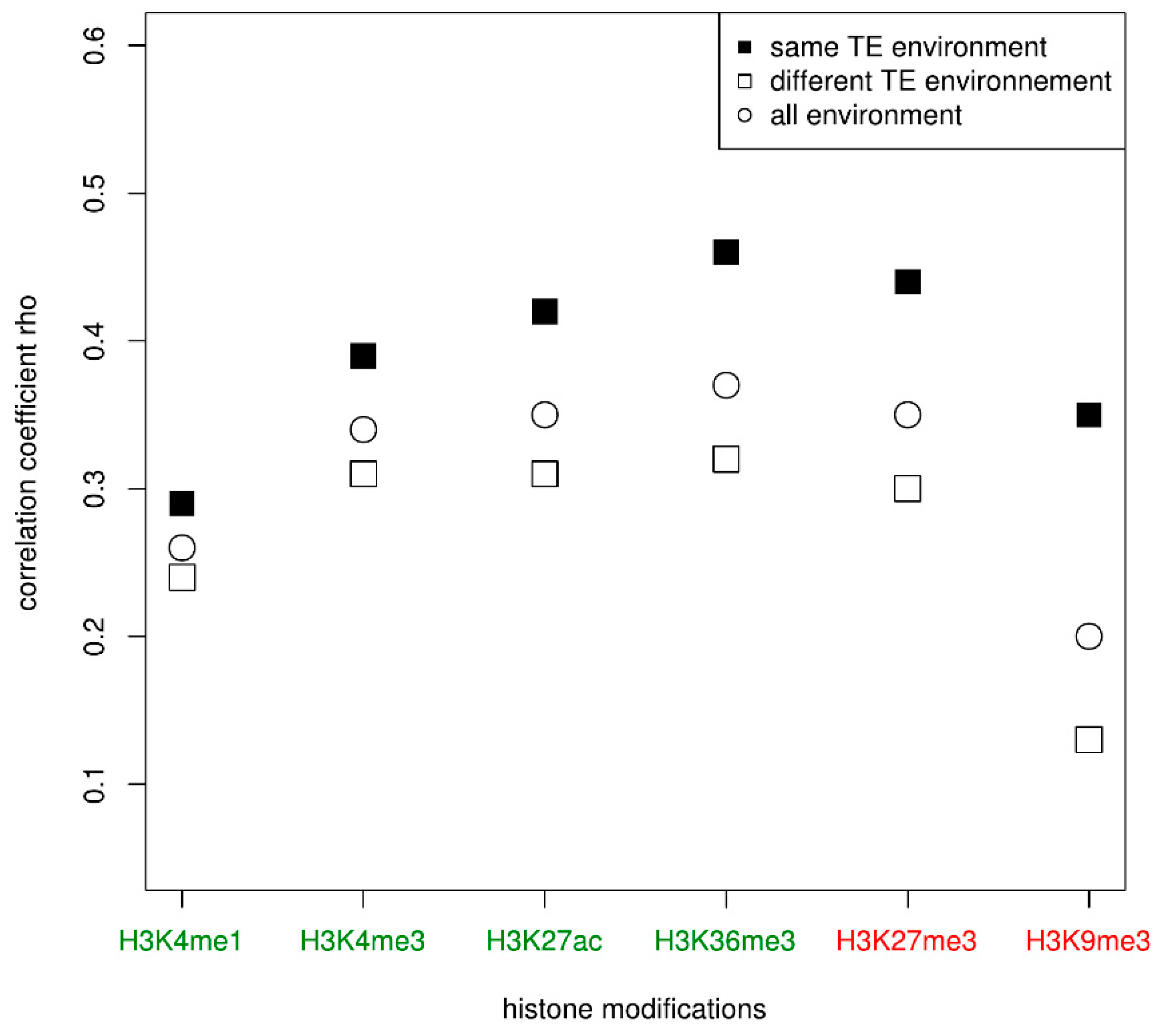
3.2. Duplicated Genes Have Similar Histone Modification Enrichment Especially If They Share a Similar TE Environment
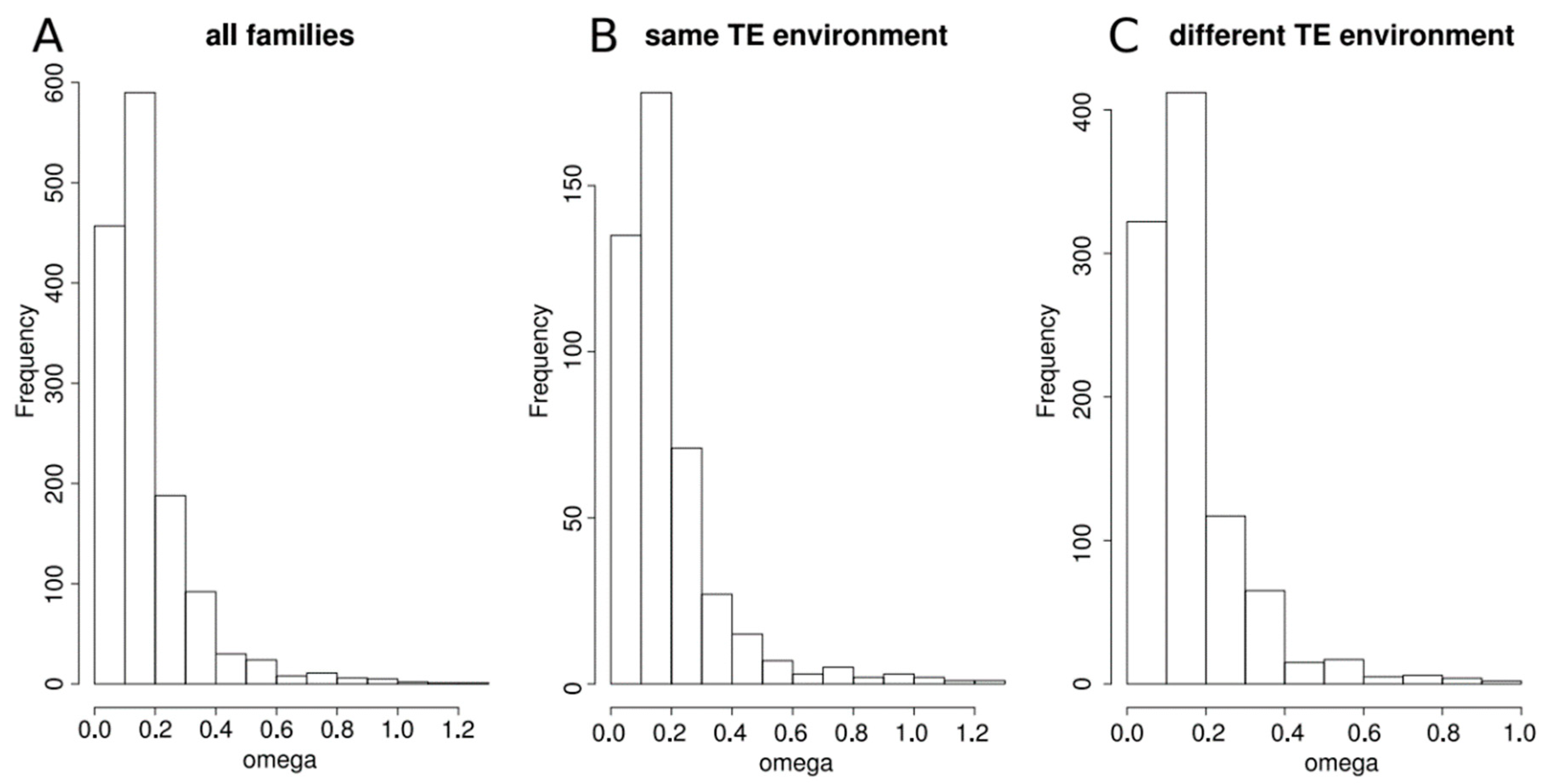
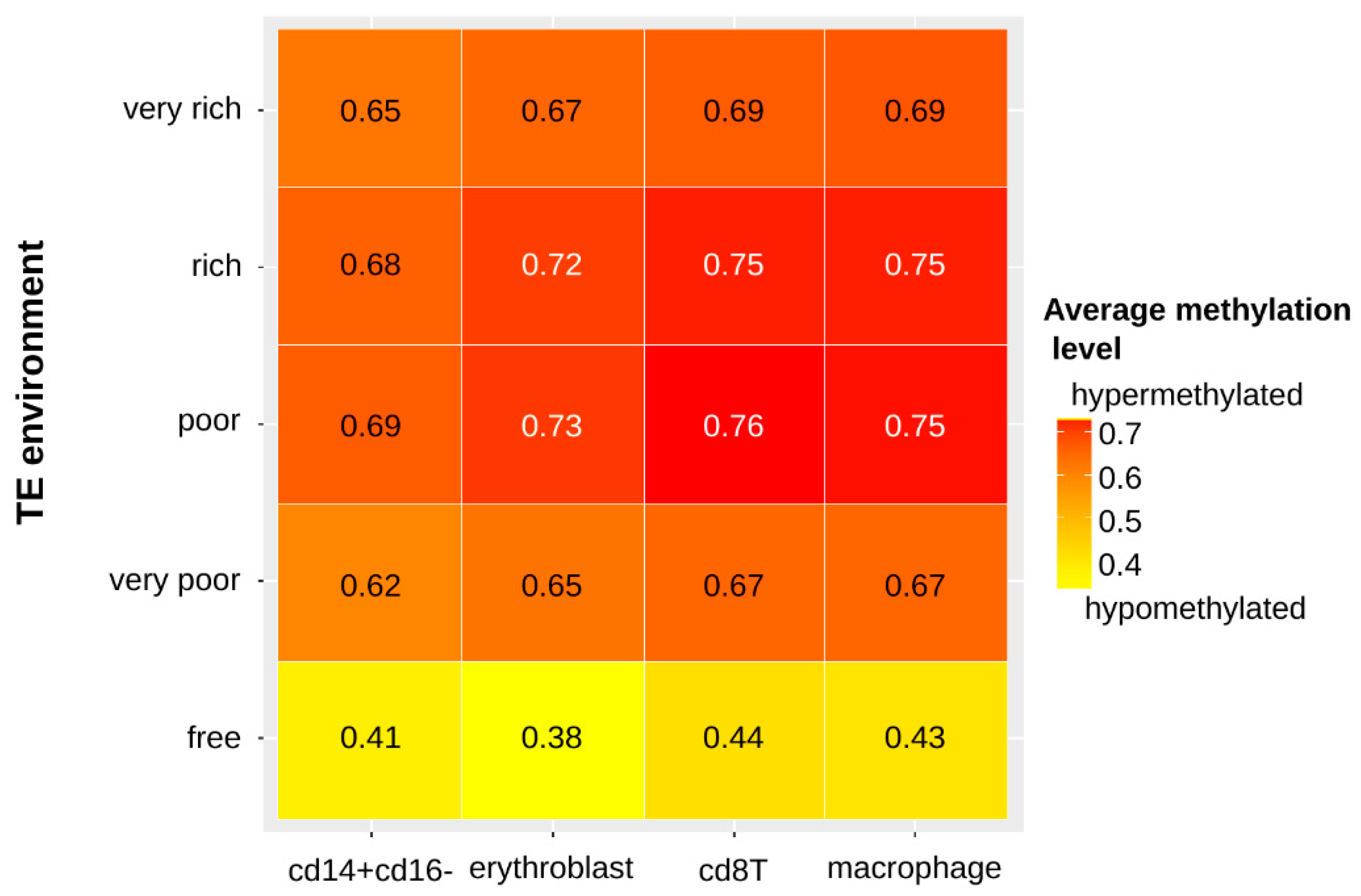
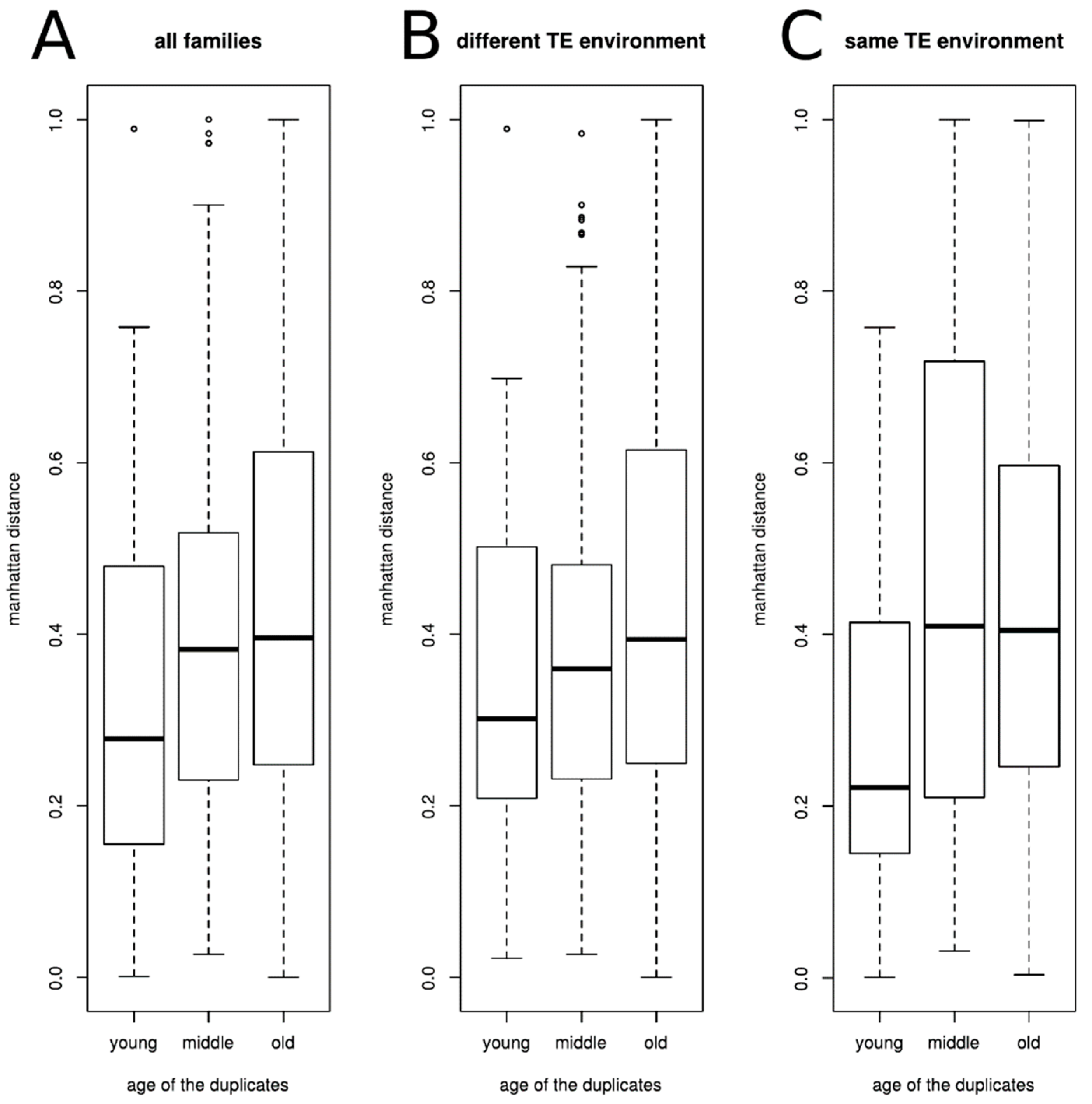
3.3. Duplicated Genes Have a Similar Methylation Level That Is Linked to Both the TE Environment Conservation and the Age of the Gene Family
3.4. The Duplicated Genes with Very Low Expression Divergence Display a High Conservation of Epigenetic Modifications and TE Environment
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Straussman, R.; Nejman, D.; Roberts, D.; Steinfeld, I.; Blum, B.; Benvenisty, N.; Simon, I.; Yakhini, Z.; Cedar, H. Developmental programming of CpG island methylation profiles in the human genome. Nat. Struct. Mol. Biol. 2009, 16, 564–571. [Google Scholar] [CrossRef] [PubMed]
- Varley, K.E.; Gertz, J.; Bowling, K.M.; Parker, S.L.; Reddy, T.E.; Pauli-Behn, F.; Cross, M.K.; Williams, B.A.; Stamatoyannopoulos, J.A.; Crawford, G.E.; et al. Dynamic DNA methylation across diverse human cell lines and tissues. Genome Res. 2013, 23, 555–567. [Google Scholar] [CrossRef]
- Ha, M.; Ng, D.W.-K.; Li, W.-H.; Chen, Z.J. Coordinated histone modifications are associated with gene expression variation within and between species. Genome Res. 2011, 21, 590–598. [Google Scholar] [CrossRef]
- Ghosh, D.; Qin, Z.S. Statistical issues in the analysis of ChIP-seq and RNA-seq data. Genes 2010, 1, 317–334. [Google Scholar] [CrossRef]
- Kucharski, R.; Maleszka, J.; Foret, S.; Maleszka, R. Nutritional control of reproductive status in honeybees via DNA methylation. Science 2008, 319, 1827–1830. [Google Scholar] [CrossRef] [PubMed]
- Chittka, A.; Wurm, Y.; Chittka, L. Epigenetics: The Making of Ant Castes. Curr. Biol. 2012, 22, R835–R838. [Google Scholar] [CrossRef] [PubMed]
- Jaenisch, R.; Bird, A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet. 2003, 33, 245–254. [Google Scholar] [CrossRef]
- Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002, 16, 6–21. [Google Scholar] [CrossRef] [PubMed]
- Weber, M.; Schübeler, D. Genomic patterns of DNA methylation: targets and function of an epigenetic mark. Curr. Opin. Cell Biol. 2007, 19, 273–280. [Google Scholar] [CrossRef]
- Jones, P.A.; Liang, G. Rethinking how DNA methylation patterns are maintained. Nat. Rev. Genet. 2009, 10, 805–811. [Google Scholar] [CrossRef]
- Bernstein, B.E.; Meissner, A.; Lander, E.S. The Mammalian Epigenome. Cell 2007, 128, 669–681. [Google Scholar] [CrossRef]
- Carthew, R.W.; Sontheimer, E.J. Origins and Mechanisms of miRNAs and siRNAs. Cell 2009, 136, 642–655. [Google Scholar] [CrossRef] [PubMed]
- Ghildiyal, M.; Zamore, P.D. Small silencing RNAs: an expanding universe. Nat. Rev. Genet. 2009, 10, 94–108. [Google Scholar] [CrossRef]
- Grant, P.A. A tale of histone modifications. Genome Biol. 2001, 2, REVIEWS0003. [Google Scholar] [CrossRef] [PubMed]
- Peterson, C.L.; Laniel, M.-A. Histones and histone modifications. Curr. Biol. 2004, 14, R546–551. [Google Scholar] [CrossRef]
- Li, B.; Carey, M.; Workman, J.L. The role of chromatin during transcription. Cell 2007, 128, 707–719. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Cancer epigenomics: DNA methylomes and histone-modification maps. Nat. Rev. Genet. 2007, 8, 286–298. [Google Scholar] [CrossRef]
- Britten, R.J.; Davidson, E.H. Gene regulation for higher cells: A theory. Science 1969, 165, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Mihola, O.; Trachtulec, Z.; Vlcek, C.; Schimenti, J.C.; Forejt, J. A mouse speciation gene encodes a meiotic histone H3 methyltransferase. Science 2009, 323, 373–375. [Google Scholar] [CrossRef] [PubMed]
- Cain, C.E.; Blekhman, R.; Marioni, J.C.; Gilad, Y. Gene expression differences among primates are associated with changes in a histone epigenetic modification. Genetics 2011, 187, 1225–1234. [Google Scholar] [CrossRef]
- Zeng, J.; Konopka, G.; Hunt, B.G.G.; Preuss, T.M.M.; Geschwind, D.; Yi, S.V.V. Divergent whole-genome methylation maps of human and chimpanzee brains reveal epigenetic basis of human regulatory evolution. Am. J. Hum. Genet. 2012, 91, 455–465. [Google Scholar] [CrossRef] [PubMed]
- Cocozza, S.; Akhtar, M.M.; Miele, G.; Monticelli, A. CpG islands undermethylation in human genomic regions under selective pressure. PLoS ONE 2011, 6, e23156. [Google Scholar] [CrossRef]
- Akhtar, M.M.; Scala, G.; Cocozza, S.; Miele, G.; Monticelli, A. (2013) CpG islands under selective pressure are enriched with H3K4me3, H3K27ac and H3K36me3 histone modifications. BMC Evol. Biol. 2013, 13, 145. [Google Scholar] [CrossRef]
- Hernando-Herraez, I.; Heyn, H.; Fernandez-Callejo, M.; Vidal, E.; Fernandez-Bellon, H.; Prado-Martinez, J.; Sharp, A.J.; Esteller, M.; Marques-Bonet, T. The interplay between DNA methylation and sequence divergence in recent human evolution. Nucleic Acids Res. 2015, 43, 8204–8214. [Google Scholar] [CrossRef]
- Kolasinska-Zwierz, P.; Down, T.; Latorre, I.; Liu, T.; Liu, X.S.; Ahringer, J. Differential chromatin marking of introns and expressed exons by H3K36me3. Nat. Genet. 2009, 41, 376–381. [Google Scholar] [CrossRef] [PubMed]
- Woo, Y.H.; Li, W.H. Evolutionary conservation of histone modifications in mammals. Mol. Biol. Evol. 2012, 29, 1757–1767. [Google Scholar] [CrossRef]
- Sarda, S.; Zeng, J.; Hunt, B.G.; Yi, S.V. The evolution of invertebrate gene body methylation. Mol. Biol. Evol. 2012, 29, 1907–1916. [Google Scholar] [CrossRef] [PubMed]
- Takuno, S.; Gaut, B.S. Gene body methylation is conserved between plant orthologs and is of evolutionary consequence. Proc. Natl. Acad. Sci. USA 2013, 110, 1797–1802. [Google Scholar] [CrossRef] [PubMed]
- Keller, T.E.; Yi, S.V. DNA methylation and evolution of duplicate genes. Proc. Natl. Acad. Sci. USA 2014, 111, 5932–5937. [Google Scholar] [CrossRef]
- Berke, L.; Sanchez-Perez, G.F.; Snel, B. Contribution of the epigenetic mark H3K27me3 to functional divergence after whole genome duplication in Arabidopsis. Genome Biol. 2012, 13, R94. [Google Scholar] [CrossRef]
- Prendergast, J.G.D.; Chambers, E.V.; Semple, C.A.M. Sequence-Level Mechanisms of Human Epigenome Evolution. Genome Biol. Evol. 2014, 6, 1758–1771. [Google Scholar] [CrossRef] [PubMed]
- Consortium International Human Genome Sequencing. Finishing the euchromatic sequence of the human genome. Nature 2004, 431, 931–945. [Google Scholar] [CrossRef] [PubMed]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [PubMed]
- Cordaux, R.; Batzer, M.A. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 2009, 10, 691–703. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, M. Functional evolution of noncoding DNA. Curr. Opin. Genet. Dev. 2002, 12, 634–639. [Google Scholar] [CrossRef]
- Kidwell, M.G.; Lisch, D.R. Transposable elements and host genome evolution. Trends Ecol. Evol. 2000, 15, 95–99. [Google Scholar] [CrossRef]
- Biémont, C.; Vieira, C. Genetics: Junk DNA as an evolutionary force. Nature 2006, 443, 521–524. [Google Scholar] [CrossRef] [PubMed]
- Medstrand, P.; van de Lagemaat, L.N.; Mager, D.L. Retroelement distributions in the human genome: Variations associated with age and proximity to genes. Genome Res. 2002, 12, 1483–1495. [Google Scholar] [CrossRef]
- Hoen, D.R.; Park, K.C.; Elrouby, N.; Yu, Z.; Mohabir, N.; Cowan, R.K.; Bureau, T.E. Transposon-mediated expansion and diversification of a family of ULP-like genes. Mol. Biol Evol. 2006, 2006 23, 1254–1268. [Google Scholar] [CrossRef]
- Juretic, N.; Hoen, D.R.; Huynh, M.L.; Harrison, P.M.; Bureau, T.E. The evolutionary fate of MULE-mediated duplications of host gene fragments in rice. Genome Res. 2005, 15, 1292–1297. [Google Scholar] [CrossRef]
- Slotkin, R.K.; Martienssen, R. Landscape of Somatic Retrotransposition in Human Cancers. Nat. Rev. Genet. 2007, 8, 272–285. [Google Scholar] [CrossRef]
- Huda, A.; Jordan, I.K. Epigenetic regulation of mammalian genomes by transposable elements. Ann. N. Y. Acad. Sci. 2009, 1178, 276–284. [Google Scholar] [CrossRef]
- Kulis, M.; Esteller, M. DNA methylation and cancer. Adv. Genet. 2010, 70, 27–56. [Google Scholar]
- Ross, J.P.; Rand, K.N.; Molloy, P.L. Hypomethylation of repeated DNA sequences in cancer. Epigenomics 2010, 2, 245–269. [Google Scholar] [CrossRef]
- Morgan, H.D.; Sutherland, H.G.; Martin, D.I.; Whitelaw, E. Epigenetic inheritance at the agouti locus in the mouse. Nat. Genet. 1999, 23, 314–318. [Google Scholar] [CrossRef]
- Rebollo, R.; Karimi, M.M.; Bilenky, M.; Gagnier, L.; Miceli-Royer, K.; Zhang, Y.; Goyal, P.; Keane, T.M.; Jones, S.; Hirst, M.; et al. Retrotransposon-induced heterochromatin spreading in the mouse revealed by insertional polymorphisms. PLoS Genet. 2011, 7, e1002301. [Google Scholar] [CrossRef] [PubMed]
- Eichten, S.R.; Ellis, N.A.; Makarevitch, I.; Yeh, C.T.; Gent, J.I.; Guo, L.; McGinnis, K.M.; Zhang, X.; Schnable, P.S.; Vaughn, M.W.; et al. Spreading of Heterochromatin Is Limited to Specific Families of Maize Retrotransposons. PLoS Genet. 2012, 8, e1003127. [Google Scholar] [CrossRef] [PubMed]
- Gendrel, A.-V.; Lippman, Z.; Yordan, C.; Colot, V.; Martienssen, R.A. Dependence of heterochromatic histone H3 methylation patterns on the Arabidopsis gene DDM1. Science 2002, 297, 1871–1873. [Google Scholar] [CrossRef] [PubMed]
- Volpe, T.A.; Kidner, C.; Hall, I.M.; Teng, G.; Grewal, S.I.S.; Martienssen, R.A. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 2002, 297, 1833–1837. [Google Scholar] [CrossRef] [PubMed]
- Lippman, Z.; Gendrel, A.-V.; Black, M.; Vaughn, M.W.; Dedhia, N.; McCombie, W.R.; Lavine, K.; Mittal, V.; May, B.; Kasschau, K.D.; et al. Role of transposable elements in heterochromatin and epigenetic control. Nature 2004, 430, 471–476. [Google Scholar] [CrossRef]
- Mirouze, M.; Vitte, C. Transposable elements, a treasure trove to decipher epigenetic variation: Insights from Arabidopsis and crop epigenomes. J. Exp. Bot. 2014, 65, 2801–2812. [Google Scholar] [CrossRef] [PubMed]
- Byun, H.-M.; Heo, K.; Mitchell, K.J.; Yang, A.S. Mono-allelic retrotransposon insertion addresses epigenetic transcriptional repression in human genome. J. Biomed. Sci. 2012, 19, 13. [Google Scholar] [CrossRef]
- Grégoire, L.; Haudry, A.; Lerat, E. The transposable element environment of human genes is associated with histone and expression changes in cancer. BMC Genomics 2016, 17, 588. [Google Scholar] [CrossRef]
- Trizzino, M.; Park, Y.; Holsbach-Beltrame, M.; Aracena, K.; Mika, K.; Caliskan, M.; Perry, G.H.; Lynch, V.J.; Brown, C.D. Transposable elements are the primary source of novelty in primate gene regulation. Genome Res. 2017, 27, 1623–1633. [Google Scholar] [CrossRef]
- Penel, S.; Arigon, A.-M.; Dufayard, J.-F.; Sertier, A.-S.; Daubin, V.; Duret, L.; Gouy, M.; Perrière, G. Databases of homologous gene families for comparative genomics. BMC Bioinform. 2009, 10 (Suppl 6), S3. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 2007, 24, 1586–1591. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Stunnenberg, H.G.; Hirst, M.; de Almeida, M.; Altucci, L.; Amin, V.; Amit, I.; Antonarakis, S.E.; Aparicio, S.; Arima, T.; International Human Epigenome Consortium; et al. The International Human Epigenome Consortium: A Blueprint for Scientific Collaboration and Discovery. Cell 2016, 167, 1145–1149. [Google Scholar] [CrossRef]
- Hart, T.; Komori, H.K.; LaMere, S.; Podshivalova, K.; Salomon, D.R. Finding the active genes in deep RNA-seq gene expression studies. BMC Genomics 2013, 14, 778. [Google Scholar] [CrossRef]
- Conesa, A.; Madrigal, P.; Tarazona, S.; Gomez-Cabrero, D.; Cervera, A.; McPherson, A.; Szcześniak, M.W.; Gaffney, D.J.; Elo, L.L.; Zhang, X.; et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 2016, 17, 13. [Google Scholar] [CrossRef]
- Bailly-Bechet, M.; Haudry, A.; Lerat, E. ‘One code to find them all’: A perl tool to conveniently parse RepeatMasker output files. Mob. DNA 2014, 5, 13. [Google Scholar] [CrossRef]
- Stewart, C.; Kural, D.; Strömberg, M.P.; Walker, J.A.; Konkel, M.K.; Stütz, A.M.; Urban, A.E.; Grubert, F.; Lam, H.Y.; Lee, W.P.; et al. A comprehensive map of mobile element insertion polymorphisms in humans. PLoS Genet. 2011, 7, e1002236. [Google Scholar] [CrossRef]
- Walter, M.; Teissandier, A.; Pérez-Palacios, R.; Bourc’his, D. An epigenetic switch ensures transposon repression upon dynamic loss of DNA methylation in embryonic stem cells. Elife 2016, 5, e11418. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: https://www.R-project.org/ (accessed on 29 January 2019).
- Rodin, S.N.; Riggs, A.D. Epigenetic Silencing May Aid Evolution by Gene Duplication. J. Mol. Evol. 2003, 56, 718–729. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Z.; Du, K.; Yu, Q.; Zhang, Y.E.; He, S. Divergent DNA Methylation Provides Insights into the Evolution of Duplicate Genes in Zebrafish. G3 2016, 6, 3581–3591. [Google Scholar] [CrossRef] [PubMed]
- McLysaght, A.; Hokamp, K.; Wolfe, K.H. Extensive genomic duplication during early chordate evolution. Nat. Genet. 2002, 31, 200–204. [Google Scholar] [CrossRef]
- Dehal, P.; Boore, J.L. Two rounds of whole genome duplication in the ancestral vertebrate. PLoS Biol. 2005, 3, e314. [Google Scholar] [CrossRef] [PubMed]
- Mendivil Ramos, O.; Ferrier, D.E.K. Mechanisms of Gene Duplication and Translocation and Progress towards Understanding Their Relative Contributions to Animal Genome Evolution. Int. J. Evol. Biol. 2012, 2012, 1–10. [Google Scholar] [CrossRef]
- Zhang, P.; Min, W.; Li, W.-H. Different age distribution patterns of human, nematode, and Arabidopsis duplicate genes. Gene 2004, 342, 263–268. [Google Scholar] [CrossRef]
- Mortada, H.; Vieira, C.; Lerat, E. Genes devoid of full-length transposable element insertions are involved in development and in the regulation of transcription in human and closely related species. J. Mol. Evol. 2010, 71, 180–191. [Google Scholar] [CrossRef]
- Nellåker, C.; Keane, T.M.; Yalcin, B.; Wong, K.; Agam, A.; Belgard, T.G.; Flint, J.; Adams, D.J.; Frankel, W.N.; Ponting, C.P. The genomic landscape shaped by selection on transposable elements across 18 mouse strains. Genome Biol. 2012, 13, R45. [Google Scholar] [CrossRef]
- Wagner, E.J.; Carpenter, P.B. Understanding the language of Lys36 methylation at histone H3. Nat. Rev. Mol. Cell Biol. 2012, 13, 115–126. [Google Scholar] [CrossRef]
- Teissandier, A.; Bourc’his, D. Gene body DNA methylation conspires with H3K36me3 to preclude aberrant transcription. EMBO J. 2017, 36, 1471–1473. [Google Scholar] [CrossRef] [PubMed]
- Kondo, Y.; Issa, J.-P.J. Enrichment for histone H3 lysine 9 methylation at Alu repeats in human cells. J. Biol. Chem. 2003, 278, 27658–27662. [Google Scholar] [CrossRef] [PubMed]
- Martens, J.H.A.; O’Sullivan, R.J.; Braunschweig, U.; Opravil, S.; Radolf, M.; Steinlein, P.; Jenuwein, T. The profile of repeat-associated histone lysine methylation states in the mouse epigenome. EMBO J. 2005, 24, 800–812. [Google Scholar] [CrossRef]
- Huda, A.; Mariño-Ramírez, L.; Jordan, I.K. Epigenetic histone modifications of human transposable elements: genome defense versus exaptation. Mob. DNA 2010, 1, 2. [Google Scholar] [CrossRef] [PubMed]
- Pauler, F.M.; Sloane, M.A.; Huang, R.; Regha, K.; Koerner, M.V.; Tamir, I.; Sommer, A.; Aszodi, A.; Jenuwein, T.; Barlow, D.P. H3K27me3 forms BLOCs over silent genes and intergenic regions and specifies a histone banding pattern on a mouse autosomal chromosome. Genome Res. 2009, 19, 221–233. [Google Scholar] [CrossRef] [PubMed]
- Li, E.; Zhang, Y. DNA methylation in mammals. Cold Spring Harb. Perspect. Biol. 2014, 6, a019133. [Google Scholar] [CrossRef]
- Zheng, D. Gene duplication in the epigenomic era. Epigenetics 2008, 3, 250–253. [Google Scholar] [CrossRef]
- Rodin, S.N.; Parkhomchuk, D.V.; Rodin, A.S.; Holmquist, G.P.; Riggs, A.D. Repositioning-dependent fate of duplicate genes. DNA Cell Biol. 2005, 24, 529–542. [Google Scholar] [CrossRef]
- Berke, L.; Snel, B. The Histone Modification H3K27me3 Is Retained after Gene Duplication and Correlates with Conserved Noncoding Sequences in Arabidopsis. Genome Biol. Evol. 2014, 6, 572–579. [Google Scholar] [CrossRef] [PubMed]
- Babarinde, I.A.; Saitou, N. Genomic Locations of Conserved Noncoding Sequences and Their Proximal Protein-Coding Genes in Mammalian Expression Dynamics. Mol. Biol. Evol. 2016, 33, 1807–1817. [Google Scholar] [CrossRef] [PubMed]
| TE Category | All Protein Coding Genes | Duplicated Genes |
|---|---|---|
| TE-free | 773 (4.08%) | 109 (3.84%) |
| TE-very-poor | 4830 (25.50%) | 713 (25.10%) |
| TE-poor | 5885 (31.08%) | 915 (32.22%) |
| TE-rich | 4848 (25.60%) | 729 (25.67%) |
| TE-very-rich | 2602 (13.74%) | 374 (13.17%) |
| TE Environment of the Second Gene | ||||||
|---|---|---|---|---|---|---|
| TE-Free | TE-Very-Poor | TE-Poor | TE-Rich | TE-Very-Rich | ||
| TE Environment of the First Gene | TE-free | 20 (1.41%) | / | / | / | / |
| TE-very-poor | 36 (2.53%) | 121 (8.52%) | / | / | / | |
| TE-poor | 18 (1.27%) | 220 (15.49%) | 169 (11.90%) | / | / | |
| TE-rich | 13 (0.91%) | 143 (10.07%) | 229 (16.13%) | 110 (7.75%) | / | |
| TE-very-rich | 2 (0.14%) | 72 (5.07%) | 110 (7.75%) | 124 (8.73%) | 33 (2.321%) | |
| Repressive Modifications | Activating Modifications | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| H3K27me3 | H3K9me3 | H3K36me3 | H3K27ac | H3K4me1 | H3K4me3 | ||||||||
| Spearman Rho | q Value | Spearman Rho | q Value | Spearman Rho | q Value | Spearman Rho | q Value | Spearman Rho | q Value | Spearman Rho | q Value | ||
| Duplicated genes | CD14+CD16− | 0.31 * | 4.525714 × 10−16 | 0.18 * | 2.767059 × 10−11 | 0.29 * | 4.525714 × 10−16 | 0.31 * | 4.525714 × 10−16 | 0.19 * | 1.496681 × 10−13 | 0.33 * | 4.525714 × 10−16 |
| erythroblast | 0.32 * | 4.525714 × 10−16 | 0.15 * | 2.088000 × 10−8 | 0.30 * | 4.525714 × 10−16 | 0.31 * | 4.525714 × 10−16 | 0.27 * | 4.525714 × 10−16 | 0.39 * | 4.525714 × 10−16 | |
| CD8T | 0.32 * | 4.525714 × 10−16 | 0.10 * | 1.878261 × 10−4 | 0.29 * | 4.525714 × 10−16 | 0.16 * | 4.189091 × 10−9 | 0.24 * | 4.525714 × 10−16 | 0.30 * | 4.525714 × 10−16 | |
| macrophage | 0.34 * | 4.525714 × 10−16 | 0.17 * | 1.369385 × 10−10 | 0.30 * | 4.525714 × 10−16 | 0.23 * | 4.525714 × 10−16 | 0.20 * | 1.224000 × 10−14 | 0.33 * | 4.525714 × 10−16 | |
| Duplicated genes with same TE environment | CD14+CD16− | 0.41 * | 4.525714 × 10−16 | 0.25 * | 4.865806 × 10−8 | 0.36 * | 8.623256 × 10−15 | 0.38 * | 4.525714 × 10−16 | 0.21 * | 7.111385 × 10−6 | 0.35 * | 1.936000 × 10−14 |
| erythroblast | 0.44 * | 4.525714 × 10−16 | 0.29 * | 4.727547 × 10−10 | 0.38 * | 4.525714 × 10−16 | 0.38 * | 4.525714 × 10−16 | 0.27 * | 4.189091 × 10−9 | 0.42 * | 4.525714 × 10−16 | |
| CD8T | 0.43 * | 4.525714 × 10−16 | 0.22 * | 1.714286 × 10−6 | 0.32 * | 4.838400 × 10−12 | 0.26 * | 3.234098 × 10−8 | 0.26 * | 2.088000 × 10−8 | 0.36 * | 5.356098 × 10−15 | |
| macrophage | 0.40 * | 4.525714 × 10−16 | 0.26 * | 1.743158 × 10−8 | 0.44 * | 4.525714 × 10−16 | 0.19 * | 3.299104 × 10−5 | 0.22 * | 1.890000 × 10−6 | 0.36 * | 5.356098 × 10−15 | |
| Duplicated genes with different TE environment | CD14+CD16− | 0.25 * | 2.368421 × 10−15 | 0.14 * | 1.963636 × 10−5 | 0.25 * | 7.645714 × 10−15 | 0.27 * | 4.525714 × 10−16 | 0.18 * | 1.998621 × 10−8 | 0.31 * | 4.525714 × 10−16 |
| erythroblast | 0.26 * | 7.297297 × 10−16 | 0.08 * | 1.454197 × 10−2 | 0.26 * | 4.525714 × 10−16 | 0.27 * | 4.525714 × 10−16 | 0.26 * | 4.940000 × 10−16 | 0.36 * | 4.525714 × 10−16 | |
| CD8T | 0.25 * | 4.098462 × 10−15 | 0.03 | 2.789000 × 10−1 | 0.28 * | 4.525714 × 10−16 | 0.11 * | 5.142857 × 10−4 | 0.22 * | 4.599184 × 10−12 | 0.26 * | 4.525714 × 10−16 | |
| macrophage | 0.31 * | 4.525714 × 10−16 | 0.12 * | 1.058824 × 10−4 | 0.22 * | 3.750000 × 10−12 | 0.24 * | 8.405217 × 10−14 | 0.18 * | 7.315714 × 10−9 | 0.31 * | 4.525714 × 10−16 | |
| Duplicated Genes | Duplicated Genes with the Same TE Environment | Duplicated Genes with a Different TE Environment | ||||
|---|---|---|---|---|---|---|
| Spearman Rho | q Value | Spearman Rho | q Value | Spearman Rho | q Value | |
| CD14+CD16− | 0.14 * | 1.911360 × 10−6 | 0.17 * | 3.354545 × 10−4 | 0.11 * | 9.217000 × 10−4 |
| erythroblast | 0.16 * | 9.492000 × 10−9 | 0.21 * | 2.280000 × 10−5 | 0.13 * | 3.475500 × 10−5 |
| CD8T | 0.15 * | 1.333800 × 10−8 | 0.22 * | 9.620000 × 10−6 | 0.12 * | 1.370400 × 10−4 |
| macrophage | 0.18 * | 6.968400 × 10−11 | 0.27 * | 1.333800 × 10−8 | 0.13 * | 4.400000× 10−5 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lannes, R.; Rizzon, C.; Lerat, E. Does the Presence of Transposable Elements Impact the Epigenetic Environment of Human Duplicated Genes? Genes 2019, 10, 249. https://doi.org/10.3390/genes10030249
Lannes R, Rizzon C, Lerat E. Does the Presence of Transposable Elements Impact the Epigenetic Environment of Human Duplicated Genes? Genes. 2019; 10(3):249. https://doi.org/10.3390/genes10030249
Chicago/Turabian StyleLannes, Romain, Carène Rizzon, and Emmanuelle Lerat. 2019. "Does the Presence of Transposable Elements Impact the Epigenetic Environment of Human Duplicated Genes?" Genes 10, no. 3: 249. https://doi.org/10.3390/genes10030249
APA StyleLannes, R., Rizzon, C., & Lerat, E. (2019). Does the Presence of Transposable Elements Impact the Epigenetic Environment of Human Duplicated Genes? Genes, 10(3), 249. https://doi.org/10.3390/genes10030249





