Decreased Spikelets 4 Encoding a Novel Tetratricopeptide Repeat Domain-Containing Protein Is Involved in DNA Repair and Spikelet Number Determination in Rice
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
2.2. Map-Based Cloning
2.3. Plasmid Construction and Plant Transformation
2.4. RNA Preparation, RT-PCR Analysis, and mRNA in-Situ Hybridization
2.5. Genotoxic Stress
3. Results and Discussion
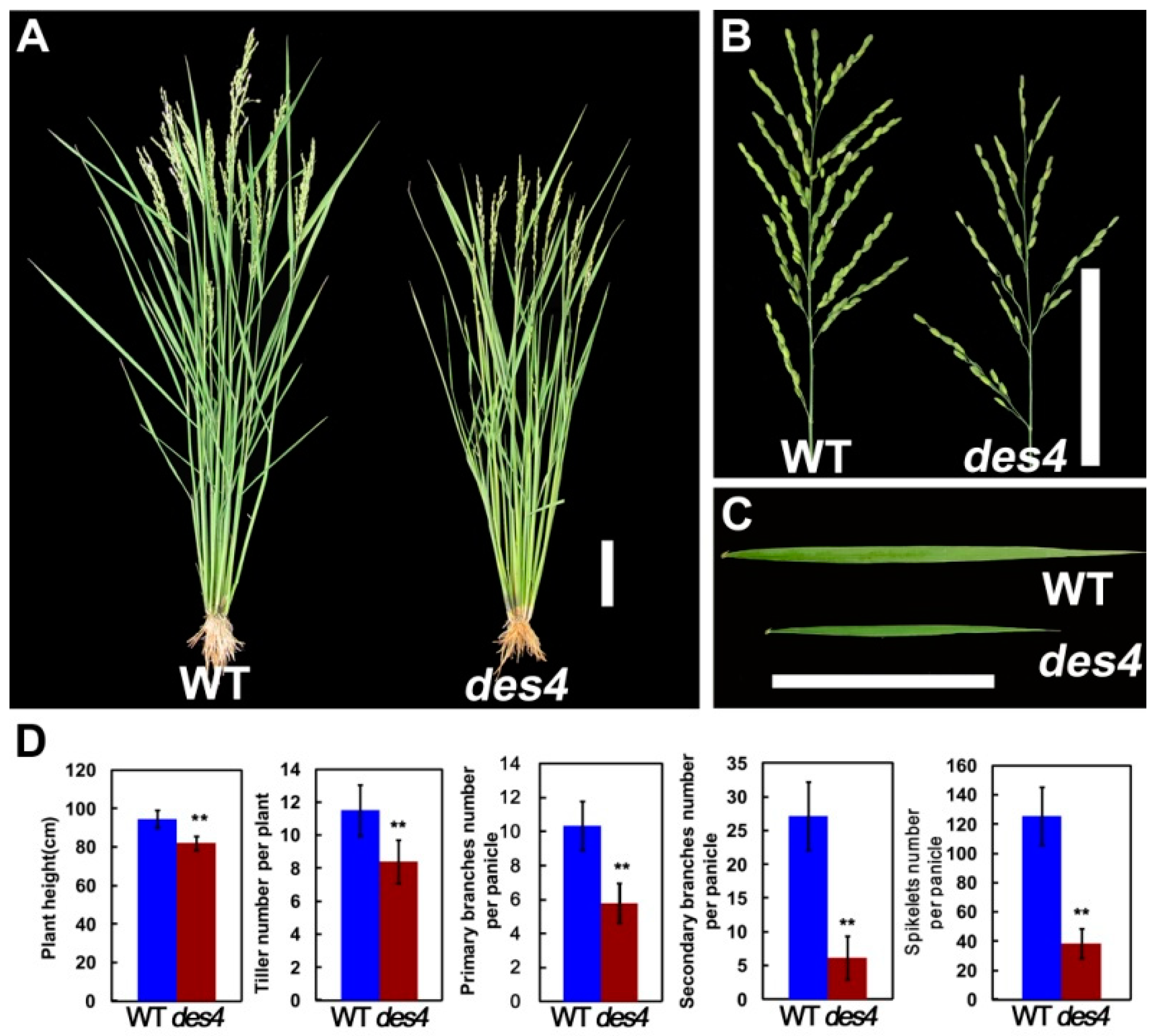
3.1. des4 Has Reduced Spikelet Number
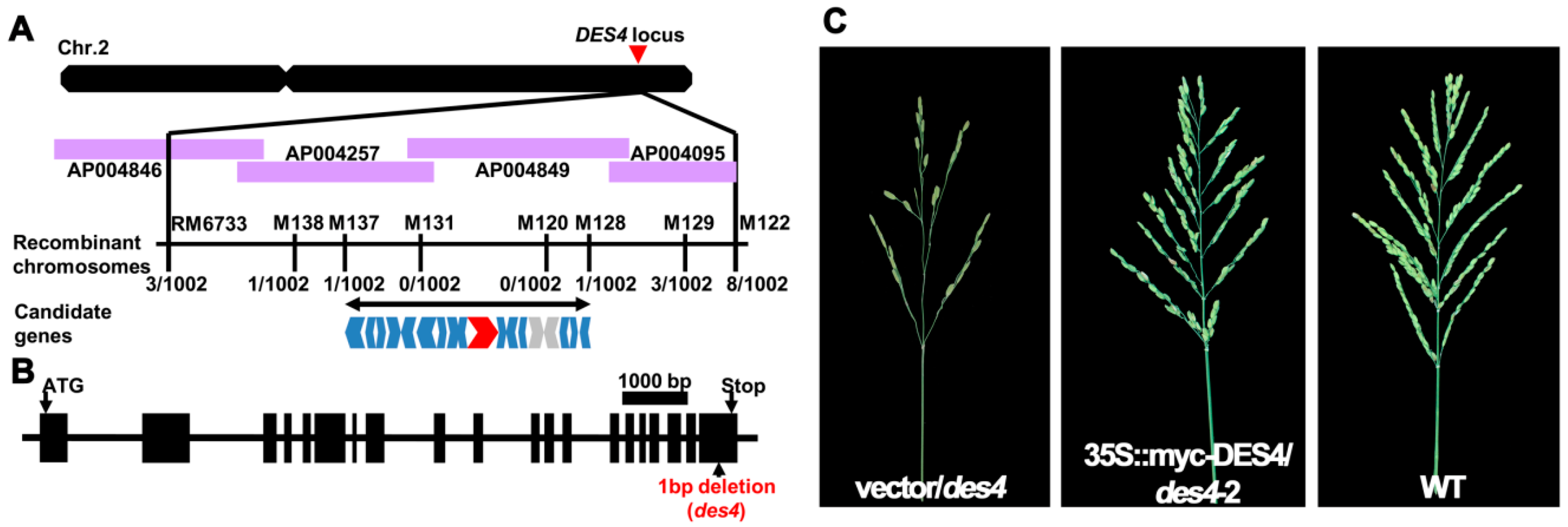
3.2. Positional Cloning of des4
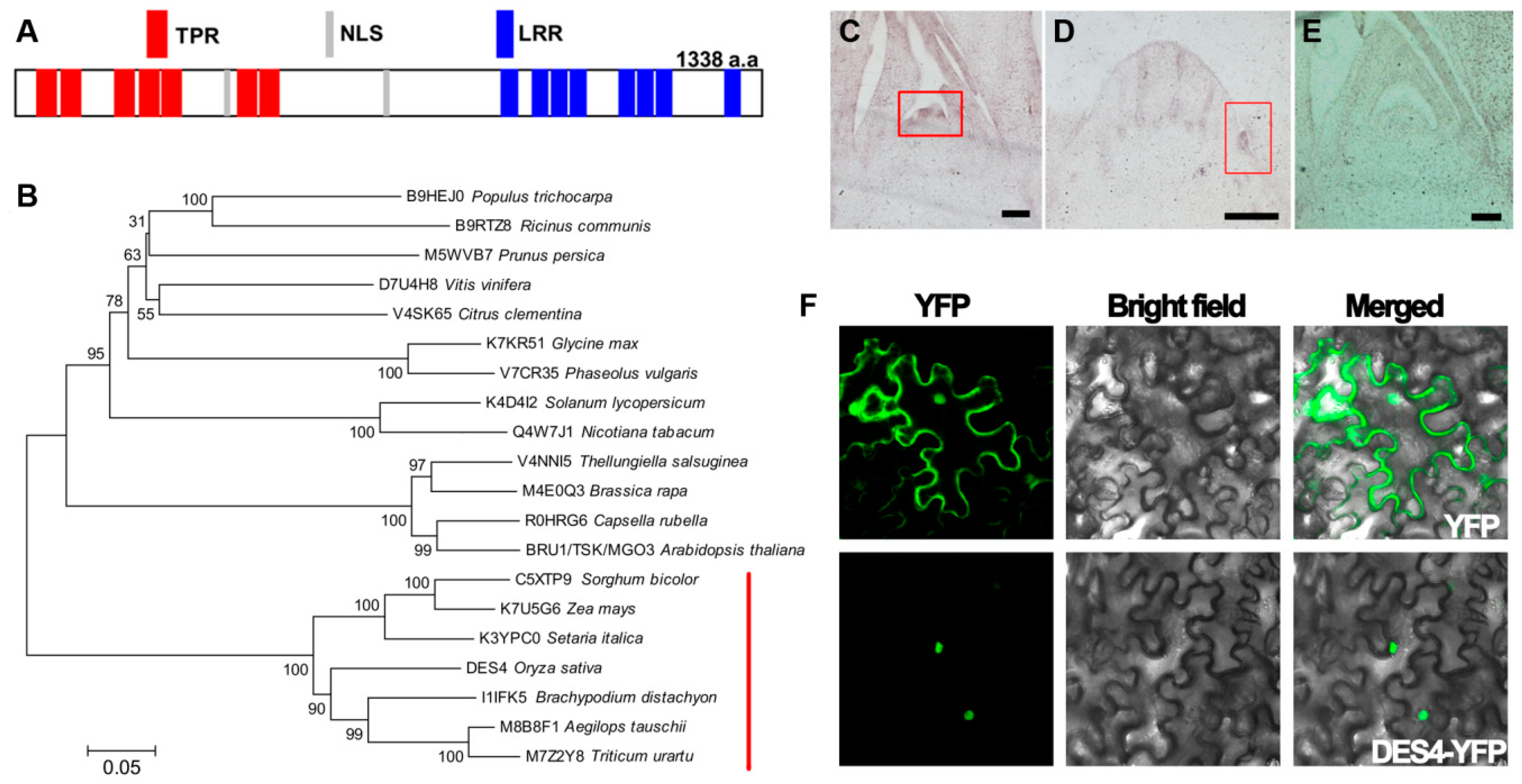
3.3. DES4 Encodes a Novel Tetratricopeptide Repeat Domain-Containing Protein
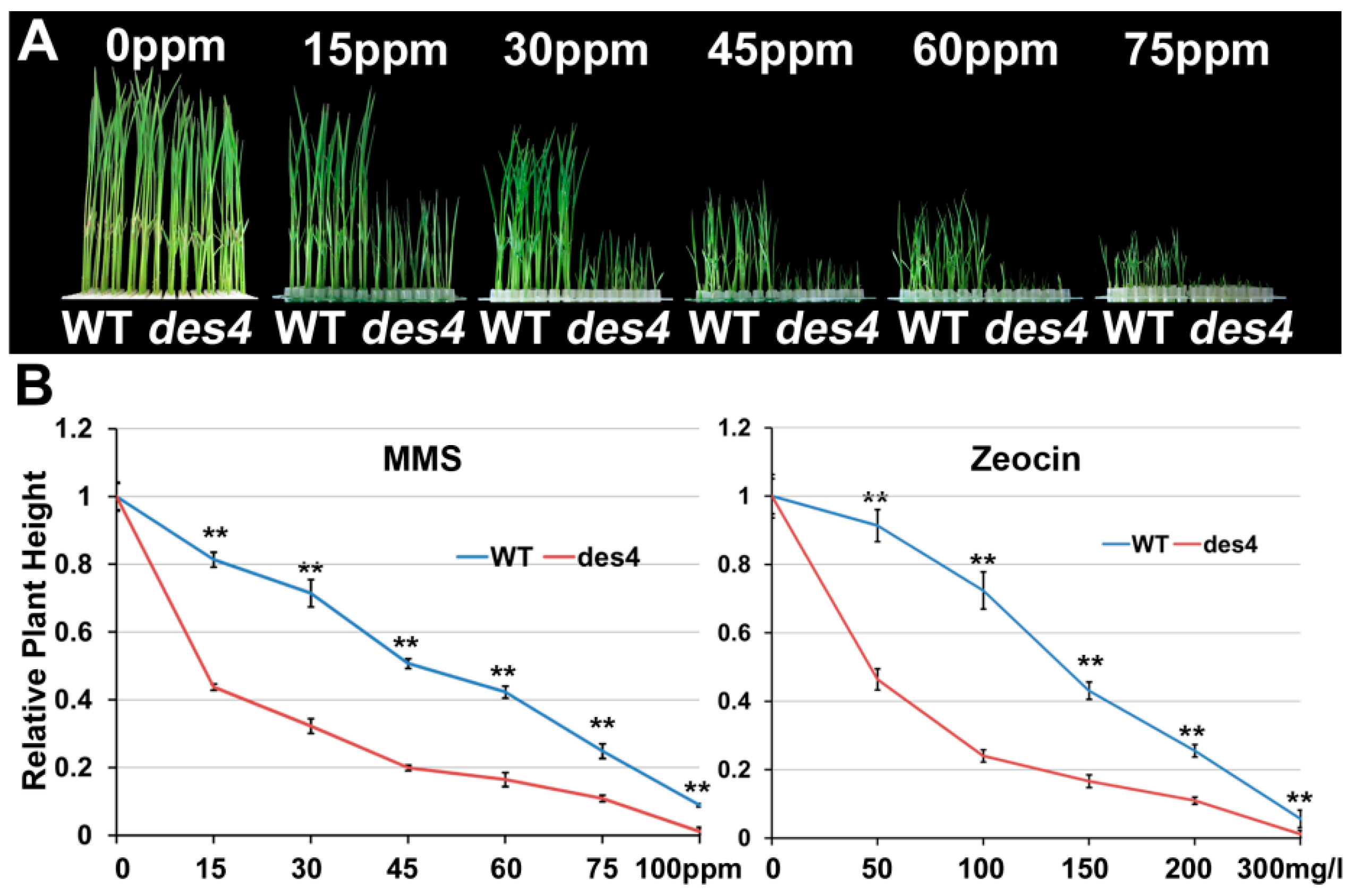
3.4. des4 Is Hypersensitive to Genotoxic Stress
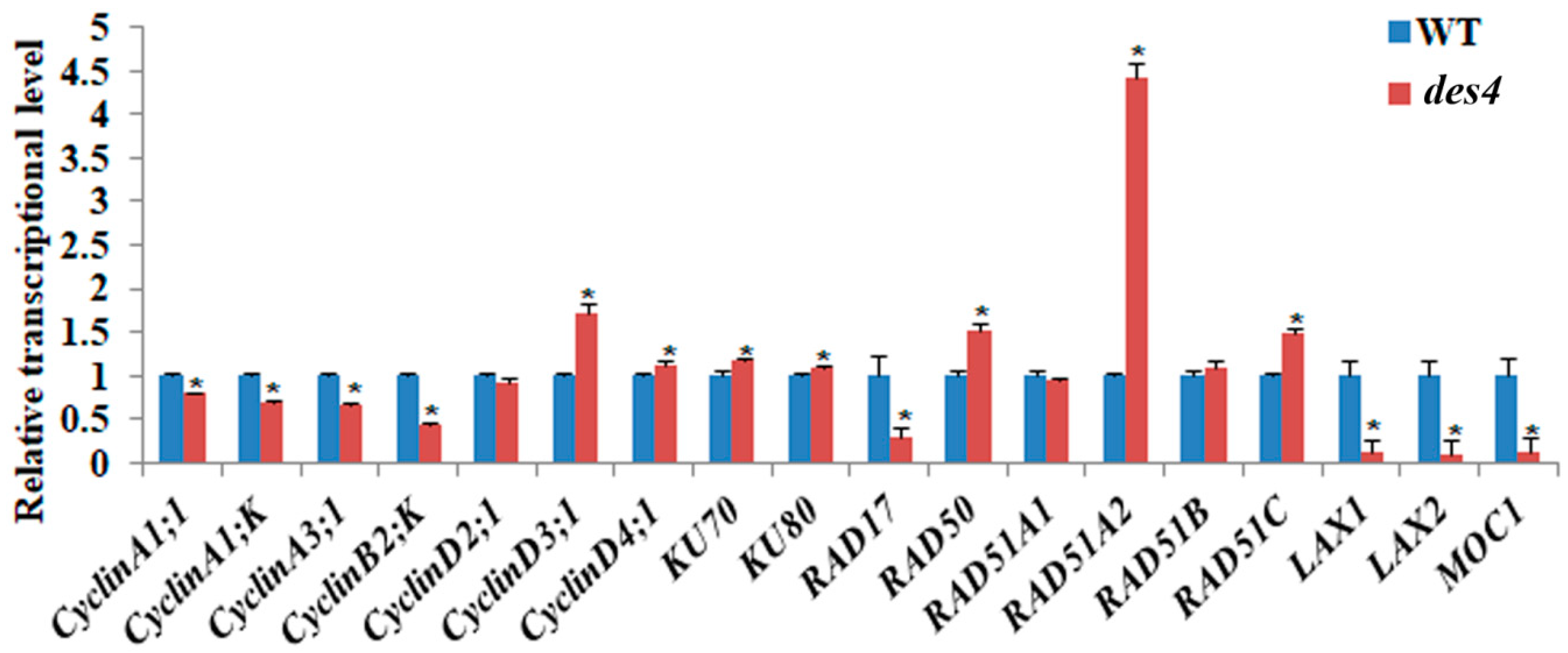
3.5. DSB Repair Genes and Spikelet Number Genes Are Differentially Regulated in des4
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhang, J.; Guo, D.; Chang, Y.; You, C.; Li, X.; Dai, X.; Weng, Q.; Zhang, J.; Chen, G.; Li, X.; et al. Non-random distribution of T-DNA insertions at various levels of the genome hierarchy as revealed by analyzing 13 804 T-DNA flanking sequences from an enhancer-trap mutant library. Plant J. Cell Mol. Biol. 2007, 49, 947–959. [Google Scholar] [CrossRef] [PubMed]
- Xing, Y.; Zhang, Q. Genetic and molecular bases of rice yield. Annu. Rev. Plant Biol. 2010, 61, 421–442. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, K.; Sunohara, H.; Nagato, Y. Developmental course of inflorescence and spikelet in rice. Jpn. J. Breed. 2004, 54, 147–156. [Google Scholar] [CrossRef]
- Bai, X.; Huang, Y.; Hu, Y.; Liu, H.; Zhang, B.; Smaczniak, C.; Hu, G.; Han, Z.; Xing, Y. Duplication of an upstream silencer of FZP increases grain yield in rice. Nat. Plants 2017, 3, 885. [Google Scholar] [CrossRef]
- Zhang, D.; Yuan, Z. Molecular control of grass inflorescence development. Annu. Rev. Plant Biol. 2014, 65, 553–578. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, J. Branching in rice. Curr. Opin. Plant Biol. 2011, 14, 94–99. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, X.; Fu, D.; Wu, C. Panicle Morphology Mutant 1 (PMM1) determines the inflorescence architecture of rice by controlling brassinosteroid biosynthesis. BMC Plant Biol. 2018, 18, 348. [Google Scholar] [CrossRef]
- Komatsu, M.; Maekawa, M.; Shimamoto, K.; Kyozuka, J. The LAX1 and FRIZZY PANICLE 2 genes determine the inflorescence architecture of rice by controlling rachis-branch and spikelet development. Dev. Biol. 2001, 231, 364–373. [Google Scholar] [CrossRef] [PubMed]
- Keishi, K.; Masahiko, M.; Shin, U.; Yuzuki, S.; Ikuyo, F.; Hironobu, O.; Ko, S.; Junko, K. LAX and SPA: Major regulators of shoot branching in rice. Proc. Natl. Acad. Sci. USA 2003, 100, 11765–11770. [Google Scholar]
- Tabuchi, H.; Zhang, Y.; Hattori, S.; Omae, M.; Shimizusato, S.; Oikawa, T.; Qian, Q.; Nishimura, M.; Kitano, H.; Xie, H. LAX PANICLE2 of rice encodes a novel nuclear protein and regulates the formation of axillary meristems. Plant Cell 2011, 23, 3276–3287. [Google Scholar] [CrossRef]
- Ashikari, M.; Sakakibara, H.; Lin, S.; Yamamoto, T.; Takashi, T.; Nishimura, A.; Angeles, E.R.; Qian, Q.; Kitano, H.; Matsuoka, M. Cytokinin oxidase regulates rice grain production. Science 2005, 309, 741–745. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Qian, Q.Z.; Sun, H.; He, S.; Luo, D.; Xia, G.; Chu, C.; Li, J.; Fu, X. Natural variation at the DEP1 locus enhances grain yield in rice. Nat. Genet. 2009, 41, 494–497. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Liu, W.; Tang, J.; Chen, J.; Tong, H.; Hu, B.; Li, C.; Fang, J.; Chen, M.; Chu, C. Rice DENSE AND ERECT PANICLE 2 is essential for determining panicle outgrowth and elongation. Cell Res. 2010, 20, 838–849. [Google Scholar] [CrossRef]
- Sun, H.Y.; Qian, Q.; Wu, K.; Wang, W.; Luo, J.J.; Wang, S.S.; Zhang, C.W.; Ma, Y.F.; Liu, Q.; Huang, X.Z. Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Sci. Found. China 2015, 46, 652. [Google Scholar] [CrossRef]
- Hiei, Y.; Komari, T.; Kubo, T. Transformation of rice mediated by Agrobacterium tumefaciens. Plant Mol. Biol. 1997, 35, 205–218. [Google Scholar] [CrossRef] [PubMed]
- Bello, B.K.; Hou, Y.; Zhao, J.; Jiao, G.; Wu, Y.; Li, Z.; Wang, Y.; Tong, X.; Wang, W.; Yuan, W.; et al. NF-YB1-YC12-bHLH144 complex directly activates Wx to regulate grain quality in rice (Oryza sativa L.). Plant Biotechnol. J. 2018. [Google Scholar] [CrossRef]
- Zhang, J.; Nallamilli, B.R.; Mujahid, H.; Peng, Z. OsMADS6 plays an essential role in endosperm nutrient accumulation and is subject to epigenetic regulation in rice (Oryza sativa). Plant J. Cell Mol. Biol. 2010, 64, 604–617. [Google Scholar] [CrossRef]
- Sikorski, R.S.; Boguski, M.S.; Goebl, M.; Hieter, P. A repeating amino acid motif in CDC23 defines a family of proteins and a new relationship among genes required for mitosis and RNA synthesis. Cell 1990, 60, 307–317. [Google Scholar] [CrossRef]
- Goebl, M.; Yanagida, M. The TPR snap helix: A novel protein repeat motif from mitosis to transcription. Trends Biochem. Sci. 1991, 16, 173. [Google Scholar] [CrossRef]
- Kobe, B.; Deisenhofer, J. Crystal structure of porcine ribonuclease inhibitor, a protein with leucine-rich repeats. Nature 1993, 366, 751–756. [Google Scholar] [CrossRef]
- Kobe, B.; Deisenhofer, J. The leucine-rich repeat: A versatile binding motif. Trends Biochem. Sci. 1994, 19, 415–421. [Google Scholar] [CrossRef]
- Guyomarc’H, S.; Vernoux, T.; Traas, J.; Zhou, D.X.; Delarue, M. MGOUN3, an Arabidopsis gene with TetratricoPeptide-Repeat-related motifs, regulates meristem cellular organization. J. Exp. Bot. 2004, 55, 673–684. [Google Scholar] [CrossRef]
- Suzuki, T.; Inagaki, S.; Nakajima, S.; Akashi, T.; Ohto, M.A.; Kobayashi, M.; Seki, M.; Shinozaki, K.; Kato, T.; Tabata, S. A novel Arabidopsis gene TONSOKU is required for proper cell arrangement in root and shoot apical meristems. Plant J. 2010, 38, 673–684. [Google Scholar] [CrossRef] [PubMed]
- Takeda, S.; Tadele, Z.; Hofmann, I.; Aline, V.P.; Angelis, K.J.; Hidetaka, K.; Takashi, A.; Tesfaye, M.; Ortrun, M.S.; et al. BRU1, a novel link between responses to DNA damage and epigenetic gene silencing in Arabidopsis. Genes Dev. 2004, 18, 782–793. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, J.L. Monofunctional alkylating agent-induced S-phase-dependent DNA damage. Mutat. Res. 1989, 216, 111–118. [Google Scholar] [CrossRef]
- Green, C.M.; Almouzni, G. When repair meets chromatin. EMBO Rep. 2002, 3, 28–33. [Google Scholar] [CrossRef] [PubMed]
- Feaver, W.J.; Svejstrup, J.Q.; Bardwell, L.; Bardwell, A.J.; Buratowski, S.; Gulyas, K.D.; Donahue, T.F.; Friedberg, E.C.; Kornberg, R.D. Dual roles of a multiprotein complex from S. cerevisiae in transcription and DNA repair. Cell 1993, 75, 1379. [Google Scholar] [CrossRef]
- Culligan, K.; Tissier, A.; Britt, A. ATR regulates a G2-phase cell-cycle checkpoint in Arabidopsis thaliana. Plant Cell 2004, 16, 1091. [Google Scholar] [CrossRef] [PubMed]
- Ricaud, L.; Proux, C.; Renou, J.-P.; Pichon, O.; Fochesato, S.; Ortet, P.; Montané, M.-H. ATM-mediated transcriptional and developmental responses to γ-rays in Arabidopsis. PLoS ONE 2007, 2, e430. [Google Scholar] [CrossRef] [PubMed]
- Culligan, K.M.; Robertson, C.E.; Julia, F.; Peter, D.; Britt, A.B. ATR and ATM play both distinct and additive roles in response to ionizing radiation. Plant J. 2010, 48, 947–961. [Google Scholar] [CrossRef] [PubMed]
- Lieber, M.R.; Karanjawala, Z.E. Ageing, repetitive genomes and DNA damage. Nat. Rev. Mol. Cell Biol. 2004, 5, 69–75. [Google Scholar] [CrossRef]
- Dynan, W.S.; Yoo, S. Interaction of Ku protein and DNA-dependent protein kinase catalytic subunit with nucleic acids. Nucleic Acids Res. 1998, 26, 1551–1559. [Google Scholar] [CrossRef]
- Morozumi, Y.; Ino, R.; Ikawa, S.; Mimida, N.; Shimizu, T.; Toki, S.; Ichikawa, H.; Shibata, T.; Kurumizaka, H. Homologous pairing activities of two rice RAD51 proteins, RAD51A1 and RAD51A2. PLoS ONE 2013, 8, e75451. [Google Scholar] [CrossRef]
- Kou, Y.; Chang, Y.; Li, X.; Xiao, J.; Wang, S. The rice RAD51C gene is required for the meiosis of both female and male gametocytes and the DNA repair of somatic cells. J. Exp. Bot. 2012, 63, 5323–5335. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ni, S.; Li, Z.; Ying, J.; Zhang, J.; Chen, H. Decreased Spikelets 4 Encoding a Novel Tetratricopeptide Repeat Domain-Containing Protein Is Involved in DNA Repair and Spikelet Number Determination in Rice. Genes 2019, 10, 214. https://doi.org/10.3390/genes10030214
Ni S, Li Z, Ying J, Zhang J, Chen H. Decreased Spikelets 4 Encoding a Novel Tetratricopeptide Repeat Domain-Containing Protein Is Involved in DNA Repair and Spikelet Number Determination in Rice. Genes. 2019; 10(3):214. https://doi.org/10.3390/genes10030214
Chicago/Turabian StyleNi, Shen, Zongzhu Li, Jiancheng Ying, Jian Zhang, and Hongqi Chen. 2019. "Decreased Spikelets 4 Encoding a Novel Tetratricopeptide Repeat Domain-Containing Protein Is Involved in DNA Repair and Spikelet Number Determination in Rice" Genes 10, no. 3: 214. https://doi.org/10.3390/genes10030214
APA StyleNi, S., Li, Z., Ying, J., Zhang, J., & Chen, H. (2019). Decreased Spikelets 4 Encoding a Novel Tetratricopeptide Repeat Domain-Containing Protein Is Involved in DNA Repair and Spikelet Number Determination in Rice. Genes, 10(3), 214. https://doi.org/10.3390/genes10030214





