RNA-Seq Transcriptome Analysis of Rice Primary Roots Reveals the Role of Flavonoids in Regulating the Rice Primary Root Growth
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Condition
2.2. Measurement of Roots
2.3. Histochemical Staining for ROS Detection of H2O2 and O2−
2.4. RNA Sequencing (RNA-seq) and Data Analysis
2.5. RNA Extraction and Real-Time Quantitative PCR (qRT-PCR)
2.6. Assays of Superoxide dismutase (SOD), Peroxidase (POD), and Catalase (CAT)Activities
2.7. DR5-GUS Staining
3. Results
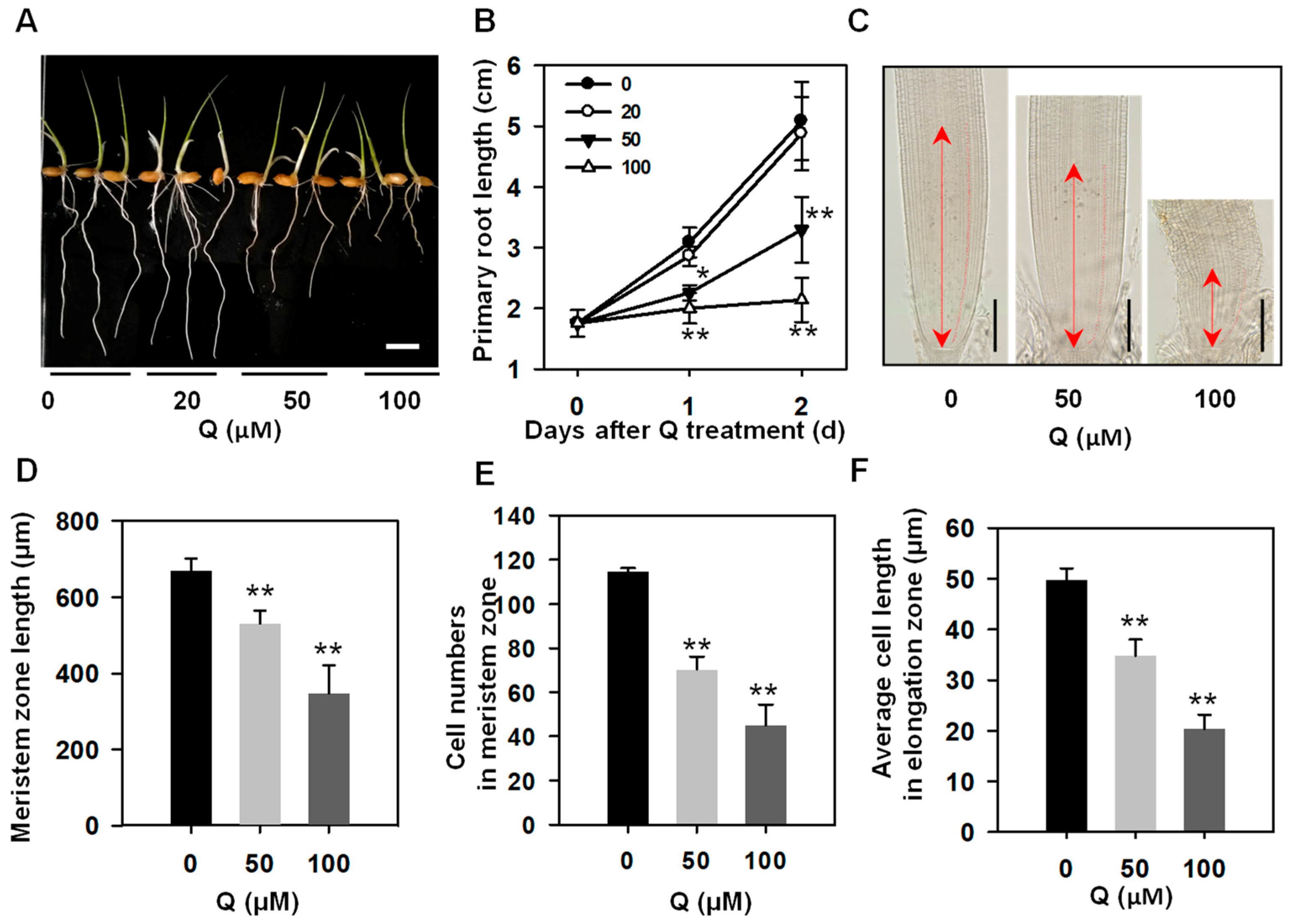
3.1. Treatment with Quercetin Inhibited Rice Root Growth
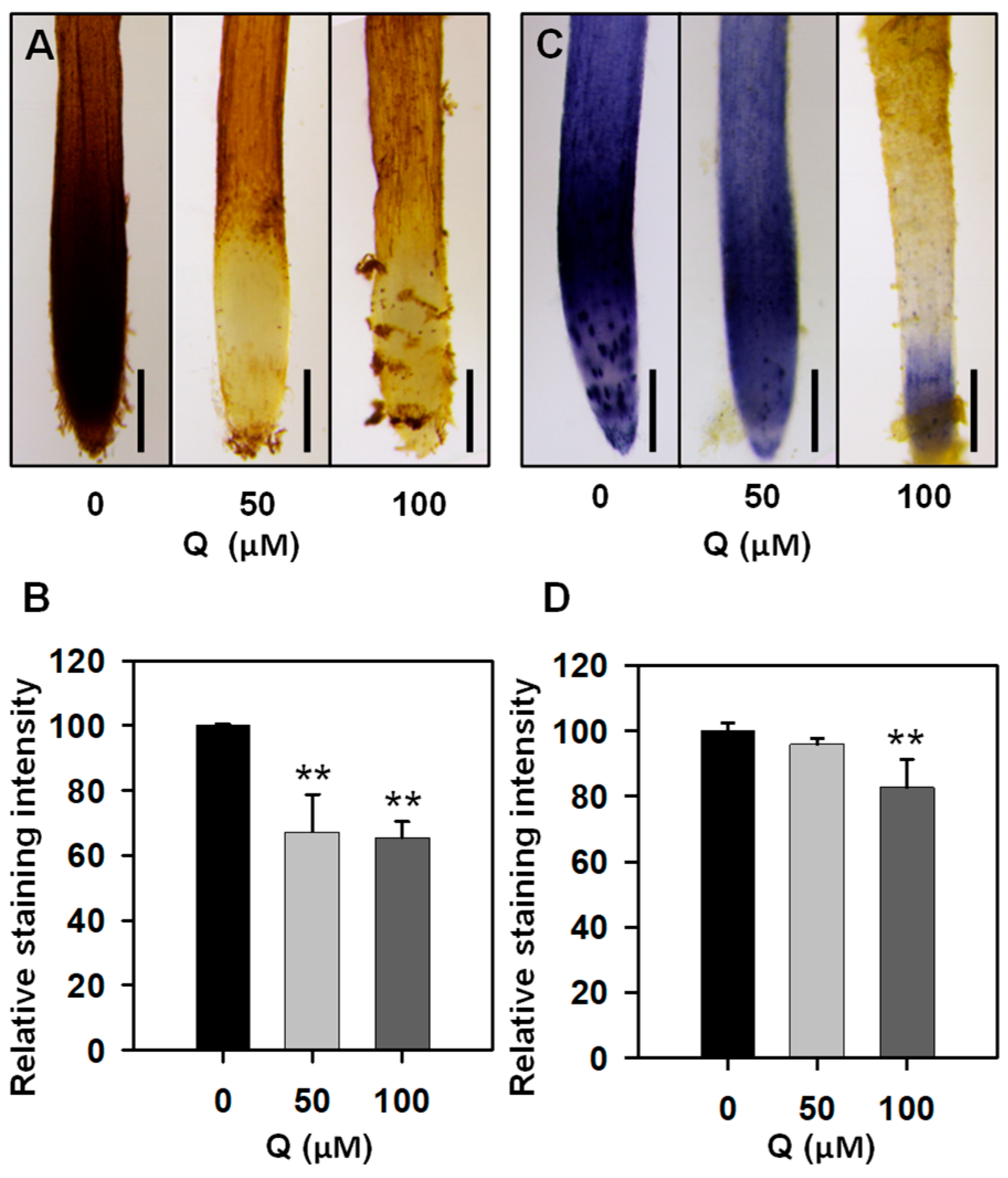
3.2. Treatment with Quercetin Reduced ROS Levels in the Root Tips
3.3. The Enriched GO Terms were Mainly Associated with Cell Wall Organization, Oxidative Stress, and Plant Hormone Signal Transduction
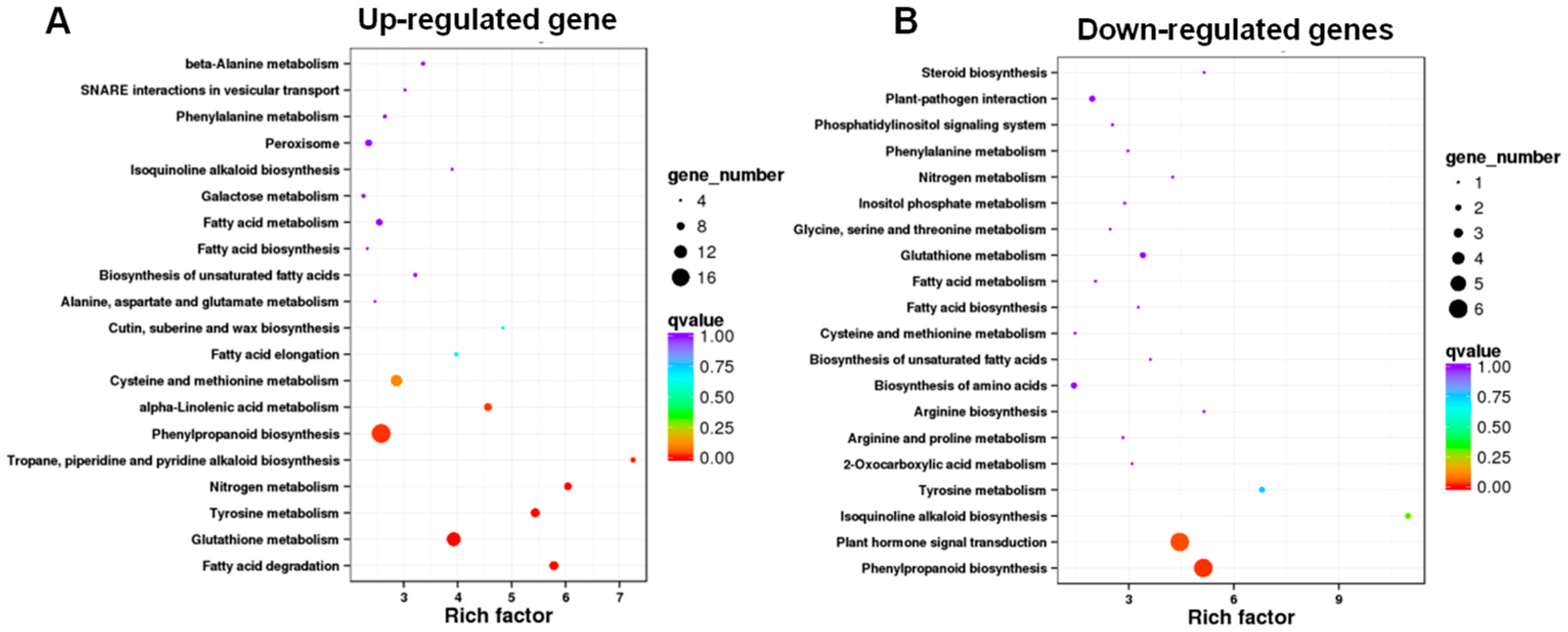
3.4. KEGG Pathway Enrichment Analysis Revealed That the Enriched DEGs Are Involved in Phenylpropanoid Biosynthesis, Glutathione Metabolism, and Plant Hormone Signal Transduction
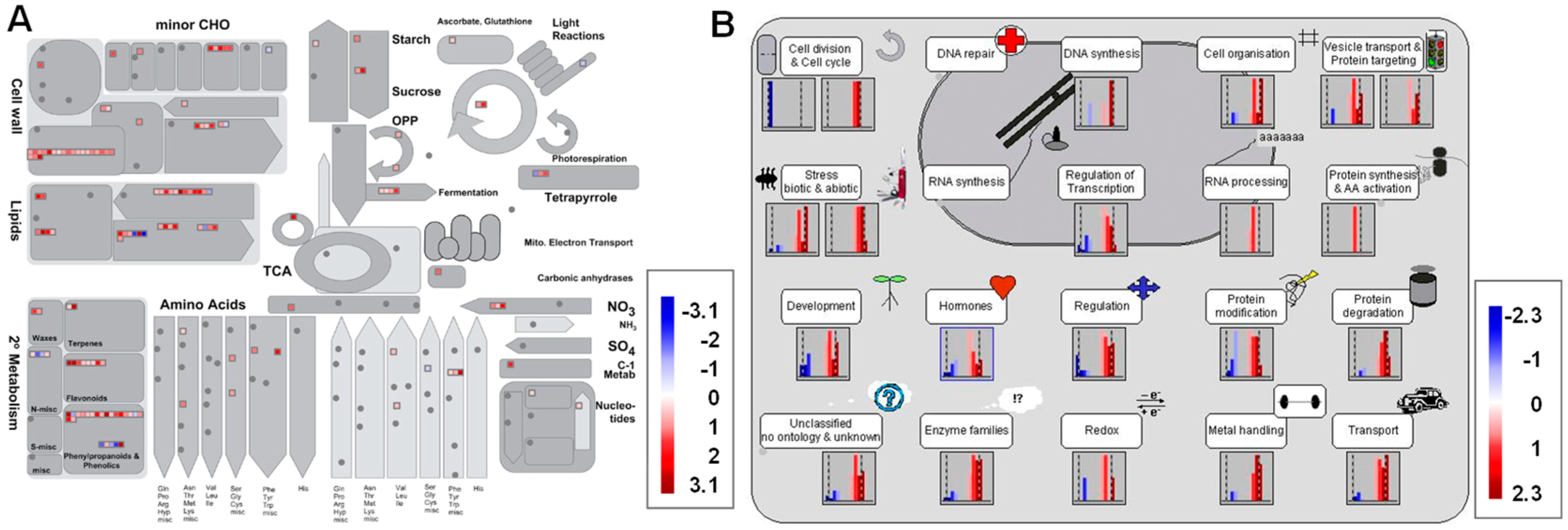
3.5. Metabolism Pathway Analysis of DEGs Using MapMan
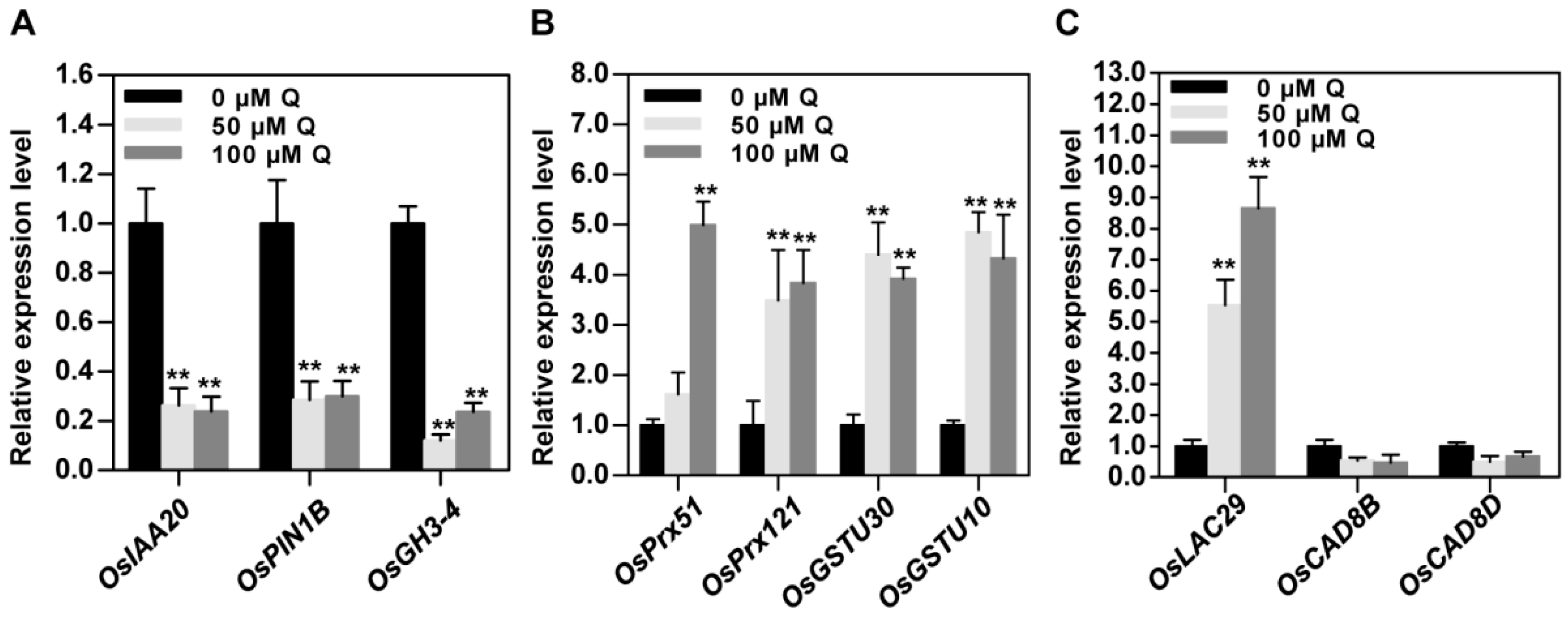
3.6. Validation of RNA-Seq Data by qRT-PCR
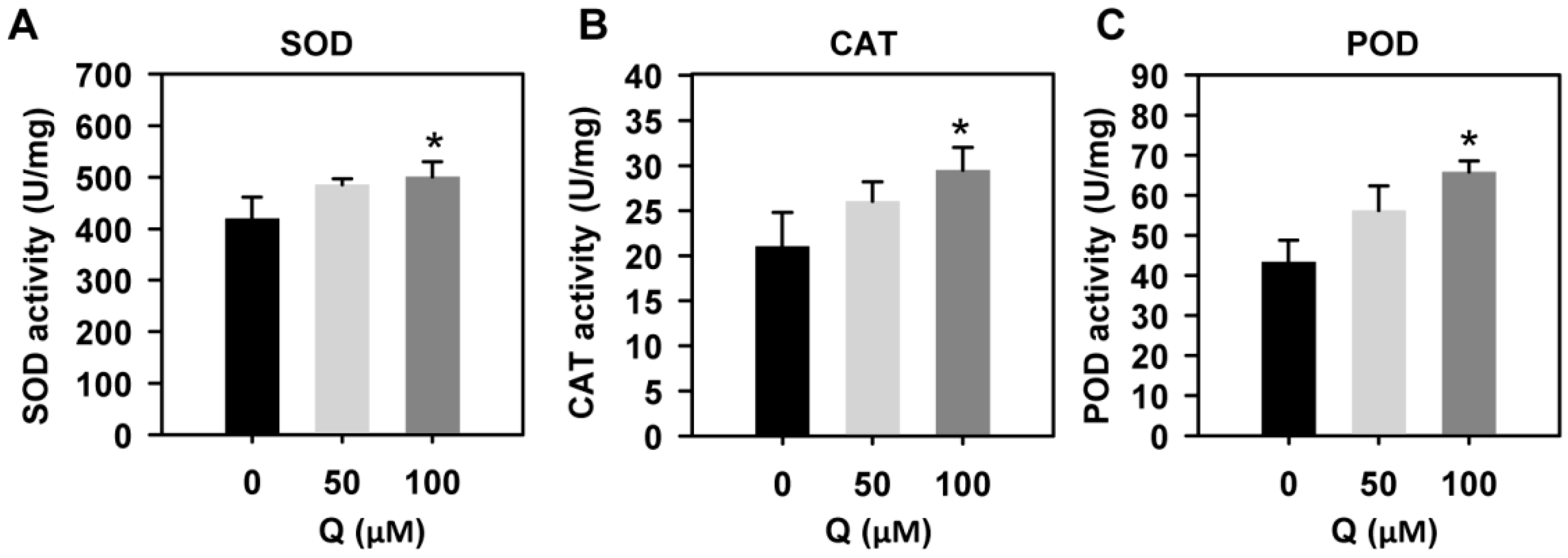
3.7. Quercetin Treatment Enhanced the Activities of Antioxidant Enzymes
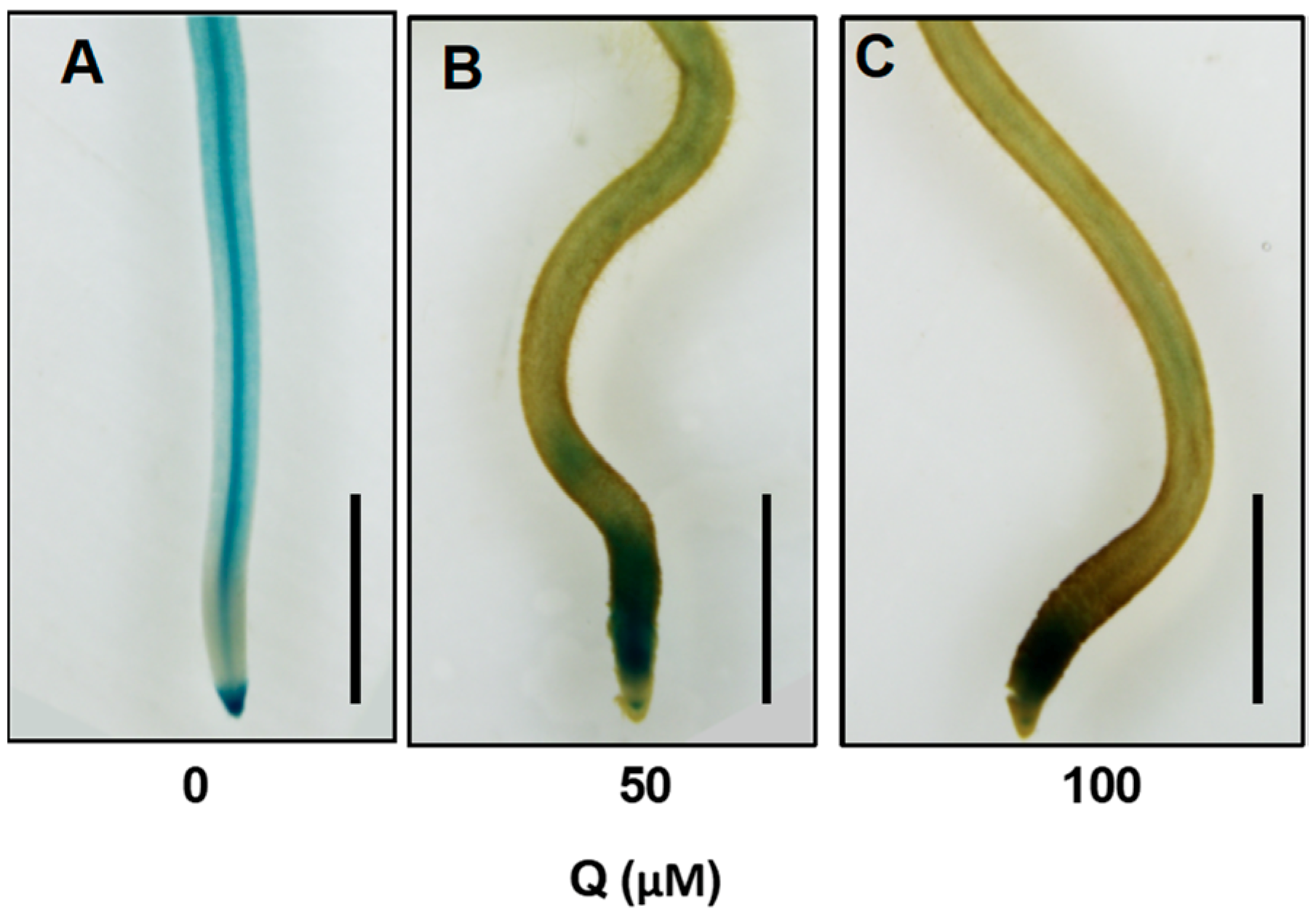
3.8. DR5:GUS Expression Pattern in the Roots Was Altered after Quercetin Treatment
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Winkel-Shirley, B. Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol. 2001, 126, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Ferreyra, M.L.F.; Rius, S.P.; Casati, P. Flavonoids: Biosynthesis, biological functions, and biotechnological applications. Front. Plant Sci. 2012, 3. [Google Scholar] [CrossRef]
- Zhao, L.; Gao, L.; Wang, H.; Chen, X.; Wang, Y.; Yang, H.; Wei, C.; Wan, X.; Xia, T. The R2R3-MYB, bHLH, WD40, and related transcription factors in flavonoid biosynthesis. Funct. Integr. Genom. 2013, 13, 75–98. [Google Scholar] [CrossRef] [PubMed]
- Afrin, S.; Nuruzzaman, M.; Zhu, J.; Luo, Z.-Y. Combinatorial interactions of MYB and bHLH in flavonoid biosynthesis and their function in plants. J. Plant Biol. Res. 2014, 3, 65–77. [Google Scholar]
- Kootstra, A. Protection from UV-B-induced DNA damage by flavonoids. Plant Mol. Biol. 1994, 26, 771–774. [Google Scholar] [CrossRef] [PubMed]
- Agati, G.; Azzarello, E.; Pollastri, S.; Tattini, M. Flavonoids as antioxidants in plants: Location and functional significance. Plant Sci. 2012, 196, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Peer, W.A.; Murphy, A.S. Flavonoids as signal molecules: Targets of flavonoid action. Sci. Flavonoids 2006, 239–268. [Google Scholar] [CrossRef]
- Watkins, J.M.; Chapman, J.M.; Muday, G.K. Abscisic acid-induced reactive oxygen species are modulated by flavonols to control stomata aperture. Plant Physiol. 2017, 175, 1807–1825. [Google Scholar] [CrossRef] [PubMed]
- Slovak, R.; Ogura, T.; Satbhai, S.B.; Ristova, D.; Busch, W. Genetic control of root growth: From genes to networks. Ann. Bot. 2016, 117, 9–24. [Google Scholar] [CrossRef] [PubMed]
- Tsukagoshi, H. Control of root growth and development by reactive oxygen species. Curr. Opin. Plant Biol. 2016, 29, 57–63. [Google Scholar] [CrossRef]
- Murphy, A.; Peer, W.A.; Taiz, L. Regulation of auxin transport by aminopeptidases and endogenous flavonoids. Planta 2000, 211, 315–324. [Google Scholar] [CrossRef]
- Peer, W.A.; Brown, D.E.; Tague, B.W.; Muday, G.K.; Taiz, L.; Murphy, A.S. Flavonoid accumulation patterns of transparent testa mutants of Arabidopsis. Plant Physiol. 2001, 126, 536–548. [Google Scholar] [CrossRef] [PubMed]
- Imin, N.; Nizamidin, M.; Wu, T.; Rolfe, B.G. Factors involved in root formation in Medicago truncatula. J. Exp. Bot. 2007, 58, 439–451. [Google Scholar] [CrossRef] [PubMed]
- Maloney, G.S.; DiNapoli, K.T.; Muday, G.K. The anthocyanin reduced tomato mutant demonstrates the role of flavonols in tomato lateral root and root hair development. Plant Physiol. 2014, 166, 614–631. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.E.; Rashotte, A.M.; Murphy, A.S.; Normanly, J.; Tague, B.W.; Peer, W.A.; Taiz, L.; Muday, G.K. Flavonoids act as negative regulators of auxin transport in vivo in Arabidopsis. Plant Physiol. 2001, 126, 524–535. [Google Scholar] [CrossRef] [PubMed]
- Buer, C.S.; Kordbacheh, F.; Truong, T.T.; Hocart, C.H.; Djordjevic, M.A. Alteration of flavonoid accumulation patterns in transparent testa mutants disturbs auxin transport, gravity responses, and imparts long-term effects on root and shoot architecture. Planta 2013, 238, 171–189. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Yang, J.; Zhang, M.; Fan, W.J.; Firon, N.; Pattanaik, S.; Yuan, L.; Zhang, P. Altered phenylpropanoid metabolism in the maize Lc-expressed sweet potato (Ipomoea batatas) affects storage root development. Sci. Rep. 2016, 6, 18645. [Google Scholar] [CrossRef]
- Peer, W.; Bandyopadhyay, A.; Blakeslee, J.J.; Makam, S.N.; Chen, R.J.; Masson, P.H.; Murphy, A.S. Variation in expression and protein localization of the PIN family of auxin efflux facilitator proteins in flavonoid mutants with altered auxin transport in Arabidopsis thaliana. Plant Cell 2004, 16, 1898–1911. [Google Scholar] [CrossRef]
- Buer, C.S.; Muday, G.K. The transparent testa4 mutation prevents flavonoid synthesis and alters auxin transport and the response of Arabidopsis roots to gravity and light. Plant Cell 2004, 16, 1191–1205. [Google Scholar] [CrossRef]
- Zeng, J.; Wang, Q.; Lin, J.; Deng, K.; Zhao, X.; Tang, D.; Liu, X. Arabidopsis cryptochrome-1 restrains lateral roots growth by inhibiting auxin transport. J. Plant Physiol. 2010, 167, 670–673. [Google Scholar] [CrossRef]
- Santelia, D.; Henrichs, S.; Vincenzetti, V.; Sauer, M.; Bigler, L.; Klein, M.; Bailly, A.; Lee, Y.; Friml, J.; Geisler, M.; et al. Flavonoids redirect PIN-mediated polar auxin fluxes during root gravitropic responses. J. Biol. Chem. 2008, 283, 31218–31226. [Google Scholar] [CrossRef]
- Silva-Navas, J.; Moreno-Risueno, M.A.; Manzano, C.; Tellez-Robledo, B.; Navarro-Neila, S.; Carrasco, V.; Pollmann, S.; Gallego, F.J.; del Pozo, J.C. Flavonols mediate root phototropism and growth through regulation of proliferation-to-differentiation transition. Plant Cell 2016, 28, 1372–1387. [Google Scholar] [CrossRef]
- Wan, J.; Zhang, P.; Wang, R.; Sun, L.; Wang, W.; Zhou, H.; Xu, J. UV-B radiation induces root bending through the flavonoid-mediated auxin pathway in Arabidopsis. Front. Plant Sci. 2018, 9, 618. [Google Scholar] [CrossRef]
- Peer, W.A.; Cheng, Y.; Murphy, A.S. Evidence of oxidative attenuation of auxin signalling. J. Exp. Bot. 2013, 64, 2629–2639. [Google Scholar] [CrossRef]
- Naoumkina, M.; Dixon, R.A. Subcellular localization of flavonoid natural products: A signaling function? Plant Signal. Behav. 2008, 3, 573–575. [Google Scholar] [CrossRef]
- Brunetti, C.; Fini, A.; Sebastiani, F.; Gori, A.; Tattini, M. Modulation of phytohormone signaling: A primary function of flavonoids in plant-environment interactions. Front. Plant Sci. 2018, 9, 1042. [Google Scholar] [CrossRef]
- Franco, D.M.; Silva, E.M.; Saldanha, L.L.; Adachi, S.A.; Schley, T.R.; Rodrigues, T.M.; Dokkedal, A.L.; Nogueira, F.T.S.; Rolim de Almeida, L.F. Flavonoids modify root growth and modulate expression of SHORT-ROOT and HD-ZIP III. J. Plant Physiol. 2015, 188, 89–95. [Google Scholar] [CrossRef]
- Saslowsky, D.E.; Warek, U.; Winkel, B.S.J. Nuclear localization of flavonoid enzymes in Arabidopsis. J. Biol. Chem. 2005, 280, 23735–23740. [Google Scholar] [CrossRef]
- Wang, L.; Feng, Z.; Wang, X.; Wang, X.; Zhang, X. DEGseq: An R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef]
- Tian, T.; Liu, Y.; Yan, H.; You, Q.; Yi, X.; Du, Z.; Xu, W.; Su, Z. agriGO v2.0: A GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 2017, 45, W122–W129. [Google Scholar] [CrossRef]
- Mao, X.; Cai, T.; Olyarchuk, J.G.; Wei, L. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 2005, 21, 3787–3793. [Google Scholar] [CrossRef]
- Thimm, O.; Blasing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Kruger, P.; Selbig, J.; Muller, L.A.; Rhee, S.Y.; Stitt, M. MAPMAN: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004, 37, 914–939. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Pandhair, V.; Sekhon, B.S. Reactive oxygen species and antioxidants in plants: An overview. J. Plant Biochem. Biot. 2006, 15, 71–78. [Google Scholar] [CrossRef]
- Qureshi, M.K.; Munir, S.; Shahzad, A.N.; Rasul, S.; Nouman, W.; Aslam, K. Role of reactive oxygen species and contribution of new players in defense mechanism under drought stress in rice. Int. J. Agric. Biol. 2018, 20, 1339–1352. [Google Scholar] [CrossRef]
- Francoz, E.; Ranocha, P.; Nguyen-Kim, H.; Jamet, E.; Burlat, V.; Dunand, C. Roles of cell wall peroxidases in plant development. Phytochemistry 2015, 112, 15–21. [Google Scholar] [CrossRef]
- Tsukagoshi, H.; Busch, W.; Benfey, P.N. Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root. Cell 2010, 143, 606–616. [Google Scholar] [CrossRef]
- Smith, A.P.; Nourizadeh, S.D.; Peer, W.A.; Xu, J.; Bandyopadhyay, A.; Murphy, A.S.; Goldsbrough, P.B. Arabidopsis AtGSTF2 is regulated by ethylene and auxin, and encodes a glutathione S-transferase that interacts with flavonoids. Plant J. 2003, 36, 433–442. [Google Scholar] [CrossRef]
- Jain, M.; Kaur, N.; Tyagi, A.K.; Khurana, J.P. The auxin-responsive GH3 gene family in rice (Oryza sativa). Funct. Integr. Genom. 2006, 6, 36–46. [Google Scholar] [CrossRef]
- Dunand, C.; Crèvecoeur, M.; Penel, C. Distribution of superoxide and hydrogen peroxide in Arabidopsis root and their influence on root development: Possible interaction with peroxidases. New Phytol. 2007, 174, 332–341. [Google Scholar] [CrossRef]
- Xu, L.; Zhao, H.; Ruan, W.; Deng, M.; Wang, F.; Peng, J.; Luo, J.; Chen, Z.; Yi, K. ABNORMAL INFLORESCENCE MERISTEM1 functions in salicylic acid biosynthesis to maintain proper reactive oxygen species levels for root meristem activity in Rice. Plant Cell 2017, 29, 560–574. [Google Scholar] [CrossRef]
- Grunewald, W.; De Smet, I.; Lewis, D.R.; Löfke, C.; Jansen, L.; Goeminne, G.; Bossche, R.V.; Karimi, M.; De Rybel, B.; Vanholme, B.; et al. Transcription factor WRKY23 assists auxin distribution patterns during Arabidopsis root development through local control on flavonol biosynthesis. Proc. Natl. Acad. Sci. USA 2012, 109, 1554–1559. [Google Scholar] [CrossRef]
- Dietz, K.J. Redox regulation of transcription factors in plant stress acclimation and development. Antioxid. Redox Signal. 2014, 21, 1356–1372. [Google Scholar] [CrossRef]
- Kuhn, B.M.; Nodzyński, T.; Errafi, S.; Bucher, R.; Gupta, S.; Aryal, B.; Dobrev, P.; Bigler, L.; Geisler, M.; Zažímalová, E.; et al. Flavonol-induced changes in PIN2 polarity and auxin transport in the Arabidopsis thaliana rol1-2 mutant require phosphatase activity. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Gayomba, S.R.; Watkins, J.M.; Muday, G.K. Flavonols regulate plant growth and development through regulation of auxin transport and cellular redox status. In Recent Advances in Polyphenol Research, 1st ed.; John Wiley & Sons, Ltd.: Chichester, UK, 2016. [Google Scholar] [CrossRef]
- Lewis, D.R.; Ramirez, M.V.; Miller, N.D.; Vallabhaneni, P.; Ray, W.K.; Helm, R.F.; Winkel, B.S.; Muday, G.K. Auxin and ethylene induce flavonol accumulation through distinct transcriptional networks. Plant Physiol. 2011, 156, 144–164. [Google Scholar] [CrossRef]
- Opassiri, R.; Pomthong, B.; Onkoksoong, T.; Akiyama, T.; Esen, A.; Ketudat Cairns, J.R. Analysis of rice glycosyl hydrolase family 1 and expression of Os4bglu12 β-glucosidase. BMC Plant Biol. 2006, 6, 33. [Google Scholar] [CrossRef]
- Cosgrove, D.J. Loosening of plant cell walls by expansins. Nature 2000, 407, 321–326. [Google Scholar] [CrossRef]
- Brunetti, C.; Di Ferdinando, M.; Fini, A.; Pollastri, S.; Tattini, M. Flavonoids as antioxidants and developmental regulators: Relative significance in plants and humans. Int. J. Mol. Sci. 2013, 14, 3540–3555. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, Y.; Zou, J.; Zheng, H.; Xu, M.; Zong, X.; Wang, L. RNA-Seq Transcriptome Analysis of Rice Primary Roots Reveals the Role of Flavonoids in Regulating the Rice Primary Root Growth. Genes 2019, 10, 213. https://doi.org/10.3390/genes10030213
Xu Y, Zou J, Zheng H, Xu M, Zong X, Wang L. RNA-Seq Transcriptome Analysis of Rice Primary Roots Reveals the Role of Flavonoids in Regulating the Rice Primary Root Growth. Genes. 2019; 10(3):213. https://doi.org/10.3390/genes10030213
Chicago/Turabian StyleXu, Yu, Junjie Zou, Hongyan Zheng, Miaoyun Xu, Xuefeng Zong, and Lei Wang. 2019. "RNA-Seq Transcriptome Analysis of Rice Primary Roots Reveals the Role of Flavonoids in Regulating the Rice Primary Root Growth" Genes 10, no. 3: 213. https://doi.org/10.3390/genes10030213
APA StyleXu, Y., Zou, J., Zheng, H., Xu, M., Zong, X., & Wang, L. (2019). RNA-Seq Transcriptome Analysis of Rice Primary Roots Reveals the Role of Flavonoids in Regulating the Rice Primary Root Growth. Genes, 10(3), 213. https://doi.org/10.3390/genes10030213






