RNA Sequencing Reveals Specific Transcriptomic Signatures Distinguishing Effects of the [SWI+] Prion and SWI1 Deletion in Yeast Saccharomyces cerevisiae
Abstract
:1. Introduction
2. Materials and Methods
2.1. Yeast Strains, Plasmids, and Cultivation Conditions
2.2. DNA Preparation and qPCR
2.3. RNA Preparation and Whole Transcriptome RNA Sequencing (RNA-Seq)
2.4. RNA-Seq Read Processing, Pseudoalignment, and Data Analysis
2.5. GO Term Overrepresentation Test
2.6. KEGG Pathway Mapping
3. Results
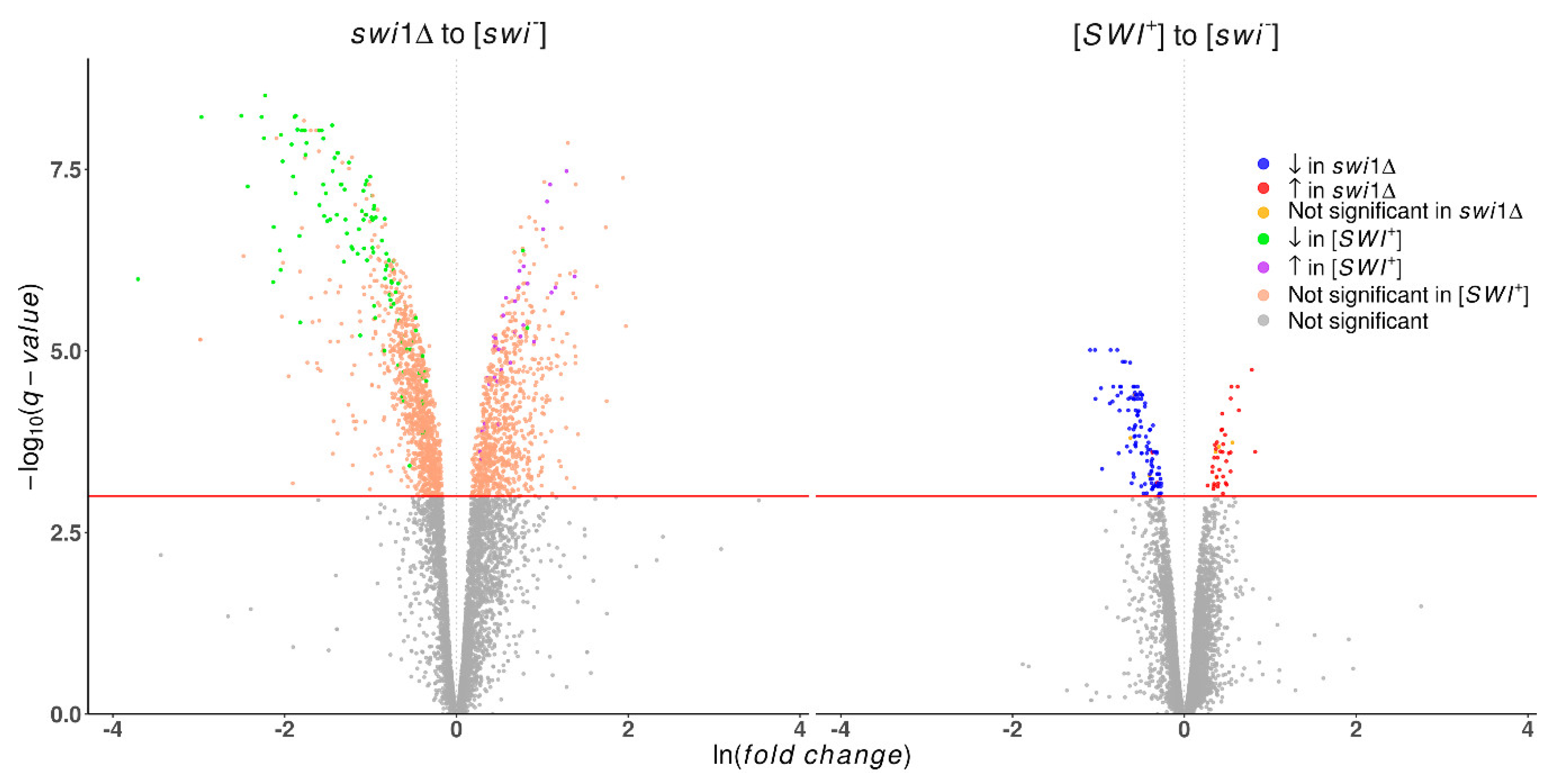
3.1. Transcriptome-Wide Effects of the Swi1 Prion Formation and Deletion of its Structural Gene are Not Equal
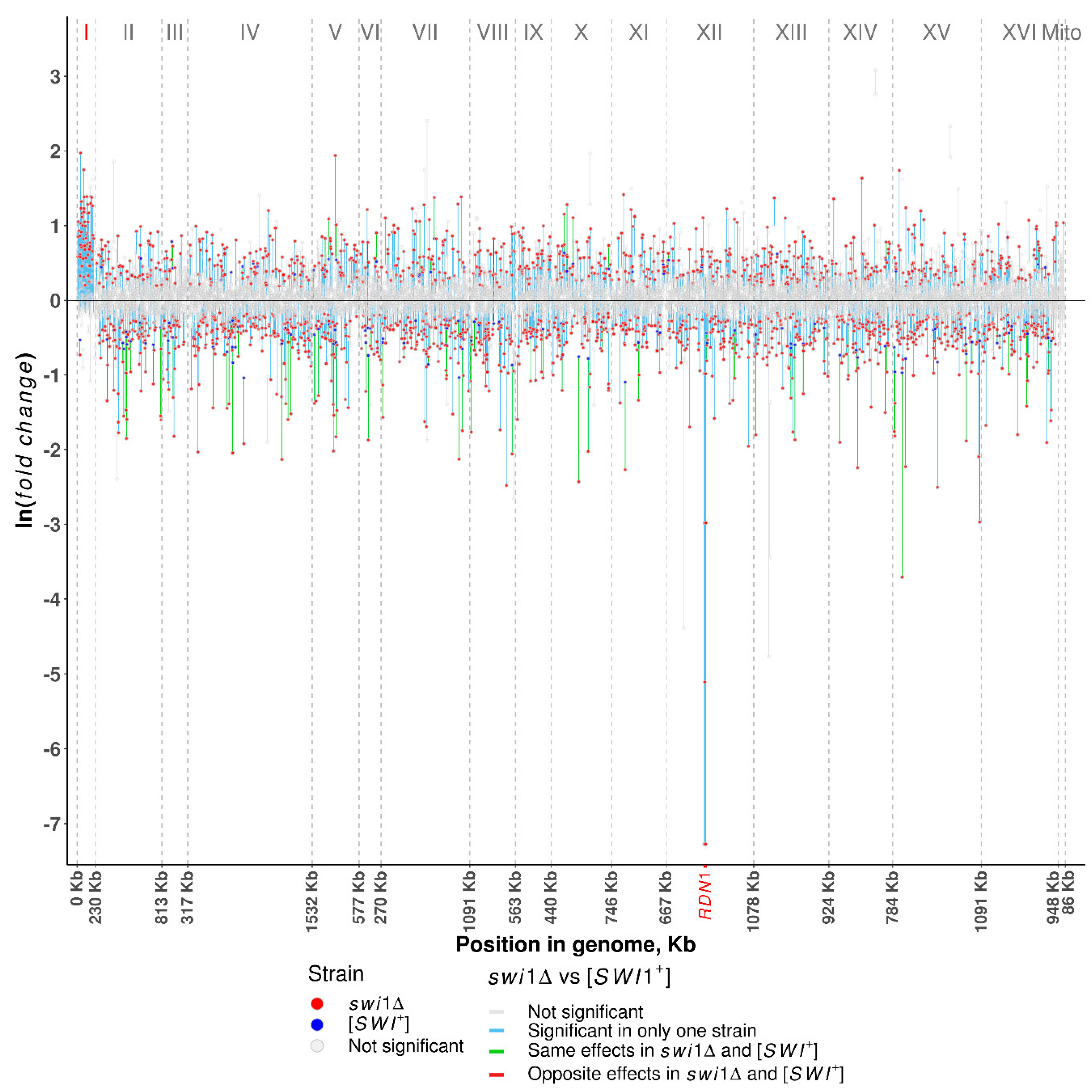
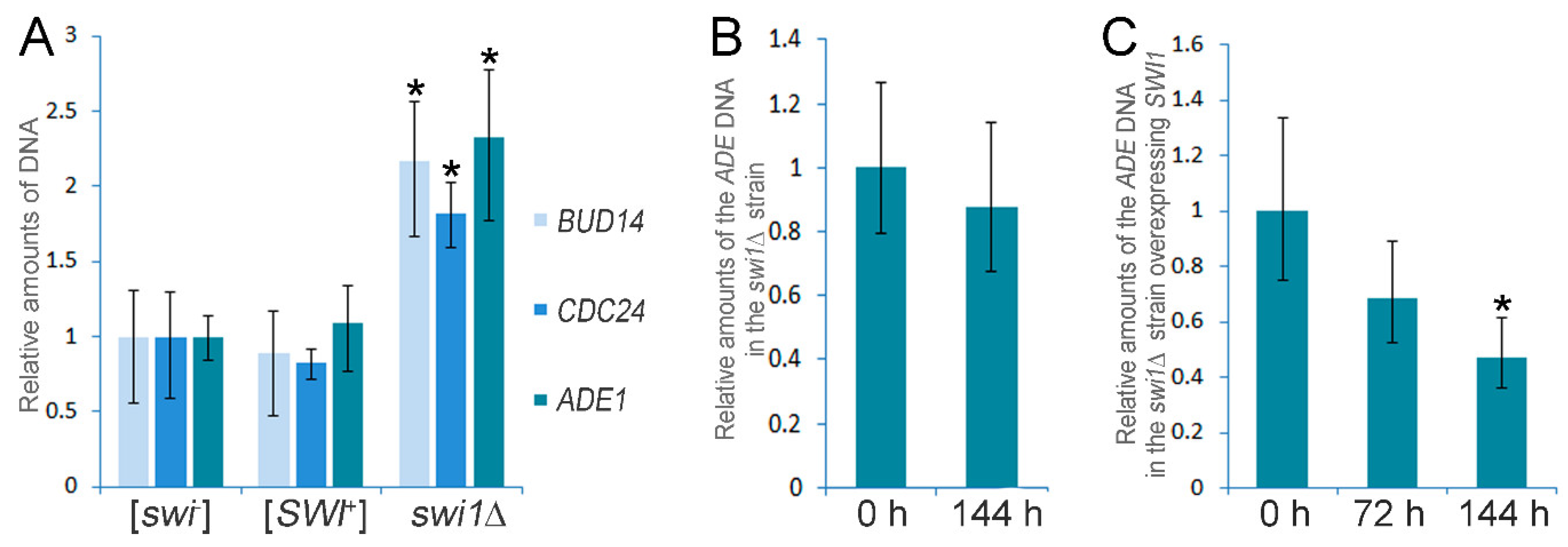
3.2. SWI1 Deletion Causes the Chromosome I Disomy
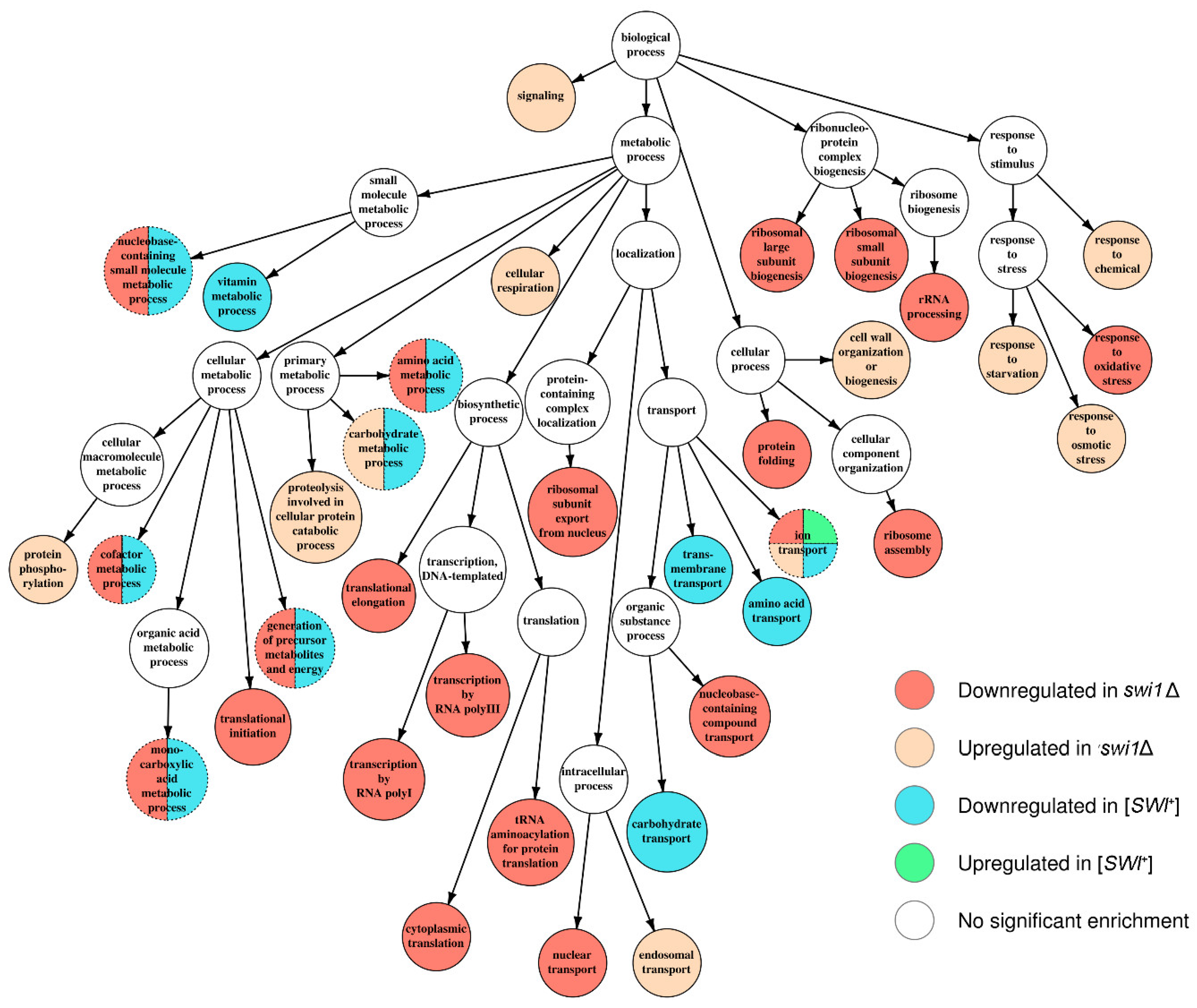
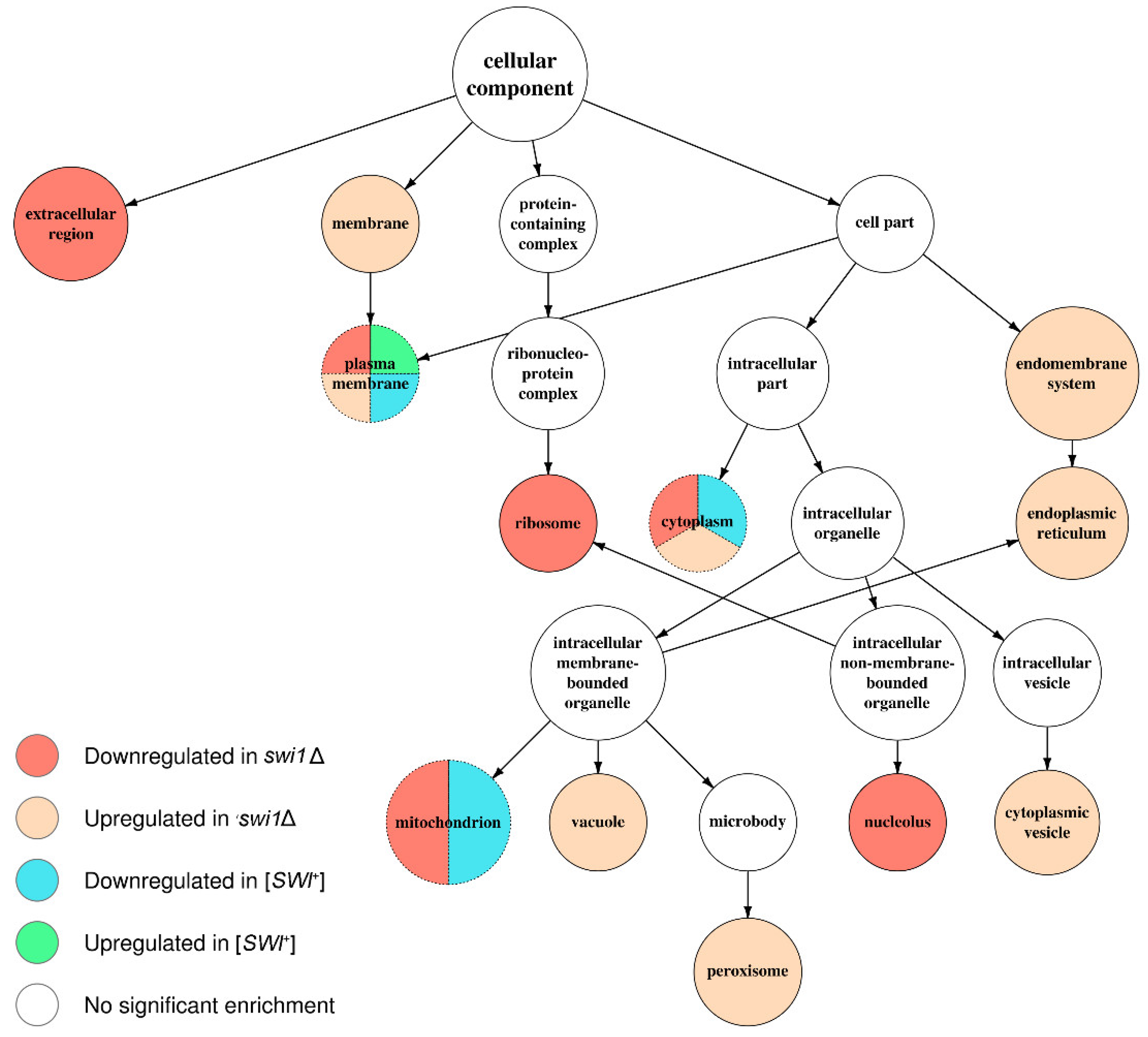
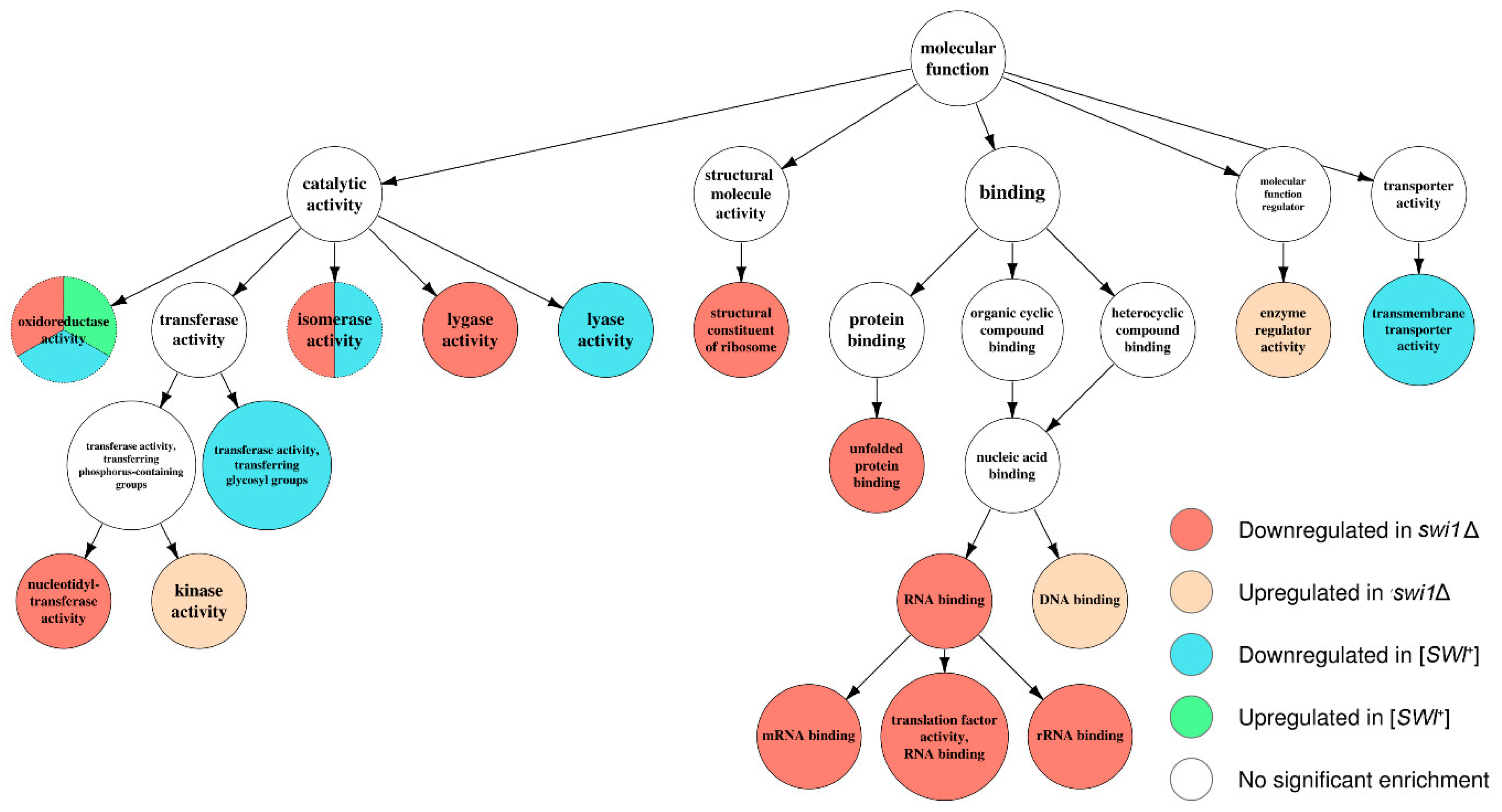
3.3. SWI1 Deletion Modulates a Higher Number of Biological Processes and Molecular Functions than the [SWI+] Prion
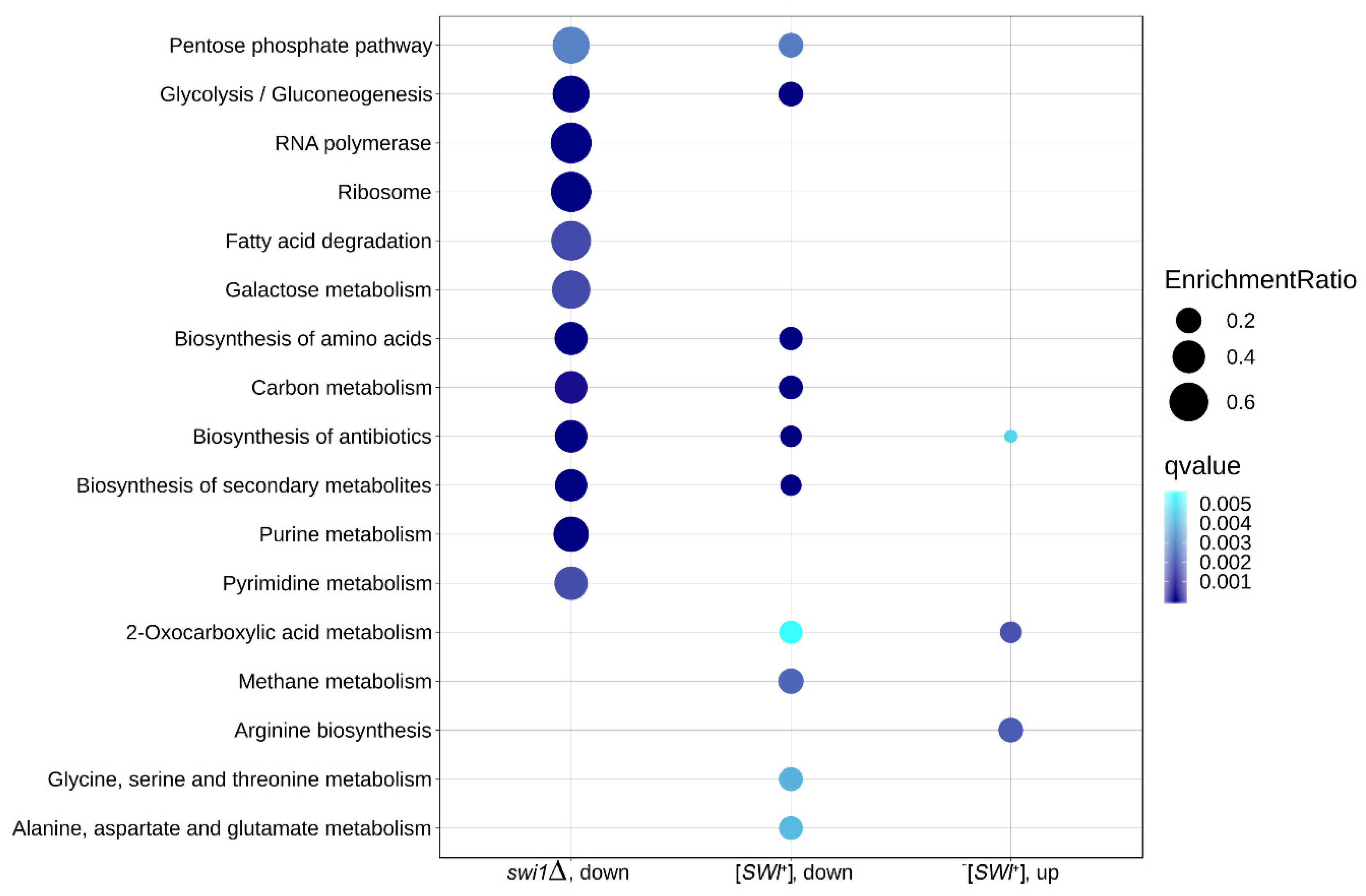
3.4. The Effects of SWI1 Deletion and [SWI+] Prion on Metabolic Pathways Correlate with Their Phenotypic Manifestations
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- McKinley, M.P.; Bolton, D.C.; Prusiner, S.B. A protease-resistant protein is a structural component of the scrapie prion. Cell 1983, 35, 57–62. [Google Scholar] [CrossRef]
- Chernova, T.A.; Wilkinson, K.D.; Chernoff, Y.O. Physiological and environmental control of yeast prions. FEMS Microbiol. Rev. 2014, 38, 326–344. [Google Scholar] [CrossRef] [PubMed]
- Wickner, R.B.; Shewmaker, F.P.; Bateman, D.A.; Edskes, H.K.; Gorkovskiy, A.; Dayani, Y.; Bezsonov, E.E. Yeast Prions: Structure, Biology, and Prion-Handling Systems. Microbiol. Mol. Biol. Rev. 2015, 79. [Google Scholar] [CrossRef] [PubMed]
- Wickner, R.B.; Kelly, A.C. Prions are affected by evolution at two levels. Cell. Mol. Life Sci. 2016, 73, 1131–1144. [Google Scholar] [CrossRef]
- Nizhnikov, A.A.; Antonets, K.S.; Bondarev, S.A.; Inge-Vechtomov, S.G.; Derkatch, I.L. Prions, amyloids, and RNA: Pieces of a puzzle. Prion 2016, 10, 182–206. [Google Scholar] [CrossRef] [PubMed]
- Patel, B.K.; Gavin-Smyth, J.; Liebman, S.W. The yeast global transcriptional co-repressor protein Cyc8 can propagate as a prion. Nat. Cell Biol. 2009, 11, 344–349. [Google Scholar] [CrossRef]
- Du, Z.; Park, K.K.-W.; Yu, H.; Fan, Q.; Li, L. Newly identified prion linked to the chromatin-remodeling factor Swi1 in Saccharomyces cerevisiae. Nat. Genet. 2008, 40, 460–465. [Google Scholar] [CrossRef]
- Wickner, R.B. [URE3] as an altered URE2 protein: Evidence for a prion analog in Saccharomyces cerevisiae. Science 1994, 264, 566–569. [Google Scholar] [CrossRef]
- Alberti, S.; Halfmann, R.; King, O.; Kapila, A.; Lindquist, S. A systematic survey identifies prions and illuminates sequence features of prionogenic proteins. Cell 2009, 137, 146–158. [Google Scholar] [CrossRef]
- Suzuki, G.; Shimazu, N.; Tanaka, M. A yeast prion, Mod5, promotes acquired drug resistance and cell survival under environmental stress. Science 2012, 336, 355–359. [Google Scholar] [CrossRef]
- Rogoza, T.; Goginashvili, A.; Rodionova, S.; Ivanov, M.; Viktorovskaya, O.; Rubel, A.; Volkov, K.; Mironova, L. Non-Mendelian determinant [ISP+] in yeast is a nuclear-residing prion form of the global transcriptional regulator Sfp1. Proc. Natl. Acad. Sci. USA 2010, 107, 10573–10577. [Google Scholar] [CrossRef]
- Michelitsch, M.D.; Weissman, J.S. A census of glutamine/asparagine-rich regions: Implications for their conserved function and the prediction of novel prions. Proc. Natl. Acad. Sci. USA 2000, 97, 11910–11915. [Google Scholar] [CrossRef]
- McGlinchey, R.P.; Kryndushkin, D.; Wickner, R.B. Suicidal [PSI+] is a lethal yeast prion. Proc. Natl. Acad. Sci. USA 2011, 108, 5337–5341. [Google Scholar] [CrossRef]
- Halfmann, R.; Jarosz, D.F.; Jones, S.K.; Chang, A.; Lancaster, A.K.; Lindquist, S. Prions are a common mechanism for phenotypic inheritance in wild yeasts. Nature 2012, 482, 363–368. [Google Scholar] [CrossRef]
- Holmes, D.L.; Lancaster, A.K.; Lindquist, S.; Halfmann, R. Heritable remodeling of yeast multicellularity by an environmentally responsive prion. Cell 2013, 153, 153–165. [Google Scholar] [CrossRef]
- Wickner, R.B.; Edskes, H.K.; Bateman, D.; Kelly, A.C.; Gorkovskiy, A. The yeast prions [PSI+] and [URE3] are molecular degenerative diseases. Prion 2011, 5, 258–262. [Google Scholar] [CrossRef]
- Wickner, R.B. Yeast and fungal prions. Cold Spring Harb. Perspect. Biol. 2016, 8, a023531. [Google Scholar] [CrossRef]
- Dechassa, M.L.; Zhang, B.; Horowitz-Scherer, R.; Persinger, J.; Woodcock, C.L.; Peterson, C.L.; Bartholomew, B. Architecture of the SWI/SNF-nucleosome complex. Mol. Cell. Biol. 2008, 28, 6010–6021. [Google Scholar] [CrossRef]
- Tang, L.; Nogales, E.; Ciferri, C. Structure and function of SWI/SNF chromatin remodeling complexes and mechanistic implications for transcription. Prog. Biophys. Mol. Biol. 2010, 102, 122–128. [Google Scholar] [CrossRef]
- Smith, C.L.; Horowitz-Scherer, R.; Flanagan, J.F.; Woodcock, C.L.; Peterson, C.L. Structural analysis of the yeast SWI/SNF chromatin remodeling complex. Nat. Struct. Biol. 2003, 10, 141–145. [Google Scholar] [CrossRef]
- Schubert, H.L.; Wittmeyer, J.; Kasten, M.M.; Hinata, K.; Rawling, D.C.; Héroux, A.; Cairns, B.R.; Hill, C.P. Structure of an actin-related subcomplex of the SWI/SNF chromatin remodeler. Proc. Natl. Acad. Sci. USA 2013, 110, 3345–3350. [Google Scholar] [CrossRef]
- Neigeborn, L.; Carlson, M. Genes affecting the regulation of SUC2 gene expression by glucose repression in Saccharomyces cerevisiae. Genetics 1984, 108, 845–858. [Google Scholar]
- Peterson, C.L.; Herskowitz, I. Characterization of the yeast SWI1, SWI2, and SWI3 genes, which encode a global activator of transcription. Cell 1992, 68, 573–583. [Google Scholar] [CrossRef]
- Sudarsanam, P.; Iyer, V.R.; Brown, P.O.; Winston, F. Whole-genome expression analysis of snf/swi mutants of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 2000, 97, 3364–3369. [Google Scholar] [CrossRef]
- Shivaswamy, S.; Iyer, V.R. Stress-dependent dynamics of global chromatin remodeling in yeast: Dual role for SWI/SNF in the heat shock stress response. Mol. Cell. Biol. 2008, 28, 2221–2234. [Google Scholar] [CrossRef]
- Dutta, A.; Gogol, M.; Kim, J.H.; Smolle, M.; Venkatesh, S.; Gilmore, J.; Florens, L.; Washburn, M.P.; Workman, J.L. Swi/Snf dynamics on stress-responsive genes is governed by competitive bromodomain interactions. Genes Dev. 2014, 28, 2314–2330. [Google Scholar] [CrossRef]
- Kadoch, C.; Hargreaves, D.C.; Hodges, C.; Elias, L.; Ho, L.; Ranish, J.; Crabtree, G.R. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet. 2013, 45, 592–601. [Google Scholar] [CrossRef]
- Breeden, L.; Nasmyth, K. Cell cycle control of the yeast HO gene: Cis- and Trans-acting regulators. Cell 1987, 48, 389–397. [Google Scholar] [CrossRef]
- Hirschhorn, J.N.; Brown, S.A.; Clark, C.D.; Winston, F. Evidence that SNF2/SWI2 and SNF5 activate transcription in yeast by altering chromatin structure. Genes Dev. 1992, 6, 2288–2298. [Google Scholar] [CrossRef]
- Dutta, A.; Sardiu, M.; Gogol, M.; Gilmore, J.; Zhang, D.; Florens, L.; Abmayr, S.M.; Washburn, M.P.; Workman, J.L. Composition and Function of Mutant Swi/Snf Complexes. Cell Rep. 2017. [Google Scholar] [CrossRef]
- Neely, K.E.; Hassan, A.H.; Brown, C.E.; Howe, L.; Workman, J.L. Transcription activator interactions with multiple SWI/SNF subunits. Mol. Cell. Biol. 2002, 22, 1615–1625. [Google Scholar] [CrossRef]
- Haber, J.E.; Garvik, B. A new gene affecting the efficiency of mating-type interconversions in homothallic strains of Saccharomyces cerevisiae. Genetics 1977, 87, 33–50. [Google Scholar]
- Taguchi, A.K.; Young, E.T. The cloning and mapping of ADR6, a gene required for sporulation and for expression of the alcohol dehydrogenase II isozyme from Saccharomyces cerevisiae. Genetics 1987, 116, 531–540. [Google Scholar]
- Du, Z.; Crow, E.T.; Kang, H.S.; Li, L. Distinct Subregions of Swi1 Manifest Striking Differences in Prion Transmission and SWI/SNF Function. Mol. Cell. Biol. 2010, 30, 4644–4655. [Google Scholar] [CrossRef]
- Nizhnikov, A.A.; Magomedova, Z.M.; Rubel, A.A.; Kondrashkina, A.M.; Inge-Vechtomov, S.G.; Galkin, A.P. Determinant has a pleiotropic phenotypic manifestation that is modulated by SUP35, SUP45, and VTS1 genes. Curr. Genet. 2012, 58, 35–47. [Google Scholar] [CrossRef]
- Du, Z.; Zhang, Y.; Li, L. The Yeast Prion [SWI+] Abolishes Multicellular Growth by Triggering Conformational Changes of Multiple Regulators Required for Flocculin Gene Expression. Cell Rep. 2015, 13, 2865–2878. [Google Scholar] [CrossRef]
- Saifitdinova, A.F.; Nizhnikov, A.A.; Lada, A.G.; Rubel, A.A.; Magomedova, Z.M.; Ignatova, V.V.; Inge-Vechtomov, S.G.; Galkin, A.P. [NSI+]: A novel non-Mendelian nonsense suppressor determinant in Saccharomyces cerevisiae. Curr. Genet. 2010, 56, 467–478. [Google Scholar] [CrossRef]
- Zhouravleva, G.; Frolova, L.; Le Goff, X.; Le Guellec, R.; Inge-Vechtomov, S.; Kisselev, L.; Philippe, M. Termination of translation in eukaryotes is governed by two interacting polypeptide chain release factors, eRF1 and eRF3. EMBO J. 1995, 14, 4065–4072. [Google Scholar] [CrossRef]
- Stansfield, I.; Jones, K.M.; Ter-Avanesyan, M.D.; Tuite, M.F. The products of the SUP45 (eRF1) and SUP35 genes interact to mediate translation termination in Saccharomyces cerevisiae. EMBO J. 1995, 14, 4365–4373. [Google Scholar] [CrossRef]
- Nizhnikov, A.A.; Kondrashkina, A.M.; Galkin, A.P. Interactions of [NSI+] prion-like determinant with SUP35 and VTS1 genes in Saccharomyces cerevisiae. Russ. J. Genet. 2013, 49, 1004–1012. [Google Scholar] [CrossRef]
- Kondrashkina, A.M.; Antonets, K.S.; Galkin, A.P.; Nizhnikov, A.A. Prion-Like Determinant [NSI+] Decreases the Expression of the SUP45 Gene in Saccharomyces cerevisiae. Mol. Biol. 2014, 48, 688–693. [Google Scholar] [CrossRef]
- Nizhnikov, A.A.; Ryzhova, T.A.; Volkov, K.V.; Zadorsky, S.P.; Sopova, J.V.; Inge-Vechtomov, S.G.; Galkin, A.P. Interaction of Prions Causes Heritable Traits in Saccharomyces cerevisiae. PLoS Genet. 2016, 12, e1006504. [Google Scholar] [CrossRef]
- Antonets, K.S.; Kliver, S.F.; Polev, D.E.; Shuvalova, A.R.; Andreeva, E.A.; Inge-Vechtomov, S.G.; Nizhnikov, A.A. Distinct mechanisms of phenotypic effects of inactivation and prionization of Swi1 protein in Saccharomyces cerevisiae. Biochemistry 2017, 82, 1147–1157. [Google Scholar] [CrossRef]
- Wach, A.; Brachat, A.; Pöhlmann, R.; Philippsen, P. New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast 1994, 10, 1793–1808. [Google Scholar] [CrossRef]
- Kaiser, C.; Michaelis, S.; Mitchell, A.; Cold Spring Harbor Laboratory. Methods in Yeast Genetics: A Cold Spring Harbor Laboratory Course Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1994; ISBN 0879694513. [Google Scholar]
- Zakharov, I.A.; Kozhin, S.A.; Kozhina, T.N.; Fedorova, I.V. Collected Methods in Genetics of the Yeast Saccharomyces; Nauka: Leningrad, Russia, 1984. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Bushnell, B. BBMap: A Fast, Accurate, Splice-Aware Aligner; Lawrence Berkeley National Lab. (LBNL): Berkeley, CA, USA, 2014. [Google Scholar]
- Andrews, S. FastQC: A quality control tool for high throughput sequence data. Babraham Bioinform. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 20 December 2018).
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 2016, 34, 525–527. [Google Scholar] [CrossRef]
- Pimentel, H.; Bray, N.L.; Puente, S.; Melsted, P.; Pachter, L. Differential analysis of RNA-seq incorporating quantification uncertainty. Nat. Methods 2017, 14, 687–690. [Google Scholar] [CrossRef]
- Hochberg, B. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Alexa, A.; Rahnenfuhrer, J. topGO: Enrichment Analysis for Gene Ontology; R Packag. Version 2.34.0; Bioconductor, 2018. [Google Scholar] [CrossRef]
- Saccharomyces Genome Database (Index of/Curation/Literature). Available online: https://downloads.yeastgenome.org/curation/literature/go_slim_mapping.tab (accessed on 20 December 2018).
- Zhang, J.D.; Wiemann, S. KEGGgraph: A graph approach to KEGG PATHWAY in R and bioconductor. Bioinformatics 2009, 25, 1470–1471. [Google Scholar] [CrossRef]
- Csárdi, G.; Nepusz, T. The igraph software package for complex network research. Inter. J. Complex Syst. 2014, 1695–1704. [Google Scholar]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. clusterProfiler: An R Package for Comparing Biological Themes Among Gene Clusters. Omics J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Ginestet, C. ggplot2: Elegant Graphics for Data Analysis. J. R. Stat. Soc. Ser. A Stat. Soc. 2011, 174, 245–246. [Google Scholar] [CrossRef]
- Luo, W.; Brouwer, C. Pathview: An R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 2013, 29, 1830–1831. [Google Scholar] [CrossRef]
- Venema, J.; Tollervey, D. Ribosome Synthesis in Saccharomyces cerevisiae. Annu. Rev. Genet. 1999, 33, 261–311. [Google Scholar] [CrossRef]
- Giaever, G.; Chu, A.M.; Ni, L.; Connelly, C.; Riles, L.; Véronneau, S.; Dow, S.; Lucau-Danila, A.; Anderson, K.; André, B.; et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 2002, 418, 387–391. [Google Scholar] [CrossRef]
- Castrillo, J.I.; Zeef, L.A.; Hoyle, D.C.; Zhang, N.; Hayes, A.; Gardner, D.C.J.; Cornell, M.J.; Petty, J.; Hakes, L.; Wardleworth, L.; et al. Growth control of the eukaryote cell: A systems biology study in yeast. J. Biol. 2007, 6, 4. [Google Scholar] [CrossRef]
- Prokhorova, I.; Altman, R.B.; Djumagulov, M.; Shrestha, J.P.; Urzhumtsev, A.; Ferguson, A.; Chang, C.-W.T.; Yusupov, M.; Blanchard, S.C.; Yusupova, G. Aminoglycoside interactions and impacts on the eukaryotic ribosome. Proc. Natl. Acad. Sci. USA 2017, 114, E10899–E10908. [Google Scholar] [CrossRef]
- Dror, V.; Winston, F. The Swi/Snf Chromatin Remodeling Complex Is Required for Ribosomal DNA and Telomeric Silencing in Saccharomyces cerevisiae. Mol. Cell. Biol. 2004, 24, 8227–8235. [Google Scholar] [CrossRef]
- Venkataramanan, S.; Douglass, S.; Galivanche, A.R.; Johnson, T.L. The chromatin remodeling complex Swi/Snf regulates splicing of meiotic transcripts in Saccharomyces cerevisiae. Nucleic Acids Res. 2017. [Google Scholar] [CrossRef]
- Brownlee, P.M.; Meisenberg, C.; Downs, J.A. The SWI/SNF chromatin remodelling complex: Its role in maintaining genome stability and preventing tumourigenesis. DNA Repair (Amst.) 2015, 32, 127–133. [Google Scholar] [CrossRef]
- Chan, P.H.W.; Lee, L.; Kim, E.; Hui, T.; Stoynov, N.; Nassar, R.; Moksa, M.; Cameron, D.M.; Hirst, M.; Gsponer, J.; et al. The [PSI+] yeast prion does not wildly affect proteome composition whereas selective pressure exerted on [PSI+] cells can promote aneuploidy. Sci. Rep. 2017, 7, 8442. [Google Scholar] [CrossRef]
- Cox, B. Psi, a cytoplasmic supperssor of supersuppressors in yeast. Heredity 1965, 20, 505–521. [Google Scholar] [CrossRef]
- Drozdova, P.; Mironova, L.; Zhouravleva, G. Haploid yeast cells undergo a reversible phenotypic switch associated with chromosome II copy number. BMC Genet. 2016, 17, 152. [Google Scholar] [CrossRef]
- Zadorsky, S.P.; Sopova, Y.V.; Andreichuk, D.Y.; Startsev, V.A.; Medvedeva, V.P.; Inge-Vechtomov, S.G. Chromosome VIII disomy influences the nonsense suppression efficiency and transition metal tolerance of the yeast Saccharomyces cerevisiae. Yeast 2015, 32, 479–497. [Google Scholar] [CrossRef]
- Oromendia, A.B.; Dodgson, S.E.; Amon, A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 2012, 26, 2696–2708. [Google Scholar] [CrossRef]
- Reddy Pannala, V.; Jayadeva Bhat, P.; Bhartiya, S.; Venkatesh, K.V. Systems biology of GAL regulon in Saccharomyces cerevisiae. Wiley Interdiscip. Rev. Syst. Biol. Med. 2010, 2, 98–106. [Google Scholar] [CrossRef]
- Timson, D.J. Galactose Metabolism in Saccharomyces cerevisiae. Dyn. Biochem. Process Biotechnol. Mol. Biol. 2007, 1, 63–73. [Google Scholar] [CrossRef]
- Holden, H.M.; Rayment, I.; Thoden, J.B. Structure and Function of Enzymes of the Leloir Pathway for Galactose Metabolism. J. Biol. Chem. 2003, 278, 43885–43888. [Google Scholar] [CrossRef]
- Proft, M.; Struhl, K. Hog1 kinase converts the Sko1-Cyc8-Tup1 repressor complex into an activator that recruits SAGA and SWI/SNF in response to osmotic stress. Mol. Cell 2002, 9, 1307–1317. [Google Scholar] [CrossRef]
- Baccarini, L.; Martínez-Montañés, F.; Rossi, S.; Proft, M.; Portela, P. PKA-chromatin association at stress responsive target genes from Saccharomyces cerevisiae. Biochim. Biophys. Acta Gene Regul. Mech. 2015, 1849, 1329–1339. [Google Scholar] [CrossRef] [PubMed]
- Nemet, J.; Vidan, N.; Sopta, M. A meta-analysis reveals complex regulatory properties at Taf14-repressed genes. BMC Genom. 2017, 18, 175. [Google Scholar] [CrossRef] [PubMed]
- Tordella, L.; Khan, S.; Hohmeyer, A.; Banito, A.; Klotz, S.; Raguz, S.; Martin, N.; Dhamarlingam, G.; Carroll, T.; González Meljem, J.M.; et al. SWI/SNF regulates a transcriptional program that induces senescence to prevent liver cancer. Genes Dev. 2016, 30, 2187–2198. [Google Scholar] [CrossRef] [PubMed]
- Karpichev, I.V.; Small, G.M. Global Regulatory Functions of Oaf1p and Pip2p (Oaf2p), Transcription Factors That Regulate Genes Encoding Peroxisomal Proteins in Saccharomyces cerevisiae. Mol. Cell. Biol. 2015, 18, 6560–6570. [Google Scholar] [CrossRef]
- Smith, J.J.; Miller, L.R.; Kreisberg, R.; Vazquez, L.; Wan, Y.; Aitchison, J.D. Environment-responsive transcription factors bind subtelomeric elements and regulate gene silencing. Mol. Syst. Biol. 2011, 7, 455. [Google Scholar] [CrossRef]
Sample Availability: Raw Illumina reads for 24 samples in total are stored in NIH NCBI SRA under accession numbers SRR8552466 through SRR8552482. The SRA repository also refers to BioProject accession PRJNA521274 and BioSamples accessions SAMN10882201, SAMN10882202 and SAMN10882203, respectively. |

| Changes in Gene Expression | Comparison | ||
|---|---|---|---|
| [SWI+] to [swi−] | swi1Δ to [swi−] | swiΔ to [SWI+] | |
| Upregulated | 40 | 822 | 409 |
| Downregulated | 119 | 1156 | 730 |
| Total | 159 | 1978 | 1139 |
| Effect or Phenotype | Manifestation in the Strain Containing | Condition | Reference | |
|---|---|---|---|---|
| [SWI+] Prion | SWI1 Deletion | |||
| Decreased vegetative growth | + | + | Media with galactose, glycerol or raffinose as carbon source and Antimycin-A | [7] |
| Nonsense suppression in the presence of mutant SUP35 (eRF3) variants | + | + | Media without adenine; in the presence of mutant eRF3 variants with decreased functional activity | [35,37,40,42] |
| Decreased vegetative growth | + | + | Media with galactose or glycerol as carbon source | [35] |
| Loss of flocculation | + | + | Stationary-phase cultures | [36] |
| Loss of invasiveness | + | + | Complete media, 6 days of incubation | [36] |
| Abolished pseudohyphal growth | + | + | SLAD media containing 4% glucose | [36] |
| Repression of the FLO1 and FLO11 expression | + | + | Complete media with glucose as carbon source | [36] |
| Increased expression of the ADE1 gene | - | + | Complete media | [43] |
| Chromosome I disomy | - | + | Complete media | This study |
| Inhibition of the ribosome biogenesis and translation | - | + | Complete media with galactose as carbon source | This study |
| Omnipotent nonsense suppression in the presence of wild-type SUP35 (eRF3) | - | + | Media without adenine or tryptophan; in the presence of the wild-type eRF3 | This study |
| Decreased expression of the SUP45 (eRF1) gene | + | - | Complete media with glucose as carbon source | [41,42] |
| Aggregation of the Mss11, Sap30, Gts1 and Msn1 transcriptional regulators | + | - | Overproduction of the YFP-fused proteins under the GAL1 promoter | [36] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Malovichko, Y.V.; Antonets, K.S.; Maslova, A.R.; Andreeva, E.A.; Inge-Vechtomov, S.G.; Nizhnikov, A.A. RNA Sequencing Reveals Specific Transcriptomic Signatures Distinguishing Effects of the [SWI+] Prion and SWI1 Deletion in Yeast Saccharomyces cerevisiae. Genes 2019, 10, 212. https://doi.org/10.3390/genes10030212
Malovichko YV, Antonets KS, Maslova AR, Andreeva EA, Inge-Vechtomov SG, Nizhnikov AA. RNA Sequencing Reveals Specific Transcriptomic Signatures Distinguishing Effects of the [SWI+] Prion and SWI1 Deletion in Yeast Saccharomyces cerevisiae. Genes. 2019; 10(3):212. https://doi.org/10.3390/genes10030212
Chicago/Turabian StyleMalovichko, Yury V., Kirill S. Antonets, Anna R. Maslova, Elena A. Andreeva, Sergey G. Inge-Vechtomov, and Anton A. Nizhnikov. 2019. "RNA Sequencing Reveals Specific Transcriptomic Signatures Distinguishing Effects of the [SWI+] Prion and SWI1 Deletion in Yeast Saccharomyces cerevisiae" Genes 10, no. 3: 212. https://doi.org/10.3390/genes10030212
APA StyleMalovichko, Y. V., Antonets, K. S., Maslova, A. R., Andreeva, E. A., Inge-Vechtomov, S. G., & Nizhnikov, A. A. (2019). RNA Sequencing Reveals Specific Transcriptomic Signatures Distinguishing Effects of the [SWI+] Prion and SWI1 Deletion in Yeast Saccharomyces cerevisiae. Genes, 10(3), 212. https://doi.org/10.3390/genes10030212






