Transcriptome Analysis of the Cecal Tonsil of Jingxing Yellow Chickens Revealed the Mechanism of Differential Resistance to Salmonella
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Animals and Experimental Design
2.3. Total RNA Isolation, cDNA Library Construction, and Sequencing
2.4. Analysis of Differentially Expressed Genes (DEGs) and Pathway Enrichment
2.5. Quantitative Real-time PCR
3. Results
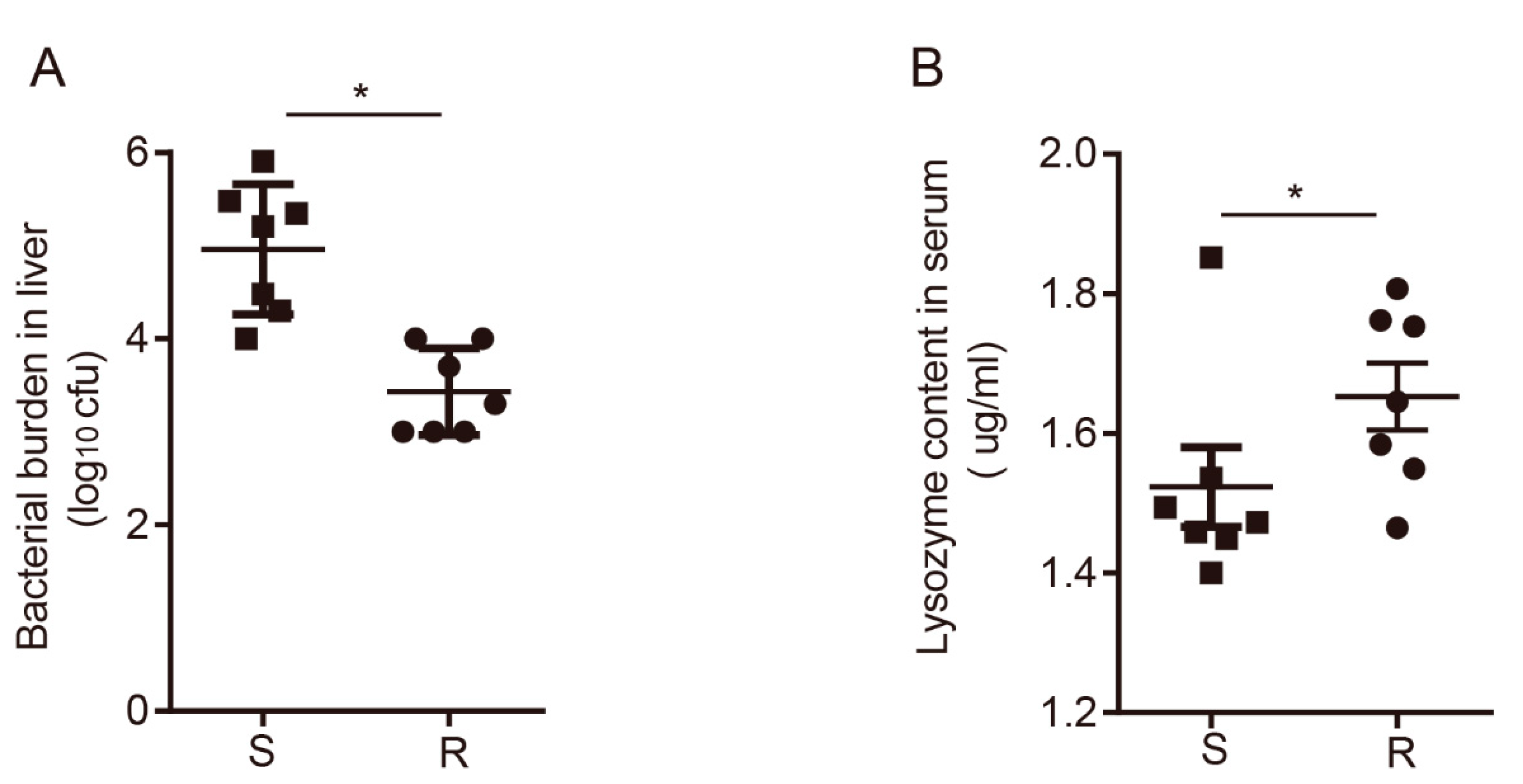
3.1. Overview of the Immune Response between Susceptible and Resistant Chickens
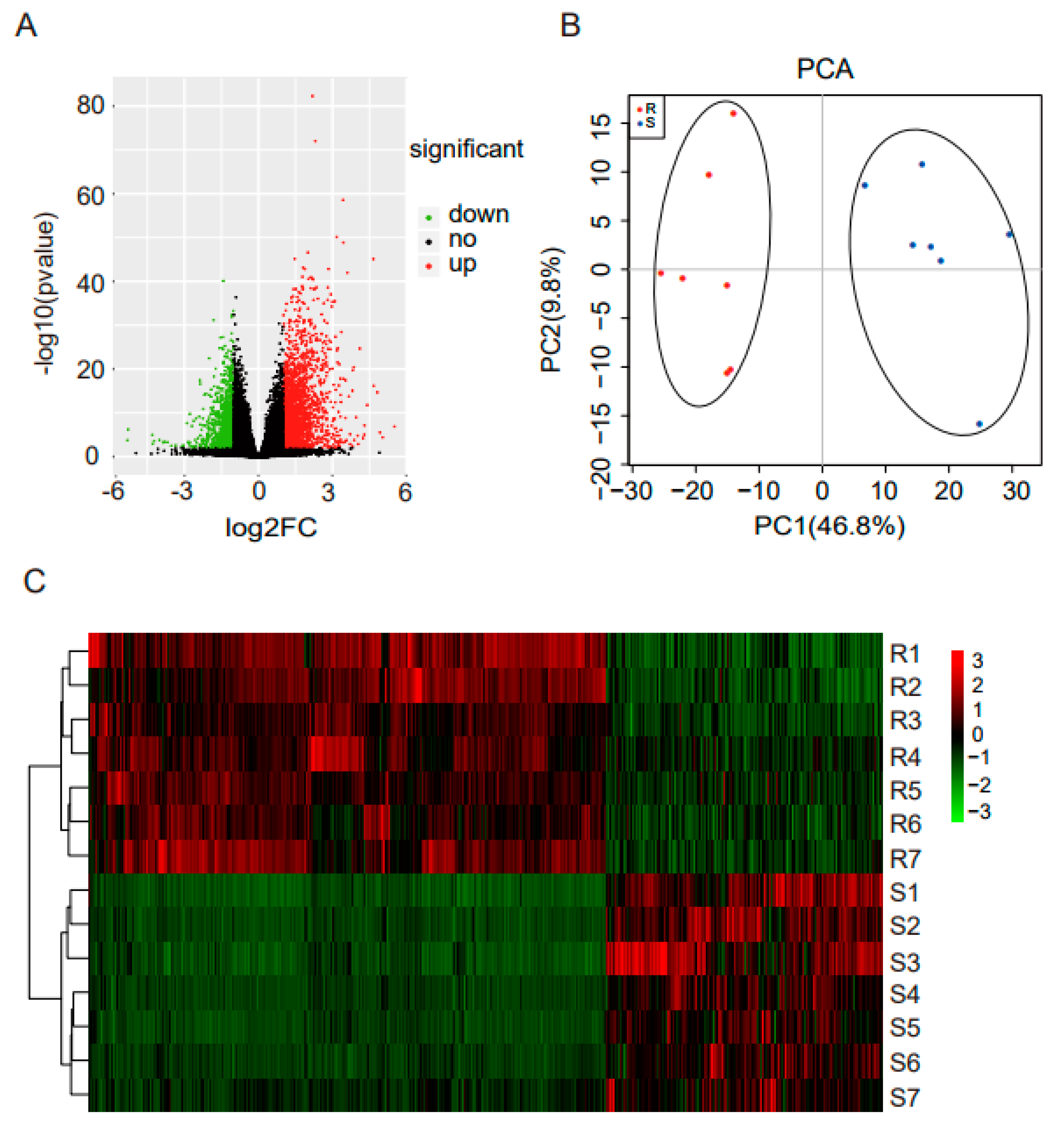
3.2. Sequencing of Cecal Tonsil Transcriptomes
3.3. Differentially Expressed mRNAs Responding to ST Infection in Cecal Tonsil
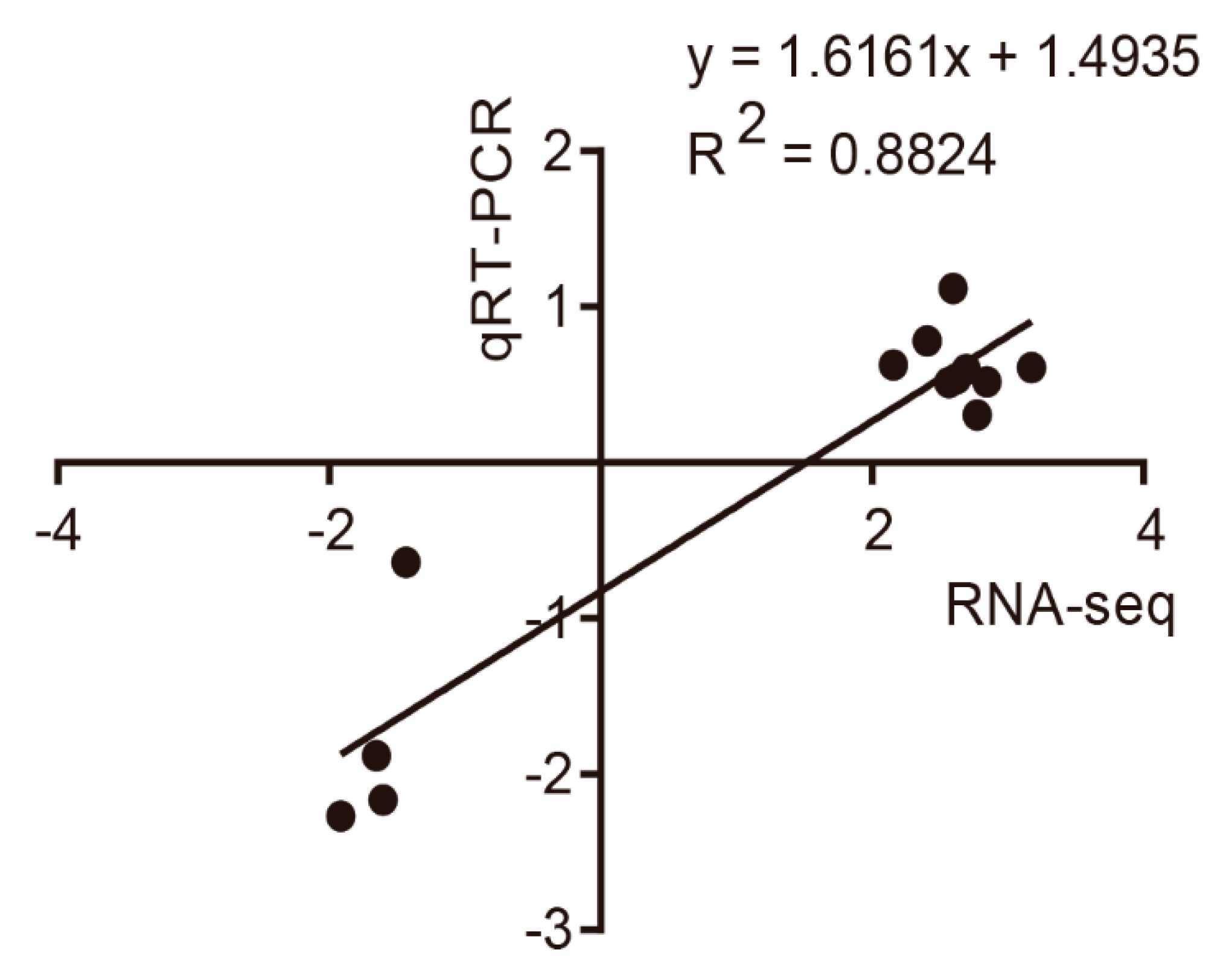
3.4. Quantitative Real-Time PCR Validation
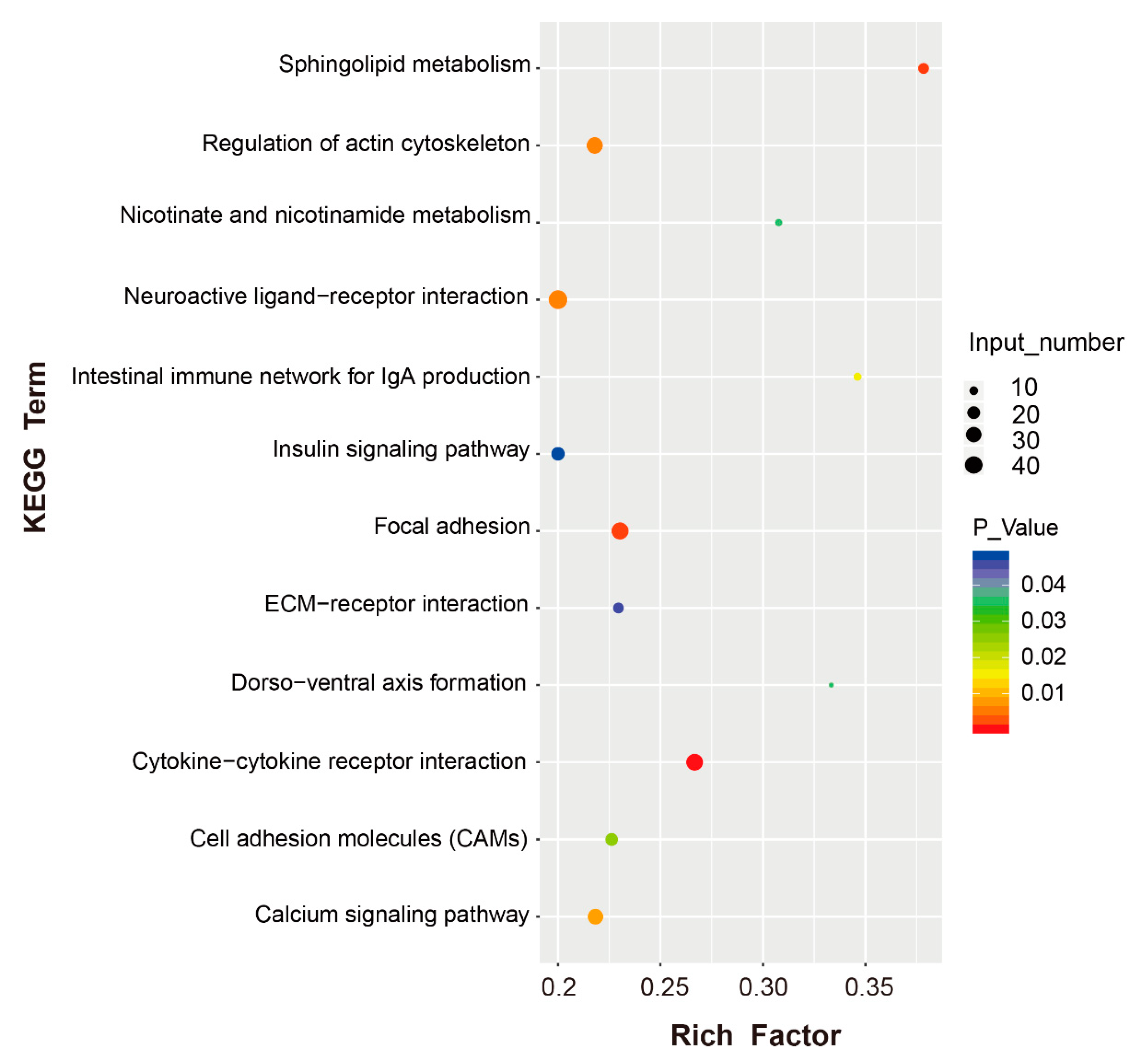
3.5. Functional Enrichment Analysis of the DEGs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Barrow, P.A.; Jones, M.A.; Smith, A.L.; Wigley, P. The long view: Salmonella—The last forty years. Avian Pathol. 2012, 41, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Zhang-Barber, L.; Turner, A.K.; Barrow, P.A. Vaccination for control of Salmonella in poultry. Vaccine 1999, 17, 2538–2545. [Google Scholar] [CrossRef]
- Snow, L.C.; Davies, R.H.; Christiansen, K.H.; Carrique-Mas, J.J.; Wales, A.D.; O’Connor, J.L.; Cook, A.J.C.; Evans, S.J. Survey of the prevalence of Salmonella species on commercial laying farms in the United Kingdom. Vet. Rec. 2007, 161, 471. [Google Scholar] [CrossRef] [PubMed]
- Bar-Shira, E.; Sklan, D.; Friedman, A. Establishment of immune competence in the avian GALT during the immediate post-hatch period. Dev. Comp. Immunol. 2003, 27, 147–157. [Google Scholar] [CrossRef]
- Bryan, T.; Nicholas, P.; Caitlyn, D.; Rafaela, P.; Mary, M.; Hassan, H.M.; Koci, M.D. Poultry Body Temperature Contributes to Invasion Control through Reduced Expression of Salmonella Pathogenicity Island 1 Genes in Salmonella enterica Serovars Typhimurium and Enteritidis. Appl. Environ. Microbiol. 2015, 81, 8192. [Google Scholar]
- Monack, D.M.; Anne, M.; Stanley, F. Persistent bacterial infections: The interface of the pathogen and the host immune system. Nat. Rev. Microbiol. 2004, 2, 747–765. [Google Scholar] [CrossRef]
- Weiss, D.S.; Raupach, B.; Takeda, K.; Akira, S.; Zychlinsky, A. Toll-like receptors are temporally involved in host defense. J. Immunol. 2004, 172, 4463–4469. [Google Scholar] [CrossRef]
- Li, P.; Xia, P.; Wen, J.; Zheng, M.; Chen, J.; Zhao, J.; Jiang, R.; Liu, R.; Zhao, G. Up-regulation of the MyD88-dependent pathway of TLR signaling in spleen and caecum of young chickens infected with Salmonella serovar Pullorum. Vet. Microbiol. 2010, 143, 346–351. [Google Scholar] [CrossRef]
- Qiu, J.W.; Xiao, C.Z.; Tian, W.; Zhang, T.Y. Effect of dietary oridonin supplementation on growth performance, gut health, and immune response of broilers infected with Salmonella pullorum. Irish Vet. J. 2018, 71, 16. [Google Scholar]
- Wieland, W.H.; Diego, O.; Aart, L.; Parmentier, H.K.; Verstegen, M.W.A.; Arjen, S. A functional polymeric immunoglobulin receptor in chicken (Gallus gallus) indicates ancient role of secretory IgA in mucosal immunity. Biochem. J. 2004, 380, 669–676. [Google Scholar] [CrossRef]
- Webb, T.; Goodman, H.C. The structure and function of immunoglobulins. Mod. Trends Immunol. 1967, 2, 151–187. [Google Scholar] [PubMed]
- Wieland, W.H.; Kruijt, L.; Jansma, A. Successive immunoglobulin and cytokine expression in the small intestine of juvenile chicken. Dev. Comp. Immunol. 2010, 34, 1254–1262. [Google Scholar]
- Bryan, C.; Yuling, L.; David, O.; Vallance, B.A.; B Brett, F. Salmonella enterica serovar Typhimurium pathogenicity island 2 is necessary for complete virulence in a mouse model of infectious enterocolitis. Infect. Immun. 2005, 73, 3219–3227. [Google Scholar]
- Rivera-Chávez, F.; Zhang, L.F.; Faber, F.; Lopez, C.A.; Byndloss, M.X.; Olsan, E.E.; Xu, G.; Velazquez, E.M.; Lebrilla, C.B.; Winter, S.E. Depletion of Butyrate-Producing Clostridia from the Gut Microbiota Drives an Aerobic Luminal Expansion of Salmonella. Cell Host Microbe 2016, 19, 443–454. [Google Scholar] [CrossRef] [PubMed]
- Byndloss, M.X.; Olsan, E.E.; Rivera-Chávez, F.; Tiffany, C.R.; Cevallos, S.A.; Lokken, K.L.; Torres, T.P.; Byndloss, A.J.; Faber, F.; Gao, Y. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 2017, 357, 570–575. [Google Scholar] [CrossRef] [PubMed]
- Setta, A.M.; Barrow, P.A.; Kaiser, P.; Jones, M.A. Early immune dynamics following infection with Salmonella enterica serovars Enteritidis, Infantis, Pullorum and Gallinarum: Cytokine and chemokine gene expression profile and cellular changes of chicken cecal tonsils. Comp. Immunol. Microbiol. Infect. Dis. 2012, 35, 397–410. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Lamont, S.J. Global gene expression profile after Salmonella enterica Serovar enteritidis challenge in two F8 advanced intercross chicken lines. Cytogenet. Genome Res. 2007, 117, 131–138. [Google Scholar] [CrossRef]
- Peng, L.; Fan, W.; Everaert, N.; Liu, R.; Li, Q.; Zheng, M.; Cui, H.; Zhao, G.; Jie, W. Messenger RNA Sequencing and Pathway Analysis Provide Novel Insights into the Susceptibility toSalmonella enteritidisInfection in Chickens. Front. Genet. 2018, 9, 256. [Google Scholar]
- Lollike, K.; Kjeldsen, L.; Sengelov, H.; Borregaard, N. Purification of lysozyme from human neutrophils, and development of an ELISA for quantification in cells and plasma. Leukemia 1995, 9, 206–209. [Google Scholar]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq--A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Ginestet, C. ggplot2: Elegant Graphics for Data Analysis. J. R. Stat. Soc. A Stat. 2011, 174, 245–246. [Google Scholar] [CrossRef]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef]
- Xie, C.; Mao, X.; Huang, J.; Ding, Y.; Wu, J.; Dong, S.; Kong, L.; Gao, G.; Li, C.Y.; Wei, L. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, W316–W322. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Lu, L.; Walker, W. Pathologic and physiologic interactions of bacteria with the gastrointestinal epithelium. Am. J. Clin. Nutr. 2001, 73, 1124S. [Google Scholar] [CrossRef]
- Schauer, D.B. Indigenous microflora: Paving the way for pathogens? Curr. Biol. 1997, 7, R75–R77. [Google Scholar] [CrossRef]
- Goosney, D.L.; Knoechel, D.G.; Finlay, B.B. Enteropathogenic, E. coli, Salmonella, and Shigella: Masters of host cell cytoskeletal exploitation. Emerg. Infect. Dis. 1999, 5, 216–223. [Google Scholar] [CrossRef]
- Gauthier, A.; Finlay, B.B. Protein translocation: Delivering virulence into the host cell. Curr. Biol. 1998, 8, R768–R770. [Google Scholar] [CrossRef]
- An, D.; Oh, S.; Olszak, T.; Neves, J.; Avci, F.; Erturk-Hasdemir, D.; Lu, X.; Zeissig, S.; Blumberg, R.; Kasper, D. Sphingolipids from a Symbiotic Microbe Regulate Homeostasis of Host Intestinal Natural Killer T Cells. Cell 2014, 156, 123–133. [Google Scholar] [CrossRef] [PubMed]
- Abdel, H.L.; Di, V.C.; Riboni, L. Fostering Inflammatory Bowel Disease: Sphingolipid Strategies to Join Forces. Mediat. Inflamm. 2016, 2016, 3827684. [Google Scholar] [CrossRef] [PubMed]
- Oliver, P. New concepts in the generation and functions of IgA. Nat. Rev. Immunol. 2012, 12, 821–832. [Google Scholar]
- Slack, E.; Balmer, M.L.; Fritz, J.H.; Hapfelmeier, S. Functional flexibility of intestinal IgA-broadening the fine line. Front. Immunol. 2012, 3, 100. [Google Scholar] [CrossRef] [PubMed]
- Mkaddem, S.B.; Christou, I.; Rossato, E.; Berthelot, L.; Lehuen, A.; Monteiro, R.C. IgA, IgA Receptors, and Their Anti-inflammatory Properties. Curr. Top. Microbiol. Immunol. 2014, 382, 221. [Google Scholar]
- Palm, N.; Dezoete, M.; Cullen, T.; Barry, N.; Stefanowski, J.; Hao, L.; Degnan, P.; Hu, J.; Peter, I.; Zhang, W. Immunoglobulin A Coating Identifies Colitogenic Bacteria in Inflammatory Bowel Disease. Cell 2014, 158, 1000–1010. [Google Scholar] [CrossRef]
- Forbes, S.J.; Marisa, E.; Mantis, N.J. Inhibition of Salmonella enterica serovar typhimurium motility and entry into epithelial cells by a protective antilipopolysaccharide monoclonal immunoglobulin A antibody. Infect. Immun. 2008, 76, 4137–4144. [Google Scholar] [CrossRef]
- Forbes, S.J.; Tia, B.; Mccarthy, E.A.; Blaise, C.; Mantis, N.J. Transient suppression of Shigella flexneri type 3 secretion by a protective O-antigen-specific monoclonal IgA. Mbio 2011, 2, 00042-11. [Google Scholar] [CrossRef]
- Freemerman, A.J.; Johnson, A.R.; Sacks, G.N.; Milner, J.J.; Kirk, E.L.; Troester, M.A.; Macintyre, A.N.; Pankuri, G.H.; Rathmell, J.C.; Liza, M. Metabolic reprogramming of macrophages: Glucose transporter 1 (GLUT1)-mediated glucose metabolism drives a proinflammatory phenotype. J. Biol. Chem. 2014, 289, 7884. [Google Scholar] [CrossRef]
- Van den Bossche, J.; O’Neill, L.A.; Menon, D. Macrophage Immunometabolism: Where Are We (Going)? Trends Immunol. 2017, 38, 395. [Google Scholar] [CrossRef] [PubMed]
- Tucey, T.M.; Verma, J.; Harrison, P.F.; Snelgrove, S.L.; Lo, T.L.; Scherer, A.K.; Barugahare, A.A.; Powell, D.R.; Wheeler, R.T.; Hickey, M.J. Glucose Homeostasis Is Important for Immune Cell Viability during Candida Challenge and Host Survival of Systemic Fungal Infection. Cell Metab. 2018, 27, 988. [Google Scholar] [CrossRef] [PubMed]
| Term | Description | Count | Corrected p-Value |
|---|---|---|---|
| GO:0023052 | signaling | 237 | 0.002 |
| GO:0007154 | cell communication | 237 | 0.002 |
| GO:0050896 | response to stimulus | 304 | 0.002 |
| GO:0007165 | signal transduction | 219 | 0.003 |
| GO:0002376 | immune system process | 98 | 0.004 |
| GO:0007155 | cell adhesion | 72 | 0.007 |
| GO:0022610 | biological adhesion | 72 | 0.007 |
| GO:0016020 | membrane | 355 | 0.007 |
| GO:0016021 | Integral component of membrane | 236 | 0.009 |
| GO:0044425 | membrane part | 276 | 0.009 |
| GO:0098602 | single organism cell adhesion | 46 | 0.100 |
| GO:0031224 | intrinsic component of membrane | 238 | 0.012 |
| GO:0016337 | single organismal cell–cell adhesion | 43 | 0.023 |
| GO:0051716 | cellular response to stimulus | 248 | 0.028 |
| GO:0006955 | immune response | 52 | 0.032 |
| GO:0005886 | plasma membrane | 165 | 0.032 |
| GO:0071944 | cell periphery | 168 | 0.043 |
| GO:0032501 | multicellular organismal process | 236 | 0.049 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, F.; Zhang, J.; Zhu, B.; Wang, J.; Wang, Q.; Zheng, M.; Wen, J.; Li, Q.; Zhao, G. Transcriptome Analysis of the Cecal Tonsil of Jingxing Yellow Chickens Revealed the Mechanism of Differential Resistance to Salmonella. Genes 2019, 10, 979. https://doi.org/10.3390/genes10120979
Wang F, Zhang J, Zhu B, Wang J, Wang Q, Zheng M, Wen J, Li Q, Zhao G. Transcriptome Analysis of the Cecal Tonsil of Jingxing Yellow Chickens Revealed the Mechanism of Differential Resistance to Salmonella. Genes. 2019; 10(12):979. https://doi.org/10.3390/genes10120979
Chicago/Turabian StyleWang, Fei, Jin Zhang, Bo Zhu, Jie Wang, Qiao Wang, Maiqing Zheng, Jie Wen, Qinghe Li, and Guiping Zhao. 2019. "Transcriptome Analysis of the Cecal Tonsil of Jingxing Yellow Chickens Revealed the Mechanism of Differential Resistance to Salmonella" Genes 10, no. 12: 979. https://doi.org/10.3390/genes10120979
APA StyleWang, F., Zhang, J., Zhu, B., Wang, J., Wang, Q., Zheng, M., Wen, J., Li, Q., & Zhao, G. (2019). Transcriptome Analysis of the Cecal Tonsil of Jingxing Yellow Chickens Revealed the Mechanism of Differential Resistance to Salmonella. Genes, 10(12), 979. https://doi.org/10.3390/genes10120979




