Abstract
ORCAE (Online Resource for Community Annotation of Eukaryotes) is a public genome annotation curation resource. ORCAE-AOCC is a branch that is dedicated to the genomes published as part of the African Orphan Crops Consortium (AOCC). The motivation behind the development of the ORCAE platform was to create a knowledge-based website where the research-community can make contributions to improve genome annotations. All changes to any given gene-model or gene description are stored, and the entire annotation history can be retrieved. Genomes can either be set to “public” or “restricted” mode; anonymous users can browse public genomes but cannot make any changes. Aside from providing a user- friendly interface to view genome annotations, the platform also includes tools and information (such as gene expression evidence) that enables authorized users to edit and validate genome annotations. The ORCAE-AOCC platform will enable various stakeholders from around the world to coordinate their efforts to annotate and study underutilized crops.
1. Introduction
According to the United Nations (2019), the number of undernourished people has increased in the past three years, with more than 820 million people still facing starvation. Moreover, the majority of these people live in developing countries. As the global population continues to grow, it is predicted that there will be 9 billion people by 2050, and the demand for food will be 70% greater than it is today [1]. Currently, out of the estimated 50,000 edible plant species, just three of them (maize, rice, and wheat) provide two-thirds of the world’s food energy intake. Technological innovations such as the development and selection of high-yield varieties, improvement of pesticide and fertilizer use, mechanization, and irrigation facilities have contributed to a global increase in the production of these grains. However, these innovations are often not readily available to the subsistence farmers throughout Africa.
Apart from maize, rice, and wheat, which are all important for Africa and African farmers, there are many other crops that are currently underutilized but have great potential for Africa. These underutilized or so-called “orphan-crops”, are ancient, neglected, or indigenous crops with limited cultivation at a global scale, and their use ranges from food, fodder, to derivatives such as oil and medicine. In addition, orphan crops are promising solutions for nutritional diversity and can reduce over-reliance on major-crops and certain agricultural practices that have a negative environmental impact, as discussed by Mayes et al. [2]. In order to fast-track the improvement of orphan-crops, either by selective breeding or genetic modification, it is essential to have reliable information about their genetic makeup.
The African Orphan Crop Consortium (AOCC) was established in 2011 to tackle hunger and malnutrition in Africa [3]. AOCC aims to facilitate the development of locally available crops to supply nutritious and high yielding varieties. AOCC has committed itself to sequence, annotate, and analyze the genomes of 101 mostly indigenous and some introduced crops [3]. Given that orphan crops are expected to advance healthy food systems, as well as genetic resources for future crops, and agricultural sustainability under climate change [4], it is anticipated that the comprehensively selected 101 species in AOCC will become an invaluable resource and broaden the diversity of our current understanding of crop genomics (Figure 1).
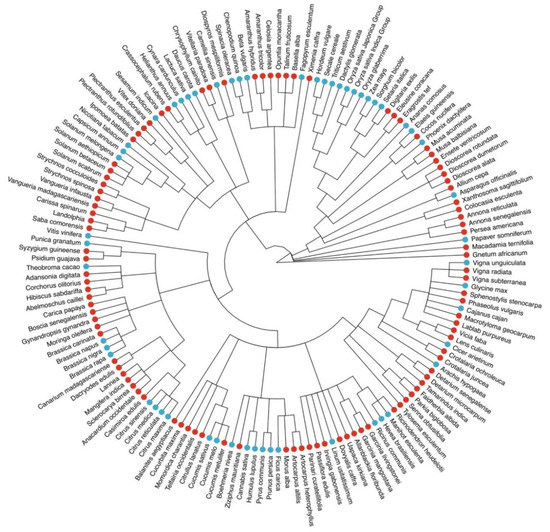

Figure 1.
Diversity of currently sequenced, publicly available crop genomes (blue dots) and the orphan crop species initially included in the African Orphan Crops Consortium list (red dots), some of which are available on ORCAE already.
The availability of high-quality crop reference genomes has already proved to be valuable to breeders, for example, by facilitating the identification of breeding targets in the genome [5]. Nevertheless, the complexity of many plant genomes, due to size, high repeat content, and polyploid ancestry has rendered their de novo genome assemblies particularly difficult [6]. Furthermore, different sequencing platforms create different challenges in the downstream bioinformatics analyses of a Next Generation Sequencing (NGS) project workflow [7,8,9]. On the other hand, long-read sequencing techniques and the recent advances of assemblers have generally increased the quality of draft genomes. However, fewer developments were made in terms of the genome annotation procedures, and some even argue that the errors of genome annotations keep propagating [9]. Here, we present a genomic resource called ORCAE-AOCC, a community-based genome annotation platform, and discuss its potential value for the scientific community.
Genome annotation mainly consists of two phases. (1) Structural annotation aims to provide information on the location of genes in the genome and the exact boundaries of exons and introns. The process often involves making use of transcript evidence [10,11] and ab initio modeling that utilizes statistical models to predict gene structures [11,12]. (2) Functional annotation aims to assign biological functions to the genes. Functional annotation is often homology-based, which means that information about the biological role of a gene in a new genome is inferred from genes with similar sequences from other genomes, where the role of the gene has been described or predicted [11]. Hypothetical genes (experimentally uncharacterized genes) or predicted gene models without any similarity found can sometimes be species-specific and their functions can be hard to deduce [13,14,15,16]. However, as more genomes are sequenced and annotated, it is becoming clear that some hypothetical genes are conserved between species [13]. Experimental evidence has shown that many hypothetical genes are indeed expressed and that they have critical biological roles [13,17,18,19]. Therefore, there is a need to focus efforts toward understanding the roles of hypothetically functional genes. Furthermore, it is also important to efficiently identify and exclude miss-annotated hypothetical genes [16].
Manual curation of genome annotations has been proven to be extremely valuable for building accurate reference gene sets in model organisms but compared to automatic annotation methods it is prohibitively expensive and is thus not widely done for non-model organisms [20]. Community- based efforts, where different researchers working on the same genome can contribute new information on gene structure and function have proven to be highly beneficial, e.g. in the case of the Vertebrate Genome Annotation Database (VEGA, [21]), which is maintained by the human and vertebrate analysis and annotation (Havana) team at the Wellcome Trust Sanger Institute (WTSI, [22]). Some of the best known community annotation platforms for plants include: The Arabidopsis Information Resource (TAIR) [23,24], the Maize Genetics and Genomics Database (Maize GDB) [25,26], The Rice Annotation Project Database (RAP-DB) [27,28,29], the International Wheat Genome Sequencing Consortium (IWGSC) [30,31] and Wheat@URGI [32,33]. Each of these platforms are dedicated to a single species that is widely grown and well-studied. Typically, these genomes have been annotated using automatic methods followed by manual curation steps [24,28]. Members of the scientific community can provide the curators of the databases above with information on newly identified genes and gene functions, which are then added when the genome annotations are updated.
2. ORCAE-AOCC and the Currently Deployed Genomes
Referred to as the Online Resource for Community Annotation of Eukaryotes, ORCAE [34] was developed and launched in 2012 to facilitate the manual curation of gene models, functional annotations, and improvement of annotation quality by genome consortia [35]. ORCAE was chosen as a central platform for the annotation of the grapevine (Vitis vinifera) as part of the International Grapevine Genome Project [36], In recent years, ORCAE has also been used by other communities resulting in several high-profile publications including the genomes of the seagrass (Zostera marina) [37], olive tree (Olea europaea var. sylvestris) [38], and sea lettuce (Ulva mutabilis) [39]. The system was designed with a wiki-like style editing mode for the community to refine information about the gene models, such as the gene structures and the definition of gene function. ORCAE also seamlessly integrates with GenomeView [40] to allow manual editing of the structure of a given gene model, with the aid of RNA-Seq or (expressed sequence tag) EST evidence. After editing the gene structures, the system will perform a number of checks in order to validate the modified gene structure before it is committed back to the database.
On the gene page (Figure 2), ORCAE displays the alignment of homologs retrieved from other public databases for a given gene, as well as other evidential information such as EST alignments and expression profiles. The built-in backend utility will search the detail properties of the protein sequence and provide protein domain information using InterProScan [41]. As an information collector, ORCAE allows expert annotators from the consortium to assess and update the information of each gene. Additionally, the functional description of each gene is transferred from the trusted reference database by homology and summarized using tools such as Automated Assignment of Human Readable Descriptions (AHRD) [42]. The system will automatically update the information of the gene after any modification. The final quality of the annotation depends not only on the initial deployed ab initio prediction but also on the effort of the consortium. Although it is hard to be scaled in all genome projects, it still provides a way to improve the quality of gene prediction and avoid the propagation of false positively predicted structures.
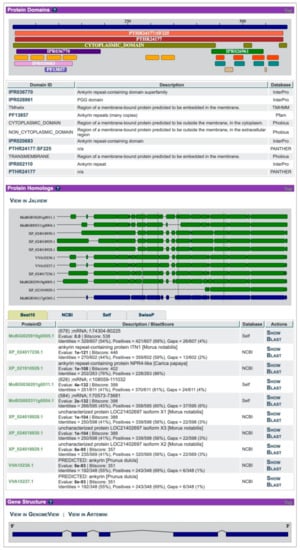
Figure 2.
Part of the gene page in ORCAE-AOCC. Screenshot taken from [43].
In order to incorporate the incoming genomes from the AOCC project into the ORCAE platform, while maintaning focus on the project, we launched ORCAE-AOCC [43], a dedicated genome portal for the AOCC consortium (Figure 3). ORCAE-AOCC currently contains five published genomes from the consortium: Faidherbia albida, Moringa oleifera, Sclerocarya birrea, Lablab purpureus, and Vigna radiata [44] the remaining genomes will be added as soon as their sequencing and annotation are complete. Apart from the curation of functional and structural annotation, users can also visit the portal of the desired genome in order to download the latest version of the annotations, the coding sequences (CDS) and the predicted protein sequences. Users can also perform BLAST [45] searches against CDS, protein, or the genomic sequence from within the portal. For on-going and restricted genome projects, one can request an account from the coordinator of the genome project to join the curation process prior to publication.
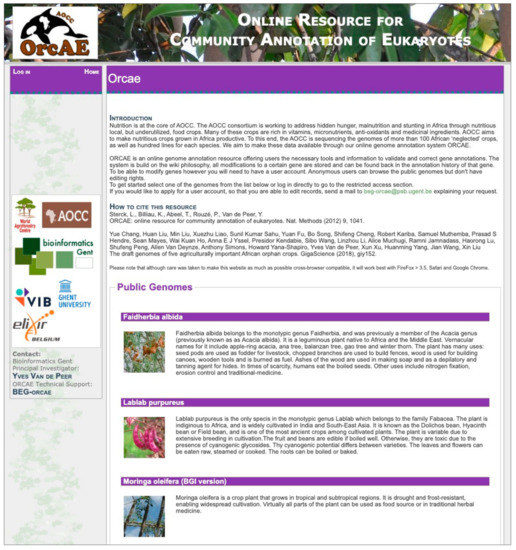
Figure 3.
An overview of the ORCAE-AOCC genome portal.
3. Concluding Remarks and Future Perspectives
ORCAE-AOCC is a platform that acts as a portal where the genomes that are sequenced as part of the AOCC initiative can be accessed by the broader research community. It also facilities collaborative efforts to improve the annotations and serves as a central repository for up to date versions of the genomes. ORCAE-AOCC also contains functionalities such as BLAST (Basic Local Alignment Search Tool) [46], the visualization of gene expression profiles, and pre-computed functional information. Additional African orphan crop genomes will be added when they will become available.
Author Contributions
Conceptualization L.S. and Y.V.d.P.; software, L.S. and S.-M.K.; resources, A.E.J.Y. and S.-M.K.; original draft preparation S.-M.K. and A.E.J.Y.; writing A.E.J.Y. and S.-M.K.; review and editing, L.S. and Y.V.d.P.; supervision L.S. and Y.V.d.P.
Funding
Anna E. J. Yssel acknowledges the University of Pretoria for providing her with a postdoctoral fellowship (Director of Research and Innovation cost centre A0C827).
Acknowledgments
We thank Thomas Van Parys for setting up the initial instance of ORCAE-AOCC.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Godfray, H.C.J.; Beddington, J.R.; Crute, I.R.; Haddad, L.; Lawrence, D.; Muir, J.F.; Pretty, J.; Robinson, S.; Thomas, S.M.; Toulmin, C. Food security: The challenge of feeding 9 billion people. Science 2010, 327, 812–818. [Google Scholar] [CrossRef] [PubMed]
- Mayes, S.; Massawe, F.J.; Alderson, P.G.; Roberts, J.A.; Azam-Ali, S.N.; Hermann, M. The potential for underutilized crops to improve security of food production. J. Exp. Bot. 2012, 63, 1075–1079. [Google Scholar] [CrossRef] [PubMed]
- African Orphan Crops Consortium—Healthy Africa through Nutritious, Diverse and Local Food Crops. Available online: http://africanorphancrops.org/ (accessed on 19 November 2019).
- Mabhaudhi, T.; Chimonyo, V.G.P.; Hlahla, S.; Massawe, F.; Mayes, S.; Nhamo, L.; Modi, A.T. Prospects of orphan crops in climate change. Planta 2019, 250, 695–708. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Scheben, A.; Edwards, D. Advances in Integrating Genomics and Bioinformatics in the Plant. Breeding Pipeline. Agriculture-Basel 2018, 8, 75. [Google Scholar] [CrossRef]
- Claros, M.G.; Bautista, R.; Guerrero-Fernández, D.; Benzerki, H.; Seoane, P.; Fernández-Pozo, N. Why assembling plant genome sequences is so challenging. Biology 2012, 1, 439–459. [Google Scholar] [CrossRef]
- Watson, M.; Warr, A. Errors in long-read assemblies can critically affect protein prediction. Nat. Biotechnol. 2019, 37, 124–126. [Google Scholar] [CrossRef]
- Vaattovaara, A.; Leppälä, J.; Salojärvi, J.; Wrzaczek, M. High-throughput sequencing data and the impact of plant gene annotation quality. J. Exp. Bot. 2019, 70, 1069–1076. [Google Scholar] [CrossRef]
- Salzberg, S.L. Next-generation genome annotation: We still struggle to get it right. Genome Biol. 2019, 20, 92. [Google Scholar] [CrossRef]
- Stanke, M.; Diekhans, M.; Baertsch, R.; Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 2008, 24, 637–644. [Google Scholar] [CrossRef]
- Bolger, M.E.; Arsova, B.; Usadel, B. Plant genome and transcriptome annotations: From misconceptions to simple solutions. Brief. Bioinform. 2018, 19, 437–449. [Google Scholar] [CrossRef]
- Sleator, R.D. An overview of the current status of eukaryote gene prediction strategies. Gene 2010, 461, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Kolker, E.; Makarova, K.S.; Shabalina, S.; Picone, A.F.; Purvine, S.; Holzman, T.; Cherny, T.; Armbruster, D.; Munson, R.S., Jr.; Kolesov, G.; et al. Identification and functional analysis of ‘hypothetical’ genes expressed in Haemophilus influenzae. Nucleic Acids Res. 2004, 32, 2353–2361. [Google Scholar] [CrossRef] [PubMed]
- Kolker, E.; Picone, A.F.; Galperin, M.Y.; Romine, M.F.; Higdon, R.; Makarova, K.S.; Kolker, N.; Anderson, G.A.; Qiu, X.; Auberry, K.J.; et al. Global profiling of Shewanella oneidensis MR-1: Expression of hypothetical genes and improved functional annotations. Proc. Natl. Acad. Sci. USA 2005, 102, 2099–2104. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, S.; Zhu, W.; Hamilton, J.; Lin, H.; Campbell, M.; Childs, K.; Thibaud-Nissen, F.; Malek, R.L.; Lee, Y.; Zheng, L.; et al. The TIGR Rice Genome Annotation Resource: Improvements and new features. Nucleic Acids Res. 2007, 35, D883–D887. [Google Scholar] [CrossRef]
- Jiang, S.Y.; Christoffels, A.; Ramamoorthy, R.; Ramachandran, S. Expansion mechanisms and functional annotations of hypothetical genes in the rice genome. Plant Physiol. 2009, 150, 1997–2008. [Google Scholar] [CrossRef][Green Version]
- Naveed, M.; Chaudhry, Z.; Ali, Z.; Amjad, M. Annotation and curation of hypothetical proteins: Prioritizing targets for experimental study. Adv. Life Sci. 2018, 5, 73–87. [Google Scholar]
- Xiao, Y.L.; Malik, M.; Whitelaw, C.A.; Town, C.D. Cloning and sequencing of cDNAs for hypothetical genes from chromosome 2 of Arabidopsis. Plant Physiol. 2002, 130, 2118–2128. [Google Scholar] [CrossRef][Green Version]
- Xiao, Y.L.; Smith, S.R.; Ishmael, N.; Redman, J.C.; Kumar, N.; Monaghan, E.L.; Ayele, M.; Haas, B.J.; Wu, H.C.; Town, C.D. Analysis of the cDNAs of hypothetical genes on Arabidopsis chromosome 2 reveals numerous transcript variants. Plant Physiol. 2005, 139, 1323–1337. [Google Scholar] [CrossRef][Green Version]
- Loveland, J.E.; Gilbert, J.G.; Griffiths, E.; Harrow, J.L. Community gene annotation in practice. Database 2012, 2012, bas009. [Google Scholar] [CrossRef]
- VEGA. Available online: http://vega.sanger.ac.uk (accessed on 19 November 2019).
- WTSI. Available online: http://www.sanger.ac.uk (accessed on 19 November 2019).
- TAIR. Available online: http://arabidopsis.org (accessed on 19 November 2019).
- Lamesch, P.; Berardini, T.Z.; Li, D.; Swarbreck, D.; Wilks, C.; Sasidharan, R.; Muller, R.; Dreher, K.; Alexander, D.L.; Garcia-Hernandez, M.; et al. The Arabidopsis Information Resource (TAIR): Improved gene annotation and new tools. Nucleic. Acids Res. 2012, 40, D1202–D1210. [Google Scholar] [CrossRef]
- Maize GDB. Available online: https://www.maizegdb.org (accessed on 19 November 2019).
- Portwood, J.L.; Woodhouse, M.R.; Cannon, E.K.; Gardiner, J.M.; Harper, L.C.; Schaeffer, M.L.; Walsh, J.R.; Sen, T.Z.; Cho, K.T.; Schott, D.A.; et al. MaizeGDB 2018: The maize multi-genome genetics and genomics database. Nucleic Acids Res. 2019, 47, D1146–D1154. [Google Scholar] [CrossRef] [PubMed]
- RAP-DB. Available online: https://rapdb.dna.affrc.go.jp (accessed on 19 November 2019).
- Sakai, H.; Lee, S.S.; Tanaka, T.; Numa, H.; Kim, J.; Kawahara, Y.; Wakimoto, H.; Yang, C.; Iwamoto, M.; Abe, T.; et al. Rice Annotation Project Database (RAP-DB): An integrative and interactive database for rice genomics. Plant Cell Physiol. 2013, 54, e6. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, Y.; de la Bastide, M.; Hamilton, J.P.; Kanamori, H.; McCombie, W.R.; Ouyang, S.; Schwartz, D.C.; Tanaka, T.; Wu, J.; Zhou, S.; et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 2013, 6, 4. [Google Scholar] [CrossRef] [PubMed]
- IWGSC. Available online: https://www.wheatgenome.org (accessed on 19 November 2019).
- Appels, R.; Eversole, K.; Stein, N.; Feuillet, C.; Keller, B.; Rogers, J.; Pozniak, C.J.; Choulet, F.; Distelfeld, A.; Poland, J.; et al. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 2018, 361. [Google Scholar] [CrossRef]
- Wheat@URGI. Available online: https://wheat-urgi.versailles.inra.fr (accessed on 19 November 2019).
- Alaux, M.; Rogers, J.; Letellier, T.; Flores, R.; Alfama, F.; Pommier, C.; Mohellibi, N.; Durand, S.; Kimmel, E.; Michotey, C.; et al. Linking the International Wheat Genome Sequencing Consortium bread wheat reference genome sequence to wheat genetic and phenomic data. Genome Biol. 2018, 19, 111. [Google Scholar] [CrossRef]
- ORCAE. Available online: https://bioinformatics.psb.ugent.be/orcae/ (accessed on 19 November 2019).
- Sterck, L.; Billiau, K.; Abeel, T.; Rouze, P.; Van de Peer, Y. ORCAE: Online resource for community annotation of eukaryotes. Nat. Methods 2012, 9, 1041. [Google Scholar] [CrossRef]
- Grimplet, J.; Adam-Blondon, A.F.; Bert, P.F.; Bitz, O.; Cantu, D.; Davies, C.; Delrot, S.; Pezzotti, M.; Rombauts, S.; Cramer, G.R. The grapevine gene nomenclature system. BMC Genomics 2014, 15, 1077. [Google Scholar] [CrossRef]
- Olsen, J.L.; Rouzé, P.; Verhelst, B.; Lin, Y.C.; Bayer, T.; Collen, J.; Dattolo, E.; De Paoli, E.; Dittami, S.; Maumus, F.; et al. The genome of the seagrass Zostera marina reveals angiosperm adaptation to the sea. Nature 2016, 530, 331–335. [Google Scholar] [CrossRef]
- Unver, T.; Wu, Z.; Sterck, L.; Turktas, M.; Lohaus, R.; Li, Z.; Yang, M.; He, L.; Deng, T.; Escalante, F.J.; et al. Genome of wild olive and the evolution of oil biosynthesis. Proc. Natl. Acad. Sci. USA 2017, 114, E9413–E9422. [Google Scholar] [CrossRef]
- De Clerck, O.; Kao, S.M.; Bogaert, K.A.; Blomme, J.; Foflonker, F.; Kwantes, M.; Vancaester, E.; Vanderstraeten, L.; Aydogdu, E.; Boesger, J.; et al. Insights into the Evolution of Multicellularity from the Sea Lettuce Genome. Curr. Biol. 2018, 28, 2921–2933. [Google Scholar] [CrossRef]
- Abeel, T.; Van Parys, T.; Saeys, Y.; Galagan, J.; Van de Peer, Y. GenomeView: A next-generation genome browser. Nucleic Acids Res. 2012, 40, e12. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Heiko, S. AHRD. Available online: https://github.com/groupschoof/AHRD (accessed on 19 November 2019).
- ORCAE-AOCC. Available online: https://bioinformatics.psb.ugent.be/orcae/aocc/ (accessed on 19 November 2019).
- Chang, Y.; Liu, H.; Liu, M.; Liao, X.; Sahu, S.K.; Fu, Y.; Song, B.; Cheng, S.; Kariba, R.; Muthemba, S.; et al. The draft genomes of five agriculturally important African orphan crops. Gigascience 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).