Dissecting the Regulatory Network of Leaf Premature Senescence in Maize (Zea mays L.) Using Transcriptome Analysis of ZmELS5 Mutant
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. RNA Extraction
2.3. Transcriptome Analysis
2.4. Quantitative Reverse Transcription PCR (RT-qPCR) Validation
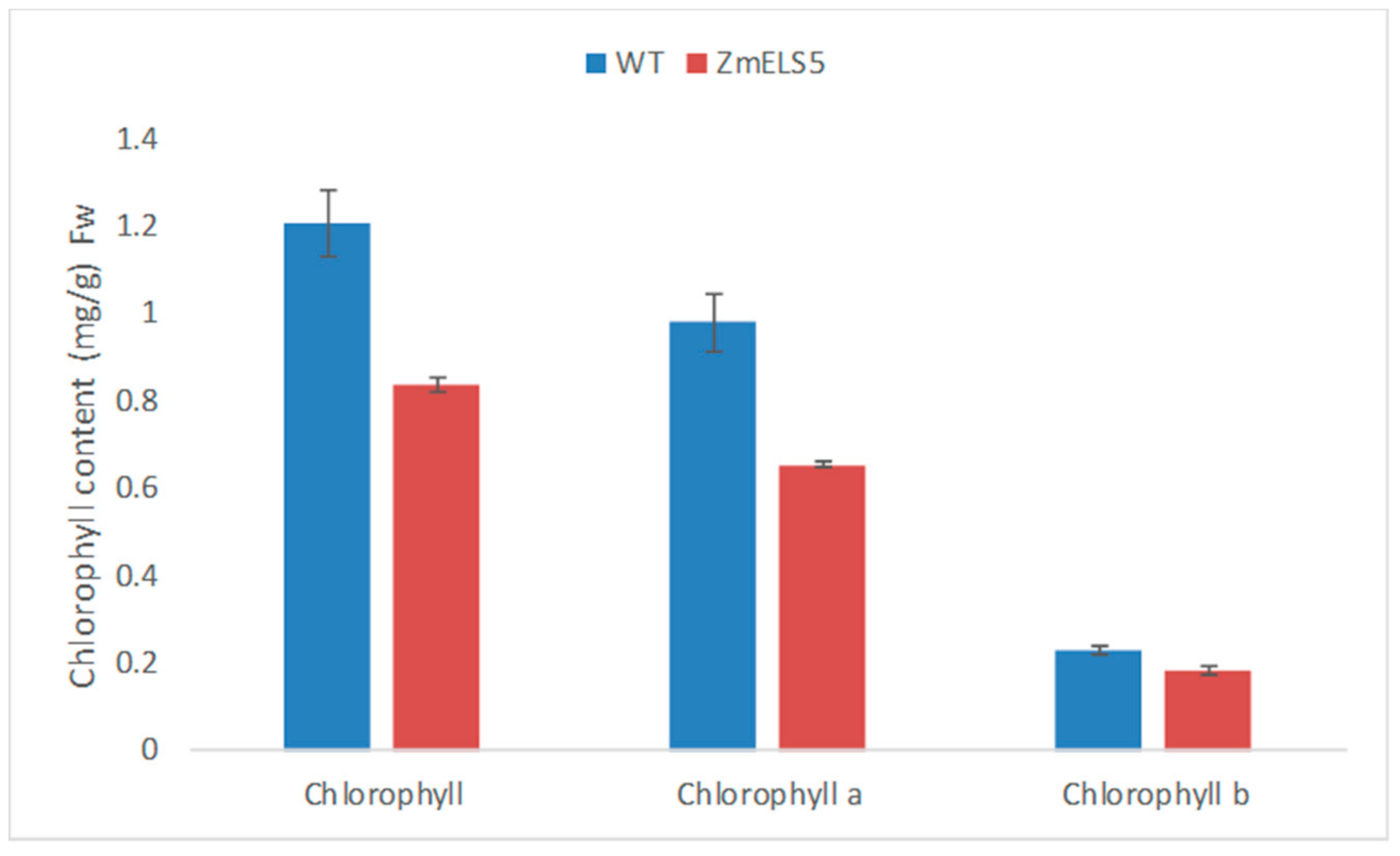
2.5. Determination of Chlorophyll Content
2.6. Putative Senescence Regulatory Network
3. Results
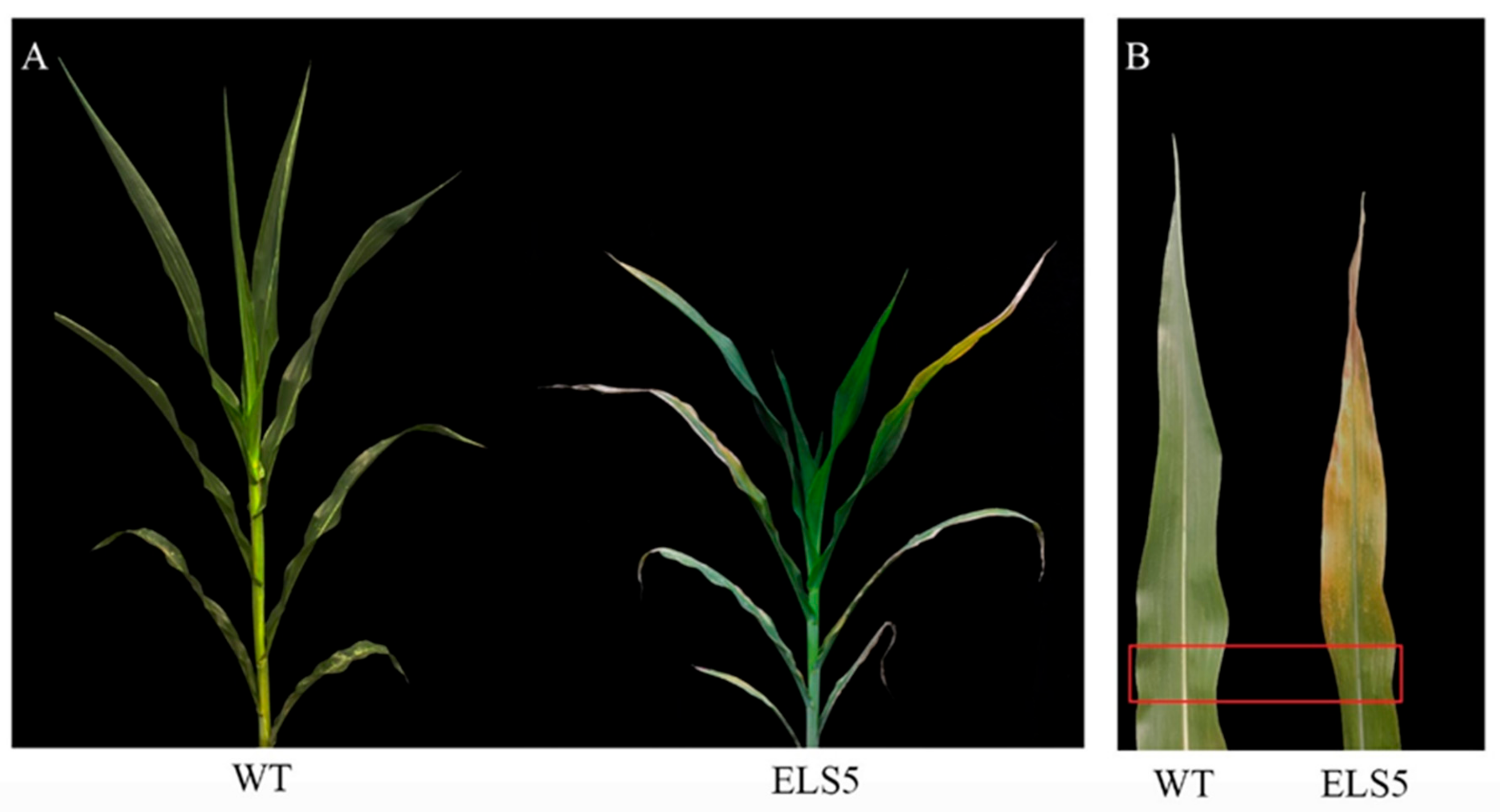
3.1. Characterization of Maize Premature-Senescence Mutant ZmELS5
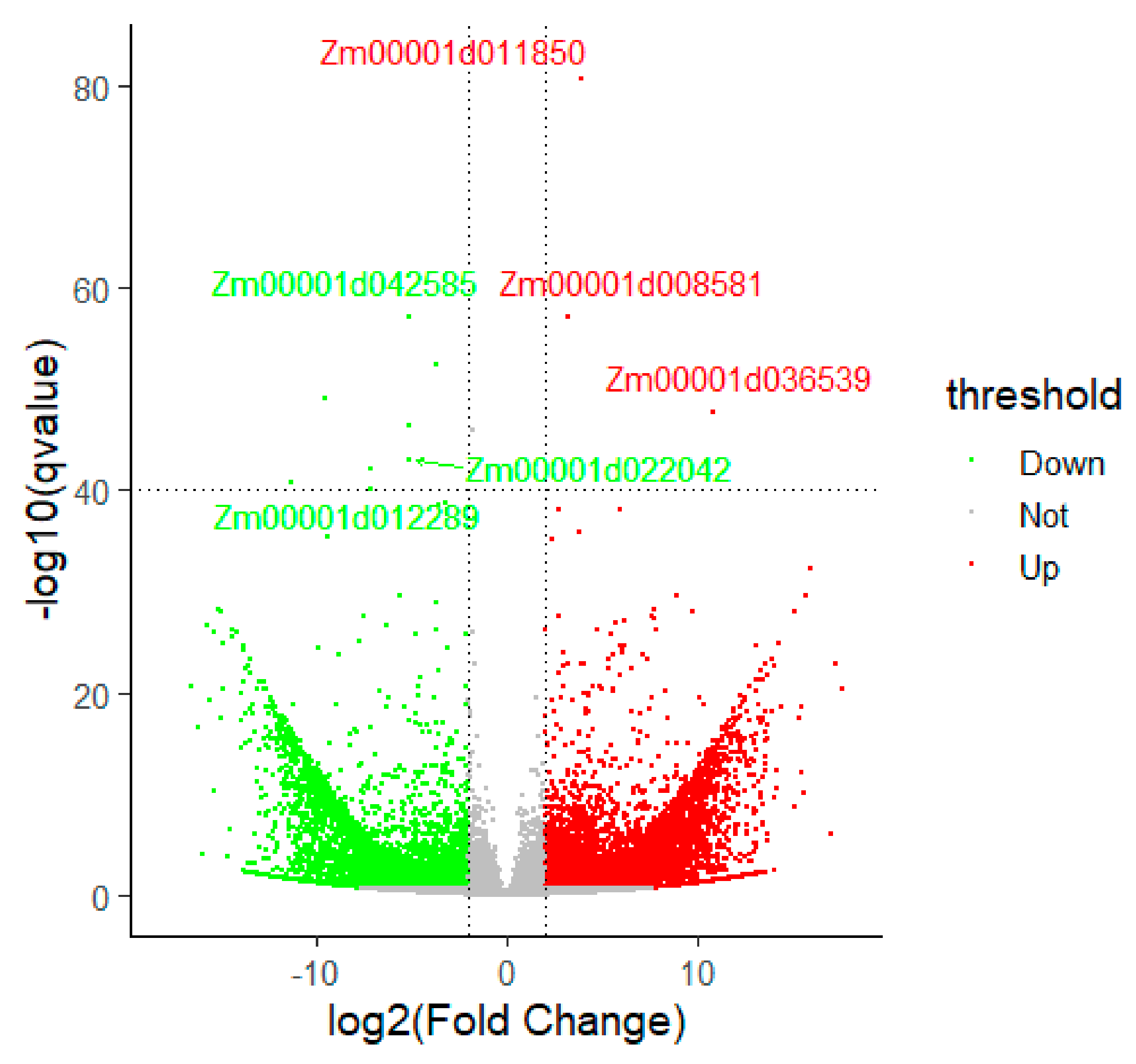
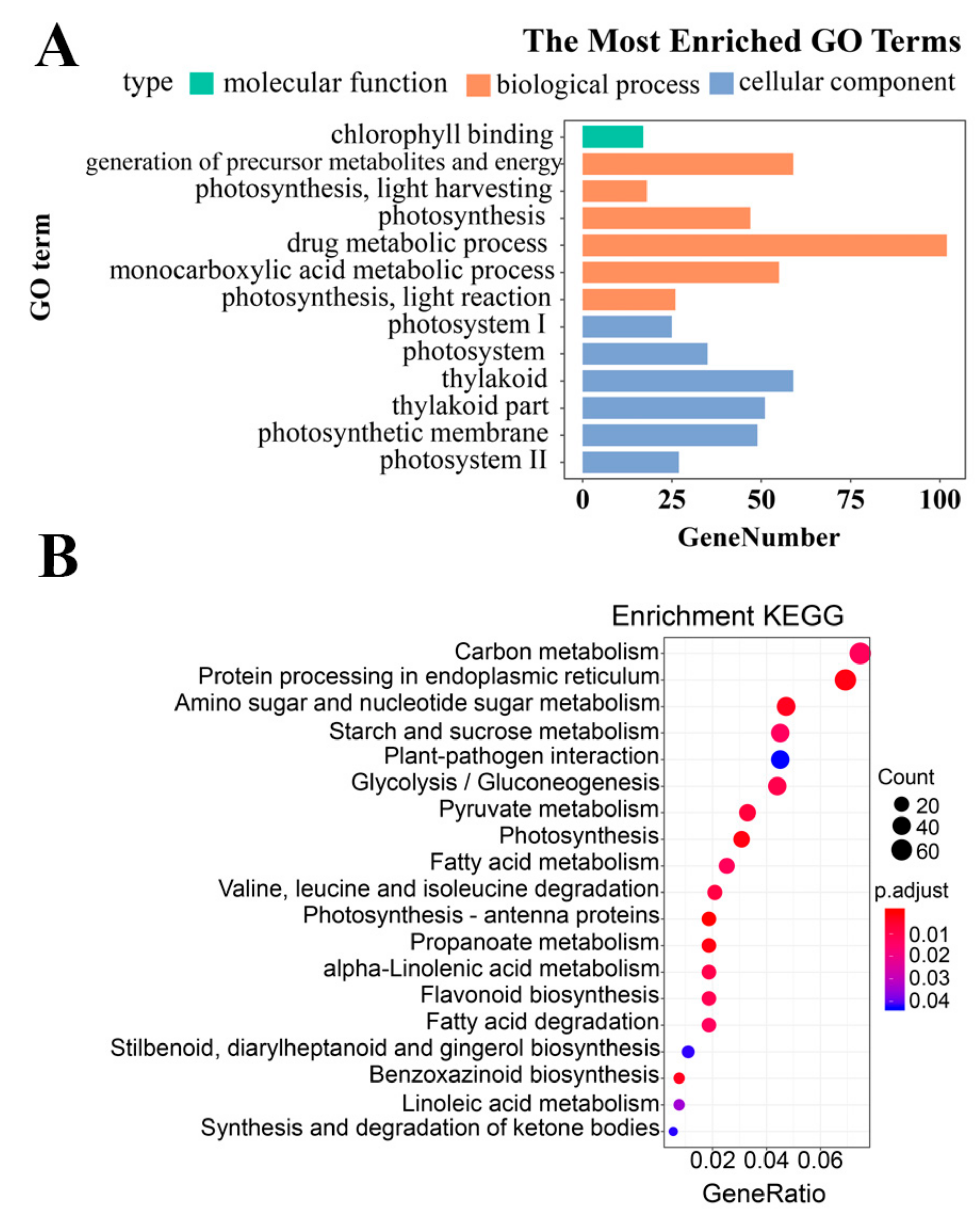
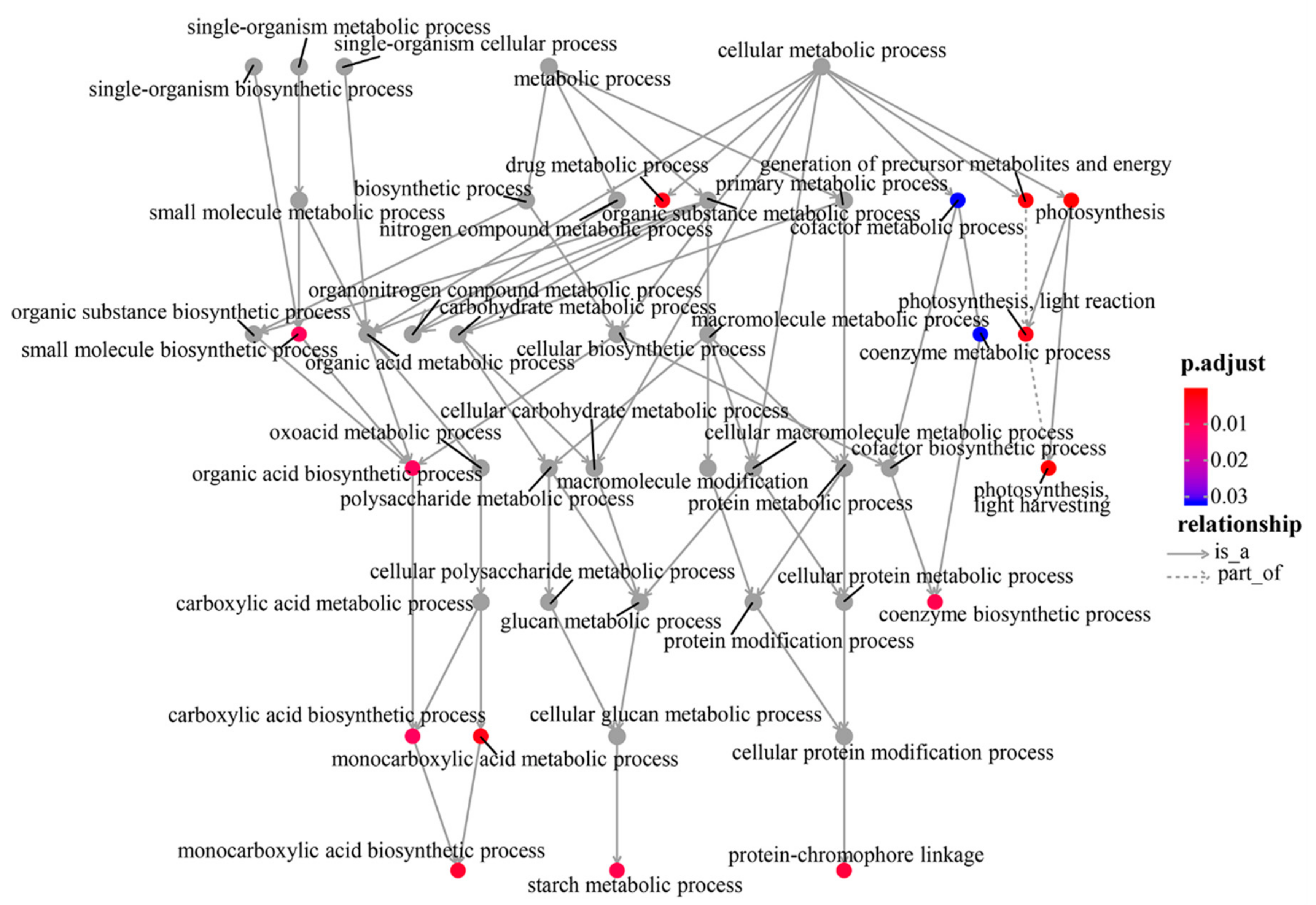
3.2. Differentially Expressed Gene Identification
3.3. Characterizing the Expression of Transcription Factor Genes in Leaf Senescence
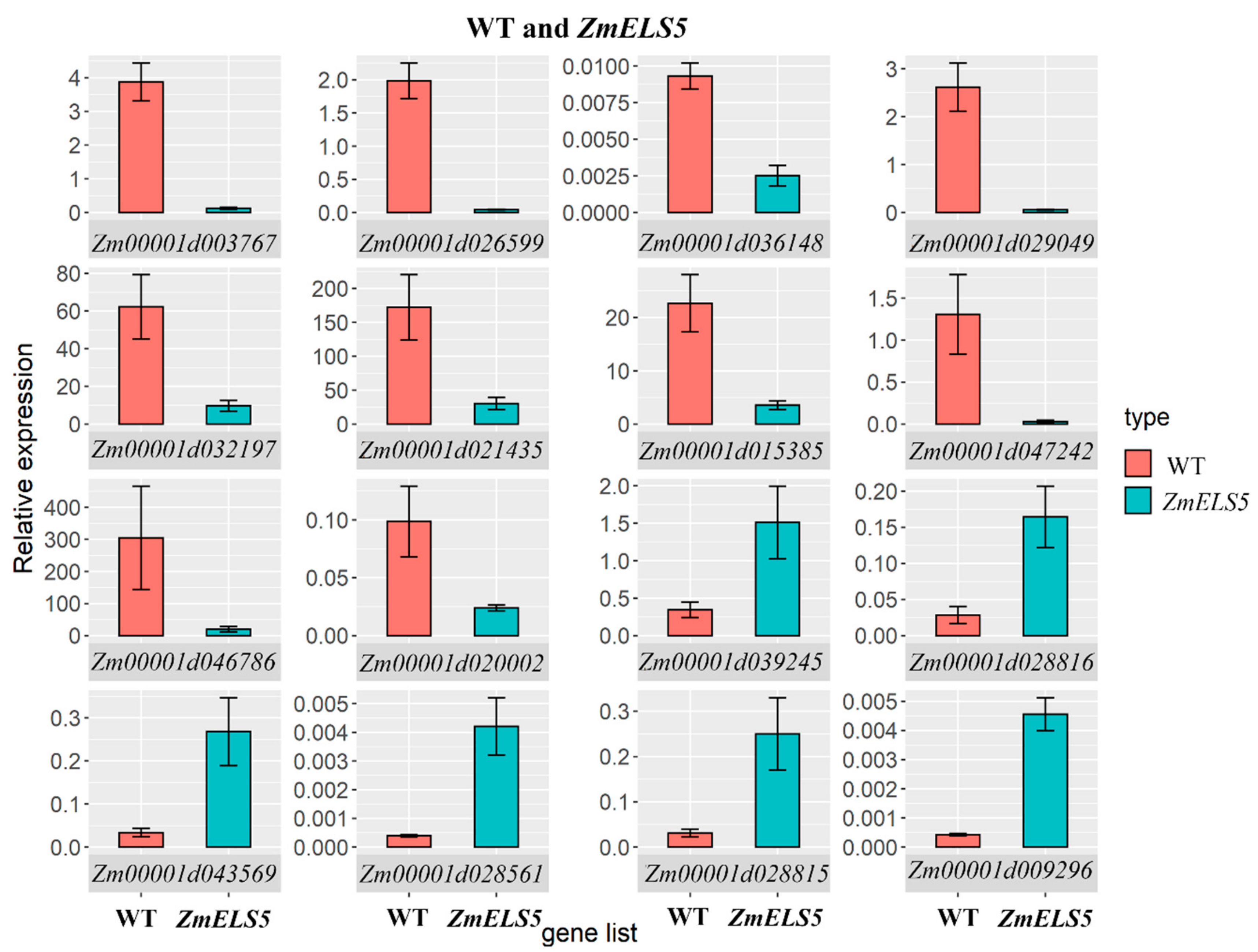
3.4. Verifying Differential Gene Expression Profiles Using RT-qPCR
4. Discussion
4.1. Key Genes in ZmELS5 Leaf Premature Senescence
4.2. Transcription Factors Involved in the Regulation of ZmELS5 Leaf Senescence
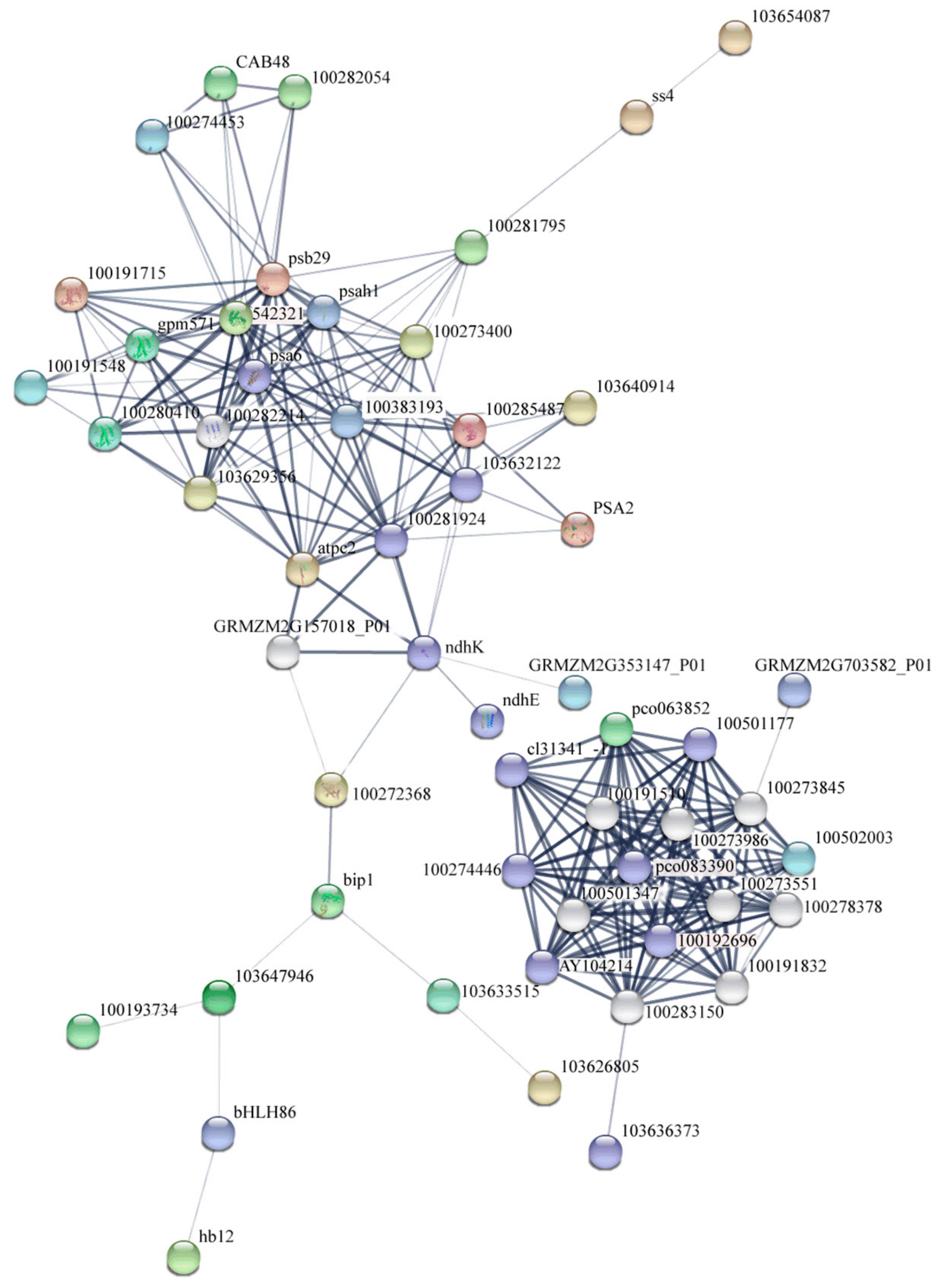
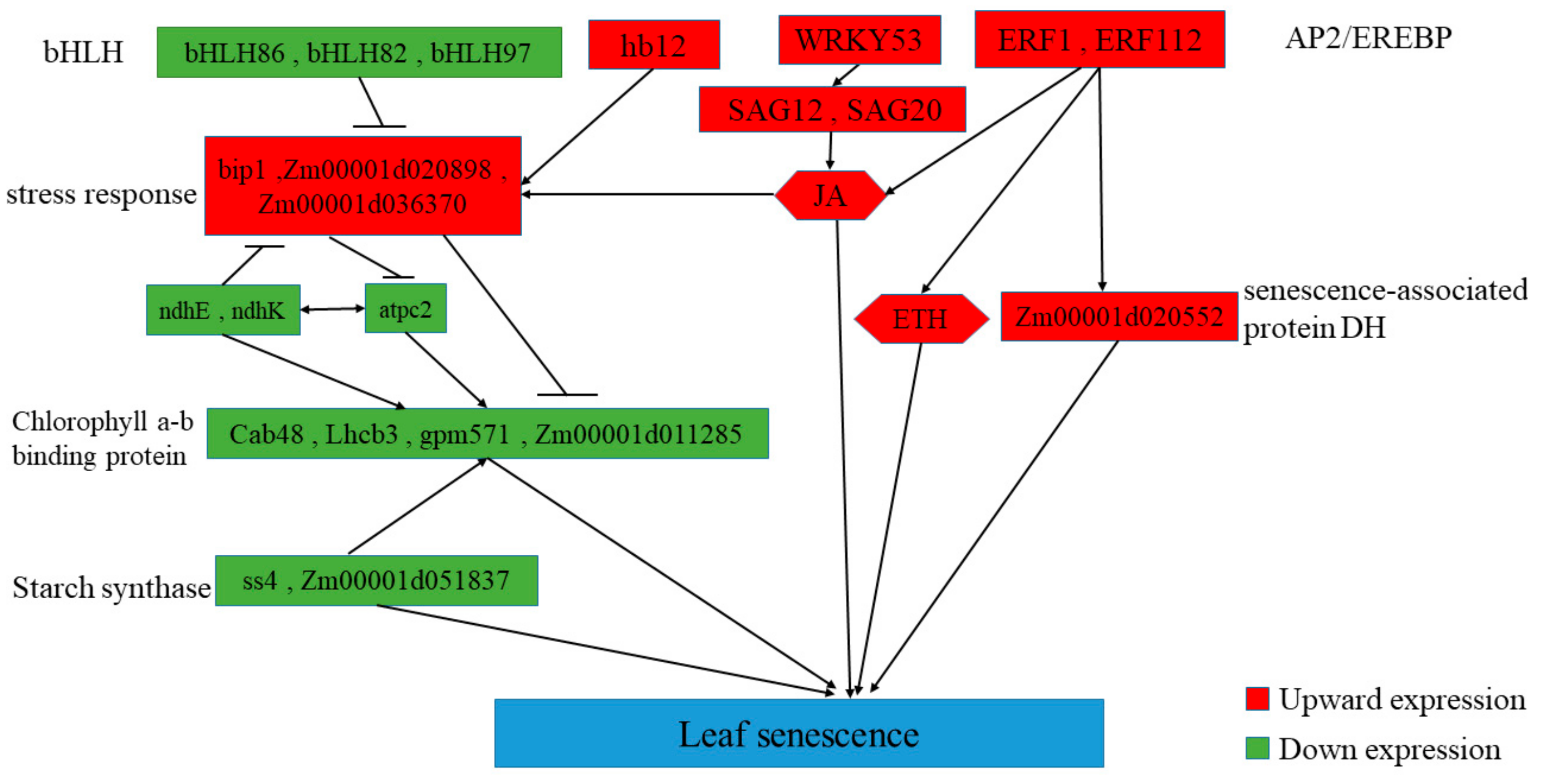
4.3. Predicted Regulatory Networks of Maize Leaf Senescence
4.4. SAG Mutant Identification and the Dissection of Related Regulatory Mechanisms
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Woo, H.R.; Masclaux-Daubresse, C.; Lim, P.O. Plant senescence: How plants know when and how to die. J. Exp. Bot. 2018, 69, 715–718. [Google Scholar] [CrossRef] [PubMed]
- Hollmann, J.; Gregersen, P.L.; Krupinska, K. Identification of predominant genes involved in regulation and execution of senescence-associated nitrogen remobilization in flag leaves of field grown barley. J. Exp. Bot. 2014, 65, 3963–3973. [Google Scholar] [CrossRef] [PubMed]
- Kohl, S.; Hollmann, J.; Blattner, F.R.; Radchuk, V.; Andersch, F.; Steuernagel, B.; Schmutzer, T.; Scholz, U.; Krupinska, K.; Weber, H.; et al. A putative role for amino acid permeases in sink-source communication of barley tissues uncovered by RNA-seq. BMC Plant Biol. 2012, 12, 154. [Google Scholar]
- Ra, R. Selectable traits to increase crop photosynthesis and yield of grain crops. J. Exp. Bot. 2000, 51, 447–458. [Google Scholar]
- Wu, X.; Ding, D.; Shi, C.; Xue, Y.; Zhang, Z.; Tang, G.; Tang, J. microRNA-dependent gene regulatory networks in maize leaf senescence. BMC Plant Biol. 2016, 16, 73. [Google Scholar] [CrossRef] [PubMed]
- Robson, P.; Mos, M.; Clifton-Brown, J.; Donnison, I. Phenotypic Variation in Senescence in Miscanthus: Towards Optimising Biomass Quality and Quantity. BioEnergy Res. 2012, 5, 95–105. [Google Scholar] [CrossRef]
- Gregersen, P.L.; Culetic, A.; Boschian, L.; Krupinska, K. Plant senescence and crop productivity. Plant Mol. Biol. 2013, 82, 603–622. [Google Scholar] [CrossRef]
- Khan, M.; Rozhon, W.; Poppenberger, B. The role of hormones in the aging of plants—A mini-review. Gerontology 2014, 60, 49–55. [Google Scholar] [CrossRef]
- Li, H.; Wang, S.; Li, B.; Tong, Y.; Yang, X.; Li, Z. Comparative study on physiological traits related with grain filling and photosynthesis of flag leaf in early aging and normal near--isogenic lines of common wheat. Acta Agron. Sin. 2006, 32, 1642–1648. [Google Scholar]
- Pleuel, H.; Ojanpera, K.; Danielsson, H.; Sild, E.; Gelang, J.; Wallin, G.; Skärby, L.; Sellden, G.; Fl, P. Effects of Ozone on Leaf Senescence in Spring Wheat-Possible Consequences for Grain Yield; Food and Agriculture Organization: Rome, Italy, 1997; Volume 37. [Google Scholar]
- Koyama, T. The roles of ethylene and transcription factors in the regulation of onset of leaf senescence. Front. Plant Sci. 2014, 5, 650. [Google Scholar] [CrossRef]
- Sarwat, M.; Naqvi, A.R.; Ahmad, P.; Ashraf, M.; Akram, N.A. Phytohormones and microRNAs as sensors and regulators of leaf senescence: Assigning macro roles to small molecules. Biotechnol. Adv. 2013, 31, 1153–1171. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, B.; Du, M.; Eneji, A.E.; Wang, B.; Duan, L.; Li, Z.; Tian, X. Mechanism of phytohormone involvement in feedback regulation of cotton leaf senescence induced by potassium deficiency. J. Exp. Bot. 2012, 63, 5887–5901. [Google Scholar] [CrossRef] [PubMed]
- Quirino, B.F.; Noh, Y.-S.; Himelblau, E.; Amasino, R.M. Molecular aspects of leaf senescence. Trends Plant Sci. 2000, 5, 278–282. [Google Scholar] [CrossRef]
- Lim, P.O.; Kim, H.J.; Gil Nam, H. Leaf Senescence. Annu. Rev. Plant Biol. 2007, 58, 115–136. [Google Scholar] [CrossRef]
- Thomas, H. Senescence, ageing and death of the whole plant. New Phytol. 2013, 197, 696–711. [Google Scholar] [CrossRef]
- Schippers, J.H.M. Transcriptional networks in leaf senescence. Curr. Opin. Plant Biol. 2015, 27, 77–83. [Google Scholar] [CrossRef]
- Lee, E.A.; Tollenaar, M. Physiological Basis of Successful Breeding Strategies for Maize Grain Yield. Crop Sci. 2007, 47, S202–S215. [Google Scholar] [CrossRef]
- Wingler, A.; Masclaux-Daubresse, C.; Fischer, A.M. Sugars, senescence, and ageing in plants and heterotrophic organisms. J. Exp. Bot. 2009, 60, 1063–1066. [Google Scholar] [CrossRef]
- Thomas, H.; Ougham, H. The stay-green trait. J. Exp. Bot. 2014, 65, 3889–3900. [Google Scholar] [CrossRef]
- Breeze, E.; Harrison, E.; McHattie, S.; Hughes, L.; Hickman, R.; Hill, C.; Kiddle, S.; Kim, Y.-S.; Penfold, C.A.; Jenkins, D. High-resolution temporal profiling of transcripts during Arabidopsis leaf senescence reveals a distinct chronology of processes and regulation. Plant Cell 2011, 23, 873–894. [Google Scholar] [CrossRef]
- Buchanan-Wollaston, V.; Earl, S.; Harrison, E.; Mathas, E.; Navabpour, S.; Page, T.; Pink, D. The molecular analysis of leaf senescence—A genomics approach. Plant Biotechnol. J. 2003, 1, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.-Y.; Hu, W.-J.; Luo, H.; Xia, Y.; Zhao, Y.; Wang, L.-D.; Zhang, L.-M.; Luo, J.-C.; Jing, H.-C. Transcriptome profiling of developmental leaf senescence in sorghum (Sorghum bicolor). Plant Mol. Biol. 2016, 92, 555–580. [Google Scholar] [CrossRef] [PubMed]
- Zentgraf, U.; Doll, J.; Riester, L. Live and Let Die: The Core Circadian Oscillator Coordinates Plant Life History and Pilots Leaf Senescence. Mol. Plant 2018, 11, 351–353. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kim, J.H.; Lyu, J.I.; Woo, H.R.; Lim, P.O. New insights into the regulation of leaf senescence in Arabidopsis. J. Exp. Bot. 2018, 69, 787–799. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Woo, H.R.; Nam, H.G. Toward Systems Understanding of Leaf Senescence: An Integrated Multi-Omics Perspective on Leaf Senescence Research. Mol. Plant 2016, 9, 813–825. [Google Scholar] [CrossRef] [PubMed]
- Podzimska-Sroka, D.; O’Shea, C.; Gregersen, P.L.; Skriver, K. NAC Transcription Factors in Senescence: From Molecular Structure to Function in Crops. Plants 2015, 4, 412–448. [Google Scholar] [CrossRef]
- Woo, H.R.; Kim, H.J.; Nam, H.G.; Lim, P.O. Plant leaf senescence and death - regulation by multiple layers of control and implications for aging in general. J. Cell Sci. 2013, 126, 4823–4833. [Google Scholar] [CrossRef]
- Li, Z.H.; Zhao, Y.; Liu, X.C.; Peng, J.Y.; Guo, H.W.; Luo, J.C. LSD 2.0: An update of the leaf senescence database. Nucleic Acids Res. 2014, 42, D1200–D1205. [Google Scholar] [CrossRef]
- Duvick, D.N. Genetic contributions to yield gains of US hybrid maize, 1930 to 1980. In Genetic Contributions to Yield Gains of Five Major Crop Plants; CSSA Special Publication Number 7; CSSA: Fitchburg, WI, USA; ASA: Madison, WI, USA, 1984; pp. 15–47. [Google Scholar]
- Tollenaar, M.; Wu, J. Yield improvement in temperate maize is attributable to greater stress tolerance. Crop Sci. 1999, 39, 1597–1604. [Google Scholar] [CrossRef]
- Guo, Y.; Gan, S.-S. Translational researches on leaf senescence for enhancing plant productivity and quality. J. Exp. Bot. 2014, 65, 3901–3913. [Google Scholar] [CrossRef]
- Jehanzeb, M.; Zheng, X.; Miao, Y. The role of the S40 gene family in leaf senescence. Int. J. Mol. Sci. 2017, 18, 2152. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.; Wang, Y.; Zhu, Y.; Tang, J.; Hu, B.; Liu, L.; Ou, S.; Wu, H.; Sun, X.; Chu, J. OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. Proc. Natl. Acad. Sci. USA 2014, 111, 10013–10018. [Google Scholar] [CrossRef] [PubMed]
- Sekhon, R.S.; Childs, K.L.; Santoro, N.; Foster, C.E.; Buell, C.R.; de Leon, N.; Kaeppler, S.M. Transcriptional and metabolic analysis of senescence induced by preventing pollination in maize. Plant Physiol. 2012, 159, 1730–1744. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Geng, M.; Shen, P.; Chen, X.; Li, Y.; Wen, X. Effect of post-silking drought stress on the expression profiles of genes involved in carbon and nitrogen metabolism during leaf senescence in maize (Zea mays L.). Plant Physiol. Biochem. PPB 2019, 135, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Li, M.; Tian, L.; Wang, S.; Wu, L.; Ku, L.; Zhang, J.; Song, X.; Liu, H.; Chen, Y. Global transcriptome analysis of the maize (Zea mays L.) inbred line 08LF during leaf senescence initiated by pollination-prevention. PLoS ONE 2017, 12, e0185838. [Google Scholar] [CrossRef]
- Zhang, W.Y.; Xu, Y.C.; Li, W.L.; Yang, L.; Yue, X.; Zhang, X.S.; Zhao, X.Y. Transcriptional analyses of natural leaf senescence in maize. PLoS ONE 2014, 9, e115617. [Google Scholar] [CrossRef]
- Wu, L.; Wang, S.; Tian, L.; Wu, L.; Li, M.; Zhang, J.; Li, P.; Zhang, W.; Chen, Y. Comparative proteomic analysis of the maize responses to early leaf senescence induced by preventing pollination. J. Proteom. 2018, 177, 75–87. [Google Scholar] [CrossRef]
- Wei, S.; Wang, X.; Zhang, J.; Liu, P.; Zhao, B.; Li, G.; Dong, S. The role of nitrogen in leaf senescence of summer maize and analysis of underlying mechanisms using comparative proteomics. Plant Sci. Int. J. Exp. Plant Biol. 2015, 233, 72–81. [Google Scholar] [CrossRef]
- Wang, J.; Guan, H.; Dong, R.; Liu, C.; Liu, Q.; Liu, T.; Wang, L.; He, C. Overexpression of maize sucrose non-fermenting-1-related protein kinase 1 genes, ZmSnRK1s, causes alteration in carbon metabolism and leaf senescence in Arabidopsis thaliana. Gene 2019, 691, 34–44. [Google Scholar] [CrossRef]
- Luo, K.; Deng, W.; Xiao, Y.; Zheng, X.; Li, Y.; Pei, Y. Leaf senescence is delayed in tobacco plants expressing the maize knotted1 gene under the control of a wound-inducible promoter. Plant Cell Rep. 2006, 25, 1246–1254. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Landmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Kim, D.; Pertea, G.M.; Leek, J.T.; Salzberg, S.L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 2016, 11, 1650–1667. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Zhu, A.; Ibrahim, J.G.; Love, M.I. Heavy-tailed prior distributions for sequence count data: Removing the noise and preserving large differences. Bioinformatics 2019, 35, 2084–2092. [Google Scholar] [CrossRef]
- Yu, G.C.; Wang, L.G.; Han, Y.Y.; He, Q.Y. clusterProfiler: An R Package for Comparing Biological Themes Among Gene Clusters. Omics 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Morgan, M.; Carlson, M.; Tenenbaum, D.; Arora, S. AnnotationHub: Client to access AnnotationHub resources. R Package Version 2017, 2. Available online: https://rdrr.io/bioc/AnnotationHub/ (accessed on 19 November 2019).
- Yu, G. clusterProfiler: An universal enrichment tool for functional and comparative study. bioRxiv 2018, 256784. [Google Scholar] [CrossRef]
- Yu, C.-P.; Chen, S.C.-C.; Chang, Y.-M.; Liu, W.-Y.; Lin, H.-H.; Lin, J.-J.; Chen, H.J.; Lu, Y.-J.; Wu, Y.-H.; Lu, M.-Y.J. Transcriptome dynamics of developing maize leaves and genomewide prediction of cis elements and their cognate transcription factors. Proc. Natl. Acad. Sci. USA 2015, 112, E2477–E2486. [Google Scholar] [CrossRef]
- Tian, T.; You, Q.; Yan, H.Y.; Xu, W.Y.; Su, Z. MCENet: A database for maize conditional co-expression network and network characterization collaborated with multi-dimensional omics levels. J. Genet. Genom. 2018, 45, 351–360. [Google Scholar] [CrossRef]
- Wang, W.; Dai, Y.; Wang, M.; Yang, W.; Zhao, D. Transcriptome Dynamics of Double Recessive Mutant, o2o2o16o16, Reveals the Transcriptional Mechanisms in the Increase of Its Lysine and Tryptophan Content in Maize. Genes 2019, 10, 316. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Porra, R.J. The chequered history of the development and use of simultaneous equations for the accurate determination of chlorophylls a and b. Photosynth. Res. 2002, 73, 149–156. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2018, 47, D607–D613. [Google Scholar] [CrossRef]
- Yoshida, S. Molecular regulation of leaf senescence. Curr. Opin. Plant Biol. 2003, 6, 79–84. [Google Scholar] [CrossRef]
- Hollander-Czytko, H.; Grabowski, J.; Sandorf, I.; Weckermann, K.; Weiler, E.W. Tocopherol content and activities of tyrosine aminotransferase and cystine lyase in Arabidopsis under stress conditions. J. Plant Physiol. 2005, 162, 767–770. [Google Scholar] [CrossRef]
- Kesari, P.; Patil, D.N.; Kumar, P.; Tomar, S.; Sharma, A.K.; Kumar, P. Structural and functional evolution of chitinase-like proteins from plants. Proteomics 2015, 15, 1693–1705. [Google Scholar] [CrossRef]
- Mazorra-Manzano, M.A.; Tanaka, T.; Dee, D.R.; Yada, R.Y. Structure-function characterization of the recombinant aspartic proteinase A1 from Arabidopsis thaliana. Phytochemistry 2010, 71, 515–523. [Google Scholar] [CrossRef]
- Xu, H.; Xu, W.; Xi, H.; Ma, W.; He, Z.; Ma, M. The ER luminal binding protein (BiP) alleviates Cd(2+)-induced programmed cell death through endoplasmic reticulum stress-cell death signaling pathway in tobacco cells. J. Plant Physiol. 2013, 170, 1434–1441. [Google Scholar] [CrossRef]
- John, C.F.; Morris, K.; Jordan, B.R.; Thomas, B.; A-H-Mackerness, S. Ultraviolet-B exposure leads to up-regulation of senescence-associated genes in Arabidopsis thaliana. J. Exp. Bot. 2001, 52, 1367–1373. [Google Scholar] [CrossRef]
- Jiang, Y.; Deyholos, M.K. Functional characterization of Arabidopsis NaCl-inducible WRKY25 and WRKY33 transcription factors in abiotic stresses. Plant Mol. Biol. 2009, 69, 91–105. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.M.; An, J.; Han, H.J.; Kim, S.H.; Lim, C.O.; Yun, D.J.; Chung, W.S. ZAT11, a zinc finger transcription factor, is a negative regulator of nickel ion tolerance in Arabidopsis. Plant Cell Rep. 2014, 33, 2015–2021. [Google Scholar] [CrossRef] [PubMed]
- Dubos, C.; Stracke, R.; Grotewold, E.; Weisshaar, B.; Martin, C.; Lepiniec, L. MYB transcription factors in Arabidopsis. Trends Plant Sci. 2010, 15, 573–581. [Google Scholar] [CrossRef] [PubMed]
- Majumdar, R.; Rajasekaran, K.; Sickler, C.; Lebar, M.; Musungu, B.M.; Fakhoury, A.M.; Payne, G.A.; Geisler, M.; Carter-Wientjes, C.; Wei, Q.; et al. The Pathogenesis-Related Maize Seed (PRms) Gene Plays a Role in Resistance to Aspergillus flavus Infection and Aflatoxin Contamination. Front. Plant Sci. 2017, 8, 1758. [Google Scholar] [CrossRef]
- Hawkins, L.K.; Warburton, M.L.; Tang, J.D.; Tomashek, J.; Alves Oliveira, D.; Ogunola, O.F.; Smith, J.S.; Williams, W.P. Survey of Candidate Genes for Maize Resistance to Infection by Aspergillus flavus and/or Aflatoxin Contamination. Toxins 2018, 10, 61. [Google Scholar] [CrossRef]
- Yuan, G.; He, X.; Li, H.; Xiang, K.; Liu, L.; Zou, C.; Lin, H.; Zhang, Z.; Pan, G. Transcriptomic responses in resistant and susceptible maize infected with Fusarium graminearum. Crop J. 2019, 2214–5141. [Google Scholar] [CrossRef]
- Yang, Q.; Balint-Kurti, P.; Xu, M. Quantitative Disease Resistance: Dissection and Adoption in Maize. Mol. Plant 2017, 10, 402–413. [Google Scholar] [CrossRef]
- Balazadeh, S.; Riaño-Pachón, D.M.; Mueller-Roeber, B. Transcription factors regulating leaf senescence in Arabidopsis thaliana. Plant Biol. 2008, 10, 63–75. [Google Scholar] [CrossRef]
- Wei, L.; Guo, Y. Transcriptome, transcription factors and transcriptional regulation of leaf senescence. J. Bioinform. Comp. Genom. 2014, 1, 1. [Google Scholar]
- Rushton, D.L.; Tripathi, P.; Rabara, R.C.; Lin, J.; Ringler, P.; Boken, A.K.; Langum, T.J.; Smidt, L.; Boomsma, D.D.; Emme, N.J. WRKY transcription factors: Key components in abscisic acid signalling. Plant Biotechnol. J. 2012, 10, 2–11. [Google Scholar] [CrossRef]
- Miao, Y.; Laun, T.; Zimmermann, P.; Zentgraf, U. Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol. Biol. 2004, 55, 853–867. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, Z.; Wang, X.; Wang, J.; Fan, K.; Li, Z.; Lin, W. DELLA proteins negatively regulate dark-induced senescence and chlorophyll degradation in Arabidopsis through interaction with the transcription factor WRKY6. Plant Cell Rep. 2018, 37, 981–992. [Google Scholar] [CrossRef] [PubMed]
- Gregersen, P.L.; Holm, P.B. Transcriptome analysis of senescence in the flag leaf of wheat (Triticum aestivum L.). Plant Biotechnol. J. 2007, 5, 192–206. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Woo, H.R.; Kim, J.; Lim, P.O.; Lee, I.C.; Choi, S.H.; Hwang, D.; Nam, H.G. Trifurcate Feed-Forward Regulation of Age-Dependent Cell Death Involving miR164 in Arabidopsis. Science 2009, 323, 1053–1057. [Google Scholar] [CrossRef]
- Balazadeh, S.; Kwasniewski, M.; Caldana, C.; Mehrnia, M.; Zanor, M.I.; Xue, G.-P.; Mueller-Roeber, B. ORS1, an H2O2-Responsive NAC Transcription Factor, Controls Senescence in Arabidopsis thaliana. Mol. Plant 2011, 4, 346–360. [Google Scholar] [CrossRef]
- Qi, T.; Wang, J.; Huang, H.; Liu, B.; Gao, H.; Liu, Y.; Song, S.; Xie, D. Regulation of Jasmonate-Induced Leaf Senescence by Antagonism between bHLH Subgroup IIIe and IIId Factors in Arabidopsis. Plant Cell 2015, 27, 1634–1649. [Google Scholar] [CrossRef]
- Song, S.; Huang, H.; Wang, J.; Liu, B.; Qi, T.; Xie, D. MYC5 is Involved in Jasmonate-Regulated Plant Growth, Leaf Senescence and Defense Responses. Plant Cell Physiol. 2017, 58, 1752–1763. [Google Scholar] [CrossRef]
- Huang, C.-K.; Lo, P.-C.; Huang, L.-F.; Wu, S.-J.; Yeh, C.-H.; Lu, C.-A. A single-repeat MYB transcription repressor, MYBH, participates in regulation of leaf senescence in Arabidopsis. Plant Mol. Biol. 2015, 88, 269–286. [Google Scholar] [CrossRef]
- Moschen, S.; Bengoa Luoni, S.; Di Rienzo, J.A.; Caro, M.D.P.; Tohge, T.; Watanabe, M.; Hollmann, J.; González, S.; Rivarola, M.; García-García, F.; et al. Integrating transcriptomic and metabolomic analysis to understand natural leaf senescence in sunflower. Plant Biotechnol. J. 2016, 14, 719–734. [Google Scholar] [CrossRef]
- Bengoa Luoni, S.; Astigueta, F.H.; Nicosia, S.; Moschen, S.; Fernandez, P.; Heinz, R. Transcription Factors Associated with Leaf Senescence in Crops. Plants 2019, 8, 411. [Google Scholar] [CrossRef]
- Ullah, A.; Akbar, A.; Yang, X. Jasmonic Acid (JA)-Mediated Signaling in Leaf Senescence. In Senescence Signalling and Control in Plants; Sarwat, M., Tuteja, N., Eds.; Academic Press: Cambridge, MA, USA, 2019; Chapter 7; pp. 111–123. [Google Scholar]
- Zhang, Y.; Wang, Y.; Wei, H.; Li, N.; Tian, W.; Chong, K.; Wang, L. Circadian Evening Complex Represses Jasmonate-Induced Leaf Senescence in Arabidopsis. Mol. Plant 2018, 11, 326–337. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Jiang, Y.; Han, X.; Wang, H.; Pan, J.; Yu, D. Jasmonate regulates leaf senescence and tolerance to cold stress: Crosstalk with other phytohormones. J. Exp. Bot. 2017, 68, 1361–1369. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Provart, N.J.; Glazebrook, J.; Katagiri, F.; Chang, H.-S.; Eulgem, T.; Mauch, F.; Luan, S.; Zou, G.; Whitham, S.A.; et al. Expression Profile Matrix of Arabidopsis Transcription Factor Genes Suggests Their Putative Functions in Response to Environmental Stresses. Plant Cell 2002, 14, 559–574. [Google Scholar] [CrossRef] [PubMed]
- Martín, M.; Casano, L.M.; Sabater, B. Identification of the Product of ndhA Gene as a Thylakoid Protein Synthesized in Response to Photooxidative Treatment. Plant Cell Physiol. 1996, 37, 293–298. [Google Scholar] [CrossRef]
- Davis, G.A.; Kanazawa, A.; Schöttler, M.A.; Kohzuma, K.; Froehlich, J.E.; Rutherford, A.W.; Satoh-Cruz, M.; Minhas, D.; Tietz, S.; Dhingra, A.; et al. Limitations to photosynthesis by proton motive force-induced photosystem II photodamage. ELife 2016, 5, e16921. [Google Scholar] [CrossRef]
- Gámez-Arjona, F.M.; Raynaud, S.; Ragel, P.; Mérida, Á. Starch synthase 4 is located in the thylakoid membrane and interacts with plastoglobule-associated proteins in Arabidopsis. Plant J. 2014, 80, 305–316. [Google Scholar] [CrossRef]
- Trobacher, C.P. Ethylene and programmed cell death in plants. Botany 2009, 87, 757–769. [Google Scholar] [CrossRef]
- Grosskinsky, D.K.; Syaifullah, S.J.; Roitsch, T. Integration of multi-omics techniques and physiological phenotyping within a holistic phenomics approach to study senescence in model and crop plants. J. Exp. Bot. 2018, 69, 825–844. [Google Scholar] [CrossRef]

| Sample | Raw Data Reads | Reads After Filter | Clean Reads Rate (%) | Mapped Reads | Mapped Rate (%) | Unique Mapped Reads | Unique Mapped Rate (%) |
|---|---|---|---|---|---|---|---|
| ZmELS 5-1 | 40,957,988 | 39,400,718 | 96.20 | 35,598,549 | 90.35 | 33,880,944 | 85.99 |
| ZmELS 5-2 | 40,996,812 | 39,317,518 | 95.90 | 35,749,713 | 91.18 | 34,040,734 | 86.58 |
| ZmELS 5-3 | 42,892,702 | 41,233,326 | 96.13 | 37,786,220 | 91.64 | 36,022,486 | 87.36 |
| WT-1 | 42,805,416 | 41,244,366 | 96.35 | 37,421,013 | 90.73 | 35,435,962 | 85.92 |
| WT-2 | 40,873,684 | 39,408,286 | 96.41 | 36,511,777 | 92.65 | 35,173,768 | 89.25 |
| WT-3 | 42,401,032 | 40,813,916 | 96.26 | 37,254,943 | 91.28 | 35,383,646 | 86.70 |
| Gene ID | BaseMean | Log2 Fold Change | p-Value | Homologous Arabidopsis Genes | Chr | Annotation of Maize | Source |
|---|---|---|---|---|---|---|---|
| ZemaCp012 | 13.17 | −2 | 0.001891 | ATCG00210 | Pt | cytochrome b6/f complex subunit N | GSEA_CC |
| Zm00001d039040 | 170,043.95 | −2.02 | 0.000026 | AT2G34420 | 6 | light harvesting complex mesophyll 7 | GO_BP,GSEA_CC |
| ZemaCp019 | 1122.6 | −2.03 | 0.000237 | ATCG00130 | Pt | ATPase subunit I | GSEA_CC |
| Zm00001d032197 | 39,958.88 | −2.11 | 0.005955 | AT3G47470 | 1 | Chlorophyll a-b binding protein 4 chloroplastic | GO_BP,GSEA_CC |
| Zm00001d032331 | 1541.9 | −2.15 | 8.35 × 10−8 | AT3G01480 | 1 | peptidyl-prolyl cis-trans isomerase | GSEA_CC |
| ZemaCp041 | 48.41 | −2.18 | 0.000427 | ATCG00590 | Pt | cytochrome b6/f complex subunit VI | GSEA_CC |
| Zm00001d050403 | 23,382.97 | −2.19 | 0.006146 | AT3G47470 | 4 | chlorophyll a-b binding protein 4 | GO_BP,GSEA_CC |
| Zm00001d018157 | 5238.44 | −2.22 | 7.28 × 10−7 | AT1G45474 | 5 | light harvesting complex a/b protein 4 | GO_BP,GSEA_CC |
| Zm00001d020030 | 522.23 | −2.23 | 0.003533 | 7 | putative NAD(P)H dehydrogenase subunit CRR3 chloroplastic | GSEA_CC | |
| ZemaCp042 | 108.17 | −2.24 | 0.002738 | ATCG00600 | Pt | cytochrome B6-F complex subunit 5 | GSEA_CC |
| ZemaCp043 | 18.23 | −2.27 | 0.000883 | ATCG00630 | Pt | photosystem I subunit IX | GSEA_CC |
| ZemaCp088 | 28.03 | −2.28 | 0.002185 | ATCG01070 | Pt | NADH dehydrogenase subunit 4L | GSEA_CC |
| Zm00001d038984 | 57,330.44 | −2.31 | 0.001118 | AT3G16140 | 6 | photosystem I H subunit 1 | GSEA_CC |
| Zm00001d030762 | 19,126.21 | −2.33 | 0.001473 | AT5G08050 | 1 | GSEA_CC | |
| Zm00001d040242 | 401.87 | −2.36 | 0.001072 | AT1G71480 | 3 | Nuclear transport factor 2 (NTF2) family protein | GSEA_CC |
| Zm00001d031738 | 2102.21 | −2.38 | 0.005397 | AT2G34860 | 1 | photosystem I | GSEA_CC |
| Zm00001d009877 | 4724.03 | −2.43 | 0.007421 | AT3G46780 | 8 | Protein plastid transcriptionally active 16 chloroplastic | GSEA_CC |
| ZemaCp011 | 119.68 | −2.45 | 2.44 × 10−8 | ATCG00220 | Pt | photosystem II protein M | GSEA_CC |
| Zm00001d021763 | 119,242.53 | −2.49 | 0.00516 | AT2G40100 | 7 | photosystem II subunit 29 | GO_BP,GSEA_CC |
| Zm00001d023757 | 6515.79 | −2.52 | 0.000234 | AT5G21430 | 10 | NAD(P)H-quinone oxidoreductase subunit U chloroplastic | GSEA_CC |
| Zm00001d021620 | 23,073.49 | −2.61 | 8.91 × 10−12 | AT4G04640 | 7 | ATP synthase chloroplast subunit 2 | GSEA_CC |
| Zm00001d006663 | 2511.32 | −2.69 | 0.000695 | AT3G61470 | 2 | chlorophyll a-b binding protein | GO_BP,GSEA_CC |
| Zm00001d026599 | 22,411.85 | −2.81 | 0.004233 | AT1G15820 | 10 | light harvesting chlorophyll a/b binding protein 6 | GO_BP,GSEA_CC |
| Zm00001d006587 | 18,274.92 | −2.87 | 0.004411 | AT3G08940 | 2 | Chlorophyll a-b binding protein CP29.1 chloroplastic | GO_BP,GSEA_CC |
| Zm00001d009589 | 26,091.22 | −2.93 | 0.0013 | AT2G34420 | 8 | light harvesting chlorophyll a/b binding protein 3 | GO_BP,GSEA_CC |
| Zm00001d045620 | 935.16 | −3.4 | 0.007634 | 9 | Plastocyanin major isoform chloroplastic | GSEA_CC | |
| Zm00001d044396 | 137.84 | −3.76 | 2.25 × 10−5 | AT2G34430 | 3 | chlorophyll a-b binding protein of LHCII type 1 | GO_BP,GSEA_CC |
| Zm00001d044399 | 1341.4 | −3.92 | 0.000709 | AT2G34430 | 3 | chlorophyll a-b binding protein 2 | GO_BP,GSEA_CC |
| Zm00001d011285 | 6415.27 | −4.45 | 3.32 × 10−7 | AT2G34420 | 8 | light harvesting chlorophyll a/b binding protein 1 | GO_BP,GSEA_CC |
| Zm00001d044401 | 1802.37 | −4.75 | 2.21 × 10−7 | AT2G34430 | 3 | chlorophyll a-b binding protein of LHCII type 1 | GO_BP,GSEA_CC |
| Zm00001d044402 | 2213.89 | −4.75 | 1.11 × 10−6 | AT2G34430 | 3 | chlorophyll a-b binding protein of LHCII type 1 | GO_BP,GSEA_CC |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chai, M.; Guo, Z.; Shi, X.; Li, Y.; Tang, J.; Zhang, Z. Dissecting the Regulatory Network of Leaf Premature Senescence in Maize (Zea mays L.) Using Transcriptome Analysis of ZmELS5 Mutant. Genes 2019, 10, 944. https://doi.org/10.3390/genes10110944
Chai M, Guo Z, Shi X, Li Y, Tang J, Zhang Z. Dissecting the Regulatory Network of Leaf Premature Senescence in Maize (Zea mays L.) Using Transcriptome Analysis of ZmELS5 Mutant. Genes. 2019; 10(11):944. https://doi.org/10.3390/genes10110944
Chicago/Turabian StyleChai, Mao, Zhanyong Guo, Xia Shi, Yingbo Li, Jihua Tang, and Zhanhui Zhang. 2019. "Dissecting the Regulatory Network of Leaf Premature Senescence in Maize (Zea mays L.) Using Transcriptome Analysis of ZmELS5 Mutant" Genes 10, no. 11: 944. https://doi.org/10.3390/genes10110944
APA StyleChai, M., Guo, Z., Shi, X., Li, Y., Tang, J., & Zhang, Z. (2019). Dissecting the Regulatory Network of Leaf Premature Senescence in Maize (Zea mays L.) Using Transcriptome Analysis of ZmELS5 Mutant. Genes, 10(11), 944. https://doi.org/10.3390/genes10110944






