Melanoma-Bearing Libechov Minipig (MeLiM): The Unique Swine Model of Hereditary Metastatic Melanoma
Abstract
1. Introduction
2. Human Melanoma
2.1. Incidence
2.2. Risk Factors
2.3. Genetic Background in Melanoma
2.3.1. Germline Mutations in Familial Melanoma
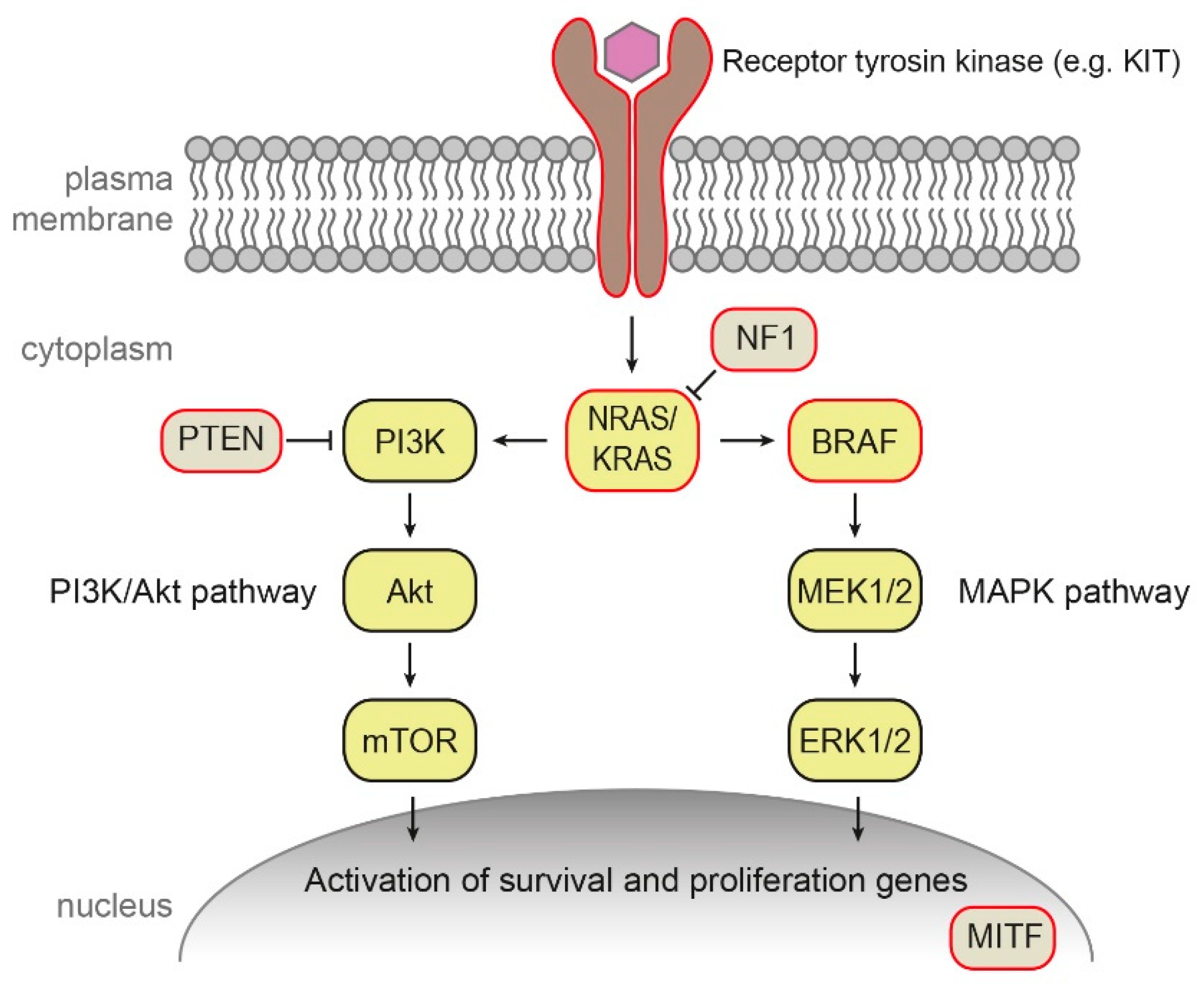
2.3.2. Somatic Mutations in Sporadic Melanoma
2.4. Regression
2.5. Therapy of Melanoma
3. Animal Models
3.1. Non-Mammalian Models
3.2. Mammalian Models
3.2.1. Mouse Models
3.2.2. Dog Models
3.2.3. Equine Models
4. Swine Melanoma Models
4.1. Sinclair Miniature Swine
4.2. Munich Miniature Swine Troll
5. The Melanoma-Bearing Libechov Minipig
5.1. Development of the MeLiM Model
5.2. Histopathological, Biochemical, and Immunohistochemical Characterization
5.3. MeLiM Melanoma Progression and Spontaneous Regression
5.4. Genetic Findings
5.5. Hematological Findings
5.6. Immunological Findings
5.7. Skin Microbiome
5.8. Experimental Therapy of MeLiM Melanoma by Tumor Devitalization
6. Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
Abbreviations
References
- Cummins, D.L.; Cummins, J.M.; Pantle, H.; Silverman, M.A.; Leonard, A.L.; Chanmugam, A. Cutaneous Malignant Melanoma. Mayo Clin. Proc. 2006, 81, 500–507. [Google Scholar] [CrossRef] [PubMed]
- Lomas, A.; Leonardi Bee, J.; Bath Hextall, F. A Systematic Review of Worldwide Incidence of Nonmelanoma Skin Cancer. Br. J. Dermatol. 2012, 166, 1069–1080. [Google Scholar] [CrossRef] [PubMed]
- Craythorne, E.; Al Niami, F. Skin Cancer. Medicine (Baltimore) 2017, 45, 431–434. [Google Scholar] [CrossRef]
- Garcovich, S.; Colloca, G.; Sollena, P.; Andrea, B.; Balducci, L.; Cho, W.C.; Bernabei, R.; Peris, K. Skin Cancer Epidemics in the Elderly as An Emerging Issue in Geriatric Oncology. Aging Dis. 2017, 8, 643–661. [Google Scholar] [CrossRef]
- Cichorek, M.; Wachulska, M.; Stasiewicz, A.; Tyminska, A. Skin Melanocytes: Biology and Development. Postepy Dermatol. Alergol. 2013, 30, 30–41. [Google Scholar] [CrossRef]
- McElearney, S.T.; Dengel, L.T.; Vaughters, A.B.R.; Patterson, J.W.; McGahren, E.D.; Slingluff, C.L. Neonatal Congenital Malignant Melanoma with Lymph Node Metastasis. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2009, 27, 2726–2728. [Google Scholar] [CrossRef]
- Perniciaro, C. Dermatopathologic Variants of Malignant Melanoma. Mayo Clin. Proc. 1997, 72, 273–279. [Google Scholar] [CrossRef]
- O’Brien, K.; Bhatia, A.; Tsen, F.; Chen, M.; Wong, A.K.; Woodley, D.T.; Li, W. Identification of the Critical Therapeutic Entity in Secreted Hsp90α that Promotes Wound Healing in Newly Re-Standardized Healthy and Diabetic Pig Models. PLoS ONE 2014, 9, e113956. [Google Scholar] [CrossRef]
- Summerfield, A.; Meurens, F.; Ricklin, M.E. The Immunology of the Porcine Skin and its Value as a Model for Human Skin. Mol. Immunol. 2015, 66, 14–21. [Google Scholar] [CrossRef] [PubMed]
- Erdmann, F.; Lortet Tieulent, J.; Schuz, J.; Zeeb, H.; Greinert, R.; Breitbart, E.W.; Bray, F. International Trends in the Incidence of Malignant Melanoma 1953–2008-Are Recent Generations at Higher or Lower Risk? Int. J. Cancer 2013, 132, 385–400. [Google Scholar] [CrossRef]
- Eggermont, A.M.; Spatz, A.; Robert, C. Cutaneous Melanoma. Lancet 2014, 383, 816–827. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer Statistics, 2019. CA Cancer J. Clin. 2019, 69, 7–34. [Google Scholar] [CrossRef] [PubMed]
- Rastrelli, M.; Tropea, S.; Rossi, C.R.; Alaibac, M. Melanoma: Epidemiology, Risk Factors, Pathogenesis, Diagnosis and Classification. Vivo Athens Greece 2014, 28, 1005–1011. [Google Scholar]
- Berwick, M.; Buller, D.B.; Cust, A.; Gallagher, R.; Lee, T.K.; Meyskens, F.; Pandey, S.; Thomas, N.E.; Veierod, M.B.; Ward, S. Melanoma Epidemiology and Prevention. Cancer Treat. Res. 2016, 167, 17–49. [Google Scholar] [PubMed]
- Leachman, S.A.; Lucero, O.M.; Sampson, J.E.; Cassidy, P.; Bruno, W.; Queirolo, P.; Ghiorzo, P. Identification, Genetic Testing, and Management of Hereditary Melanoma. Cancer Metastasis Rev. 2017, 36, 77–90. [Google Scholar] [CrossRef]
- Duggleby, W.F.; Stoll, H.; Priore, R.L.; Greenwald, P.; Graham, S. A Genetic Analysis of Melanoma--Polygenic Inheritance as a Threshold Trait. Am. J. Epidemiol. 1981, 114, 63–72. [Google Scholar] [CrossRef]
- Dracopoli, N.C.; Alhadeff, B.; Houghton, A.N.; Old, L.J. Loss of Heterozygosity at Autosomal and X-Linked Loci During Tumor Progression in a Patient with Melanoma. Cancer Res. 1987, 47, 3995–4000. [Google Scholar]
- Fountain, J.W.; Karayiorgou, M.; Ernstoff, M.S.; Kirkwood, J.M.; Vlock, D.R.; Titus-Ernstoff, L.; Bouchard, B.; Vijayasaradhi, S.; Houghton, A.N.; Lahti, J. Homozygous Deletions within Human Chromosome Band 9p21 in Melanoma. Proc. Natl. Acad. Sci. USA 1992, 89, 10557–10561. [Google Scholar] [CrossRef]
- Nancarrow, D.J.; Mann, G.J.; Holland, E.A.; Walker, G.J.; Beaton, S.C.; Walters, M.K.; Luxford, C.; Palmer, J.M.; Donald, J.A.; Weber, J.L. Confirmation of Chromosome 9p Linkage in Familial Melanoma. Am. J. Hum. Genet. 1993, 53, 936–942. [Google Scholar]
- Kamb, A.; Shattuck Eidens, D.; Eeles, R.; Liu, Q.; Gruis, N.A.; Ding, W.; Hussey, C.; Tran, T.; Miki, Y.; Weaver Feldhaus, J. Analysis of the p16 Gene (CDKN2) as a Candidate for the Chromosome 9p Melanoma Susceptibility Locus. Nat. Genet. 1994, 8, 23–26. [Google Scholar] [CrossRef]
- Borg, A.; Johannsson, U.; Johannsson, O.; Hakansson, S.; Westerdahl, J.; Masback, A.; Olsson, H.; Ingvar, C. Novel Germline p16 Mutation in Familial Malignant Melanoma in Southern Sweden. Cancer Res. 1996, 56, 2497–2500. [Google Scholar] [PubMed]
- FitzGerald, M.G.; Harkin, D.P.; Silva Arrieta, S.; MacDonald, D.J.; Lucchina, L.C.; Unsal, H.; O’Neill, E.; Koh, J.; Finkelstein, D.M.; Isselbacher, K.J.; et al. Prevalence of Germ-Line Mutations in p16, p19ARF, and CDK4 in Familial Melanoma: Analysis of a Clinic-Based Population. Proc. Natl. Acad. Sci. USA 1996, 93, 8541–8545. [Google Scholar] [CrossRef]
- Harland, M.; Meloni, R.; Gruis, N.; Pinney, E.; Brookes, S.; Spurr, N.K.; Frischauf, A.M.; Bataille, V.; Peters, G.; Cuzick, J.; et al. Germline Mutations of the CDKN2 Gene in UK Melanoma Families. Hum. Mol. Genet. 1997, 6, 2061–2067. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Soufir, N.; Avril, M.F.; Chompret, A.; Demenais, F.; Bombled, J.; Spatz, A.; Stoppa Lyonnet, D.; Benard, J.; Bressac De Paillerets, B. Prevalence of p16 and CDK4 Germline Mutations in 48 Melanoma-Prone Families in France. The French Familial Melanoma Study Group. Hum. Mol. Genet. 1998, 7, 209–216. [Google Scholar] [CrossRef]
- Aitken, J.; Welch, J.; Duffy, D.; Milligan, A.; Green, A.; Martin, N.; Hayward, N. CDKN2A Variants in a Population-Based Sample of Queensland Families with Melanoma. J. Natl. Cancer Inst. 1999, 91, 446–452. [Google Scholar] [CrossRef]
- Quelle, D.E.; Zindy, F.; Ashmun, R.A.; Sherr, C.J. Alternative Reading Frames of the INK4a Tumor Suppressor Gene Encode two Unrelated Proteins Capable of Inducing Cell Cycle Arrest. Cell 1995, 83, 993–1000. [Google Scholar] [PubMed]
- Stott, F.J.; Bates, S.; James, M.C.; McConnell, B.B.; Starborg, M.; Brookes, S.; Palmero, I.; Ryan, K.; Hara, E.; Vousden, K.H.; et al. The Alternative Product from the Human CDKN2A Locus, p14(ARF), Participates in a Regulatory Feedback Loop with p53 and MDM2. EMBO J. 1998, 17, 5001–5014. [Google Scholar] [CrossRef]
- Goldstein, A.M.; Chan, M.; Harland, M.; Hayward, N.K.; Demenais, F.; Bishop, D.T.; Azizi, E.; Bergman, W.; Bianchi Scarra, G.; Bruno, W.; et al. Features Associated with Germline CDKN2A Mutations: A GenoMEL Study of Melanoma-Prone Families from Three Continents. J. Med. Genet. 2007, 44, 99–106. [Google Scholar] [CrossRef]
- Rossi, M.; Pellegrini, C.; Cardelli, L.; Ciciarelli, V.; Di Nardo, L.; Fargnoli, M.C. Familial Melanoma: Diagnostic and Management Implications. Dermatol. Pract. Concept. 2019, 9, 10–16. [Google Scholar] [CrossRef]
- Harland, M.; Cust, A.E.; Badenas, C.; Chang, Y.M.; Holland, E.A.; Aguilera, P.; Aitken, J.F.; Armstrong, B.K.; Barrett, J.H.; Carrera, C.; et al. Prevalence and Predictors of Germline CDKN2A Mutations for Melanoma Cases from Australia, Spain and the United Kingdom. Hered. Cancer Clin. Pract. 2014, 12, 20. [Google Scholar] [CrossRef]
- Wolfel, T.; Hauer, M.; Schneider, J.; Serrano, M.; Wolfel, C.; Klehmann Hieb, E.; De Plaen, E.; Hankeln, T.; Meyer Zum Buschenfelde, K.H.; Beach, D. A p16INK4a-Insensitive CDK4 Mutant Targeted by Cytolytic T Lymphocytes in a Human Melanoma. Science 1995, 269, 1281–1284. [Google Scholar] [CrossRef] [PubMed]
- Zuo, L.; Weger, J.; Yang, Q.; Goldstein, A.M.; Tucker, M.A.; Walker, G.J.; Hayward, N.; Dracopoli, N.C. Germline Mutations in the p16INK4a Binding Domain of cdk4 in Familial Melanoma. Nat. Genet. 1996, 12, 97–99. [Google Scholar] [CrossRef]
- Puntervoll, H.E.; Yang, X.R.; Vetti, H.H.; Bachmann, I.M.; Avril, M.F.; Benfodda, M.; Catricala, C.; Dalle, S.; Duval Modeste, A.B.; Ghiorzo, P.; et al. Melanoma Prone Families with CDK4 Germline Mutation: Phenotypic Profile and Associations with MC1R Variants. J. Med. Genet. 2013, 50, 264–270. [Google Scholar] [CrossRef]
- Harbour, J.W.; Onken, M.D.; Roberson, E.D.O.; Duan, S.; Cao, L.; Worley, L.A.; Council, M.L.; Matatall, K.A.; Helms, C.; Bowcock, A.M. Frequent Mutation of BAP1 in Metastasizing Uveal Melanomas. Science 2010, 330, 1410–1413. [Google Scholar] [CrossRef] [PubMed]
- Wiesner, T.; Obenauf, A.C.; Murali, R.; Fried, I.; Griewank, K.G.; Ulz, P.; Windpassinger, C.; Wackernagel, W.; Loy, S.; Wolf, I.; et al. Germline Mutations in BAP1 Predispose to Melanocytic Tumors. Nat. Genet. 2011, 43, 1018–1021. [Google Scholar] [CrossRef] [PubMed]
- Njauw, C.N.J.; Kim, I.; Piris, A.; Gabree, M.; Taylor, M.; Lane, A.M.; DeAngelis, M.M.; Gragoudas, E.; Duncan, L.M.; Tsao, H. Germline BAP1 Inactivation is Preferentially Associated with Metastatic Ocular Melanoma and Cutaneous-Ocular Melanoma Families. PLoS ONE 2012, 7, e35295. [Google Scholar] [CrossRef]
- Murali, R.; Wilmott, J.S.; Jakrot, V.; Al Ahmadie, H.A.; Wiesner, T.; McCarthy, S.W.; Thompson, J.F.; Scolyer, R.A. BAP1 Expression in Cutaneous Melanoma: A Pilot Study. Pathol. J. PCPA 2013, 45, 606–609. [Google Scholar] [CrossRef]
- Ismail, I.H.; Davidson, R.; Gagne, J.P.; Xu, Z.Z.; Poirier, G.G.; Hendzel, M.J. Germline Mutations in BAP1 Impair its Function in DNA Double-Strand Break Repair. Cancer Res. 2014, 74, 4282–4294. [Google Scholar] [CrossRef]
- Soura, E.; Eliades, P.J.; Shannon, K.; Stratigos, A.J.; Tsao, H. Hereditary Melanoma: Update on Syndromes and Management: Emerging Melanoma Cancer Complexes and Genetic Counseling. J. Am. Acad. Dermatol. 2016, 74, 411–420. [Google Scholar] [CrossRef]
- Horn, S.; Figl, A.; Rachakonda, P.S.; Fischer, C.; Sucker, A.; Gast, A.; Kadel, S.; Moll, I.; Nagore, E.; Hemminki, K.; et al. TERT Promoter Mutations in Familial and Sporadic Melanoma. Science 2013, 339, 959–961. [Google Scholar] [CrossRef]
- Robles Espinoza, C.D.; Harland, M.; Ramsay, A.J.; Aoude, L.G.; Quesada, V.; Ding, Z.; Pooley, K.A.; Pritchard, A.L.; Tiffen, J.C.; Petljak, M.; et al. POT1 Loss-of-Function Variants Predispose to Familial Melanoma. Nat. Genet. 2014, 46, 478–481. [Google Scholar] [CrossRef]
- Shi, J.; Yang, X.R.; Ballew, B.; Rotunno, M.; Calista, D.; Fargnoli, M.C.; Ghiorzo, P.; Bressac De Paillerets, B.; Nagore, E.; Avril, M.F.; et al. Rare Missense Variants in POT1 Predispose to Familial Cutaneous Malignant Melanoma. Nat. Genet. 2014, 46, 482–486. [Google Scholar] [CrossRef]
- Potrony, M.; Puig Butille, J.A.; Ribera Sola, M.; Iyer, V.; Robles Espinoza, C.D.; Aguilera, P.; Carrera, C.; Malvehy, J.; Badenas, C.; Landi, M.T.; et al. POT1 Germline Mutations but Not TERT Promoter Mutations are Implicated in Melanoma Susceptibility in a Large Cohort of Spanish Melanoma Families. Br. J. Dermatol. 2019, 181, 105–113. [Google Scholar] [CrossRef]
- Aoude, L.G.; Pritchard, A.L.; Robles Espinoza, C.D.; Wadt, K.; Harland, M.; Choi, J.; Gartside, M.; Quesada, V.; Johansson, P.; Palmer, J.M.; et al. Nonsense Mutations in the Shelterin Complex Genes ACD and TERF2IP in Familial Melanoma. J. Natl. Cancer Inst. 2015, 107, dju408. [Google Scholar] [CrossRef]
- Hodgkinson, C.A.; Moore, K.J.; Nakayama, A.; Steingrímsson, E.; Copeland, N.G.; Jenkins, N.A.; Arnheiter, H. Mutations at the mouse microphthalmia locus are associated with defects in a gene encoding a novel basic-helix-loop-helix-zipper protein. Cell 1993, 74, 395–404. [Google Scholar] [CrossRef]
- Yasumoto, K.; Yokoyama, K.; Shibata, K.; Tomita, Y.; Shibahara, S. Microphthalmia Associated Transcription Factor as a Regulator for Melanocyte-Specific Transcription of the Human Tyrosinase Gene. Mol. Cell. Biol. 1994, 14, 8058–8070. [Google Scholar] [CrossRef] [PubMed]
- Dynek, J.N.; Chan, S.M.; Liu, J.; Zha, J.; Fairbrother, W.J.; Vucic, D. Microphthalmia-Associated Transcription Factor is a Critical Transcriptional Regulator of Melanoma Inhibitor of Apoptosis in Melanomas. Cancer Res. 2008, 68, 3124–3132. [Google Scholar] [CrossRef]
- Garraway, L.A.; Widlund, H.R.; Rubin, M.A.; Getz, G.; Berger, A.J.; Ramaswamy, S.; Beroukhim, R.; Milner, D.A.; Granter, S.R.; Du, J.; et al. Integrative Genomic Analyses Identify MITF as a Lineage Survival Oncogene Amplified in Malignant Melanoma. Nature 2005, 436, 117–122. [Google Scholar] [CrossRef] [PubMed]
- Bertolotto, C.; Lesueur, F.; Giuliano, S.; Strub, T.; De Lichy, M.; Bille, K.; Dessen, P.; D’Hayer, B.; Mohamdi, H.; Remenieras, A.; et al. A SUMOylation-Defective MITF Germline Mutation Predisposes to Melanoma and Renal Carcinoma. Nature 2011, 480, 94–98. [Google Scholar] [CrossRef] [PubMed]
- Palmer, J.S.; Duffy, D.L.; Box, N.F.; Aitken, J.F.; O’Gorman, L.E.; Green, A.C.; Hayward, N.K.; Martin, N.G.; Sturm, R.A. Melanocortin-1 Receptor Polymorphisms and Risk of Melanoma: Is the Association Explained Solely by Pigmentation Phenotype? Am. J. Hum. Genet. 2000, 66, 176–186. [Google Scholar] [CrossRef]
- Kennedy, C.; Ter Huurne, J.; Berkhout, M.; Gruis, N.; Bastiaens, M.; Bergman, W.; Willemze, R.; Bavinck, J.N. Melanocortin 1 Receptor (MC1R) Gene Variants are Associated with an Increased Risk for Cutaneous Melanoma which is Largely Independent of Skin Type and Hair Color. J. Invest. Dermatol. 2001, 117, 294–300. [Google Scholar] [CrossRef] [PubMed]
- Box, N.F.; Duffy, D.L.; Chen, W.; Stark, M.; Martin, N.G.; Sturm, R.A.; Hayward, N.K. MC1R Genotype Modifies Risk of Melanoma in Families Segregating CDKN2A Mutations. Am. J. Hum. Genet. 2001, 69, 765–773. [Google Scholar] [CrossRef] [PubMed]
- Paolino, G.; Moliterni, E.; Corsetti, P.; Didona, D.; Bottoni, U.; Calvieri, S.; Mattozzi, C. Vitamin D and Melanoma: State of the Art and Possible Therapeutic Uses. G. Ital. Dermatol. E Venereol. Organo Uff. Soc. Ital. Dermatol. E Sifilogr. 2019, 154, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Slominski, A.T.; Brozyna, A.A.; Zmijewski, M.A.; Jozwicki, W.; Jetten, A.M.; Mason, R.S.; Tuckey, R.C.; Elmets, C.A. Vitamin D Signaling and Melanoma: Role of Vitamin D and its Receptors in Melanoma Progression and Management. Lab. Investig. J. Tech. Methods Pathol. 2017, 97, 706–724. [Google Scholar] [CrossRef]
- Hutchinson, P.E.; Osborne, J.E.; Lear, J.T.; Smith, A.G.; Bowers, P.W.; Morris, P.N.; Jones, P.W.; York, C.; Strange, R.C.; Fryer, A.A. Vitamin D Receptor Polymorphisms are Associated with Altered Prognosis in Patients with Malignant Melanoma. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2000, 6, 498–504. [Google Scholar]
- Shahbazi, M.; Pravica, V.; Nasreen, N.; Fakhoury, H.; Fryer, A.A.; Strange, R.C.; Hutchinson, P.E.; Osborne, J.E.; Lear, J.T.; Smith, A.G.; et al. Association between Functional Polymorphism in EGF Gene and Malignant Melanoma. Lancet Lond. Engl. 2002, 359, 397–401. [Google Scholar] [CrossRef]
- Hayward, N.K. Genetics of Melanoma Predisposition. Oncogene 2003, 22, 3053–3062. [Google Scholar] [CrossRef]
- Fargnoli, M.C.; Argenziano, G.; Zalaudek, I.; Peris, K. High-And Low-Penetrance Cutaneous Melanoma Susceptibility Genes. Expert Rev. Anticancer Ther. 2006, 6, 657–670. [Google Scholar] [CrossRef]
- Aoude, L.G.; Wadt, K.A.W.; Pritchard, A.L.; Hayward, N.K. Genetics of Familial Melanoma: 20 Years after CDKN2A. Pigment Cell Melanoma Res. 2015, 28, 148–160. [Google Scholar] [CrossRef]
- Bennett, D.C. Genetics of Melanoma Progression: The Rise and Fall of Cell Senescence. Pigment Cell Melanoma Res. 2016, 29, 122–140. [Google Scholar] [CrossRef]
- Roberts, M.R.; Asgari, M.M.; Toland, A.E. Genome-Wide Association Studies and Polygenic Risk Scores for Skin Cancer: Clinically Useful Yet? Br. J. Dermatol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Pho, L.N.; Leachman, S.A. Genetics of Pigmentation and Melanoma Predisposition. G. Ital. Dermatol. E Venereol. Organo Uff. Soc. Ital. Dermatol. E Sifilogr. 2010, 145, 37–45. [Google Scholar]
- Scherer, D.; Kumar, R. Genetics of Pigmentation in Skin Cancer-A Review. Mutat. Res. 2010, 705, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Ritterhouse, L.L.; Barletta, J.A. BRAF V600E Mutation-Specific Antibody: A Review. Semin. Diagn. Pathol. 2015, 32, 400–408. [Google Scholar] [CrossRef] [PubMed]
- Kunz, M. Oncogenes in Melanoma: An Update. Eur. J. Cell Biol. 2014, 93, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Reddy, B.Y.; Miller, D.M.; Tsao, H. Somatic Driver Mutations in Melanoma. Cancer 2017, 123, 2104–2117. [Google Scholar] [CrossRef]
- Cicenas, J.; Tamosaitis, L.; Kvederaviciute, K.; Tarvydas, R.; Staniute, G.; Kalyan, K.; Meskinyte Kausiliene, E.; Stankevicius, V.; Valius, M. KRAS, NRAS and BRAF Mutations in Colorectal Cancer and Melanoma. Med. Oncol. Northwood Lond. Engl. 2017, 34, 26. [Google Scholar] [CrossRef]
- Rabbie, R.; Ferguson, P.; Molina Aguilar, C.; Adams, D.J.; Robles Espinoza, C.D. Melanoma Subtypes: Genomic Profiles, Prognostic Molecular Markers and Therapeutic Possibilities. J. Pathol. 2019, 247, 539–551. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Network Genomic Classification of Cutaneous Melanoma. Cell 2015, 161, 1681–1696. [CrossRef]
- Hartman, R.I.; Lin, J.Y. Cutaneous Melanoma A Review in Detection, Staging, and Management. Hematol. Oncol. Clin. N. Am. 2019, 33, 25–38. [Google Scholar] [CrossRef]
- Everson, T.C. Spontaneous Regression of Cancer. Ann. N. Y. Acad. Sci. 1964, 114, 721–735. [Google Scholar] [CrossRef] [PubMed]
- Maio, M. Melanoma as a Model Tumour for Immuno-Oncology. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2012, 23, viii10–viii14. [Google Scholar] [CrossRef] [PubMed]
- Kalialis, L.V.; Drzewiecki, K.T.; Klyver, H. Spontaneous Regression of Metastases from Melanoma: Review of the Literature. Melanoma Res. 2009, 19, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Margaritescu, I.; Chiriţa, A.D.; Vasilescu, F. Completely Regressed Primary Cutaneous Melanoma-Difficulties in Diagnosis and Classification. Rom. J. Morphol. Embryol. Rev. Roum. Morphol. Embryol. 2014, 55, 635–642. [Google Scholar]
- Cole, W.H.; Everson, T.C. Spontaneous Regression of Cancer: Preliminary Report. Ann. Surg. 1956, 144, 366–383. [Google Scholar] [PubMed]
- Bourneuf, E. The MeLiM Minipig: An Original Spontaneous Model to Explore Cutaneous Melanoma Genetic Basis. Front. Genet. 2017, 8, 146. [Google Scholar] [CrossRef]
- Cervinkova, M.; Kucerova, P.; Cizkova, J. Spontaneous Regression of Malignant Melanoma-Is it Based on the Interplay between Host Immune System and Melanoma Antigens? Anticancer. Drugs 2017, 28, 819–830. [Google Scholar] [CrossRef]
- Ribero, S.; Moscarella, E.; Ferrara, G.; Piana, S.; Argenziano, G.; Longo, C. Regression in Cutaneous Melanoma: A Comprehensive Review from Diagnosis to Prognosis. J. Eur. Acad. Dermatol. Venereol. JEADV 2016, 30, 2030–2037. [Google Scholar] [CrossRef]
- Kaur, C.; Thomas, R.J.; Desai, N.; Green, M.A.; Lovell, D.; Powell, B.W.E.M.; Cook, M.G. The Correlation of Regression in Primary Melanoma with sentinel Lymph Node Status. J. Clin. Pathol. 2008, 61, 297–300. [Google Scholar] [CrossRef]
- Crompton, J.G.; Gilbert, E.; Brady, M.S. Clinical Implications of the Eighth Edition of the American Joint Committee on Cancer Melanoma Staging. J. Surg. Oncol. 2019, 119, 168–174. [Google Scholar] [CrossRef]
- Kozar, I.; Margue, C.; Rothengatter, S.; Haan, C.; Kreis, S. Many Ways to Resistance: How Melanoma Cells Evade Targeted Therapies. Biochim. Biophys. Acta Rev. Cancer 2019, 1871, 313–322. [Google Scholar] [CrossRef] [PubMed]
- Lorentzen, H.F. Targeted Therapy for Malignant Melanoma. Curr. Opin. Pharmacol. 2019, 46, 116–121. [Google Scholar] [CrossRef] [PubMed]
- Glitza Oliva, I.C.; Alqusairi, R. Immunotherapy for Melanoma. Adv. Exp. Med. Biol. 2018, 995, 43–63. [Google Scholar] [PubMed]
- Margolis, N.; Markovits, E.; Markel, G. Reprogramming Lymphocytes for the Treatment of Melanoma: From Biology to Therapy. Adv. Drug Deliv. Rev. 2019, 141, 104–124. [Google Scholar] [CrossRef] [PubMed]
- Weiss, S.A.; Wolchok, J.D.; Sznol, M. Immunotherapy of Melanoma: Facts and Hopes. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2019, 25, 5191–5201. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Liu, X.; Yang, J.; Zhang, M.; Jin, H.; Ma, X.; Shi, H. Combination of Immunotherapy With Targeted Therapy: Theory and Practice in Metastatic Melanoma. Front. Immunol. 2019, 10, 990. [Google Scholar] [CrossRef] [PubMed]
- Stern, H.M.; Zon, L.I. Cancer Genetics and Drug Discovery in the Zebrafish. Nat. Rev. Cancer 2003, 3, 533–539. [Google Scholar] [CrossRef]
- Schartl, M.; Walter, R.B. Xiphophorus and Medaka Cancer Models. Adv. Exp. Med. Biol. 2016, 916, 531–552. [Google Scholar]
- Bootorabi, F.; Manouchehri, H.; Changizi, R.; Barker, H.; Palazzo, E.; Saltari, A.; Parikka, M.; Pincelli, C.; Aspatwar, A. Zebrafish as a Model Organism for the Development of Drugs for Skin Cancer. Int. J. Mol. Sci. 2017, 18, 1550. [Google Scholar] [CrossRef]
- Cagan, R.L.; Zon, L.I.; White, R.M. Modeling Cancer with Flies and Fish. Dev. Cell 2019, 49, 317–324. [Google Scholar] [CrossRef]
- Bennett, D.; Lyulcheva, E.; Cobbe, N. Drosophila as a Potential Model for Ocular Tumors. Ocul. Oncol. Pathol. 2015, 1, 190–199. [Google Scholar] [CrossRef] [PubMed]
- Hanratty, W.P.; Ryerse, J.S. A Genetic Melanotic Neoplasm of Drosophila Melanogaster. Dev. Biol. 1981, 83, 238–249. [Google Scholar] [CrossRef]
- Anders, F. Contributions of the Gordon-Kosswig Melanoma System to the Present Concept of Neoplasia. Pigment Cell Res. 1991, 4, 7–29. [Google Scholar] [CrossRef] [PubMed]
- Setlow, R.B.; Woodhead, A.D.; Grist, E. Animal Model for Ultraviolet Radiation-Induced Melanoma: Platyfish-Swordtail Hybrid. Proc. Natl. Acad. Sci. USA 1989, 86, 8922–8926. [Google Scholar] [CrossRef]
- Wood, S.R.; Berwick, M.; Ley, R.D.; Walter, R.B.; Setlow, R.B.; Timmins, G.S. UV Causation of Melanoma in Xiphophorus is Dominated by Melanin Photosensitized Oxidant Production. Proc. Natl. Acad. Sci. USA 2006, 103, 4111–4115. [Google Scholar] [CrossRef]
- Wittbrodt, J.; Lammers, R.; Malitschek, B.; Ullrich, A.; Schartl, M. The Xmrk Receptor Tyrosine Kinase is Activated in Xiphophorus Malignant Melanoma. EMBO J. 1992, 11, 4239–4246. [Google Scholar] [CrossRef]
- Patton, E.E.; Widlund, H.R.; Kutok, J.L.; Kopani, K.R.; Amatruda, J.F.; Murphey, R.D.; Berghmans, S.; Mayhall, E.A.; Traver, D.; Fletcher, C.D.M.; et al. BRAF Mutations are Sufficient to Promote Nevi Formation and Cooperate with p53 in the Genesis of Melanoma. Curr. Biol. CB 2005, 15, 249–254. [Google Scholar] [CrossRef]
- Dovey, M.; White, R.M.; Zon, L.I. Oncogenic NRAS Cooperates with p53 Loss to Generate Melanoma in Zebrafish. Zebrafish 2009, 6, 397–404. [Google Scholar] [CrossRef]
- Santoriello, C.; Zon, L.I. Hooked! Modeling Human Disease in Zebrafish. J. Clin. Invest. 2012, 122, 2337–2343. [Google Scholar] [CrossRef]
- Scahill, C.M.; Digby, Z.; Sealy, I.M.; Wojciechowska, S.; White, R.J.; Collins, J.E.; Stemple, D.L.; Bartke, T.; Mathers, M.E.; Patton, E.E.; et al. Loss of the Chromatin Modifier Kdm2aa Causes BrafV600E-Independent Spontaneous Melanoma in Zebrafish. PLoS Genetics 2017, 13, e1006959. [Google Scholar] [CrossRef]
- Stoletov, K.; Klemke, R. Catch of the Day: Zebrafish as a Human Cancer Model. Oncogene 2008, 27, 4509–4520. [Google Scholar] [CrossRef] [PubMed]
- Ablain, J.; Zon, L.I. Of Fish and Men: Using Zebrafish to Fight Human Diseases. Trends Cell Biol. 2013, 23, 584–586. [Google Scholar] [CrossRef]
- Haldi, M.; Ton, C.; Seng, W.L.; McGrath, P. Human Melanoma Cells Transplanted into Zebrafish Proliferate, Migrate, Produce Melanin, form Masses and Stimulate Angiogenesis in Zebrafish. Angiogenesis 2006, 9, 139–151. [Google Scholar] [CrossRef] [PubMed]
- Heilmann, S.; Ratnakumar, K.; Langdon, E.; Kansler, E.; Kim, I.; Campbell, N.R.; Perry, E.; McMahon, A.; Kaufman, C.; Van Rooijen, E.; et al. A Quantitative System for Studying Metastasis Using Transparent Zebrafish. Cancer Res. 2015, 75, 4272–4282. [Google Scholar] [CrossRef] [PubMed]
- Schartl, M.; Wilde, B.; Laisney, J.A.G.C.; Taniguchi, Y.; Takeda, S.; Meierjohann, S. A Mutated EGFR is Sufficient to Induce Malignant Melanoma with Genetic Background-Dependent Histopathologies. J. Invest. Dermatol. 2010, 130, 249–258. [Google Scholar] [CrossRef] [PubMed]
- Schartl, M.; Kneitz, S.; Wilde, B.; Wagner, T.; Henkel, C.V.; Spaink, H.P.; Meierjohann, S. Conserved Expression Signatures between Medaka and Human Pigment Cell Tumors. PLoS ONE 2012, 7, e37880. [Google Scholar] [CrossRef]
- Levine, N.; Queen, L.; Chalom, A.A.; Daniels, L.J. Animal Model of Intracutaneous Melanoma. J. Invest. Dermatol. 1982, 78, 191–193. [Google Scholar] [CrossRef]
- Berkelhammer, J.; Oxenhandler, R.W. Evaluation of Premalignant and Malignant Lesions During the Induction of Mouse Melanomas. Cancer Res. 1987, 47, 1251–1254. [Google Scholar]
- Bradl, M.; Klein Szanto, A.; Porter, S.; Mintz, B. Malignant Melanoma in Transgenic Mice. Proc. Natl. Acad. Sci. USA 1991, 88, 164–168. [Google Scholar] [CrossRef]
- Mintz, B.; Silvers, W.K. Transgenic Mouse Model of Malignant Skin Melanoma. Proc. Natl. Acad. Sci. USA 1993, 90, 8817–8821. [Google Scholar] [CrossRef]
- Gattoni Celli, S.; Byers, R.H.; Calorini, L.; Ferrone, S. Organ-Specific Metastases in Melanoma: Experimental Animal Models. Pigment Cell Res. 1993, 6, 381–384. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhu, H.; Wetzel, W.J.; Philbert, M.A. Spontaneous Melanocytosis in Transgenic Mice. J. Invest. Dermatol. 1996, 106, 1145–1151. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Reuhl, K.; Zhang, X.; Botha, R.; Ryan, K.; Wei, J.; Chen, S. Development of Heritable Melanoma in Transgenic Mice. J. Invest. Dermatol. 1998, 110, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Bobek, V.; Kolostova, K.; Pinterova, D.; Kacprzak, G.; Adamiak, J.; Kolodziej, J.; Boubelik, M.; Kubecova, M.; Hoffman, R.M. A Clinically Relevant, Syngeneic Model of Spontaneous, Highly Metastatic B16 Mouse Melanoma. Anticancer Res. 2010, 5, 4799–4803. [Google Scholar]
- Saleh, J. Murine Models of Melanoma. Pathol. Res. Pract. 2018, 214, 1235–1238. [Google Scholar] [CrossRef]
- Kuzu, O.F.; Nguyen, F.D.; Noory, M.A.; Sharma, A. Current State of Animal (Mouse) Modeling in Melanoma Research. Cancer Growth Metastasis 2015, 8, 81–94. [Google Scholar] [CrossRef]
- Harris, A.L.; Joseph, R.W.; Copland, J.A. Patient-Derived Tumor Xenograft Models for Melanoma Drug Discovery. Expert Opin. Drug Discov. 2016, 11, 895–906. [Google Scholar] [CrossRef]
- Choi, Y.; Lee, S.; Kim, K.; Kim, S.H.; Chung, Y.J.; Lee, C. Studying Cancer Immunotherapy Using Patient-Derived Xenografts (PDXs) in Humanized Mice. Exp. Mol. Med. 2018, 50, 1–9. [Google Scholar] [CrossRef]
- McKinney, A.J.; Holmen, S.L. Animal Models of Melanoma: A Somatic Cell Gene Delivery Mouse Model Allows Rapid Evaluation of Genes Implicated in Human Melanoma. Chin. J. Cancer 2011, 30, 153–162. [Google Scholar] [CrossRef]
- Perez Guijarro, E.; Day, C.P.; Merlino, G.; Zaidi, M.R. Genetically Engineered Mouse Models of Melanoma. Cancer 2017, 123, 2089–2103. [Google Scholar] [CrossRef]
- Niu, Y.; Liang, S. Mammalian Models Based on RCAS-TVA Technique. Zool. Res. 2008, 29, 335–345. [Google Scholar]
- Dankort, D.; Curley, D.P.; Cartlidge, R.A.; Nelson, B.; Karnezis, A.N.; Damsky, W.E.; You, M.J.; DePinho, R.A.; McMahon, M.; Bosenberg, M. Braf(V600E) Cooperates with Pten Loss to Induce Metastatic Melanoma. Nat. Genet. 2009, 41, 544–552. [Google Scholar] [CrossRef] [PubMed]
- Klein Szanto, A.J.; Silvers, W.K.; Mintz, B. Ultraviolet Radiation-Induced Malignant Skin Melanoma in Melanoma-Susceptible Transgenic Mice. Cancer Res. 1994, 54, 4569–4572. [Google Scholar] [PubMed]
- Broome Powell, M.; Gause, P.R.; Hyman, P.; Gregus, J.; Lluria Prevatt, M.; Nagle, R.; Bowden, G.T. Induction of Melanoma in TPras Transgenic Mice. Carcinogenesis 1999, 20, 1747–1753. [Google Scholar] [CrossRef]
- Ley, R.D. Animal Models of Ultraviolet Radiation (UVR)-Induced Cutaneous Melanoma. Front. Biosci. J. Virtual Libr. 2002, 7, d1531–d1534. [Google Scholar]
- Larue, L. Origin of Mouse Melanomas. J. Invest. Dermatol. 2012, 132, 2135–2136. [Google Scholar] [CrossRef]
- Goldschmidt, M.H. Pigmented Lesions of the Skin. Clin. Dermatol. 1994, 12, 507–514. [Google Scholar] [CrossRef]
- Modiano, J.F.; Ritt, M.G.; Wojcieszyn, J. The Molecular Basis of Canine Melanoma: Pathogenesis and Trends in Diagnosis and Therapy. J. Vet. Intern. Med. 1999, 13, 163–174. [Google Scholar] [CrossRef]
- Prouteau, A.; Andre, C. Canine Melanomas as Models for Human Melanomas: Clinical, Histological, and Genetic Comparison. Genes 2019, 10, 501. [Google Scholar] [CrossRef]
- Koenig, A.; Bianco, S.R.; Fosmire, S.; Wojcieszyn, J.; Modiano, J.F. Expression and Significance of p53, rb, p21/waf-1, p16/ink-4a, and PTEN Tumor Suppressors in Canine Melanoma. Vet. Pathol. 2002, 39, 458–472. [Google Scholar] [CrossRef]
- Gillard, M.; Cadieu, E.; De Brito, C.; Abadie, J.; Vergier, B.; Devauchelle, P.; Degorce, F.; Dreano, S.; Primot, A.; Dorso, L.; et al. Naturally Occurring Melanomas in Dogs as Models for Non-UV Pathways of Human Melanomas. Pigment Cell Melanoma Res. 2014, 27, 90–102. [Google Scholar] [CrossRef] [PubMed]
- Pisamai, S.; Rungsipipat, A.; Kalpravidh, C.; Suriyaphol, G. Gene Expression Profiles of Cell Adhesion Molecules, Matrix Metalloproteinases and Their Tissue Inhibitors in Canine Oral Tumors. Res. Vet. Sci. 2017, 113, 94–100. [Google Scholar] [CrossRef] [PubMed]
- Brachelente, C.; Cappelli, K.; Capomaccio, S.; Porcellato, I.; Silvestri, S.; Bongiovanni, L.; De Maria, R.; Verini Supplizi, A.; Mechelli, L.; Sforna, M. Transcriptome Analysis of Canine Cutaneous Melanoma and Melanocytoma Reveals a Modulation of Genes Regulating Extracellular Matrix Metabolism and Cell Cycle. Sci. Rep. 2017, 7, 6386. [Google Scholar] [CrossRef] [PubMed]
- Hendricks, W.P.D.; Zismann, V.; Sivaprakasam, K.; Legendre, C.; Poorman, K.; Tembe, W.; Perdigones, N.; Kiefer, J.; Liang, W.; DeLuca, V.; et al. Somatic Inactivating PTPRJ Mutations and Dysregulated Pathways Identified in Canine Malignant Melanoma by Integrated Comparative Genomic Analysis. PLoS Genetics 2018, 14. [Google Scholar] [CrossRef] [PubMed]
- Valentine, B.A. Equine Melanocytic Tumors: A Retrospective Study of 53 Horses (1988 to 1991). J. Vet. Intern. Med. 1995, 9, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Fleury, C.; Berard, F.; Balme, B.; Thomas, L. The Study of Cutaneous Melanomas in Camargue-Type Gray-Skinned Horses (1): Clinical-Pathological Characterization. Pigment Cell Res. 2000, 13, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Vostry, L.; Hofmanova, B.; Vydrova, H.; Pribyl, J.; Majzlik, I. Estimation of Genetic Parameters for Melanoma in the Old Kladruber Horse. Czech J. Anim. Sci. 2012, 57, 75–82. [Google Scholar] [CrossRef]
- Curik, I.; Druml, T.; Seltenhammer, M.; Sundstrom, E.; Pielberg, G.R.; Andersson, L.; Solkner, J. Complex Inheritance of Melanoma and Pigmentation of Coat and Skin in Grey Horses. PLoS Genetics 2013, 9, e1003248. [Google Scholar] [CrossRef]
- Campagne, C.; Jule, S.; Bernex, F.; Estrada, M.; Aubin Houzelstein, G.; Panthier, J.J.; Egidy, G. RACK1, a Clue to the Diagnosis of Cutaneous Melanomas in Horses. BMC Vet. Res. 2012, 8, 95. [Google Scholar] [CrossRef]
- Thirloway, L.; Rudolph, R.; Leipold, H.W. Malignant Melanomas in a Duroc Boar. J. Am. Vet. Med. Assoc. 1977, 170, 345–347. [Google Scholar]
- Fisher, L.F.; Olander, H.J. Spontaneous Neoplasms of Pigs-A Study of 31 Cases. J. Comp. Pathol. 1978, 88, 505–517. [Google Scholar] [CrossRef]
- Baba, A.I.; Gaboreanu, M.; Rotaru, O.; Kwieczinsky, R. Malignant Melanomas in Farm Animals. Morphol. Embryol. (Bucur.) 1983, 29, 191–194. [Google Scholar] [PubMed]
- Perez, J.; Garcia, P.M.; Bautista, M.J.; Millan, Y.; Ordas, J.; De Las Mulas, J.M. Immunohistochemical Characterization of Tumor Cells and Inflammatory Infiltrate Associated with Cutaneous Melanocytic Tumors of Duroc and Iberian Swine. Vet. Pathol. 2002, 39, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Bundza, A.; Feltmate, T.E. Melanocytic Cutaneous Lesions and Melanotic Regional Lymph Nodes in Slaughter Swine. Can. J. Vet. Res. Rev. Can. Rech. Vet. 1990, 54, 301–304. [Google Scholar]
- Hordinsky, M.K.; Ruth, G.; King, R. Inheritance of Melanocytic Tumors in Duroc Swine. J. Hered. 1985, 76, 385–386. [Google Scholar]
- Strafuss, A.C.; Dommert, A.R.; Tumbleson, M.E.; Middleton, C.C. Cutaneous Melanoma in Miniature Swine. Lab. Anim. Care 1968, 18, 165–169. [Google Scholar]
- Millikan, L.E.; Boylon, J.L.; Hook, R.R.; Manning, P.J. Melanoma in Sinclair Swine: A New Animal Model. J. Invest. Dermatol. 1974, 62, 20–30. [Google Scholar] [CrossRef]
- Hook, R.R.; Aultman, M.D.; Adelstein, E.H.; Oxenhandler, R.W.; Millikan, L.E.; Middleton, C.C. Influence of Selective Breeding on the Incidence of Melanomas in Sinclair Miniature Swine. Int. J. Cancer 1979, 24, 668–672. [Google Scholar] [CrossRef]
- Manning, P.J.; Millikan, L.E.; Cox, V.S.; Carey, K.D.; Hook, R.R. Congenital Cutaneous and Visceral Melanomas of Sinclair Miniature Swine: Three Case Reports. J. Natl. Cancer Inst. 1974, 52, 1559–1566. [Google Scholar] [CrossRef]
- Oxenhandler, R.W.; Adelstein, E.H.; Haigh, J.P.; Hook, R.R.; Clark, W.H. Malignant Melanoma in the Sinclair Miniature Swine: An Autopsy Study of 60 Cases. Am. J. Pathol. 1979, 96, 707–720. [Google Scholar]
- Hook, R.R.; Berkelhammer, J.; Oxenhandler, R.W. Melanoma: Sinclair Swine Melanoma. Am. J. Pathol. 1982, 108, 130–133. [Google Scholar] [PubMed]
- Misfeldt, M.L.; Grimm, D.R. Sinclair Miniature Swine: An Animal Model of Human Melanoma. Vet. Immunol. Immunopathol. 1994, 43, 167–175. [Google Scholar] [CrossRef]
- Greene, J.F.; Townsend, J.S.; Amoss, M.S. Histopathology of Regression in Sinclair Swine Model of Melanoma. Lab. Investig. J. Tech. Methods Pathol. 1994, 71, 17–24. [Google Scholar]
- Morgan, C.D.; Measel, J.W.; Amoss, M.S.; Rao, A.; Greene, J.F. Immunophenotypic Characterization of Tumor Infiltrating Lymphocytes and Peripheral Blood Lymphocytes Isolated from Melanomatous and Non-Melanomatous Sinclair Miniature Swine. Vet. Immunol. Immunopathol. 1996, 55, 189–203. [Google Scholar] [CrossRef]
- Cui, J.; Chen, D.; Misfeldt, M.L.; Swinfard, R.W.; Bystryn, J.C. Antimelanoma Antibodies in Swine with Spontaneously Regressing Melanoma. Pigment Cell Res. 1995, 8, 60–63. [Google Scholar] [CrossRef]
- Gossett, R.; Kier, A.B.; Schroeder, F.; McConkey, D.; Fadok, V.; Amoss, M.S. Cycloheximide-Induced Apoptosis in Melanoma Cells Derived from Regressing Cutaneous Tumours of SINCLAIR Swine. J. Comp. Pathol. 1996, 115, 353–372. [Google Scholar] [CrossRef]
- Pathak, S.; Multani, A.S.; McConkey, D.J.; Imam, A.S.; Amoss, M.S. Spontaneous Regression of Cutaneous Melanoma in Sinclair Swine is Associated with Defective Telomerase Activity and Extensive Telomere Erosion. Int. J. Oncol. 2000, 17, 1219–1243. [Google Scholar] [CrossRef]
- Tissot, R.G.; Beattie, C.W.; Amoss, M.S. The Swine Leucocyte Antigen (SLA) Complex and Sinclair Swine Cutaneous Malignant Melanoma. Anim. Genet. 1989, 20, 51–57. [Google Scholar] [CrossRef]
- Tissot, R.G.; Beattie, C.W.; Amoss, M.S. Inheritance of Sinclair Swine Cutaneous Malignant Melanoma. Cancer Res. 1987, 47, 5542–5545. [Google Scholar]
- Blangero, J.; Tissot, R.G.; Beattie, C.W.; Amoss, M.S. Genetic Determinants of Cutaneous Malignant Melanoma in Sinclair Swine. Br. J. Cancer 1996, 73, 667–671. [Google Scholar] [CrossRef][Green Version]
- Pathak, S.; Amoss, M.S. Genetic Predisposition and Specific Chromosomal Defects Associated with Sinclair Swine Malignant Melanomas. Int. J. Oncol. 1997, 11, 53–57. [Google Scholar] [CrossRef] [PubMed]
- Muller, S.; Wanke, R.; Distl, O. Inheritance of Melanocytic Lesions and Their Association with the White Colour Phenotype in Miniature Swine. J. Anim. Breed. Genet. 2001, 118, 275–283. [Google Scholar] [CrossRef]
- Muller, S.; Wanke, R.; Distl, O. Segregation of Pigment Cell Anomalies in Munich Miniature Swine (MMS) Troll Crossed with German Landrace. DTW Dtsch. Tierarztl. Wochenschr. 1995, 102, 391–394. [Google Scholar] [PubMed]
- Buttner, M.; Wanke, R.; Obermann, B. Natural Killer (NK) Activity of Porcine Blood Lymphocytes Against Allogeneic Melanoma Target Cells. Vet. Immunol. Immunopathol. 1991, 29, 89–103. [Google Scholar] [CrossRef]
- Dieckhoff, B.; Puhlmann, J.; Buscher, K.; Hafner Marx, A.; Herbach, N.; Bannert, N.; Buttner, M.; Wanke, R.; Kurth, R.; Denner, J. Expression of Porcine Endogenous Retroviruses (PERVs) in Melanomas of Munich Miniature Swine (MMS) Troll. Vet. Microbiol. 2007, 123, 53–68. [Google Scholar] [CrossRef]
- Buscher, K.; Trefzer, U.; Hofmann, M.; Sterry, W.; Kurth, R.; Denner, J. Expression of Human Endogenous Retrovirus K in Melanomas and Melanoma Cell Lines. Cancer Res. 2005, 65, 4172–4180. [Google Scholar] [CrossRef]
- Gonzalez Cao, M.; Iduma, P.; Karachaliou, N.; Santarpia, M.; Blanco, J.; Rosell, R. Human Endogenous Retroviruses and Cancer. Cancer Biol. Med. 2016, 13, 483–488. [Google Scholar]
- Glodek, P. Breeding Program and Population Standards of the Goettingen Miniature Swine. In Swine in Biomedical Research; Tumbleson, M.E., Ed.; Plenum Press: New York, NY, USA, 1986; Volume 1, pp. 23–28. [Google Scholar]
- Horak, V.; Fortyn, K.; Hruban, V.; Klaudy, J. Hereditary Melanoblastoma in Miniature Pigs and its Successful Therapy by Devitalization Technique. Cell. Mol. Biol. Noisy Gd. Fr. 1999, 45, 1119–1129. [Google Scholar]
- Hruban, V.; Horak, V.; Fortyn, K.; Hradecky, J.; Klaudy, J.; Smith, D.M.; Reisnerova, H.; Majzlik, I. Inheritance of Malignant Melanoma in the MeLiM Strain of Miniature Pigs. Vet. Med. (Praha) 2004, 49, 453–459. [Google Scholar] [CrossRef]
- Vincent Naulleau, S.; Le Chalony, C.; Leplat, J.J.; Bouet, S.; Bailly, C.; Spatz, A.; Vielh, P.; Avril, M.F.; Tricaud, Y.; Gruand, J.; et al. Clinical and Histopathological Characterization of Cutaneous Melanomas in the Melanoblastoma-Bearing Libechov Minipig Model. Pigment Cell Melanoma Res. 2004, 17, 24–35. [Google Scholar] [CrossRef]
- Fortyn, K.; Hruban, V.; Horak, V. Treatment of Malignant Melanoma. Br. J. Surg. 1994, 81, 146–147. [Google Scholar] [CrossRef] [PubMed]
- Fortyn, K.; Hruban, V.; Horak, V.; Tichy, J. Exceptional Occurrence and Extent of Malignant Melanoma in Pig. Vet. Med. (Praha) 1998, 43, 87–91. [Google Scholar]
- Fortyn, K.; Hruban, V.; Horak, V.; Hradecky, J.; Tichy, J. Melanoblastoma in Laboratory Minipigs: A Model for Studying Human Malignant Melan6oma. Vet. Med. (Praha) 1994, 39, 597–604. [Google Scholar]
- Al Shaer, M.; Gollapudi, D.; Papageorgio, C. Melanoma Biomarkers: Vox Clamantis in Deserto (Review). Oncol. Lett. 2010, 1, 399–405. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pohlreich, P.; Stribrna, J.; Kleibl, Z.; Horak, V.; Klaudy, J. Detection of Neoplastic Cells in Blood of Miniature Pigs with Hereditary Melanoma. Vet. Med. (Praha) 2001, 46, 199–204. [Google Scholar] [CrossRef]
- De Souza, L.M.; Robertson, B.M.; Robertson, G.P. Future of Circulating Tumor Cells in the Melanoma Clinical and Research Laboratory Settings. Cancer Lett. 2017, 392, 60–70. [Google Scholar] [CrossRef]
- Rodic, S.; Mihalcioiu, C.; Saleh, R.R. Detection Methods of Circulating Tumor Cells in Cutaneous Melanoma: A Systematic Review. Crit. Rev. Oncol. Hematol. 2014, 91, 74–92. [Google Scholar] [CrossRef]
- Borovansky, J.; Horak, V.; Elleder, M.; Fortyn, K.; Smit, N.P.; Kolb, A.M. Biochemical Characterization of a New Melanoma Model-The Minipig MeLiM Strain. Melanoma Res. 2003, 13, 543–548. [Google Scholar] [CrossRef]
- Millikan, L.E.; Hook, R.R.; Manning, P.J. Immunobiology of Melanoma. Gross and Ultrastructural Studies in a New Melanoma Model: The Sinclair Swine. Yale J. Biol. Med. 1973, 46, 631–645. [Google Scholar]
- Hunter, J.A.; Zaynoun, S.; Paterson, W.D.; Bleehen, S.S.; Mackie, R.; Cochran, A.J. Cellular Fine Structure in the Invasive Nodules of Different Histogenetic Types of Malignant Melanoma. Br. J. Dermatol. 1978, 98, 255–272. [Google Scholar] [CrossRef]
- Borovansky, J. Quantitative Parameters of Melanomas Differentiation. Neoplasma 1978, 25, 349–352. [Google Scholar] [PubMed]
- Ochi, Y.; Atsumi, S.; Aoyagi, T.; Umezawa, K. Inhibition of Tumor Cell Invasion in the Boyden Chamber Assay by a Mannosidase Inhibitor, Mannostatin A. Anticancer Res. 1993, 13, 1421–1424. [Google Scholar] [PubMed]
- Borovansky, J.; Hach, P. Disparate behaviour of Two Melanosomal Enzymes (α-Mannosidase and γ-Glutamyltransferase). Cell. Mol. Biol. Noisy Gd. Fr. 1999, 45, 1047–1052. [Google Scholar]
- Egidy, G.; Jule, S.; Bosse, P.; Bernex, F.; Geffrotin, C.; Vincent Naulleau, S.; Horak, V.; Sastre Garau, X.; Panthier, J.J. Transcription Analysis in the MeLiM Swine Model Identifies RACK1 as a Potential Marker of Malignancy for Human Melanocytic Proliferation. Mol. Cancer 2008, 7, 34. [Google Scholar] [CrossRef]
- Xia, J.; Wang, Y.; Li, F.; Wang, J.; Mu, Y.; Mei, X.; Li, X.; Zhu, W.; Jin, X.; Yu, K. Expression of Microphthalmia Transcription Factor, S100 Protein, and HMB-45 in Malignant Melanoma and Pigmented Nevi. Biomed. Rep. 2016, 5, 327–331. [Google Scholar] [CrossRef]
- Planska, D.; Burocziova, M.; Strnadel, J.; Horak, V. Immunohistochemical Analysis of Collagen IV and Laminin Expression in Spontaneous Melanoma Regression in the Melanoma-Bearing Libechov Minipig. Acta Histochem. Cytochem. 2015, 48, 15–26. [Google Scholar] [CrossRef][Green Version]
- Geffrotin, C.; Horak, V.; Crechet, F.; Tricaud, Y.; Lethias, C.; Vincent Naulleau, S.; Vielh, P. Opposite Regulation of Tenascin-C and Tenascin-X in MeLiM Swine Heritable Cutaneous Malignant Melanoma. Biochim. Biophys. Acta BBA Gen. Subj. 2000, 1524, 196–202. [Google Scholar] [CrossRef]
- Planska, D.; Kovalska, J.; Cizkova, J.; Horak, V. Tissue Rebuilding During Spontaneous Regression of Melanoma in the Melanoma-bearing Libechov Minipig. Anticancer Res. 2018, 38, 4629–4636. [Google Scholar] [CrossRef]
- Engbring, J.A.; Kleinman, H.K. The Basement Membrane Matrix in Malignancy. J. Pathol. 2003, 200, 465–470. [Google Scholar] [CrossRef]
- Pasco, S.; Ramont, L.; Maquart, F.X.; Monboisse, J.C. Control of Melanoma Progression by Various Matrikines from Basement Membrane Macromolecules. Crit. Rev. Oncol. Hematol. 2004, 49, 221–233. [Google Scholar] [CrossRef]
- Hofmann, U.B.; Houben, R.; Brocker, E.B.; Becker, J.C. Role of Matrix Metalloproteinases in Melanoma Cell Invasion. Biochimie 2005, 87, 307–314. [Google Scholar] [CrossRef] [PubMed]
- Kaariainen, E.; Nummela, P.; Soikkeli, J.; Yin, M.; Lukk, M.; Jahkola, T.; Virolainen, S.; Ora, A.; Ukkonen, E.; Saksela, O.; et al. Switch to an Invasive Growth Phase in Melanoma is Associated with Tenascin-C, Fibronectin, and Procollagen-I Forming Specific Channel Structures for Invasion. J. Pathol. 2006, 210, 181–191. [Google Scholar] [CrossRef]
- Shao, H.; Kirkwood, J.M.; Wells, A. Tenascin-C Signaling in Melanoma. Cell Adhes. Migr. 2014, 9, 125–130. [Google Scholar] [CrossRef]
- Anyz, J.; Vyslouzilova, L.; Vaculovic, T.; Tvrdonova, M.; Kanicky, V.; Haase, H.; Horak, V.; Stepankova, O.; Heger, Z.; Adam, V. Spatial Mapping of Metals in Tissue-Sections Using Combination of Mass-Spectrometry and Histology Through Image Registration. Sci. Rep. 2017, 7, 40169. [Google Scholar] [CrossRef] [PubMed]
- Gorodetsky, R.; Sheskin, J.; Weinreb, A. Iron, Copper, and Zinc Concentrations in Normal Skin and in Various Nonmalignant and Malignant Lesions. Int. J. Dermatol. 1986, 25, 440–445. [Google Scholar] [CrossRef]
- Guran, R.; Vanickova, L.; Horak, V.; Krizkova, S.; Michalek, P.; Heger, Z.; Zitka, O.; Adam, V. MALDI MSI of MeLiM Melanoma: Searching for Differences in Protein Profiles. PLoS ONE 2017, 12, e0189305. [Google Scholar] [CrossRef]
- Krizkova, S.; Fabrik, I.; Adam, V.; Kukacka, J.; Prusa, R.; Chavis, G.J.; Trnkova, L.; Strnadel, J.; Horak, V.; Kizek, R. Utilizing of Adsorptive Transfer Stripping Technique Brdicka Reaction for Determination of Metallothioneins Level in Melanoma Cells, Blood Serum and Tissues. Sensors 2008, 8, 3106–3122. [Google Scholar] [CrossRef]
- Weinlich, G. Metallothionein-Overexpression as a Prognostic Marker in Melanoma. G. Ital. Dermatol. E Venereol. Organo Uff. Soc. Ital. Dermatol. E Sifilogr. 2009, 144, 27–38. [Google Scholar]
- Cizkova, J.; Erbanova, M.; Sochor, J.; Jindrova, A.; Strnadova, K.; Horak, V. Relationship Between Haematological Profile and Progression or Spontaneous Regression of Melanoma in the Melanoma-Bearing Libechov Minipigs. Vet. J. Lond. Engl. 1997 2019, 249, 1–9. [Google Scholar]
- Blanc, F.; Crechet, F.; Bruneau, N.; Piton, G.; Leplat, J.J.; Andreoletti, F.; Egidy, G.; Vincent Naulleau, S.; Bourneuf, E. Impact of a CD4 Gene Haplotype on the Immune Response in Minipigs. Immunogenetics 2018, 70, 209–222. [Google Scholar] [CrossRef]
- Du, Z.Q.; Vincent Naulleau, S.; Gilbert, H.; Vignoles, F.; Crechet, F.; Shimogiri, T.; Yasue, H.; Leplat, J.J.; Bouet, S.; Gruand, J.; et al. Detection of Novel Quantitative Trait Loci for Cutaneous Melanoma by Genome-Wide Scan in the MeLiM Swine Model. Int. J. Cancer 2007, 120, 303–320. [Google Scholar] [PubMed]
- Le Chalony, C.; Renard, C.; Vincent Naulleau, S.; Crechet, F.; Leplat, J.J.; Tricaud, Y.; Horak, V.; Gruand, J.; Le Roy, P.; Frelat, G.; et al. CDKN2A Region Polymorphism and Genetic Susceptibility to Melanoma in the Melim Swine Model of Familial Melanoma. Int. J. Cancer 2003, 103, 631–635. [Google Scholar] [CrossRef] [PubMed]
- Geffrotin, C.; Crechet, F.; Le Roy, P.; Le Chalony, C.; Leplat, J.J.; Iannuccelli, N.; Barbosa, A.; Renard, C.; Gruand, J.; Milan, D.; et al. Identification of Five Chromosomal Regions Involved in Predisposition to Melanoma by Genome-Wide Scan in the MeLiM Swine Model. Int. J. Cancer 2004, 110, 39–50. [Google Scholar] [CrossRef] [PubMed]
- Bourneuf, E.; Du, Z.Q.; Estelle, J.; Gilbert, H.; Crechet, F.; Piton, G.; Milan, D.; Geffrotin, C.; Lathrop, M.; Demenais, F.; et al. Genetic and Functional Evaluation of MITF as a Candidate Gene for Cutaneous Melanoma Predisposition in Pigs. Mamm. Genome Off. J. Int. Mamm. Genome Soc. 2011, 22, 602–612. [Google Scholar] [CrossRef]
- Fernandez Rodriguez, A.; Estelle, J.; Blin, A.; Munoz, M.; Crechet, F.; Demenais, F.; Vincent Naulleau, S.; Bourneuf, E. KIT and Melanoma Predisposition in Pigs: Sequence Variants and Association Analysis. Anim. Genet. 2014, 45, 445–448. [Google Scholar] [CrossRef]
- Rambow, F.; Piton, G.; Bouet, S.; Leplat, J.J.; Baulande, S.; Marrau, A.; Stam, M.; Horak, V.; Vincent Naulleau, S. Gene Expression Signature for Spontaneous Cancer Regression in Melanoma Pigs. Neoplasia 2008, 10, 714. [Google Scholar] [CrossRef]
- Rambow, F.; Malek, O.; Geffrotin, C.; Leplat, J.J.; Bouet, S.; Piton, G.; Hugot, K.; Bevilacqua, C.; Horak, V.; Vincent Naulleau, S. Identification of Differentially Expressed Genes in Spontaneously Regressing Melanoma Using the MeLiM Swine Model. Pigment Cell Melanoma Res. 2008, 21, 147–161. [Google Scholar] [CrossRef]
- Bourneuf, E.; Estelle, J.; Blin, A.; Crechet, F.; del Pilar Schneider, M.; Gilbert, H.; Brossard, M.; Vaysse, A.; Lathrop, M.; Vincent Naulleau, S.; et al. New Susceptibility Loci for Cutaneous Melanoma Risk and Progression Revealed Using a Porcine Model. Oncotarget 2018, 9, 27682–27697. [Google Scholar] [CrossRef][Green Version]
- Sun, X.; Gao, L.; Chien, H.Y.; Li, W.C.; Zhao, J. The Regulation and Function of the NUAK Family. J. Mol. Endocrinol. 2013, 51, R15–R22. [Google Scholar] [CrossRef]
- Xu, X.; Hussain, W.M.; Vijai, J.; Offit, K.; Rubin, M.A.; Demichelis, F.; Klein, R.J. Variants at IRX4 as Prostate Cancer Expression Quantitative Trait Loci. Eur. J. Hum. Genet. EJHG 2014, 22, 558–563. [Google Scholar] [CrossRef]
- Sanchez Tena, S.; Cubillos Rojas, M.; Schneider, T.; Rosa, J.L. Functional and Pathological Relevance of HERC Family Proteins: A Decade Later. Cell. Mol. Life Sci. 2016, 73, 1955–1968. [Google Scholar] [CrossRef] [PubMed]
- Fattore, L.; Costantini, S.; Malpicci, D.; Ruggiero, C.F.; Ascierto, P.A.; Croce, C.M.; Mancini, R.; Ciliberto, G. MicroRNAs in Melanoma Development and Resistance to Target Therapy. Oncotarget 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Baco, M.; Chu, C.Y.; Bouet, S.; Rogel Gaillard, C.; Bourneuf, E.; Le Provost, F.; Chu, C.Y.; Vincent Naulleau, S. Analysis of Melanoma-Related microRNAs Expression During the Spontaneous Regression of Cutaneous Melanomas in MeLiM Pigs. Pigment Cell Melanoma Res. 2014, 27, 668–670. [Google Scholar] [CrossRef] [PubMed]
- So, K.A.; Hong, J.H.; Jin, H.M.; Kim, J.W.; Song, J.Y.; Lee, J.K.; Lee, N.W. The Prognostic Significance of Preoperative Leukocytosis in Epithelial Ovarian Carcinoma: A Retrospective Cohort Study. Gynecol. Oncol. 2014, 132, 551–555. [Google Scholar] [CrossRef] [PubMed]
- Azab, B.; Mohammad, F.; Shah, N.; Vonfrolio, S.; Lu, W.; Kedia, S.; Bloom, S.W. The Value of the Pretreatment Neutrophil Lymphocyte Ratio vs. Platelet Lymphocyte Ratio in Predicting the Long-Term Survival in Colorectal Cancer. Cancer Biomark. Sect. Dis. Markers 2014, 14, 303–312. [Google Scholar] [CrossRef]
- Zhang, H.; Xia, H.; Zhang, L.; Zhang, B.; Yue, D.; Wang, C. Clinical Significance of Preoperative Neutrophil-Lymphocyte vs Platelet-Lymphocyte Ratio in Primary Operable Patients with Non-Small Cell Lung Cancer. Am. J. Surg. 2015, 210, 526–535. [Google Scholar] [CrossRef]
- Feng, L.; Gu, S.; Wang, P.; Chen, H.; Chen, Z.; Meng, Z.; Liu, L. White Blood Cell and Granulocyte Counts Are Independent Predictive Factors for Prognosis of Advanced Pancreatic Caner. Gastroenterol. Res. Pract. 2018, 2018, 8096234. [Google Scholar] [CrossRef]
- Cananzi, F.C.M.; Dalgleish, A.; Mudan, S. Surgical Management of Intraabdominal Metastases from Melanoma: Role of the Neutrophil to Lymphocyte Ratio as a Potential Prognostic Factor. World J. Surg. 2014, 38, 1542–1550. [Google Scholar] [CrossRef]
- Cassidy, M.R.; Wolchok, R.E.; Zheng, J.; Panageas, K.S.; Wolchok, J.D.; Coit, D.; Postow, M.A.; Ariyan, C. Neutrophil to Lymphocyte Ratio is Associated With Outcome During Ipilimumab Treatment. EBioMedicine 2017, 18, 56–61. [Google Scholar] [CrossRef]
- Davis, J.L.; Ripley, R.T.; Frankel, T.L.; Maric, I.; Lozier, J.N.; Rosenberg, S.A. Paraneoplastic Granulocytosis in Metastatic Melanoma. Melanoma Res. 2010, 20, 326–329. [Google Scholar] [CrossRef]
- Ferrucci, P.F.; Gandini, S.; Battaglia, A.; Alfieri, S.; Di Giacomo, A.M.; Giannarelli, D.; Cappellini, G.C.A.; De Galitiis, F.; Marchetti, P.; Amato, G.; et al. Baseline Neutrophil-To-Lymphocyte Ratio is Associated with Outcome of Ipilimumab-Treated Metastatic Melanoma Patients. Br. J. Cancer 2015, 112, 1904–1910. [Google Scholar] [CrossRef] [PubMed]
- Wade, R.G.; Robinson, A.V.; Lo, M.C.I.; Keeble, C.; Marples, M.; Dewar, D.J.; Moncrieff, M.D.S.; Peach, H. Baseline Neutrophil-Lymphocyte and Platelet-Lymphocyte Ratios as Biomarkers of Survival in Cutaneous Melanoma: A Multicenter Cohort Study. Ann. Surg. Oncol. 2018, 25, 3341–3349. [Google Scholar] [CrossRef] [PubMed]
- Rachidi, S.; Kaur, M.; Lautenschlaeger, T.; Li, Z. Platelet Count Correlates with Stage and Predicts Survival in Melanoma. Platelets 2019, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Tas, F.; Erturk, K. Anemia in Cutaneous Malignant Melanoma: Low Blood Hemoglobin Level is Associated with Nodal Involvement, Metastatic Disease, and Worse Survival. Nutr. Cancer 2018, 70, 236–240. [Google Scholar] [CrossRef]
- Busti, F.; Marchi, G.; Ugolini, S.; Castagna, A.; Girelli, D. Anemia and Iron Deficiency in Cancer Patients: Role of Iron Replacement Therapy. Pharm. Basel Switz. 2018, 11. [Google Scholar] [CrossRef]
- Weinstein, D.A. Inappropriate Expression of Hepcidin is Associated with Iron Refractory Anemia: Implications for the Anemia of Chronic Disease. Blood 2002, 100, 3776–3781. [Google Scholar] [CrossRef]
- Porto, G. Iron Overload and Immunity. World J. Gastroenterol. 2007, 13, 4707. [Google Scholar] [CrossRef]
- Vyoral, D.; Petrak, J. Hepcidin: A Direct Link Between Iron Metabolism and Immunity. Int. J. Biochem. Cell Biol. 2005, 37, 1768–1773. [Google Scholar] [CrossRef]
- Gonzalez, H.; Hagerling, C.; Werb, Z. Roles of the Immune System in Cancer: From Tumor Initiation to Metastatic Progression. Genes Dev. 2018, 32, 1267–1284. [Google Scholar] [CrossRef]
- Antohe, M.; Nedelcu, R.I.; Nichita, L.; Popp, C.G.; Cioplea, M.; Brinzea, A.; Hodorogea, A.; Calinescu, A.; Balaban, M.; Ion, D.A.; et al. Tumor Infiltrating Lymphocytes: The Regulator of Melanoma Evolution. Oncol. Lett. 2019, 17, 4155–4161. [Google Scholar] [CrossRef]
- Clemente, C.G.; Mihm, M.C.; Bufalino, R.; Zurrida, S.; Collini, P.; Cascinelli, N. Prognostic Value of Tumor Infiltrating Lymphocytes in the Vertical Growth Phase of Primary Cutaneous Melanoma. Cancer 1996, 77, 1303–1310. [Google Scholar] [CrossRef]
- Van Houdt, I.S.; Sluijter, B.J.R.; Moesbergen, L.M.; Vos, W.M.; De Gruijl, T.D.; Molenkamp, B.G.; Van Den Eertwegh, A.J.M.; Hooijberg, E.; Van Leeuwen, P.A.M.; Meijer, C.J.L.M.; et al. Favorable Outcome in Clinically Stage II Melanoma Patients is Associated with the Presence of Activated Tumor Infiltrating T-Lymphocytes and Preserved MHC Class I Antigen Expression. Int. J. Cancer 2008, 123, 609–615. [Google Scholar] [CrossRef] [PubMed]
- Rohaan, M.W.; Van Den Berg, J.H.; Kvistborg, P.; Haanen, J.B.A.G. Adoptive Transfer of Tumor-Infiltrating lymphocytes in Melanoma: A Viable Treatment Option. J. Immunother. Cancer 2018, 6, 102. [Google Scholar] [CrossRef] [PubMed]
- Cizkova, J.; Sinkorova, Z.; Strnadova, K.; Cervinkova, M.; Horak, V.; Sinkora, J.; Stepanova, K.; Sinkora, M. The Role of αβ T-Cells in Spontaneous Regression of Melanoma Tumors in Swine. Dev. Comp. Immunol. 2019, 92, 60–68. [Google Scholar] [CrossRef]
- Overgaard, N.H.; Jung, J.W.; Steptoe, R.J.; Wells, J.W. CD4+/CD8+ Double-Positive T Cells: More Than Just a Developmental Stage? J. Leukoc. Biol. 2015, 97, 31–38. [Google Scholar] [CrossRef]
- Pomorska Mol, M.; Markowska Daniel, I. AGE-Dependent Changes in Relative and Absolute Size of Lymphocyte Subsets in the Blood of Pigs from Birth to Slaughter. Bull. Vet. Inst. Pulawy 2011, 55, 305–310. [Google Scholar]
- Bagot, M.; Echchakir, H.; Mami Chouaib, F.; Delfau Larue, M.H.; Charue, D.; Bernheim, A.; Chouaib, S.; Boumsell, L.; Bensussan, A. Isolation of Tumor-Specific Cytotoxic CD4+ and CD4+CD8dim+ T-Cell Clones Infiltrating a Cutaneous T-Cell Lymphoma. Blood 1998, 91, 4331–4341. [Google Scholar] [CrossRef]
- De Marchi, S.U.; Stinco, G.; Errichetti, E.; Bonin, S.; Di Meo, N.; Trevisan, G. The Influence of the Coexpression of CD4 and CD8 in Cutaneous Lesions on Prognosis of Mycosis Fungoides: A Preliminary Study. J. Skin Cancer 2014, 2014, 624143. [Google Scholar] [CrossRef]
- Desfrançois, J.; Moreau Aubry, A.; Vignard, V.; Godet, Y.; Khammari, A.; Dreno, B.; Jotereau, F.; Gervois, N. Double Positive CD4CD8 Alphabeta T Cells: A New Tumor-Reactive Population in Human Melanomas. PLoS ONE 2010, 5, e8437. [Google Scholar] [CrossRef]
- Roy, S.; Trinchieri, G. Microbiota: A Key Orchestrator of Cancer Therapy. Nat. Rev. Cancer 2017, 17, 271–285. [Google Scholar] [CrossRef]
- McQuade, J.L.; Daniel, C.R.; Helmink, B.A.; Wargo, J.A. Modulating the Microbiome to Improve Therapeutic Response in Cancer. Lancet Oncol. 2019, 20, e77–e91. [Google Scholar] [CrossRef]
- Warner, A.B.; McQuade, J.L. Modifiable Host Factors in Melanoma: Emerging Evidence for Obesity, Diet, Exercise, and the Microbiome. Curr. Oncol. Rep. 2019, 21, 72. [Google Scholar] [CrossRef] [PubMed]
- Gopalakrishnan, V.; Spencer, C.N.; Nezi, L.; Reuben, A.; Andrews, M.C.; Karpinets, T.V.; Prieto, P.A.; Vicente, D.; Hoffman, K.; Wei, S.C.; et al. Gut Microbiome Modulates Response to Anti-PD-1 Immunotherapy in Melanoma Patients. Science 2018, 359, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Salava, A.; Aho, V.; Pereira, P.; Koskinen, K.; Paulin, L.; Auvinen, P.; Lauerma, A. Skin Microbiome in Melanomas and Melanocytic Nevi. Eur. J. Dermatol. EJD 2016, 26, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Svejstil, R.; Salmonova, H.; Cizkova, J. Analysis of Cutaneous Microbiota of Piglets with Hereditary Melanoma. Sci. Agric. Bohem. 2018, 49, 285–290. [Google Scholar] [CrossRef]
- Mrazek, J.; Mekadim, C.; Kucerova, P.; Svejstil, R.; Salmonova, H.; Vlasakova, J.; Tarasova, R.; Cizkova, J.; Cervinkova, M. Melanoma-Related Changes in Skin Microbiome. Folia Microbiol. (Praha) 2019, 64, 435–442. [Google Scholar] [CrossRef]
- Zhou, Z.; Chen, J.; Yao, H.; Hu, H. Fusobacterium and Colorectal Cancer. Front. Oncol. 2018, 8, 371. [Google Scholar] [CrossRef]
- Brennan, C.A.; Garrett, W.S. Fusobacterium Nucleatum-Symbiont, Opportunist and Oncobacterium. Nat. Rev. Microbiol. 2019, 17, 156–166. [Google Scholar] [CrossRef]
- Yu, L.C.H.; Wei, S.C.; Ni, Y.H. Impact of Microbiota in Colorectal Carcinogenesis: Lessons from Experimental Models. Intest. Res. 2018, 16, 346–357. [Google Scholar] [CrossRef]
- Fortyn, K.; Hradecky, J.; Pazdera, J.; Klaudy, J.; Hruban, V.; Dvorak, P.; Matousek, J.; Tichy, J.; Kolin, V. Experimental Elimination of Various Intestinal Segments by Means of Devascularization (Devitalization). Z. Exp. Chir. Transplant. Kunstl. Organe Organ Sekt. Exp. Chir. Ges. Chir. DDR 1985, 18, 34–41. [Google Scholar]
- Fortyn, K.; Hradecky, J.; Pazdera, J.; Klaudy, J.; Hruban, V.; Dvorak, P.; Matousek, J.; Tichy, J.; Kolin, V. Small and Large Intestine Devascularization (Devitalization) and Potentials in the Therapeutic Use of this Operative Method. Z. Exp. Chir. Transplant. Kunstl. Organe Organ Sekt. Exp. Chir. Ges. Chir. DDR 1985, 18, 42–50. [Google Scholar]
- Vasek, P.; Krajnik, J.; Kopsky, D.J.; Kalina, V.; Frydrych, M. Autologous Tumor Immunizing Devascularization of an Invasive Colorectal Cancer: A Case Report and Literature Review. Mol. Clin. Oncol. 2016, 5, 521–526. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Fortyn, K.; Hradecky, J.; Hruban, V.; Horak, V.; Dvorak, P.; Tichy, J. Morphology of Regressive Changes in the Kidney Following Experimental Ischaemia. Int. Urol. Nephrol. 1987, 19, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Fortyn, K.; Hruban, V.; Hradecky, J.; Tichy, J.; Dvorak, P.; Horak, V. A Technique of the Segmental Devitalization of Kidneys in Experiment. Z. Exp. Chir. Transplant. Kunstl. Organe Organ Sekt. Exp. Chir. Ges. Chir. DDR 1988, 21, 275–280. [Google Scholar]
- Fortyn, K.; Hruban, V.; Hradecky, J.; Tichy, J.; Dvorak, P.; Horak, V. The Devitalization of the Stomach by a Devascularization Technique. Acta Chir. Hung. 1988, 29, 163–172. [Google Scholar]
- Fortyn, K.; Hruban, V.; Hradecky, J.; Tichy, J.; Dvorak, P.; Horak, V. Experimental Devascularization (Devitalization) of the Rectum and Sigmoideum. Z. Exp. Chir. Transplant. Kunstl. Organe Organ Sekt. Exp. Chir. Ges. Chir. DDR 1989, 22, 173–179. [Google Scholar]
- Horak, V.; Moravkova, A.; Strnadel, J.; Hradecky, J.; Usvald, D.; Vannucci, L. Devitalization as a Special Surgical Tumour Treatment Inducing Anti-Cancer Response–An Experimental Study in Two Animal Models. In Proceedings of the CIMT Cancer Immunotherapy 6th Annual Meeting, Mainz, Germany, 15–16 May 2008. [Google Scholar]
- Srivastava, P.K.; Udono, H.; Blachere, N.E.; Li, Z. Heat Shock Proteins Transfer Peptides During Antigen Processing and CTL Priming. Immunogenetics 1994, 39, 93–98. [Google Scholar] [CrossRef]
- Binder, R.J.; Blachere, N.E.; Srivastava, P.K. Heat Shock Protein-Chaperoned Peptides but not Free Peptides Introduced into the Cytosol are Presented Efficiently by Major Histocompatibility Complex I Molecules. J. Biol. Chem. 2001, 276, 17163–17171. [Google Scholar] [CrossRef]
- Singh Jasuja, H.; Hilf, N.; Arnold Schild, D.; Schild, H. The Role of Heat Shock Proteins and Their Receptors in the Activation of the Immune System. Biol. Chem. 2001, 382, 629–636. [Google Scholar] [CrossRef]
- Hoos, A.; Levey, D.L. Vaccination with Heat Shock Protein-Peptide Complexes: From Basic Science to Clinical Applications. Expert Rev. Vaccines 2003, 2, 369–379. [Google Scholar] [CrossRef]
- Testori, A.; Richards, J.; Whitman, E.; Mann, G.B.; Lutzky, J.; Camacho, L.; Parmiani, G.; Tosti, G.; Kirkwood, J.M.; Hoos, A.; et al. Phase III Comparison of Vitespen, an Autologous Tumor-Derived Heat Shock Protein gp96 Peptide Complex Vaccine, with Physician’s Choice of Treatment for Stage IV Melanoma: The C-100-21 Study Group. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2008, 26, 955–962. [Google Scholar] [CrossRef] [PubMed]
- Tosti, G.; Di Pietro, A.; Ferrucci, P.F.; Testori, A. HSPPC-96 Vaccine in Metastatic Melanoma Patients: From the State of the Art to a Possible Future. Expert Rev. Vaccines 2009, 8, 1513–1526. [Google Scholar] [CrossRef] [PubMed]
- Di Pietro, A.; Tosti, G.; Ferrucci, P.F.; Testori, A. The Immunological era in Melanoma Treatment: New Challenges for Heat Shock Protein-Based Vaccine in the Advanced Disease. Expert Opin. Biol. Ther. 2011, 11, 1395–1407. [Google Scholar] [CrossRef] [PubMed]
- Shevtsov, M.; Multhoff, G. Heat Shock Protein-Peptide and HSP-Based Immunotherapies for the Treatment of Cancer. Front. Immunol. 2016, 7, 171. [Google Scholar] [CrossRef] [PubMed]
- Prather, R.S.; Shen, M.; Dai, Y. Genetically Modified Pigs for Medicine and Agriculture. Biotechnol. Genet. Eng. Rev. 2008, 25, 245–265. [Google Scholar]
- Flisikowska, T.; Kind, A.; Schnieke, A. Genetically Modified Pigs to Model Human Diseases. J. Appl. Genet. 2014, 55, 53–64. [Google Scholar] [CrossRef]
- Watson, A.L.; Carlson, D.F.; Largaespada, D.A.; Hackett, P.B.; Fahrenkrug, S.C. Engineered Swine Models of Cancer. Front. Genet. 2016, 7, 78. [Google Scholar] [CrossRef]
- Schachtschneider, K.M.; Schwind, R.M.; Newson, J.; Kinachtchouk, N.; Rizko, M.; Mendoza Elias, N.; Grippo, P.; Principe, D.R.; Park, A.; Overgaard, N.H.; et al. The Oncopig Cancer Model: An Innovative Large Animal Translational Oncology Platform. Front. Oncol. 2017, 7, 190. [Google Scholar] [CrossRef]
- Vodicka, P.; Smetana, K.; Dvorankova, B.; Emerick, T.; Xu, Y.Z.; Ourednik, J.; Ourednik, V.; Motlik, J. The Miniature Pig as an Animal Model in Biomedical Research. Ann. N. Y. Acad. Sci. 2005, 1049, 161–171. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Horak, V.; Palanova, A.; Cizkova, J.; Miltrova, V.; Vodicka, P.; Kupcova Skalnikova, H. Melanoma-Bearing Libechov Minipig (MeLiM): The Unique Swine Model of Hereditary Metastatic Melanoma. Genes 2019, 10, 915. https://doi.org/10.3390/genes10110915
Horak V, Palanova A, Cizkova J, Miltrova V, Vodicka P, Kupcova Skalnikova H. Melanoma-Bearing Libechov Minipig (MeLiM): The Unique Swine Model of Hereditary Metastatic Melanoma. Genes. 2019; 10(11):915. https://doi.org/10.3390/genes10110915
Chicago/Turabian StyleHorak, Vratislav, Anna Palanova, Jana Cizkova, Veronika Miltrova, Petr Vodicka, and Helena Kupcova Skalnikova. 2019. "Melanoma-Bearing Libechov Minipig (MeLiM): The Unique Swine Model of Hereditary Metastatic Melanoma" Genes 10, no. 11: 915. https://doi.org/10.3390/genes10110915
APA StyleHorak, V., Palanova, A., Cizkova, J., Miltrova, V., Vodicka, P., & Kupcova Skalnikova, H. (2019). Melanoma-Bearing Libechov Minipig (MeLiM): The Unique Swine Model of Hereditary Metastatic Melanoma. Genes, 10(11), 915. https://doi.org/10.3390/genes10110915




