A Double-Edged Sword: Aneuploidy is a Prevalent Strategy in Fungal Adaptation
Abstract
1. Introduction
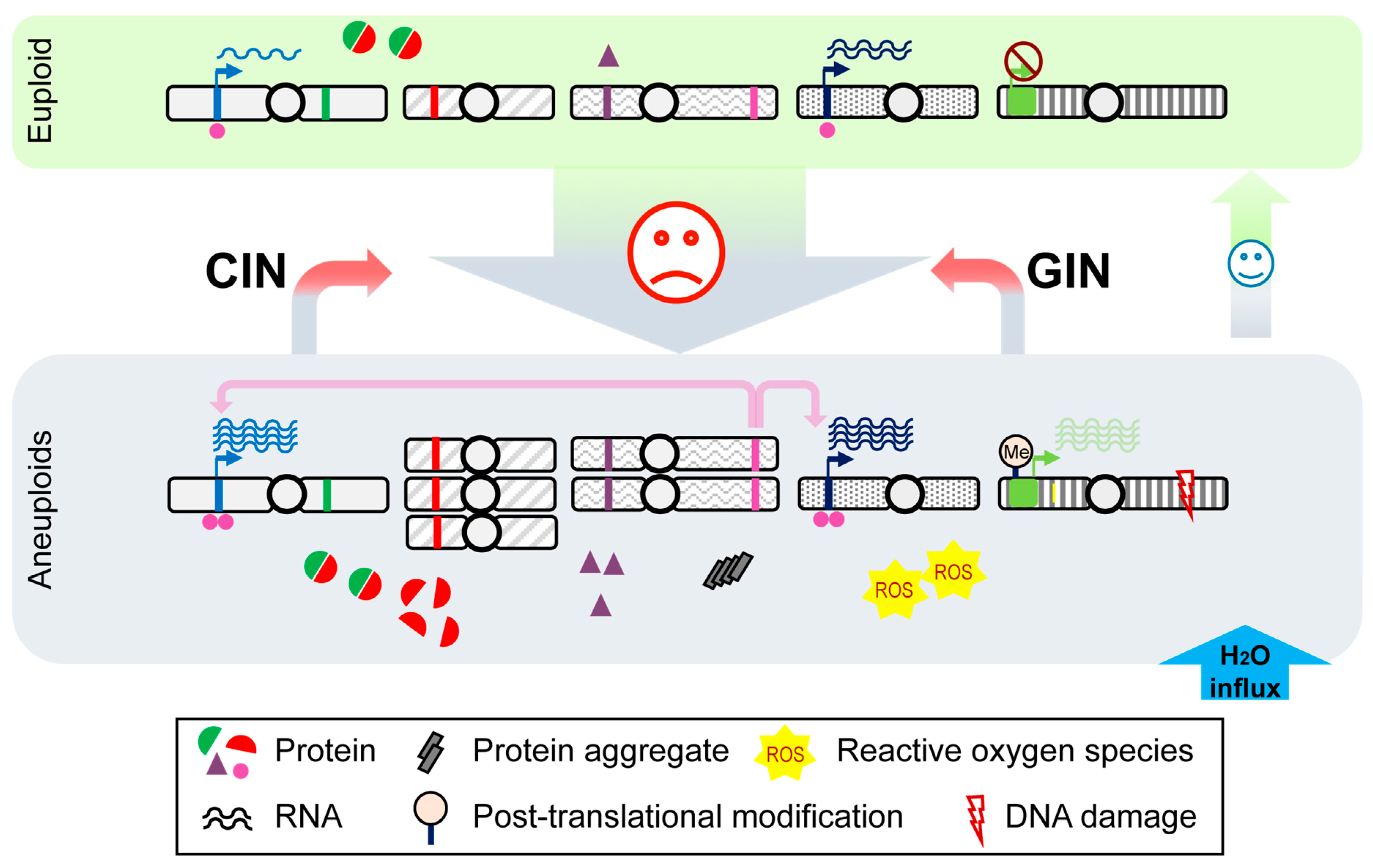
2. Cellular Impacts of Genome Aneuploidization
3. Aneuploid Fungi Are Widespread in Diverse Environments
4. Gene Copy Number Variations on Aneuploid Chromosomes Drive Adaptive Phenotypes
5. Future Perspective
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Torres, E.M.; Sokolsky, T.; Tucker, C.M.; Chan, L.Y.; Boselli, M.; Dunham, M.J.; Amon, A. Effects of aneuploidy on cellular physiology and cell division in haploid yeast. Science 2007, 317, 916–924. [Google Scholar] [CrossRef] [PubMed]
- Williams, B.R.; Prabhu, V.R.; Hunter, K.E.; Glazier, C.M.; Whittaker, C.A.; Housman, D.E.; Amon, A. Aneuploidy affects proliferation and spontaneous immortalization in mammalian cells. Science 2008, 322, 703–709. [Google Scholar] [CrossRef] [PubMed]
- Stingele, S.; Stoehr, G.; Peplowska, K.; Cox, J.; Mann, M.; Storchova, Z. Global analysis of genome, transcriptome and proteome reveals the response to aneuploidy in human cells. Mol. Syst. Biol. 2012, 8, 608. [Google Scholar] [CrossRef] [PubMed]
- Segal, D.J.; McCoy, E.E. Studies on Down’s syndrome in tissue culture. I. Growth rates and protein contents of fibroblast cultures. J. Cell. Physiol. 1974, 83, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Gogendeau, D.; Siudeja, K.; Gambarotto, D.; Pennetier, C.; Bardin, A.J.; Basto, R. Aneuploidy causes premature differentiation of neural and intestinal stem cells. Nat. Commun. 2015, 6, 8894. [Google Scholar] [CrossRef] [PubMed]
- Pavelka, N.; Rancati, G.; Zhu, J.; Bradford, W.D.; Saraf, A.; Florens, L.; Sanderson, B.W.; Hattem, G.L.; Li, R. Aneuploidy confers quantitative proteome changes and phenotypic variation in budding yeast. Nature 2010, 468, 321–325. [Google Scholar] [CrossRef] [PubMed]
- Blakeslee, A.F.; Belling, J.; Farnham, M.E. Chromosomal duplication and mendelian phenomena in datura mutants. Science 1920, 52, 388–390. [Google Scholar] [CrossRef]
- Satina, S.; Blakeslee, A.F.; Avery, A.G. Balanced and unbalanced haploids in datura. J. Hered. 1937, 28, 193–202. [Google Scholar] [CrossRef]
- Roper, R.J.; Reeves, R.H. Understanding the basis for Down syndrome phenotypes. PLoS Genet. 2006, 2, e50. [Google Scholar] [CrossRef]
- Beroukhim, R.; Mermel, C.H.; Porter, D.; Wei, G.; Raychaudhuri, S.; Donovan, J.; Barretina, J.; Boehm, J.S.; Dobson, J.; Urashima, M.; et al. The landscape of somatic copy-number alteration across human cancers. Nature 2010, 463, 899–905. [Google Scholar] [CrossRef]
- Lejeune, J. Etude des chromosomes somatiques de neuf enfants mongoliens. C. R. Acad. Sci. 1959, 248, 1721–1722. [Google Scholar]
- Gordon, D.J.; Resio, B.; Pellman, D. Causes and consequences of aneuploidy in cancer. Nat. Rev. Genet. 2012, 13, 189–203. [Google Scholar] [CrossRef] [PubMed]
- Hanks, S.; Coleman, K.; Reid, S.; Plaja, A.; Firth, H.; Fitzpatrick, D.; Kidd, A.; Méhes, K.; Nash, R.; Robin, N.; et al. Constitutional aneuploidy and cancer predisposition caused by biallelic mutations in BUB1B. Nat. Genet. 2004, 36, 1159–1161. [Google Scholar] [CrossRef] [PubMed]
- Duncan, A.W.; Hanlon Newell, A.E.; Smith, L.; Wilson, E.M.; Olson, S.B.; Thayer, M.J.; Strom, S.C.; Grompe, M. Frequent aneuploidy among normal human hepatocytes. Gastroenterology 2012, 142, 25–28. [Google Scholar] [CrossRef] [PubMed]
- Knouse, K.A.; Wu, J.; Whittaker, C.A.; Amon, A. Single cell sequencing reveals low levels of aneuploidy across mammalian tissues. Proc. Natl. Acad. Sci. USA 2014, 111, 13409–13414. [Google Scholar] [CrossRef] [PubMed]
- Rehen, S.K.; McConnell, M.J.; Kaushal, D.; Kingsbury, M.A.; Yang, A.H.; Chun, J. Chromosomal variation in neurons of the developing and adult mammalian nervous system. Proc. Natl. Acad. Sci. USA 2001, 98, 13361–13366. [Google Scholar] [CrossRef] [PubMed]
- Rehen, S.K.; Yung, Y.C.; McCreight, M.P.; Kaushal, D.; Yang, A.H.; Almeida, B.S.V.; Kingsbury, M.A.; Cabral, K.M.S.; McConnell, M.J.; Anliker, B.; et al. Constitutional aneuploidy in the normal human brain. J. Neurosci. 2005, 25, 2176–2180. [Google Scholar] [CrossRef] [PubMed]
- Selmecki, A.; Forche, A.; Berman, J. Aneuploidy and isochromosome formation in drug-resistant Candida albicans. Science 2006, 313, 367–370. [Google Scholar] [CrossRef]
- Hirakawa, M.P.; Martinez, D.A.; Sakthikumar, S.; Anderson, M.Z.; Berlin, A.; Gujja, S.; Zeng, Q.; Zisson, E.; Wang, J.M.; Greenberg, J.M.; et al. Genetic and phenotypic intra-species variation in Candida albicans. Genome Res. 2015, 25, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Peter, J.; De Chiara, M.; Friedrich, A.; Yue, J.-X.; Pflieger, D.; Bergström, A.; Sigwalt, A.; Barre, B.; Freel, K.; Llored, A.; et al. Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature 2018, 556, 339–344. [Google Scholar] [CrossRef] [PubMed]
- Tan, Z.; Hays, M.; Cromie, G.A.; Jeffery, E.W.; Scott, A.C.; Ahyong, V.; Sirr, A.; Skupin, A.; Dudley, A.M. Aneuploidy underlies a multicellular phenotypic switch. Proc. Natl. Acad. Sci. USA 2013, 110, 12367–12372. [Google Scholar] [CrossRef] [PubMed]
- Bennett, R.J.; Forche, A.; Berman, J. Rapid mechanisms for generating genome diversity: Whole ploidy shifts, aneuploidy, and loss of heterozygosity. Cold Spring Harb. Perspect. Med. 2014, 4, a019604. [Google Scholar] [CrossRef] [PubMed]
- Sionov, E.; Lee, H.; Chang, Y.C.; Kwon-Chung, K.J. Cryptococcus neoformans overcomes stress of azole drugs by formation of disomy in specific multiple chromosomes. PLoS Pathog. 2010, 6, e1000848. [Google Scholar] [CrossRef] [PubMed]
- Berman, J. Ploidy plasticity: A rapid and reversible strategy for adaptation to stress. FEMS Yeast Res. 2016, 16, fow020. [Google Scholar] [CrossRef]
- Torres, E.M.; Dephoure, N.; Panneerselvam, A.; Tucker, C.M.; Whittaker, C.A.; Gygi, S.P.; Dunham, M.J.; Amon, A. Identification of aneuploidy-tolerating mutations. Cell 2010, 143, 71–83. [Google Scholar] [CrossRef]
- Sheltzer, J.M.; Blank, H.M.; Pfau, S.J.; Tange, Y.; George, B.M.; Humpton, T.J.; Brito, I.L.; Hiraoka, Y.; Niwa, O.; Amon, A. Aneuploidy drives genomic instability in yeast. Science 2011, 333, 1026–1030. [Google Scholar] [CrossRef]
- Sheltzer, J.M.; Torres, E.M.; Dunham, M.J.; Amon, A. Transcriptional consequences of aneuploidy. Proc. Natl. Acad. Sci. USA 2012, 109, 12644–12649. [Google Scholar] [CrossRef]
- Oromendia, A.B.; Dodgson, S.E.; Amon, A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 2012, 26, 2696–2708. [Google Scholar] [CrossRef]
- Thorburn, R.R.; Gonzalez, C.; Brar, G.A.; Christen, S.; Carlile, T.M.; Ingolia, N.T.; Sauer, U.; Weissman, J.S.; Amon, A. Aneuploid yeast strains exhibit defects in cell growth and passage through START. Mol. Biol. Cell 2013, 24, 1274–1289. [Google Scholar] [CrossRef]
- Dodgson, S.E.; Santaguida, S.; Kim, S.; Sheltzer, J.; Amon, A. The pleiotropic deubiquitinase Ubp3 confers aneuploidy tolerance. Genes Dev. 2016, 30, 2259–2271. [Google Scholar] [CrossRef]
- Tsai, H.-J.; Nelliat, A.R.; Choudhury, M.I.; Kucharavy, A.; Bradford, W.D.; Cook, M.E.; Kim, J.; Mair, D.B.; Sun, S.X.; Schatz, M.C.; et al. Hypo-osmotic-like stress underlies general cellular defects of aneuploidy. Nature 2019, 570, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Dephoure, N.; Hwang, S.; O’Sullivan, C.; Dodgson, S.E.; Gygi, S.P.; Amon, A.; Torres, E.M. Quantitative proteomic analysis reveals posttranslational responses to aneuploidy in yeast. eLife 2014, 3, e03023. [Google Scholar] [CrossRef] [PubMed]
- Blank, H.M.; Sheltzer, J.M.; Meehl, C.M.; Amon, A. Mitotic entry in the presence of DNA damage is a widespread property of aneuploidy in yeast. Mol. Biol. Cell 2015, 26, 1440–1451. [Google Scholar] [CrossRef] [PubMed]
- Taggart, J.C.; Li, G.-W. Production of Protein-Complex Components Is Stoichiometric and Lacks General Feedback Regulation in Eukaryotes. Cell Syst. 2018, 7, 580–589.e4. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, B.; Solomon, F. Phenotypic consequences of tubulin overproduction in Saccharomyces cerevisiae: Differences between alpha-tubulin and beta-tubulin. Mol. Cell. Biol. 1990, 10, 5295–5304. [Google Scholar] [CrossRef] [PubMed]
- Gasch, A.P.; Spellman, P.T.; Kao, C.M.; Carmel-Harel, O.; Eisen, M.B.; Storz, G.; Botstein, D.; Brown, P.O. Genomic expression programs in the response of yeast cells to environmental changes. Mol. Biol. Cell 2000, 11, 4241–4257. [Google Scholar] [CrossRef]
- Cromie, G.A.; Tan, Z.; Hays, M.; Jeffery, E.W.; Dudley, A.M. Dissecting Gene Expression Changes Accompanying a Ploidy-Based Phenotypic Switch. G3 2017, 7, 233–246. [Google Scholar] [CrossRef] [PubMed]
- Hose, J.; Yong, C.M.; Sardi, M.; Wang, Z.; Newton, M.A.; Gasch, A.P. Dosage compensation can buffer copy-number variation in wild yeast. eLife 2015, 4, e05462. [Google Scholar] [CrossRef]
- Wertheimer, N.B.; Stone, N.; Berman, J. Ploidy dynamics and evolvability in fungi. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2016, 371, 20150461. [Google Scholar] [CrossRef]
- Zhu, Y.O.; Siegal, M.L.; Hall, D.W.; Petrov, D.A. Precise estimates of mutation rate and spectrum in yeast. Proc. Natl. Acad. Sci. USA 2014, 111, E2310–E2318. [Google Scholar] [CrossRef]
- Ezov, T.K.; Boger-Nadjar, E.; Frenkel, Z.; Katsperovski, I.; Kemeny, S.; Nevo, E.; Korol, A.; Kashi, Y. Molecular-genetic biodiversity in a natural population of the yeast Saccharomyces cerevisiae from “Evolution Canyon”: Microsatellite polymorphism, ploidy and controversial sexual status. Genetics 2006, 174, 1455–1468. [Google Scholar] [CrossRef] [PubMed]
- Lidzbarsky, G.A.; Shkolnik, T.; Nevo, E. Adaptive response to DNA-damaging agents in natural Saccharomyces cerevisiae populations from “Evolution Canyon”, Mt. Carmel, Israel. PLoS ONE 2009, 4, e5914. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.-L.; Lai, H.-Y.; Tung, S.-Y.; Leu, J.-Y. Dynamic large-scale chromosomal rearrangements fuel rapid adaptation in yeast populations. PLoS Genet. 2013, 9, e1003232. [Google Scholar] [CrossRef] [PubMed]
- Forche, A. Large-Scale Chromosomal Changes and Associated Fitness Consequences in Pathogenic Fungi. Curr. Fungal Infect. Rep. 2014, 8, 163–170. [Google Scholar] [CrossRef] [PubMed]
- van den Bossche, H.; Marichal, P.; Odds, F.C.; Le Jeune, L.; Coene, M.C. Characterization of an azole-resistant Candida glabrata isolate. Antimicrob. Agents Chemother. 1992, 36, 2602–2610. [Google Scholar] [CrossRef] [PubMed]
- Marichal, P.; Vanden Bossche, H.; Odds, F.C.; Nobels, G.; Warnock, D.W.; Timmerman, V.; Van Broeckhoven, C.; Fay, S.; Mose-Larsen, P. Molecular biological characterization of an azole-resistant Candida glabrata isolate. Antimicrob. Agents Chemother. 1997, 41, 2229–2237. [Google Scholar] [CrossRef] [PubMed]
- Poláková, S.; Blume, C.; Zárate, J.A.; Mentel, M.; Jørck-Ramberg, D.; Stenderup, J.; Piskur, J. Formation of new chromosomes as a virulence mechanism in yeast Candida glabrata. Proc. Natl. Acad. Sci. USA 2009, 106, 2688–2693. [Google Scholar] [CrossRef]
- Selmecki, A.; Gerami-Nejad, M.; Paulson, C.; Forche, A.; Berman, J. An isochromosome confers drug resistance in vivo by amplification of two genes, ERG11 and TAC1. Mol. Microbiol. 2008, 68, 624–641. [Google Scholar] [CrossRef]
- Ford, C.B.; Funt, J.M.; Abbey, D.; Issi, L.; Guiducci, C.; Martinez, D.A.; Delorey, T.; Li, B.Y.; White, T.C.; Cuomo, C.; et al. The evolution of drug resistance in clinical isolates of Candida albicans. eLife 2015, 4, e00662. [Google Scholar] [CrossRef]
- Selmecki, A.M.; Dulmage, K.; Cowen, L.E.; Anderson, J.B.; Berman, J. Acquisition of aneuploidy provides increased fitness during the evolution of antifungal drug resistance. PLoS Genet. 2009, 5, e1000705. [Google Scholar] [CrossRef]
- Bravo Ruiz, G.; Ross, Z.K.; Holmes, E.; Schelenz, S.; Gow, N.A.R.; Lorenz, A. Rapid and extensive karyotype diversification in haploid clinical Candida auris isolates. Curr. Genet. 2019, 65, 1217–1228. [Google Scholar] [CrossRef] [PubMed]
- Ngamskulrungroj, P.; Chang, Y.; Hansen, B.; Bugge, C.; Fischer, E.; Kwon-Chung, K.J. Characterization of the chromosome 4 genes that affect fluconazole-induced disomy formation in Cryptococcus neoformans. PLoS ONE 2012, 7, e33022. [Google Scholar] [CrossRef] [PubMed]
- Sionov, E.; Chang, Y.C.; Kwon-Chung, K.J. Azole heteroresistance in Cryptococcus neoformans: Emergence of resistant clones with chromosomal disomy in the mouse brain during fluconazole treatment. Antimicrob. Agents Chemother. 2013, 57, 5127–5130. [Google Scholar] [CrossRef] [PubMed]
- Gerstein, A.C.; Fu, M.S.; Mukaremera, L.; Li, Z.; Ormerod, K.L.; Fraser, J.A.; Berman, J.; Nielsen, K. Polyploid titan cells produce haploid and aneuploid progeny to promote stress adaptation. mBio 2015, 6, e01340-15. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.O.; Sherlock, G.; Petrov, D.A. Whole Genome Analysis of 132 Clinical Saccharomyces cerevisiae Strains Reveals Extensive Ploidy Variation. G3 2016, 6, 2421–2434. [Google Scholar] [CrossRef] [PubMed]
- Forche, A.; Solis, N.V.; Swidergall, M.; Thomas, R.; Guyer, A.; Beach, A.; Cromie, G.A.; Le, G.T.; Lowell, E.; Pavelka, N.; et al. Selection of Candida albicans trisomy during oropharyngeal infection results in a commensal-like phenotype. PLoS Genet. 2019, 15, e1008137. [Google Scholar] [CrossRef]
- Hughes, T.R.; Roberts, C.J.; Dai, H.; Jones, A.R.; Meyer, M.R.; Slade, D.; Burchard, J.; Dow, S.; Ward, T.R.; Kidd, M.J.; et al. Widespread aneuploidy revealed by DNA microarray expression profiling. Nat. Genet. 2000, 25, 333–337. [Google Scholar] [CrossRef]
- Gasch, A.P.; Huang, M.; Metzner, S.; Botstein, D.; Elledge, S.J.; Brown, P.O. Genomic expression responses to DNA-damaging agents and the regulatory role of the yeast ATR homolog Mec1p. Mol. Biol. Cell 2001, 12, 2987–3003. [Google Scholar] [CrossRef]
- Gresham, D.; Desai, M.M.; Tucker, C.M.; Jenq, H.T.; Pai, D.A.; Ward, A.; DeSevo, C.G.; Botstein, D.; Dunham, M.J. The repertoire and dynamics of evolutionary adaptations to controlled nutrient-limited environments in yeast. PLoS Genet. 2008, 4, e1000303. [Google Scholar] [CrossRef]
- Rancati, G.; Pavelka, N.; Fleharty, B.; Noll, A.; Trimble, R.; Walton, K.; Perera, A.; Staehling-Hampton, K.; Seidel, C.W.; Li, R. Aneuploidy underlies rapid adaptive evolution of yeast cells deprived of a conserved cytokinesis motor. Cell 2008, 135, 879–893. [Google Scholar] [CrossRef]
- Chen, G.; Bradford, W.D.; Seidel, C.W.; Li, R. Hsp90 stress potentiates rapid cellular adaptation through induction of aneuploidy. Nature 2012, 482, 246–250. [Google Scholar] [CrossRef] [PubMed]
- Yona, A.H.; Manor, Y.S.; Herbst, R.H.; Romano, G.H.; Mitchell, A.; Kupiec, M.; Pilpel, Y.; Dahan, O. Chromosomal duplication is a transient evolutionary solution to stress. Proc. Natl. Acad. Sci. USA 2012, 109, 21010–21015. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.K.; Liu, T.; Parreiras, L.S.; Williams, D.L.; Wohlbach, D.J.; Bice, B.D.; Ong, I.M.; Breuer, R.J.; Qin, L.; Busalacchi, D.; et al. Harnessing genetic diversity in Saccharomyces cerevisiae for fermentation of xylose in hydrolysates of alkaline hydrogen peroxide-pretreated biomass. Appl. Environ. Microbiol. 2014, 80, 540–554. [Google Scholar] [CrossRef] [PubMed]
- Sunshine, A.B.; Payen, C.; Ong, G.T.; Liachko, I.; Tan, K.M.; Dunham, M.J. The fitness consequences of aneuploidy are driven by condition-dependent gene effects. PLoS Biol. 2015, 13, e1002155. [Google Scholar] [CrossRef] [PubMed]
- Gerstein, A.C.; Ono, J.; Lo, D.S.; Campbell, M.L.; Kuzmin, A.; Otto, S.P. Too much of a good thing: The unique and repeated paths toward copper adaptation. Genetics 2015, 199, 555–571. [Google Scholar] [CrossRef] [PubMed]
- Selmecki, A.M.; Maruvka, Y.E.; Richmond, P.A.; Guillet, M.; Shoresh, N.; Sorenson, A.L.; De, S.; Kishony, R.; Michor, F.; Dowell, R.; et al. Polyploidy can drive rapid adaptation in yeast. Nature 2015, 519, 349–352. [Google Scholar] [CrossRef] [PubMed]
- Sirr, A.; Cromie, G.A.; Jeffery, E.W.; Gilbert, T.L.; Ludlow, C.L.; Scott, A.C.; Dudley, A.M. Allelic variation, aneuploidy, and nongenetic mechanisms suppress a monogenic trait in yeast. Genetics 2015, 199, 247–262. [Google Scholar] [CrossRef] [PubMed]
- Voordeckers, K.; Kominek, J.; Das, A.; Espinosa-Cantú, A.; De Maeyer, D.; Arslan, A.; Van Pee, M.; van der Zande, E.; Meert, W.; Yang, Y.; et al. Adaptation to High Ethanol Reveals Complex Evolutionary Pathways. PLoS Genet. 2015, 11, e1005635. [Google Scholar] [CrossRef]
- Millet, C.; Makovets, S. Aneuploidy as a mechanism of adaptation to telomerase insufficiency. Curr. Genet. 2016, 62, 557–564. [Google Scholar] [CrossRef]
- Hope, E.A.; Amorosi, C.J.; Miller, A.W.; Dang, K.; Heil, C.S.; Dunham, M.J. Experimental Evolution Reveals Favored Adaptive Routes to Cell Aggregation in Yeast. Genetics 2017, 206, 1153–1167. [Google Scholar] [CrossRef] [PubMed]
- Beaupere, C.; Dinatto, L.; Wasko, B.M.; Chen, R.B.; VanValkenburg, L.; Kiflezghi, M.G.; Lee, M.B.; Promislow, D.E.L.; Dang, W.; Kaeberlein, M.; et al. Genetic screen identifies adaptive aneuploidy as a key mediator of ER stress resistance in yeast. Proc. Natl. Acad. Sci. USA 2018, 115, 9586–9591. [Google Scholar] [CrossRef] [PubMed]
- Lauer, S.; Avecilla, G.; Spealman, P.; Sethia, G.; Brandt, N.; Levy, S.F.; Gresham, D. Single-cell copy number variant detection reveals the dynamics and diversity of adaptation. PLoS Biol. 2018, 16, e3000069. [Google Scholar] [CrossRef] [PubMed]
- Dunham, M.J.; Badrane, H.; Ferea, T.; Adams, J.; Brown, P.O.; Rosenzweig, F.; Botstein, D. Characteristic genome rearrangements in experimental evolution of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 2002, 99, 16144–16149. [Google Scholar] [CrossRef] [PubMed]
- Pavelka, N.; Rancati, G.; Li, R. Dr Jekyll and Mr Hyde: Role of aneuploidy in cellular adaptation and cancer. Curr. Opin. Cell Biol. 2010, 22, 809–815. [Google Scholar] [CrossRef]
- Yang, F.; Kravets, A.; Bethlendy, G.; Welle, S.; Rustchenko, E. Chromosome 5 monosomy of Candida albicans controls susceptibility to various toxic agents, including major antifungals. Antimicrob. Agents Chemother. 2013, 57, 5026–5036. [Google Scholar] [CrossRef]
- Yang, F.; Teoh, F.; Tan, A.S.M.; Cao, Y.; Pavelka, N.; Berman, J. Aneuploidy Enables Cross-Adaptation to Unrelated Drugs. Mol. Biol. Evol. 2019, 36, 1768–1782. [Google Scholar] [CrossRef]
- Mount, H.O.; Revie, N.M.; Todd, R.T.; Anstett, K.; Collins, C.; Costanzo, M.; Boone, C.; Robbins, N.; Selmecki, A.; Cowen, L.E. Global analysis of genetic circuitry and adaptive mechanisms enabling resistance to the azole antifungal drugs. PLoS Genet. 2018, 14, e1007319. [Google Scholar] [CrossRef]
- Ma, L.-J.; van der Does, H.C.; Borkovich, K.A.; Coleman, J.J.; Daboussi, M.-J.; Di Pietro, A.; Dufresne, M.; Freitag, M.; Grabherr, M.; Henrissat, B.; et al. Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 2010, 464, 367–373. [Google Scholar] [CrossRef]
- Miao, V.P.; Covert, S.F.; VanEtten, H.D. A fungal gene for antibiotic resistance on a dispensable (“B”) chromosome. Science 1991, 254, 1773–1776. [Google Scholar] [CrossRef]
- van den Broek, M.; Bolat, I.; Nijkamp, J.F.; Ramos, E.; Luttik, M.A.H.; Koopman, F.; Geertman, J.M.; de Ridder, D.; Pronk, J.T.; Daran, J.-M. Chromosomal Copy Number Variation in Saccharomyces pastorianus Is Evidence for Extensive Genome Dynamics in Industrial Lager Brewing Strains. Appl. Environ. Microbiol. 2015, 81, 6253–6267. [Google Scholar] [CrossRef]
- Morard, M.; Macías, L.G.; Adam, A.C.; Lairón-Peris, M.; Pérez-Torrado, R.; Toft, C.; Barrio, E. Aneuploidy and Ethanol Tolerance in Saccharomyces cerevisiae. Front. Genet. 2019, 10, 82. [Google Scholar] [CrossRef] [PubMed]
- Guijo, S.; Mauricio, J.C.; Salmon, J.M.; Ortega, J.M. Determination of the relative ploidy in different Saccharomyces cerevisiae strains used for fermentation and “flor”film ageing of dry sherry-type wines. Yeast 1997, 13, 101–117. [Google Scholar] [CrossRef]
- Zhang, K.; Zhang, L.-J.; Fang, Y.-H.; Jin, X.-N.; Qi, L.; Wu, X.-C.; Zheng, D.-Q. Genomic structural variation contributes to phenotypic change of industrial bioethanol yeast Saccharomyces cerevisiae. FEMS Yeast Res. 2016, 16, fov118. [Google Scholar] [CrossRef] [PubMed]
- Will, J.L.; Kim, H.S.; Clarke, J.; Painter, J.C.; Fay, J.C.; Gasch, A.P. Incipient balancing selection through adaptive loss of aquaporins in natural Saccharomyces cerevisiae populations. PLoS Genet. 2010, 6, e1000893. [Google Scholar] [CrossRef] [PubMed]
- Coste, A.; Turner, V.; Ischer, F.; Morschhäuser, J.; Forche, A.; Selmecki, A.; Berman, J.; Bille, J.; Sanglard, D. A mutation in Tac1p, a transcription factor regulating CDR1 and CDR2, is coupled with loss of heterozygosity at chromosome 5 to mediate antifungal resistance in Candida albicans. Genetics 2006, 172, 2139–2156. [Google Scholar] [CrossRef] [PubMed]
- Coste, A.; Selmecki, A.; Forche, A.; Diogo, D.; Bougnoux, M.-E.; d’Enfert, C.; Berman, J.; Sanglard, D. Genotypic evolution of azole resistance mechanisms in sequential Candida albicans isolates. Eukaryot. Cell 2007, 6, 1889–1904. [Google Scholar] [CrossRef] [PubMed]
- Perea, S.; López-Ribot, J.L.; Kirkpatrick, W.R.; McAtee, R.K.; Santillán, R.A.; Martínez, M.; Calabrese, D.; Sanglard, D.; Patterson, T.F. Prevalence of molecular mechanisms of resistance to azole antifungal agents in Candida albicans strains displaying high-level fluconazole resistance isolated from human immunodeficiency virus-infected patients. Antimicrob. Agents Chemother. 2001, 45, 2676–2684. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.T.; Znaidi, S.; Barker, K.S.; Xu, L.; Homayouni, R.; Saidane, S.; Morschhäuser, J.; Nantel, A.; Raymond, M.; Rogers, P.D. Genome-wide expression and location analyses of the Candida albicans Tac1p regulon. Eukaryot. Cell 2007, 6, 2122–2138. [Google Scholar] [CrossRef] [PubMed]
- Harrison, B.D.; Hashemi, J.; Bibi, M.; Pulver, R.; Bavli, D.; Nahmias, Y.; Wellington, M.; Sapiro, G.; Berman, J. A tetraploid intermediate precedes aneuploid formation in yeasts exposed to fluconazole. PLoS Biol. 2014, 12, e1001815. [Google Scholar] [CrossRef]
- Muller, H.; Thierry, A.; Coppée, J.-Y.; Gouyette, C.; Hennequin, C.; Sismeiro, O.; Talla, E.; Dujon, B.; Fairhead, C. Genomic polymorphism in the population of Candida glabrata: Gene copy-number variation and chromosomal translocations. Fungal Genet. Biol. 2009, 46, 264–276. [Google Scholar] [CrossRef]
- Chang, Y.C.; Khanal Lamichhane, A.; Kwon-Chung, K.J. Cryptococcus neoformans, Unlike Candida albicans, Forms Aneuploid Clones Directly from Uninucleated Cells under Fluconazole Stress. mBio 2018, 9, e01290-18. [Google Scholar] [CrossRef] [PubMed]
- Altamirano, S.; Fang, D.; Simmons, C.; Sridhar, S.; Wu, P.; Sanyal, K.; Kozubowski, L. Fluconazole-Induced Ploidy Change in Cryptococcus neoformans Results from the Uncoupling of Cell Growth and Nuclear Division. mSphere 2017, 2, e00205-17. [Google Scholar] [CrossRef] [PubMed]
- Bennett, R.J.; Johnson, A.D. Completion of a parasexual cycle in Candida albicans by induced chromosome loss in tetraploid strains. EMBO J. 2003, 22, 2505–2515. [Google Scholar] [CrossRef] [PubMed]
- Forche, A.; Alby, K.; Schaefer, D.; Johnson, A.D.; Berman, J.; Bennett, R.J. The parasexual cycle in Candida albicans provides an alternative pathway to meiosis for the formation of recombinant strains. PLoS Biol. 2008, 6, e110. [Google Scholar] [CrossRef] [PubMed]
- Berman, J.; Hadany, L. Does stress induce (para) sex? Implications for Candida albicans evolution. Trends Genet. 2012, 28, 197–203. [Google Scholar] [CrossRef]
- Hickman, M.A.; Paulson, C.; Dudley, A.; Berman, J. Parasexual Ploidy Reduction Drives Population Heterogeneity through Random and Transient Aneuploidy in Candida albicans. Genetics 2015, 200, 781–794. [Google Scholar] [CrossRef] [PubMed]
- Todd, R.T.; Wikoff, T.D.; Forche, A.; Selmecki, A. Genome plasticity in Candida albicans is driven by long repeat sequences. eLife 2019, 8, e45954. [Google Scholar] [CrossRef]
- Payen, C.; Di Rienzi, S.C.; Ong, G.T.; Pogachar, J.L.; Sanchez, J.C.; Sunshine, A.B.; Raghuraman, M.K.; Brewer, B.J.; Dunham, M.J. The dynamics of diverse segmental amplifications in populations of Saccharomyces cerevisiae adapting to strong selection. G3 2014, 4, 399–409. [Google Scholar] [CrossRef]
- Möller, M.; Stukenbrock, E.H. Evolution and genome architecture in fungal plant pathogens. Nat. Rev. Microbiol. 2017, 15, 756–771. [Google Scholar] [CrossRef]
- Hu, J.; Shrestha, S.; Zhou, Y.; Liu, X.; Lamour, K. Dynamic Extreme Aneuploidy (DEA) in the vegetable pathogen Phytophthora capsici sheds light on instant evolution and intractability. BioRxiv 2018, 29778. [Google Scholar] [CrossRef]
- Kasuga, T.; Bui, M.; Bernhardt, E.; Swiecki, T.; Aram, K.; Cano, L.M.; Webber, J.; Brasier, C.; Press, C.; Grünwald, N.J.; et al. Host-induced aneuploidy and phenotypic diversification in the Sudden Oak Death pathogen Phytophthora ramorum. BMC Genom. 2016, 17, 385. [Google Scholar] [CrossRef] [PubMed]
- Mulla, W.A.; Seidel, C.W.; Zhu, J.; Tsai, H.-J.; Smith, S.E.; Singh, P.; Bradford, W.D.; McCroskey, S.; Nelliat, A.R.; Conkright, J.; et al. Aneuploidy as a cause of impaired chromatin silencing and mating-type specification in budding yeast. eLife 2017, 6, e27991. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Yong, M.Y.J.; Yurieva, M.; Srinivasan, K.G.; Liu, J.; Lim, J.S.Y.; Poidinger, M.; Wright, G.D.; Zolezzi, F.; Choi, H.; et al. Gene Essentiality Is a Quantitative Property Linked to Cellular Evolvability. Cell 2015, 163, 1388–1399. [Google Scholar] [CrossRef] [PubMed]
- Beach, R.R.; Ricci-Tam, C.; Brennan, C.M.; Moomau, C.A.; Hsu, P.-H.; Hua, B.; Silberman, R.E.; Springer, M.; Amon, A. Aneuploidy Causes Non-genetic Individuality. Cell 2017, 169, 229–242.e21. [Google Scholar] [CrossRef] [PubMed]
- Stirling, P.C.; Bloom, M.S.; Solanki-Patil, T.; Smith, S.; Sipahimalani, P.; Li, Z.; Kofoed, M.; Ben-Aroya, S.; Myung, K.; Hieter, P. The complete spectrum of yeast chromosome instability genes identifies candidate CIN cancer genes and functional roles for ASTRA complex components. PLoS Genet. 2011, 7, e1002057. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Pavelka, N.; Bradford, W.D.; Rancati, G.; Li, R. Karyotypic determinants of chromosome instability in aneuploid budding yeast. PLoS Genet. 2012, 8, e1002719. [Google Scholar] [CrossRef] [PubMed]
- Duffy, S.; Fam, H.K.; Wang, Y.K.; Styles, E.B.; Kim, J.-H.; Ang, J.S.; Singh, T.; Larionov, V.; Shah, S.P.; Andrews, B.; et al. Overexpression screens identify conserved dosage chromosome instability genes in yeast and human cancer. Proc. Natl. Acad. Sci. USA 2016, 113, 9967–9976. [Google Scholar] [CrossRef]
- Tutaj, H.; Pogoda, E.; Tomala, K.; Korona, R. Gene overexpression screen for chromosome instability in yeast primarily identifies cell cycle progression genes. Curr. Genet. 2019, 65, 483–492. [Google Scholar] [CrossRef]
- Stirling, P.C.; Crisp, M.J.; Basrai, M.A.; Tucker, C.M.; Dunham, M.J.; Spencer, F.A.; Hieter, P. Mutability and mutational spectrum of chromosome transmission fidelity genes. Chromosoma 2012, 121, 263–275. [Google Scholar] [CrossRef]
- Zhu, J.; Heinecke, D.; Mulla, W.A.; Bradford, W.D.; Rubinstein, B.; Box, A.; Haug, J.S.; Li, R. Single-Cell Based Quantitative Assay of Chromosome Transmission Fidelity. G3 2015, 5, 1043–1056. [Google Scholar] [CrossRef] [PubMed]
- Ravichandran, M.C.; Fink, S.; Clarke, M.N.; Hofer, F.C.; Campbell, C.S. Genetic interactions between specific chromosome copy number alterations dictate complex aneuploidy patterns. Genes Dev. 2018, 32, 1485–1498. [Google Scholar] [CrossRef] [PubMed]
- Tange, Y.; Kurabayashi, A.; Goto, B.; Hoe, K.-L.; Kim, D.-U.; Park, H.-O.; Hayles, J.; Chikashige, Y.; Tsutumi, C.; Hiraoka, Y.; et al. The CCR4-NOT complex is implicated in the viability of aneuploid yeasts. PLoS Genet. 2012, 8, e1002776. [Google Scholar] [CrossRef] [PubMed]
- Dodgson, S.E.; Kim, S.; Costanzo, M.; Baryshnikova, A.; Morse, D.L.; Kaiser, C.A.; Boone, C.; Amon, A. Chromosome-Specific and Global Effects of Aneuploidy in Saccharomyces cerevisiae. Genetics 2016, 202, 1395–1409. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Mulla, W.A.; Kucharavy, A.; Tsai, H.-J.; Rubinstein, B.; Conkright, J.; McCroskey, S.; Bradford, W.D.; Weems, L.; Haug, J.S.; et al. Targeting the adaptability of heterogeneous aneuploids. Cell 2015, 160, 771–784. [Google Scholar] [CrossRef] [PubMed]
| Species | Strain Details | Selected Phenotype | Aneuploid Chromosomes | Implicated Genes | Aneuploidy Transience | Reference |
|---|---|---|---|---|---|---|
| C. albicans | Clinical isolates | Fluconazole resistance | V (+1) * | ERG11, TAC1 | Selmecki et al. (2008) [48] | |
| In vivo OPC mouse model | Host defense during OPC | VI (+1) | Forche et al. (2019) [56] | |||
| Laboratory strains | Cisplatin resistance | V (−1), II (+1), II (+2) | Yang et al. (2013) [75], Yang et al. (2019) [76] | |||
| Hydroxyurea resistance | II (+1) | Yang et al. (2019) [76] | ||||
| Suppressor of RGD1 deletion | III (+1) *, VII (+1) | NPR2 | Mount et al. (2018) [77] | |||
| Multiple clinical isolates | Fluconazole resistance | V (+1) *, III (+1) * | ERG11, MRR1, CDR1, CDR2 | Confirmed | Ford et al. (2015) [49] | |
| C. neoformans | Clinical isolates | Fluconazole heteroresistance | I (+1), IV (+1) | ERG11, AFR1 | Sionov et al. (2010) [23], Sionov et al. (2013) [53] | |
| Laboratory strains | Fluconazole heteroresistance | I (+1), IV (+1) | ERG11, AFR1, SEY1, GLO3 | Ngamskulrungroj et al. (2012) [52] | ||
| F. oxysporum | Plant pathogen | Increased pathogenecity | XIV (+1) ** | SIX1, SIX3, ORX1 | Ma et al. (2010) [78] | |
| N. haematococca | Plant pathogen | Increased pathogenicity | CD (+1) ** | PDA6 | Miao et al. (1991) [79] | |
| S. pastorianus | Industrial lager strains | Flocculation | I (+1), IV (+1), X (+1), XII (+2) | LgFLO, FLO1, FLO5, FLO10 | Van den Borek et al. (2015) [80] | |
| Increased diacetyl synthesis | X (+1), XII (+2) | ILV5, ILV3 | Van den Borek et al. (2015) [80] | |||
| S. cerevisiae | Clinical isolates | Host survival | variable across strain phylogeny | Zhu et al. (2016) [55] | ||
| Environmental isolates | Copper tolerance | II (+1), VII (+1), VIII (+1) | CUP1, CUP2, SCO1, SCO2 | Ezov et al. (2006) [41], Chang et al. (2013) [43] | ||
| Freeze-thaw tolerance | XII (+1) | AQY2 | Hose et al. (2015) [38] | |||
| Industrial isolates | Ethanol tolerance | III (+1) | Morard et al. (2019) [81] | |||
| Industrial wine strain | Increased ethanol oxidation | VII (+1), XIII (+2) | ADH2, ADH3 | Guijo et al. (1997) [82] | ||
| Industrial beer strain | Increased ethanol yield | XI (+1) | Zhang et al. (2015) [83] | |||
| Laboratory strains | 4-NQO resistance | IV (+1) | ATR1 | Pavelka et al. (2010) [6] | ||
| Benomyl resistance | XII (−1) | Chen et al. (2012) [61] | ||||
| Copper tolerance | II (+1), VIII (+1) | CUP1, SCO1, SCO2 | Gerstein et al. (2015) [65] | |||
| Ethanol tolerance | III (+1), VIII (+1) | Voordeckers et al. (2015) [68] | ||||
| Ferulic acid tolerance | XIV (+1) | Sato et al. (2014) [63] | ||||
| Flocculation | I (+1) | FLO1 | Hope et al. (2017) [70] | |||
| Fluconazole resistance | VIII (+1) | ERG11 | Chen et al. (2012) [61] | |||
| Galactose tolerance | VIII (+1) | GAL80 | Sirr et al. (2015) [67] | |||
| Glucose-limited growth | I (+1), III (+1), V (+1) *, XIV (−1) | Speculated | Gresham et al. (2008) [59] | |||
| Nitrogen-limited growth (glutamine) | XI, XI (+1–+4) * | GAP1 | Lauer et al. (2018) [62] | |||
| Heat shock tolerance | III (+1) | 17 genes | Confirmed | Yona et al. (2012) [62] | ||
| High pH tolerance | V (+1) | Confirmed | Yona et al. (2012) [62] | |||
| Phosphate-limited growth | IV (+1), VI (+1), X (+1), XIII (+2), XVI (+1) | Speculated | Gresham et al. (2008) [59] | |||
| Raffinose growth | XIII (+1) | Selmecki et al. (2015) [66] | ||||
| Radicicol resistance | XV (+1) | STL1, PDR5 | Chen et al. (2012) [61] | |||
| Suppressors of MEC1 deficiency | IV (+1) | RNR1 | Gasch et al. (2001) [58] | |||
| Suppressors of MYO1 deletion | XIII (+1), XVI (+1) | RLM1, MKK2 | Rancati et al. (2008) [60] | |||
| Suppressors of RPS24A and RNR1 deletion | IX (+1) | RPS24B, RNR3 | Hughes et al. (2000) [57] | |||
| Suppressors of telomerase insufficiency | VIII (−1) | PRP8, UTP9, KOG1, SCH9 | Millet et al. (2016) [69] | |||
| Tunicamycin resistance | XVI (−1), II (+1) | ALG7, PRE7, YBR085C-A | Chen et al. (2012) [61], Beaupere et al. (2018) [71] | |||
| Xylose utilization | I (−1) | Sato et al. (2014) [63] | ||||
| S. paradoxus | Environmental isolates | Freeze-thaw tolerance | XII (+1) | AQY2 | Will et al. (2010) [84] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsai, H.-J.; Nelliat, A. A Double-Edged Sword: Aneuploidy is a Prevalent Strategy in Fungal Adaptation. Genes 2019, 10, 787. https://doi.org/10.3390/genes10100787
Tsai H-J, Nelliat A. A Double-Edged Sword: Aneuploidy is a Prevalent Strategy in Fungal Adaptation. Genes. 2019; 10(10):787. https://doi.org/10.3390/genes10100787
Chicago/Turabian StyleTsai, Hung-Ji, and Anjali Nelliat. 2019. "A Double-Edged Sword: Aneuploidy is a Prevalent Strategy in Fungal Adaptation" Genes 10, no. 10: 787. https://doi.org/10.3390/genes10100787
APA StyleTsai, H.-J., & Nelliat, A. (2019). A Double-Edged Sword: Aneuploidy is a Prevalent Strategy in Fungal Adaptation. Genes, 10(10), 787. https://doi.org/10.3390/genes10100787





