Convergent Evolution of the Seed Shattering Trait
Abstract
1. Introduction
2. Histological Modifications That Underlie Seed Shattering in Crops
3. Genetic Control of Seed Shattering in the Model Species A. thaliana and in Relevant Crops
4. Conclusions
Author Contributions
Funding
References
- Darwin, C.R. On the Origin of Species by Means of Natural Selection; John Murray: London, UK, 1859. [Google Scholar]
- Hammer, K. Das Domestikations syndrom. Die Kulturpflanze 1984, 32, 11–34. [Google Scholar] [CrossRef]
- Gepts, P.; Debouck, D.G. Origin, domestication, and evolution of the common bean (Phaseolus vulgaris L.). In Common Beans: Research for Crop Improvement; Voysest, O., Van Schoonhoven, A., Eds.; CAB: Oxon, UK, 1991; pp. 7–53. [Google Scholar]
- Gu, Q.; Ferrándiz, C.; Yanofsky, M.F.; Martienssen, R. The FRUITFULLMADS-box gene mediates cell differentiation during Arabidopsis fruit development. Development 1998, 125, 1509–1517. [Google Scholar] [PubMed]
- Liljegren, S.J.; Ditta, G.S.; Eshed, Y.; Savidge, B.; Bowman, J.; Yanofsky, M.F. SHATTERPROOF MADS-box genes control seed dispersal in Arabidopsis. Nature 2000, 404, 766–770. [Google Scholar] [CrossRef] [PubMed]
- Rajani, S.; Sundaresan, V. The Arabidopsis myc-bHLH gene ALCATRAZ enables cell separation in fruit dehiscence. Curr. Biol. 2001, 11, 1914–1922. [Google Scholar] [CrossRef]
- Roeder, A.H.K.; Ferrándiz, C.; Yanofsky, M.F. The role of the REPLUMLESS homeodomain protein in patterning the Arabidopsis fruit. Curr. Biol. 2003, 13, 1630–1635. [Google Scholar] [CrossRef] [PubMed]
- Liljegren, S.J.; Roeder, A.H.; Kempin, S.A.; Gremski, K.; Østergaard, L.; Guimil, S.; Reyes, D.K.; Yanofsky, M.F. Control of fruit patterning in Arabidopsis by INDEHISCENT. Cell 2004, 116, 843–853. [Google Scholar] [CrossRef]
- Mitsuda, N.; Ohme-Takagi, M. NAC transcription factors NST1 and NST3 regulate pod dehiscence in a partially redundant manner by promoting secondary wall formation after the establishment of tissue identity. Plant J. 2008, 56, 768–778. [Google Scholar] [CrossRef]
- Konishi, S.; Izawa, T.; Lin, S.Y.; Ebana, K.; Fukuta, Y.; Sasaki, T.; Yano, M.A. SNP caused loss of seed shattering during rice domestication. Science 2006, 312, 1392–1396. [Google Scholar] [CrossRef]
- Li, C.; Zhou, A.; Sang, T. Rice domestication by reducing shattering. Science 2006, 311, 1936–1939. [Google Scholar] [CrossRef]
- Zhou, Y.; Lu, D.; Li, C.; Luo, J.; Zhu, B.F.; Zhu, J.; Shangguan, Y.; Wang, Z.; Sang, T.; Zhou, B.; et al. Genetic control of seed shattering in rice by the APETALA2 transcription factor shattering abortion1. Plant Cell 2012, 24, 1034–1048. [Google Scholar] [CrossRef]
- Yoon, J.; Cho, L.H.; Kim, S.L.; Choi, H.; Koh, H.J.; An, G. The BEL1 type homeobox gene SH5 induces seed shattering by enhancing abscission-zone development and inhibiting lignin biosynthesis. Plant J. 2014, 79, 717–728. [Google Scholar] [CrossRef] [PubMed]
- Pourkheirandish, M.; Hensel, G.; Kilian, B.; Senthil, N.; Chen, G.; Sameri, M.; Azhaguvel, P.; Sakuma, S.; Dhanagond, S.; Sharma, R.; et al. Evolution of the grain dispersal system in barley. Cell 2015, 162, 527–539. [Google Scholar] [CrossRef] [PubMed]
- Funatsuki, H.; Suzuki, M.; Hirose, A.; Inaba, H.; Yamada, T.; Hajika, M.; Komatsu, K.; Katayama, T.; Sayama, T.; Ishimoto, M.; et al. Molecular basis of a shattering resistance boosting global dissemination of soybean. Proc. Natl. Acad. Sci. USA 2014, 111, 17797–17802. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Yang, X.; Liu, J.; Wang, B.H.; Liu, B.L.; Wang, Y.Z. Pod dehiscence resistance associated with domestication is mediated by a NAC gene in soybean. Nat. Commun. 2014, 5, 3352. [Google Scholar] [CrossRef] [PubMed]
- Suanum, W.; Somta, P.; Kongjaimun, A.; Yimran, T.; Kaga, A.; Tomooka, N.; Takahashi, Y.; Srinives, P. Co-localization of QTLs for pod fiber content and pod shattering in F2 and backcross populations between yardlong bean and wild cowpea. Mol. Breed. 2016, 36, 1–11. [Google Scholar] [CrossRef]
- Lo, S.; Muñoz-Amatriaín, M.; Boukar, O.; Herniter, I.; Cisse, N.; Guo, Y.-N.; Roberts, P.A.; Xu, S.; Fatokun, C.; Close, T.J. Identification of QTL controlling domestication-related traits in cowpea (Vigna unguiculata L. Walp). Sci. Rep. 2018, 8, 6261. [Google Scholar] [CrossRef] [PubMed]
- Fourquin, C.; Cerro, C.; Victoria, F.C.; Vialette-Guiraud, A.; Oliveira, A.C.; Ferrandiz, C. A change in SHATTERPROOF protein lies at the origin of a fruit morphological novelty and a new strategy for seed dispersal in Medicago genus. Plant Physiol. 2013, 162, 907–917. [Google Scholar] [CrossRef]
- Koinange, E.M.S.; Singh, S.P.; Gepts, P. Genetic control of the domestication syndrome in common bean. Crop Sci. 1996, 36, 1037–1045. [Google Scholar] [CrossRef]
- Hagerty, C.H.; Cuesta-Marcos, A.; Cregan, P.; Song, Q.; McClean, P.E.; Myers, J.R. Mapping snap bean pod and color traits, in a dry bean × snap bean recombinant inbred population. J. Am. Soc. Hortic. Sci. 2016, 141, 131–138. [Google Scholar]
- Rau, D.; Murgia, M.L.; Rodriguez, M.; Bitocchi, E.; Bellucci, E.; Fois, D.; Albani, D.; Nanni, L.; Gioia, T.; Santo, D.; et al. Genomic dissection of pod shattering in common bean: Mutations at nonorthologous loci at the basis of convergent phenotypic evolution under domestication of leguminous species. Plant J. 2018. [Google Scholar] [CrossRef]
- Nanni, L.; Bitocchi, E.; Bellucci, E.; Rossi, M.; Rau, D.; Attene, G.; Gepts, P.; Papa, R. Nucleotide diversity of a genomic sequence similar to SHATTERPROOF (PvSHP1) in domesticated and wild common bean (Phaseolus vulgaris L.). Theor. Appl. Genet. 2011, 123, 1341–1357. [Google Scholar] [CrossRef] [PubMed]
- Gioia, T.; Logozzo, G.; Kami, J.; Spagnoletti Zeuli, P.; Gepts, P. Identification and characterization of a homolog to the Arabidopsis INDEHISCENT gene in common bean. J. Hered. 2013, 104, 273–286. [Google Scholar] [CrossRef] [PubMed]
- Hofhuis, H.; Moulton, D.; Lessinnes, T.; Routier-Kierzkowska, A.-L.; Bomphrey, R.J.; Mosca, G.; Reinhardt, H.; Sarchet, P.; Gan, X.; Tsiantis, M.; et al. Morphomechanical innovation drives explosive seed dispersal. Cell 2016, 166, 222–233. [Google Scholar] [CrossRef] [PubMed]
- Prakken, R. Inheritance of colours and pod characters in Phaseolus vulgaris L. Genetica 1934, 16, 177–296. [Google Scholar] [CrossRef]
- Murgia, M.L.; Attene, G.; Rodriguez, M.; Bitocchi, E.; Bellucci, E.; Fois, D.; Nanni, L.; Gioia, T.; Albani, D.M.; Papa, R.; et al. A comprehensive phenotypic investigation of the “pod-shattering syndrome” in common bean. Front. Plant Sci. 2017, 8, 251. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, M.; Kay, P.; Wilson, S.; Swain, S.M. ARABIDOPSIS DEHISCENCE ZONE POLYGALACTURONASE1 (ADPG1), ADPG2, and QUARTET2 are polygalacturonases required for cell separation during reproductive development in Arabidopsis. Plant Cell 2009, 21, 216–233. [Google Scholar] [CrossRef] [PubMed]
- Ripoll, J.J.; Roeder, A.H.; Ditta, G.S.; Yanofsky, M.F. A novel role for the floral homeotic gene APETALA2 during Arabidopsis fruit development. Development 2011, 138, 5167–5176. [Google Scholar] [CrossRef]
- Mao, L.; Begum, D.; Chuang, H.; Budlman, M.A.; Szymkowiak, E.J.; Irish, E.E.; Wing, R.A. JOINTLESS is a MADS-box gene controlling tomato flower abscission zone development. Nature 2000, 406, 910–913. [Google Scholar] [CrossRef]
- Vrebalov, J.; Pan, I.L.; Arroyo, A.J.; McQuinn, R.; Chung, M.; Poole, M.; Rose, J.; Seymour, G.; Grandillo, S.; Giovannoni, J.; et al. Fleshy fruit expansion and ripening are regulated by the tomato SHATTERPROOF gene TAGL1. Plant Cell 2009, 21, 3041–3062. [Google Scholar] [CrossRef]
- Nakano, T.; Kimbara, J.; Fujisawa, M.; Kitagawa, M.; Ihashi, N.; Maeda, H.; Kasumi, T.; Ito, Y. MACROCALYX and JOINTLESS interact in the transcriptional regulation of tomato fruit abscission zone development. Plant Physiol. 2012, 158, 439–450. [Google Scholar] [CrossRef]
- Lin, Z.; Griffith, M.E.; Li, X.; Zhu, Z.; Tan, L.; Fu, Y.; Zhang, W.; Wang, X.; Xie, D.; Sun, C. Origin of seed shattering in rice (Oryza sativa L.). Planta 2007, 226, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Liu, X.; Wang, M.; Meyer, R.S.; Luo, X.; Ndjiondjop, M.; Tan, L.; Zhang, J.; Wu, J.; Cai, H.; et al. A single-nucleotide polymorphism causes smaller grain size and loss of seed shattering during African rice domestication. Nat. Plants 2017, 3, 17064. [Google Scholar] [CrossRef] [PubMed]
- Lv, S.; Wu, W.; Wang, M.; Meyer, R.S.; Ndjiondjop, M.; Tan, L.; Zhou, H.; Zhang, J.; Fu, Y.; Cai, H.; et al. Genetic control of seed shattering during African rice domestication. Nat. Plants 2018, 4, 331–337. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Li, X.; Shannon, L.M.; Yeh, C.T.; Wang, M.L.; Bai, G.; Peng, Z.; Li, J.; Trick, H.N.; Clemente, T.E.; et al. Parallel domestication of the Shattering1 genes in cereals. Nat. Genet. 2012, 44, 720–724. [Google Scholar] [CrossRef] [PubMed]
- Ferrándiz, C. Regulation of fruit dehiscence in Arabidopsis. J. Exp. Bot. 2002, 53, 2031–2038. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Wang, Y.-Z. Seed shattering: From models to crops. Front. Plant Sci. 2015, 6, 476. [Google Scholar] [CrossRef] [PubMed]
- Ferrándiz, C.; Liljegren, S.J.; Yanofsky, M.F. Negative regulation of the SHATTERPROOF genes by FRUITFULL during Arabidopsis fruit development. Science 2000, 289, 436–438. [Google Scholar] [CrossRef]
- Sorefan, K.; Girin, T.; Liljegren, S.J.; Ljung, K.; Robles, P.; Galvan-Ampudia, C.S.; Offringa, R.; Friml, J.; Yanofsky, M.F.; Østergaard, L. A regulated auxin minimum is required for seed dispersal in Arabidopsis. Nature 2009, 459, 583–586. [Google Scholar] [CrossRef]
- Girin, T.; Paicu, T.; Stephenson, P.; Fuentes, S.; Korner, E.; O’Brien, M.; Sorefan, K.; Wood, T.A.; Balanzá, V.; Ferrándiz, C.; et al. INDEHISCENT and SPATULA interact to specify carpel and valve margin tissue and thus promote seed dispersal in Arabidopsis. Plant Cell 2011, 23, 3641–3653. [Google Scholar] [CrossRef]
- Marsch-Martinez, N.; Ramos-Cruz, D.; Reyes-Olalde, J.; Lozano-Sotomayor, P.; Zuniga Mayo, V.M.; Folter, S. The role of cytokinin during Arabidopsis gynoecia and fruit morphogenesis and patterning. Plant J. 2012, 72, 222–234. [Google Scholar] [CrossRef]
- Shima, Y.; Kitagawa, M.; Fujisawa, M.; Nakano, T.; Kato, H.; Kimbara, J.; Kasumi, T.; Ito, Y. Tomato FRUITFULL homologues actin fruit ripening via forming MADS-box transcription factor complexes with RIN. Plant Mol. Biol. 2013, 82, 427–438. [Google Scholar] [CrossRef] [PubMed]
- Bemer, M.; Karlova, R.; Ballester, A.R.; Tikunov, Y.M.; Bovy, A.G.; Wolters-Arts, M.; Rossetto Pde, B.; Angenent, G.C.; de Maagd, R.A. The tomato FRUITFULL homologs TDR4/FUL1 and MBP7/FUL2 regulate ethylene independent aspects of fruit ripening. Plant Cell 2012, 24, 4437–4451. [Google Scholar] [CrossRef] [PubMed]
- Vrebalov, J.; Ruezinsky, D.; Padmanabhan, V.; White, R.; Medrano, D.; Drake, R.; Schuch, W.; Giovannoni, J. A MADS-box gene necessary for fruit ripening at the tomato ripening inhibitor (rin) locus. Science 2002, 296, 343–346. [Google Scholar] [CrossRef] [PubMed]
- Lamprecht, H. Beitrage zur genetic von Phaseolus vulgaris. Hereditas 1932, 16, 169–211. [Google Scholar] [CrossRef]
- Ballester, P.; Ferràndiz, C. Shattering fruits: Variations on a dehiscent theme. Curr. Opin. Plant Biol. 2017, 35, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Bitocchi, E.; Rau, D.; Bellucci, E.; Rodriguez, M.; Murgia, M.L.; Gioia, T.; Santo, D.; Nanni, L.; Attene, G.; Papa, R. Beans (Phaseolus ssp.) as a Model for Understanding Crop Evolution. Front. Plant Sci. 2017, 8, 722. [Google Scholar] [CrossRef] [PubMed]
- Bitocchi, E.; Bellucci, E.; Giardini, A.; Rau, D.; Rodriguez, M.; Biagetti, E.; Santilocchi, R.; Spagnoletti Zeuli, P.; Gioia, T.; Logozzo, G.; et al. Molecular analysis of the parallel domestication of the common bean (Phaseolus vulgaris) in Mesoamerica and the Andes. New Phytol. 2013, 197, 300–313. [Google Scholar] [CrossRef] [PubMed]
- McClean, P.E.; Bett, K.; Stonehouse, R.; Lee, R.; Pflieger, S.; Moghaddam, S.M.; Geffroy, V.; Miklas, P.; Mamidi, S. White seed color in common bean (Phaseolus vulgaris) results from convergent evolution in the P (pigment) gene. New Phytol. 2018, 219, 1112–1123. [Google Scholar] [CrossRef]
| Species | Histological Modification | Observed Phenotype | Reference |
|---|---|---|---|
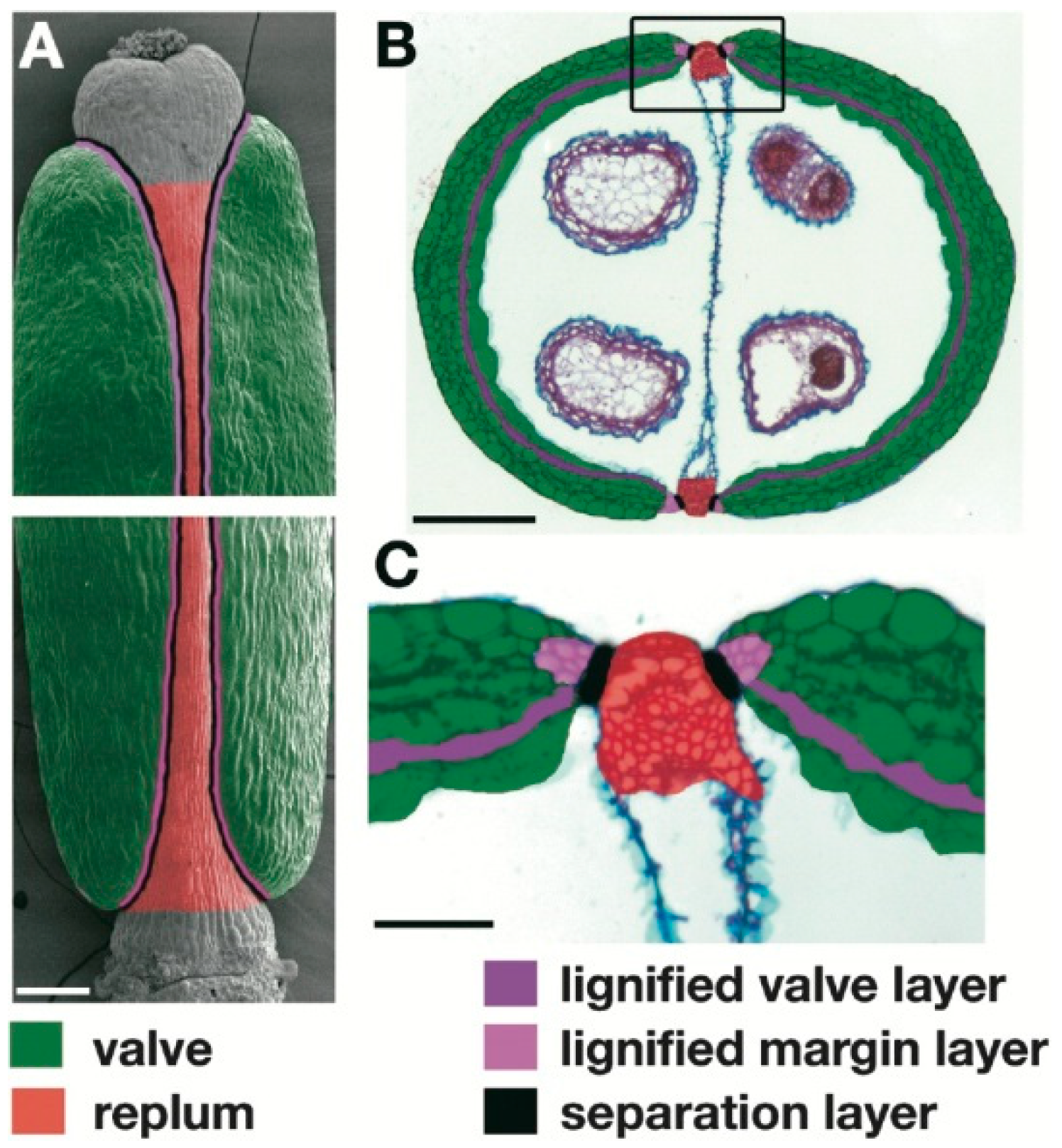
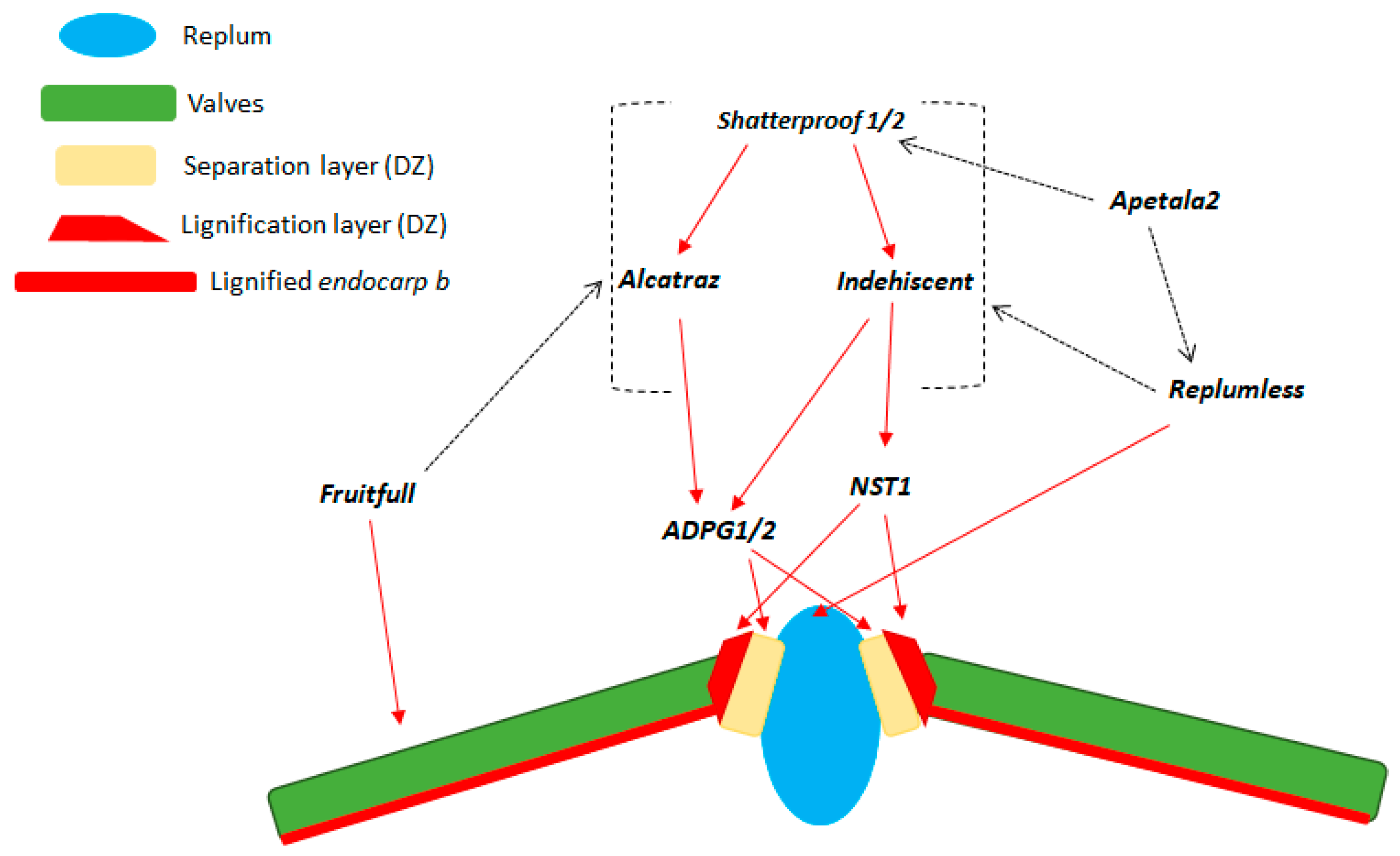
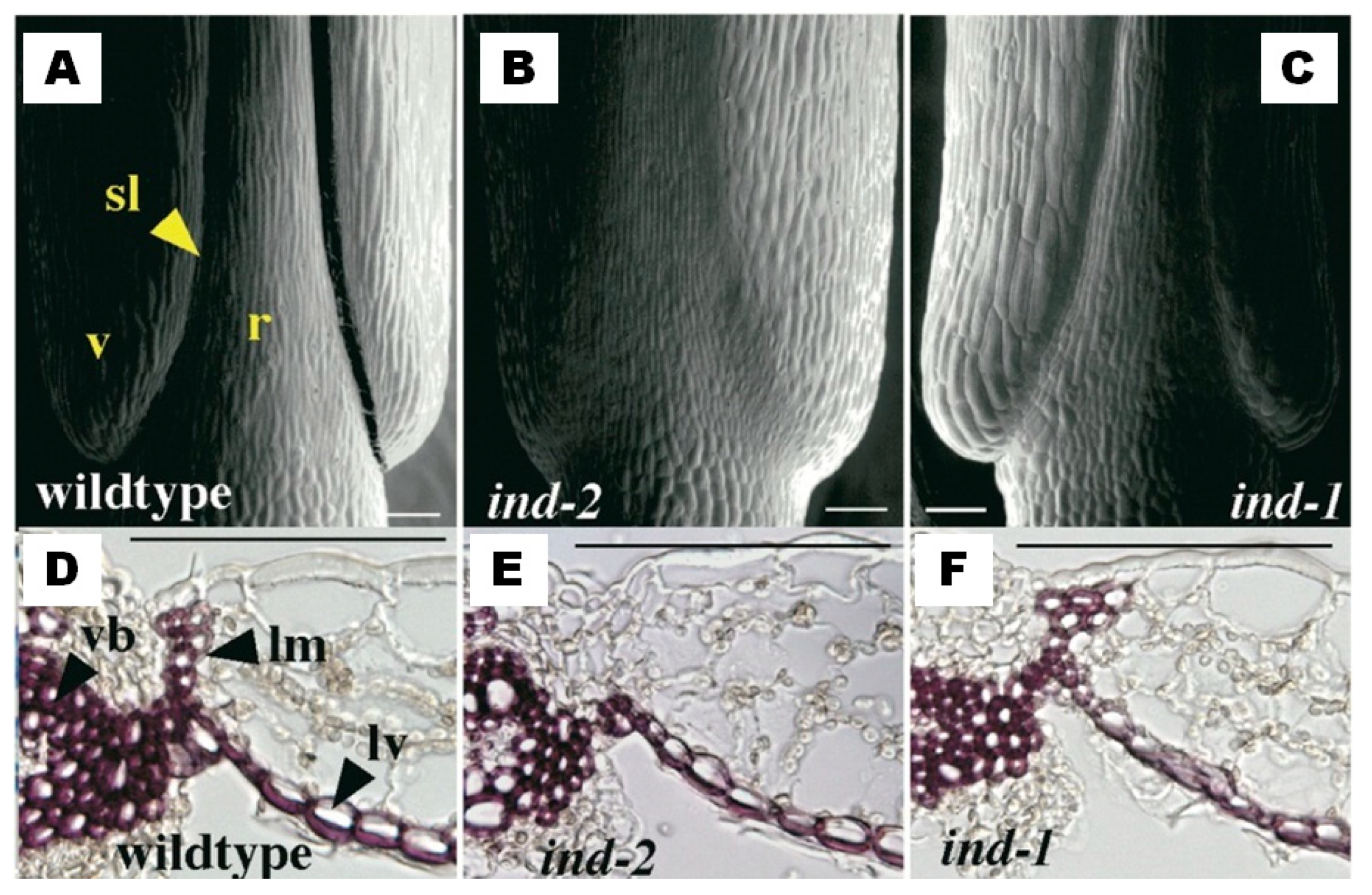
| A. thaliana | Lignification of the valve margin cells | Increase of the mechanical tension at the valve margin in the dry silique, and seed shattering | Liljegren et al. [5] Liljegren et al. [8] |
| A. thaliana | Presence of a functional abscission layer (i.e., separation layer) between valves and replum | Separation of the valves from the replum in the dry silique, and seed shattering | Rajani and Sundaresan [6] |
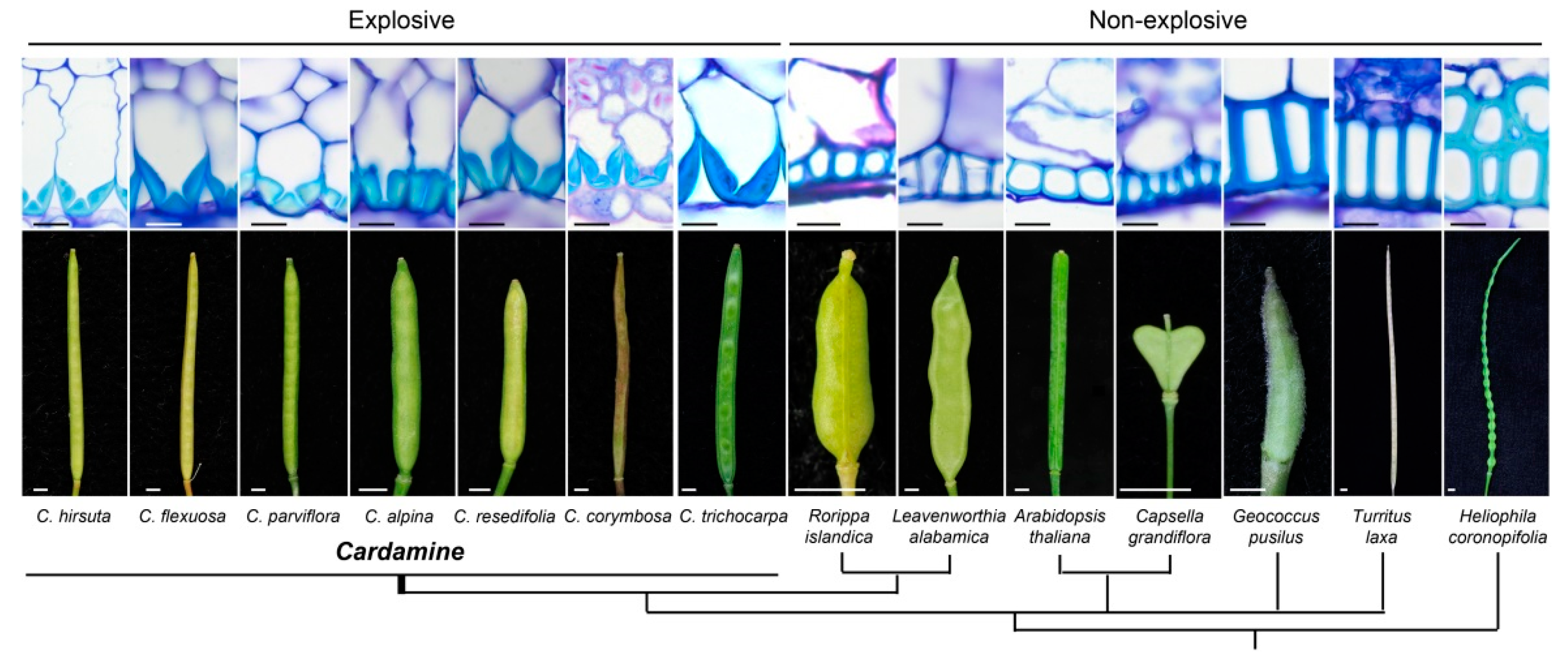
| C. hirsuta | Asymmetric lignin deposition in the endocarp b cell walls (internal valve layer) | Explosive seed shattering and valve curling | Hofhuis et al. [25] |
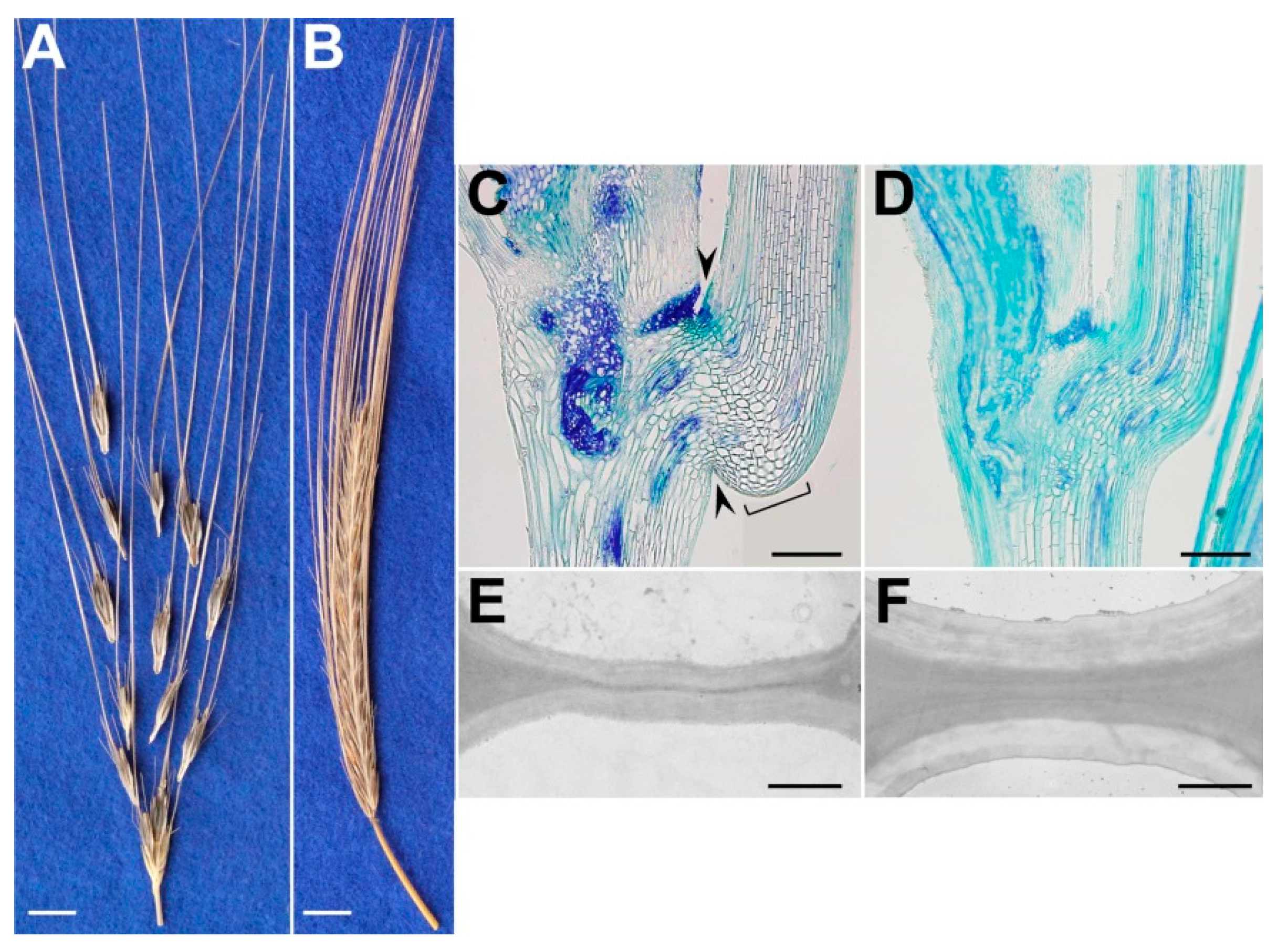
| H. spontaneum; H. vulgare | Reduced cell-wall thickness of cells of the separation layer/Cell wall thickening of cells of the separation layer | Brittle-rachis (disarticulation of the spikelet from the rachis)/Non-brittle rachis | Pourkheirandish et al. [14] |
| O. nivara; O. sativa | Presence of a continuous abscission cell layer at the junction between the pedicel and the flower/discontinuous or absent separation layer | Detachment of the grain from the pedicel and easy threshing of the panicle/reduction or loss of seed shattering | Konishi et al. [10] Li et al. [11] Zhou et al. [12] Yoon et al. [13] |
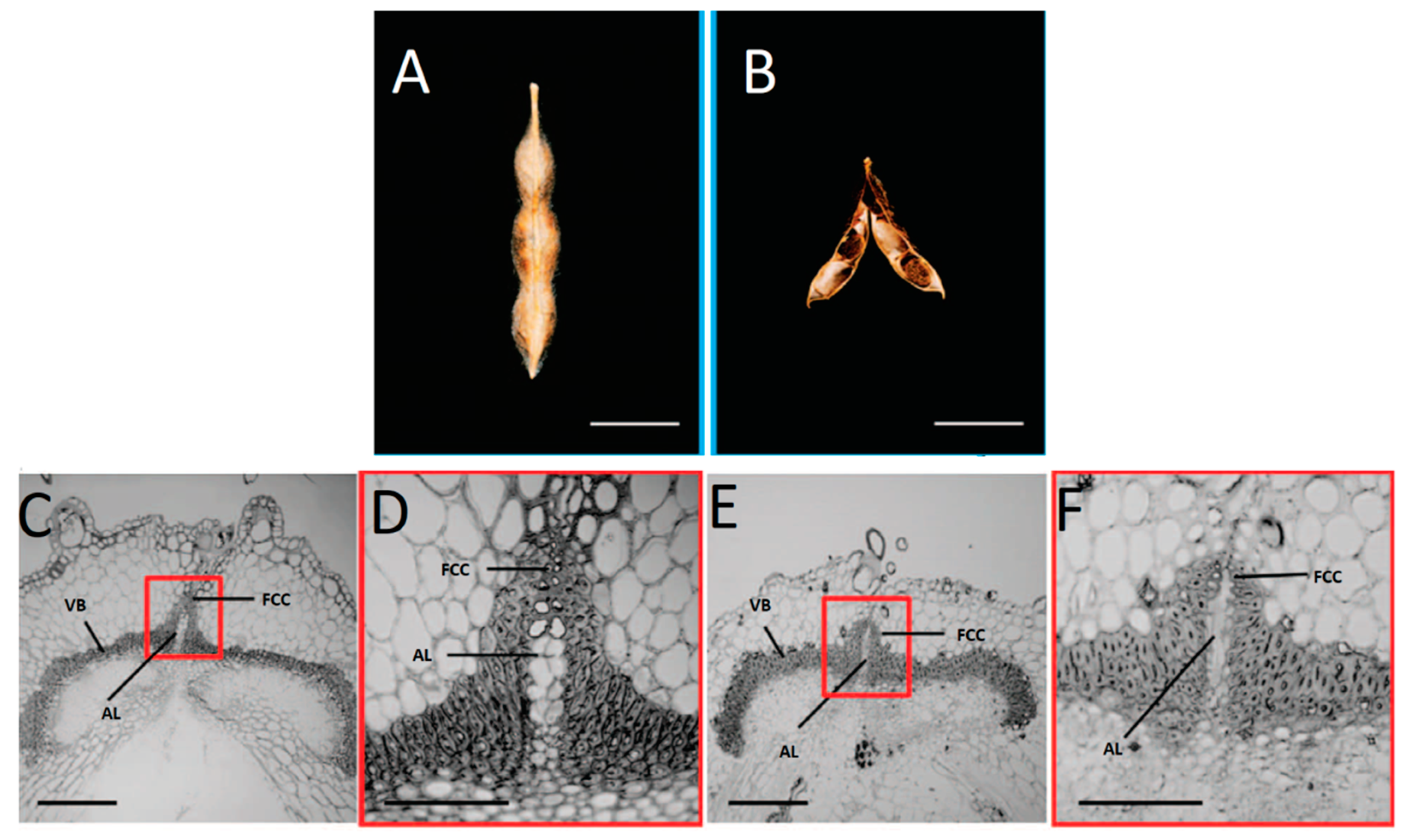
| G.max | Secondary cell-wall thickening in the fiber cap cells of the ventral suture (dehiscence zone) | Resistance to pod shattering | Dong et al. [16] |
| G. soja; G. max | Presence of an internal lignified valve layer | Increase of pod shattering | Funatsuki et al. [15] |
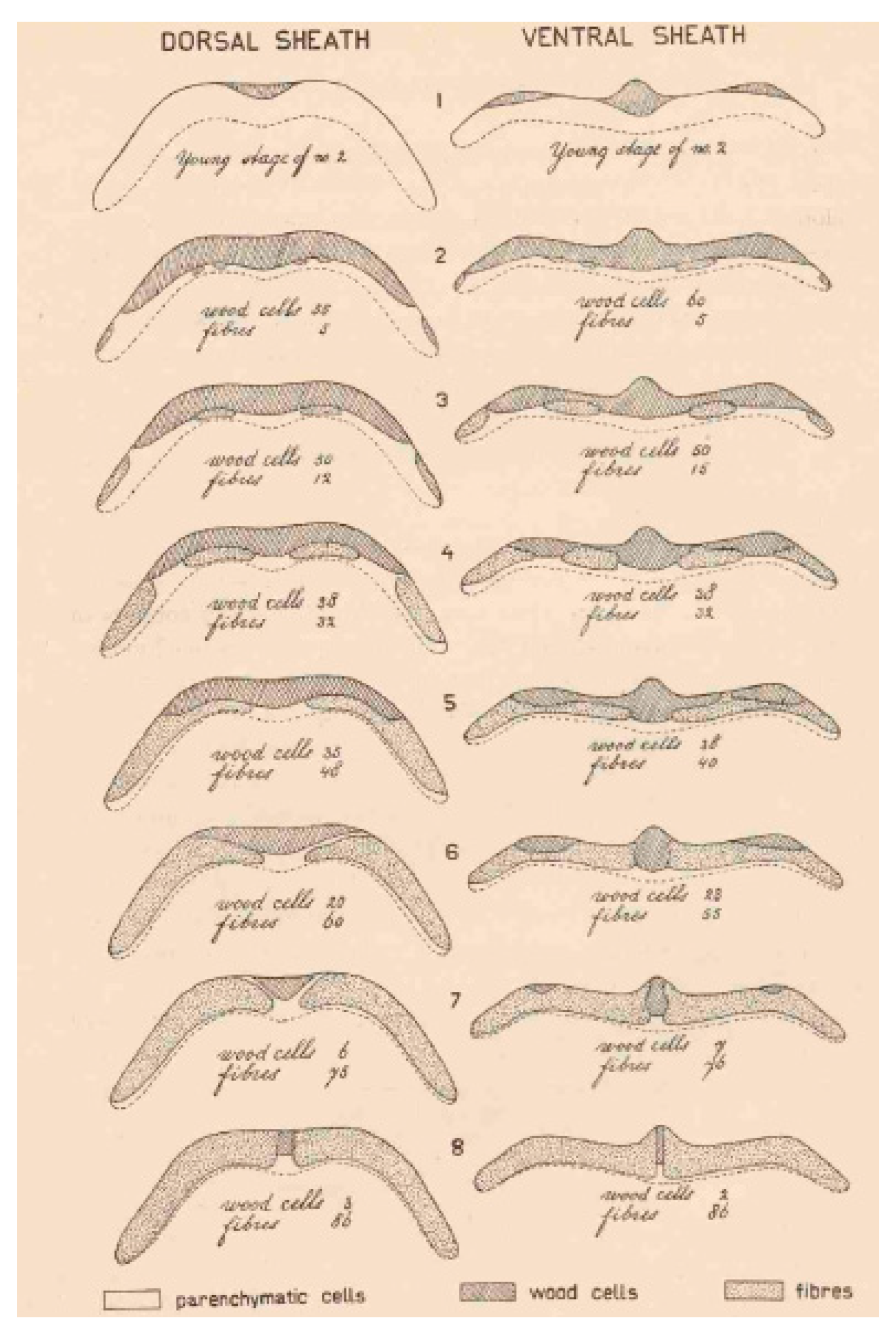
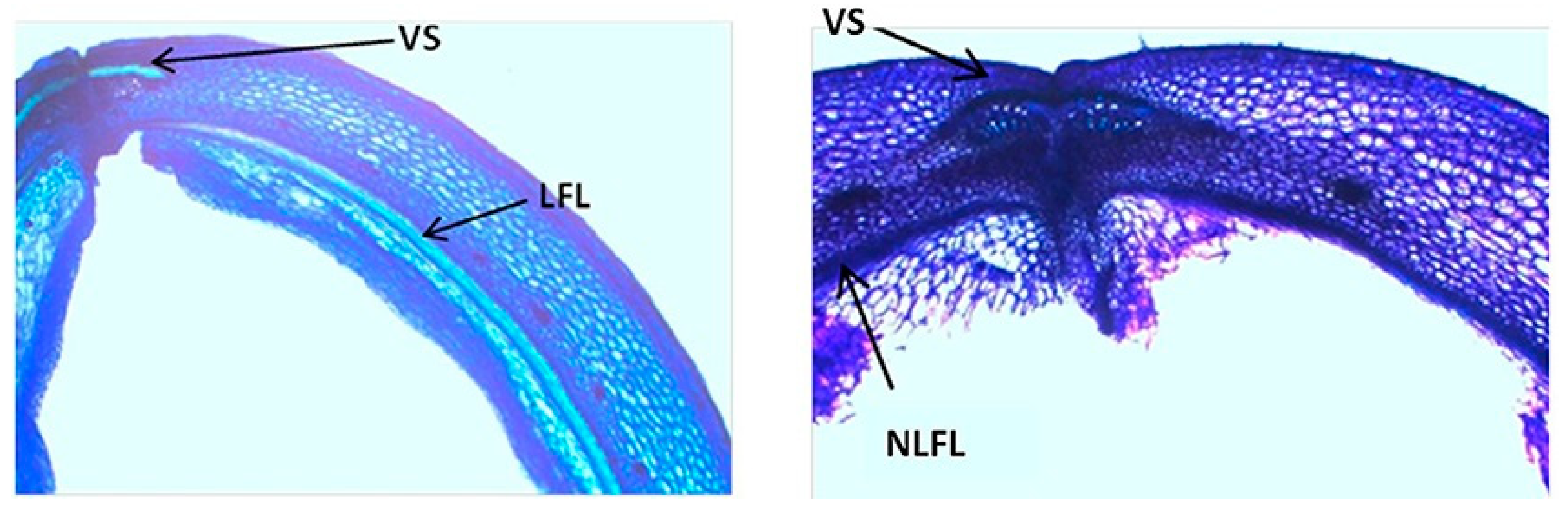
| P. vulgaris | Strong lignin deposition in the cells that surround the dehiscence zone | Increase of pod shattering | Murgia et al. [27] |
| P. vulgaris | Presence of an internal lignified valve layer | Increase of pod shattering | Murgia et al. [27] |
| Species | Gene | Chromosome | Notes/Other Names | Reference |
|---|---|---|---|---|
| A. thaliana | AT3G58780 | 3 | Shatterproof-1 | Liljegren et al. [5] |
| A. thaliana | AT2G42830 | 2 | Shatterproof-2 | Liljegren et al. [5] |
| A. thaliana | AT4G00120 | 4 | INDEHISCENT | Liljegren et al. [8] |
| A. thaliana | AT5G67110 | 5 | ALCATRAZ | Rajani and Sundaresan [6] |
| A. thaliana | AT5G60910 | 5 | FRUITFULL, AGL8 | Gu et al. [4] |
| A. thaliana | AT5G02030 | 5 | REPLUMLESS | Roeder et al. [7] |
| A. thaliana | AT1G32770 | 1 | NST3, SND1 | Mitsuda and Ohme-Takagi [9] |
| A. thaliana | AT2G46770 | 2 | NST1 | Mitsuda and Ohme-Takagi [9] |
| A. thaliana | AT3G57510 | 3 | ADPG1 | Ogawa et al. [28] |
| A. thaliana | AT2G41850 | 2 | ADPG2 | Ogawa et al. [28] |
| A. thaliana | AT4G36920.1 | 4 | APETALA 2 | Ripoll et al. [29] |
| G. max | Glyma.16G019400 | 16 | Shatt 1-5 | Dong et al. [16] |
| G. max | Glyma.16G141400 | 16 | PDH1 | Funatsuki et al. [15] |
| S. lycopersicum | Solyc11g010570 | 11 | JOINTLESS | Mao et al. [30] |
| S. lycopersicum | Solyc02g071730 | 2 | TAGL1 | Vrebalov et al. [31] |
| S. lycopersicum | Solyc05g056620 | 5 | MACROCALYX | Nakano et al. [32] |
| O. sativa | LOC_Os01g62920 | 1 | qSH1 | Konishi et al. [10] |
| O. sativa | LOC_Os05g38120 | 5 | SH5 | Yoon et al. [13] |
| O. sativa | LOC_Os04g55560 | 4 | Shattering Abortion1, SHAT1 | Zhou et al. [12] |
| O. sativa | LOC_Os04g57530 | 4 | Shattering4, SHA1 | Li et al. [11], Lin et al. [33] |
| O. glaberrima | ORGLA04G0254300 | 4 | OgSH4 | Wu et al. [34] |
| O. barthii | 3 | ObSH3 | Lv et al. [35] | |
| S. bicolor | Sobic.001G152901 | 1 | Sh1 | Lin et al. [36] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Di Vittori, V.; Gioia, T.; Rodriguez, M.; Bellucci, E.; Bitocchi, E.; Nanni, L.; Attene, G.; Rau, D.; Papa, R. Convergent Evolution of the Seed Shattering Trait. Genes 2019, 10, 68. https://doi.org/10.3390/genes10010068
Di Vittori V, Gioia T, Rodriguez M, Bellucci E, Bitocchi E, Nanni L, Attene G, Rau D, Papa R. Convergent Evolution of the Seed Shattering Trait. Genes. 2019; 10(1):68. https://doi.org/10.3390/genes10010068
Chicago/Turabian StyleDi Vittori, Valerio, Tania Gioia, Monica Rodriguez, Elisa Bellucci, Elena Bitocchi, Laura Nanni, Giovanna Attene, Domenico Rau, and Roberto Papa. 2019. "Convergent Evolution of the Seed Shattering Trait" Genes 10, no. 1: 68. https://doi.org/10.3390/genes10010068
APA StyleDi Vittori, V., Gioia, T., Rodriguez, M., Bellucci, E., Bitocchi, E., Nanni, L., Attene, G., Rau, D., & Papa, R. (2019). Convergent Evolution of the Seed Shattering Trait. Genes, 10(1), 68. https://doi.org/10.3390/genes10010068








