1. Introduction
T-cell acute lymphoblastic leukemia (T-ALL) results from the malignant transformation of T-cell progenitors that cause diffuse infiltration of the bone marrow by immature T-cell lymphoblasts.
High-dose multi-agent chemotherapy regimens can cure 80% of pediatric patients, but at least 20% of pediatric and 40% of adult T-ALL patients eventually relapse, leading to poor prognosis. Unfortunately, the specific mechanisms mediating escape from therapy, disease progression, and leukemia relapse remain largely unknown [
1].
T-cell transformation is characterized by an accumulation of genomic alterations that generate oncogenes and inactivate tumor suppressor genes, contributing to uncontrolled cell proliferation and cell cycle progression, differentiation arrest, and abnormal cellular metabolism. A central role in T-cell transformation is played by NOTCH1, which is a key transmembrane transcription factor involved in T-cell fate specification and development and is activated in over 60% of cases [
2]. Upon interaction with its ligands, NOTCH1 undergoes a conformational change that leads to two sequential proteolytic cleavages, first by an ADAM metalloprotease and subsequently by the γ-secretase complex. This final cleavage releases the intracellular domains of NOTCH1 (ICN1) from the membrane, allowing its translocation to the nucleus, where it activates gene expression via association with CBF1, Suppressor of Hairless, and Lag-1 (CSL). Patients carrying mutations in the
NOTCH1 gene constitutively express high levels of ICN1. The NOTCH1 oncogenic program can be therapeutically targeted by small-molecule γ-secretase inhibitors (GSIs), which effectively block NOTCH1 activation via the inhibition of a critical intramembrane proteolytic cleavage that is required for NOTCH1 signaling, making NOTCH1 signaling an important therapeutic target in T-ALL. During T-cell transformation, high levels of activated Notch1 in murine T-cell progenitor models impair T-cell maturation, leading to the accumulation of CD4pos/CD8pos cells, promote thymic-independent T-cell development, and ultimately lead to T-cell leukemia [
3]. In fact, Notch directly regulates pre-T-cell antigen receptor α (
Ptcra), interleukin 7 receptor (
Il7r), and
Igf1r genes [
4,
5,
6]. Moreover, NOTCH1 directly upregulates genes that control anabolic metabolism, including those involved in biosynthesis, protein translation, and nucleotide and amino acid metabolism, mainly through direct transcriptional regulation of the
MYC oncogene [
7,
8].
Non-coding RNAs (ncRNAs) have emerged as crucial players in post-transcriptional gene regulation. Among the ncRNAs are microRNAs (miRNAs), which control target mRNAs through degradation or translational repression and are reported to regulate different biological processes, including development, differentiation, and cancer [
9,
10]. Recently, miRNAs that may play critical roles in the NOTCH signaling pathway have been identified using different approaches, from genetic screens to miRNA profiling, by comparing normal T-cell subsets with NOTCH1-driven leukemia [
11,
12,
13,
14]. However, little is currently known about miRNAs that are regulated in therapeutic contexts, such as NOTCH1 blockage with gamma-secretase inhibitors. Using T-ALL cell lines and inhibiting NOTCH1 in vitro, Guascott et al. identified only a few miRNAs, mainly because of the heterogeneity among the analyzed cell lines [
15]. In our study, we took advantage of a mouse model of NOTCH1-induced T-cell leukemia that is strictly dependent on this oncogene and performed in vivo NOTCH1 inhibition using a gamma-secretase inhibitor. This analysis allowed us to identify novel miRNAs that may act in concert with NOTCH1 to play a role in in vivo T-ALL progression. We focused our research on miR-22-3p, one of the most significantly modulated miRNAs whose function in T-ALL is still ill defined.
2. Materials and Methods
2.1. Mouse Models of NOTCH1-Induced T-ALL
As previously reported [
3,
16], retrovirus-mediated overexpression of activated NOTCH1 alleles in hematopoietic lineage-negative progenitors induces primarily ectopic T-cell development and secondary T-cell leukemia. Different
NOTCH1 alleles can recapitulate T-cell leukemia in the mouse: the HD-ΔPEST allele contains a mutation in the HD (heterodimerization) domain (L1601P) and a deletion in the PEST (proline (P), glutamic acid (E), serine (S), and threonine (T)) domain (ΔPEST) that closely resembles a human
NOTCH1 mutation, and the ΔE allele presents a truncated NOTCH1 that resembles NOTCH1 translocation found in about 1–3% of patients. Both alleles are sensitive to gamma-secretase inhibitors.
We generated NOTCH1-induced tumors using both HD-ΔPEST and ΔE alleles, as previously described [
16,
17]. Tumor-bearing mice were euthanized, and primary tumor cells were extracted from their spleens. These tumor cells were then re-injected into sub-lethally irradiated mice (4 Gy) to generate secondary NOTCH1-induced T-ALL tumors. When these mice showed signs of leukemia development, groups of mice were randomized and injected intraperitoneally (i.p.) with three doses of dibenzazepine (DBZ) (5 mg/kg), which is a potent GSI, or Dimethyl sulfoxide (DMSO, vehicle) at 8 h intervals. Each experimental group consisted of at least three animals. After this treatment, mice were sacrificed, and T-leukemia cells were isolated from infiltrated spleens to perform molecular analyses. Procedures involving animals and their care conformed with institutional guidelines that comply with national and international laws and policies (EEC Council Directive 86/609, OJ L 358, 12 206 December 1987). All mice were monitored daily, and animals showing overt signs of disease or excessive weight loss were euthanized following Institutional Animal Care and Use Committee guidelines. These experiments were authorized by the Italian Ministry of Health (authorization n° 136/2014 released the 28th of March 2014).
2.2. Microarray Experiments and Bioinformatic Analyses
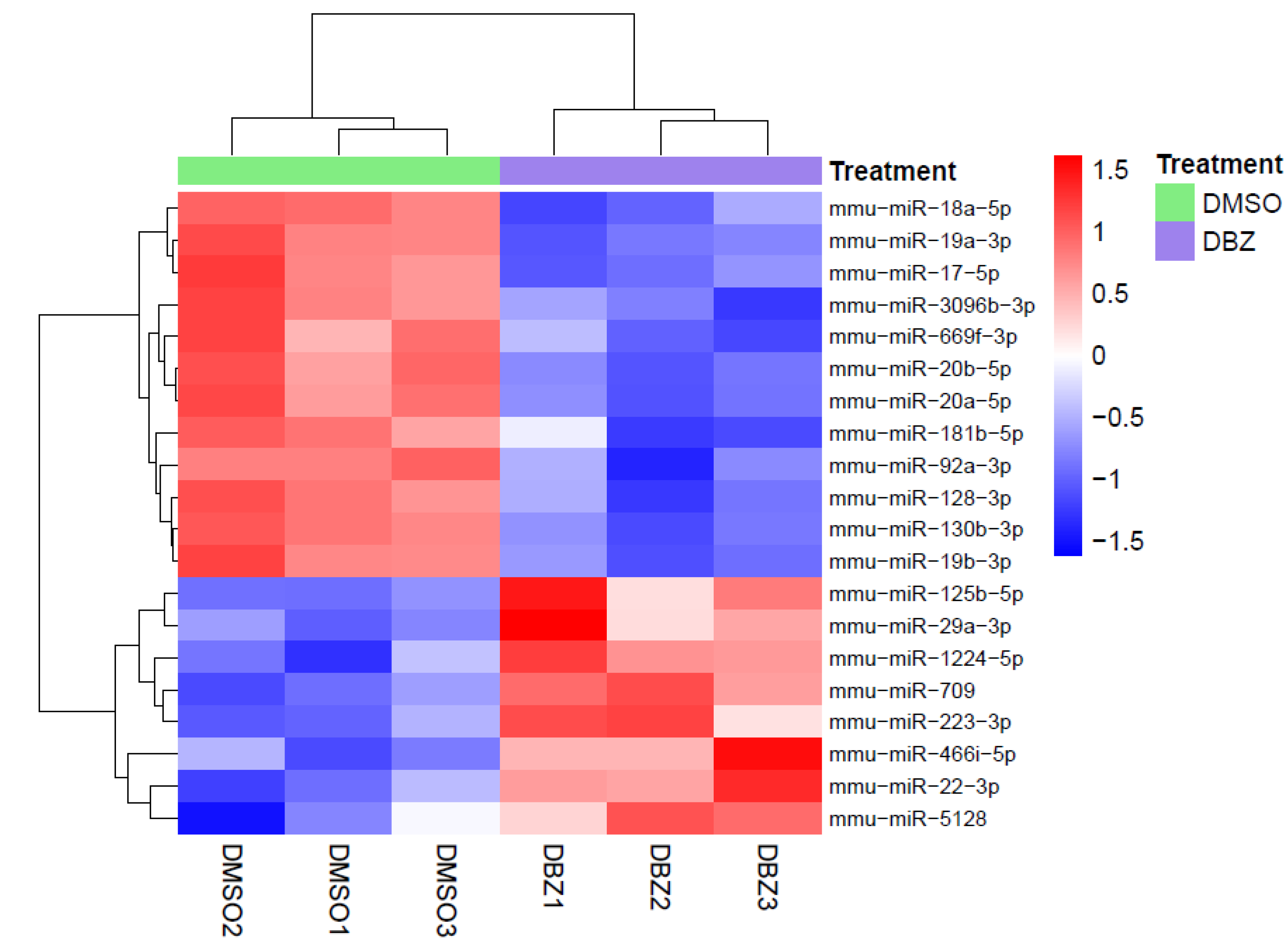
Total RNA from the spleens of NOTCH1 (HD-∆PEST)-induced T-leukemia-bearing mice treated with DBZ (n = 3) or DMSO (n = 3) was extracted using TRIzol according to the manufacturer’s instructions. RNA quality and purity were assessed using an Agilent Bioanalyzer 2100 (Agilent Technologies, Waldbronn, Germany) at CRIBI Biotechnology Center (University of Padova). Only RNA samples that passed quality control were used to perform gene expression profiling, which was carried out using SurePrint G3 Mouse GE 8 × 60 K (Agilent Technologies). MiRNA expression profiling was performed using mouse miRNA 8 × 60 K release 19.0 (Agilent Technologies). Microarray slides were scanned using an Agilent microarray scanner system (G2505C for gene and G2565CA for miRNA expression) following the manufacturer’s instructions. Feature Extraction software version 10.7.3.1 (Agilent Technologies) (including the optimal grid file, 028005_D_F_20100804 for gene and 046065_D_F_20121223 for miRNA expression microarrays) was used with the recommended settings to quantify hybridization signals and produce quality control reports and raw data for bioinformatic analyses.
Bioinformatic analysis of gene expression microarray data was performed in the R/Bioconductor statistical environment using the limma package. Data were preprocessed using the ‘Normexp’ background correction method with an offset of 16, and normalization between arrays was executed with the quantile method. Differential expression analysis was performed with a linear model, and gene-wide moderated t-statistics were obtained by empirical Bayes shrinkage. The Benjamini–Hochberg method was used to correct for multiple testing using a strict false discovery rate (FDR) cut-off of 0.01. Gene set enrichment analysis (GSEA) was performed to evaluate the functional significance of curated gene sets. Genes were ranked by decreasing moderated t-statistics, and GSEAPreranked was run with default parameters. Gene sets in the H collection of the Molecular Signatures Database (MSigDB) v7.0, consisting of 50 hallmark gene sets, were tested for significance.
For miRNA analysis, raw data were loaded and preprocessed in the R statistical environment using the AgiMicroRna Bioconductor library. Preprocessing was performed using the robust multi-array average (RMA) algorithm to yield a summary measure of the miRNA expression using a linear model that accounts for the probe affinity effect. As suggested by López-Romero [
18,
19], background correction was omitted as it can increase the false-positive detection of fold changes in low-expressed miRNAs. Differential expression analysis was performed using the limma Bioconductor package, followed by multiple testing correction using the Benjamini–Hochberg method. MiRNAs with absolute fold change (FC) > 1.3 and FDR < 0.05 were considered differentially expressed.
Transcriptome profiling of the MOLT4 cell line overexpressing miR-22-3p (n = 3) or the empty vector (n = 3) was carried out using the human Clariom™ S Assay (ThermoFisher Scientific, Waltham, MA, USA). Microarray data were analyzed in the R/Bioconductor statistical environment using the RMA algorithm for normalization and the limma package for differential expression analysis. Gene set enrichment analysis was performed to test the significance of gene ontology (GO) gene sets in the MSigDB v7.0 C5 collection corresponding to the molecular function (MF) and biological process (BP) ontologies.
Raw microarray data, together with the description of experiments, protocols and results of differential expression analysis, have been deposited in the ArrayExpress database (
www.ebi.ac.uk/arrayexpress) under accession numbers E-MTAB-9278 (mouse miRNA arrays), E-MTAB-9279 (mouse gene expression arrays) and E-MTAB-9280 (human gene expression arrays).
2.3. In Vivo Experiments with Xenografts
Primary T-ALL samples were obtained at the time of diagnosis with informed consent from the Oncohematology Laboratory, Department of Woman and Child Health, University of Padova, according to the guidelines of the local ethics committee. Patient-derived xenografts (PDX) were previously established and characterized for
NOTCH1 and
FBXW7 mutations by Dr. Stefano Indraccolo at the Veneto Institute of Oncology, IRCCS, Padua [
20,
21]. PDX cells were thawed, and 10 × 10
6 viable T-ALL cells (in 400 µL of PBS) were injected intravenously (i.v.) in 6–8-week-old NOD.Cg-
Prkdcscid Il2rgtm1Wjl/SzJ (NSG) immunodeficient mice. The degree of T-ALL engraftment in mice was monitored by periodic blood collection and flow cytometric analysis of human CD45. When mice showed signs of leukemia development, they were randomized and injected intraperitoneally (i.p.) three times, 8 h apart, with either DBZ (5 mg/kg) or DMSO (vehicle). Each experimental group consisted of at least
n = 3 mice. After this treatment, mice were sacrificed, and T-leukemia cells were isolated from fully infiltrated spleens to obtain RNA and protein lysates.
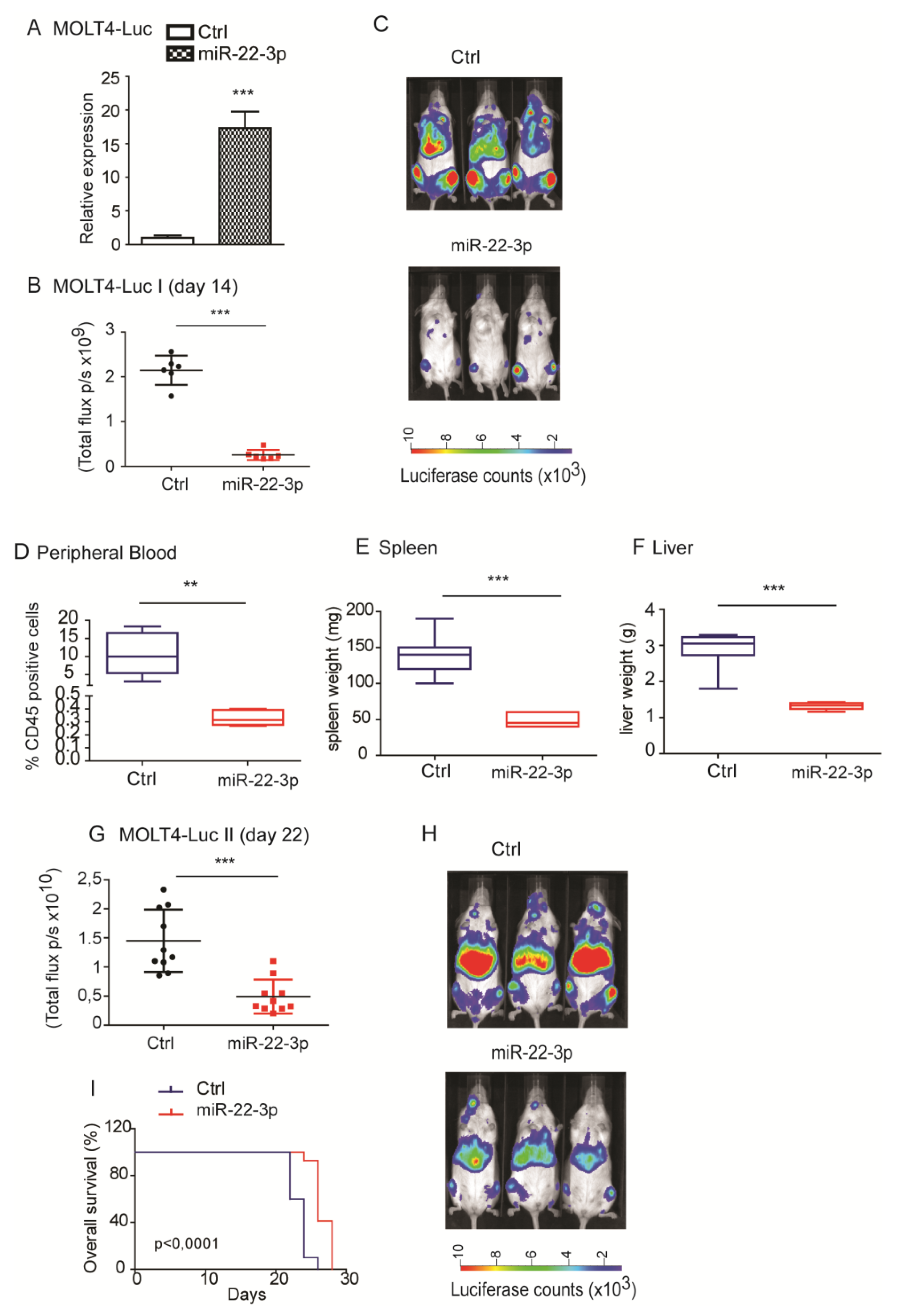
To evaluate the in vivo effects of miR-22-3p overexpression, MOLT4 and Jurkat-E6 cells that were previously engineered to express the luciferase reporter gene (FUW-Luc-mCherry-puro or MIGR1-mCherry-Luc2) were transduced with pLenti-III-GFP pre-hsa-miR-22-3p or pLenti-III-GFP empty vector control.
For in vivo experiments, 5 × 106 miR-22-3p-overexpressing or control MOLT4 cells expressing luciferase were i.v. injected into 6–8-week-old female NSG mice. After a 7-day window for tumor engraftment, we evaluated disease progression by bioimaging using the IVIS Spectrum in vivo imaging system (Xenogen Corporation, Caliper Life Sciences, Waltham, MA, USA). Tumor bioluminescence was quantified by integrating the total photonic flux (photons/s, p/s) through a region encircling each mouse as determined by the Living Image software package (Xenogen Corporation). In order to evaluate tumor infiltration, all mice were sacrificed at the same time point, and flow cytometry analysis of CD45-positive cells was used to monitor tumor load in peripheral blood, spleen, and liver tissues. These experiments were authorized by the Italian Ministry of Health (authorization n° 618/2016-PR released the 17th of June 2016).
2.4. Statistical Analysis
The results are expressed as mean value ± standard deviation (SD). Statistical testing was performed using Student’s t-test (two-tailed, unpaired). For in vivo experiments, survival curves were estimated using the Kaplan–Meier method, and the difference between them was evaluated using the log-rank test (GraphPad Prism, GraphPad Software, San Diego, CA, USA)). Statistically significant differences are defined as * p < 0.05, ** p < 0.01, *** p < 0.001.
4. Discussion
T-ALL is an aggressive hematologic tumor that accounts for 10–15% of pediatric and 25% of adult ALL cases. The introduction of intensive combination chemotherapy protocols has led to significant improvements in survival; however, these aggressive regimens are non-specific and very often associated with acute toxicity. The discovery that more than 60% of T-ALL patients harbor mutations in the
NOTCH1 gene has generated new perspectives for the treatment of T-ALL through the use of GSIs or small peptides/antibodies that block NOTCH1 signaling and its regulators. Among the different modulators of NOTCH1 signaling, miRNAs have been shown to play an important role in tumor progression, suggesting the possibility of combining NOTCH1 inhibition with miRNA-based therapy. In this study, we undertook an in vivo approach to identify critical miRNAs that are regulated in the context of NOTCH1 inhibition and that may exert an important function in concert with NOTCH1 in established tumors. We thus used murine NOTCH1-induced T-cell tumors treated in vivo with a highly active GSI to generate paired miRNA and gene expression profiles. Gene expression analysis found numerous genes that were significantly down- and upregulated upon DBZ treatment, in line with previously described results [
17]. Importantly, GSEA showed that the gene sets related to NOTCH signaling and MYC targets were significantly downregulated following NOTCH1 inhibition, confirming the efficacy of our experimental model [
7,
8].
In accordance with previous studies, we identified several components of the miR-17-92 cluster at the top of the list of NOTCH1 downregulated miRNAs, thus reinforcing the relevance of this cluster in the context of NOTCH1-induced T-ALL [
11,
13]. In fact, dual translocations that simultaneously affect the 17-92 cluster where miR-19 is located and NOTCH1 highlight the oncogenic importance of this interaction in T-ALL [
11]. Notably, from our analysis, miR-19a was the most significantly downregulated miRNA upon NOTCH1 inhibition, further supporting its crucial role as an oncomiR in NOTCH1-driven T-ALL. Among the upregulated miRNAs was miR-709, which was previously reported as a tumor suppressor miRNA in the context of NOTCH1-induced leukemia [
12]. According to this study, miR-709 represses crucial oncogenes such as Myc, Akt, and Ras-GRF1, playing an important role in murine leukemia initiation and maintenance. Within the miRNAs upregulated upon NOTCH1 inhibition, we also found two conserved miRNAs previously described in the context of T-ALL [
23]: miR-29a-3p and miR-223-3p [
22,
23]. In particular, miR-29 has been reported to function as a tumor suppressor in a NOTCH1-driven T-ALL model and found to be significantly downregulated in T-ALL samples relative to normal thymocytes through the regulation of key genes involved in epigenetic regulatory mechanisms [
24,
25]. According to our data, in the context of NOTCH1 inhibition, miR-29a-3p showed inconsistent results among human T-ALL cells, exhibiting both upregulation and downregulation without a specific trend. miR-223-3p has been described as an oncomiR in T-ALL because it was found to be highly expressed in T-ALL samples and to cooperate with other miRNAs in regulating key tumor suppressors implicated in T-ALL [
13,
14]. In our dataset, miR-223-3p was upregulated upon NOTCH1 inhibition, apparently in contrast to its reported oncogenic function. However, our data are in line with another study in which miR-223-3p was found to be upregulated upon NOTCH1 inhibition, implying a complex function for miR-223-3p in T-ALL that may depend on the cell context [
15].
Within the miRNAs upregulated upon NOTCH1 inhibition, we also found significant upregulation of miR-22-3p, which has been previously reported as a putative tumor suppressor in acute myeloid leukemia (AML) but never explored in T-ALL, especially downstream of NOTCH1 [
22]. We confirmed miR-22-3p modulation upon NOTCH1 inhibition, not only in different NOTCH1-induced tumors, but also in human specimens, such as T-ALL cell lines and PDX samples.
In the last couple of years, miR-22-3p has emerged as a crucial regulator of neoplastic progression. Initially, it was shown that miR-22-3p-overexpressing transgenic mice develop myelodysplastic syndrome, which subsequently progresses to AML [
26]. In contrast to these results, Jiang X et al. demonstrated that the forced expression of miR-22-3p significantly suppressed in vitro AML cell viability and growth and substantially inhibited in vivo leukemia development and maintenance [
27]. According to these results, miR-22-3p targeted multiple oncogenes, including CRTC1, FLT3, and MYCBP, and thus repressed the CREB and MYC pathways. In line with these data, another study showed that miR-22-3p is a tumor suppressor in AML induced by PU.1, an important transcription factor of monocyte/macrophage differentiation, and reintroduction of miR-22-3p in AML blasts reversed the blocking of differentiation and the inhibition of cell growth [
27].
Our data showed that miR-22-3p was repressed in human T-ALL cell lines and PDX samples relative to normal thymocytes. Interestingly, Ghisi M. and collaborators identified miR-22-3p amongst the miRNAs upregulated during normal T-cell development, indicating a different mechanism of regulation of miR-22-3p during the maturation of T cells [
28].
Importantly, as previously shown in AML, we demonstrated that overexpression of miR-22-3p in T-ALL cell lines carrying constitutively active NOTCH1 inhibited single-cell growth in soft agar, affecting both the number and size of the colonies. Moreover, miR-22-3p significantly impaired tumor growth in vivo when overexpressed in human T-ALL cells, reducing the percentage of tumor cells in the blood and the size and weight of the spleen and liver.
The striking effects observed in MOLT4 cells overexpressing miR-22-3p can, in part, be explained by the effects of miR-22-3p overexpression on NOTCH1 target genes. Using qPCR, we found that crucial targets downstream of NOTCH1 were significantly downregulated upon miR-22-3p overexpression. For instance, HES1, a known repressor downstream of NOTCH1 that is involved in the survival and growth of T-ALL cells [
29], was downregulated in all three analyzed cell lines, and in MOLT4 cells, the corresponding decrease could also be detected at the protein level. Similar results were also obtained in MOLT4 cells for IGF1R, an important regulator of growth and leukemic stem cell activity [
6]. It is plausible that these regulations may affect T-ALL cell growth in vivo, where the tumor microenvironment, in concert with NOTCH1 signaling, is a critical factor for tumor establishment and growth.
Overall, these data demonstrate that high miR-22-3p levels can impact tumor progression, possibly affecting the leukemic stem cell compartment in supportive niches and leading to delayed tumor growth.