Evaluation of Glycosylated PTGS2 in Colorectal Cancer for NSAIDS-Based Adjuvant Therapy
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Specimens Collection and Processing
2.3. Cell Lines
2.4. Western Blot
2.5. Immunohistochemistry
2.6. IL1β ELISA
2.7. Statistics
3. Results
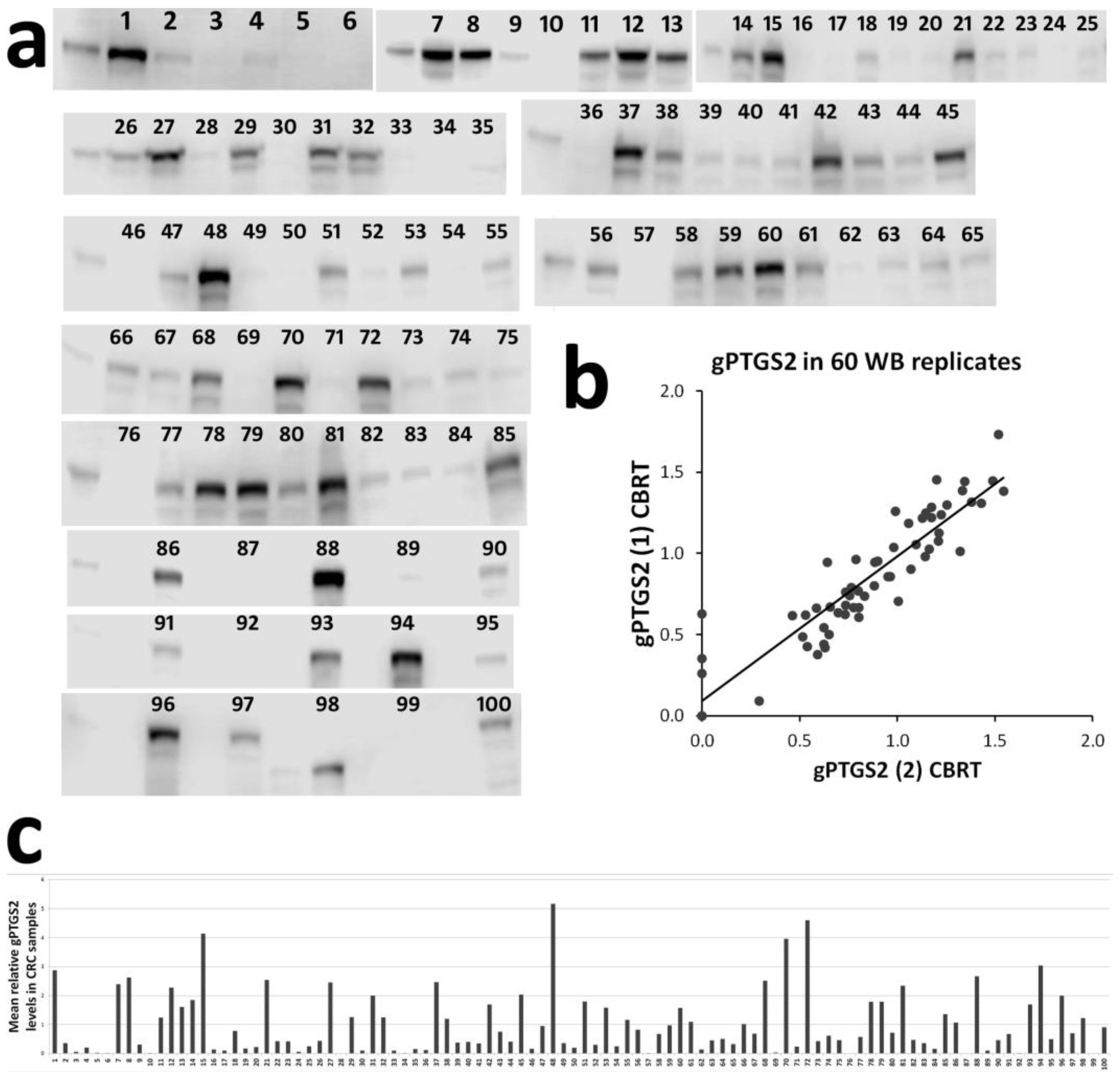
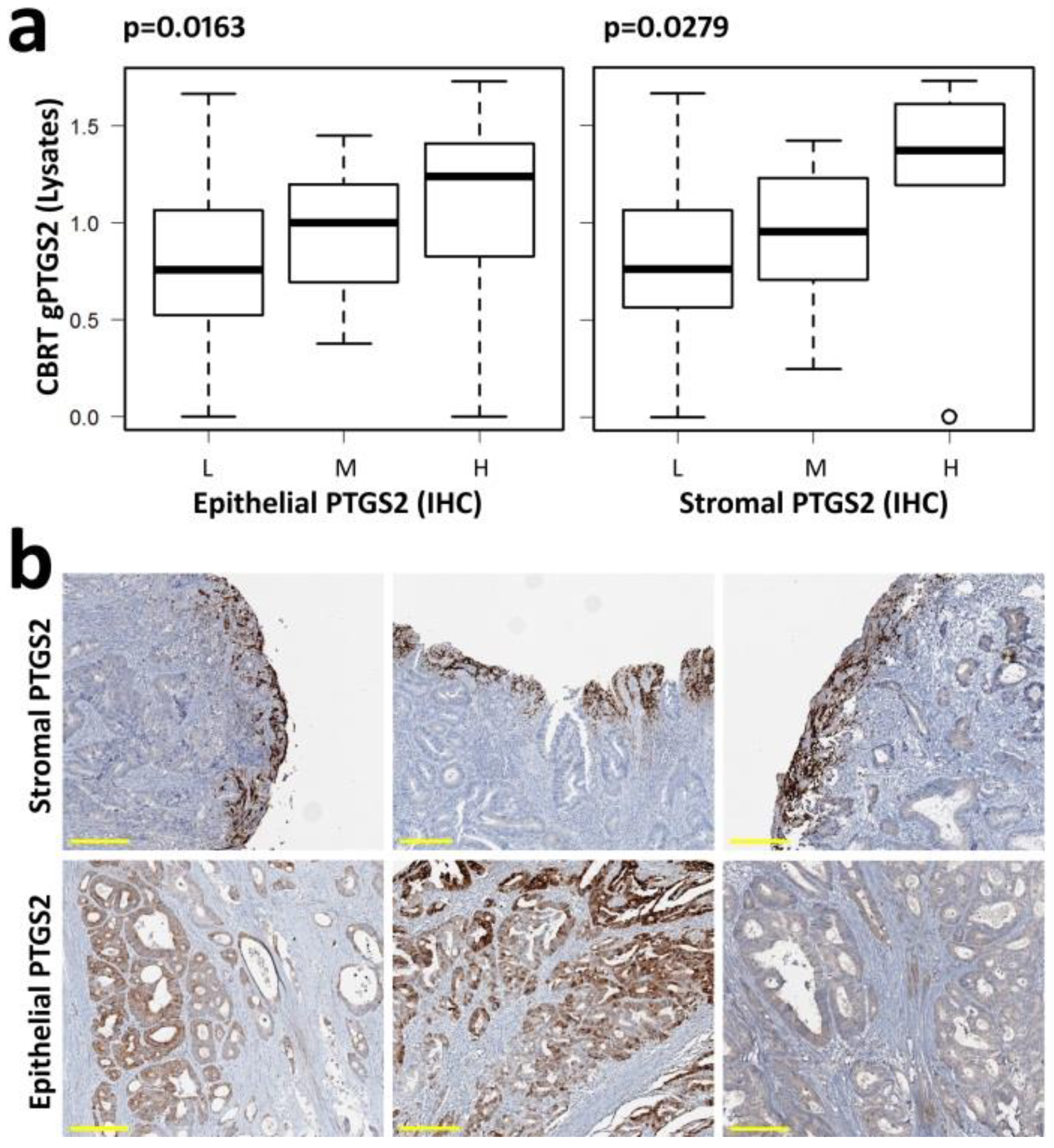
3.1. gPTGS2 Quantification in 100 CRC Lysates and its Relation to Tissue PTGS2
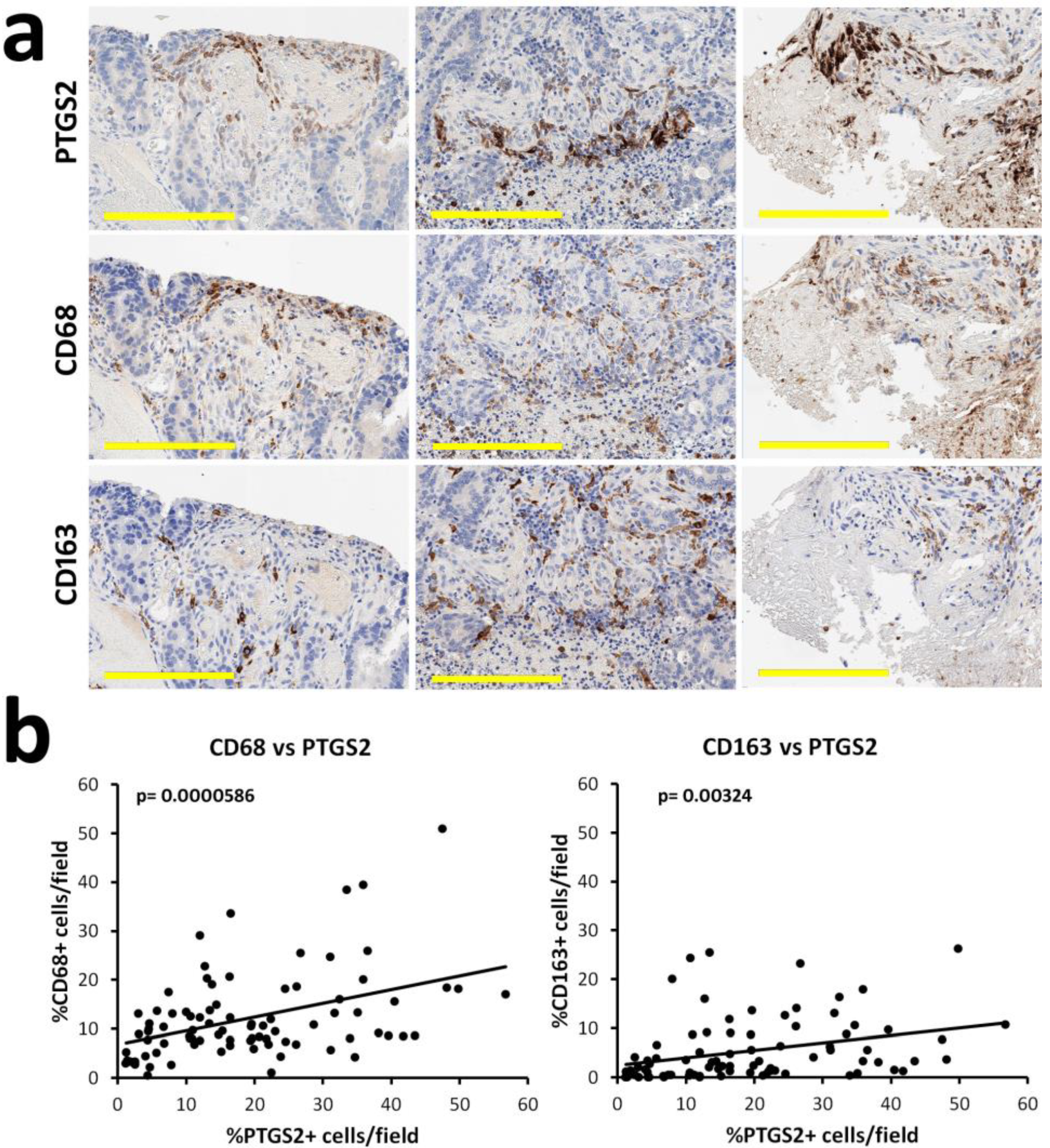
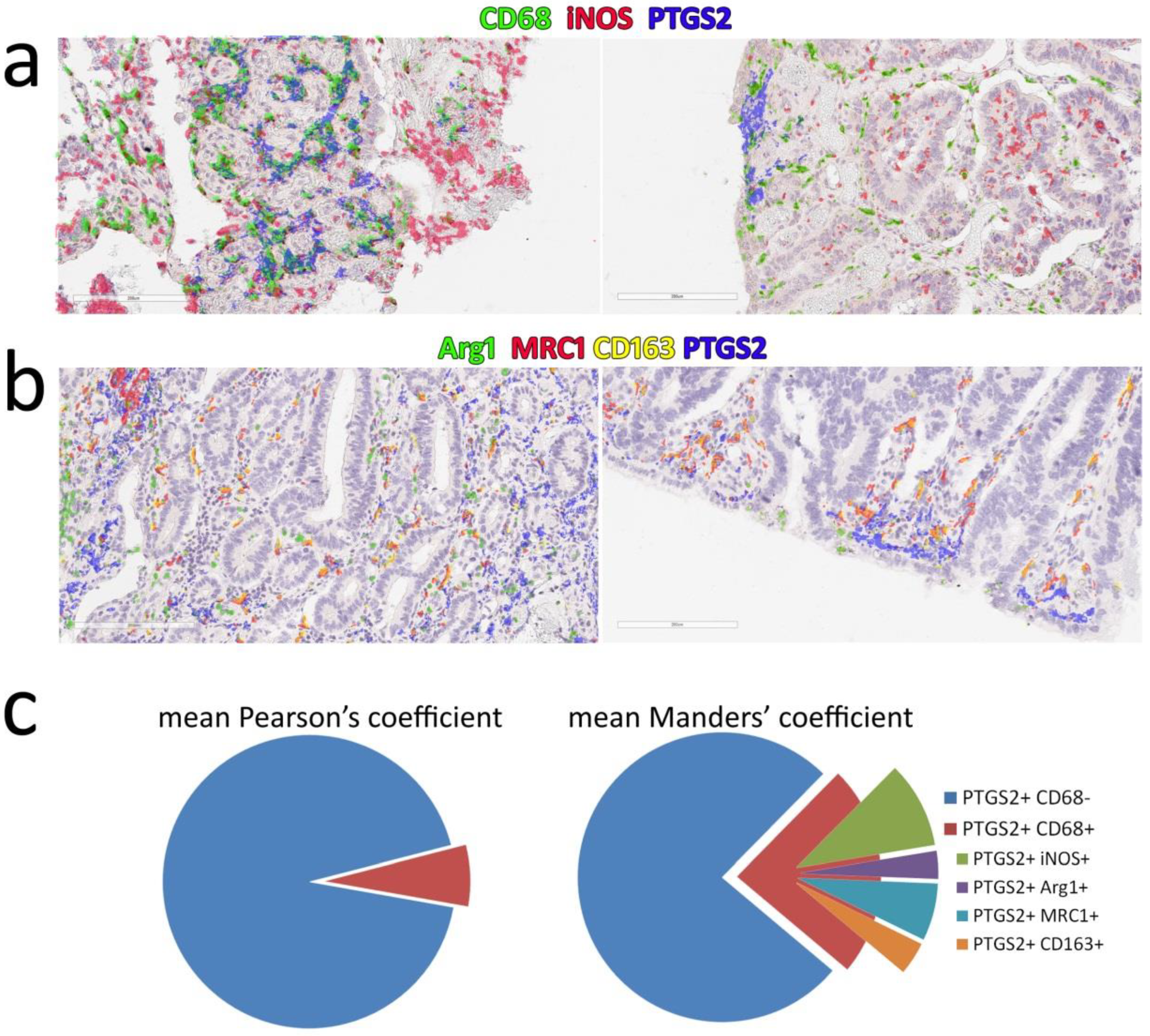
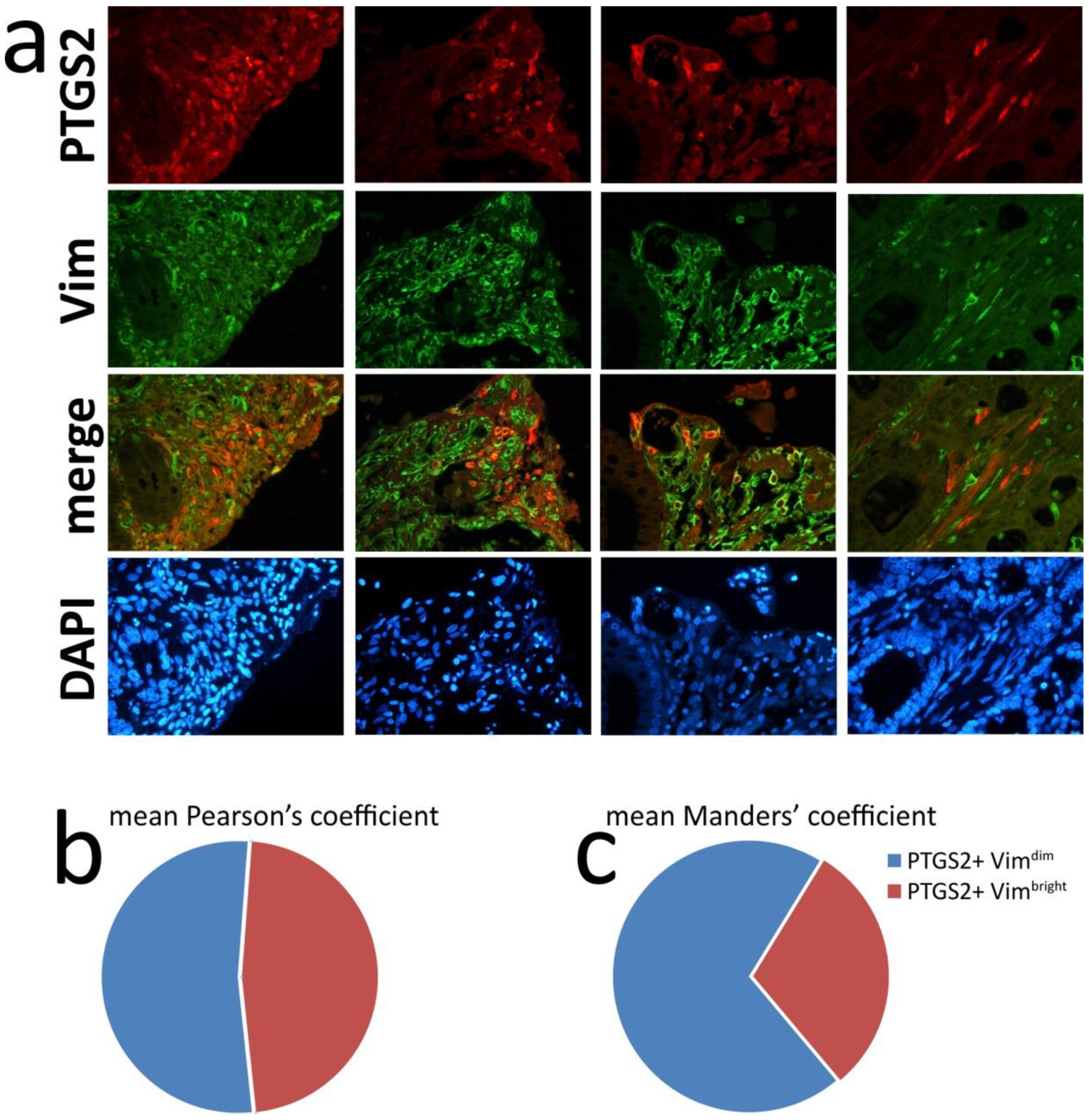
3.2. Identification of gPTGS2 Positive Cells in the Stromal Component
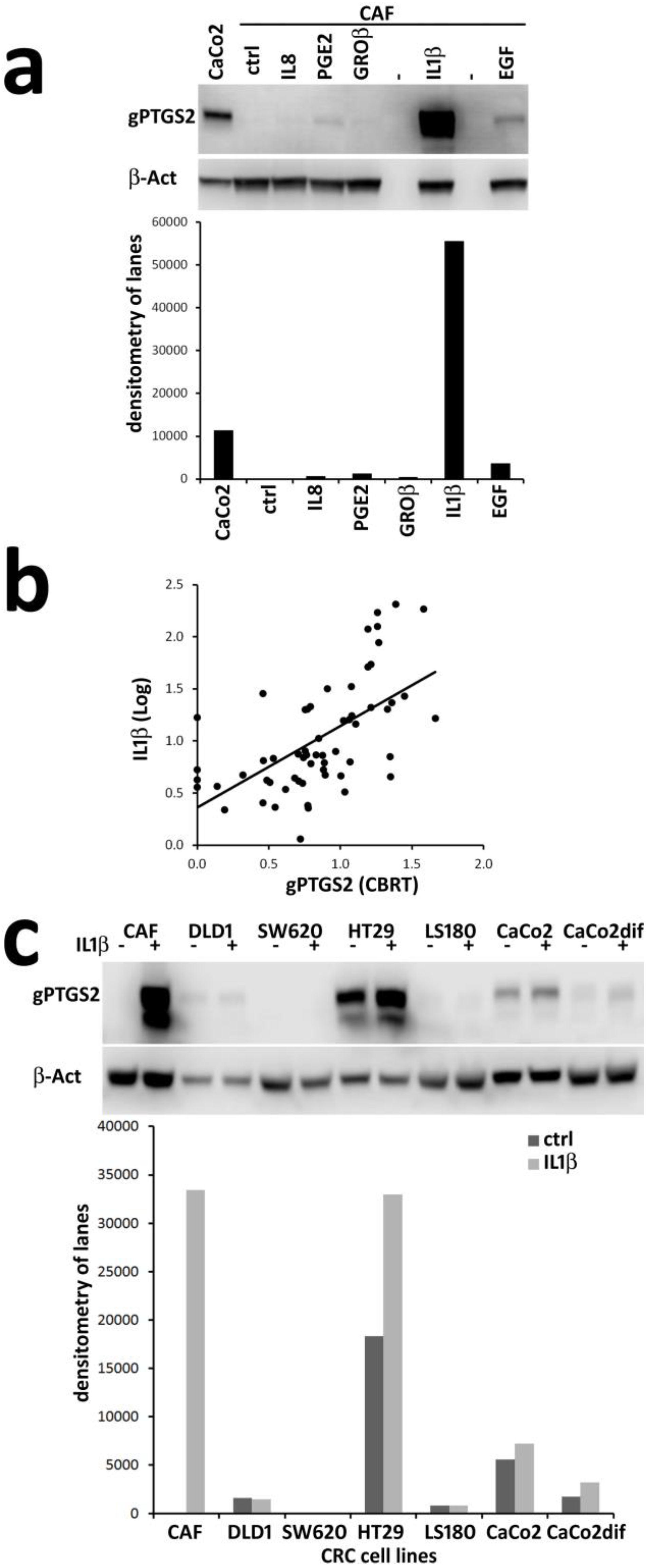
3.3. In Vitro, IL1β is a Powerful Inducers of PTGS2 in CAF and Correlates to gPTGS2 Levels in Tissue Lysates
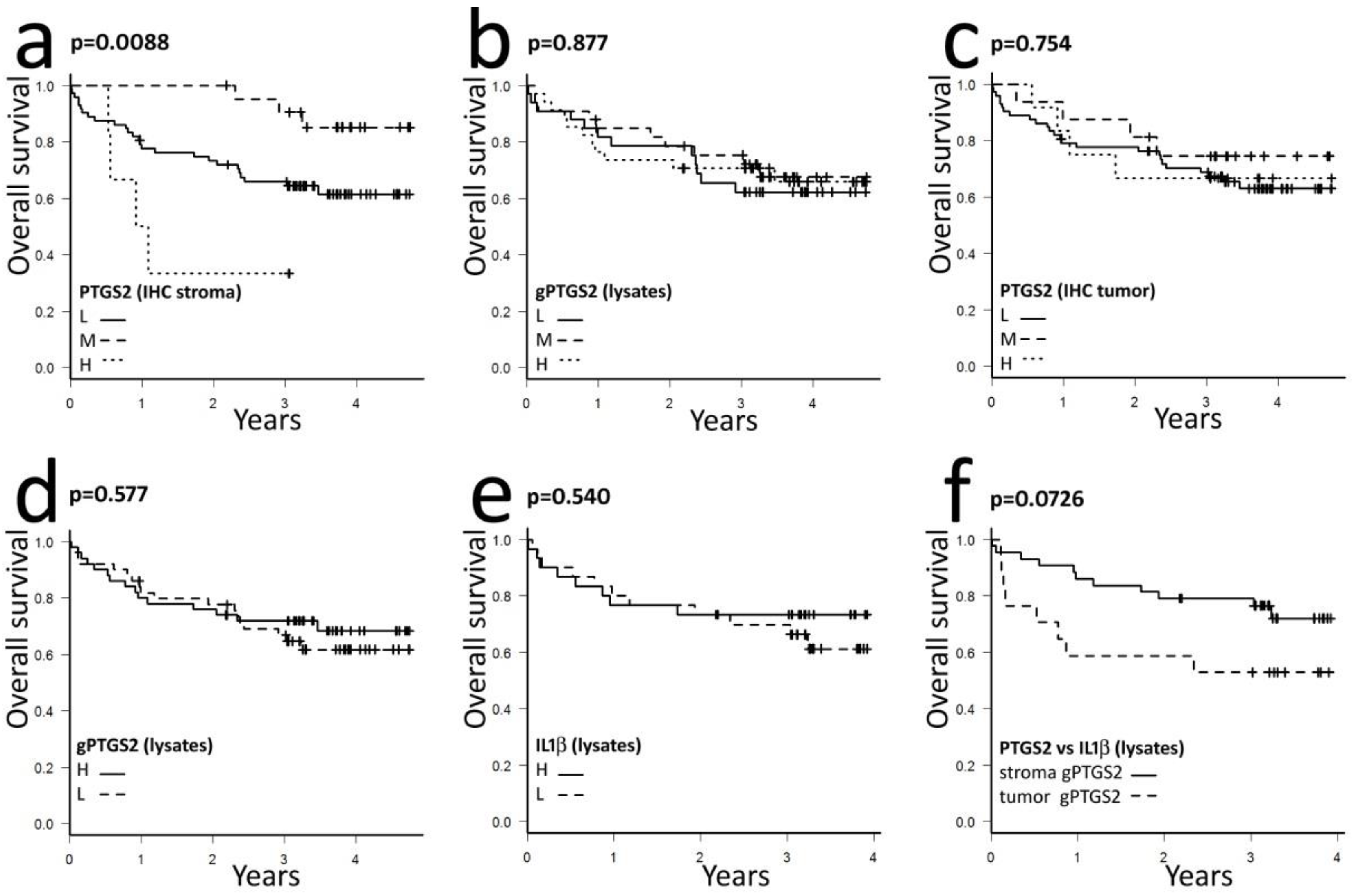
3.4. Effects of Stromal PTGS2 on Patients Prognosis
4. Discussion
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Benelli, R.; Venè, R.; Ferrari, N. Prostaglandin-endoperoxide synthase 2 (cyclooxygenase-2), a complex target for colorectal cancer prevention and therapy. Transl. Res. 2018, 196, 42–61. [Google Scholar] [CrossRef] [PubMed]
- Jaén, R.I.; Prieto, P.; Casado, M.; Martín-Sanz, P.; Boscá, L. Post-translational modifications of prostaglandin-endoperoxide synthase 2 in colorectal cancer: An update. World J. Gastroenterol. 2018, 24, 5454–5461. [Google Scholar] [CrossRef] [PubMed]
- Arber, N.; Eagle, C.J.; Spicak, J.; Rácz, I.; Dite, P.; Hajer, J.; Zavoral, M.; Lechuga, M.J.; Gerletti, P.; Tang, J.; et al. Celecoxib for the Prevention of Colorectal Adenomatous Polyps. N. Engl. J. Med. 2006, 355, 885–895. [Google Scholar] [CrossRef]
- Bertagnolli, M.M.; Eagle, C.J.; Zauber, A.G.; Redston, M.; Solomon, S.D.; Kim, K.; Tang, J.; Rosenstein, R.B.; Wittes, J.; Corle, D.; et al. Celecoxib for the Prevention of Sporadic Colorectal Adenomas. N. Engl. J. Med. 2006, 355, 873–884. [Google Scholar] [CrossRef] [PubMed]
- Ng, K.; Meyerhardt, J.A.; Chan, A.T.; Sato, K.; Chan, J.A.; Niedzwiecki, D.; Saltz, L.B.; Mayer, R.J.; Benson, A.B.; Schaefer, P.L.; et al. Aspirin and COX-2 inhibitor use in patients with stage III colon cancer. J. Natl. Cancer Inst. 2015, 107, 345. [Google Scholar] [CrossRef] [PubMed]
- Friis, S.; Riis, A.H.; Erichsen, R.; Baron, J.A.; Sørensen, H.T. Low-Dose Aspirin or Nonsteroidal Anti-inflammatory Drug Use and Colorectal Cancer Risk: A Population-Based, Case-Control Study. Ann. Intern. Med. 2015, 163, 347–355. [Google Scholar] [CrossRef] [PubMed]
- Hua, X.; Adams, S.V.; Phipps, A.I.; Cohen, S.A.; Burnett-Hartman, A.; Hardikar, S.; Newcomb, P.A. Pre- and Post-Diagnostic Non-Steroidal Anti-Inflammatory Drug Use and Colorectal Cancer Survival in Seattle Colon Cancer Family Registry. Cancer Epidemiol. Biomarkers Prev. 2016, 25, 559. [Google Scholar] [CrossRef]
- Midgley, R.S.; McConkey, C.C.; Johnstone, E.C.; Dunn, J.A.; Smith, J.L.; Grumett, S.A.; Julier, P.; Iveson, C.; Yanagisawa, Y.; Warren, B.; et al. Phase III randomized trial assessing rofecoxib in the adjuvant setting of colorectal cancer: Final results of the VICTOR trial. J. Clin. Oncol. 2010, 28, 4575–4580. [Google Scholar] [CrossRef]
- Fux, R.; Schwab, M.; Thon, K.-P.; Gleiter, C.H.; Fritz, P. Cyclooxygenase-2 Expression in Human Colorectal Cancer Is Unrelated to Overall Patient Survival. Clin. Cancer Res. 2005, 11, 4754–4760. [Google Scholar] [CrossRef]
- Ogino, S.; Kirkner, G.J.; Nosho, K.; Irahara, N.; Kure, S.; Shima, K.; Hazra, A.; Chan, A.T.; Dehari, R.; Giovannucci, E.L.; et al. Cyclooxygenase-2 expression is an independent predictor of poor prognosis in colon cancer. Clin. Cancer Res. 2008, 14, 8221–8227. [Google Scholar] [CrossRef]
- Soumaoro, L.T.; Uetake, H.; Higuchi, T.; Takagi, Y.; Enomoto, M.; Sugihara, K. Cyclooxygenase-2 expression: A significant prognostic indicator for patients with colorectal cancer. Clin. Cancer Res. 2004, 10, 8465–8471. [Google Scholar] [CrossRef] [PubMed]
- Conti, J.; Thomas, G. The Role of Tumour Stroma in Colorectal Cancer Invasion and Metastasis. Cancers 2011, 3, 2160–2168. [Google Scholar] [CrossRef] [PubMed]
- Mei, Z.; Liu, Y.; Liu, C.; Cui, A.; Liang, Z.; Wang, G.; Peng, H.; Cui, L.; Li, C. Tumour-infiltrating inflammation and prognosis in colorectal cancer: Systematic review and meta-analysis. Br. J. Cancer 2014, 110, 1595–1605. [Google Scholar] [CrossRef] [PubMed]
- Roseweir, A.K.; Park, J.H.; Hoorn, S.; Powell, A.G.; Roxburgh, C.S.; McMillan, D.C.; Horgan, P.G.; Vermeulen, L.; Edwards, J. Abstract 4615: Phenotypic subtypes successfully stratify prognosis of patient with colorectal cancer: A step towards precision medicine. Cancer Res. 2018, 78, 4615. [Google Scholar]
- Benelli, R.; Venè, R.; Minghelli, S.; Carlone, S.; Gatteschi, B.; Ferrari, N. Celecoxib induces proliferation and Amphiregulin production in colon subepithelial myofibroblasts, activating erk1-2 signaling in synergy with EGFR. Cancer Lett. 2013, 328, 73–82. [Google Scholar] [CrossRef]
- Chapple, K.S.; Cartwright, E.J.; Hawcroft, G.; Tisbury, A.; Bonifer, C.; Scott, N.; Windsor, A.C.; Guillou, P.J.; Markham, A.F.; Coletta, P.L.; et al. Localization of cyclooxygenase-2 in human sporadic colorectal adenomas. Am. J. Pathol. 2000, 156, 545–553. [Google Scholar] [CrossRef]
- Bamba, H.; Ota, S.; Kato, A.; Adachi, A.; Itoyama, S.; Matsuzaki, F. High expression of cyclooxygenase-2 in macrophages of human colonic adenoma. Int. J. Cancer 1999, 83, 470–475. [Google Scholar] [CrossRef]
- Heusinkveld, M.; van der Burg, S.H. Identification and manipulation of tumor associated macrophages in human cancers. J. Transl. Med. 2011, 9, 216. [Google Scholar] [CrossRef]
- Nakanishi, Y.; Nakatsuji, M.; Seno, H.; Ishizu, S.; Akitake-Kawano, R.; Kanda, K.; Ueo, T.; Komekado, H.; Kawada, M.; Minami, M.; et al. COX-2 inhibition alters the phenotype of tumor-associated macrophages from M2 to M1 in ApcMin/+ mouse polyps. Carcinogenesis 2011, 32, 1333–1339. [Google Scholar] [CrossRef]
- Sheppe, A.E.F.; Kummari, E.; Walker, A.; Richards, A.; Hui, W.W.; Lee, J.H.; Mangum, L.; Borazjani, A.; Ross, M.K.; Edelmann, M.J. PGE2 Augments Inflammasome Activation and M1 Polarization in Macrophages Infected With Salmonella Typhimurium and Yersinia enterocolitica. Front. Microbiol 2018, 9, 2447. [Google Scholar] [CrossRef]
- Adegboyega, P.A.; Ololade, O.; Saada, J.; Mifflin, R.; Di Mari, J.F.; Powell, D.W. Subepithelial myofibroblasts express cyclooxygenase-2 in colorectal tubular adenomas. Clin. Cancer Res. 2004, 10, 5870–5879. [Google Scholar] [CrossRef] [PubMed]
- Sonoshita, M.; Takaku, K.; Oshima, M.; Sugihara, K.; Taketo, M.M. Cyclooxygenase-2 Expression in Fibroblasts and Endothelial Cells of Intestinal Polyps. Cancer Res. 2002, 62, 6846–6849. [Google Scholar] [PubMed]
- Ramsay, R.G.; Ciznadija, D.; Vanevski, M.; Mantamadiotis, T. Transcriptional regulation of cyclo-oxygenase expression: Three pillars of control. Int. J. Immunopathol. Pharmacol 2003, 16, 59–67. [Google Scholar] [PubMed]
- Ning, Y.; Lenz, H.-J. Targeting IL-8 in colorectal cancer. Expert Opin. Ther. Tar. 2012, 16, 491–497. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Huang, J.; Zhu, H.; Ju, S.; Wang, H.; Wang, X. Overexpression of GRO-β is associated with an unfavorable outcome in colorectal cancer. Oncol. Lett. 2016, 11, 2391–2397. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, D.; Xia, D.; DuBois, R.N. The Crosstalk of PTGS2 and EGF Signaling Pathways in Colorectal Cancer. Cancers 2011, 3, 3894–3908. [Google Scholar] [CrossRef] [PubMed]
- Martinez, F.O.; Gordon, S.; Locati, M.; Mantovani, A. Transcriptional Profiling of the Human Monocyte-to-Macrophage Differentiation and Polarization: New Molecules and Patterns of Gene Expression. J. Immunol. 2006, 177, 7303–7311. [Google Scholar] [CrossRef]
- Voronov, E.; Apte, R.N. IL-1 in Colon Inflammation, Colon Carcinogenesis and Invasiveness of Colon Cancer. Cancer Microenviron. 2015, 8, 187–200. [Google Scholar] [CrossRef]
- Cui, G.; Yuan, A.; Sun, Z.; Zheng, W.; Pang, Z. IL-1β/IL-6 network in the tumor microenvironment of human colorectal cancer. Pathol. Res. Pract. 2018, 214, 986–992. [Google Scholar] [CrossRef]
- Dahmus, J.D.; Kotler, D.L.; Kastenberg, D.M.; Kistler, C.A. The gut microbiome and colorectal cancer: A review of bacterial pathogenesis. J. Gastrointest. Oncol. 2018, 9, 769–777. [Google Scholar] [CrossRef]
- Abdulamir, A.S.; Hafidh, R.R.; Bakar, F.A. Molecular detection, quantification, and isolation of Streptococcus gallolyticus bacteria colonizing colorectal tumors: Inflammation-driven potential of carcinogenesis via IL-1, COX-2, and IL-8. Mol. Cancer 2010, 9, 249. [Google Scholar] [CrossRef] [PubMed]
- Castells, A.; Payá, A.; Alenda, C.; Rodríguez-Moranta, F.; Agrelo, R.; Andreu, M.; Piñol, V.; Castellví-Bel, S.; Jover, R.; Llor, X.; et al. Cyclooxygenase 2 expression in colorectal cancer with DNA mismatch repair deficiency. Clin. Cancer Res. 2006, 12, 1686–1692. [Google Scholar] [CrossRef] [PubMed]
- Karnes, W.E.; Shattuck-Brandt, R.; Burgart, L.J.; DuBois, R.N.; Tester, D.J.; Cunningham, J.M.; Kim, C.Y.; McDonnell, S.K.; Schaid, D.J.; Thibodeau, S.N. Reduced COX-2 protein in colorectal cancer with defective mismatch repair. Cancer Res. 1998, 58, 5473–5477. [Google Scholar] [PubMed]
- Domingo, E.; Church, D.N.; Sieber, O.; Ramamoorthy, R.; Yanagisawa, Y.; Johnstone, E.; Davidson, B.; Kerr, D.J.; Tomlinson, I.P.M.; Midgley, R. Evaluation of PIK3CA Mutation As a Predictor of Benefit From Nonsteroidal Anti-Inflammatory Drug Therapy in Colorectal Cancer. JCO 2013, 31, 4297–4305. [Google Scholar] [CrossRef]
| Clinicopathologic Parameters | All | PTGS2 < Median | PTGS2 > Median | p |
|---|---|---|---|---|
| TOTAL | 100 | 50 | 50 | - |
| SEX | 0.689 | |||
| Male | 49 | 26 | 23 | |
| Female | 51 | 24 | 27 | |
| AGE (surgery) | 0.317 | |||
| ≤70 | 47 | 22 | 28 | |
| >70 | 53 | 28 | 22 | |
| TUMOR LOCATION * | 0.965 | |||
| Proximal colon | 43 | 21 | 22 | |
| Distal colon | 36 | 18 | 18 | |
| Rectum | 21 | 11 | 10 | |
| STAGE (UICC-2009) | 0.974 | |||
| I | 14 | 7 | 7 | |
| II | 34 | 16 | 18 | |
| III | 39 | 20 | 19 | |
| IV | 13 | 7 | 6 | |
| TUMOR GRADE | 0.479 | |||
| well | 5 | 2 | 4 | |
| moderate | 75 | 40 | 35 | |
| poor | 19 | 8 | 11 | |
| JASS SCORE | 0.719 | |||
| I | 17 | 7 | 10 | |
| II | 14 | 6 | 8 | |
| III | 32 | 18 | 14 | |
| IV | 37 | 19 | 18 | |
| PERINEURAL INVASION | 0.387 | |||
| PNI 0 | 69 | 37 | 32 | |
| PNI 1 | 31 | 13 | 18 | |
| MICROSAT. INSTAB. ** | 0.158 | |||
| MSS | 72 | 40 | 32 | |
| MSI | 15 | 5 | 10 | |
| IL1β *** | 0.002 | |||
| IL1β<median | 30 | 21 | 8 | |
| IL1β>median | 30 | 10 | 22 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Venè, R.; Costa, D.; Augugliaro, R.; Carlone, S.; Scabini, S.; Casoni Pattacini, G.; Boggio, M.; Zupo, S.; Grillo, F.; Mastracci, L.; et al. Evaluation of Glycosylated PTGS2 in Colorectal Cancer for NSAIDS-Based Adjuvant Therapy. Cells 2020, 9, 683. https://doi.org/10.3390/cells9030683
Venè R, Costa D, Augugliaro R, Carlone S, Scabini S, Casoni Pattacini G, Boggio M, Zupo S, Grillo F, Mastracci L, et al. Evaluation of Glycosylated PTGS2 in Colorectal Cancer for NSAIDS-Based Adjuvant Therapy. Cells. 2020; 9(3):683. https://doi.org/10.3390/cells9030683
Chicago/Turabian StyleVenè, Roberta, Delfina Costa, Raffaella Augugliaro, Sebastiano Carlone, Stefano Scabini, Gianmaria Casoni Pattacini, Maurizio Boggio, Simonetta Zupo, Federica Grillo, Luca Mastracci, and et al. 2020. "Evaluation of Glycosylated PTGS2 in Colorectal Cancer for NSAIDS-Based Adjuvant Therapy" Cells 9, no. 3: 683. https://doi.org/10.3390/cells9030683
APA StyleVenè, R., Costa, D., Augugliaro, R., Carlone, S., Scabini, S., Casoni Pattacini, G., Boggio, M., Zupo, S., Grillo, F., Mastracci, L., Pitto, F., Minghelli, S., Ferrari, N., Tosetti, F., Romairone, E., Mingari, M. C., Poggi, A., & Benelli, R. (2020). Evaluation of Glycosylated PTGS2 in Colorectal Cancer for NSAIDS-Based Adjuvant Therapy. Cells, 9(3), 683. https://doi.org/10.3390/cells9030683








