Computational Detection of Breast Cancer Invasiveness with DNA Methylation Biomarkers
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials: DNA Methylation Datasets
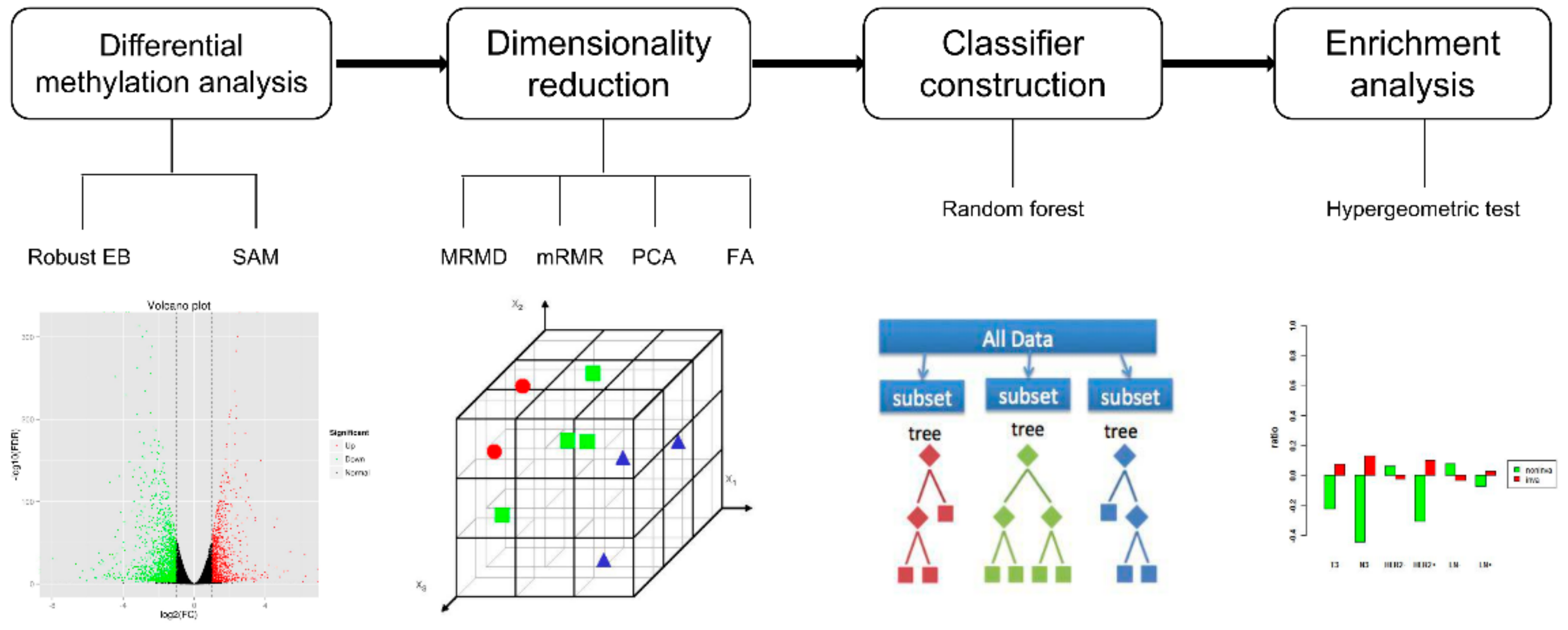
2.2. Methods: Study Design
2.2.1. Robust Empirical Bayes
2.2.2. Significance Analysis of Microarrays
2.2.3. Maximum Relevance Maximum Distance
2.2.4. Minimal Redundancy Maximal Relevance
2.2.5. Principal Component Analysis
2.2.6. Factor Analysis
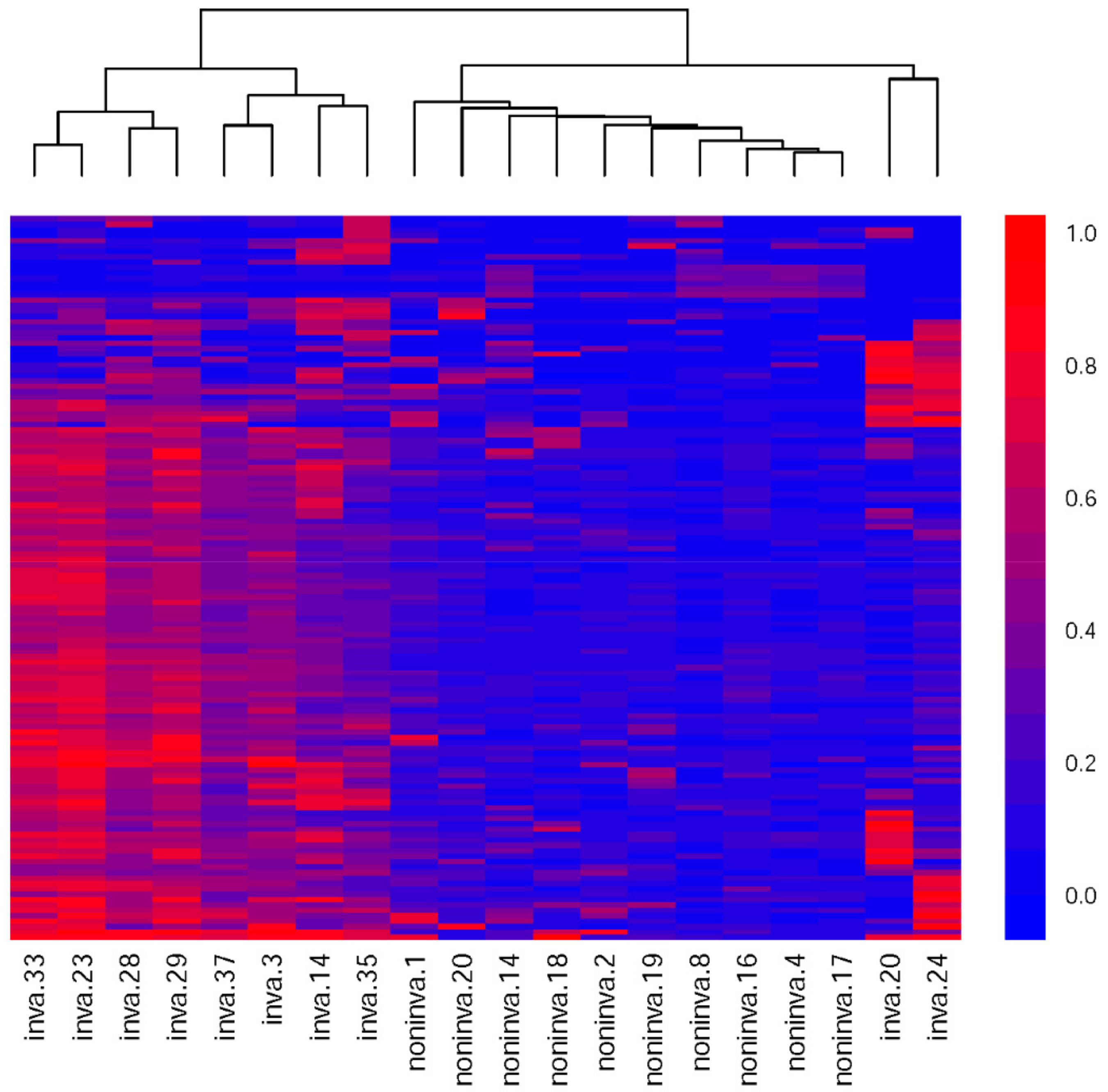
2.2.7. Unsupervised Hierarchical Clustering
2.2.8. Constructing the DNA Methylation-Based Invasiveness Classifier
2.2.9. Hypergeometric Test
3. Results
3.1. Feature Selection
3.2. Development of Classifier Based on DNA Methylation Biomarkers
3.3. TCGA Beast Cancer Cohort Confirms the Performance of Classifiers
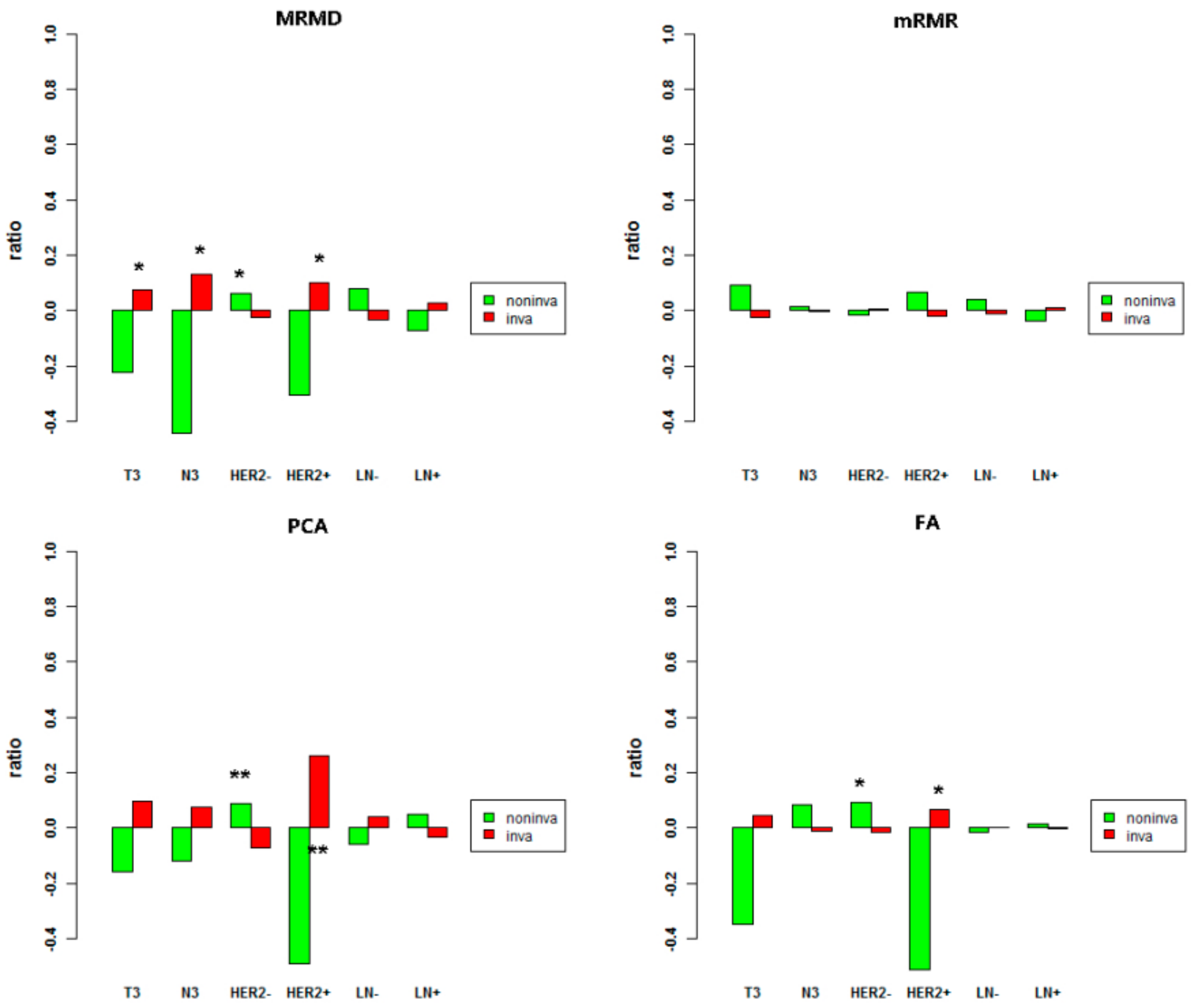
3.4. Methylation Differences between the Invasive and Noninvasive Groups
3.5. Genes Related to Metastasis
3.6. Website BMMP
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Perou, C.M.; Sørlie, T.; Eisen, M.B.; Van, d.R.M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A. Molecular portraits of human breast tumors. Nature 2012, 490, 747–752. [Google Scholar] [CrossRef]
- Sorlie, T.; Tibshirani, R.; Parker, J.; Hastie, T.; Marron, J.S.; Nobel, A.; Deng, S.; Johnsen, H.; Pesich, R.; Geisler, S. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc. Natl. Acad. Sci. USA 2003, 100, 8418–8423. [Google Scholar] [CrossRef] [PubMed]
- Nones, K.; Johnson, J.; Newell, F.; Patch, A.; Thorne, H.; Kazakoff, S.; de Luca, X.; Parsons, M.; Ferguson, K.; Reid, L. Whole-genome sequencing reveals clinically relevant insights into the aetiology of familial breast cancers. Ann. Oncol. 2019, 30, 1071–1079. [Google Scholar] [CrossRef]
- Wang, J.H.; Yang, X.; Cai, H.M.; Tan, W.C.; Jin, C.Z.; Li, L. Discrimination of Breast Cancer with Microcalcifications on Mammography by Deep Learning. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef]
- Doornebal, C.W.; Klarenbeek, S.; Braumuller, T.M.; Klijn, C.N.; Ciampricotti, M.; Hau, C.S.; Hollmann, M.W.; Jonkers, J.; Visser, K.E.D. A Preclinical Mouse Model of Invasive Lobular Breast Cancer Metastasis. Cancer Res. 2013, 73, 353–363. [Google Scholar] [CrossRef] [PubMed]
- Kozłowski, J.; Kozłowska, A.; Kocki, J. Breast cancer metastasis - insight into selected molecular mechanisms of the phenomenon. Postȩpy Hig. I Med. Doświadczalnej 2014, 69, 447–451. [Google Scholar] [CrossRef]
- Fingleton, B. Molecular targets in metastasis: Lessons from genomic approaches. Cancer Genom. Proteom. 2007, 4, 211–221. [Google Scholar]
- Fokas, E.; Engenhart-Cabillic, R.; Daniilidis, K.; Rose, F.; An, H.X. Metastasis: The seed and soil theory gains identity. Cancer Metastasis Rev. 2007, 26, 705–715. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmark of cancer. Cell 2000, 100, 57–71. [Google Scholar] [CrossRef]
- Poste, G.; Fidler, I.J. The pathogenesis of cancer metastasis. Nature 1980, 283, 139–146. [Google Scholar] [CrossRef]
- Du, X.Q.; Li, X.R.; Li, W.; Yan, Y.T.; Zhang, Y.P. Identification and Analysis of Cancer Diagnosis Using Probabilistic Classification Vector Machines with Feature Selection. Curr. Bioinform. 2018, 13, 625–632. [Google Scholar] [CrossRef]
- Liu, H.; Luo, L.B.; Cheng, Z.Z.; Sun, J.J.; Guan, J.H.; Zheng, J.; Zhou, S.G. Group-sparse Modeling Drug-kinase Networks for Predicting Combinatorial Drug Sensitivity in Cancer Cells. Curr. Bioinform. 2018, 13, 437–443. [Google Scholar] [CrossRef]
- Ring, B.Z.; Ross, D.T. Predicting the sites of metastases. Genome Biol. 2005, 6, 241. [Google Scholar] [CrossRef][Green Version]
- Ma, Q.; Reeves, J.H.; Liberles, D.A.; Yu, L.; Chang, Z.; Zhao, J.; Cui, J.; Xu, Y.; Liu, L. A phylogenetic model for understanding the effect of gene duplication on cancer progression. Nucleic Acids Res. 2013, 42, 2870–2878. [Google Scholar] [CrossRef]
- Ellsworth, R.E.; Seebach, J.; Field, L.A.; Heckman, C.; Kane, J.; Hooke, J.A.; Love, B.; Shriver, C.D. A gene expression signature that defines breast cancer metastases. Clin. Exp. Metastasis 2009, 26, 205–213. [Google Scholar] [CrossRef]
- Feng, Y.; Sun, B.; Li, X.; Zhang, L.; Niu, Y.; Xiao, C.; Ning, L.; Fang, Z.; Wang, Y.; Zhang, L. Differentially expressed genes between primary cancer and paired lymph node metastases predict clinical outcome of node-positive breast cancer patients. Breast Cancer Res. Treat. 2007, 103, 319–329. [Google Scholar] [CrossRef]
- Hao, X.; Sun, B.; Hu, L.; Lähdesmäki, H.; Dunmire, V.; Feng, Y.; Zhang, S.W.; Wang, H.; Wu, C.; Wang, H. Differential gene and protein expression in primary breast malignancies and their lymph node metastases as revealed by combined cDNA microarray and tissue microarray analysis. Cancer 2010, 100, 1110–1122. [Google Scholar] [CrossRef]
- Suzuki, M.; Tarin, D. Gene expression profiling of human lymph node metastases and matched primary breast carcinomas: Clinical implications. Mol. Oncol. 2008, 1, 172–180. [Google Scholar] [CrossRef]
- Weigelt, B.; Glas, A.M.; Wessels, L.F.; Witteveen, A.T.; Peterse, J.L.; van’t Veer, L.J. Gene expression profiles of primary breast tumors maintained in distant metastases. Proc. Natl. Acad. Sci. USA 2003, 100, 15901–15905. [Google Scholar] [CrossRef]
- Ren, G.H.; Cao, Y.T.; Wen, S.P.; Huang, T.W.; Zeng, Z.G. A modified Elman neural network with a new learning rate scheme. Neurocomputing 2018, 286, 11–18. [Google Scholar] [CrossRef]
- Cheng, L.; Jiang, Y.; Ju, H.; Sun, J.; Peng, J.; Zhou, M.; Hu, Y. InfAcrOnt: Calculating cross-ontology term similarities using information flow by a random walk. Bmc Genom. 2018, 19, 919. [Google Scholar] [CrossRef]
- Cheng, L.; Hu, Y.; Sun, J.; Zhou, M.; Jiang, Q. DincRNA: A comprehensive web-based bioinformatics toolkit for exploring disease associations and ncRNA function. Bioinformatics 2018, 34, 1953–1956. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.; Li, X.; Guo, H.; Ji, F.; Ye, L.; Ma, X.; Cheng, W. Identification of Bone Metastasis-associated Genes of Gastric Cancer by Genome-wide Transcriptional Profiling. Curr. Bioinform. 2019, 14, 62–69. [Google Scholar] [CrossRef]
- Bianchini, G.; Iwamoto, T.; Qi, Y.; Coutant, C.; Shiang, C.Y.; Wang, B.; Santarpia, L.; Valero, V.; Hortobagyi, G.N.; Symmans, W.F. Prognostic and therapeutic implications of distinct kinase expression patterns in different subtypes of breast cancer. Cancer Res. 2010, 70, 8852. [Google Scholar] [CrossRef] [PubMed]
- Xin, Z.; Ma, Q.; Ren, S.; Wang, G.; Li, F. The understanding of circular RNAs as special triggers in carcinogenesis. Brief. Funct. Genom. 2017, 16, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Breiman, L. Bagging Predictors. Mach. Learn. 1996, 24, 123–140. [Google Scholar] [CrossRef]
- Xu, Y.; Wang, Y.; Luo, J.; Zhao, W.; Zhou, X. Deep learning of the splicing (epi)genetic code reveals a novel candidate mechanism linking histone modifications to ESC fate decision. Nucleic Acids Res. 2017, 45, 12100–12112. [Google Scholar] [CrossRef]
- Xu, M.Z.; Zhao, Z.M.; Zhang, X.P.; Gao, A.Q.; Wu, S.Y.; Wang, J.Y. Synstable Fusion: A Network-Based Algorithm for Estimating Driver Genes in Fusion Structures. Molecules 2018, 23, 2055. [Google Scholar] [CrossRef]
- Cheng, L.; Wang, P.; Tian, R.; Wang, S.; Guo, Q.; Luo, M.; Zhou, W.; Liu, G.; Jiang, H.; Jiang, Q. LncRNA2Target v2.0: A comprehensive database for target genes of lncRNAs in human and mouse. Nucleic Acids Res. 2019, 47, D140–D144. [Google Scholar] [CrossRef]
- Tang, W.; Wan, S.; Yang, Z.; Teschendorff, A.E.; Zou, Q. Tumor origin detection with tissue-specific miRNA and DNA methylation markers. Bioinformatics 2018, 34, 398–406. [Google Scholar] [CrossRef]
- Liao, Z.J.; Li, D.P.; Wang, X.R.; Li, L.S.; Zou, Q. Cancer Diagnosis Through IsomiR Expression with Machine Learning Method. Curr. Bioinform. 2018, 13, 57–63. [Google Scholar] [CrossRef]
- Zeng, W.; Wang, F.; Ma, Y.; Liang, X.C.; Chen, P. Dysfunctional Mechanism of Liver Cancer Mediated by Transcription Factor and Non-coding RNA. Curr. Bioinform. 2019, 14, 100–107. [Google Scholar] [CrossRef]
- Liu, G.; Jiang, Q. Alzheimer’s disease CD33 rs3865444 variant does not contribute to cognitive performance. Proc. Natl. Acad. Sci. USA 2016, 113, E1589–E1590. [Google Scholar] [CrossRef]
- Liu, G.; Zhao, Y.; Jin, S.; Hu, Y.; Wang, T.; Tian, R.; Han, Z.; Xu, D.; Jiang, Q. Circulating vitamin E levels and Alzheimer’s disease: A Mendelian randomization study. Neurobiol. Aging 2018, 72, 189.e181–189.e189. [Google Scholar] [CrossRef]
- Xu, A.D.; Chen, J.Z.; Peng, H.; Han, G.Q.; Cai, H.M. Simultaneous Interrogation of Cancer Omics to Identify Subtypes With Significant Clinical Differences. Front. Genet. 2019, 10, 17. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Han, G.; Xu, A.; Cai, H. Identification of Multidimensional Regulatory Modules through Multi-graph Matching with Network Constraints. IEEE Trans. Bio-Med. Eng. 2019. [Google Scholar] [CrossRef]
- Jiang, Q.; Wang, J.; Wang, Y.; Ma, R.; Wu, X.; Li, Y. TF2LncRNA: Identifying common transcription factors for a list of lncRNA genes from ChIP-Seq data. Biomed. Res. Int 2014, 2014, 317642. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Zhou, X. Applications of Single-Cell Sequencing for Multiomics. Methods Mol. Biol 2018, 1754, 327–374. [Google Scholar] [CrossRef] [PubMed]
- Kalimutho, M.; Nones, K.; Srihari, S.; Duijf, P.H.; Waddell, N.; Khanna, K.K. Patterns of genomic instability in breast cancer. Trends Pharmacol. Sci. 2019, 40, 198–211. [Google Scholar] [CrossRef] [PubMed]
- Duijf, P.H.; Nanayakkara, D.; Nones, K.; Srihari, S.; Kalimutho, M.; Khanna, K.K. Mechanisms of genomic instability in breast cancer. Trends Mol. Med. 2019, 25, 595–611. [Google Scholar] [CrossRef]
- Mundbjerg, K.; Chopra, S.; Alemozaffar, M.; Duymich, C.; Lakshminarasimhan, R.; Nichols, P.W.; Aron, M.; Siegmund, K.D.; Ukimura, O.; Aron, M. Identifying aggressive prostate cancer foci using a DNA methylation classifier. Genome Biol. 2017, 18, 3. [Google Scholar] [CrossRef] [PubMed]
- Hatada, I. The Epigenomics of Cancer. In An Omics Perspective on Cancer Research; Cho, W., Ed.; Springer: Dordrecht, The Netherlands, 2010; pp. 51–67. [Google Scholar] [CrossRef]
- Cui, J.; Yin, Y.; Ma, Q.; Wang, G.; Olman, V.; Zhang, Y.; Chou, W.C.; Hong, C.S.; Zhang, C.; Cao, S. Comprehensive characterization of the genomic alterations in human gastric cancer. Int. J. Cancer 2015, 137, 86–95. [Google Scholar] [CrossRef] [PubMed]
- Nones, K.; Waddell, N.; Song, S.; Patch, A.M.; Miller, D.; Johns, A.; Wu, J.; Kassahn, K.S.; Wood, D.; Bailey, P. Genome-wide DNA methylation patterns in pancreatic ductal adenocarcinoma reveal epigenetic deregulation of SLIT-ROBO, ITGA2 and MET signaling. Int. J. Cancer 2014, 135, 1110–1118. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Luo, X.; Wang, J.; Wan, J.; Xia, S.; Zhu, H.; Qian, J.; Wang, Y. MeDReaders: A database for transcription factors that bind to methylated DNA. Nucleic Acids Res. 2018, 46, D146–D151. [Google Scholar] [CrossRef] [PubMed]
- Chiam, K.; Ricciardelli, C.; Bianco-Miotto, T. Epigenetic biomarkers in prostate cancer: Current and future uses. Cancer Lett. 2014, 342, 248–256. [Google Scholar] [CrossRef]
- Vitale, A.M.; Matigian, N.A.; Cristino, A.S.; Nones, K.; Ravishankar, S.; Bellette, B.; Fan, Y.; Wood, S.A.; Wolvetang, E.; Mackay-Sim, A. DNA methylation in schizophrenia in different patient-derived cell types. npj Schizophrenia 2017, 3, 1–11. [Google Scholar] [CrossRef]
- Fisher, B.; Bauer, M.; Wickerham, D.L.; Redmond, C.K.; Fisher, E.R.; Cruz, A.B.; Foster, R.; Gardner, B.; Lerner, H.; Margolese, R. Relation of number of positive axillary nodes to the prognosis of patients with primary breast cancer. An NSABP update. Cancer 2015, 52, 1551–1557. [Google Scholar] [CrossRef]
- Reyngold, M.; Turcan, S.; Giri, D.; Kannan, K.; Walsh, L.A.; Viale, A.; Drobnjak, M.; Vahdat, L.T.; Lee, W.; Chan, T.A. Remodeling of the Methylation Landscape in Breast Cancer Metastasis. PLoS ONE 2014, 9, e103896. [Google Scholar] [CrossRef]
- Jones, L.R.; Young, W.; Divine, G.; Datta, I.; Chen, K.M.; Ozog, D.; Worsham, M.J. Genome-Wide Scan for Methylation Profiles in Breast Cancer. Dis. Markers 2015, 2015, 943176. [Google Scholar] [CrossRef]
- Sandoval, J.; Heyn, H.; Moran, S.; Serra-Musach, J.; Pujana, M.A.; Bibikova, M.; Esteller, M. Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics 2016, 6, 692–702. [Google Scholar] [CrossRef]
- Phipson, B.; Lee, S.; Majewski, I.J.; Alexander, W.S.; Smyth, G.K. Robust hyperparameter estimation protects against hypervariable genes and improves power to detect differential expression. Ann. Appl. Stat. 2016, 10, 946. [Google Scholar] [CrossRef] [PubMed]
- Smyth, G.K. Linear Models and Empirical Bayes Methods for Assessing Differential Expression in Microarray Experiments. Stat. Appl Genet. Mol. Biol. 2004, 3, Article3. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Jolma, A.; Yan, J.; Whitington, T.; Toivonen, J.; Nitta, K.R.; Rastas, P.; Morgunova, E.; Enge, M.; Taipale, M.; Wei, G.; et al. DNA-binding specificities of human transcription factors. Cell 2013, 152, 327–339. [Google Scholar] [CrossRef] [PubMed]
- Zou, Q.; Zeng, J.; Cao, L.; Ji, R. A novel features ranking metric with application to scalable visual and bioinformatics data classification. Neurocomputing 2016, 173, 346–354. [Google Scholar] [CrossRef]
- Xu, Y.; Guo, M.; Liu, X.; Wang, C.; Liu, Y.; Liu, G. Identify bilayer modules via pseudo-3D clustering: Applications to miRNA-gene bilayer networks. Nucleic Acids Res. 2016, 44, e152. [Google Scholar] [CrossRef]
- Cheng, L.; Sun, J.; Xu, W.Y.; Dong, L.X.; Hu, Y.; Zhou, M. OAHG: An integrated resource for annotating human genes with multi-level ontologies. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef]
- Cheng, L.; Jiang, Y.; Wang, Z.; Shi, H.; Sun, J.; Yang, H.; Zhang, S.; Hu, Y.; Zhou, M. DisSim: An online system for exploring significant similar diseases and exhibiting potential therapeutic drugs. Sci. Rep. 2016, 6, 30024. [Google Scholar] [CrossRef]
- Peng, H.; Long, F.; Ding, C. Feature Selection Based on Mutual Information: Criteria of Max-Dependency, Max-Relevance, and Min-Redundancy. Ieee Trans. Pattern Anal. Mach. Intell. 2005, 27, 1226–1238. [Google Scholar] [CrossRef]
- Ding, C.; Peng, H. Minimum Redundancy Feature Selection from Microarray Gene Expression Data. J. Bioinform. Comput. Biol. 2005, 3, 185–205. [Google Scholar] [CrossRef]
- Tang, Y.; Liu, D.; Wang, Z.; Wen, T.; Deng, L. A boosting approach for prediction of protein-RNA binding residues. Bmc Bioinform. 2017, 18, 465. [Google Scholar] [CrossRef]
- Liu, B. BioSeq-Analysis: A platform for DNA, RNA and protein sequence analysis based on machine learning approaches. Brief. Bioinform. 2017, 20, 1280–1294. [Google Scholar] [CrossRef]
- Dao, F.Y.; Lv, H.; Wang, F.; Feng, C.Q.; Ding, H.; Chen, W.; Lin, H. Identify origin of replication in Saccharomyces cerevisiae using two-step feature selection technique. Bioinformatics 2019, 35, 2075–2083. [Google Scholar] [CrossRef]
- Chen, W.; Feng, P.; Liu, T.; Jin, D. Recent advances in machine learning methods for predicting heat shock proteins. Curr. Drug Metab. 2018, 20, 224–228. [Google Scholar] [CrossRef]
- Tipping, M.E.; Bishop, C.M. Probabilistic Principal Component Analysis. J. R. Stat. Soc. 1999, 61, 611–622. [Google Scholar] [CrossRef]
- Minka, T.P. Automatic choice of dimensionality for PCA. In Proceedings of the International Conference on Neural Information Processing Systems, Montreal, QC, Canada, 29 December 2000; pp. 577–583. [Google Scholar]
- Pedregosa, F.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; Vanderplas, J. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2013, 12, 2825–2830. [Google Scholar]
- Gorsuch, R.L. Factor Analysis; Lawrence Erlbaum Associates: Mahwah, NJ, USA, 1983. [Google Scholar]
- Chen, S.; Doolen, G.D. Lattice Boltzmann method for fluid flows. Annu. Rev. Fluid Mech. 1998, 30, 329–364. [Google Scholar] [CrossRef]
- Johnson, N.L.; Kotz, S.; Kemp, A.W. Univariate Discrete Distributions; John Wiley & Sons: Hoboken, NJ, USA, 1992. [Google Scholar]
- Haffner, M.C.; Mosbruger, T.; Esopi, D.M.; Fedor, H.; Heaphy, C.M.; Walker, D.A.; Adejola, N.; Gürel, M.; Hicks, J.; Meeker, A.K. Tracking the clonal origin of lethal prostate cancer. J. Clin. Investig. 2013, 123, 4918–4922. [Google Scholar] [CrossRef]
- Marino, N.; Woditschka, S.; Reed, L.T.; Nakayama, J.; Mayer, M.; Wetzel, M.; Steeg, P.S. Breast Cancer Metastasis. Am. J. Pathol. 2013, 183, 1084–1095. [Google Scholar] [CrossRef]
- Zheng, G.; Ma, Y.; Zou, Y.; Yin, A.; Li, W.; Dong, D. HCMDB: The human cancer metastasis database. Nucleic Acids Res. 2018, 46, D950–D955. [Google Scholar] [CrossRef]
- Wang, C.; Yuan, L. BRCA Methylation Metastasis Prediction. Available online: http://server.malab.cn/BMMP/ (accessed on 30 January 2020).
- Jahid, M.J.; Huang, T.H.; Ruan, J. A personalized committee classification approach to improving prediction of breast cancer metastasis. Bioinformatics 2014, 30, 1858–1866. [Google Scholar] [CrossRef][Green Version]
- Zeng, X.X.; Liu, L.; Lu, L.Y.; Zou, Q. Prediction of potential disease-associated microRNAs using structural perturbation method. Bioinformatics 2018, 34, 2425–2432. [Google Scholar] [CrossRef]
- Ding, Y.; Tang, J.; Guo, F. Identification of drug-target interactions via multiple information integration. Inf. Sci. 2017, 418, 546–560. [Google Scholar] [CrossRef]
- Xiao, Y.; Zhang, J.; Deng, L. Prediction of lncRNA-protein interactions using HeteSim scores based on heterogeneous networks. Sci. Rep. 2017, 7, 3664. [Google Scholar] [CrossRef]
- Zhang, X.; Zou, Q.; Rodriguez-Paton, A.; Zeng, X.X. Meta-Path Methods for Prioritizing Candidate Disease miRNAs. IEEE-Acm Trans. Comput. Biol. Bioinform. 2019, 16, 283–291. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Q.; Wang, G.; Jin, S.; Li, Y.; Wang, Y. Predicting human microRNA-disease associations based on support vector machine. Int J. Data Min. Bioinform 2013, 8, 282–293. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Li, S.C.; Du, P.; Wang, L. Probabilistic Models for Capturing More Physicochemical Properties on Protein-Protein Interface. J. Chem. Inf. Modeling 2014, 54, 1798–1809. [Google Scholar] [CrossRef]
- Ding, Y.; Tang, J.; Guo, F. Identification of Protein-Ligand Binding Sites by Sequence Information and Ensemble Classifier. J. Chem. Inf. Modeling 2017, 57, 3149–3161. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Wang, Y.; Feng, W.; Wang, X.; Yang, J.Y.; Zhao, Y.; Wang, Y.; Liu, Y. Transcription factor and microRNA regulation in androgen-dependent and -independent prostate cancer cells. Bmc Genom. 2008, 9 Suppl 2, S22. [Google Scholar] [CrossRef]
- Wang, G.; Wang, Y.; Teng, M.; Zhang, D.; Li, L.; Liu, Y. Signal transducers and activators of transcription-1 (STAT1) regulates microRNA transcription in interferon gamma-stimulated HeLa cells. PLoS ONE 2010, 5, e11794. [Google Scholar] [CrossRef]
- Cabarle, F.G.C.; Adorna, H.N.; Jiang, M.; Zeng, X. Spiking Neural P Systems With Scheduled Synapses. IEEE Trans. Nanobioscience 2017, 16, 792–801. [Google Scholar] [CrossRef] [PubMed]
- Dong, M.H.; Wen, S.P.; Zeng, Z.G.; Yan, Z.; Huang, T.W. Sparse fully convolutional network for face labeling. Neurocomputing 2019, 331, 465–472. [Google Scholar] [CrossRef]
- Li, Z.L.; Dong, M.H.; Wen, S.P.; Hu, X.; Zhou, P.; Zeng, Z.G. CLU-CNNs: Object detection for medical images. Neurocomputing 2019, 350, 53–59. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, F.; Juan, L. MicroRNA Promoter Identification in Arabidopsis Using Multiple Histone Markers. Biomed. Res. Int 2015, 2015, 861402. [Google Scholar] [CrossRef] [PubMed]
- Zou, Q.; Li, J.; Song, L.; Zeng, X.; Wang, G. Similarity computation strategies in the microRNA-disease network: A Survey. Brief. Funct. Genom. 2016, 15, 55–64. [Google Scholar] [CrossRef]
- Zeng, X.; Ding, N.; Rodríguezpatón, A.; Quan, Z.J.B.M.G. Probability-based collaborative filtering model for predicting gene–disease associations. Bmc Med. Genom. 2017, 10, 76. [Google Scholar] [CrossRef]
- Ding, Y.; Tang, J.; Guo, F. Identification of Protein-Protein Interactions via a Novel Matrix-Based Sequence Representation Model with Amino Acid Contact Information. Int. J. Mol. Sci. 2016, 17, 1623. [Google Scholar] [CrossRef]
- Cheng, L.; Yang, H.; Zhao, H.; Pei, X.; Shi, H.; Sun, J.; Zhang, Y.; Wang, Z.; Zhou, M. MetSigDis: A manually curated resource for the metabolic signatures of diseases. Brief. Bioinform. 2017, 20, 203–209. [Google Scholar] [CrossRef]
- Pan, Y.W.; Wang, Z.; Zhan, W.; Deng, L. Computational identification of binding energy hot spots in protein-RNA complexes using an ensemble approach. Bioinformatics 2018, 34, 1473–1480. [Google Scholar] [CrossRef]
- Liu, G.; Jin, S.; Hu, Y.; Jiang, Q. Disease status affects the association between rs4813620 and the expression of Alzheimer’s disease susceptibility gene TRIB3. Proc. Natl. Acad. Sci. USA 2018, 115, E10519–E10520. [Google Scholar] [CrossRef]
- Liu, G.; Hu, Y.; Han, Z.; Jin, S.; Jiang, Q. Genetic variant rs17185536 regulates SIM1 gene expression in human brain hypothalamus. Proc. Natl. Acad. Sci. USA 2019, 116, 3347–3348. [Google Scholar] [CrossRef]
- Wang, X.; Yu, B.; Ma, A.; Chen, C.; Liu, B.; Ma, Q. Protein–protein interaction sites prediction by ensemble random forests with synthetic minority oversampling technique. Bioinformatics 2018, 35, 2395–2402. [Google Scholar] [CrossRef]
- Liu, G.; Wang, T.; Tian, R.; Hu, Y.; Han, Z.; Wang, P.; Zhou, W.; Ren, P.; Zong, J.; Jin, S.; et al. Alzheimer’s Disease Risk Variant rs2373115 Regulates GAB2 and NARS2 Expression in Human Brain Tissues. J. Mol. Neurosci. 2018, 66, 37–43. [Google Scholar] [CrossRef]
- Liu, G.; Xu, Y.; Jiang, Y.; Zhang, L.; Feng, R.; Jiang, Q. PICALM rs3851179 Variant Confers Susceptibility to Alzheimer’s Disease in Chinese Population. Mol. Neurobiol. 2017, 54, 3131–3136. [Google Scholar] [CrossRef]
| Normal | Invasiveness | ||
|---|---|---|---|
| MRMD | Number of CpG | 14 | 134 |
| Training Accuracy | 97% | 93.6% | |
| Testing Accuracy | 96.9% | 549/217 | |
| mRMR | Number of CpG | 12 | 5 |
| Training Accuracy | 99% | 100% | |
| Testing Accuracy | 96.9% | 611/165 | |
| PCA | Number of CpG | 8 | 3 |
| Training Accuracy | 99% | 95% | |
| Testing Accuracy | 91% | 454/312 | |
| FA | Number of CpG | 80 | 60 |
| Training Accuracy | 99% | 93.3% | |
| Testing Accuracy | 94.8% | 664/102 |
| Gene | Description |
|---|---|
| ABCC5 | ABCC5 functions as a mediator of breast cancer skeletal metastasis. |
| ASCL2 | Functions as a suppressor of colorectal cancer metastasis by down-regulating the ASCL2-CXCR4 signaling axis. |
| BNIP3 | “BNIP3 deletion can be used as a prognostic marker of tumor progression to metastasis in human triple-negative breast cancer” |
| FLI1 | “This study for the first time identifies FLI1 as a clinically and functionally important target gene of metastasis, providing a rationale for developing FLI1 inhibitors in the treatment of breast cancer.” |
| ITGA6 | “The role of PTHrP in breast cancer growth and metastasis may be mediated via upregulation of integrin alpha6beta4 expression and Akt activation, with consequent inactivation of GSK-3.” |
| MPL | “In migrating cancer stem cells isolated from primary human colorectal cancers, CD110(+) and CDCP1(+) subpopulations mediate organ-specific lung and liver metastasis.” |
| NCOR2 | “Thyroid hormone receptors induce TRAIL expression, and TRAIL thus synthesized acts in concert with simultaneously synthesized Bcl-xL to promote metastasis” |
| RHOB | “RHOB belongs to a novel class of ““genes of recurrence”“ that have a dual role in metastasis and treatment resistance.” |
| SLITRK3 | SLITRK3 expression is a highly significant predictor of gastrointestinal stromal tumor recurrence and metastasis. |
| SND1 | “SND1 is a novel MTDH-interacting protein and has shown that it is a functionally and clinically significant mediator of metastasis.” |
| TRPS1 | “TRPS1 plays a crucial role in osteosarcoma angiogenesis, metastasis and clinical surgical stage.” |
| WWOX | WWOX is associated with tumorigenicity and metastasis of head and neck and gastric signet-ring cell carcinoma. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, C.; Zhao, N.; Yuan, L.; Liu, X. Computational Detection of Breast Cancer Invasiveness with DNA Methylation Biomarkers. Cells 2020, 9, 326. https://doi.org/10.3390/cells9020326
Wang C, Zhao N, Yuan L, Liu X. Computational Detection of Breast Cancer Invasiveness with DNA Methylation Biomarkers. Cells. 2020; 9(2):326. https://doi.org/10.3390/cells9020326
Chicago/Turabian StyleWang, Chunyu, Ning Zhao, Linlin Yuan, and Xiaoyan Liu. 2020. "Computational Detection of Breast Cancer Invasiveness with DNA Methylation Biomarkers" Cells 9, no. 2: 326. https://doi.org/10.3390/cells9020326
APA StyleWang, C., Zhao, N., Yuan, L., & Liu, X. (2020). Computational Detection of Breast Cancer Invasiveness with DNA Methylation Biomarkers. Cells, 9(2), 326. https://doi.org/10.3390/cells9020326






