The Role of Brachypodium distachyon Wall-Associated Kinases (WAKs) in Cell Expansion and Stress Responses
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. RNAseq Analysis of B. distachyon Coleoptiles
2.3. Protein Sequence and Phylogenetic Analysis
2.4. Quantitative PCR and Data Analysis
2.5. PCR Amplification and Cloning of Genes of Interest
2.6. Expression of Recombinant Proteins in E. coli and Protein Purification
2.7. Kinase Activity Assay with Pro-Q Staining
2.8. Cell Wall Polysaccharide Extraction and In Vitro Binding Assay
3. Results
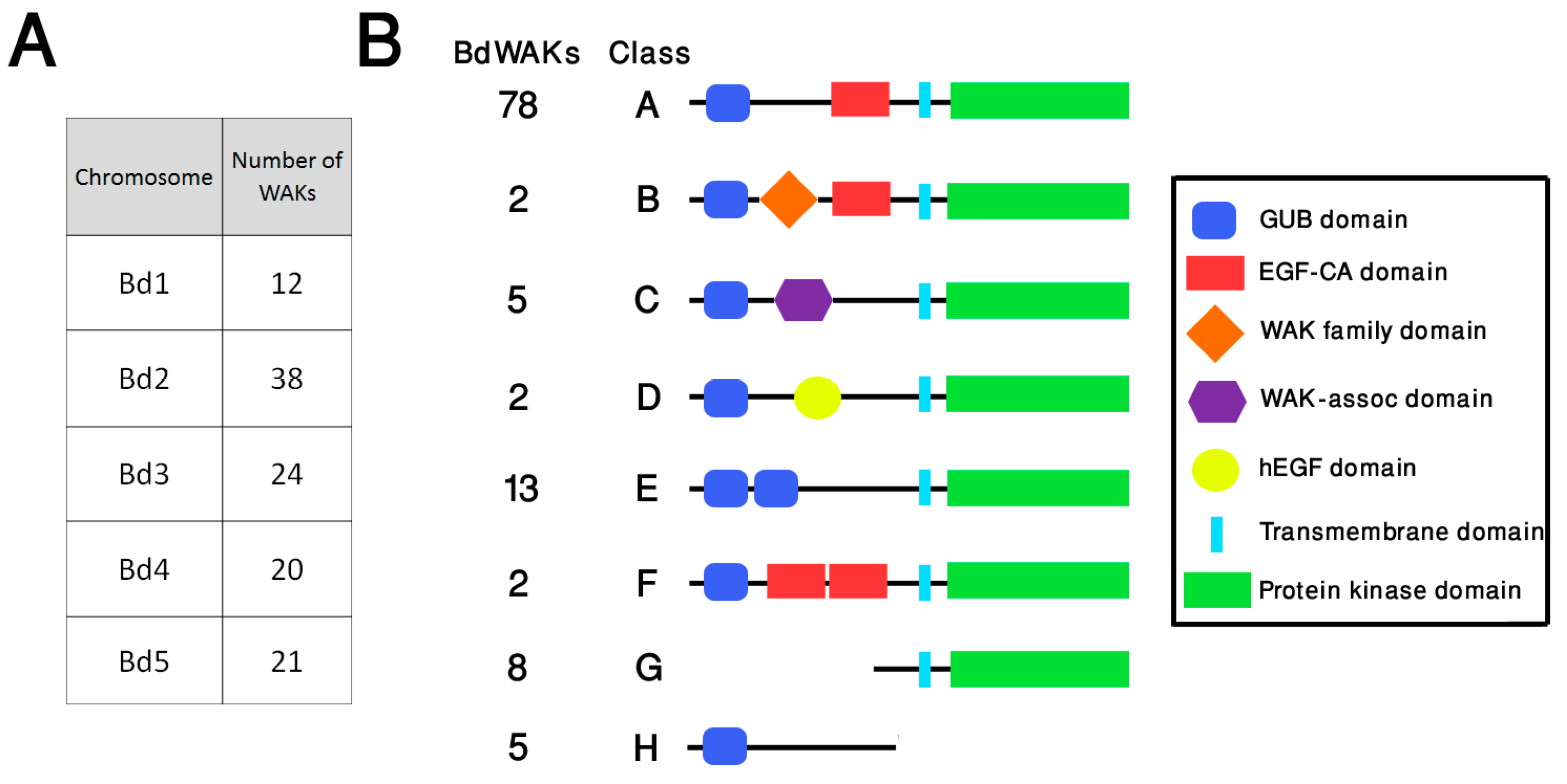
3.1. Identification and Analysis of the BdWAK Family
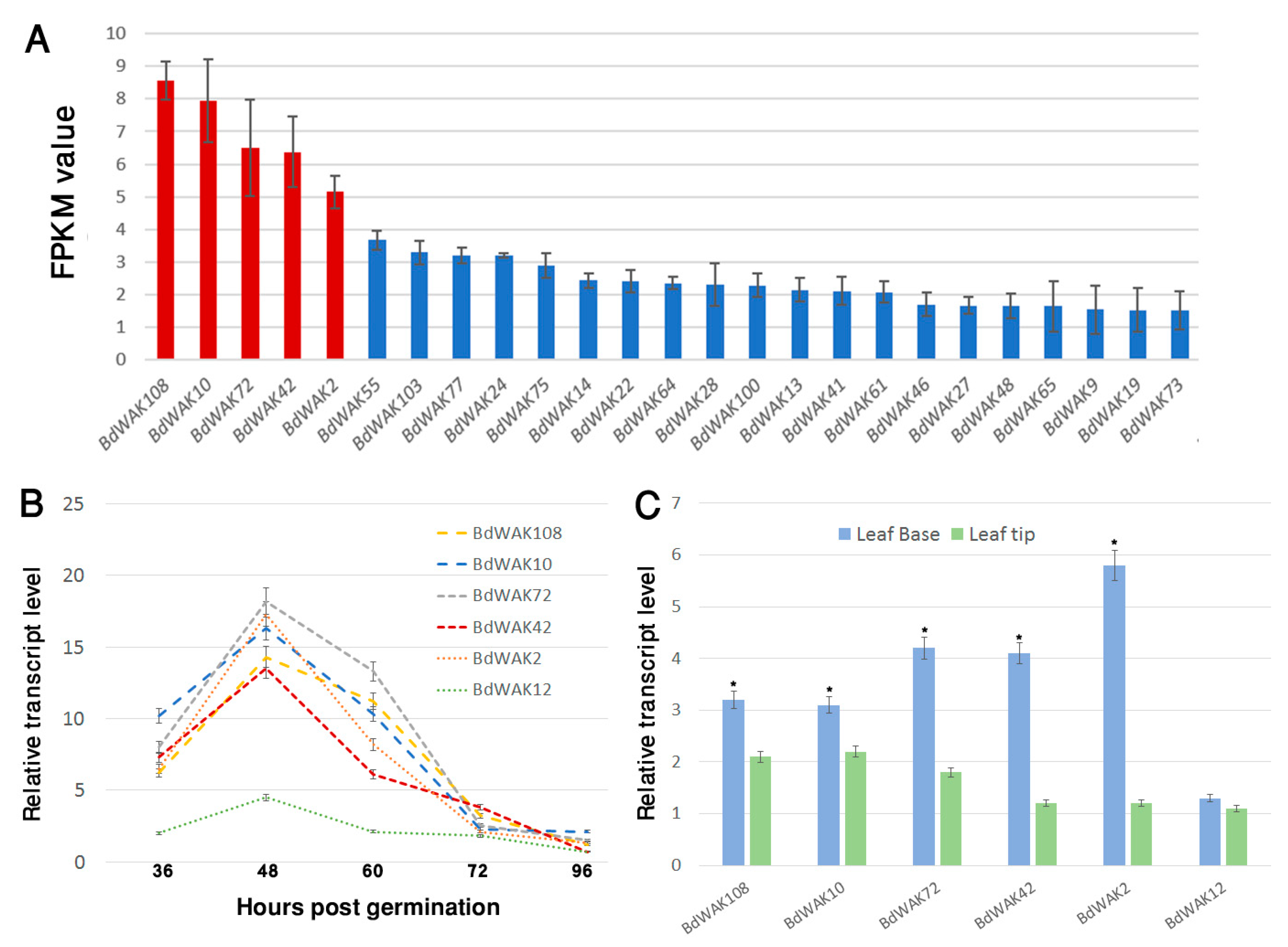
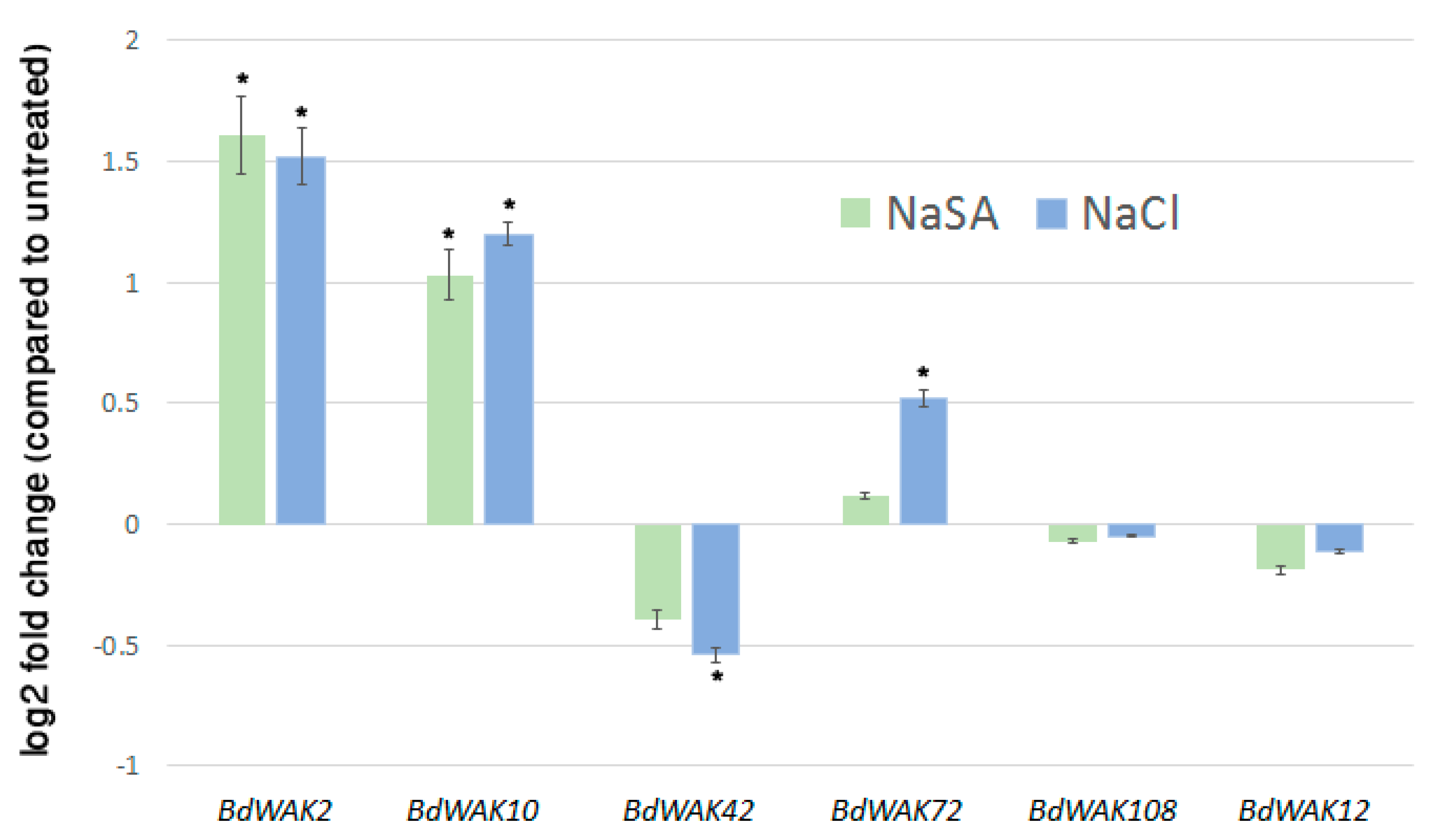
3.2. Expression Profiling of B. distachyon WAK Genes
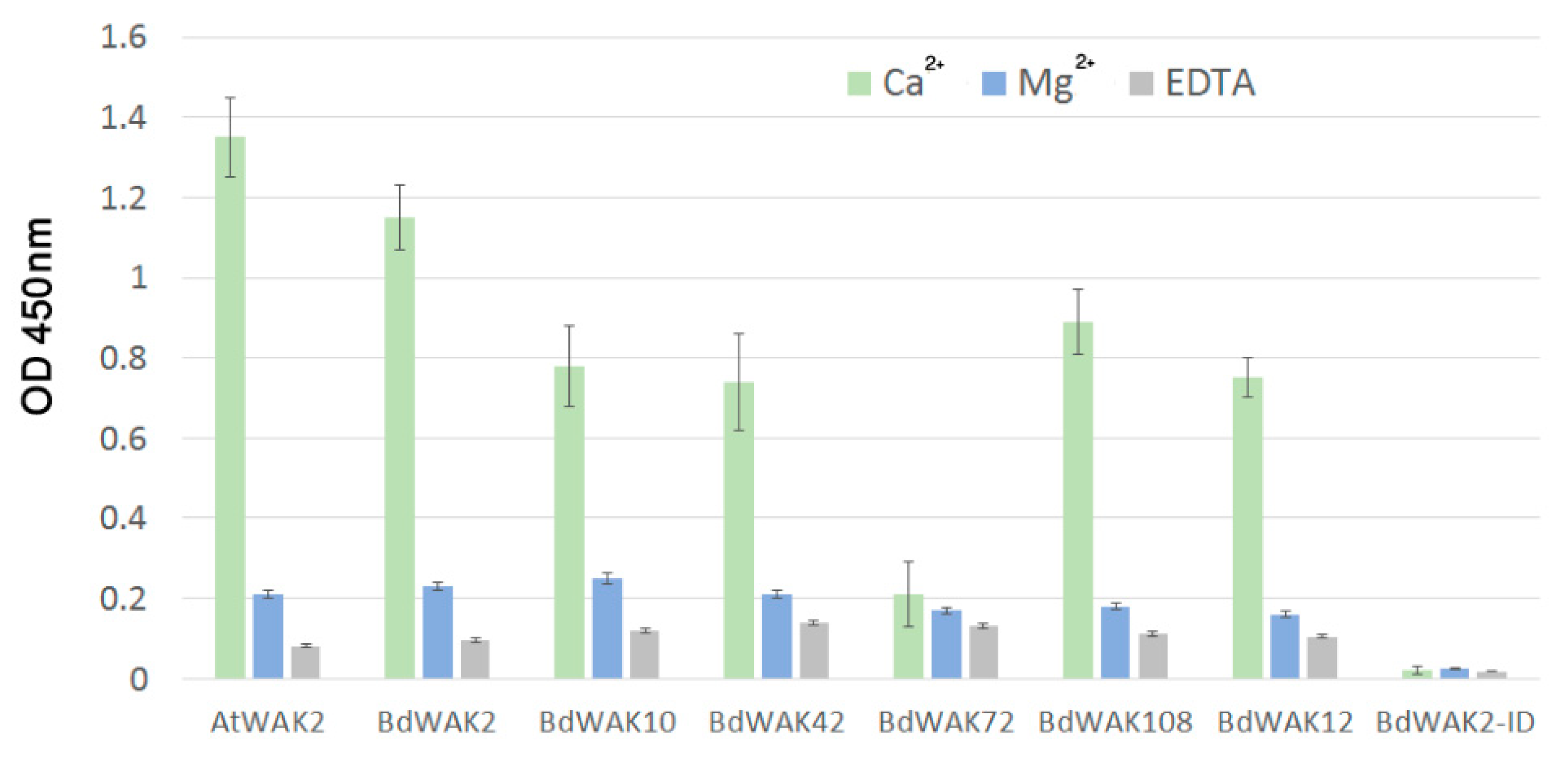
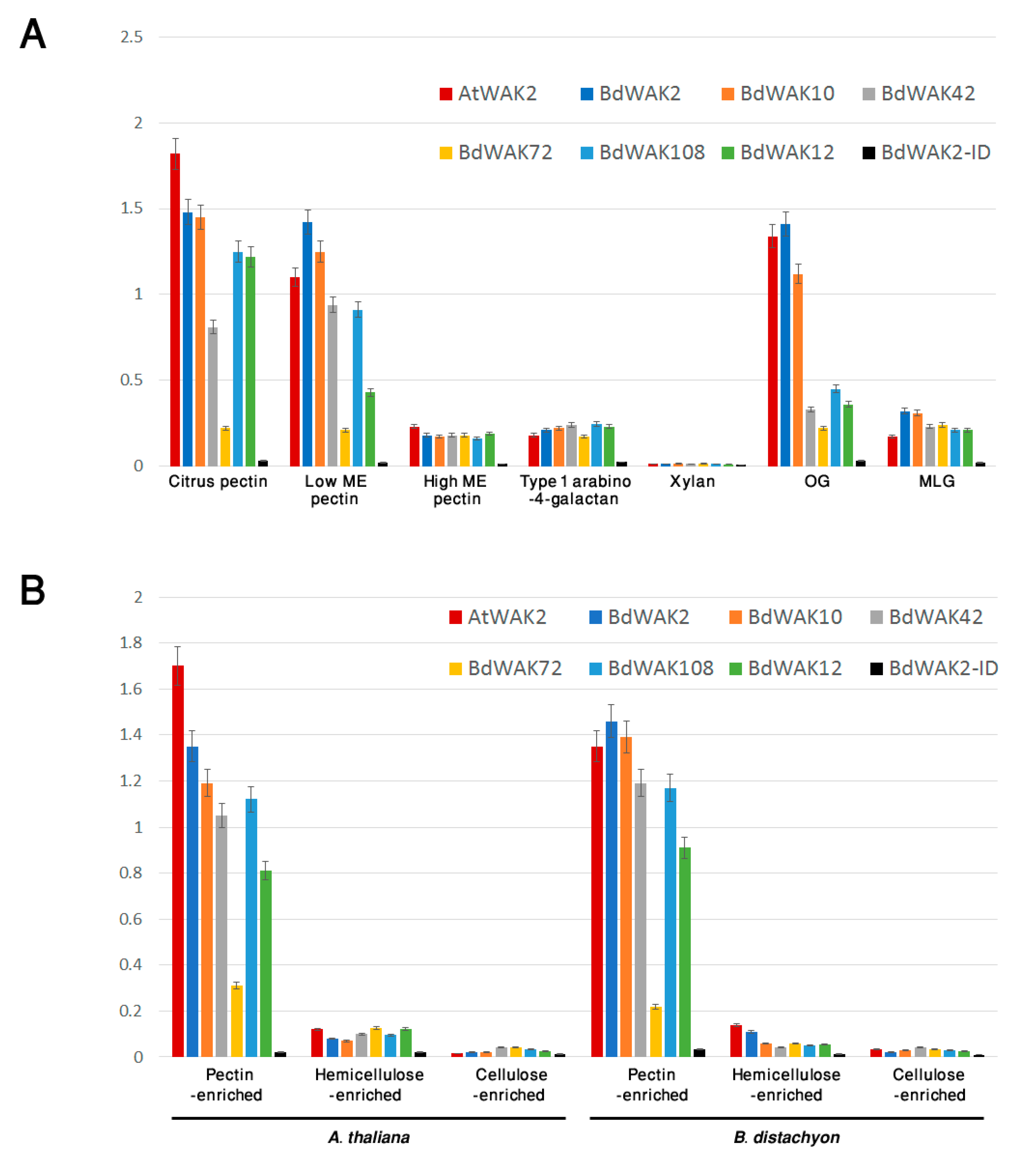
3.3. BdWAK Extracellular Domain In Vitro Binding Assay
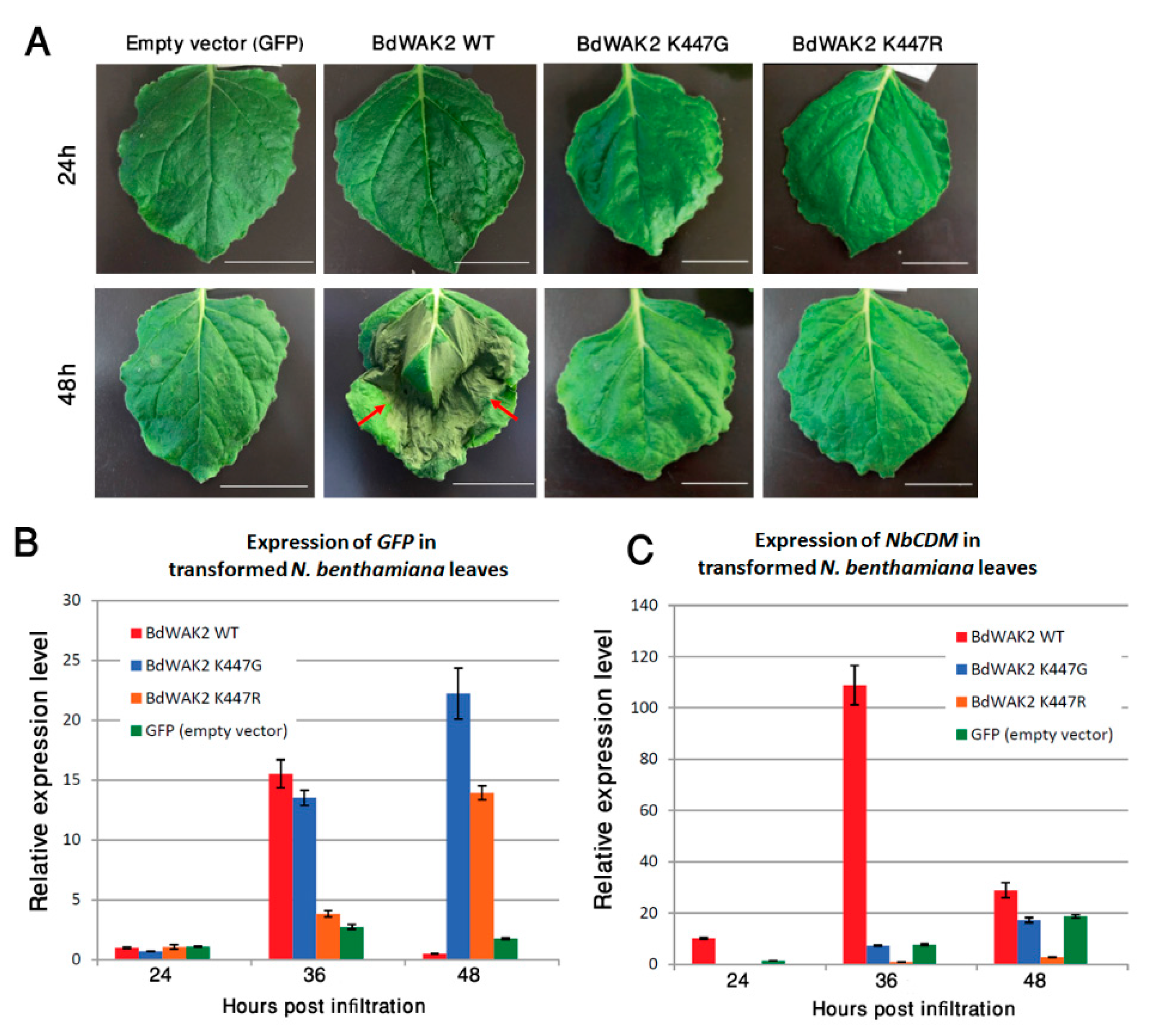
3.4. BdWAK2 Triggers Cell Death in N. benthamiana Leaves
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Novaković, L.; Guo, T.; Bacic, A.; Sampathkumar, A.; Johnson, K.L. Hitting the wall-sensing and signaling pathways involved in plant cell wall remodeling in response to abiotic stress. Plants 2018, 7, 89. [Google Scholar] [CrossRef] [PubMed]
- Monaghan, J.; Zipfel, C. Plant pattern recognition receptor complexes at the plasma membrane. Curr. Opin. Plant Biol. 2012, 15, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Zipfel, C. Plant pattern-recognition receptors. Trends Immunol. 2014, 35, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef] [PubMed]
- Levin, D.E. Regulation of cell wall biogenesis in Saccharomyces cerevisiae: The cell wall integrity signaling pathway. Genetics 2011, 189, 1145–1175. [Google Scholar] [CrossRef] [PubMed]
- Hamann, T. Plant cell wall integrity maintenance as an essential component of biotic stress response mechanisms. Front. Plant Sci. 2012, 3, 1–5. [Google Scholar] [CrossRef]
- Wolf, S. Plant cell wall signalling and receptor-like kinases. Biochem. J. 2017, 474, 471–492. [Google Scholar] [CrossRef]
- Williamson, R. Alignment of Cortical Microtubules by Anisotropic Wall Stresses. Funct. Plant Biol. 1990, 17, 601–613. [Google Scholar] [CrossRef]
- Vogel, V.; Sheetz, M.P. Local force and geometry sensing regulate cell functions. Nat. Rev. Mol. Cell Biol. 2006, 7, 265–275. [Google Scholar] [CrossRef]
- Wolf, S.; Hematy, K.; Höfte, H. Growth control and cell wall signaling in plants. Annu. Rev. Plant Biol. 2012, 63, 381–407. [Google Scholar] [CrossRef]
- Ringli, C. Monitoring the outside: Cell wall-sensing mechanisms. Plant Physiol. 2010, 153, 1445–1452. [Google Scholar] [CrossRef] [PubMed]
- Kohorn, B.D.; Kohorn, S.L. The cell wall-associated kinases, WAKs, as pectin receptors. Front. Plant Sci. 2012, 3, 80–84. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.L.; Rahman, A.; Baskin, T.I.; Kieber, J.J. Two leucine-rich repeat receptor kinases mediate signaling, linking cell wall biosynthesis and ACC synthase in arabidopsis. Plant Cell 2008, 20, 3065–3079. [Google Scholar] [CrossRef] [PubMed]
- Harpaz-Saad, S.; Western, T.L.; Kieber, J.J. The FEI2-SOS5 pathway and CELLULOSE SYNTHASE 5 are required for cellulose biosynthesis in the Arabidopsis seed coat and affect pectin mucilage structure. Plant Signal. Behav. 2012, 7, 285–288. [Google Scholar] [CrossRef]
- Hématy, K.; Sado, P.E.; Van Tuinen, A.; Rochange, S.; Desnos, T.; Balzergue, S.; Pelletier, S.; Renou, J.P.; Höfte, H. A receptor-like kinase mediates the response of arabidopsis cells to the inhibition of cellulose synthesis. Curr. Biol. 2007, 17, 922–931. [Google Scholar] [CrossRef]
- He, Z.H.; Fujiki, M.; Kohorn, B.D. A cell wall-associated, receptor-like protein kinase. J. Biol. Chem. 1996, 271, 19789–19793. [Google Scholar] [CrossRef]
- Wagner, T.A.; Kohorn, B.D. Wall-associated kinases are expressed throughout plant development and are required for cell expansion. Plant Cell 2001, 13, 303–318. [Google Scholar] [CrossRef]
- Decreux, A.; Messiaen, J. Wall-associated kinase WAK1 interacts with cell wall pectins in a calcium-induced conformation. Plant Cell Physiol. 2005, 46, 268–278. [Google Scholar] [CrossRef]
- Verica, J.A.; He, Z.H. The cell wall-associated kinase (WAK) andWAK-like kinase gene family. Plant Physiol. 2002, 129, 455–459. [Google Scholar] [CrossRef]
- He, Z.H.; Cheesem, I.; He, D.; Kohorn, B.D. A cluster of five cell wall-associated receptor kinase genes, Wak1–5, are expressed in specific organs of Arabidopsis. Plant Mol. Biol. 1999, 39, 1189–1196. [Google Scholar] [CrossRef]
- He, Z.; He, D.; Kohorn, B.D. Requirement for the induced expression of a cell wall associated receptor kinase for survival during the pathogen response. Plant J. 1998, 14, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Verica, J.A.; Chae, L.; Tong, H.; Ingmire, P.; He, Z.-H. Tissue-Specific and Developmentally Regulated Expression of a Cluster of Tandemly Arrayed Cell Wall-Associated Kinase-Like Kinase Genes in Arabidopsis. Plant Physiol. 2003, 133, 1732–1746. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Tong, H.; Selby, J.; DeWitt, J.; Peng, X.; He, Z.H. Involvement of a cell wall-associated kinase, WAKL4, in arabidopsis mineral responses. Plant Physiol. 2005, 139, 1704–1716. [Google Scholar] [CrossRef] [PubMed]
- Decreux, A.; Thomas, A.; Spies, B.; Brasseur, R.; Cutsem, P.; Messiaen, J. In vitro characterization of the homogalacturonan-binding domain of the wall-associated kinase WAK1 using site-directed mutagenesis. Phytochemistry 2006, 67, 1068–1079. [Google Scholar] [CrossRef]
- Brutus, A.; Sicilia, F.; Macone, A.; Cervone, F.; Lorenzo, G.D. A domain swap approach reveals a role of the plant wall-associated kinase 1 (WAK1) as a receptor of oligogalacturonides. Proc. Natl. Acad. Sci. USA 2010, 107, 9452–9457. [Google Scholar] [CrossRef]
- Yang, K.; Qi, L.; Zhang, Z. Isolation and characterization of a novel wall-associated kinase gene TaWAK5 in wheat (Triticum aestivum). Crop. J. 2014, 2, 255–266. [Google Scholar] [CrossRef]
- Zuo, W.; Chao, Q.; Zhang, N.; Ye, J.; Tan, G.; Li, B.; Xing, Y.; Zhang, B.; Liu, H.; A Fengler, K.; et al. A maize wall-associated kinase confers quantitative resistance to head smut. Nat. Genet. 2015, 47, 151–157. [Google Scholar] [CrossRef]
- Li, H.; Zhou, S.-Y.; Zhao, W.-S.; Su, S.-C.; Peng, Y.-L. A novel wall-associated receptor-like protein kinase gene, OsWAK1, plays important roles in rice blast disease resistance. Plant Mol. Biol. 2008, 69, 337–346. [Google Scholar] [CrossRef]
- Hu, W.; Lv, Y.; Lei, W.; Li, X.; Chen, Y.; Zheng, L.; Xia, Y.; Shen, Z. Cloning and characterization of the Oryza sativa wall-associated kinase gene OsWAK11 and its transcriptional response to abiotic stresses. Plant Soil 2014, 384, 335–346. [Google Scholar] [CrossRef]
- Wang, N.; Huang, H.J.; Ren, S.T.; Li, J.J.; Sun, Y.; Sun, D.Y.; Zhang, S.Q. The rice wall-associated receptor-like kinase gene OsDEES1 plays a role in female gametophyte development. Plant Physiol. 2012, 160, 696–707. [Google Scholar] [CrossRef]
- Kaur, R.; Singh, K.; Singh, J. A root-specific wall-associated kinase gene, HvWAK1, regulates root growth and is highly divergent in barley and other cereals. Funct. Integr. Genom. 2013, 13, 167–177. [Google Scholar] [CrossRef] [PubMed]
- Dardick, C.; Schwessinger, B.; Ronald, P. Non-arginine-aspartate (non-RD) kinases are associated with innate immune receptors that recognize conserved microbial signatures. Curr. Opin. Plant Biol. 2012, 15, 358–366. [Google Scholar] [CrossRef] [PubMed]
- Bacic, A.; Harris, P.J.; Stone, B.A. Structure and function of plant cell walls. Biochem. Plants 1988, 14, 297–371. [Google Scholar]
- Vogel, J.P. Unique aspects of the grass cell wall. Curr. Opin. Plant Biol. 2008, 11, 301–307. [Google Scholar] [CrossRef]
- Doblin, M.S.; Pettolino, F.; Bacic, A. Plant cell walls: the skeleton of the plant world. Funct. Plant Biol. 2010, 37, 357–381. [Google Scholar] [CrossRef]
- Doblin, M.S.; Johnson, K.L.; Humphries, J.; Newbigin, E.J.; Bacic, A. Are designer plant cell walls a realistic aspiration or will the plasticity of the plant’s metabolism win out? Curr. Opin. Biotechnol. 2014, 26, 108–114. [Google Scholar] [CrossRef]
- Zhang, N.; Zhang, B.; Zuo, W.; Xing, Y.; Konlasuk, S.; Tan, G.; Zhang, Q.; Ye, J.; Xu, M. Cytological and Molecular Characterization of ZmWAK-Mediated Head-Smut Resistance in Maize. Mol. Plant Microbe. Interact. 2017, 30, 455–465. [Google Scholar] [CrossRef]
- Hoagland, D.R. The Water-Culture Method for Growing Plants without Soil; Arnon, D.I., Ed.; College of Agriculture, University of California: Berkeley, CA, USA, 1950. [Google Scholar]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Altschul, S.; Gish, W.; Miller, W.; Myers, E.; Lipman, D. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Jukes, T.H.; Cantor, C.R. Evolution of protein molecules. In Mammalian Protein Metabolism; Munro, H.N., Ed.; Academic Press: New York, NY, USA, 1969; pp. 21–132. [Google Scholar]
- Burton, R.A.; Shirley, N.J.; King, B.; Harvey, A.J.; Fincher, G.B. The CesA gene family of barley. quantitative analysis of transcripts reveals two groups of co-expressed genes. Plant Physiol. 2004, 134, 224–236. [Google Scholar] [CrossRef]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Geldner, N.; Hyman, D.L.; Wang, X.; Schumacher, K.; Chory, J. Endosomal signaling of plant steroid receptor kinase BRI1. Genes Dev. 2007, 21, 1598–1602. [Google Scholar] [CrossRef]
- Grefen, C.; Donald, N.; Hashimoto, K.; Kudla, J.; Schumacher, K.; Blatt, M. A ubiquitin-10 promoter-based vector set for fluorescent protein tagging facilitates temporal stability and native protein distribution in transient and stable expression studies. Plant J. 2010, 64, 355–365. [Google Scholar] [CrossRef] [PubMed]
- Hartley, J.L.; Temple, G.F.; Brasch, M.A. DNA cloning using in vitro site-specific recombination. Genome Res. 2000, 10, 1788–1795. [Google Scholar] [CrossRef]
- Taylor, I.; Seitz, K.; Bennewitz, S.; Walker, J.C. A simple in vitro method to measure autophosphorylation of protein kinases. Plant Methods 2013, 9, 22. [Google Scholar] [CrossRef]
- Pettolino, F.A.; Walsh, C.; Fincher, G.B.; Bacic, A. Determining the polysaccharide composition of plant cell walls. Nat. Protoc. 2012, 7, 1590–1607. [Google Scholar] [CrossRef]
- Woodward, J.; Phillips, D.; Fincher, G. Water-soluble (1 → 3,1 → 4)-β-d-glucans from barley (Hordeum vulgare) endosperm. IV. Comparison of 40°C and 65°C soluble fractions. Carbohydr. Polym. 1988, 8, 85–97. [Google Scholar] [CrossRef]
- Kohorn, B.D.; Hoon, D.; Minkoff, B.B.; Sussman, M.R.; Kohorn, S.L. Rapid oligo-galacturonide induced changes in protein phosphorylation in arabidopsis. Mol. Cell. Proteom. 2016, 15, 1351–1359. [Google Scholar] [CrossRef] [PubMed]
- Wouters, M.A.; Rigoutsos, I.; Chu, C.K.; Feng, L.L.; Sparrow, D.B.; Dunwoodie, S.L. Evolution of distinct EGF domains with specific functions. Protein Sci. 2005, 14, 1091–1103. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Chen, C.; Ling, M.; Meng, L.; Singh, J.; Jiang, N.; Deng, X.-W.; He, Z.-H.; Lemaux, P.G. Evolutionary expansion, gene structure, and expression of the rice wall-associated kinase gene family. Plant Physiol. 2005, 139, 1107–1124. [Google Scholar] [CrossRef] [PubMed]
- Esau, K. Plant Anatomy, 2nd ed.; John Wiley and Sons: New York, NY, USA, 1965. [Google Scholar]
- Handakumbura, P.P.; Matos, D.A.; Osmont, K.S.; Harrington, M.J.; Heo, K.; Kafle, K.; Kim, S.; Baskin, T.I.; Hazen, S.P. Perturbation of brachypodium distachyon CELLULOSE SYNTHASE A4 or 7 results in abnormal cell walls. BMC Plant Biol. 2013, 13, 131. [Google Scholar] [CrossRef]
- Brabham, C.; Singh, A.; Stork, J.; Rong, Y.; Kumar, I.; Kikuchi, K.; Yingling, Y.G.; Brutnell, T.P.; Rose, J.K.C.; DeBolt, S. Biochemical and physiological flexibility accompanies reduced cellulose biosynthesis in Brachypodium cesa1S830N. AoB PLANTS 2019, 11, plz041. [Google Scholar] [CrossRef]
- Sivaguru, M.; Ezaki, B.; He, Z.-H.; Tong, H.; Osawa, H.; Baluška, F.; Volkmann, D.; Matsumoto, H. Aluminum-induced gene expression and protein localization of a cell wall-associated receptor kinase in arabidopsis. Plant Physiol. 2003, 132, 2256–2266. [Google Scholar] [CrossRef]
- Kohorn, B.D.; Johansen, S.; Shishido, A.; Todorova, T.; Martinez, R.; Defeo, E.; Obregon, P. Pectin activation of MAP kinase and gene expression is WAK2 dependent. Plant J. 2009, 60, 974–982. [Google Scholar] [CrossRef]
- Liners, F.; Letesson, J.J.; Didembourg, C.; Van Cutsem, P. Monoclonal antibodies against pectin. Plant Physiol. 1989, 91, 1419–1424. [Google Scholar] [CrossRef]
- Hanks, S.K.; Hunter, T. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification 1. FASEB J. 1995, 9, 576–596. [Google Scholar] [CrossRef]
- Hanks, S.K.; Quinn, A.M.; Hunter, T. The protein kinase family: conserved features and deduced phylogeny of the catalytic domains. Science 1988, 241, 42–52. [Google Scholar] [CrossRef]
- Kim, M.; Lim, J.H.; Ahn, C.S.; Park, K.; Kim, G.T.; Kim, W.T.; Pai, H.S. Mitochondria-associated hexokinases play a role in the control of programmed cell death in nicotiana benthamiana. Plant Cell 2006, 18, 2341–2355. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, R.K.; Aguirre, J.A.; Singh, J. Genome-wide analysis of wall associated kinase (WAK) gene family in barley. Genomics 2020, in press. [Google Scholar] [CrossRef] [PubMed]
- Tocquard, K.; Placette, C.L.; Auguin, D.; Muries, B.; Bronner, G.; Lopez, D.; Fumanal, B.; Franchel, J.; Bourgerie, S.; Maury, S.; et al. In silico study of wall-associated kinase family reveals large-scale genomic expansion potentially connected with functional diversification in populus. Tree Genet. Genomes 2014, 10, 1135–1147. [Google Scholar] [CrossRef]
- Rao, Z.; Handford, P.; Mayhew, M.; Knott, V.; Brownlee, G.G.; StuartZ, D. The structure of a Ca2+-binding epidermal growth factor-like domain: Its role in protein-protein interactions. Cell 1995, 82, 131–141. [Google Scholar] [CrossRef]
- Burstin, J. Differential expression of two barley XET-related genes during coleoptile growth. J. Exp. Bot. 2000, 51, 847–852. [Google Scholar] [CrossRef][Green Version]
- Lockhart, J.A. An analysis of irreversible plant cell elongation. J. Theor. Biol. 1965, 8, 264–275. [Google Scholar] [CrossRef]
- Kohorn, B.D.; Kobayashi, M.; Johansen, S.; Riese, J.; Huang, L.F.; Koch, K.; Fu, S.; Dotson, A.; Byers, N. An arabidopsis cell wall-associated kinase required for invertase activity and cell growth. Plant J. 2006, 46, 307–316. [Google Scholar] [CrossRef]
- Kakei, Y.; Mochida, K.; Sakurai, T.; Yoshida, T.; Shinozaki, K.; Shimada, Y. Transcriptome analysis of hormone-induced gene expression in Brachypodium distachyon. Sci. Rep. 2015, 5, 14476. [Google Scholar] [CrossRef]
- Cuvillier, O.; Pirianov, G.; Kleuser, B.; Vanek, P.G.; Coso, O.A.; Gutkind, J.S.; Spiegel, S. Suppression of ceramide-mediated programmed cell death by sphingosine-1-phosphate. Nat. Cell Biol. 1996, 381, 800–803. [Google Scholar] [CrossRef]
- Ashkenazi, A. Death receptors: Signaling and modulation. Science 1998, 281, 1305–1308. [Google Scholar] [CrossRef]
- Cross, T.G.; Scheel-Toellner, D.; Henriquez, N.V.; Deacon, E.; Salmon, M.; Lord, J.M. Serine/threonine protein kinases and apoptosis. Exp. Cell Res. 2000, 256, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Kohorn, B.D.; Kohorn, S.L.; Todorova, T.; Baptiste, G.; Stansky, K.; McCullough, M. A Dominant allele of arabidopsis pectin-binding wall-associated kinase induces a stress response suppressed by MPK6 but not MPK3 mutations. Mol. Plant 2012, 5, 841–851. [Google Scholar] [CrossRef] [PubMed]
- Malukani, K.K.; Ranjan, A.; Hota, S.J.; Patel, H.K.; Sonti, R.V. Dual activities of receptor-like kinase sWAKL21.2 induce immune responses. Plant Physiol. 2020, 183, 1345–1363. [Google Scholar] [CrossRef] [PubMed]
- Hurni, S.; Scheuermann, D.; Krattinger, S.G.; Kessel, B.; Wicker, T.; Herren, G.; Fitze, M.N.; Breen, J.; Presterl, T.; Ouzunova, M.; et al. The maize disease resistance gene Htn1 against northern corn leaf blight encodes a wall-associated receptor-like kinase. Proc. Natl. Acad. Sci. USA 2015, 112, 8780–8785. [Google Scholar] [CrossRef] [PubMed]
- Yang, P.; Praz, C.; Li, B.; Singla, J.; Robert, C.A.M.; Kessel, B.; Scheuermann, D.; Lüthi, L.; Ouzunova, M.; Erb, M.; et al. Fungal resistance mediated by maize wall-associated kinase Zm WAK–RLK 1 correlates with reduced benzoxazinoid content. New Phytol. 2018, 221, 976–987. [Google Scholar] [CrossRef]
- Lv, D.; Li, X.; Zhang, M.; Gu, A.; Zhen, S.; Wang, C.; Li, X.; Yan, Y. Large-scale phosphoproteome analysis in seedling leaves of Brachypodium distachyon L. BMC Genom. 2014, 15, 1–18. [Google Scholar] [CrossRef]
- Ridley, B.L.; A O’Neill, M.; Mohnen, D. Pectins: structure, biosynthesis, and oligogalacturonide-related signaling. Phytochemistry 2001, 57, 929–967. [Google Scholar] [CrossRef]
- Willats, W.G.T.; McCartney, L.; Mackie, W.; Knox, J.P. Pectin: cell biology and prospects for functional analysis. Plant Mol. Biol. 2001, 47, 9–27. [Google Scholar] [CrossRef]
- Wolf, S.; Greiner, S. Growth control by cell wall pectins. Protoplasma 2012, 249, 169–175. [Google Scholar] [CrossRef]
- Kohorn, B.D. Cell wall-associated kinases and pectin perception. J. Exp. Bot. 2016, 67, 489–494. [Google Scholar] [CrossRef]
- Cayrol, B.; Delteil, A.; Gobbato, E.; Kroj, T.; Morel, J.-B. Three wall-associated kinases required for rice basal immunity form protein complexes in the plasma membrane. Plant Signal. Behav. 2016, 11, e1149676. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, X.; Bacic, A.; Johnson, K.L.; Humphries, J. The Role of Brachypodium distachyon Wall-Associated Kinases (WAKs) in Cell Expansion and Stress Responses. Cells 2020, 9, 2478. https://doi.org/10.3390/cells9112478
Wu X, Bacic A, Johnson KL, Humphries J. The Role of Brachypodium distachyon Wall-Associated Kinases (WAKs) in Cell Expansion and Stress Responses. Cells. 2020; 9(11):2478. https://doi.org/10.3390/cells9112478
Chicago/Turabian StyleWu, Xingwen, Antony Bacic, Kim L. Johnson, and John Humphries. 2020. "The Role of Brachypodium distachyon Wall-Associated Kinases (WAKs) in Cell Expansion and Stress Responses" Cells 9, no. 11: 2478. https://doi.org/10.3390/cells9112478
APA StyleWu, X., Bacic, A., Johnson, K. L., & Humphries, J. (2020). The Role of Brachypodium distachyon Wall-Associated Kinases (WAKs) in Cell Expansion and Stress Responses. Cells, 9(11), 2478. https://doi.org/10.3390/cells9112478






