Influenza A Hemagglutinin Passage Bias Sites and Host Specificity Mutations
Abstract
1. Introduction
2. Materials and Methods
2.1. Sequence and Passage Data Sources and Analysis
2.2. Same Strain Different Passage (SSDP) Analysis
2.3. Pair Analysis
3. Results
3.1. Structural Overview of Passage Bias Sites in HA Across Different Influenza A Subtypes
3.2. Epistasis of Site Pairs Differs by Passage Types
3.3. Comparing HA Passage Bias with Host Specificity Mutations
3.3.1. Passage Adaptations cannot Easily Predict Directionality of Host Specific Adaptation Mutations of Specific Strains in Ferret-Adaptation Studies
3.3.2. Passage Adaptation Sites Overlap with Previously Known H5 Sites That Affect Host Specificity
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Neumann, G.; Kawaoka, Y. Transmission of Influenza A Viruses. Virology 2015, 479, 234–246. [Google Scholar] [CrossRef] [PubMed]
- Reperant, L.A.; Kuiken, T.; Osterhaus, A.D. Adaptive pathways of zoonotic influenza viruses: From exposure to establishment in humans. Vaccine 2012, 30, 4419–4434. [Google Scholar] [CrossRef] [PubMed]
- Richard, M.; Fouchier, R.A. Influenza A virus transmission via respiratory aerosols or droplets as it relates to pandemic potential. FEMS Microbiol. Rev. 2015, 40, 68–85. [Google Scholar] [CrossRef] [PubMed]
- Peiris, M.; Yen, H.-L. Animal and human influenzas. Rev. Sci. Tech. Int. Off. Epizoot. 2014, 33, 539–553. [Google Scholar] [CrossRef]
- Yoon, S.-W.; Webby, R.J.; Webster, R.G. Evolution and Ecology of Influenza A Viruses. Curr. Top. Microbiol. Immunol. 2014, 385, 359–375. [Google Scholar]
- Stevens, J.; Blixt, O.; Tumpey, T.M.; Taubenberger, J.K.; Paulson, J.C.; Wilson, I.A. Structure and Receptor Specificity of the Hemagglutinin from an H5N1 Influenza Virus. Science 2006, 312, 404–410. [Google Scholar] [CrossRef] [PubMed]
- Imai, M.; Watanabe, T.; Hatta, M.; Das, S.C.; Ozawa, M.; Shinya, K.; Zhong, G.; Hanson, A.; Katsura, H.; Watanabe, S.; et al. Experimental adaptation of an influenza H5 haemagglutinin (HA) confers respiratory droplet transmission to a reassortant H5 HA/H1N1 virus in ferrets. Nature 2012, 486, 420–428. [Google Scholar] [CrossRef]
- Herfst, S.; Schrauwen, E.J.A.; Linster, M.; Chutinimitkul, S.; De Wit, E.; Munster, V.J.; Sorrell, E.M.; Bestebroer, T.M.; Burke, D.F.; Smith, D.J.; et al. Airborne Transmission of Influenza A/H5N1 Virus Between Ferrets. Science 2012, 336, 1534–1541. [Google Scholar] [CrossRef]
- Duprex, W.P.; Fouchier, R.A.M.; Imperiale, M.J.; Lipsitch, M.; Relman, D.A. Gain-of-function experiments: time for a real debate. Nat. Rev. Microbiol. 2015, 13, 58–64. [Google Scholar] [CrossRef]
- Tharakaraman, K.; Raman, R.; Viswanathan, K.; Stebbins, N.W.; Jayaraman, A.; Krishnan, A.; Sasisekharan, V.; Sasisekharan, R. Structural Determinants for Naturally Evolving H5N1 Hemagglutinin to Switch its Receptor Specificity. Cell 2013, 153, 1475–1485. [Google Scholar] [CrossRef]
- Bush, R.M.; Smith, C.B.; Cox, N.J.; Fitch, W.M. Effects of passage history and sampling bias on phylogenetic reconstruction of human influenza A evolution. Proc. Natl. Acad. Sci. USA 2000, 97, 6974–6980. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Deng, Q.; Ng, S.H.; Lee, R.T.C.; Maurer-Stroh, S.; Zhai, W. Dynamic Convergent Evolution Drives the Passage Adaptation across 48 Years’ History of H3N2 Influenza Evolution. Mol. Boil. Evol. 2016, 33, 3133–3143. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Russell, C.A.; Kasson, P.M.; Donis, R.O.; Riley, S.; Dunbar, J.; Rambaut, A.; Asher, J.; Burke, S.; Davis, C.T.; Garten, R.J.; et al. Improving pandemic influenza risk assessment. eLife 2014, 3, e03883. [Google Scholar] [CrossRef] [PubMed]
- Trock, S.C.; Burke, S.A.; Cox, N.J. Development of Framework for Assessing Influenza Virus Pandemic Risk. Emerg. Infect. Dis. 2015, 21, 1372–1378. [Google Scholar] [CrossRef] [PubMed]
- Russell, C.A.; Fonville, J.M.; Brown, A.E.X.; Burke, D.F.; Smith, D.L.; James, S.L.; Herfst, S.; Van Boheemen, S.; Linster, M.; Schrauwen, E.J.; et al. The potential for respiratory droplet transmissible A/H5N1 influenza virus to evolve in a mammalian host. Science 2012, 336, 1541–1547. [Google Scholar] [CrossRef]
- Lipsitch, M.; Barclay, W.; Raman, R.; Russell, C.J.; Belser, J.A.; Cobey, S.; Kasson, P.M.; Lloyd-Smith, J.O.; Maurer-Stroh, S.; Riley, S.; et al. Viral factors in influenza pandemic risk assessment. eLife 2016, 5, e18491. [Google Scholar] [CrossRef] [PubMed]
- Maurer-Stroh, S.; Li, Y.; Bastien, N.; Gunalan, V.; Lee, R.T.C.; Eisenhaber, F.; Booth, T.F. Potential Human Adaptation Mutation of Influenza A(H5N1) Virus, Canada. Emerg. Infect. Dis. 2014, 20, 1580–1582. [Google Scholar] [CrossRef]
- DuPai, C.D.; McWhite, C.D.; Smith, C.B.; Garten, R.; Maurer-Stroh, S.; Wilke, C.O. Influenza passaging annotations: what they tell us and why we should listen. Virus Evol. 2019, 5, vez016. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT: iterative refinement and additional methods. Methods Mol. Biol. Clifton NJ 2014, 1079, 131–146. [Google Scholar]
- Therneau, T. A Package for Survival Analysis in S. Version 2.38. 2015. [Google Scholar]
- Zaraket, H.; Bridges, O.A.; Duan, S.; Baranovich, T.; Yoon, S.-W.; Reed, M.L.; Salomon, R.; Webby, R.J.; Webster, R.G.; Russell, C.J. Increased Acid Stability of the Hemagglutinin Protein Enhances H5N1 Influenza Virus Growth in the Upper Respiratory Tract but Is Insufficient for Transmission in Ferrets. J. Virol. 2013, 87, 9911–9922. [Google Scholar] [CrossRef]
- Shelton, H.; Roberts, K.L.; Molesti, E.; Temperton, N.; Barclay, W.S. Mutations in haemagglutinin that affect receptor binding and pH stability increase replication of a PR8 influenza virus with H5 HA in the upper respiratory tract of ferrets and may contribute to transmissibility. J. Gen. Virol. 2013, 94, 1220–1229. [Google Scholar] [CrossRef]
- Weltman, J.K.; Skowron, G.; Loriot, G.B. Influenza A H5N1 hemagglutinin cleavable signal sequence substitutions. Biochem. Biophys. Res. Commun. 2007, 352, 177–180. [Google Scholar] [CrossRef] [PubMed]
- H5N1 Genetic Changes Inventory. Available online: https://www.cdc.gov/flu/avianflu/h5n1/inventory.htm (accessed on 28 May 2019).
- Guarnaccia, T.; Carolan, L.A.; Maurer-Stroh, S.; Lee, R.T.C.; Job, E.; Reading, P.C.; Petrie, S.; McCaw, J.M.; McVernon, J.; Hurt, A.C.; et al. Antigenic Drift of the Pandemic 2009 A(H1N1) Influenza Virus in a Ferret Model. PLoS Pathog. 2013, 9, e1003354. [Google Scholar] [CrossRef] [PubMed]
- Burnet, F.; Bull, D.H. Changes in influenza virus associated with adaptation to passage in chick embryos. Aust. J. Exp. Boil. Med. Sci. 1943, 21, 55–69. [Google Scholar] [CrossRef]
- Gatherer, D. Passage in egg culture is a major cause of apparent positive selection in influenza B hemagglutinin. J. Med. Virol. 2010, 82, 123–127. [Google Scholar] [CrossRef] [PubMed]
- Subbarao, K. The Critical Interspecies Transmission Barrier at the Animal–Human Interface. Trop. Med. Infect. Dis. 2019, 4, 72. [Google Scholar] [CrossRef]
- Skowronski, D.M.; Janjua, N.Z.; De Serres, G.; Sabaiduc, S.; Eshaghi, A.; Dickinson, J.A.; Fonseca, K.; Winter, A.-L.; Gubbay, J.B.; Krajden, M.; et al. Low 2012–13 Influenza Vaccine Effectiveness Associated with Mutation in the Egg-Adapted H3N2 Vaccine Strain Not Antigenic Drift in Circulating Viruses. PLoS ONE 2014, 9, e92153. [Google Scholar] [CrossRef]
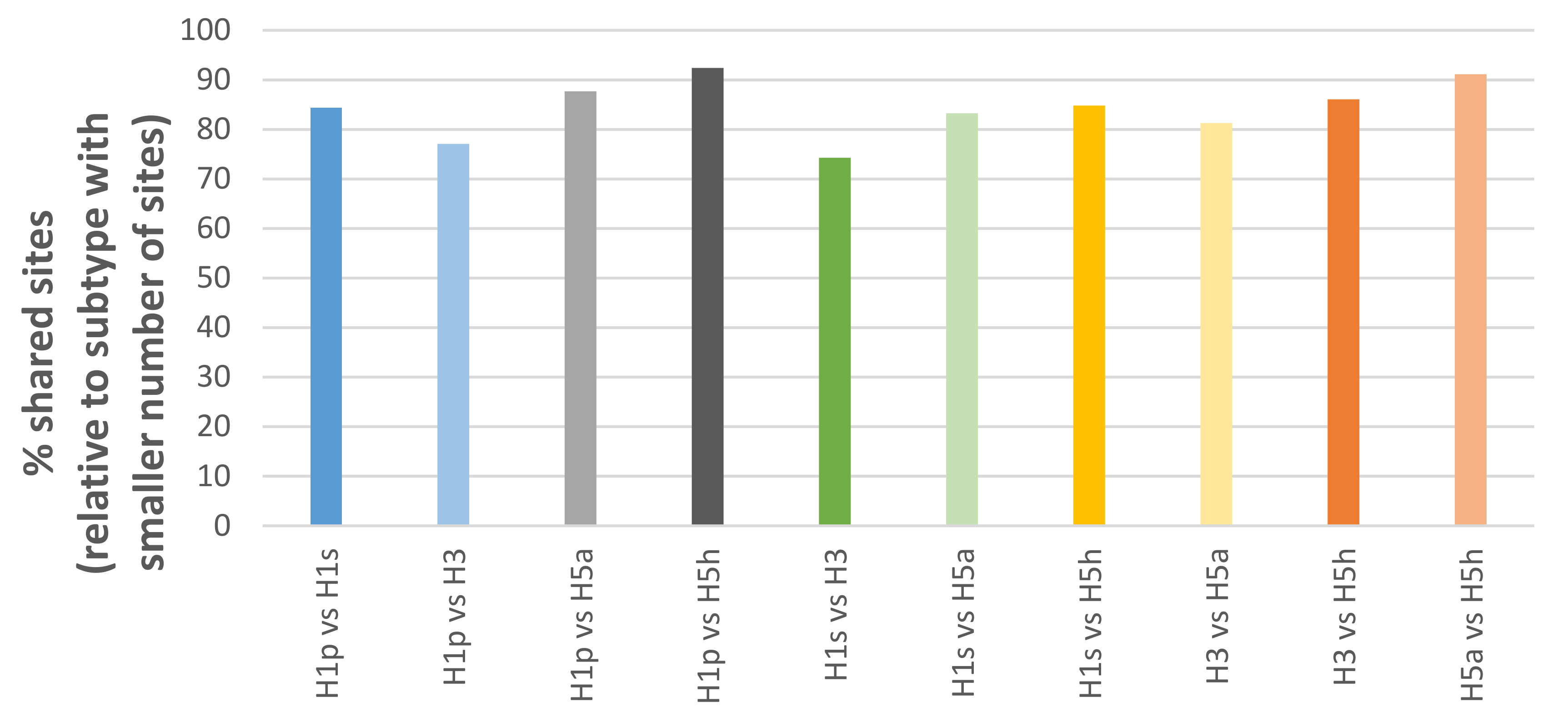
| Subtype | EGG | MDCK | SIAT | ORI | SUM |
|---|---|---|---|---|---|
| H1 pandemic (≥2009) | 1312 | 18,059 | 2103 | 13,004 | 34,478 |
| H1 seasonal (<2009) | 606 | 2610 | 145 | 342 | 3703 |
| H3 seasonal | 1405 | 9663 | 10,423 | 17,641 | 39,132 |
| H5 (human) | 234 | 36 | 0 | 14 | 284 |
| H5 (avian) | 1439 | 16 | 0 | 186 | 1641 |
| SUM | 4996 | 30,384 | 12,671 | 31,187 | 79,238 |
| Structural Regions | Sites (H3 Numbering, Starting After Signal Peptide) | Sites (H1pdm Absolute Numbering, Starting at Leading Methionine) | # Occurrence in 18 Possible Subtype + Cell Combinations (4 H1p, 4 H1s, 4 H3, 3 H5 Avian and 3 H5 Human) | Geometric Mean of Odds Ratios from all Subtype/Cell Combinations |
|---|---|---|---|---|
| Globular Head (RBS) | 155 186 187 189 193 194 225 226 227 | 169 200 201 203 207 208 239 240 241 | 15 10 14 16 17 13 12 10 13 | 8.31 6.03 4.11 23.25 11.92 7.59 6.65 16.05 10.73 |
| Globular Head (Around RBS) | 131 132 133 137 140 141 144 156 157 158 159 188 196 198 218 221 222 230 | 144 145 146 151 154 155 158 170 171 172 173 202 210 212 232 235 236 244 | 16 17 15 13 16 11 18 12 7 15 17 17 14 14 13 11 11 12 | 11.42 9.25 13.47 10.68 11.92 3.91 81.24 11.11 3.36 12.97 31.17 20.84 8.40 19.90 3.62 3.19 9.97 3.7 |
| Globular Head (Others) | 77 80 81 82 101 125 163 167 260 262 263 | 85 88 89 91 111 138 177 181 274 277 278 | 12 17 14 15 18 18 14 10 13 16 10 | 5.9 35.95 6.27 18.62 18.65 24.19 12.41 3.97 5.02 9.43 4.45 |
| pH Structural Change site | 45 46 53 54a * 273 275 279 285 | 52 53 60 62 288 290 294 300 | 13 14 16 11 14 17 12 14 | 7.08 9.02 15.26 7.99 12.36 14.31 2.87 11.54 |
| N-terminal (Signal Peptide) | −9 −8 −7 −6 −5 −4 −3 −2 | 9 10 11 12 13 14 15 16 | 15 10 16 16 13 15 13 15 | 12.59 4.92 12.53 10.95 10.44 12.89 12.33 13.36 |
| Position 1 | Position 2 | Log Odds Ratio | P-Value |
|---|---|---|---|
| H1N1 ORI vs. EGG in H1 Absolute Numbering (H1pdm09 Absolute Numbering in Bracket) | |||
| 13 (13) | 101 (101) | 1.69 | 1.11 × 10−10 |
| H3N2 MDCK vs. EGG in H3 Absolute Numbering (H1pdm09 Absolute Numbering in Bracket) | |||
| 19 (gap) | 176 (174) | 1.20 | 2.22 × 10−15 |
| 19 (gap) | 210 (208) | −1.98 | 4.17 × 10−8 |
| 154 (152) | 158 (156) | 1.31 | 5.70 × 10−6 |
| 160 (158) | 176 (174) | 1.99 | 0 |
| 160 (158) | 210 (208) | −3.26 | 4.81 × 10−6 |
| 175 (173) | 176 (174) | 0.90 | 8.23 × 10−11 |
| H3N2 ORI vs. EGG in H3 absolute numbering (H1pdm09 absolute numbering in bracket) | |||
| 19 (gap) | 176 (174) | 3.00 | 0 |
| 19 (gap) | 210 (208) | −1.87 | 5.75 × 10−8 |
| 19 (gap) | 241 (239) | 1.57 | 2.82 × 10−7 |
| 19 (gap) | 327 (326) | 1.55 | 5.88 × 10−7 |
| 19 (gap) | 505 (504) | 1.78 | 5.69 × 10−7 |
| 130 (127) | 176 (174) | 5.85 | 9.10 × 10−6 |
| 144 (141) | 176 (174) | −0.69 | 5.79 × 10−7 |
| 154 (152) | 158 (156) | 4.15 | 5.41 × 10−8 |
| 154 (152) | 160 (158) | −1.01 | 1.32 × 10−7 |
| 154 (152) | 176 (174) | −0.95 | 4.60 × 10−8 |
| 154 (152) | 327 (326) | −0.99 | 6.46 × 10−8 |
| 158 (156) | 176 (174) | −0.76 | 2.02 × 10−9 |
| 158 (156) | 327 (326) | −0.68 | 4.27 × 10−7 |
| 160 (158) | 176 (174) | 4.18 | 0 |
| 160 (158) | 210 (208) | −2.85 | 1.14 × 10−6 |
| 160 (158) | 241 (239) | 3.11 | 1.38 × 10−6 |
| 175 (173) | 176 (174) | 2.27 | 0 |
| 175 (173) | 210 (208) | −1.23 | 4.46 × 10−6 |
| 175 (173) | 327 (326) | 1.12 | 7.11 × 10−6 |
| 175 (173) | 505 (504) | 1.28 | 2.13 × 10−6 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, R.T.C.; Chang, H.-H.; Russell, C.A.; Lipsitch, M.; Maurer-Stroh, S. Influenza A Hemagglutinin Passage Bias Sites and Host Specificity Mutations. Cells 2019, 8, 958. https://doi.org/10.3390/cells8090958
Lee RTC, Chang H-H, Russell CA, Lipsitch M, Maurer-Stroh S. Influenza A Hemagglutinin Passage Bias Sites and Host Specificity Mutations. Cells. 2019; 8(9):958. https://doi.org/10.3390/cells8090958
Chicago/Turabian StyleLee, Raphael T. C., Hsiao-Han Chang, Colin A. Russell, Marc Lipsitch, and Sebastian Maurer-Stroh. 2019. "Influenza A Hemagglutinin Passage Bias Sites and Host Specificity Mutations" Cells 8, no. 9: 958. https://doi.org/10.3390/cells8090958
APA StyleLee, R. T. C., Chang, H.-H., Russell, C. A., Lipsitch, M., & Maurer-Stroh, S. (2019). Influenza A Hemagglutinin Passage Bias Sites and Host Specificity Mutations. Cells, 8(9), 958. https://doi.org/10.3390/cells8090958





