1. Introduction
HIV encodes pathogenic proteins, such as gp120, Nef, and Tat, that modulate cellular architecture and behavior. Such modulations are implicated in HIV-induced pathological processes, including immune activation that persist during combination antiretroviral therapy (cART) and contribute to serious non-AIDS events. Although cART has dramatically reduced HIV/AIDS-related pathologies and mortality [
1], use of illicit substances (mostly psychostimulants) is a major barrier to combating the HIV pandemic [
2,
3,
4,
5]. Psychostimulants such as cocaine have been linked to exacerbated HIV disease progression and HIV-associated disorders [
6,
7,
8,
9,
10,
11,
12,
13]. In addition to its ability to promote risky behavior [
14], cocaine impairs antiviral mechanisms [
8,
15], thus increasing the risk of HIV acquisition. The combination of behavioral alteration and psychostimulant-mediated impairment of antiviral mechanisms continues to be a major obstacle in combating the global HIV/AIDS pandemic. The risk of exacerbated HIV disease progression and/or HIV-associated disorders among those who use psychostimulants and who are also infected with HIV is present in those adherent to cART [
16,
17,
18].
Aside from the brain, peripheral tissues, including lymphocytes, monocytes, and the male urogenital organs, are responsive to psychostimulants due to the presence of dopamine transporters (DAT) and dopamine receptors (DR) [
19,
20,
21]. In particular, DRD1 and DRD2 are expressed in male genital tissues such as the testis [
22] and cocaine induces ultrastructural changes in the testis [
23] and negatively affects testicular physiology as well as spermatogenic processes [
23,
24]. Similar to its function in the central nervous system, the function of dopamine in myeloid cells is mediated primarily by DRs, which are expressed in human monocytes and macrophages [
25,
26,
27]. Myeloid cell DRs are functional and have been implicated in HIV infection and substance use disorders [
28]. Although peripheral cells have been linked to increased viral replication in the presence of psychostimulants [
29,
30], how HIV and/or psychostimulants alter monocyte function is not completely understood.
Recently, acellular mechanisms regulating host functions have been discovered to occur through extracellular vesicles, in particular, exosomes, which are conveyors of bio-information [
31,
32,
33]. Exosomes have been implicated in the modulation of immune responses [
34,
35] and microbial pathogenesis, including HIV infection [
36,
37,
38,
39,
40,
41,
42,
43,
44,
45]. Other biological processes, such as extracellular matrix (ECM) reorganization, epithelial barrier regulation, inflammatory cell recruitment, microglial migration [
46], and regulation of HIV transcription [
44], have been associated with exosomes [
47]. Given that exosomes are released by various cell types into all body fluids [
35,
36,
48,
49,
50,
51,
52,
53], it is likely that HIV infection and/or psychostimulant-mediated effects on peripheral tissues may be imprinted in exosomes and that such exosomes may reprogram host gene expression and function. In a recent study, we showed that SEs from HIV-uninfected donors who do not use psychostimulants selectively modifies HIV-induced activation of host transcription factors [
44].
In the present study, our goal was to evaluate the effect of HIV infection, psychostimulant use, and co-occurring HIV/psychostimulant use on SE-mediated regulation of monocyte function. We used monocytes as a model because monocytes are present in nearly every tissue, including the brain that has little or no T cell colonization. Moreover, monocytes differentiate into HIV target cells—dendritic cells and macrophages. Finally, monocytes are the first cells recruited to sites of inflammation, are one of the immune cell types present in semen [
54,
55], and are important target cells for mucosal HIV transmission [
56], as well as HIV-associated neurocognitive disorders [
57].
2. Materials and Methods
2.1. Ethics
This study involves the use of existing human specimens (semen) and, therefore, is not human subjects’ research. De-identified semen samples were obtained from participants in the Multicenter AIDS Cohort Study (MACS), a prospective cohort study of the natural history of HIV infection in men who have sex with men which was initiated in 1984 in 4 US sites and obtained semen samples from study participants semiannually from 1984 to 1987. The semen samples were stored at −80 °C until analysis in the present study. The participants included HIV− and HIV+ men who, at the time of collection, reported using or not using illicit substances. Studies were conducted according to University regulations approved by The University of Iowa and Stony Brook University Institutional Review Boards (IRB # 201608703). HIV-1-negative donors had no history of HIV, hepatitis B virus (HBV), or hepatitis C virus (HCV) infections. HIV-1-infected donors were ART-naive.
2.2. Semen Samples
A total of 64 samples from four clinical groups (HIV-uninfected and not illicit substance users, HIV−Drug−; HIV-uninfected and self-reported illicit substance users, HIV-Drug+; HIV-infected and not illicit substance users, HIV+Drug−; and HIV-infected and self-reported use of illicit substances, HIV+Drug+). A participant was classified as an illicit substance user only if they reported using cocaine (taken by any route); in other words, if a participant reported using other substances without cocaine, they were excluded (
Table 1). Sixteen participants in each group were analyzed. The samples were received frozen on dry ice from the MACS. The samples were collected between 1984 and 1987, and participants were between 20 and 65 years old.
2.3. Cells
U937 monocytic cells were obtained from the American Type Culture Collection (ATCC) and maintained in complete Roswell Park Memorial Institute (RPMI) media (Corning, Thermofisher, Grand Island, NY, USA). HIV-1 LAV-infected HeLa CD4+ cells from which HIV secretome was collected were obtained from the National Institutes of Health (NIH) Aids Reagent Program and maintained in complete Dulbecco’s Modified Eagle Medium (DMEM) media. RPMI and DMEM media were supplemented with 10% fetal bovine serum (FBS) (Atlanta Biologicals, Flowery Branch, GA, USA) that was exosome-depleted by ultracentrifugation (100,000× g, 2 h, 4 °C), 1% Penicillin-streptomycin (Thermofisher, Grand Island, NY, USA), 1 µg/mL Amphotericin B (Thermofisher, Grand Island, NY, USA), 2 mM sodium pyruvate (Corning, Corning, NY, USA), 1% of glutamate (Thermofisher, Grand Island, NY, USA), and 10 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) buffer (Fisher Biotech, Fair Lawn, NJ, USA) at pH 8. NucBlue™ Live ReadyProbes™ reagent was purchased from EasyProbes (Thermofisher, Grand Island, NY, USA). Cell Viability Imaging Kit (Blue/Green) was obtained from Genecopoeia (Rockville, MD, USA), and Type I collagen was purchased from Corning (Corning, NY, USA).
2.4. Isolation of Exosomes
64 semen samples from four clinical groups (n = 16/group) were liquefied at room temperature for 30 min and subsequently centrifuged at 10,000×
g for 30 min to remove cellular debris and large vesicles. Clarified seminal plasmas were transferred to new tubes. For Nano Tracking Analysis (NTA) experiments, six pools of samples in each group, each pool from 2 participants (100 µL/sample), were used. Samples were pooled to obtain sufficient volume needed for efficient separation and analysis. For the rest of the experiments, 4 pooled samples (n = 16, 50 µL/sample) per clinical group were used. Exosomes were purified by size exclusion chromatography (SEC), where clarified seminal plasma was loaded onto Sephadex G-50 fine beads (GE-Healthcare, Pittsburgh, PA, USA) packed in a 22 cm × 1 cm Econo-column (Bio-Rad, Hercules, CA, USA). Elution was achieved by gravity using Phosphate Buffered Saline (PBS, Corning, NY, USA). Fractions of 200 µL were collected, and elution profiles were determined by absorbance measurements at 280 nm and 600 nm. The first peak which corresponds to semen exosomes (SE) was collected, and the protein content was measured by the Bradford Assay (Bio-Rad, Hercules, CA, USA). Of note, HIV could not be efficiently separated from semen exosomes using the Optiprep (Iodixanol)-based density gradient centrifugation method. While a good gradient prior to centrifugation was obtained, a satisfactory purification was not achieved due to the fact that the gold-standard exosomal marker AChE, as well as the exosomal markers CD9, CD63, and HSP70, along with the viral protein reverse transcriptase (RT) were found across the gradients. This is not surprising since HIV and exosomes overlap in size, density, and charge, and HIV is known to incorporate exosomal markers such as CD9, CD81 [
58], and CD63 [
59], while exosomes in turn also contain viral proteins [
60] and RNA [
61]. Immunocapture purification could not be used either because this mechanism depends on the use of antibodies against either host or viral proteins which are present in exosomes and HIV. Moreover, the “release” mechanism of exosomes trapped on the antibody-bead complex was inefficient. Thus, the inclusion of exosomal proteins in HIV and HIV proteins in exosomes hindered separation of these vesicles but also highlighted the need to assess the vesicles in their near-native state to understand their effect on host cells.
2.5. Nanoparticle Tracking Analysis (NTA)
Exosome size and concentration were measured by NTA using ZetaView PMX 110 (Particle Metrix, Mebane, NC, USA) and the corresponding software ZetaView v8.04.02. Samples were diluted appropriately in ultrapure water and measured under the same settings (temperature 25 °C, sensitivity 92, shutter speed 70, and frame rate 30 fps). Data acquisition for size and concentration was performed in triplicate measurements, and each replicate corresponded to 11 positions with two cycles of reading at each position. The system was aligned and calibrated with 102-nm polystyrene standard beads. After automated analysis of the 11 positions and removal of any outlier position, the median number (X50) was used to report the particle size. The measured concentration was normalized to the volume of plasma and reported in particles/mL of seminal plasma. For zeta potential, measurements were performed in ultrapure water (pH 5.8) and data were acquired in quintuplicate. Each replicate corresponded to two cycles of reading.
2.6. Transmission Electron Microscopy (TEM)
Microscopic analysis of exosome samples was performed as previously described [
36,
38]: 200 µL of purified SE were buffer exchanged with Tris buffer (pH = 7.5, 1 M) and concentrated through a 0.5-mL centrifugal filter (10,000 NMWL) into 50 µL; 10 µL of concentrated SE was applied on to carbon-coated copper grids (Pellco Easiglow, 0.2 mpar, 30 mA, 40 s, negative) and allowed to sit for 30 s. Excess samples were removed with filter paper. The grids were washed with distilled deionized water (ddH
2O) twice, stained with 0.7% Uranyl Formate solution for 20 s, and then allowed to air dry. Images were viewed and collected using a FEI Tecnai12 BioTwinG 2 electron microscope. The samples were captured with an AMT XR-60 CCD Digital Camera system. The size of particles from TEM images were quantified by ImageJ.
Microscopic analysis of exosome samples was performed as previously described [
36,
38]: 200 µL of purified SE were buffer exchanged with Tris buffer (pH = 7.5, 1 M) and concentrated through a 0.5-mL centrifugal filter (10,000 NMWL) into 50 µL; 10 µL of concentrated SE was applied on to carbon-coated copper grids (Pellco Easiglow, 0.2 mpar, 30 mA, 40 s, negative) and allowed to sit for 30 s. Excess samples were removed with filter paper. The grids were washed with distilled deionized water (ddH
2O) twice, stained with 0.7% Uranyl Formate solution for 20 s, and then allowed to air dry. Images were viewed and collected using a FEI Tecnai12 BioTwinG 2 electron microscope. The samples were captured with an AMT XR-60 CCD Digital Camera system. The size of particles from TEM images were quantified by ImageJ.
2.7. Reverse Transcriptase (RT) Assay
HIV RT activity was determined with an EnzCheck Reverse Transcriptase Assay kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. Briefly, 30 µg (~6 × 109 particles) of purified SE (6 pools of 2 donors each for HIV+Drug− and HIV+Drug+ groups) were lysed with 6 µL Triton X-100 in a total volume of 50 µL per well, to which 20 µL of poly(A)-oligo(dT) in a polymerization buffer were added. Assay was performed in a 96-well black plate in triplicates. An equivalent volume of PBS was used as the negative control. RT standard curve was prepared with serial dilution (0, 0.625, 1.25, 2.5, and 5 µg/mL) of Murine Leukemia Virus (MLV) recombinant RT.
2.8. ELISA Assays
HIV p24 ELISA (Xpressbio, Frederick, MD, USA) and cocaine ELISA (Abnova, Taipei, Taiwan, China) were conducted by following the manufacturers’ protocols. Briefly, for HIV p24 ELISA, a total of 30 µg purified SE (~6 × 109 particles) from HIV+ groups were tested in 6 pools of 2 donors each. An equivalent volume of PBS was used as the negative control. The same procedure was adopted for cocaine metabolite ELISA, with Drug+ groups being tested (6 pools of 2 donors each per group) in triplicate. The detection limit of the ELISA kit for HIV p24 and cocaine were 1.7 pg/mL and 1 ng/mL, respectively.
2.9. RNA Purification
Collagen coating of tissue culture plates was described previously [
62]. Briefly, 6-well tissue culture plates were pre-coated with 50 µg/mL of collagen for 2 h at 37 °C, after which 1 mL of 2 mg/mL bovine serum albumin (BSA, Research Products International, Mount Prospect, IL, USA) was added to block nonspecific sites. Two million U937 cells treated with vehicle (PBS) or 100 µg/mL of SE from each of the four clinical groups were plated and incubated for 18 h at 37 °C and 5% CO
2. Each treatment included three replicates. Subsequently, total RNA was extracted using the miRNeasy Mini Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. An on-column DNAse digestion step (RNase-Free DNase set, Qiagen) was added after the first buffer wash step. The yield, quality, and size distribution of RNA isolated from the cells were determined using the Bioanalyzer instrument (Agilent, Santa Clara, CA, USA). Six hundred ng of the RNA from each treatment group was applied to an RNA Nano Chip, and the RNA profiles were detected and analyzed on the Agilent 2100 Bioanalyzer with 2100 Bioanalyzer expert software (v B.02.08.S1648 (SR 1)). The electropherogram traces and “gel-like” images were exported from the instrument’s software and presented in
Supplementary Figure S1. Isolated RNA was used for microarray analysis or for cDNA synthesis and subsequent real-time quantitative PCR (RT-qPCR) analysis.
2.10. Microarray Analysis, Data Mining, and Data Visualization
150 ng of total RNA was prepared for microarray analysis using the GeneChip™ WT PLUS Reagent Kit (Applied Biosystems, Foster City, CA) according to manufacturer’s protocol. The samples were hybridized (16 h) to Clariom™ S Human Arrays (Applied Biosystems, Foster City, CA) in a GeneChip™ Hybridization Oven 645 (Applied Biosystems™). The arrays were washed and stained using the GeneChip™ Hybridization, Wash and Stain Kit (Applied Biosystems, Foster City, CA) in a GeneChip™ Fluidics Station 450 according to manufacturer’s protocol. The arrays were scanned in a GeneChip™ Scanner 3000 7G (Applied Biosystems, Foster City, CA). Quality control and initial analysis, including scatterplots of differentially expressed genes (DEGs), Venn analysis, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis, were performed using Transcriptome Analysis Console (TAC) v 4.0.0.25 (Applied Biosystems, Foster City, CA). Clustered heatmaps were plotted using heatmapper [
63] (
www.heatmapper.ca), with the average linkage method and Euclidean distance measurement method. The lists of SE, SE-Drug, and SE-HIV DEGs that were obtained from a TAC analysis were subjected to data mining in a Web-based Gene Set Analysis Toolkit (WebGestalt [
64],
www.webgestalt.org), from which biological process and molecular function and cellular component gene ontology (GO) terms were obtained.
2.11. Primer Design and Real-Time Quantitative PCR (RT-qPCR) Data Validation
2.12. Collagen Adhesion Assay
Flat-bottom 96-well plate were pre-coated with 50 µL of 50 µg/mL of Type I collagen (Corning, Corning, NY, USA) for 2 h at 37 °C; 40 µL of 2 mg/mL Bovine Serum Albumin (BSA) was used to block the nonspecific sites. Incoming U937 cells (10,000 cells/well) treated with 100 µg/mL of SE or with equivalent volume of vehicle (PBS control) were added to the pre-coated wells and allowed to adhere for 18 h at 37 °C. Non-adhered cells were gently washed off with PBS three times. The adhered cells were labeled with NucBlue™ for 20 min at room temperature, and the wells were imaged (4× objective) in their entirety in the DAPI channel (360 nm/460 nm excitation/emission) using a Lionheart FX Automated Microscope (BioTek, Winooski, VT, USA). The captured images were then stitched, and the cell numbers were calculated using Gen5 ImagePrime. Values were represented as the number of total adhered cells in each well. Each treatment included four repeated wells.
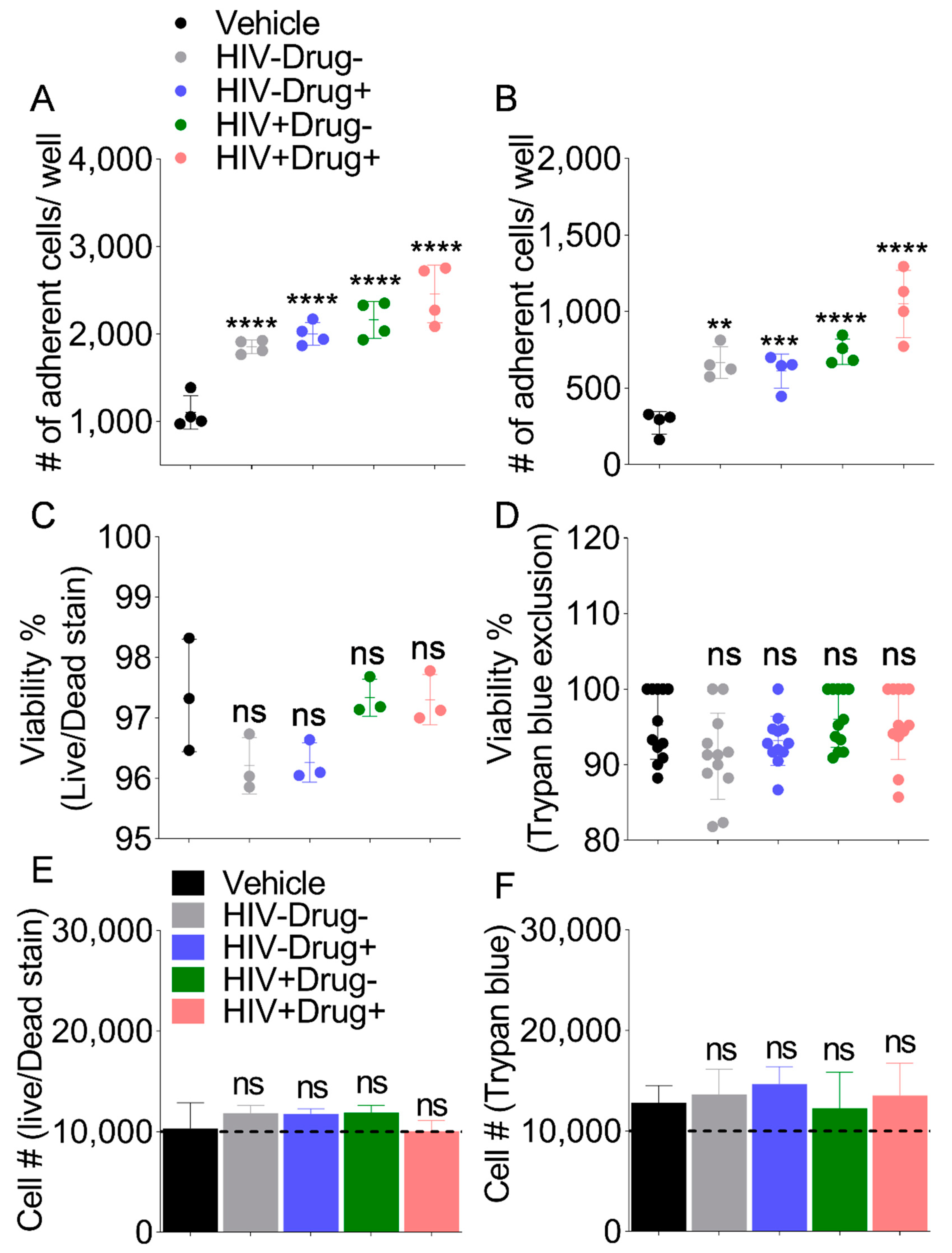
2.13. Evaluation of Cell Viability and Proliferation
A total of 10,000 U937 cells per well were seeded in a collagen-coated 96-well plate with 100 µg/mL of SE or equivalent volume of PBS for 18 h at 37 °C. All cells were collected after treatment and tested for viability by the Trypan Blue (Life Technologies, Carlsbad, CA, USA) exclusion and Live/Dead Cell Stain (Cell Viability Imaging Kit, GeneCopoeia, Rockville, MD, USA) methods. Cell proliferation was determined by counting the total number of live cells. The experiments were repeated 3 times, and each experiment included three replicates.
2.14. Immunofluorescence-Based Analysis of Cytoskeletal Changes and Focal Adhesion
U937 cells were plated (10,000 cells/well) on a 96-well glass bottom dish (Cellvis, Mountain View, CA, USA) coated with Type I collagen and treated with 100 µg/mL of respective SE. The plate was centrifuged at 200×
g for 8 min to facilitate cellular adherence to the bottom of the well and incubated at 37 °C for 18 h. Following incubation, cells were washed with PBS and fixed with 4% paraformaldehyde (PFA) in PBS for 15 min. Cells were then permeabilized by incubation in 0.1% TritonX-100 for 10 min. AlexaFluor 594 Phalloidin (Thermofisher, Grand Island, NY, USA) and Alexa Fluor 488 Vinculin (Thermofisher, Grand Island, NY, USA) were applied in a 1:40 dilution for 1 h, followed by a 5-min DAPI stain. Images were acquired using a Lionheart FX Automated Microscope (Biotek, Winooski, VT, USA). Representative 10× and 60× images were acquired manually for five fields of view per well. Image procession was performed using Gen5 ImagePrime. Quantification of cellular size, area, and circularity was performed by Gen5 ImagePrime via masking of phalloidin. A circularity metric was created by inputting the equation
, where
is the circularity,
is the area of the cell, and
is the perimeter of the cell (
https://imagej.nih.gov/ij/plugins/circularity.html).
2.15. Colocalization Analysis
60× fluorescent images of U937 cells treated with vehicle or SE (100 µg/mL) and stained with Alexa Fluor 594 Phalloidin and Alexa Fluor 488 Vinculin captured on a Lionheart FX Automated Microscope were imported into ImageJ (
http://imagej.nih.gov/) for colocalization analysis. An open source ImageJ plugin “EzColocalization” (
http://sites.imagej.net/EzColocalization/plugins/) was used to quantify colocalization of actin (phalloidin) and vinculin at regions of cell–cell contact and membrane protrusions. Regions of interests (ROIs) were selected via an ROI manager. Using EzColocalization, Pearson correlation coefficient (PCC) and Threshold overlap score (TOS, linear) quantifications were performed for 7 representative fields of view per SE treatment and vehicle. PCC and TOS (linear) values for each ROI were exported into GraphPad Prism for further analysis. One-way ANOVA was performed to determine the significance of SE treatment relative to the vehicle. Colocalization heatmaps were generated using the ImageJ plugin “Colocalization Colormap” (
https://sites.google.com/site/colocalizationcolormap/home).
2.16. Chemotaxis
Migration assays were conducted in a 10-well chemotaxis chamber (Neuroprobe Inc., Gaithersburg, MD, USA). Basal chambers were filled with media containing 0% FBS (serum-free), 30% FBS, or conditioned media from HIV-1 LAV-infected HeLa CD4+ cells (HIV secretome). A polycarbonate polyvinylpyrrolidone-free filter with a pore size of 5 µm was then placed over the lower chambers, and 285 µL of U937 cell suspensions (500,000 cells per well) that were pretreated for 24 h with either vehicle or 100 µg/mL SE from the 4 clinical groups in equal volumes of serum-free media were placed on the filter. The chambers were incubated for an additional 20 h at 37 °C in a 5% CO2 incubator. The apical chamber cells were carefully harvested, membranes were thoroughly rinsed, and basal chamber cells were harvested by piercing the membrane in the basal chamber. Cell suspensions were mixed with Trypan Blue dye, and total cell numbers and viability were quantified via hemocytometer counting.
2.17. Gelatin and Casein Zymography
After the 24 h serum starvation for migration assays, the conditioned media was harvested on ice. Following a 2000× g centrifuge step for 10 min, the media was mixed with a 4× Laemmli sample buffer (Bio-rad, Hercules, CA, USA) with the absence of boiling or 2-mercaptoethanol. Ten percent SDS-PAGE gels (0.75 mm thick) containing 0.1% gelatin in the resolving gel were prepared. Equal volumes of samples were loaded into the lanes, and electrophoresis was performed (Mini-PROTEAN Bio-Rad). Gels were removed from their cassettes, rinsed in distilled water, and incubated with a 1× Zymogram Renaturation Buffer (Bio-Rad) for 30 min with gentle agitation to remove SDS and to renature the proteins. Gels were then transferred to a 1× Zymogram Development Buffer (Bio-rad) for 30 min at room temperature, followed by replacement with fresh development buffer and incubated for 24 h at 37 °C to allow proteolytic digestion of the gelatin substrate. Gels were then rinsed with distilled water and stained with Coomassie blue for 30 min. Destaining was carried out with 50% methanol and 10% acetic for 1 h. Zones of gelatin degradation were imaged using an Odyssey CLx Imaging system (LI-COR Biosciences, Lincoln, NE, USA). The area of destained bands (zones of gelatin degradation) was then measured with ImageJ analyzing software and normalized to the value of vehicle treated samples. B-casein zymography was performed as described for gelatin zymography, aside from the inclusion of a 40 mA gel pre-running step performed prior to sample loading.
2.18. Statistical Analysis
The expression analysis settings for the microarray analysis were as follows: Gene-Level Fold Change < −2 or > 2, Gene-Level p-Value < 0.05, and ebayes ANOVA Method. The matrix correlation analysis was performed using GraphPad Prism software (v 8.1.2). Graphpad Prism was also used to plot all the graphs and to determine the statistical significance in this study. For a two-group comparison, unpaired t-test with Welch’s correction was used to determine the differences between the groups. For a four-group comparison, ordinary one-way ANOVA test with Dunnett’s correction was used in this study to determine the differences between SE groups as compared to HIV−Drug−. * p < 0.05, ** p < 0.01, *** p < 0.005, **** p < 0.001, and ns, nonsignificant.
2.19. Data Availability
The authors declare that all data supporting the findings of this study are available within the article. Microarray data have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE129506.
4. Discussion
In this study, we show that SEs are present in the semen of HIV-uninfected and HIV-infected participants independent of psychostimulants use. We also highlight the association of viral protein—RT and cocaine metabolite—benzoylecgonine with SE and show that SEs from study participants reprogrammed monocyte gene expression, morphometrics, and function.
Transcriptomics analysis of monocytes plated atop collagen and treated with SE from HIV−/+ and Drug−/+ groups showed that, in general, SE contain factors, such as COL16A1 and MMPs known to regulate cell adhesion [
81,
82,
83] (
Figure 2F and
Figure 3C), cell chemotaxis (
Figure 4A), and metalloprotease activity (
Figure 4B). Furthermore, SE-Drug suppressed the expression of HUS1B, linked to the induction of cell death [
85]. Amongst other observations was the result that SE-HIV suppressed the expression of TSC1, a protein known to maintain HIV in a latent state via the AKT-mTORC1 pathway [
87]. These transcriptomic results support the observation that SE-Drug and SE-HIV promote monocyte adhesion, cytoskeletal reorganization, and chemotactic migration.
Categorization of DEGs in each treatment into GO and KEGG functional terms identified a common theme—FA as a potential SE-regulated function (
Table 8). FAs are large protein complexes that physically connect the ECM to the cytoskeleton of the cells and are integral to cell adhesion, signaling, actin cytoskeleton dynamics, and cell migration while SE induced monocyte adhesion to collagen, HIV infection, and/or psychostimulant use resulted in the secretion of SE-Drug and SE-HIV that potentiated monocyte adhesion to collagen (
Figure 5). Indeed, HIV infection of lymphocytes and monocytes resulted in increased adhesion of the infected cells to vascular endothelium and ECM molecules [
95], and treatment of monocytes with HIV Tat protein increased monocyte adhesion to endothelial monolayers [
96].
Analyses of monocyte morphometrics and migration revealed that, in the presence of HIV infection and psychostimulant use, SE profoundly altered monocytes by increasing membrane ruffling and formation of filopodium-like structures. These changes resulted in variable modification of cell size and cell area by the different SEs; however, SE from HIV+Drug+ participants induced the most significant changes. These observations suggest that the ability of SE to regulate actin cytoskeleton dynamics, formation of membrane structures, FA, AJ, and localization of F-actin and vinculin to structures similar to the leading edge of migrating cells [
97] and at cell–cell junctions depend on the microenvironment secreting the SE. On the basis of these observations, it is therefore possible that exosomes may modify FAs and AJs in response to changes in their microenvironment.
While FA bridges cells to ECM, AJ links neighboring cells and the actin–myosin cytoskeleton. These processes play a role in physiologic and pathologic signaling and cell migration and invasion during morphogenetics, tissue repair, and barrier disruption events [
98,
99]. In the center of these cell-to-cell and cell-to-ECM interactions is the actin machinery [
100], which we found profoundly colocalized with vinculin in FA and AJ (
Figure 8). Thus, elevated vinculin incorporation in the adhesion complexes may explain the observed enhanced firm adhesion of SE-Drug- and SE-HIV-treated monocytes to collagen and migration of monocytes to HIV secretome.
The components of FA and AJ include scaffolding molecules, GTPases, and enzymes (kinases, phosphatases, proteases, and lipases). We found that, in comparison to vehicle-treated cells, monocytes treated with SE < SE-HIV < SE-Drug, in this order, released bioactive enzymes with enhanced gelatinolytic and caseinolytic activities. Amongst the enzymes identified by their molecular weight and banding pattern were gelatinolytic MMP2 and MMP9 and caseinolytic MMP7. However, other enzymes regulated by SE are yet to be identified.
MMPs are vital to normal immune response to infection because they degrade the ECM for leukocyte migration and modulate the activity of cytokines, chemokines, and defensins. However, MMPs have been implicated in the upregulation of adhesion molecules [
101] and in the immunopathology associated with tissue damage, metastasis, and microbial dissemination, such as bacterial meningitis, endotoxic shock, mycobacterial infection, and hepatitis B and HIV infection [
102]. Our observation that SE-Drug and SE-HIV induced high levels of gelatinolytic and caseinolytic MMPs points to the potential that the use of psychostimulants and infection with HIV may change the function of SE. Indeed, it has been shown that HIV infection is associated with an altered production and secretion of MMPs which contribute to HIV-induced immunopathology, dysregulation of T-cell dynamics, leukocyte trafficking, and viral dissemination [
72]. It is possible that the elevated MMP activity in HIV-infected cells may be related to HIV-associated immune activation, viral dissemination, and the development of HIV-associated diseases. In our studies, we observed that treatment of monocytes with SE from all clinical groups increased monocyte adhesion to collagen, with SE-Drug and SE-HIV producing the greatest increases (
Figure 5). Indeed, HIV infection of lymphocytes and monocytes results in increased adhesion of the infected cells to vascular endothelium and ECM molecules [
103] and treatment of monocytes with HIV Tat protein increased monocyte adhesion to endothelial monolayers [
104]. It has been shown that MMPs, especially MMP9, promotes the ability of HIV-infected mononuclear cells to traverse artificial basement membrane barriers [
105,
106]. Similar to our observation of induction of MMP expression by SE, MMP9 is one of the immediate early genes expressed after HIV infection of monocyte/macrophage [
107]. HIV proteins Tat and gp120 proteins upregulate MMP9 secretion from monocytes and T cells, while gp41-derived peptides stimulate MMP2 production [
107,
108]. Furthermore, elevated levels of MMP9 has been reported in vivo from patients with HIV infection not receiving ART [
109]. Thus, factors in SE in general and in SE-Drug and SE-HIV, in particular, may play important roles in orchestrating monocytes dynamics in HIV infection and substances abuse.
Finally, because of the potential role of MMP in HIV dissemination via transmigration of infected cells [
110] and HIV-associated pathologies (brain injury/neuronal damage, HIV-associated dementia (HAD) [
111,
112,
113], Kaposi’s sarcoma (KS) [
114,
115,
116], HIV-associated nephropathy (HIVAN) [
117], and periodontal diseases [
118,
119]), understanding the role of SE-Drug and SE-HIV in the induction of MMP enzyme activity may constitute a novel therapeutic approach for HIV infection.
Aside from the pathologic functions of monocytes, these cells perform systemic immune surveillance and maintenance of macrophage populations through constitutive migration from the bloodstream across the vascular endothelium. Thus, enhanced pathologic monocyte migration may promote disease while the consequence of losing constitutive migration may include defective cell-mediated immune responses. It remains to be determined whether alteration in monocyte morphometrics and motility induced by SE, SE-Drug, and SE-HIV will be pathologic or protective in primary monocytes. These interesting findings are limited by the possibility that HIV+Drug− and HIV+Drug+ SE may contain HIV particles because the semen samples were obtained before the advent of ART. Thus, HIV+ donors were almost certainly viremic. However, these limitations do not negate the findings of this study, since HIV particles and exosomes are naturally present in the semen of infected individuals. Hence, our findings highlight the possible events that may occur within individuals who are infected with HIV and use or do not use psychostimulants. Another caveat of this study is that the donors reported the use of multiple substances and alcohol. Thus, the observed effects may be due to one of the other substances or due to an interaction between different substances. Due to such complexities, well-controlled animal model studies are needed to determine drug-specific effects. Nevertheless, the presented data should pave the path for future works that study the effects of body fluids exosomes in the context of HIV infection and substance abuse.