Understanding Quantitative Circadian Regulations Are Crucial Towards Advancing Chronotherapy
Abstract
:1. Introduction
2. Molecular Insights of the Clockwork and the Feedback
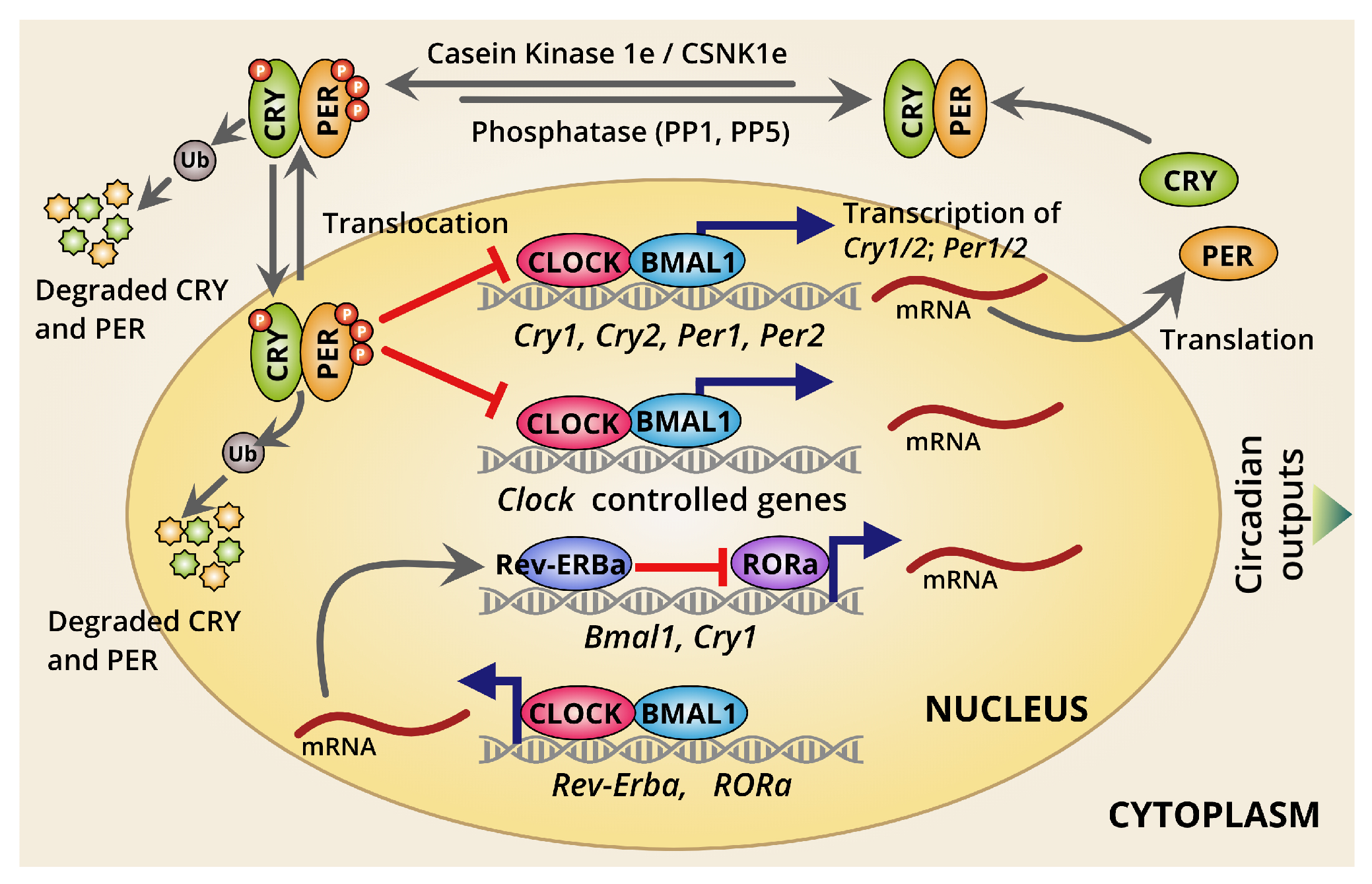
2.1. The Overview of the Mammalian Circadian Clockwork and Its Regulations
2.2. Dynamics of Circadian Transcription/Translation Feedback Loops
2.3. Different Modes of Circadian Transcriptional Regulation
3. Post-Transcriptional and Post-Translational Controls over the Molecular Oscillations
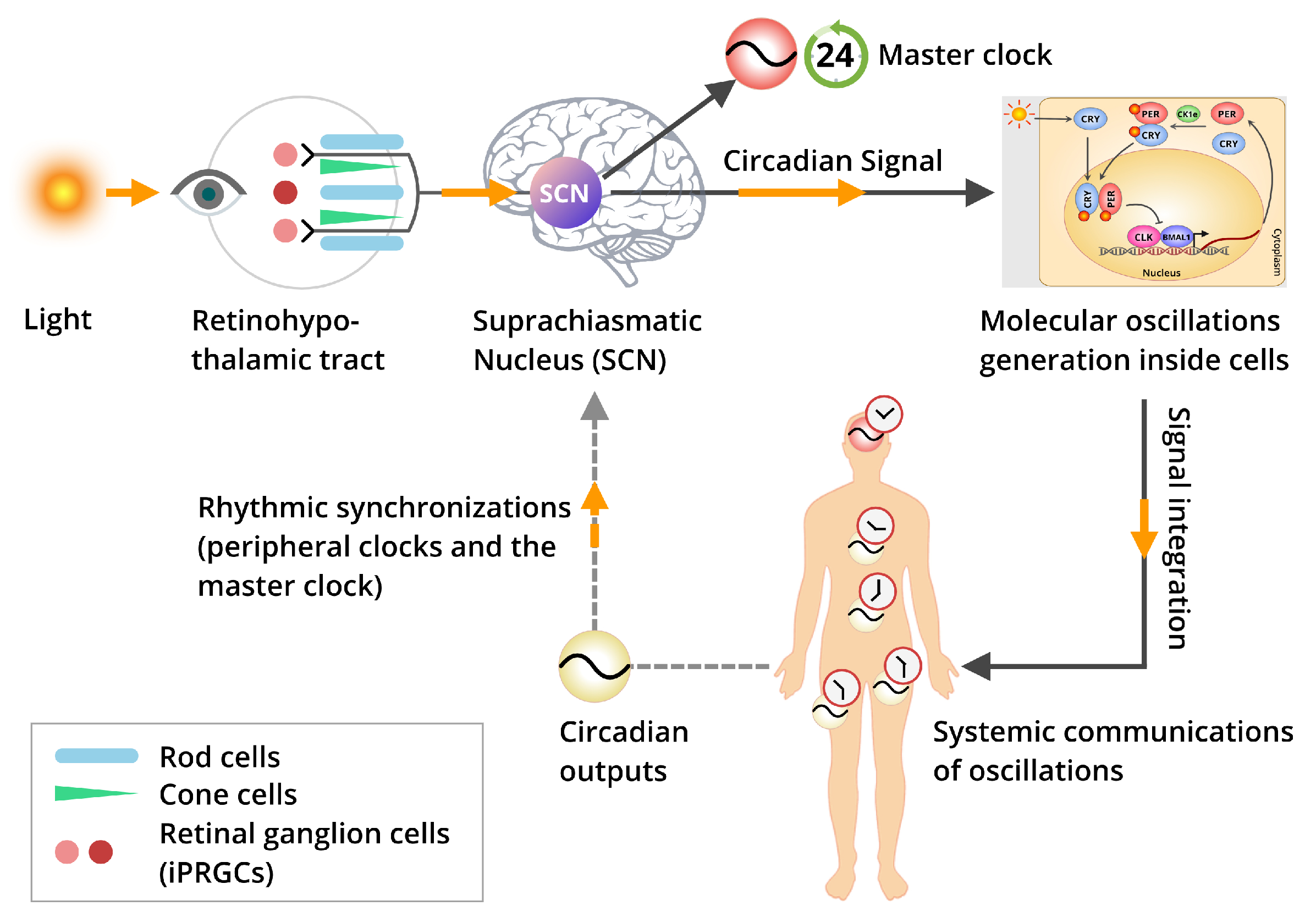
4. Light Entrainment and Synchronization of Biological Clocks
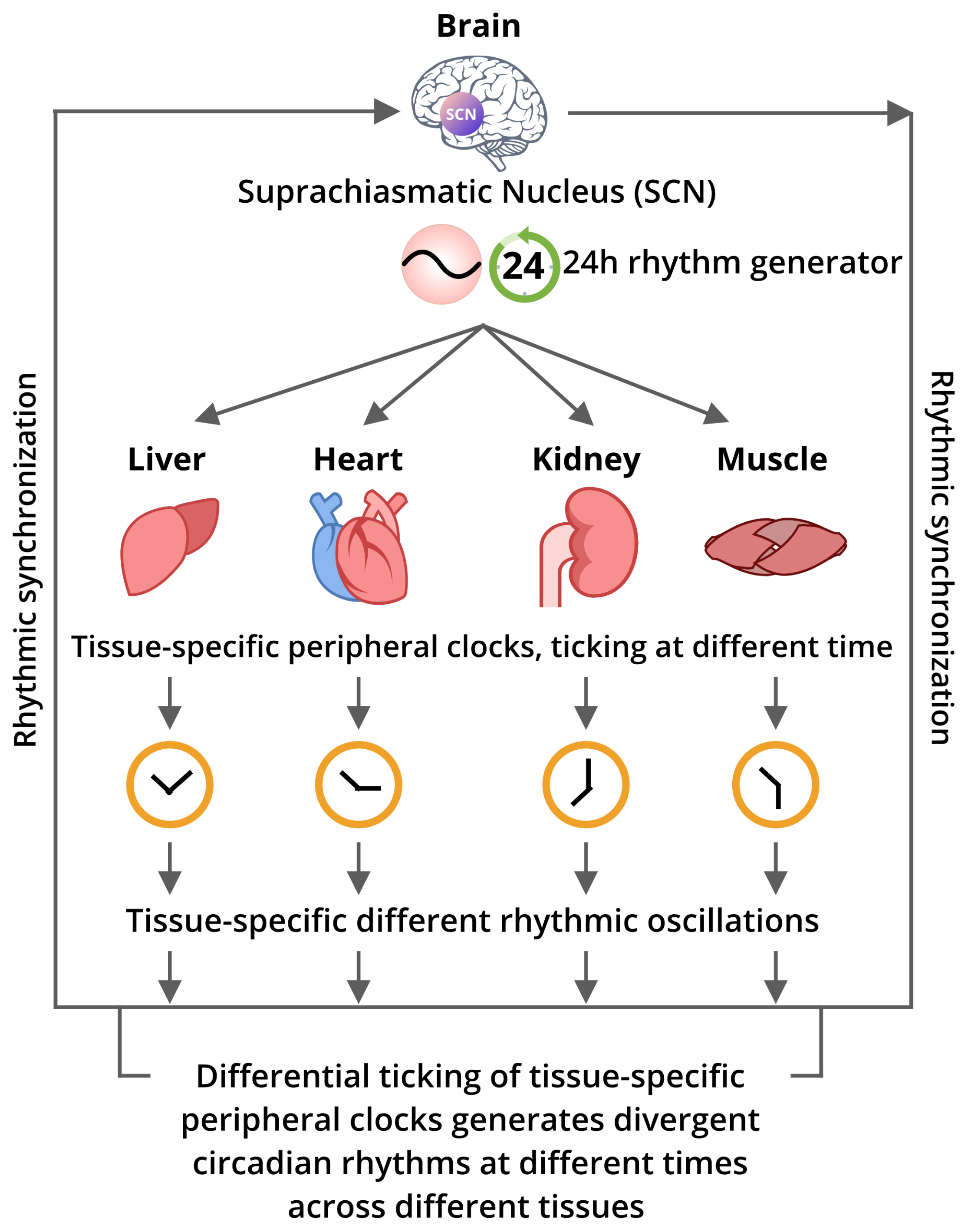
5. The Inter-Relation Among the Core Circadian Pacemaker and the Peripheral Clocks
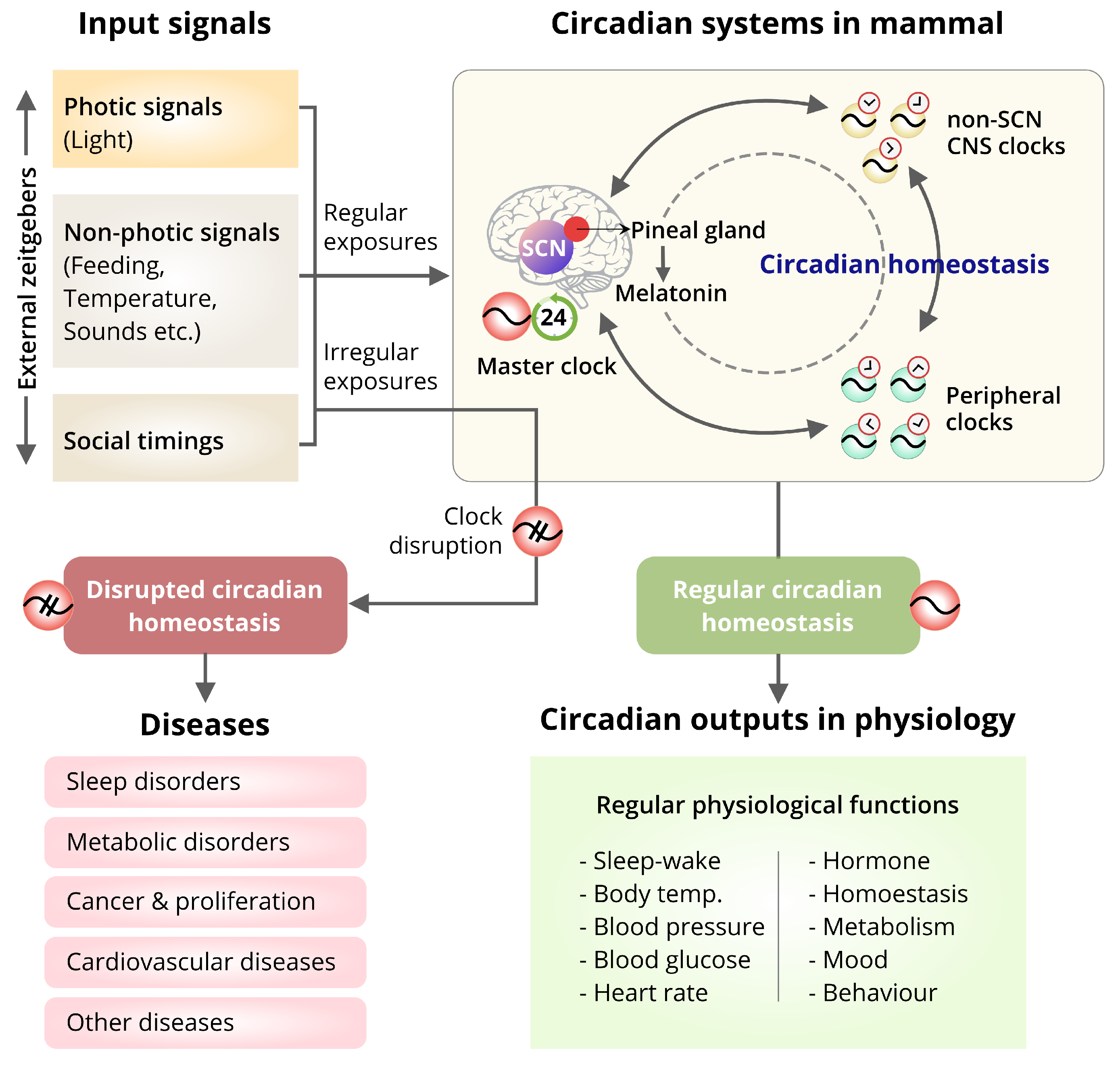
6. Circadian Biology and Human Health
6.1. Influence of the External Factors on Circadian Homeostasis and Diseases
6.2. Effect of the Misalignment of the SCN-Master Clock and Peripheral Clocks on Diseases
6.2.1. Sleep Disorders
6.2.2. Metabolic Diseases
6.2.3. Cancer
6.2.4. Cardiovascular Diseases
7. Recent Status of the Chronotherapy: A Potential Therapeutic Intervention
8. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Granados-Fuentes, D.; Herzog, E.D. The clock shop: Coupled circadian oscillators. Exp. Neurol. 2013, 243, 21–27. [Google Scholar] [CrossRef] [PubMed]
- Partch, C.L.; Green, C.B.; Takahashi, J.S. Molecular architecture of the mammalian circadian clock. Trends Cell Biol. 2014, 24, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Mure, L.S.; Le, H.D.; Benegiamo, G.; Chang, M.W.; Rios, L.; Jillani, N.; Ngotho, M.; Kariuki, T.; Dkhissi-Benyahya, O.; Cooper, H.M.; et al. Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science 2018, 359, eaao0318. [Google Scholar] [CrossRef] [PubMed]
- Koike, N.; Yoo, S.H.; Huang, H.C.; Kumar, V.; Lee, C.; Kim, T.K.; Takahashi, J.S. Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science 2012, 338, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Rey, G.; Cesbron, F.; Rougemont, J.; Reinke, H.; Brunner, M.; Naef, F. Genome-wide and phase-specific DNA-binding rhythms of BMAL1 control circadian output functions in mouse liver. PLoS Biol. 2011, 9, e1000595. [Google Scholar] [CrossRef] [PubMed]
- Schmutz, I.; Ripperger, J.A.; Baeriswyl-Aebischer, S.; Albrecht, U. The mammalian clock component PERIOD2 coordinates circadian output by interaction with nuclear receptors. Genes Dev. 2010, 24, 345–357. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lamia, K.A.; Papp, S.J.; Ruth, T.Y.; Barish, G.D.; Uhlenhaut, N.H.; Jonker, J.W.; Downes, M.; Evans, R.M. Cryptochromes mediate rhythmic repression of the glucocorticoid receptor. Nature 2011, 480, 552. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, K.; Robles, M.S.; Westerling, T.; Weitz, C.J. Feedback regulation of transcriptional termination by the mammalian circadian clock PERIOD complex. Science 2012, 337, 599–602. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.C.; Welsh, D.K.; Ko, C.H.; Tran, H.G.; Zhang, E.E.; Priest, A.A.; Buhr, E.D.; Singer, O.; Meeker, K.; Verma, I.M.; et al. Intercellular coupling confers robustness against mutations in the SCN circadian clock network. Cell 2007, 129, 605–616. [Google Scholar] [CrossRef]
- Cho, H.; Zhao, X.; Hatori, M.; Ruth, T.Y.; Barish, G.D.; Lam, M.T.; Chong, L.W.; DiTacchio, L.; Atkins, A.R.; Glass, C.K.; et al. Regulation of circadian behaviour and metabolism by REV-ERB-α and REV-ERB-β. Nature 2012, 485, 123. [Google Scholar] [CrossRef]
- Sato, T.K.; Panda, S.; Miraglia, L.J.; Reyes, T.M.; Rudic, R.D.; McNamara, P.; Naik, K.A.; FitzGerald, G.A.; Kay, S.A.; Hogenesch, J.B. A functional genomics strategy reveals Rora as a component of the mammalian circadian clock. Neuron 2004, 43, 527–537. [Google Scholar] [CrossRef] [PubMed]
- Preitner, N.; Damiola, F.; Zakany, J.; Duboule, D.; Albrecht, U.; Schibler, U. The orphan nuclear receptor REV-ERBα controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 2002, 110, 251–260. [Google Scholar] [CrossRef]
- Damiola, F.; Schibler, U. Orphan nuclear receptors, molecular clockwork, and the entrainment of peripheral oscillators. Mol. Clocks Light Signal. 2003. [Google Scholar] [CrossRef]
- Menet, J.S.; Rodriguez, J.; Abruzzi, K.C.; Rosbash, M. Nascent-Seq reveals novel features of mouse circadian transcriptional regulation. elife 2012, 1, e00011. [Google Scholar] [CrossRef] [PubMed]
- Suter, D.M.; Molina, N.; Gatfield, D.; Schneider, K.; Schibler, U.; Naef, F. Mammalian genes are transcribed with widely different bursting kinetics. Science 2011, 332, 472–474. [Google Scholar] [CrossRef] [PubMed]
- Suter, D.M.; Molina, N.; Naef, F.; Schibler, U. Origins and consequences of transcriptional discontinuity. Curr. Opin. Cell Biol. 2011, 23, 657–662. [Google Scholar] [CrossRef] [PubMed]
- Terajima, H.; Yoshitane, H.; Ozaki, H.; Suzuki, Y.; Shimba, S.; Kuroda, S.; Iwasaki, W.; Fukada, Y. ADARB1 catalyzes circadian A-to-I editing and regulates RNA rhythm. Nat. Genet. 2017, 49, 146. [Google Scholar] [CrossRef]
- Ralph, M.R.; Menaker, M. A mutation of the circadian system in golden hamsters. Science 1988, 241, 1225–1227. [Google Scholar] [CrossRef]
- Lowrey, P.L.; Shimomura, K.; Antoch, M.P.; Yamazaki, S.; Zemenides, P.D.; Ralph, M.R.; Menaker, M.; Takahashi, J.S. Positional syntenic cloning and functional characterization of the mammalian circadian mutation tau. Science 2000, 288, 483–491. [Google Scholar] [CrossRef]
- Toh, K.L.; Jones, C.R.; He, Y.; Eide, E.J.; Hinz, W.A.; Virshup, D.M.; Ptáček, L.J.; Fu, Y.H. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 2001, 291, 1040–1043. [Google Scholar] [CrossRef]
- Gallego, M.; Virshup, D.M. Post-translational modifications regulate the ticking of the circadian clock. Nat. Rev. Mol. Cell Biol. 2007, 8, 139. [Google Scholar] [CrossRef]
- Hirota, T.; Lewis, W.G.; Liu, A.C.; Lee, J.W.; Schultz, P.G.; Kay, S.A. A chemical biology approach reveals period shortening of the mammalian circadian clock by specific inhibition of GSK-3β. Proc. Natl. Acad. Sci. USA 2008, 105, 20746–20751. [Google Scholar] [CrossRef]
- Siepka, S.M.; Yoo, S.H.; Park, J.; Song, W.; Kumar, V.; Hu, Y.; Lee, C.; Takahashi, J.S. Circadian mutant Overtime reveals F-box protein FBXL3 regulation of cryptochrome and period gene expression. Cell 2007, 129, 1011–1023. [Google Scholar] [CrossRef]
- Godinho, S.I.; Maywood, E.S.; Shaw, L.; Tucci, V.; Barnard, A.R.; Busino, L.; Pagano, M.; Kendall, R.; Quwailid, M.M.; Romero, M.R.; et al. The after-hours mutant reveals a role for Fbxl3 in determining mammalian circadian period. Science 2007, 316, 897–900. [Google Scholar] [CrossRef]
- Belden, W.J.; Dunlap, J.C. SIRT1 is a circadian deacetylase for core clock components. Cell 2008, 134, 212–214. [Google Scholar] [CrossRef]
- Cardone, L.; Hirayama, J.; Giordano, F.; Tamaru, T.; Palvimo, J.J.; Sassone-Corsi, P. Circadian clock control by SUMOylation of BMAL1. Science 2005, 309, 1390–1394. [Google Scholar] [CrossRef]
- Xu, Y.; Padiath, Q.S.; Shapiro, R.E.; Jones, C.R.; Wu, S.C.; Saigoh, N.; Saigoh, K.; Ptáček, L.J.; Fu, Y.H. Functional consequences of a CKIδ mutation causing familial advanced sleep phase syndrome. Nature 2005, 434, 640. [Google Scholar] [CrossRef]
- He, Y.; Jones, C.R.; Fujiki, N.; Xu, Y.; Guo, B.; Holder, J.L.; Rossner, M.J.; Nishino, S.; Fu, Y.H. The transcriptional repressor DEC2 regulates sleep length in mammals. Science 2009, 325, 866–870. [Google Scholar] [CrossRef]
- Ebisawa, T.; Uchiyama, M.; Kajimura, N.; Mishima, K.; Kamei, Y.; Katoh, M.; Watanabe, T.; Sekimoto, M.; Shibui, K.; Kim, K.; et al. Association of structural polymorphisms in the human period3 gene with delayed sleep phase syndrome. EMBO Rep. 2001, 2, 342–346. [Google Scholar] [CrossRef]
- Partonen, T. Clock gene variants in mood and anxiety disorders. J. Neural Transm. 2012, 119, 1133–1145. [Google Scholar] [CrossRef]
- Albrecht, U. Molecular mechanisms in mood regulation involving the circadian clock. Front. Neurol. 2017, 8, 30. [Google Scholar] [CrossRef]
- De Leersnyder, H.; Claustrat, B.; Munnich, A.; Verloes, A. Circadian rhythm disorder in a rare disease: Smith–Magenis syndrome. Mol. Cell. Endocrinol. 2006, 252, 88–91. [Google Scholar] [CrossRef]
- Shi, S.Q.; Bichell, T.J.; Ihrie, R.A.; Johnson, C.H. Ube3a imprinting impairs circadian robustness in Angelman syndrome models. Curr. Biol. 2015, 25, 537–545. [Google Scholar] [CrossRef]
- Tsuchiya, Y.; Minami, Y.; Umemura, Y.; Watanabe, H.; Ono, D.; Nakamura, W.; Takahashi, T.; Honma, S.; Kondoh, G.; Matsuishi, T.; et al. Disruption of Me CP 2 attenuates circadian rhythm in CRISPR/Cas9-based Rett syndrome model mouse. Genes Cells 2015, 20, 992–1005. [Google Scholar] [CrossRef]
- Field, M.D.; Maywood, E.S.; O’Brien, J.A.; Weaver, D.R.; Reppert, S.M.; Hastings, M.H. Analysis of clock proteins in mouse SCN demonstrates phylogenetic divergence of the circadian clockwork and resetting mechanisms. Neuron 2000, 25, 437–447. [Google Scholar] [CrossRef]
- Honma, S. The mammalian circadian system: A hierarchical multi-oscillator structure for generating circadian rhythm. J. Physiol. Sci. 2018, 68, 207–219. [Google Scholar] [CrossRef]
- Abbott, K.S.; Queener, H.M.; Ostrin, L.A. The ipRGC-driven pupil response with light exposure, refractive error, and sleep. Optom. Vis. Sci. 2018, 95, 323–331. [Google Scholar] [CrossRef]
- Hatori, M.; Gronfier, C.; Van Gelder, R.N.; Bernstein, P.S.; Carreras, J.; Panda, S.; Marks, F.; Sliney, D.; Hunt, C.E.; Hirota, T.; et al. Global rise of potential health hazards caused by blue light-induced circadian disruption in modern aging societies. npj Aging Mech. Dis. 2017, 3, 9. [Google Scholar] [CrossRef]
- Chew, K.S.; Renna, J.M.; McNeill, D.S.; Fernandez, D.C.; Keenan, W.T.; Thomsen, M.B.; Ecker, J.L.; Loevinsohn, G.S.; VanDunk, C.; Vicarel, D.C.; et al. A subset of ipRGCs regulates both maturation of the circadian clock and segregation of retinogeniculate projections in mice. Elife 2017, 6, e22861. [Google Scholar] [CrossRef]
- Buijs, F.N.; León-Mercado, L.; Guzmán-Ruiz, M.; Guerrero-Vargas, N.N.; Romo-Nava, F.; Buijs, R.M. The circadian system: A regulatory feedback network of periphery and brain. Physiology 2016, 31, 170–181. [Google Scholar] [CrossRef]
- Mohawk, J.A.; Green, C.B.; Takahashi, J.S. Central and peripheral circadian clocks in mammals. Annu. Rev. Neurosci. 2012, 35, 445–462. [Google Scholar] [CrossRef]
- Eckel-Mahan, K.L.; Patel, V.R.; Mohney, R.P.; Vignola, K.S.; Baldi, P.; Sassone-Corsi, P. Coordination of the transcriptome and metabolome by the circadian clock. Proc. Natl. Acad. Sci. USA 2012, 109, 5541–5546. [Google Scholar] [CrossRef] [Green Version]
- Honma, S.; Ono, D.; Suzuki, Y.; Inagaki, N.; Yoshikawa, T.; Nakamura, W.; Honma, K.I. Suprachiasmatic nucleus: Cellular clocks and networks. In Progress in Brain Research; Elsevier: Amsterdam, The Netherlands, 2012; Volume 199, pp. 129–141. [Google Scholar]
- Balsalobre, A.; Damiola, F.; Schibler, U. A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 1998, 93, 929–937. [Google Scholar] [CrossRef]
- Panda, S. Circadian physiology of metabolism. Science 2016, 354, 1008–1015. [Google Scholar] [CrossRef] [Green Version]
- Castelo-Szekely, V.; Arpat, A.B.; Janich, P.; Gatfield, D. Translational contributions to tissue specificity in rhythmic and constitutive gene expression. Genome Biol. 2017, 18, 116. [Google Scholar] [CrossRef]
- Antle, M.C.; Silver, R. Orchestrating time: Arrangements of the brain circadian clock. Trends Neurosci. 2005, 28, 145–151. [Google Scholar] [CrossRef]
- El Cheikh Hussein, L.; Mollard, P.; Bonnefont, X. Molecular and Cellular Networks in The Suprachiasmatic Nuclei. Int. J. Mol. Sci. 2019, 20, 2052. [Google Scholar] [CrossRef]
- Lehman, M.N.; Silver, R.; Gladstone, W.; Kahn, R.M.; Gibson, M.; Bittman, E.L. Circadian rhythmicity restored by neural transplant. Immunocytochemical characterization of the graft and its integration with the host brain. J. Neurosci. 1987, 7, 1626–1638. [Google Scholar] [CrossRef] [Green Version]
- Ecker, J.L.; Dumitrescu, O.N.; Wong, K.Y.; Alam, N.M.; Chen, S.K.; LeGates, T.; Renna, J.M.; Prusky, G.T.; Berson, D.M.; Hattar, S. Melanopsin-expressing retinal ganglion-cell photoreceptors: Cellular diversity and role in pattern vision. Neuron 2010, 67, 49–60. [Google Scholar] [CrossRef]
- Golombek, D.A.; Rosenstein, R.E. Physiology of circadian entrainment. Physiol. Rev. 2010, 90, 1063–1102. [Google Scholar] [CrossRef]
- Antle, M.C.; Smith, V.M.; Sterniczuk, R.; Yamakawa, G.R.; Rakai, B.D. Physiological responses of the circadian clock to acute light exposure at night. Rev. Endocr. Metab. Disord. 2009, 10, 279–291. [Google Scholar] [CrossRef]
- Guilding, C.; Piggins, H.D. Challenging the omnipotence of the suprachiasmatic timekeeper: Are circadian oscillators present throughout the mammalian brain? Eur. J. Neurosci. 2007, 25, 3195–3216. [Google Scholar] [CrossRef]
- Colwell, C.S. Linking neural activity and molecular oscillations in the SCN. Nat. Rev. Neurosci. 2011, 12, 553. [Google Scholar] [CrossRef]
- Albrecht, U. Timing to perfection: The biology of central and peripheral circadian clocks. Neuron 2012, 74, 246–260. [Google Scholar] [CrossRef]
- Chang, A.M.; Aeschbach, D.; Duffy, J.F.; Czeisler, C.A. Evening use of light-emitting eReaders negatively affects sleep, circadian timing, and next-morning alertness. Proc. Natl. Acad. Sci. USA 2015, 112, 1232–1237. [Google Scholar] [CrossRef]
- Roenneberg, T.; Allebrandt, K.V.; Merrow, M.; Vetter, C. Social jetlag and obesity. Curr. Biol. 2012, 22, 939–943. [Google Scholar] [CrossRef]
- Schernhammer, E.S.; Laden, F.; Speizer, F.E.; Willett, W.C.; Hunter, D.J.; Kawachi, I.; Colditz, G.A. Rotating night shifts and risk of breast cancer in women participating in the nurses’ health study. J. Natl. Cancer Inst. 2001, 93, 1563–1568. [Google Scholar] [CrossRef]
- Pan, A.; Schernhammer, E.S.; Sun, Q.; Hu, F.B. Rotating night shift work and risk of type 2 diabetes: Two prospective cohort studies in women. PLoS Med. 2011, 8, e1001141. [Google Scholar] [CrossRef]
- Nakamura, T.J.; Nakamura, W.; Yamazaki, S.; Kudo, T.; Cutler, T.; Colwell, C.S.; Block, G.D. Age-related decline in circadian output. J. Neurosci. 2011, 31, 10201–10205. [Google Scholar] [CrossRef]
- Kondratov, R.V.; Kondratova, A.A.; Gorbacheva, V.Y.; Vykhovanets, O.V.; Antoch, M.P. Early aging and age-related pathologies in mice deficient in BMAL1, the core componentof the circadian clock. Genes Dev. 2006, 20, 1868–1873. [Google Scholar] [CrossRef] [Green Version]
- Fu, L.; Pelicano, H.; Liu, J.; Huang, P.; Lee, C.C. The circadian gene Period2 plays an important role in tumor suppression and DNA damage response in vivo. Cell 2002, 111, 41–50. [Google Scholar] [CrossRef]
- Panda, S.; Antoch, M.P.; Miller, B.H.; Su, A.I.; Schook, A.B.; Straume, M.; Schultz, P.G.; Kay, S.A.; Takahashi, J.S.; Hogenesch, J.B. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 2002, 109, 307–320. [Google Scholar] [CrossRef]
- Sobel, J.A.; Krier, I.; Andersin, T.; Raghav, S.; Canella, D.; Gilardi, F.; Kalantzi, A.S.; Rey, G.; Weger, B.; Gachon, F.; et al. Transcriptional regulatory logic of the diurnal cycle in the mouse liver. PLoS Biol. 2017, 15, e2001069. [Google Scholar] [CrossRef]
- Storch, K.F.; Lipan, O.; Leykin, I.; Viswanathan, N.; Davis, F.C.; Wong, W.H.; Weitz, C.J. Extensive and divergent circadian gene expression in liver and heart. Nature 2002, 417, 78. [Google Scholar] [CrossRef]
- Nguyen, T.T.; Mattick, J.S.; Yang, Q.; Orman, M.A.; Ierapetritou, M.G.; Berthiaume, F.; Androulakis, I.P. Bioinformatics analysis of transcriptional regulation of circadian genes in rat liver. BMC Bioinform. 2014, 15, 83. [Google Scholar] [CrossRef]
- Le Martelot, G.; Canella, D.; Symul, L.; Migliavacca, E.; Gilardi, F.; Liechti, R.; Martin, O.; Harshman, K.; Delorenzi, M.; Desvergne, B.; et al. Genome-wide RNA polymerase II profiles and RNA accumulation reveal kinetics of transcription and associated epigenetic changes during diurnal cycles. PLoS Biol. 2012, 10, e1001442. [Google Scholar] [CrossRef]
- Vollmers, C.; Schmitz, R.J.; Nathanson, J.; Yeo, G.; Ecker, J.R.; Panda, S. Circadian oscillations of protein-coding and regulatory RNAs in a highly dynamic mammalian liver epigenome. Cell Metab. 2012, 16, 833–845. [Google Scholar] [CrossRef]
- Yan, B.; Guan, D.; Wang, C.; Wang, J.; He, B.; Qin, J.; Boheler, K.R.; Lu, A.; Zhang, G.; Zhu, H. An integrative method to decode regulatory logics in gene transcription. Nat. Commun. 2017, 8, 1044. [Google Scholar] [CrossRef]
- Zee, P.C.; Attarian, H.; Videnovic, A. Circadian rhythm abnormalities. Contin. Lifelong Learn. Neurol. 2013, 19, 132. [Google Scholar] [CrossRef]
- Jagannath, A.; Taylor, L.; Wakaf, Z.; Vasudevan, S.R.; Foster, R.G. The genetics of circadian rhythms, sleep and health. Hum. Mol. Genet. 2017, 26, R128–R138. [Google Scholar] [CrossRef]
- Santamaria, F.; Esposito, M.; Montella, S.; Cantone, E.; Mollica, C.; De Stefano, S.; Mirra, V.; Carotenuto, M. Sleep disordered breathing and airway disease in primary ciliary dyskinesia. Respirology 2014, 19, 570–575. [Google Scholar] [CrossRef]
- Sutton, C.E.; Finlay, C.M.; Raverdeau, M.; Early, J.O.; DeCourcey, J.; Zaslona, Z.; O’Neill, L.A.; Mills, K.H.; Curtis, A.M. Loss of the molecular clock in myeloid cells exacerbates T cell-mediated CNS autoimmune disease. Nat. Commun. 2017, 8, 1923. [Google Scholar] [CrossRef]
- Baiardi, S.; Cirignotta, F.; Cicolin, A.; Garbazza, C.; D’Agostino, A.; Gambini, O.; Giordano, A.; Canevini, M.; Zambrelli, E.; Marconi, A.M.; et al. Chronobiology, sleep-related risk factors and light therapy in perinatal depression: The “Life-ON” project. BMC Psychiatry 2016, 16, 374. [Google Scholar] [CrossRef]
- Bass, J.; Lazar, M.A. Circadian time signatures of fitness and disease. Science 2016, 354, 994–999. [Google Scholar] [CrossRef] [Green Version]
- Schulz, P.; Steimer, T. Neurobiology of circadian systems. CNS Drugs 2009, 23, 3–13. [Google Scholar] [CrossRef]
- Ma, D.; Liu, T.; Chang, L.; Rui, C.; Xiao, Y.; Li, S.; Hogenesch, J.B.; Chen, Y.E.; Lin, J.D. The liver clock controls cholesterol homeostasis through Trib1 protein-mediated regulation of PCSK9/Low density lipoprotein receptor (LDLR) axis. J. Biol. Chem. 2015, 290, 31003–31012. [Google Scholar] [CrossRef]
- Sato, S.; Solanas, G.; Peixoto, F.O.; Bee, L.; Symeonidi, A.; Schmidt, M.S.; Brenner, C.; Masri, S.; Benitah, S.A.; Sassone-Corsi, P. Circadian reprogramming in the liver identifies metabolic pathways of aging. Cell 2017, 170, 664–677. [Google Scholar] [CrossRef]
- Musiek, E.S.; Holtzman, D.M. Mechanisms linking circadian clocks, sleep, and neurodegeneration. Science 2016, 354, 1004–1008. [Google Scholar] [CrossRef] [Green Version]
- Kitazawa, M. Circadian rhythms, metabolism, and insulin sensitivity: transcriptional networks in animal models. Curr. Diabetes Rep. 2013, 13, 223–228. [Google Scholar] [CrossRef]
- Papagiannakopoulos, T.; Bauer, M.R.; Davidson, S.M.; Heimann, M.; Subbaraj, L.; Bhutkar, A.; Bartlebaugh, J.; Vander Heiden, M.G.; Jacks, T. Circadian rhythm disruption promotes lung tumorigenesis. Cell Metab. 2016, 24, 324–331. [Google Scholar] [CrossRef]
- Czeisler, C.A.; Duffy, J.F.; Shanahan, T.L.; Brown, E.N.; Mitchell, J.F.; Rimmer, D.W.; Ronda, J.M.; Silva, E.J.; Allan, J.S.; Emens, J.S.; et al. Stability, precision, and near-24-h period of the human circadian pacemaker. Science 1999, 284, 2177–2181. [Google Scholar] [CrossRef]
- Sack, R.L.; Lewy, A.J.; Blood, M.L.; Keith, L.D.; Nakagawa, H. Circadian rhythm abnormalities in totally blind people: Incidence and clinical significance. J. Clin. Endocrinol. Metab. 1992, 75, 127–134. [Google Scholar]
- Sack, R.L.; Auckley, D.; Auger, R.R.; Carskadon, M.A.; Wright, K.P., Jr.; Vitiello, M.V.; Zhdanova, I.V. Circadian rhythm sleep disorders: Part II, advanced sleep phase disorder, delayed sleep phase disorder, free-running disorder, and irregular sleep/wake rhythm. Sleep 2007, 30, 1484–1501. [Google Scholar] [CrossRef]
- Quera Salva, M.A.; Hartley, S.; Léger, D.; Dauvilliers, Y.A. Non-24-h sleep–wake rhythm disorder in the totally blind: Diagnosis and management. Front. Neurol. 2017, 8, 686. [Google Scholar] [CrossRef]
- Morgenthaler, T.I.; Lee-Chiong, T.; Alessi, C.; Friedman, L.; Aurora, R.N.; Boehlecke, B.; Brown, T.; Chesson, A.L., Jr.; Kapur, V.; Maganti, R.; et al. Practice parameters for the clinical evaluation and treatment of circadian rhythm sleep disorders. Sleep 2007, 30, 1445–1459. [Google Scholar] [CrossRef]
- Edgar, D.M.; Dement, W.C.; Fuller, C.A. Effect of SCN lesions on sleep in squirrel monkeys: Evidence for opponent processes in sleep/wake regulation. J. Neurosci. 1993, 13, 1065–1079. [Google Scholar] [CrossRef]
- Pang, S.; Brown, G.; Grota, L.; Chambers, J.; Rodman, R. Determination of N-acetylserotonin and melatonin activities in the pineal gland, retina, Harderian gland, brain and serum of rats and chickens. Neuroendocrinology 1977, 23, 1–13. [Google Scholar] [CrossRef]
- Lewy, A. Melatonin and human chronobiology. In Cold Spring Harbor Symposia on Quantitative Biology; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2007; Volume 72, pp. 623–636. [Google Scholar]
- Lewy, A.J.; Emens, J.; Jackman, A.; Yuhas, K. Circadian uses of melatonin in humans. Chronobiol. Int. 2006, 23, 403–412. [Google Scholar] [CrossRef]
- Grof, E.; Grof, P.; Brown, G.M.; Arato, M.; Lane, J. Investigations of melatonin secretion in man. Prog. Neuro-Psychopharmacol. Biol. Psychiatry 1985, 9, 609–612. [Google Scholar] [CrossRef]
- Bartness, T.; Goldman, B. Mammalian pineal melatonin: A clock for all seasons. Experientia 1989, 45, 939–945. [Google Scholar] [CrossRef]
- Brown, G.M.; Pandi-Perumal, S.R.; Trakht, I.; Cardinali, D.P. Melatonin and its relevance to jet lag. Travel Med. Infect. Dis. 2009, 7, 69–81. [Google Scholar] [CrossRef]
- Claustrat, B.; Brun, J.; Chazot, G. The basic physiology and pathophysiology of melatonin. Sleep Med. Rev. 2005, 9, 11–24. [Google Scholar] [CrossRef]
- Khalsa, S.B.S.; Jewett, M.E.; Cajochen, C.; Czeisler, C.A. A phase response curve to single bright light pulses in human subjects. J. Physiol. 2003, 549, 945–952. [Google Scholar] [CrossRef]
- Thompson, A.; Batterham, A.; Jones, H.; Gregson, W.; Scott, D.; Atkinson, G. The practicality and effectiveness of supplementary bright light for reducing jet-lag in elite female athletes. Int. J. Sport. Med. 2013, 34, 582–589. [Google Scholar] [CrossRef]
- Dubocovich, M.L.; Markowska, M. Functional MT 1 and MT 2 melatonin receptors in mammals. Endocrine 2005, 27, 101–110. [Google Scholar] [CrossRef]
- Liu, C.; Weaver, D.R.; Jin, X.; Shearman, L.P.; Pieschl, R.L.; Gribkoff, V.K.; Reppert, S.M. Molecular dissection of two distinct actions of melatonin on the suprachiasmatic circadian clock. Neuron 1997, 19, 91–102. [Google Scholar] [CrossRef]
- Von Gall, C.; Stehle, J.H.; Weaver, D.R. Mammalian melatonin receptors: Molecular biology and signal transduction. Cell Tissue Res. 2002, 309, 151–162. [Google Scholar] [CrossRef]
- Hunt, A.E.; Al-Ghoul, W.M.; Gillette, M.U.; Dubocovich, M.L. Activation of MT2 melatonin receptors in rat suprachiasmatic nucleus phase advances the circadian clock. Am. J. Physiol.-Cell Physiol. 2001, 280, C110–C118. [Google Scholar] [CrossRef]
- Wan, Q.; Man, H.Y.; Liu, F.; Braunton, J.; Niznik, H.B.; Pang, S.F.; Brown, G.M.; Wang, Y.T. Differential modulation of GABA A receptor function by Mel 1a and Mel 1b receptors. Nat. Neurosci. 1999, 2, 401. [Google Scholar] [CrossRef]
- Liu, C.; Reppert, S.M. GABA synchronizes clock cells within the suprachiasmatic circadian clock. Neuron 2000, 25, 123–128. [Google Scholar] [CrossRef]
- Arendt, J. Managing jet lag: Some of the problems and possible new solutions. Sleep Med. Rev. 2009, 13, 249–256. [Google Scholar] [CrossRef]
- Arendt, J.; Skene, D.J. Melatonin as a chronobiotic. Sleep Med. Rev. 2005, 9, 25–39. [Google Scholar] [CrossRef]
- Burgess, H.J.; Sharkey, K.M.; Eastman, C.I. Bright light, dark and melatonin can promote circadian adaptation in night shift workers. Sleep Med. Rev. 2002, 6, 407–420. [Google Scholar] [CrossRef]
- Gordon, C.J.; Comas, M.; Postnova, S.; Miller, C.B.; Roy, D.; Bartlett, D.J.; Grunstein, R.R. The effect of consecutive transmeridian flights on alertness, sleep–wake cycles and sleepiness: A case study. Chronobiol. Int. 2018, 35, 1471–1480. [Google Scholar] [CrossRef]
- Shiota, M.; Sudou, M.; Ohshima, M. Using outdoor exercise to decrease jet lag in airline crewmembers. Aviat. Space Environ. Med. 1996, 67, 1155–1160. [Google Scholar]
- Fowler, P.M.; Duffield, R.; Morrow, I.; Roach, G.; Vaile, J. Effects of sleep hygiene and artificial bright light interventions on recovery from simulated international air travel. Eur. J. Appl. Physiol. 2015, 115, 541–553. [Google Scholar] [CrossRef]
- Sack, R.L. The pathophysiology of jet lag. Travel Med. Infect. Dis. 2009, 7, 102–110. [Google Scholar] [CrossRef]
- Daan, S.; Beersma, D.; Borbély, A.A. Timing of human sleep: Recovery process gated by a circadian pacemaker. Am. J. Physiol.-Regul. Integr. Comp. Physiol. 1984, 246, R161–R183. [Google Scholar] [CrossRef]
- Wong, W.S.; Fielding, R. Prevalence of insomnia among Chinese adults in Hong Kong: A population-based study. J. Sleep Res. 2011, 20, 117–126. [Google Scholar] [CrossRef]
- Holst, S.C.; Valomon, A.; Landolt, H.P. Sleep pharmacogenetics: Personalized sleep/wake therapy. Annu. Rev. Pharmacol. Toxicol. 2016, 56, 577–603. [Google Scholar] [CrossRef]
- Burke, T.M.; Markwald, R.R.; McHill, A.W.; Chinoy, E.D.; Snider, J.A.; Bessman, S.C.; Jung, C.M.; O’Neill, J.S.; Wright, K.P. Effects of caffeine on the human circadian clock in vivo and in vitro. Sci. Transl. Med. 2015, 7, 305ra146. [Google Scholar] [CrossRef]
- Oike, H.; Kobori, M.; Suzuki, T.; Ishida, N. Caffeine lengthens circadian rhythms in mice. Biochem. Biophys. Res. Commun. 2011, 410, 654–658. [Google Scholar] [CrossRef]
- Van Diepen, H.C.; Lucassen, E.A.; Yasenkov, R.; Groenen, I.; Ijzerman, A.P.; Meijer, J.H.; Deboer, T. Caffeine increases light responsiveness of the mouse circadian pacemaker. Eur. J. Neurosci. 2014, 40, 3504–3511. [Google Scholar] [CrossRef]
- Chen, G.; Van Den Pol, A.N. Adenosine modulation of calcium currents and presynaptic inhibition of GABA release in suprachiasmatic and arcuate nucleus neurons. J. Neurophysiol. 1997, 77, 3035–3047. [Google Scholar] [CrossRef]
- Conlay, L.A.; Conant, J.A.; Debros, F.; Wurtman, R. Caffeine alters plasma adenosine levels. Nature 1997, 389, 136. [Google Scholar] [CrossRef]
- O’Neill, J.S.; Maywood, E.S.; Chesham, J.E.; Takahashi, J.S.; Hastings, M.H. cAMP-dependent signaling as a core component of the mammalian circadian pacemaker. Science 2008, 320, 949–953. [Google Scholar] [CrossRef]
- Huang, M.C.; Ho, C.W.; Chen, C.H.; Liu, S.C.; Chen, C.C.; Leu, S.J. Reduced expression of circadian clock genes in male alcoholic patients. Alcohol. Clin. Exp. Res. 2010, 34, 1899–1904. [Google Scholar] [CrossRef]
- Karasek, M.; Winczyk, K. Melatonin in humans. J. Physiol. Pharmacol. 2006, 57, 19. [Google Scholar]
- Turek, F.W.; Joshu, C.; Kohsaka, A.; Lin, E.; Ivanova, G.; McDearmon, E.; Laposky, A.; Losee-Olson, S.; Easton, A.; Jensen, D.R.; et al. Obesity and metabolic syndrome in circadian Clock mutant mice. Science 2005, 308, 1043–1045. [Google Scholar] [CrossRef]
- Saper, C.B.; Scammell, T.E.; Lu, J. Hypothalamic regulation of sleep and circadian rhythms. Nature 2005, 437, 1257. [Google Scholar] [CrossRef]
- Fuller, P.M.; Gooley, J.J.; Saper, C.B. Neurobiology of the sleep/wake cycle: Sleep architecture, circadian regulation, and regulatory feedback. J. Biol. Rhythm. 2006, 21, 482–493. [Google Scholar] [CrossRef]
- Xie, L.; Kang, H.; Xu, Q.; Chen, M.J.; Liao, Y.; Thiyagarajan, M.; O’Donnell, J.; Christensen, D.J.; Nicholson, C.; Iliff, J.J.; et al. Sleep drives metabolite clearance from the adult brain. Science 2013, 342, 373–377. [Google Scholar] [CrossRef]
- Zada, D.; Bronshtein, I.; Lerer-Goldshtein, T.; Garini, Y.; Appelbaum, L. Sleep increases chromosome dynamics to enable reduction of accumulating DNA damage in single neurons. Nat. Commun. 2019, 10, 895. [Google Scholar] [CrossRef]
- Laposky, A.D.; Bass, J.; Kohsaka, A.; Turek, F.W. Sleep and circadian rhythms: Key components in the regulation of energy metabolism. FEBS Lett. 2008, 582, 142–151. [Google Scholar] [CrossRef]
- Wisor, J.P.; Pasumarthi, R.K.; Gerashchenko, D.; Thompson, C.L.; Pathak, S.; Sancar, A.; Franken, P.; Lein, E.S.; Kilduff, T.S. Sleep deprivation effects on circadian clock gene expression in the cerebral cortex parallel electroencephalographic differences among mouse strains. J. Neurosci. 2008, 28, 7193–7201. [Google Scholar] [CrossRef]
- Borbély, A.A. A two process model of sleep regulation. Hum. Neurobiol. 1982, 1, 195–204. [Google Scholar]
- Franken, P.; Dijk, D.J. Circadian clock genes and sleep homeostasis. Eur. J. Neurosci. 2009, 29, 1820–1829. [Google Scholar] [CrossRef]
- Franken, P. A role for clock genes in sleep homeostasis. Curr. Opin. Neurobiol. 2013, 23, 864–872. [Google Scholar] [CrossRef]
- Shaw, P.J.; Tononi, G.; Greenspan, R.J.; Robinson, D.F. Stress response genes protect against lethal effects of sleep deprivation in Drosophila. Nature 2002, 417, 287. [Google Scholar] [CrossRef]
- Aran, A.; Einen, M.; Lin, L.; Plazzi, G.; Nishino, S.; Mignot, E. Clinical and therapeutic aspects of childhood narcolepsy-cataplexy: A retrospective study of 51 children. Sleep 2010, 33, 1457–1464. [Google Scholar] [CrossRef]
- Peter-Derex, L.; Yammine, P.; Bastuji, H.; Croisile, B. Sleep and Alzheimer’s disease. Sleep Med. Rev. 2015, 19, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Peek, C.B.; Affinati, A.H.; Ramsey, K.M.; Kuo, H.Y.; Yu, W.; Sena, L.A.; Ilkayeva, O.; Marcheva, B.; Kobayashi, Y.; Omura, C.; et al. Circadian clock NAD+ cycle drives mitochondrial oxidative metabolism in mice. Science 2013, 342, 1243417. [Google Scholar] [CrossRef] [PubMed]
- Pekovic-Vaughan, V.; Gibbs, J.; Yoshitane, H.; Yang, N.; Pathiranage, D.; Guo, B.; Sagami, A.; Taguchi, K.; Bechtold, D.; Loudon, A.; et al. The circadian clock regulates rhythmic activation of the NRF2/glutathione-mediated antioxidant defense pathway to modulate pulmonary fibrosis. Genes Dev. 2014, 28, 548–560. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grimaldi, B.; Bellet, M.M.; Katada, S.; Astarita, G.; Hirayama, J.; Amin, R.H.; Granneman, J.G.; Piomelli, D.; Leff, T.; Sassone-Corsi, P. PER2 controls lipid metabolism by direct regulation of PPARγ. Cell Metab. 2010, 12, 509–520. [Google Scholar] [CrossRef] [PubMed]
- Buxton, O.M.; Cain, S.W.; O’Connor, S.P.; Porter, J.H.; Duffy, J.F.; Wang, W.; Czeisler, C.A.; Shea, S.A. Adverse metabolic consequences in humans of prolonged sleep restriction combined with circadian disruption. Sci. Transl. Med. 2012, 4, 129ra43. [Google Scholar] [CrossRef] [PubMed]
- McHill, A.W.; Melanson, E.L.; Higgins, J.; Connick, E.; Moehlman, T.M.; Stothard, E.R.; Wright, K.P. Impact of circadian misalignment on energy metabolism during simulated nightshift work. Proc. Natl. Acad. Sci. USA 2014, 111, 17302–17307. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arble, D.M.; Bass, J.; Laposky, A.D.; Vitaterna, M.H.; Turek, F.W. Circadian timing of food intake contributes to weight gain. Obesity 2009, 17, 2100–2102. [Google Scholar] [CrossRef] [PubMed]
- Kohsaka, A.; Laposky, A.D.; Ramsey, K.M.; Estrada, C.; Joshu, C.; Kobayashi, Y.; Turek, F.W.; Bass, J. High-fat diet disrupts behavioral and molecular circadian rhythms in mice. Cell Metab. 2007, 6, 414–421. [Google Scholar] [CrossRef]
- Hatori, M.; Vollmers, C.; Zarrinpar, A.; DiTacchio, L.; Bushong, E.A.; Gill, S.; Leblanc, M.; Chaix, A.; Joens, M.; Fitzpatrick, J.A.; et al. Time-restricted feeding without reducing caloric intake prevents metabolic diseases in mice fed a high-fat diet. Cell Metab. 2012, 15, 848–860. [Google Scholar] [CrossRef]
- Jakubowicz, D.; Froy, O.; Wainstein, J.; Boaz, M. Meal timing and composition influence ghrelin levels, appetite scores and weight loss maintenance in overweight and obese adults. Steroids 2012, 77, 323–331. [Google Scholar] [CrossRef]
- Marcheva, B.; Ramsey, K.M.; Buhr, E.D.; Kobayashi, Y.; Su, H.; Ko, C.H.; Ivanova, G.; Omura, C.; Mo, S.; Vitaterna, M.H.; et al. Disruption of the clock components CLOCK and BMAL1 leads to hypoinsulinaemia and diabetes. Nature 2010, 466, 627. [Google Scholar] [CrossRef]
- Perelis, M.; Marcheva, B.; Ramsey, K.M.; Schipma, M.J.; Hutchison, A.L.; Taguchi, A.; Peek, C.B.; Hong, H.; Huang, W.; Omura, C.; et al. Pancreatic β cell enhancers regulate rhythmic transcription of genes controlling insulin secretion. Science 2015, 350, aac4250. [Google Scholar] [CrossRef]
- Gallou-Kabani, C.; Vigé, A.; Junien, C. Lifelong circadian and epigenetic drifts in metabolic syndrome. Epigenetics 2007, 2, 137–146. [Google Scholar] [CrossRef]
- Spiegel, K.; Tasali, E.; Leproult, R.; Van Cauter, E. Effects of poor and short sleep on glucose metabolism and obesity risk. Nat. Rev. Endocrinol. 2009, 5, 253. [Google Scholar] [CrossRef]
- McIntyre, R.S. Managing weight gain in patients with severe mental illness. J. Clin. Psychiatry 2009, 70, e23. [Google Scholar] [CrossRef]
- Frank, E.; Swartz, H.A.; Kupfer, D.J. Interpersonal and social rhythm therapy: Managing the chaos of bipolar disorder. Biol. Psychiatry 2000, 48, 593–604. [Google Scholar] [CrossRef]
- Hlastala, S.A.; Frank, E. Adapting interpersonal and social rhythm therapy to the developmental needs of adolescents with bipolar disorder. Dev. Psychopathol. 2006, 18, 1267–1288. [Google Scholar] [CrossRef]
- Iwamoto, A.; Kawai, M.; Furuse, M.; Yasuo, S. Effects of chronic jet lag on the central and peripheral circadian clocks in CBA/N mice. Chronobiol. Int. 2014, 31, 189–198. [Google Scholar] [CrossRef]
- Ozturk, N.; Lee, J.H.; Gaddameedhi, S.; Sancar, A. Loss of cryptochrome reduces cancer risk in p53 mutant mice. Proc. Natl. Acad. Sci. USA 2009, 106, 2841–2846. [Google Scholar] [CrossRef]
- Thosar, S.S.; Butler, M.P.; Shea, S.A. Role of the circadian system in cardiovascular disease. J. Clin. Investig. 2018, 128, 2157–2167. [Google Scholar] [CrossRef] [Green Version]
- Curtis, A.M.; Cheng, Y.; Kapoor, S.; Reilly, D.; Price, T.S.; FitzGerald, G.A. Circadian variation of blood pressure and the vascular response to asynchronous stress. Proc. Natl. Acad. Sci. USA 2007, 104, 3450–3455. [Google Scholar] [CrossRef] [Green Version]
- Reinke, H.; Asher, G. Crosstalk between metabolism and circadian clocks. Nat. Rev. Mol. Cell Biol. 2019, 20, 227–241. [Google Scholar] [CrossRef]
- Yamada, Y.; Forger, D. Multiscale complexity in the mammalian circadian clock. Curr. Opin. Genet. Dev. 2010, 20, 626–633. [Google Scholar] [CrossRef] [Green Version]
- Susaki, E.A.; Ukai, H.; Ueda, H.R. Next-generation mammalian genetics toward organism-level systems biology. NPJ Syst. Biol. Appl. 2017, 3, 15. [Google Scholar] [CrossRef]
- Ballesta, A.; Innominato, P.F.; Dallmann, R.; Rand, D.A.; Levi, F.A. Systems chronotherapeutics. Pharmacol. Rev. 2017, 69, 161–199. [Google Scholar] [CrossRef]
- Lieverse, R.; Van Someren, E.J.; Nielen, M.M.; Uitdehaag, B.M.; Smit, J.H.; Hoogendijk, W.J. Bright Light Treatment in Elderly patients with nonseasonal Major Depressive Disorder. Arch. Gen. Psychiatry 2011, 68, 61–70. [Google Scholar] [CrossRef]
- Weitzman, E.D.; Czeisler, C.A.; Coleman, R.M.; Spielman, A.J.; Zimmerman, J.C.; Dement, W.; Pollak, C.P. Delayed sleep phase syndrome: A chronobiological disorder with sleep-onset insomnia. Arch. Gen. Psychiatry 1981, 38, 737–746. [Google Scholar] [CrossRef]
- Barion, A.; Zee, P.C. A clinical approach to circadian rhythm sleep disorders. Sleep Med. 2007, 8, 566–577. [Google Scholar] [CrossRef] [Green Version]
- Rosenthal, N.E.; Joseph-Vanderpool, J.R.; Levendosky, A.A.; Johnston, S.H.; Allen, R.; Kelly, K.A.; Souetre, E.; Schultz, P.M.; Starz, K.E. Phase-shifting effects of bright morning light as treatment for delayed sleep phase syndrome. Sleep 1990, 13, 354–361. [Google Scholar]
- Chesson, A.L.; Wise, M.; Davila, D.; Johnson, S.; Littner, M.; Anderson, W.M.; Hartse, K.; Rafecas, J. Practice parameters for the treatment of restless legs syndrome and periodic limb movement disorder. Sleep 1999, 22, 961–968. [Google Scholar] [CrossRef]
- Levi, F.; Zidani, R.; Misset, J.L. Randomised multicentre trial of chronotherapy with oxaliplatin, fluorouracil, and folinic acid in metastatic colorectal cancer. Lancet 1997, 350, 681–686. [Google Scholar] [CrossRef]
- Lévi, F.; Schibler, U. Circadian rhythms: Mechanisms and therapeutic implications. Annu. Rev. Pharmacol. Toxicol. 2007, 47, 593–628. [Google Scholar] [CrossRef]
- Kirschbaum, I.; Straub, J.; Gest, S.; Holtmann, M.; Legenbauer, T. Short-term effects of wake-and bright light therapy on sleep in depressed youth. Chronobiol. Int. 2018, 35, 101–110. [Google Scholar] [CrossRef]
- Suzuki, M.; Dallaspezia, S.; Locatelli, C.; Uchiyama, M.; Colombo, C.; Benedetti, F. Does early response predict subsequent remission in bipolar depression treated with repeated sleep deprivation combined with light therapy and lithium? J. Affect. Disord. 2018, 229, 371–376. [Google Scholar] [CrossRef]
- Bais, B.; Kamperman, A.M.; van der Zwaag, M.D.; Dieleman, G.C.; van der Vliet, H.W.H.; Bijma, H.H.; Lieverse, R.; Hoogendijk, W.J.; Lambregtse-van den Berg, M.P. Bright light therapy in pregnant women with major depressive disorder: Study protocol for a randomized, double-blind, controlled clinical trial. BMC Psychiatry 2016, 16, 381. [Google Scholar] [CrossRef]
- Calhoun, S.L.; Fernandez-Mendoza, J.; Vgontzas, A.N.; Liao, D.; Bixler, E.O. Prevalence of insomnia symptoms in a general population sample of young children and preadolescents: Gender effects. Sleep Med. 2014, 15, 91–95. [Google Scholar] [CrossRef]
- Stranges, S.; Tigbe, W.; Gómez-Olivé, F.X.; Thorogood, M.; Kandala, N.B. Sleep problems: An emerging global epidemic? Findings from the INDEPTH WHO-SAGE study among more than 40,000 older adults from 8 countries across Africa and Asia. Sleep 2012, 35, 1173–1181. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chowdhury, D.; Wang, C.; Lu, A.-P.; Zhu, H.-L. Understanding Quantitative Circadian Regulations Are Crucial Towards Advancing Chronotherapy. Cells 2019, 8, 883. https://doi.org/10.3390/cells8080883
Chowdhury D, Wang C, Lu A-P, Zhu H-L. Understanding Quantitative Circadian Regulations Are Crucial Towards Advancing Chronotherapy. Cells. 2019; 8(8):883. https://doi.org/10.3390/cells8080883
Chicago/Turabian StyleChowdhury, Debajyoti, Chao Wang, Ai-Ping Lu, and Hai-Long Zhu. 2019. "Understanding Quantitative Circadian Regulations Are Crucial Towards Advancing Chronotherapy" Cells 8, no. 8: 883. https://doi.org/10.3390/cells8080883




