Construction and Comprehensive Analysis of a Molecular Association Network via lncRNA–miRNA–Disease–Drug–Protein Graph
Abstract
:1. Introduction
2. Materials and Methods
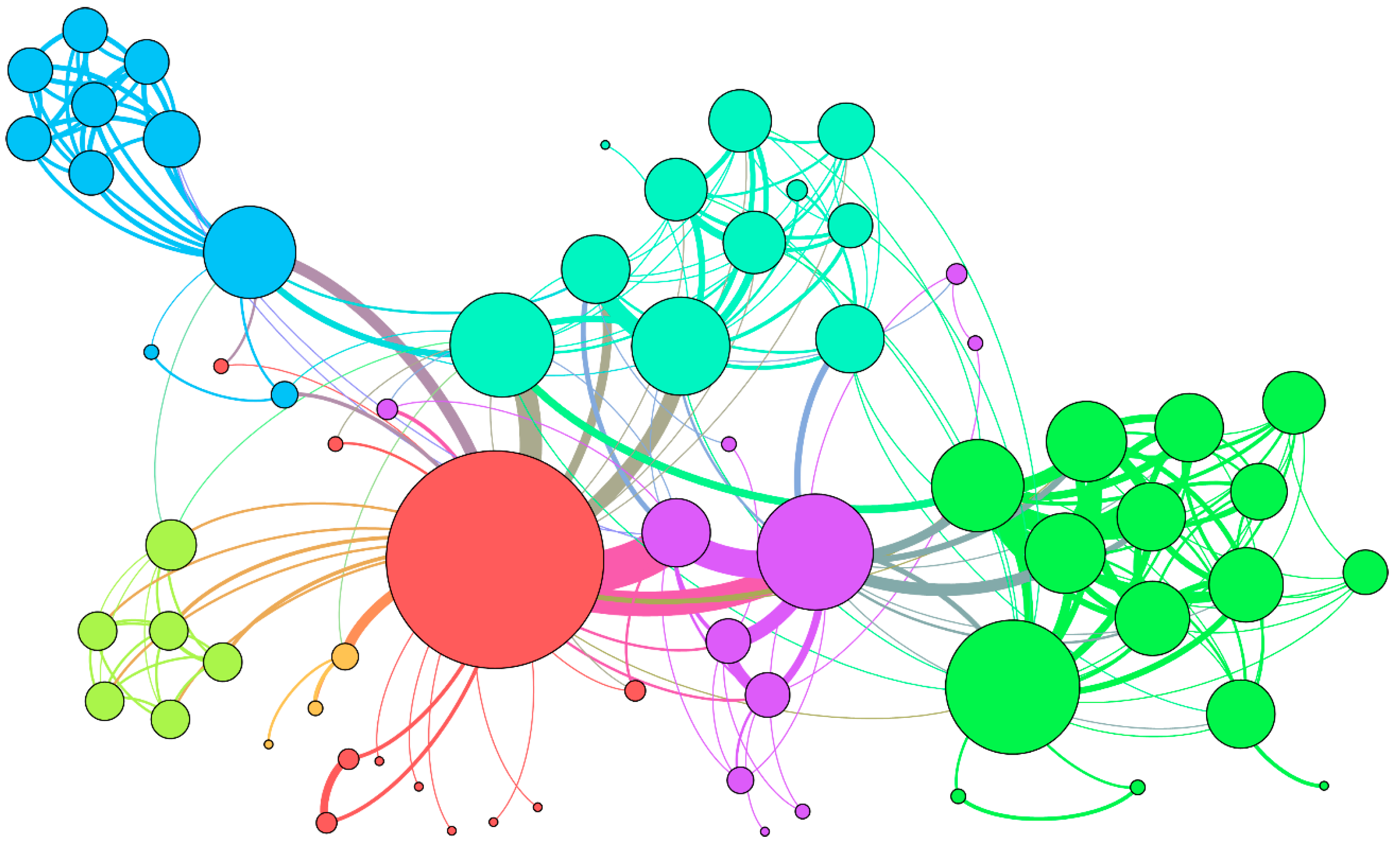
2.1. Construction of the Molecular Association Network
2.2. NcRNA and Protein Sequence
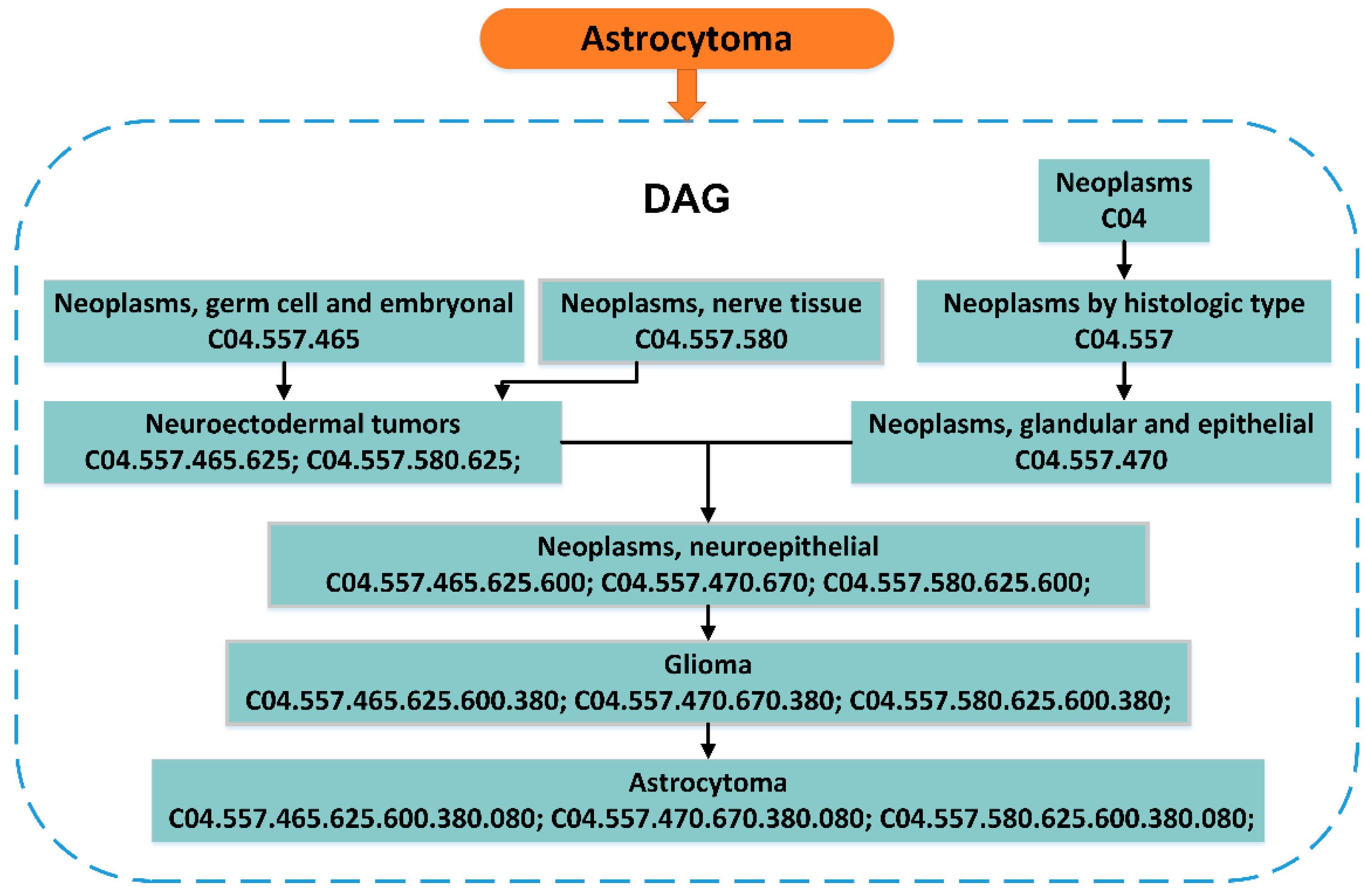
2.3. Disease MeSH Descriptors and Directed Acyclic Graph
2.4. Drug Molecular Fingerprint
2.5. Stacked Autoencoder
2.6. Node Representation
3. Results and Discussion
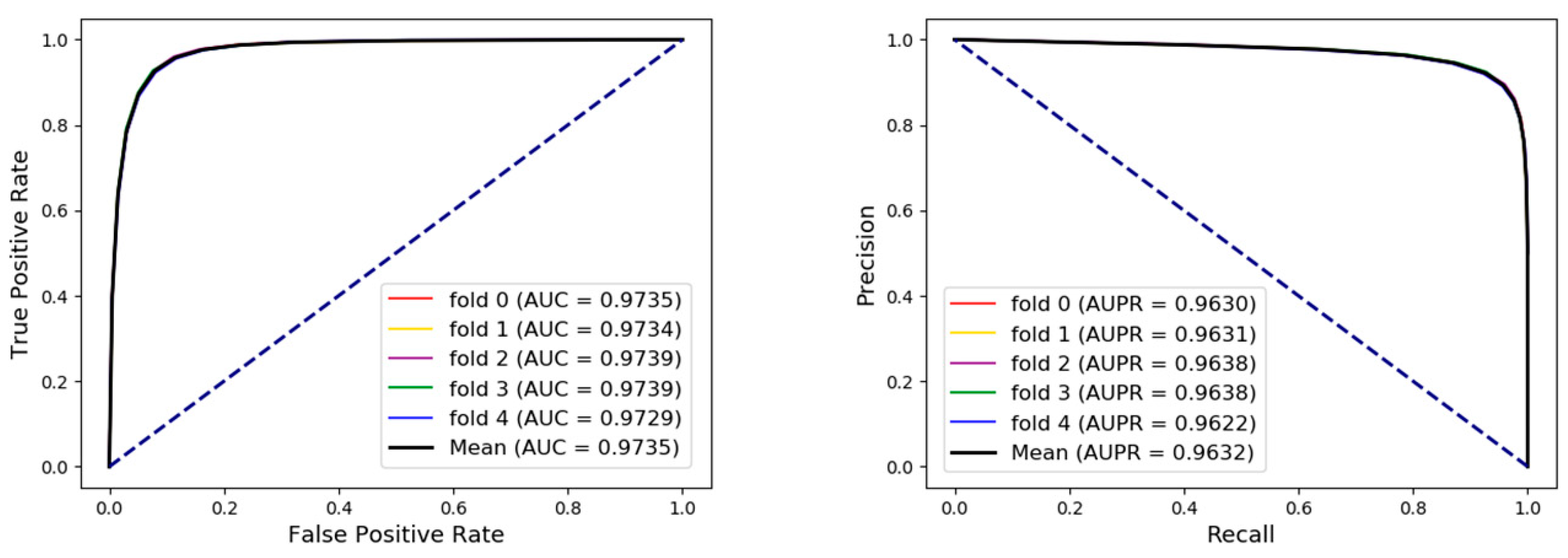
3.1. Evaluate the Five-Fold Cross Validation Performance of Our Method
3.2. Comparison of Different Feature Extraction Methods
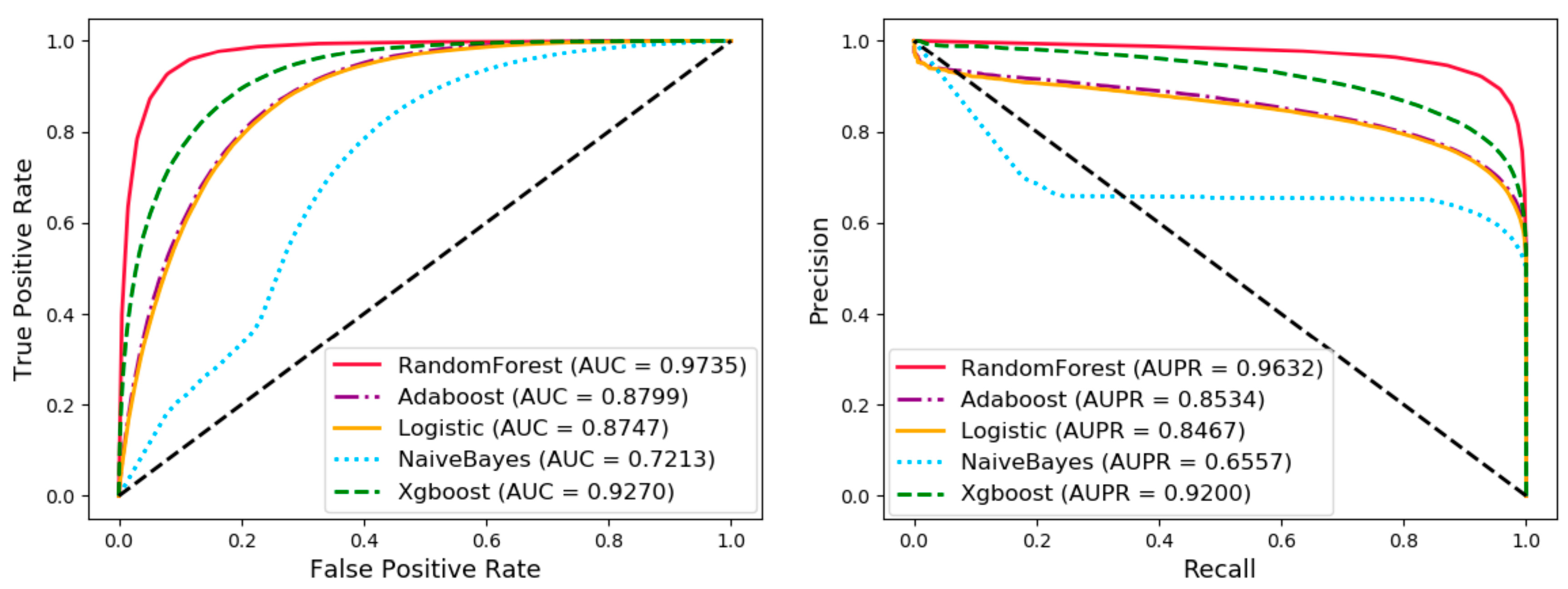
3.3. Comparison of Different Classifiers
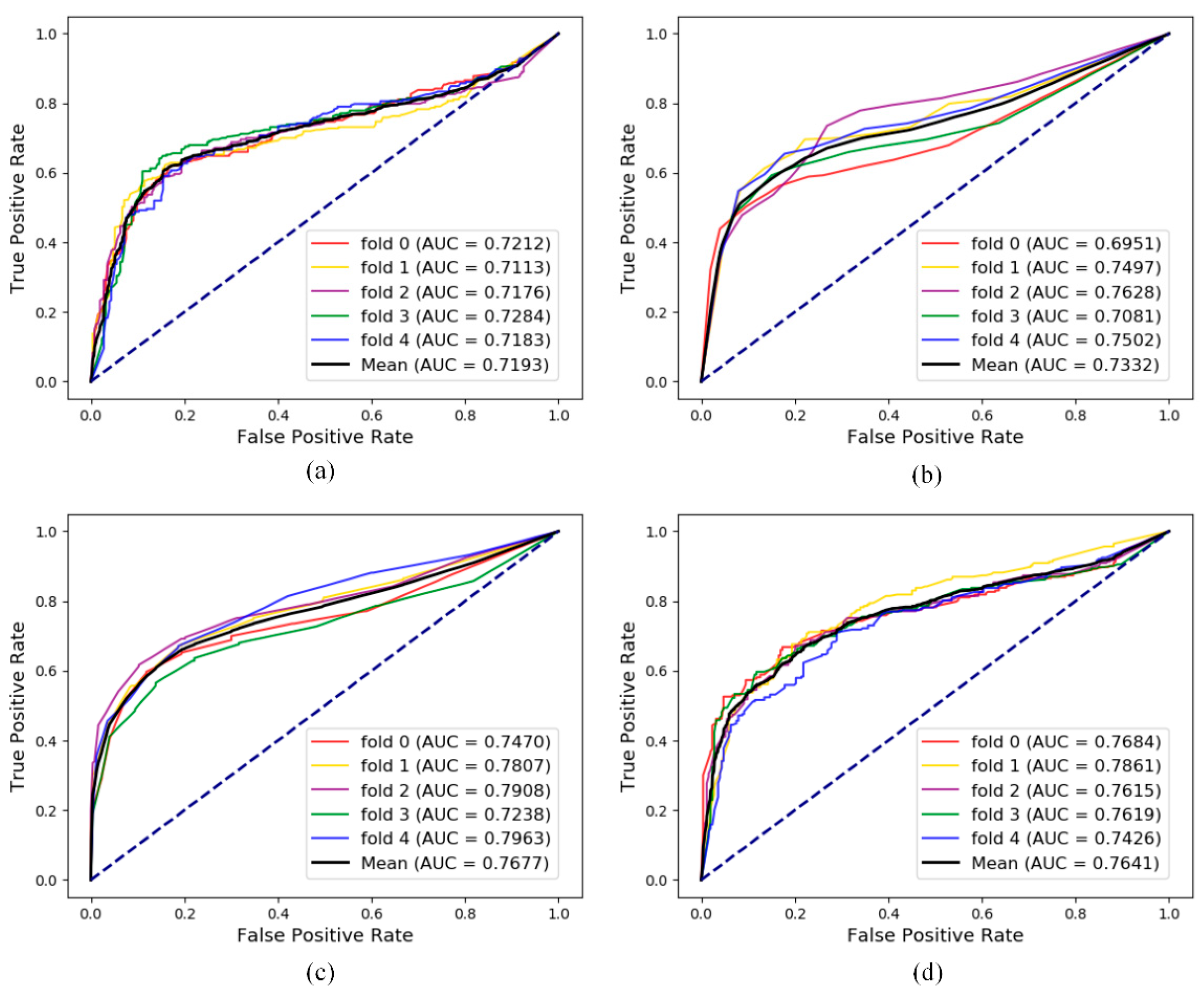
3.4. Additional Comparison Experiment for lncRNA-Disease Association Prediction
3.5. Analysis Based on a Specific miRNA, lncRNA, and Protein
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Ambros, V. MicroRNA pathways in flies and worms: Growth, death, fat, stress and timing. Cell 2003, 113, 673–676. [Google Scholar] [CrossRef]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Bonetta, L. Protein–protein interactions: Interactome under construction. Nature 2010, 468, 851. [Google Scholar] [CrossRef] [PubMed]
- Skrabanek, L.; Saini, H.K.; Bader, G.D.; Enright, A.J. Computational prediction of protein–protein interactions. Mol. Biotechnol. 2008, 38, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Ashburn, T.T.; Thor, K.B. Drug repositioning: Identifying and developing new uses for existing drugs. Nat. Rev. Drug Discov. 2004, 3, 673. [Google Scholar] [CrossRef] [PubMed]
- Collins, S.R.; Kemmeren, P.; Zhao, X.C.; Greenblatt, J.F.; Spencer, F.; Holstege, F.C.; Weissman, J.S.; Krogan, N.J. Toward a comprehensive atlas of the physical interactome of Saccharomyces cerevisiae. Mol. Cell. Proteom. 2007, 6, 439–450. [Google Scholar] [CrossRef] [PubMed]
- Yan, B.; Wang, Z.H.; Guo, J.T. The research strategies for probing the function of long noncoding RNAs. Genomics 2012, 99, 76–80. [Google Scholar] [CrossRef] [PubMed]
- Fields, S.; Song, O.K. A novel genetic system to detect protein–protein interactions. Nature 1989, 340, 245. [Google Scholar] [CrossRef]
- Chen, G.; Wang, Z.; Wang, D.; Qiu, C.; Liu, M.; Chen, X.; Zhang, Q.; Yan, G.; Cui, Q. LncRNADisease: A database for long-non-coding RNA-associated diseases. Nucleic Acids Res. 2012, 41, D983–D986. [Google Scholar] [CrossRef]
- Huang, Z.; Shi, J.; Gao, Y.; Cui, C.; Zhang, S.; Li, J.; Zhou, Y.; Cui, Q. HMDD v3. 0: A database for experimentally supported human microRNA–disease associations. Nucleic Acids Res. 2018, 47, D1013–D1017. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P. The STRING database in 2017: Quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2016. [Google Scholar] [CrossRef] [PubMed]
- Yi, H.C.; You, Z.H.; Zhou, X.; Cheng, L.; Li, X.; Jiang, T.H.; Chen, Z.H. ACP-DL: A Deep Learning Long Short-Term Memory Model to Predict Anticancer Peptides Using High Efficiency Feature Representation. Mol. Ther. Nucleic Acids 2019, 17, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; You, Z.H.; Yang, S.; Li, X.; Jiang, T.H.; Zhou, X. A High Efficient Biological Language Model for Predicting Protein–Protein Interactions. Cells 2019, 8, 122. [Google Scholar] [CrossRef] [PubMed]
- Yi, H.C.; You, Z.H.; Huang, D.S.; Li, X.; Jiang, T.H.; Li, L.P. A deep learning framework for robust and accurate prediction of ncRNA-protein interactions using evolutionary information. Mol. Ther. Nucleic Acids 2018, 11, 337–344. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; You, Z.H.; Chen, X.; Yan, X.; Liu, G.; Zhang, W. Rfdt: A rotation forest-based predictor for predicting drug-target interactions using drug structure and protein sequence information. Curr. Protein Pept. Sci. 2018, 19, 445–454. [Google Scholar] [CrossRef] [PubMed]
- You, Z.H.; Lei, Y.K.; Zhu, L.; Xia, J.; Wang, B. Prediction of protein-protein interactions from amino acid sequences with ensemble extreme learning machines and principal component analysis. BMC Bioinform. 2013, 14, S10. [Google Scholar] [CrossRef]
- Chen, X.; Xie, D.; Wang, L.; Zhao, Q.; You, Z.H.; Liu, H. BNPMDA: Bipartite network projection for MiRNA—Disease association prediction. Bioinformatics 2018, 34, 3178–3186. [Google Scholar] [CrossRef]
- Li, J.Q.; Rong, Z.H.; Chen, X.; Yan, G.Y.; You, Z H. MCMDA: Matrix completion for MiRNA-disease association prediction. Oncotarget 2017, 8, 21187. [Google Scholar]
- Chen, X.; Zhang, D.H.; You, Z.H. A heterogeneous label propagation approach to explore the potential associations between miRNA and disease. J. Transl. Med. 2018, 16, 348. [Google Scholar] [CrossRef]
- Peng, J.; Hui, W.; Li, Q.; Chen, B.; Jiang, Q.; Wei, Z.; Shang, X. A learning-based framework for miRNA-disease association prediction using neural networks. bioRxiv 2018. [Google Scholar] [CrossRef]
- Hrdlickova, B.; de Almeida, R.C.; Borek, Z.; Withoff, S. Genetic variation in the non-coding genome: Involvement of micro-RNAs and long non-coding RNAs in disease. Biochim. Biophys. Acta Mol. Basis Dis. 2014, 1842, 1910–1922. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barabási, A.L.; Oltvai, Z.N. Network biology: Understanding the cell’s functional organization. Nat. Rev. Genet. 2004, 5, 101–113. [Google Scholar] [CrossRef] [PubMed]
- Miao, Y.R.; Liu, W.; Zhang, Q.; Guo, A.Y. lncRNASNP2: An updated database of functional SNPs and mutations in human and mouse lncRNAs. Nucleic Acids Res. 2017, 46, D276–D280. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.H.; Shrestha, S.; Yang, C.D.; Chang, N.W.; Lin, Y.L.; Liao, K.W.; Huang, W.C.; Sun, T.H.; Tu, S.J.; Lee, W.H. miRTarBase update 2018: A resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2017, 46, D296–D302. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Wang, P.; Tian, R.; Wang, S.; Guo, Q.; Luo, M.; Zhou, W.; Liu, G.; Jiang, H.; Jiang, Q. LncRNA2Target v2.0: A comprehensive database for target genes of lncRNAs in human and mouse. Nucleic Acids Res. 2018, 47, D140–D144. [Google Scholar] [CrossRef] [PubMed]
- Piñero, J.; Bravo, À.; Queralt-Rosinach, N.; Gutiérrez-Sacristán, A.; Deu-Pons, J.; Centeno, E.; García-García, J.; Sanz, F.; Furlong, L.I. DisGeNET: A comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 2016. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2017, 46, D1074–D1082. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.P.; Grondin, C.J.; Johnson, R.J.; Sciaky, D.; McMorran, R.; Wiegers, J.; Wiegers, T.C.; Mattingly, C.J. The comparative toxicogenomics database: Update 2019. Nucleic Acids Res. 2018, 47, D948–D954. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Birgaoanu, M.; Griffiths-Jones, S. miRBase: From microRNA sequences to function. Nucleic Acids Res. 2018, 47, D155–D162. [Google Scholar] [CrossRef] [PubMed]
- Fang, S.; Zhang, L.; Guo, J.; Niu, Y.; Wu, Y.; Li, H.; Zhao, L.; Li, X.; Teng, X.; Sun, X. NONCODEV5: A comprehensive annotation database for long non-coding RNAs. Nucleic Acids Res. 2017, 46, D308–D314. [Google Scholar] [CrossRef]
- Shen, J.; Zhang, J.; Luo, X.; Zhu, W.; Yu, K.; Chen, K.; Li, Y.; Jiang, H. Predicting protein–protein interactions based only on sequences information. Proc. Natl. Acad. Sci. USA 2007, 104, 4337–4341. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Wang, J.; Lu, M.; Song, F.; Cui, Q. Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 2010, 26, 1644–1650. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, J.; Qu, M.; Wang, M.; Zhang, M.; Yan, J.; Mei, Q. Line: Large-scale information network embedding. In Proceedings of the 24th International Conference on World Wide Web, Florence, Italy, 18–22 May 2015; International World Wide Web Conferences Steering Committee: Geneva, Switzerland, 2015; pp. 1067–1077. [Google Scholar]
- van Laarhoven, T.; Nabuurs, S.B.; Marchiori, E. Gaussian interaction profile kernels for predicting drug–target interaction. Bioinformatics 2011, 27, 3036–3043. [Google Scholar] [CrossRef] [PubMed]
- Chen, X. Predicting lncRNA-disease associations and constructing lncRNA functional similarity network based on the information of miRNA. Sci. Rep. 2015, 5, 13186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cui, T.; Zhang, L.; Huang, Y.; Yi, Y.; Tan, P.; Zhao, Y.; Hu, Y.; Xu, L.; Li, E.; Wang, D. MNDR v2.0: An updated resource of ncRNA–disease associations in mammals. Nucleic Acids Res. 2017, 46, D371–D374. [Google Scholar] [CrossRef] [PubMed]



| Relationship Type | Database | Number of Associations |
|---|---|---|
| miRNA–lncRNA | lncRNASNP2 [23] | 8374 |
| miRNA–disease | HMDD [10] | 16,427 |
| miRNA–protein | miRTarBase [24] | 4944 |
| lncRNA–disease | LncRNADisease [9] lncRNASNP2 [23] | 1264 |
| lncRNA–protein Protein–disease Drug–protein Drug–disease Protein–protein | LncRNA2Target [25] DisGeNET [26] DrugBank [27] CTD [28] STRING [11] | 690 25,087 11,107 18,416 19,237 |
| Total | N/A | 105,546 |
| Node | Number of Nodes |
|---|---|
| Disease | 2062 |
| LncRNA | 769 |
| MiRNA | 1023 |
| Protein | 1649 |
| Drug | 1025 |
| Total | 6528 |
| Fold | Acc. (%) | Sen. (%) | Spec. (%) | Prec. (%) | MCC (%) | AUC (%) |
|---|---|---|---|---|---|---|
| 0 | 92.25 | 92.68 | 91.82 | 91.89 | 84.51 | 97.35 |
| 1 | 92.43 | 92.52 | 92.35 | 92.36 | 84.87 | 97.34 |
| 2 | 92.49 | 92.84 | 92.13 | 92.19 | 84.98 | 97.39 |
| 3 | 92.58 | 92.75 | 92.42 | 92.44 | 85.16 | 97.39 |
| 4 | 92.13 | 92.28 | 91.98 | 92.01 | 84.26 | 97.29 |
| Average | 92.38± 0.18 | 92.61± 0.22 | 92.14± 0.25 | 92.18± 0.23 | 84.76± 0.37 | 97.35± 0.04 |
| Feature | Acc. (%) | Sen. (%) | Spec. (%) | Prec. (%) | MCC (%) | AUC (%) |
|---|---|---|---|---|---|---|
| Attribute | 88.62 ± 0.14 | 91.48 ± 0.13 | 85.76 ± 0.2 | 86.53 ± 0.17 | 77.37 ± 0.28 | 94.47 ± 0.11 |
| Behavior | 90.7 ± 0.14 | 88.84 ± 0.15 | 92.56 ± 0.19 | 92.27 ± 0.19 | 81.45 ± 0.29 | 96.26 ± 0.05 |
| Both | 92.38± 0.18 | 92.61± 0.22 | 92.14± 0.25 | 92.18± 0.23 | 84.76± 0.37 | 97.35± 0.04 |
| Classifier | Acc. (%) | Sen. (%) | Spec. (%) | Prec. (%) | MCC (%) | AUC (%) |
|---|---|---|---|---|---|---|
| Adaboost | 80.03 ± 0.29 | 80.91 ± 0.3 | 79.14 ± 0.43 | 79.51 ± 0.36 | 60.07 ± 0.58 | 87.99 ± 0.28 |
| Logistic | 79.92 ± 0.29 | 82.78 ± 0.29 | 77.06 ± 0.49 | 78.3 ± 0.37 | 59.94 ± 0.57 | 87.47 ± 0.26 |
| Naive Bayes | 55.93 ± 0.15 | 24.83 ± 0.24 | 87.04 ± 0.32 | 65.7 ± 0.5 | 15.15 ± 0.41 | 72.13 ± 0.34 |
| XGBoost | 84.37 ± 1.3 | 82.89 ± 2.96 | 85.85 ± 0.56 | 85.42 ± 0.37 | 68.8 ± 2.58 | 92.7 ± 0.66 |
| Random Forest | 92.38± 0.18 | 92.61± 0.22 | 92.14± 0.25 | 92.18± 0.23 | 84.76± 0.37 | 97.35± 0.04 |
| Number | Disease Name | Probability | Evidence |
|---|---|---|---|
| 1 | Melanoma | 0.85 | LncRNADisease |
| 2 | Cervical cancer | 0.85 | LncRNADisease |
| 3 | Rheumatoid arthritis | 0.85 | MNDR 2.0 |
| 4 | Hepatocellular carcinoma | 0.85 | MNDR 2.0, LncRNADisease |
| 5 | Myelodysplastic syndrome | 0.846551724 | Unconfirmed |
| 6 | Schizophrenia | 0.842857143 | Unconfirmed |
| 7 | Chronic lymphocytic leukemia | 0.840909091 | Unconfirmed |
| 8 | Diffuse large b-cell lymphoma | 0.837460815 | Unconfirmed |
| 9 | Cardiac hypertrophy | 0.833766234 | Unconfirmed |
| 10 | Digeorge syndrome | 0.8 | Unconfirmed |
| 11 | Multiple sclerosis | 0.8 | Unconfirmed |
| 12 | Acute promyelocytic leukemia | 0.786551724 | Unconfirmed |
| 13 | Autism spectrum disorder | 0.75 | Unconfirmed |
| 14 | Colorectal cancer | 0.75 | MNDR 2.0, LncRNADisease |
| 15 | Osteosarcoma | 0.75 | MNDR 2.0, LncRNADisease |
| 16 | Squamous cell carcinoma | 0.75 | MNDR 2.0, LncRNADisease |
| 17 | Atherosclerosis | 0.75 | LncRNADisease |
| 18 | Glioblastoma | 0.75 | LncRNADisease |
| 19 | Pituitary adenoma | 0.75 | LncRNADisease |
| 20 | Pre-eclampsia | 0.75 | LncRNADisease |
| LncRNA–miRNA Pairs | Associated Proteins Respectively | PPI Network Edges |
|---|---|---|
| NONHSAT007662.2/hsa-miR-205-5p | 70/27 | 1066 |
| NONHSAT017460.2/hsa-miR-148a-3p | 74/28 | 877 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, Z.-H.; Yi, H.-C.; You, Z.-H. Construction and Comprehensive Analysis of a Molecular Association Network via lncRNA–miRNA–Disease–Drug–Protein Graph. Cells 2019, 8, 866. https://doi.org/10.3390/cells8080866
Guo Z-H, Yi H-C, You Z-H. Construction and Comprehensive Analysis of a Molecular Association Network via lncRNA–miRNA–Disease–Drug–Protein Graph. Cells. 2019; 8(8):866. https://doi.org/10.3390/cells8080866
Chicago/Turabian StyleGuo, Zhen-Hao, Hai-Cheng Yi, and Zhu-Hong You. 2019. "Construction and Comprehensive Analysis of a Molecular Association Network via lncRNA–miRNA–Disease–Drug–Protein Graph" Cells 8, no. 8: 866. https://doi.org/10.3390/cells8080866






