The Transcription Factor Elf3 Is Essential for a Successful Mesenchymal to Epithelial Transition
Abstract
1. Introduction
2. Materials and Methods
2.1. Microarray Data Analysis and Correlation Study
2.2. Cell Culture and Induction of EMT and MET
2.3. siRNA Knockdown and mRNA Expression Analysis
2.4. Luciferase Reporter Assays
2.5. Expression and Reporter Vectors
2.6. Chromatin Immunoprecipitation (ChIP)
2.7. Immunofluorescence Labeling and Confocal Microscopy
2.8. SA-β-GAL Assay
2.9. RT-PCR
2.10. Fluorescence-Activated Cell Sorting (FACS)
2.11. Western Blot
2.12. Wound Healing Assay
2.13. Statistical Analysis
3. Results
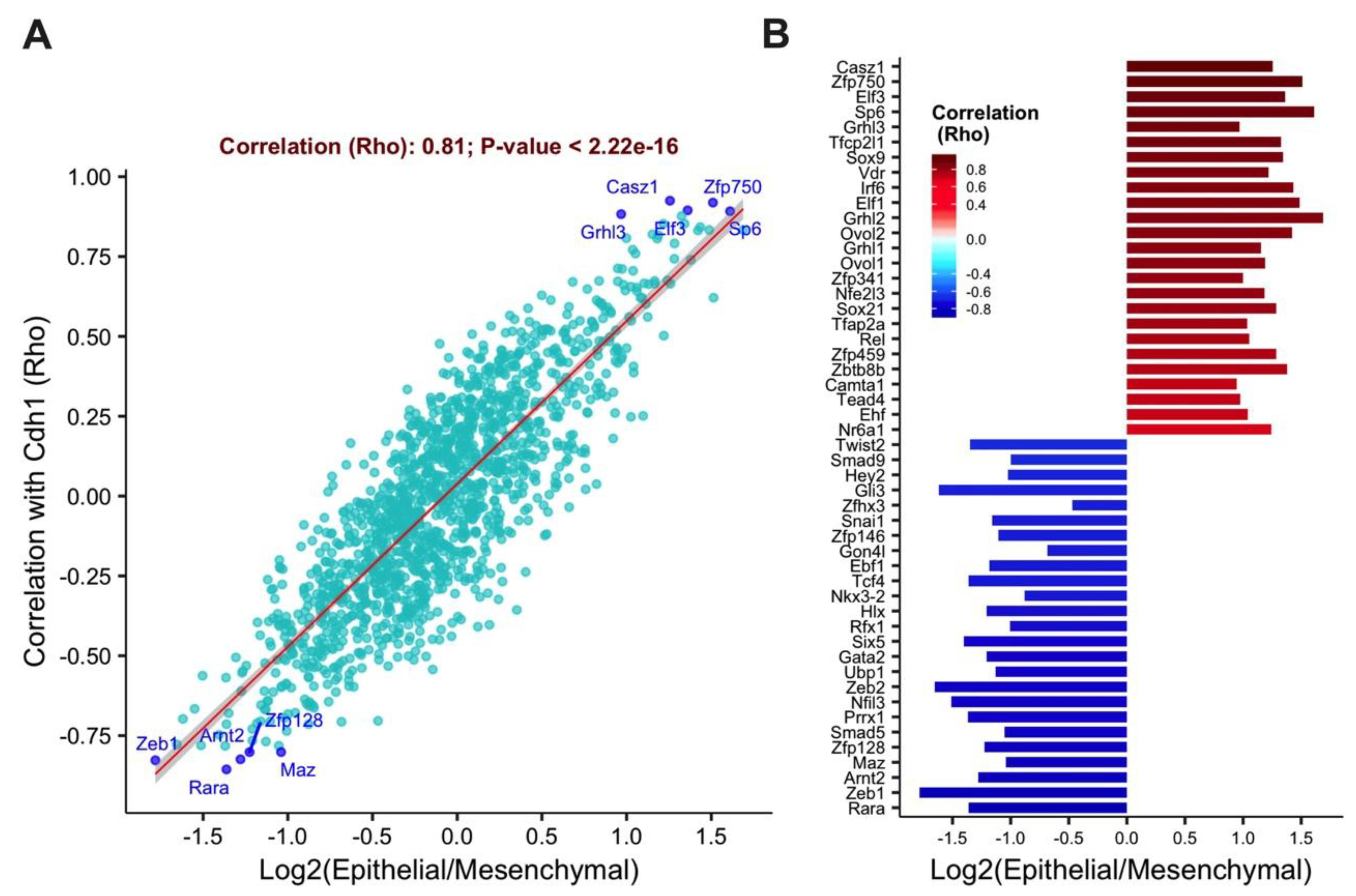
3.1. The Transcription Factors Elf3 and Grhl3 Are Highly Correlated with the Epithelial State
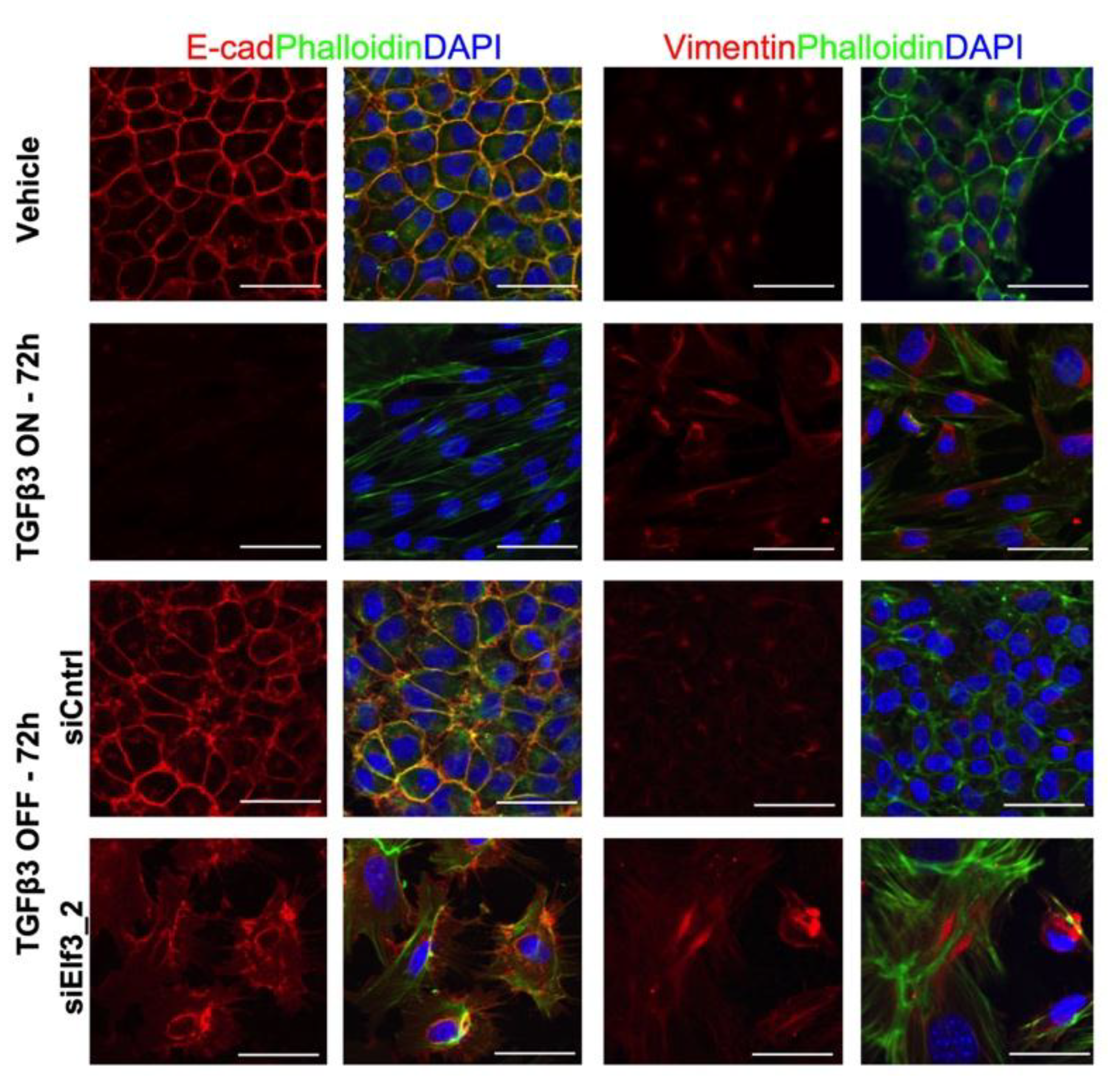
3.2. Elf3 Is Essential for MET
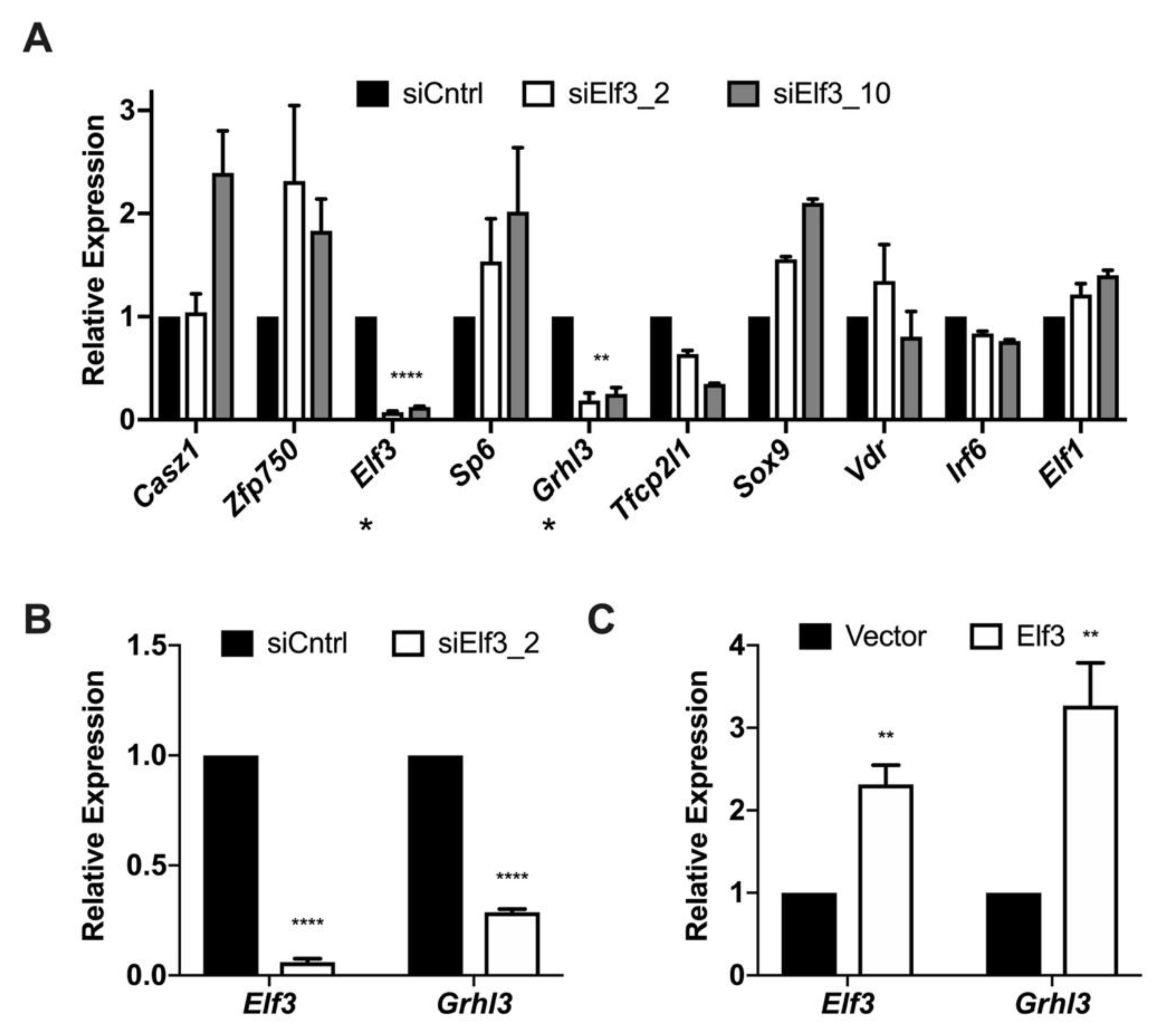
3.3. Elf3 Regulates the Expression of Grhl3
3.4. The Transcription Factor Elf3 Activates the Promoter of Grhl3
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 2009, 119, 1420–1428. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Xing, T.; Yang, Z.; Dudek, R.; Lu, Q.; Chen, Y.H. Epithelial Mesenchymal Transition in Embryonic Development, Tissue Repair and Cancer: A Comprehensive Overview. J. Clin. Med. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Lamouille, S.; Xu, J.; Derynck, R. Molecular mechanisms of epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 2014, 15, 178–196. [Google Scholar] [CrossRef] [PubMed]
- Pei, D.; Shu, X.; Gassama-Diagne, A.; Thiery, J.P. Mesenchymal-epithelial transition in development and reprogramming. Nat. Cell Biol. 2019, 21, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Nieto, M.A. Epithelial-Mesenchymal Transitions in development and disease: Old views and new perspectives. Int J. Dev. Biol. 2009, 53, 1541–1547. [Google Scholar] [CrossRef] [PubMed]
- Brabletz, T. To differentiate or not--routes towards metastasis. Nat. Rev. Cancer 2012, 12, 425–436. [Google Scholar] [CrossRef] [PubMed]
- Savagner, P. Epithelial-mesenchymal transitions: From cell plasticity to concept elasticity. Curr. Top. Dev. Biol. 2015, 112, 273–300. [Google Scholar] [CrossRef]
- Thiery, J.P.; Acloque, H.; Huang, R.Y.; Nieto, M.A. Epithelial-mesenchymal transitions in development and disease. Cell 2009, 139, 871–890. [Google Scholar] [CrossRef]
- Stemmler, M.P. Cadherins in development and cancer. Mol. Biosyst. 2008, 4, 835–850. [Google Scholar] [CrossRef]
- Wells, A.; Yates, C.; Shepard, C.R. E-cadherin as an indicator of mesenchymal to epithelial reverting transitions during the metastatic seeding of disseminated carcinomas. Clin. Exp. Metastasis 2008, 25, 621–628. [Google Scholar] [CrossRef]
- Lombaerts, M.; van Wezel, T.; Philippo, K.; Dierssen, J.W.; Zimmerman, R.M.; Oosting, J.; van Eijk, R.; Eilers, P.H.; van de Water, B.; Cornelisse, C.J.; et al. E-cadherin transcriptional downregulation by promoter methylation but not mutation is related to epithelial-to-mesenchymal transition in breast cancer cell lines. Br. J. Cancer 2006, 94, 661–671. [Google Scholar] [CrossRef] [PubMed]
- Bolos, V.; Peinado, H.; Perez-Moreno, M.A.; Fraga, M.F.; Esteller, M.; Cano, A. The transcription factor Slug represses E-cadherin expression and induces epithelial to mesenchymal transitions: A comparison with Snail and E47 repressors. J. Cell Sci. 2003, 116, 499–511. [Google Scholar] [CrossRef] [PubMed]
- Puisieux, A.; Brabletz, T.; Caramel, J. Oncogenic roles of EMT-inducing transcription factors. Nat. Cell Biol. 2014, 16, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Gomez, S.J.; Maziveyi, M.; Alahari, S.K. Regulation of epithelial-mesenchymal transition through epigenetic and post-translational modifications. Mol. Cancer 2016, 15, 18. [Google Scholar] [CrossRef] [PubMed]
- Canel, M.; Serrels, A.; Frame, M.C.; Brunton, V.G. E-cadherin-integrin crosstalk in cancer invasion and metastasis. J. Cell Sci. 2013, 126, 393–401. [Google Scholar] [CrossRef] [PubMed]
- Yee, D.S.; Tang, Y.; Li, X.; Liu, Z.; Guo, Y.; Ghaffar, S.; McQueen, P.; Atreya, D.; Xie, J.; Simoneau, A.R.; et al. The Wnt inhibitory factor 1 restoration in prostate cancer cells was associated with reduced tumor growth, decreased capacity of cell migration and invasion and a reversal of epithelial to mesenchymal transition. Mol. Cancer 2010, 9, 162. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zheng, Y.W.; Sano, Y.; Taniguchi, H. Evidence for mesenchymal-epithelial transition associated with mouse hepatic stem cell differentiation. PLoS ONE 2011, 6, e17092. [Google Scholar] [CrossRef] [PubMed]
- Almeida, M.S.; Bray, S.J. Regulation of post-embryonic neuroblasts by Drosophila Grainyhead. Mech. Dev. 2005, 122, 1282–1293. [Google Scholar] [CrossRef]
- Delva, E.; Kowalczyk, A.P. Regulation of cadherin trafficking. Traffic 2009, 10, 259–267. [Google Scholar] [CrossRef]
- Wheelock, M.J.; Shintani, Y.; Maeda, M.; Fukumoto, Y.; Johnson, K.R. Cadherin switching. J. Cell Sci. 2008, 121, 727–735. [Google Scholar] [CrossRef]
- Schafer, G.; Narasimha, M.; Vogelsang, E.; Leptin, M. Cadherin switching during the formation and differentiation of the Drosophila mesoderm - implications for epithelial-to-mesenchymal transitions. J. Cell Sci. 2014, 127, 1511–1522. [Google Scholar] [CrossRef] [PubMed]
- Gheldof, A.; Berx, G. Cadherins and epithelial-to-mesenchymal transition. Prog. Mol. Biol. Transl. Sci. 2013, 116, 317–336. [Google Scholar] [CrossRef] [PubMed]
- Stemmler, M.P.; Hecht, A.; Kinzel, B.; Kemler, R. Analysis of regulatory elements of E-cadherin with reporter gene constructs in transgenic mouse embryos. Dev. Dyn 2003, 227, 238–245. [Google Scholar] [CrossRef] [PubMed]
- Stemmler, M.P.; Hecht, A.; Kemler, R. E-cadherin intron 2 contains cis-regulatory elements essential for gene expression. Development 2005, 132, 965–976. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Alotaibi, H.; Basilicata, M.F.; Shehwana, H.; Kosowan, T.; Schreck, I.; Braeutigam, C.; Konu, O.; Brabletz, T.; Stemmler, M.P. Enhancer cooperativity as a novel mechanism underlying the transcriptional regulation of E-cadherin during mesenchymal to epithelial transition. Biochim. Et Biophys. Acta-Gene Regul. Mech. 2015, 1849, 731–742. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Liang, J.; Ni, S.; Zhou, T.; Qing, X.; Li, H.; He, W.; Chen, J.; Li, F.; Zhuang, Q.; et al. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 2010, 7, 51–63. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, A.; Loughner, C.L.; Swamynathan, S.; Swamynathan, S.K. KLF4 Plays an Essential Role in Corneal Epithelial Homeostasis by Promoting Epithelial Cell Fate and Suppressing Epithelial-Mesenchymal Transition. Investig. Ophthalmol. Vis. Sci. 2017, 58, 2785–2795. [Google Scholar] [CrossRef] [PubMed]
- Varma, S.; Cao, Y.; Tagne, J.B.; Lakshminarayanan, M.; Li, J.; Friedman, T.B.; Morell, R.J.; Warburton, D.; Kotton, D.N.; Ramirez, M.I. The transcription factors Grainyhead-like 2 and NK2-homeobox 1 form a regulatory loop that coordinates lung epithelial cell morphogenesis and differentiation. J. Biol. Chem. 2012, 287, 37282–37295. [Google Scholar] [CrossRef]
- Xiang, X.; Deng, Z.; Zhuang, X.; Ju, S.; Mu, J.; Jiang, H.; Zhang, L.; Yan, J.; Miller, D.; Zhang, H.G. Grhl2 determines the epithelial phenotype of breast cancers and promotes tumor progression. PLoS ONE 2012, 7, e50781. [Google Scholar] [CrossRef]
- Chen, W.; Xiao Liu, Z.; Oh, J.E.; Shin, K.H.; Kim, R.H.; Jiang, M.; Park, N.H.; Kang, M.K. Grainyhead-like 2 (GRHL2) inhibits keratinocyte differentiation through epigenetic mechanism. Cell Death Dis. 2012, 3, e450. [Google Scholar] [CrossRef]
- Werth, M.; Walentin, K.; Aue, A.; Schonheit, J.; Wuebken, A.; Pode-Shakked, N.; Vilianovitch, L.; Erdmann, B.; Dekel, B.; Bader, M.; et al. The transcription factor grainyhead-like 2 regulates the molecular composition of the epithelial apical junctional complex. Development 2010, 137, 3835–3845. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, B.S.; Irizarry, R.A. A framework for oligonucleotide microarray preprocessing. Bioinformatics 2010, 26, 2363–2367. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing. 2018. Available online: ftp://ftp.uvigo.es/CRAN/web/packages/dplR/vignettes/intro-dplR.pdf (accessed on 25 February 2019).
- Yasrebi, H. Comparative study of joint analysis of microarray gene expression data in survival prediction and risk assessment of breast cancer patients. Brief. Bioinform. 2016, 17, 771–785. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, J.W. mogene10sttranscriptcluster.db: Affymetrix Mogene10 Annotation Data (Chip Mogene10sttranscriptcluster). 2017. Available online: https://bioconductor.org/packages/release/data/annotation/html/mogene10sttranscriptcluster.db.html (accessed on 25 February 2019).
- Hu, H.; Miao, Y.R.; Jia, L.H.; Yu, Q.Y.; Zhang, Q.; Guo, A.Y. AnimalTFDB 3.0: A comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res. 2019, 47, D33–D38. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- Lichti, U.; Anders, J.; Yuspa, S.H. Isolation and short-term culture of primary keratinocytes, hair follicle populations and dermal cells from newborn mice and keratinocytes from adult mice for in vitro analysis and for grafting to immunodeficient mice. Nat. Protoc. 2008, 3, 799–810. [Google Scholar] [CrossRef] [PubMed]
- Alotaibi, H.; Yaman, E.; Salvatore, D.; Di Dato, V.; Telkoparan, P.; Di Lauro, R.; Tazebay, U.H. Intronic elements in the Na+/I- symporter gene (NIS) interact with retinoic acid receptors and mediate initiation of transcription. Nucleic Acids Res. 2010, 38, 3172–3185. [Google Scholar] [CrossRef] [PubMed]
- Albino, D.; Longoni, N.; Curti, L.; Mello-Grand, M.; Pinton, S.; Civenni, G.; Thalmann, G.; D’Ambrosio, G.; Sarti, M.; Sessa, F.; et al. ESE3/EHF controls epithelial cell differentiation and its loss leads to prostate tumors with mesenchymal and stem-like features. Cancer Res. 2012, 72, 2889–2900. [Google Scholar] [CrossRef]
- Huan, H.; Wen, X.; Chen, X.; Wu, L.; Liu, W.; Habib, N.A.; Bie, P.; Xia, F. C/EBPalpha Short-Activating RNA Suppresses Metastasis of Hepatocellular Carcinoma through Inhibiting EGFR/beta-Catenin Signaling Mediated EMT. PLoS ONE 2016, 11, e0153117. [Google Scholar] [CrossRef]
- Mistry, D.S.; Chen, Y.; Sen, G.L. Progenitor function in self-renewing human epidermis is maintained by the exosome. Cell Stem Cell 2012, 11, 127–135. [Google Scholar] [CrossRef]
- Roca, H.; Hernandez, J.; Weidner, S.; McEachin, R.C.; Fuller, D.; Sud, S.; Schumann, T.; Wilkinson, J.E.; Zaslavsky, A.; Li, H.; et al. Transcription factors OVOL1 and OVOL2 induce the mesenchymal to epithelial transition in human cancer. PLoS ONE 2013, 8, e76773. [Google Scholar] [CrossRef] [PubMed]
- Xue, P.; Fu, J.; Zhou, Y. The Aryl Hydrocarbon Receptor and Tumor Immunity. Front. Immunol. 2018, 9, 286. [Google Scholar] [CrossRef] [PubMed]
- Cooper, C.D.; Newman, J.A.; Gileadi, O. Recent advances in the structural molecular biology of Ets transcription factors: Interactions, interfaces and inhibition. Biochem. Soc. Trans. 2014, 42, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, N.; Yoshida, S.; Araie, M.; Handa, H.; Nabeshima, Y. Ets family transcription factor ESE-1 is expressed in corneal epithelial cells and is involved in their differentiation. Mech. Dev. 2000, 97, 27–34. [Google Scholar] [CrossRef]
- Andreoli, J.M.; Jang, S.I.; Chung, E.; Coticchia, C.M.; Steinert, P.M.; Markova, N.G. The expression of a novel, epithelium-specific ets transcription factor is restricted to the most differentiated layers in the epidermis. Nucleic Acids Res. 1997, 25, 4287–4295. [Google Scholar] [CrossRef] [PubMed]
- Hollenhorst, P.C.; McIntosh, L.P.; Graves, B.J. Genomic and biochemical insights into the specificity of ETS transcription factors. Annu. Rev. Biochem. 2011, 80, 437–471. [Google Scholar] [CrossRef]
- Otero, M.; Plumb, D.A.; Tsuchimochi, K.; Dragomir, C.L.; Hashimoto, K.; Peng, H.; Olivotto, E.; Bevilacqua, M.; Tan, L.; Yang, Z.; et al. E74-like factor 3 (ELF3) impacts on matrix metalloproteinase 13 (MMP13) transcriptional control in articular chondrocytes under proinflammatory stress. J. Biol. Chem. 2012, 287, 3559–3572. [Google Scholar] [CrossRef]
- Yordy, J.S.; Muise-Helmericks, R.C. Signal transduction and the Ets family of transcription factors. Oncogene 2000, 19, 6503–6513. [Google Scholar] [CrossRef]
- Hsu, T.; Trojanowska, M.; Watson, D.K. Ets proteins in biological control and cancer. J. Cell Biochem. 2004, 91, 896–903. [Google Scholar] [CrossRef]
- Li, R.; Pei, H.; Watson, D.K. Regulation of Ets function by protein - protein interactions. Oncogene 2000, 19, 6514–6523. [Google Scholar] [CrossRef]
- Maroulakou, I.G.; Bowe, D.B. Expression and function of Ets transcription factors in mammalian development: A regulatory network. Oncogene 2000, 19, 6432–6442. [Google Scholar] [CrossRef] [PubMed]
- Oikawa, T. ETS transcription factors: Possible targets for cancer therapy. Cancer Sci. 2004, 95, 626–633. [Google Scholar] [CrossRef] [PubMed]
- Wasylyk, B.; Hahn, S.L.; Giovane, A. The Ets family of transcription factors. Eur. J. Biochem. 1993, 211, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Oettgen, P.; Carter, K.C.; Augustus, M.; Barcinski, M.; Boltax, J.; Kunsch, C.; Libermann, T.A. The novel epithelial-specific Ets transcription factor gene ESX maps to human chromosome 1q32.1. Genomics 1997, 45, 456–457. [Google Scholar] [CrossRef] [PubMed]
- Luk, I.Y.; Reehorst, C.M.; Mariadason, J.M. ELF3, ELF5, EHF and SPDEF Transcription Factors in Tissue Homeostasis and Cancer. Molecules 2018, 23. [Google Scholar] [CrossRef] [PubMed]
- Oliver, J.R.; Kushwah, R.; Hu, J. Multiple roles of the epithelium-specific ETS transcription factor, ESE-1, in development and disease. Lab. Investig. 2012, 92, 320–330. [Google Scholar] [CrossRef] [PubMed]
- Kushwah, R.; Oliver, J.R.; Wu, J.; Chang, Z.; Hu, J. Elf3 regulates allergic airway inflammation by controlling dendritic cell-driven T cell differentiation. J. Immunol. 2011, 187, 4639–4653. [Google Scholar] [CrossRef] [PubMed]
- Gajulapalli, V.N.; Samanthapudi, V.S.; Pulaganti, M.; Khumukcham, S.S.; Malisetty, V.L.; Guruprasad, L.; Chitta, S.K.; Manavathi, B. A transcriptional repressive role for epithelial-specific ETS factor ELF3 on oestrogen receptor alpha in breast cancer cells. Biochem. J. 2016, 473, 1047–1061. [Google Scholar] [CrossRef] [PubMed]
- Longoni, N.; Sarti, M.; Albino, D.; Civenni, G.; Malek, A.; Ortelli, E.; Pinton, S.; Mello-Grand, M.; Ostano, P.; D’Ambrosio, G.; et al. ETS transcription factor ESE1/ELF3 orchestrates a positive feedback loop that constitutively activates NF-kappaB and drives prostate cancer progression. Cancer Res. 2013, 73, 4533–4547. [Google Scholar] [CrossRef] [PubMed]
- Tymms, M.J.; Ng, A.Y.; Thomas, R.S.; Schutte, B.C.; Zhou, J.; Eyre, H.J.; Sutherland, G.R.; Seth, A.; Rosenberg, M.; Papas, T.; et al. A novel epithelial-expressed ETS gene, ELF3: Human and murine cDNA sequences, murine genomic organization, human mapping to 1q32.2 and expression in tissues and cancer. Oncogene 1997, 15, 2449–2462. [Google Scholar] [CrossRef]
- Yu, Z.; Xu, J.; Liu, J.; Wu, J.; Lee, C.M.; Yu, L.; Hu, J. Epithelium-Specific Ets-Like Transcription Factor 1, ESE-1, Regulates ICAM-1 Expression in Cultured Lung Epithelial Cell Lines. Mediat. Inflamm. 2015, 2015, 547928. [Google Scholar] [CrossRef] [PubMed]
- Kopp, J.L.; Wilder, P.J.; Desler, M.; Kim, J.H.; Hou, J.; Nowling, T.; Rizzino, A. Unique and selective effects of five Ets family members, Elf3, Ets1, Ets2, PEA3, and PU.1, on the promoter of the type II transforming growth factor-beta receptor gene. J. Biol. Chem. 2004, 279, 19407–19420. [Google Scholar] [CrossRef] [PubMed]
- Ng, A.Y.; Waring, P.; Ristevski, S.; Wang, C.; Wilson, T.; Pritchard, M.; Hertzog, P.; Kola, I. Inactivation of the transcription factor Elf3 in mice results in dysmorphogenesis and altered differentiation of intestinal epithelium. Gastroenterology 2002, 122, 1455–1466. [Google Scholar] [CrossRef] [PubMed]
- Bock, M.; Hinley, J.; Schmitt, C.; Wahlicht, T.; Kramer, S.; Southgate, J. Identification of ELF3 as an early transcriptional regulator of human urothelium. Dev. Biol. 2014, 386, 321–330. [Google Scholar] [CrossRef] [PubMed]
- Scholz, G.M.; Sulaiman, N.S.; Al Baiiaty, S.; Kwa, M.Q.; Reynolds, E.C. A novel regulatory relationship between RIPK4 and ELF3 in keratinocytes. Cell Signal. 2016, 28, 1916–1922. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Cheng, P.; Wang, J.; Qiu, X.; Zhang, X.; Xu, L.; Liu, Y.; Qin, S. IRF6 Is Directly Regulated by ZEB1 and ELF3, and Predicts a Favorable Prognosis in Gastric Cancer. Front. Oncol. 2019, 9, 220. [Google Scholar] [CrossRef] [PubMed]
- Sayan, A.E.; Griffiths, T.R.; Pal, R.; Browne, G.J.; Ruddick, A.; Yagci, T.; Edwards, R.; Mayer, N.J.; Qazi, H.; Goyal, S.; et al. SIP1 protein protects cells from DNA damage-induced apoptosis and has independent prognostic value in bladder cancer. Proc. Natl. Acad. Sci. USA 2009, 106, 14884–14889. [Google Scholar] [CrossRef] [PubMed]
- Onder, T.T.; Gupta, P.B.; Mani, S.A.; Yang, J.; Lander, E.S.; Weinberg, R.A. Loss of E-cadherin promotes metastasis via multiple downstream transcriptional pathways. Cancer Res. 2008, 68, 3645–3654. [Google Scholar] [CrossRef] [PubMed]
- Elston, M.S.; Gill, A.J.; Conaglen, J.V.; Clarkson, A.; Cook, R.J.; Little, N.S.; Robinson, B.G.; Clifton-Bligh, R.J.; McDonald, K.L. Nuclear accumulation of e-cadherin correlates with loss of cytoplasmic membrane staining and invasion in pituitary adenomas. J. Clin. Endocrinol. Metab. 2009, 94, 1436–1442. [Google Scholar] [CrossRef]
- Thurmond, J.; Goodman, J.L.; Strelets, V.B.; Attrill, H.; Gramates, L.S.; Marygold, S.J.; Matthews, B.B.; Millburn, G.; Antonazzo, G.; Trovisco, V.; et al. FlyBase 2.0: The next generation. Nucleic Acids Res. 2019, 47, D759–D765. [Google Scholar] [CrossRef]
- Schober, M.; Rebay, I.; Perrimon, N. Function of the ETS transcription factor Yan in border cell migration. Development 2005, 132, 3493–3504. [Google Scholar] [CrossRef] [PubMed]
- Bray, S.J.; Kafatos, F.C. Developmental function of Elf-1: An essential transcription factor during embryogenesis in Drosophila. Genes Dev. 1991, 5, 1672–1683. [Google Scholar] [CrossRef] [PubMed]
- Boffelli, D.; McAuliffe, J.; Ovcharenko, D.; Lewis, K.D.; Ovcharenko, I.; Pachter, L.; Rubin, E.M. Phylogenetic shadowing of primate sequences to find functional regions of the human genome. Science 2003, 299, 1391–1394. [Google Scholar] [CrossRef] [PubMed]
- Dubchak, I.; Brudno, M.; Loots, G.G.; Pachter, L.; Mayor, C.; Rubin, E.M.; Frazer, K.A. Active conservation of noncoding sequences revealed by three-way species comparisons. Genome Res. 2000, 10, 1304–1306. [Google Scholar] [CrossRef] [PubMed]
- Duan, Z.; Li, Y.; Li, L. Promoting epithelial-to-mesenchymal transition by D-kynurenine via activating aryl hydrocarbon receptor. Mol. Cell Biochem. 2018, 448, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Qu, Y.; Li, X.; Sui, F.; Yao, D.; Yang, Q.; Shi, B.; Ji, M.; Hou, P. Increased expression of EHF via gene amplification contributes to the activation of HER family signaling and associates with poor survival in gastric cancer. Cell Death Dis. 2016, 7, e2442. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.K.; Han, H.Q.; Zhang, H.J.; Xu, M.; Li, L.; Chen, L.; Xiang, T.; Feng, Q.S.; Kang, T.; Qian, C.N.; et al. OVOL2 links stemness and metastasis via fine-tuning epithelial-mesenchymal transition in nasopharyngeal carcinoma. Theranostics 2018, 8, 2202–2216. [Google Scholar] [CrossRef]
- Wang, Z.H.; Li, Z.; Hu, M.; Yang, Q.J.; Yan, S.; Wu, R.S.; Li, B.A.; Guo, M. Ovol2 gene inhibits the Epithelial-to-Mesenchymal Transition in lung adenocarcinoma by transcriptionally repressing Twist1. Gene 2017, 600, 1–8. [Google Scholar] [CrossRef]
- Watanabe, K.; Liu, Y.; Noguchi, S.; Murray, M.; Chang, J.C.; Kishima, M.; Nishimura, H.; Hashimoto, K.; Minoda, A.; Suzuki, H. OVOL2 induces mesenchymal-to-epithelial transition in fibroblasts and enhances cell-state reprogramming towards epithelial lineages. Sci. Rep. 2019, 9, 6490. [Google Scholar] [CrossRef]
- Chakrabarti, R.; Hwang, J.; Andres Blanco, M.; Wei, Y.; Lukacisin, M.; Romano, R.A.; Smalley, K.; Liu, S.; Yang, Q.; Ibrahim, T.; et al. Elf5 inhibits the epithelial-mesenchymal transition in mammary gland development and breast cancer metastasis by transcriptionally repressing Snail2. Nat. Cell Biol. 2012, 14, 1212–1222. [Google Scholar] [CrossRef]
- Watanabe, K.; Villarreal-Ponce, A.; Sun, P.; Salmans, M.L.; Fallahi, M.; Andersen, B.; Dai, X. Mammary morphogenesis and regeneration require the inhibition of EMT at terminal end buds by Ovol2 transcriptional repressor. Dev. Cell 2014, 29, 59–74. [Google Scholar] [CrossRef] [PubMed]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sengez, B.; Aygün, I.; Shehwana, H.; Toyran, N.; Tercan Avci, S.; Konu, O.; Stemmler, M.P.; Alotaibi, H. The Transcription Factor Elf3 Is Essential for a Successful Mesenchymal to Epithelial Transition. Cells 2019, 8, 858. https://doi.org/10.3390/cells8080858
Sengez B, Aygün I, Shehwana H, Toyran N, Tercan Avci S, Konu O, Stemmler MP, Alotaibi H. The Transcription Factor Elf3 Is Essential for a Successful Mesenchymal to Epithelial Transition. Cells. 2019; 8(8):858. https://doi.org/10.3390/cells8080858
Chicago/Turabian StyleSengez, Burcu, Ilkin Aygün, Huma Shehwana, Neslihan Toyran, Sanem Tercan Avci, Ozlen Konu, Marc P. Stemmler, and Hani Alotaibi. 2019. "The Transcription Factor Elf3 Is Essential for a Successful Mesenchymal to Epithelial Transition" Cells 8, no. 8: 858. https://doi.org/10.3390/cells8080858
APA StyleSengez, B., Aygün, I., Shehwana, H., Toyran, N., Tercan Avci, S., Konu, O., Stemmler, M. P., & Alotaibi, H. (2019). The Transcription Factor Elf3 Is Essential for a Successful Mesenchymal to Epithelial Transition. Cells, 8(8), 858. https://doi.org/10.3390/cells8080858







