The Ubiquitin Moiety of Ubi1 Is Required for Productive Expression of Ribosomal Protein eL40 in Saccharomyces cerevisiae
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strains and Microbiological Methods
2.2. Plasmids
2.3. Polysome Analysis and Sucrose Gradient Fractionation
2.4. Western Blot Analysis and Antibodies
2.5. Analysis of Aggregated Proteins
2.6. Reproducibility
3. Results
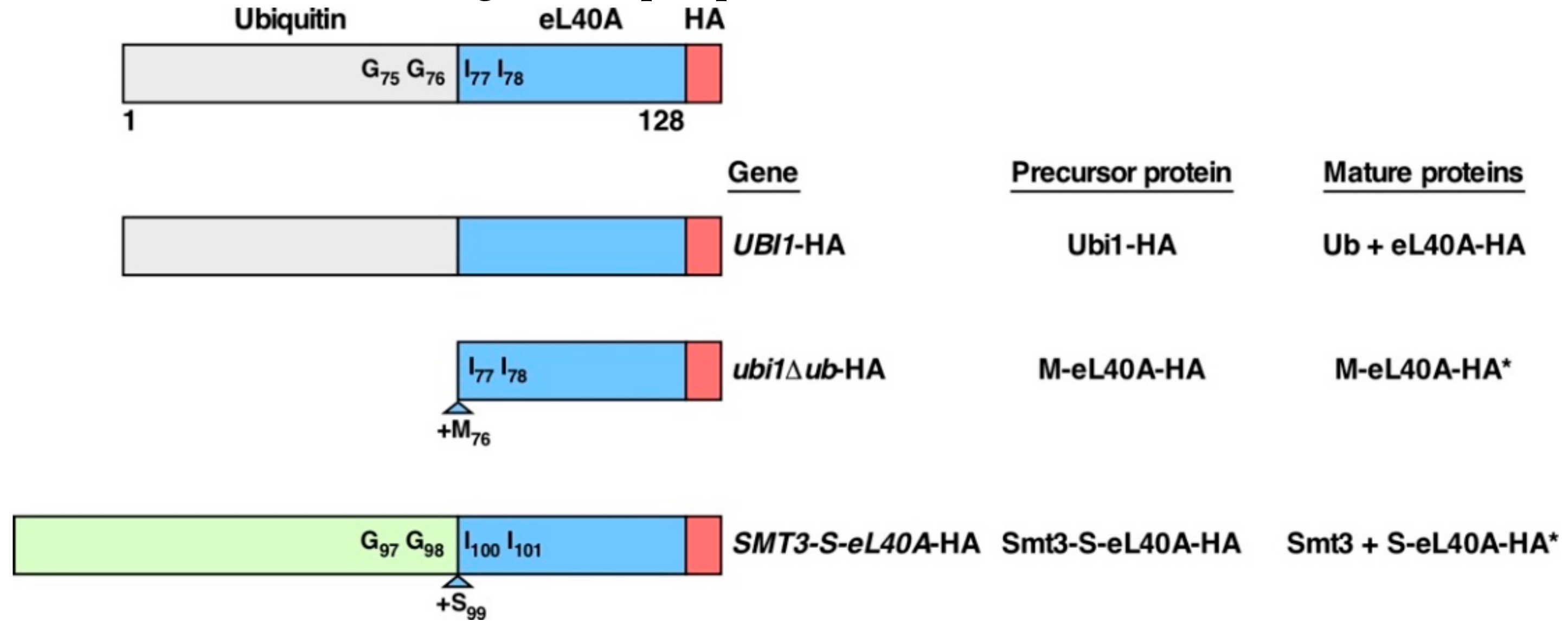
3.1. Generation of ubi1 Mutants for Phenotypic Analysis
3.2. The ubi1∆ub Ubiquitin Deletion Mutant Displays a Slow-Growth Phenotype
3.3. The SMT3-S-eL40A ubi2∆ Mutant Displays a Slow-Growth Phenotype
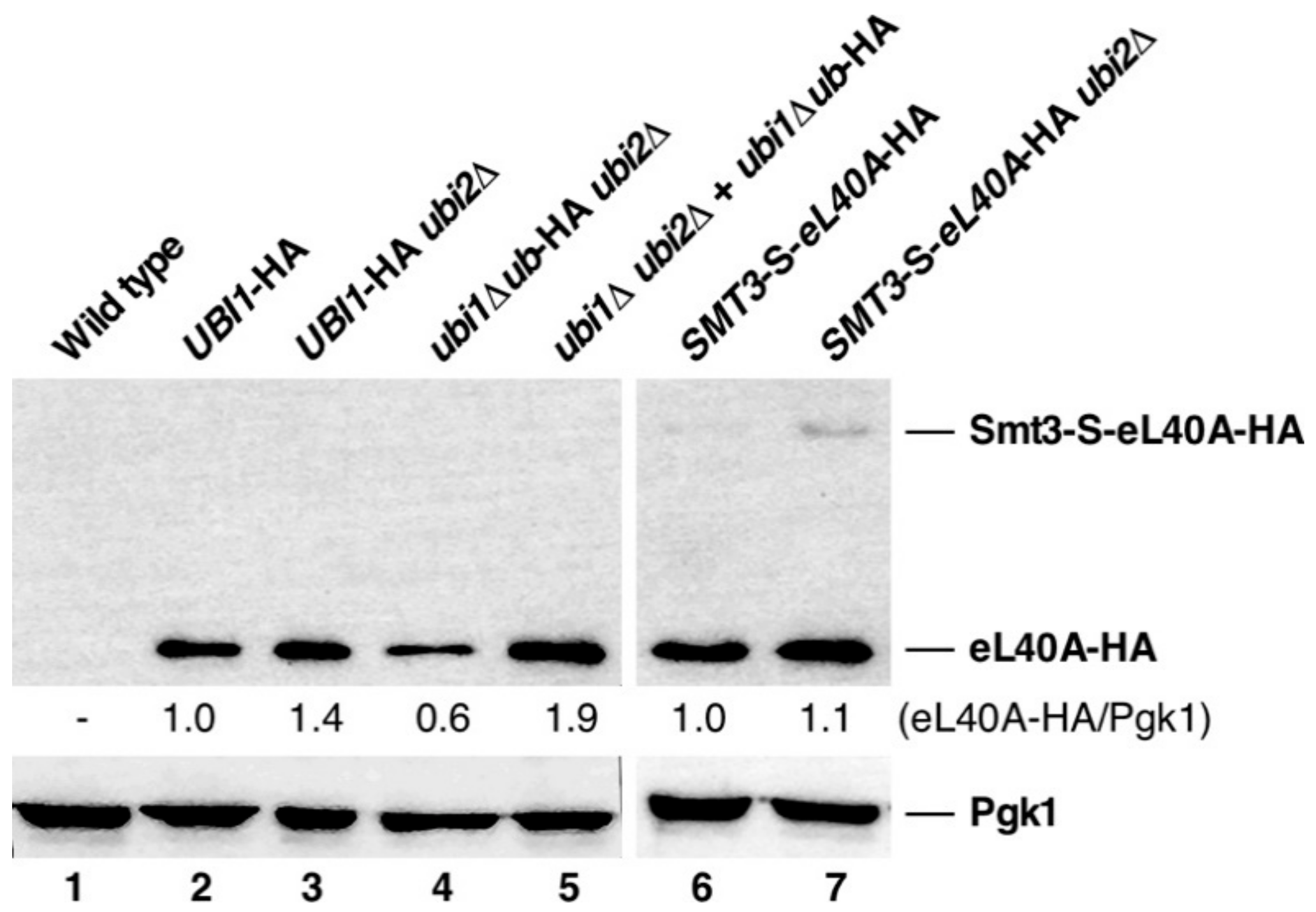
3.4. Contribution of the Ubiquitin and SUMO Moieties to the Expression of eL40A
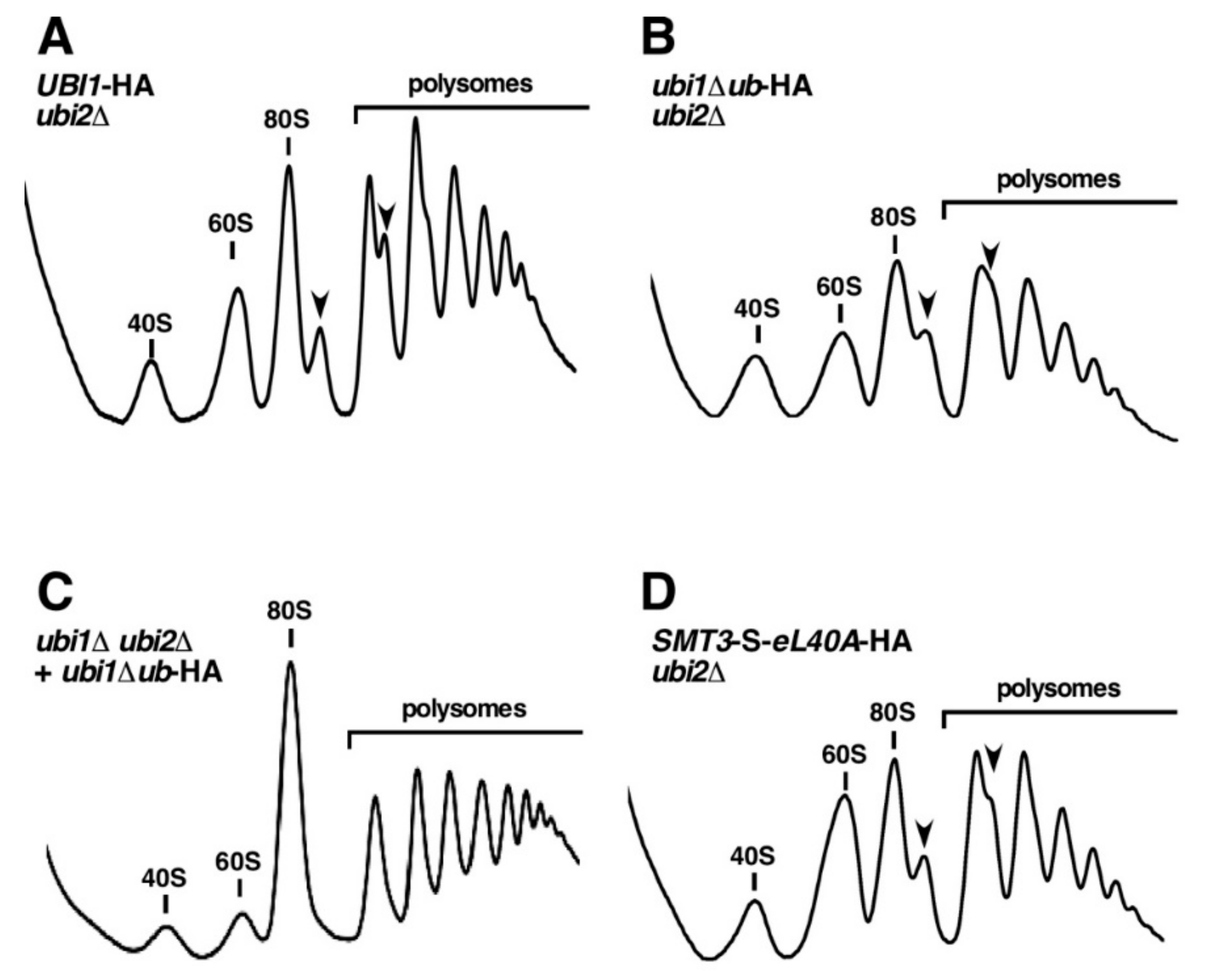
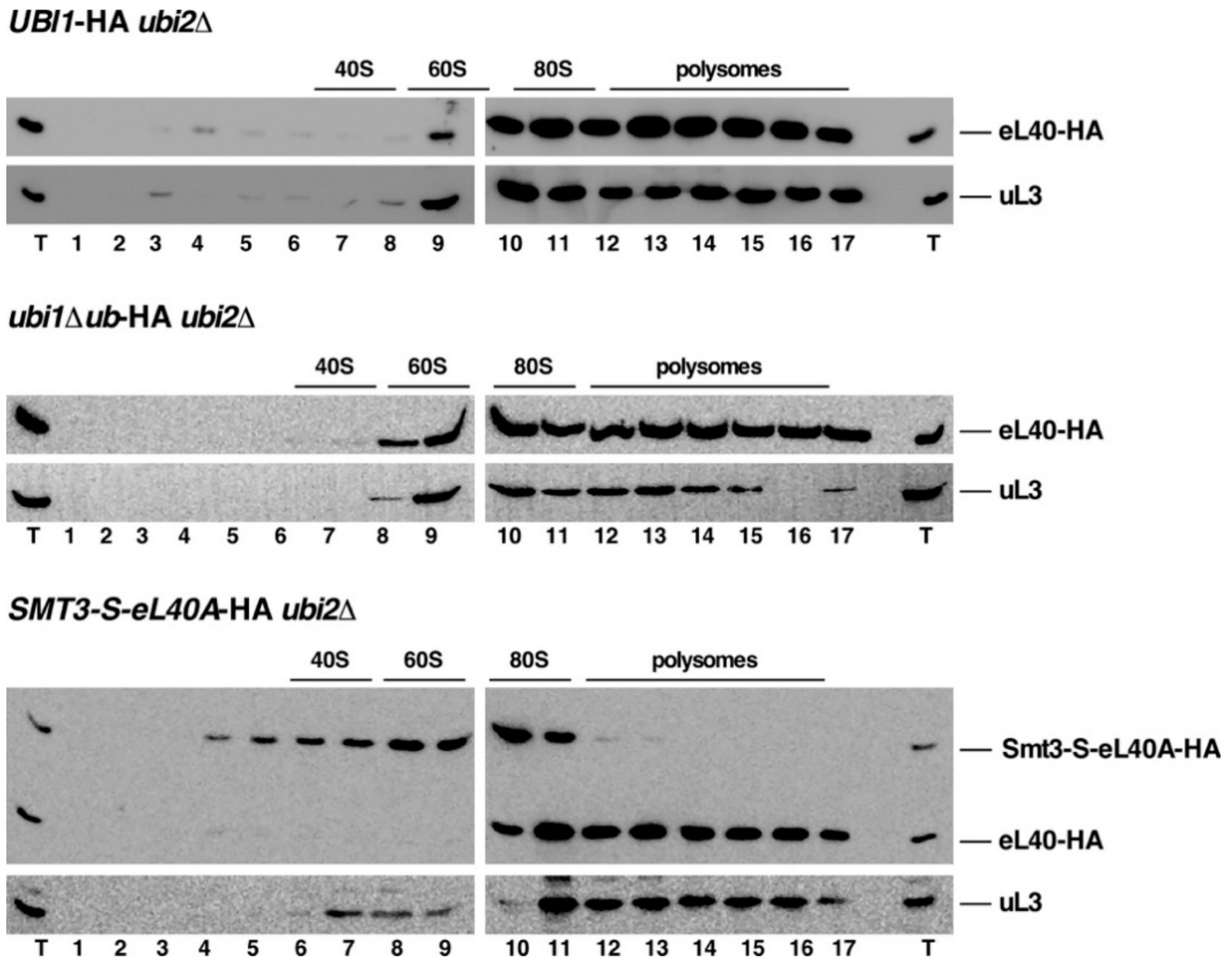
3.5. The Genomic ubi1∆ub-HA and SMT3-S-eL40A-HA Alleles Affect the Functional Integrity of 60S r-Subunits
3.6. In Trans Expression of Free Ubiquitin Ffails to Rescue the Deficiencies of the ubi1∆ub-HA ubi2∆ and SMT3-S-eL40A-HA ubi2∆ Mutants
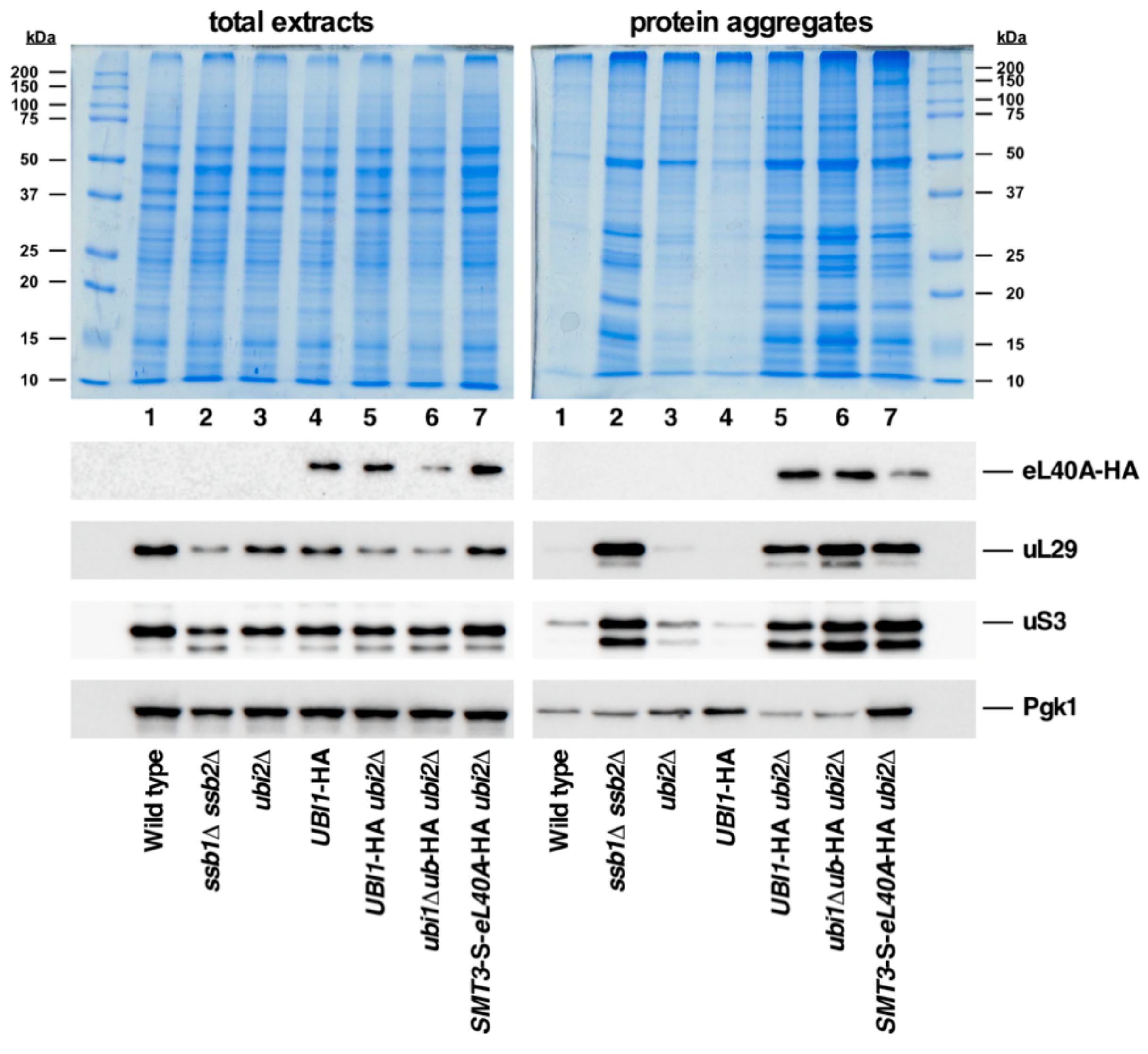
3.7. Ubiquitin and SUMO Modestly Prevent eL40 or eS31 Protein Aggregation
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Iyer, L.M.; Burroughs, A.M.; Aravind, L. The prokaryotic antecedents of the ubiquitin-signaling system and the early evolution of ubiquitin-like beta-grasp domains. Genome Biol. 2006, 7, R60. [Google Scholar] [CrossRef] [PubMed]
- Kwon, Y.T.; Ciechanover, A. The ubiquitin code in the ubiquitin-proteasome system and autophagy. Trends Biochem. Sci. 2017, 42, 873–886. [Google Scholar] [CrossRef] [PubMed]
- Pickart, C.M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 2001, 70, 503–533. [Google Scholar] [CrossRef] [PubMed]
- Clague, M.J.; Urbé, S. Ubiquitin: Same molecule, different degradation pathways. Cell 2010, 143, 682–685. [Google Scholar] [CrossRef] [PubMed]
- Amerik, A.Y.; Hochstrasser, M. Mechanism and function of deubiquitinating enzymes. Biochim. Biophys. Acta 2004, 1695, 189–207. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Archibald, J.M.; Teh, E.M.; Keeling, P.J. Novel ubiquitin fusion proteins: Ribosomal protein P1 and actin. J. Mol. Biol. 2003, 328, 771–778. [Google Scholar] [CrossRef]
- Gemayel, R.; Yang, Y.; Dzialo, M.C.; Kominek, J.; Vowinckel, J.; Saels, V.; Van Huffel, L.; van der Zande, E.; Ralser, M.; Steensels, J.; et al. Variable repeats in the eukaryotic polyubiquitin gene ubi4 modulate proteostasis and stress survival. Nat. Commun. 2017, 8, 397. [Google Scholar] [CrossRef] [PubMed]
- Özkaynak, E.; Finley, D.; Solomon, M.J.; Varshavsky, A. The yeast ubiquitin genes: A family of natural gene fusions. EMBO J. 1987, 6, 1429–1439. [Google Scholar] [CrossRef]
- Baker, R.T.; Board, P.G. The human ubiquitin gene family: Structure of a gene and pseudogenes from the Ub B subfamily. Nucleic Acids Res. 1987, 15, 443–463. [Google Scholar] [CrossRef]
- Callis, J.; Carpenter, T.; Sun, C.W.; Vierstra, R.D. Structure and evolution of genes encoding polyubiquitin and ubiquitin-like proteins in Arabidopsis thaliana ecotype Columbia. Genetics 1995, 139, 921–939. [Google Scholar]
- Krebber, H.; Wöstmann, C.; Bakker-Grunwald, T. Evidence for the existence of a single ubiquitin gene in Giardia lamblia. FEBS Lett. 1994, 343, 234–236. [Google Scholar] [CrossRef]
- Wöstmann, C.; Liakopoulos, D.; Ciechanover, A.; Bakker-Grunwald, T. Characterization of ubiquitin genes and -transcripts and demonstration of a ubiquitin-conjugating system in Entamoeba histolytica. Mol. Biochem. Parasitol. 1996, 82, 81–90. [Google Scholar] [CrossRef]
- Sibbald, S.J.; Hopkins, J.F.; Filloramo, G.V.; Archibald, J.M. Ubiquitin fusion proteins in algae: Implications for cell biology and the spread of photosynthesis. BMC Genom. 2019, 20, 38. [Google Scholar] [CrossRef] [PubMed]
- Finley, D.; Bartel, B.; Varshavsky, A. The tails of ubiquitin precursors are ribosomal proteins whose fusion to ubiquitin facilitates ribosome biogenesis. Nature 1989, 338, 394–401. [Google Scholar] [CrossRef] [PubMed]
- Finley, D.; Özkaynak, E.; Varshavsky, A. The yeast polyubiquitin gene is essential for resistance to high temperatures, starvation, and other stresses. Cell 1987, 48, 1035–1046. [Google Scholar] [CrossRef]
- Treger, J.M.; Heichman, K.A.; McEntee, K. Expression of the yeast UB14 gene increases in response to DNA-damaging agents and in meiosis. Mol. Cell. Biol. 1988, 8, 1132–1136. [Google Scholar] [CrossRef]
- Simon, J.R.; Treger, J.M.; McEntee, K. Multiple independent regulatory pathways control UBI4 expression after heat shock in Saccharomyces cerevisiae. Mol. Microbiol. 1999, 31, 823–832. [Google Scholar] [CrossRef]
- Cheng, L.; Watt, R.; Piper, P.W. Polyubiquitin gene expression contributes to oxidative stress resistance in respiratory yeast (Saccharomyces cerevisiae). Mol. Gen. Genet. 1994, 243, 358–362. [Google Scholar] [CrossRef]
- MacDiarmid, C.W.; Taggart, J.; Jeong, J.; Kerdsomboon, K.; Eide, D.J. Activation of the yeast UBI4 polyubiquitin gene by Zap1 transcription factor via an intragenic promoter is critical for Zinc-deficient growth. J. Biol. Chem. 2016, 291, 18880–18896. [Google Scholar] [CrossRef]
- Fraser, J.; Luu, H.A.; Neculcea, J.; Thomas, D.Y.; Storms, R.K. Ubiquitin gene expression: Response to environmental changes. Curr. Genet. 1991, 20, 17–23. [Google Scholar] [CrossRef]
- Pickart, C.M.; Eddins, M.J. Ubiquitin: Structures, functions, mechanisms. Biochim. Biophys. Acta 2004, 1695, 55–72. [Google Scholar] [CrossRef] [PubMed]
- Johnson, E.S. Protein modification by SUMO. Annu. Rev. Biochem. 2004, 73, 355–382. [Google Scholar] [CrossRef] [PubMed]
- Ciechanover, A.; Schwartz, A.L. The ubiquitin-mediated proteolytic pathway: Mechanisms of recognition of the proteolytic substrate and involvement in the degradation of native cellular proteins. FASEB J. 1994, 8, 182–191. [Google Scholar] [CrossRef] [PubMed]
- Müller, S.; Hoege, C.; Pyrowolakis, G.; Jentsch, S. SUMO, ubiquitin’s mysterious cousin. Nat. Rev. Mol. Cell Biol. 2001, 2, 202–210. [Google Scholar] [CrossRef] [PubMed]
- Peroutka Iii, R.J.; Orcutt, S.J.; Strickler, J.E.; Butt, T.R. SUMO fusion technology for enhanced protein expression and purification in prokaryotes and eukaryotes. Methods Mol. Biol. 2011, 705, 15–30. [Google Scholar] [PubMed]
- Lee, C.D.; Sun, H.C.; Hu, S.M.; Chiu, C.F.; Homhuan, A.; Liang, S.M.; Leng, C.H.; Wang, T.F. An improved SUMO fusion protein system for effective production of native proteins. Protein Sci. 2008, 17, 1241–1248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Butt, T.R.; Jonnalagadda, S.; Monia, B.P.; Sternberg, E.J.; Marsh, J.A.; Stadel, J.M.; Ecker, D.J.; Crooke, S.T. Ubiquitin fusion augments the yield of cloned gene products in Escherichia coli. Proc. Natl. Acad. Sci. USA 1989, 86, 2540–2544. [Google Scholar] [CrossRef]
- Ecker, D.J.; Stadel, J.M.; Butt, T.R.; Marsh, J.A.; Monia, B.P.; Powers, D.A.; Gorman, J.A.; Clark, P.E.; Warren, F.; Shatzman, A.; et al. Increasing gene expression in yeast by fusion to ubiquitin. J. Biol. Chem. 1989, 264, 7715–7719. [Google Scholar]
- Baker, R.T. Protein expression using ubiquitin fusion and cleavage. Curr. Opin. Biotechnol. 1996, 7, 541–546. [Google Scholar] [CrossRef]
- Graciet, E.; Hu, R.G.; Piatkov, K.; Rhee, J.H.; Schwarz, E.M.; Varshavsky, A. Aminoacyl-transferases and the N-end rule pathway of prokaryotic/eukaryotic specificity in a human pathogen. Proc. Natl. Acad. Sci. USA 2006, 103, 3078–3083. [Google Scholar] [CrossRef] [Green Version]
- Fernández-Pevida, A.; Martín-Villanueva, S.; Murat, G.; Lacombe, T.; Kressler, D.; de la Cruz, J. The eukaryote-specific N-terminal extension of ribosomal protein S31 contributes to the assembly and function of 40S ribosomal subunits. Nucleic Acids Res. 2016, 44, 7777–7791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, Q.; Zhu, X.; Qi, J.; An, W.; Lan, P.; Tan, D.; Chen, R.; Wang, B.; Zheng, S.; Zhang, C.; et al. Molecular architecture of the 90S small subunit pre-ribosome. eLife 2017, 6, e22086. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Pevida, A.; Rodríguez-Galán, O.; Díaz-Quintana, A.; Kressler, D.; de la Cruz, J. Yeast ribosomal protein L40 assembles late into precursor 60S ribosomes and is required for their cytoplasmic maturation. J. Biol. Chem. 2012, 287, 38390–38407. [Google Scholar] [CrossRef] [PubMed]
- Kruiswijk, T.; Planta, R.J.; Krop, J.M. The course of the assembly of ribosomal subunits in yeast. Biochim. Biophys. Acta 1978, 517, 378–389. [Google Scholar] [CrossRef]
- Lacombe, T.; García-Gómez, J.J.; de la Cruz, J.; Roser, D.; Hurt, E.; Linder, P.; Kressler, D. Linear ubiquitin fusion to Rps31 and its subsequent cleavage are required for the efficient production and functional integrity of 40S ribosomal subunits. Mol. Microbiol. 2009, 72, 69–84. [Google Scholar] [CrossRef] [PubMed]
- Martín-Villanueva, S.; Fernández-Pevida, A.; Fernández-Fernández, J.; Kressler, D.; de la Cruz, J. Ubiquitin release from eL40 is required for cytoplasmic maturation and function of 60S ribosomal subunits in Saccharomyces cerevisiae. FEBS J. 2019. [Google Scholar] [CrossRef] [PubMed]
- Thomas, B.J.; Rothstein, R. Elevated recombination rates in transcriptionally active DNA. Cell 1989, 56, 619–630. [Google Scholar] [CrossRef]
- Klockner, C.; Schneider, M.; Lutz, S.; Jani, D.; Kressler, D.; Stewart, M.; Hurt, E.; Kohler, A. Mutational uncoupling of the role of Sus1 in nuclear pore complex targeting of an mRNA export complex and histone H2B deubiquitination. J. Biol. Chem. 2009, 284, 12049–12056. [Google Scholar] [CrossRef]
- Burke, D.; Dawson, D.; Stearns, T. Methods in Yeast Genetics: A Cold Spring Harbor Laboratory Course Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2000. [Google Scholar]
- Gietz, D.; St., Jean, A.; Woods, R.A.; Schiestl, R.H. Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res. 1992, 20, 1425. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1989. [Google Scholar]
- de la Cruz, J.; Kressler, D.; Tollervey, D.; Linder, P. Dob1p (Mtr4p) is a putative ATP-dependent RNA helicase required for the 3’ end formation of 5.8S rRNA in Saccharomyces cerevisiae. EMBO J. 1998, 17, 1128–1140. [Google Scholar] [CrossRef]
- Gietz, R.D.; Sugino, A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 1988, 74, 527–534. [Google Scholar] [CrossRef]
- Kressler, D.; de la Cruz, J.; Rojo, M.; Linder, P. Fal1p is an essential DEAD-box protein involved in 40S-ribosomal-subunit biogenesis in Saccharomyces cerevisiae. Mol. Cell. Biol. 1997, 17, 7283–7294. [Google Scholar] [CrossRef] [PubMed]
- Foiani, M.; Cigan, A.M.; Paddon, C.J.; Harashima, S.; Hinnebusch, A.G. GCD2, a translational repressor of the GCN4 gene, has a general function in the initiation of protein synthesis in Saccharomyces cerevisiae. Mol. Cell. Biol. 1991, 11, 3203–3216. [Google Scholar] [CrossRef] [PubMed]
- De la Cruz, J.; Kressler, D.; Rojo, M.; Tollervey, D.; Linder, P. Spb4p, an essential putative RNA helicase, is required for a late step in the assembly of 60S ribosomal subunits in Saccharomyces cerevisiae. RNA 1998, 4, 1268–1281. [Google Scholar] [CrossRef] [PubMed]
- Yaffe, M.P.; Schatz, G. Two nuclear mutations that block mitochondrial protein import in yeast. Proc. Natl. Acad. Sci. USA 1984, 81, 4819–4823. [Google Scholar] [CrossRef] [PubMed]
- Vilardell, J.; Warner, J.R. Ribosomal protein L32 of Saccharomyces cerevisiae influences both the splicing of its own transcript and the processing of rRNA. Mol. Cell. Biol. 1997, 17, 1959–1965. [Google Scholar] [CrossRef] [PubMed]
- Frey, S.; Pool, M.; Seedorf, M. Scp160p, an RNA-binding, polysome-associated protein, localizes to the endoplasmic reticulum of Saccharomyces cerevisiae in a microtubule-dependent manner. J. Biol. Chem. 2001, 276, 15905–15912. [Google Scholar] [CrossRef]
- Koplin, A.; Preissler, S.; Ilina, Y.; Koch, M.; Scior, A.; Erhardt, M.; Deuerling, E. A dual function for chaperones SSB-RAC and the NAC nascent polypeptide-associated complex on ribosomes. J. Cell Biol. 2010, 189, 57–68. [Google Scholar] [CrossRef]
- Panasenko, O.O.; Collart, M.A. Presence of Not5 and ubiquitinated Rps7A in polysome fractions depends upon the Not4 E3 ligase. Mol. Microbiol. 2012, 83, 640–653. [Google Scholar] [CrossRef]
- van der Veen, A.G.; Ploegh, H.L. Ubiquitin-like proteins. Annu. Rev. Biochem. 2012, 81, 323–357. [Google Scholar] [CrossRef]
- Gareau, J.R.; Lima, C.D. The SUMO pathway: Emerging mechanisms that shape specificity, conjugation and recognition. Nat. Rev. Mol. Cell Biol. 2010, 11, 861–871. [Google Scholar] [CrossRef] [PubMed]
- Kemmler, S.; Occhipinti, L.; Veisu, M.; Panse, V.G. Yvh1 is required for a late maturation step in the 60S biogenesis pathway. J. Cell Biol. 2009, 186, 863–880. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodríguez-Mateos, M.; García-Gómez, J.J.; Francisco-Velilla, R.; Remacha, M.; de la Cruz, J.; Ballesta, J.P.G. Role and dynamics of the ribosomal protein P0 and its related trans-acting factor Mrt4 during ribosome assembly in Saccharomyces cerevisiae. Nucleic Acids Res. 2009, 37, 7519–7532. [Google Scholar] [CrossRef] [PubMed]
- Eisinger, D.P.; Dick, F.A.; Trumpower, B.L. Qsr1p, a 60S ribosomal subunit protein, is required for joining of 40S and 60S subunits. Mol. Cell. Biol. 1997, 17, 5136–5145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tye, B.W.; Commins, N.; Ryazanova, L.V.; Wuhr, M.; Springer, M.; Pincus, D.; Churchman, L.S. Proteotoxicity from aberrant ribosome biogenesis compromises cell fitness. elife 2019, 8, e43002. [Google Scholar] [CrossRef] [PubMed]
- Grou, C.P.; Pinto, M.P.; Mendes, A.V.; Domingues, P.; Azevedo, J.E. The de novo synthesis of ubiquitin: Identification of deubiquitinases acting on ubiquitin precursors. Sci. Rep. 2015, 5, 12836. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, M.; Oshima, S.; Maeyashiki, C.; Nibe, Y.; Otsubo, K.; Matsuzawa, Y.; Nemoto, Y.; Nagaishi, T.; Okamoto, R.; Tsuchiya, K.; et al. The ubiquitin hybrid gene UBA52 regulates ubiquitination of ribosome and sustains embryonic development. Sci. Rep. 2016, 6, 36780. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maicas, E.; Pluthero, F.G.; Friesen, J.D. The accumulation of three yeast ribosomal proteins under conditions of excess mRNA is determined primarily by fast protein decay. Mol. Cell. Biol. 1988, 8, 169–175. [Google Scholar] [CrossRef]
- Warner, J.R.; Mitra, G.; Schwindinger, W.F.; Studeny, M.; Fried, H.M. Saccharomyces cerevisiae coordinates accumulation of yeast ribosomal proteins by modulating mRNA splicing, translational initiation, and protein turnover. Mol. Cell. Biol. 1985, 5, 1512–1521. [Google Scholar] [CrossRef] [PubMed]
- Sung, M.K.; Reitsma, J.M.; Sweredoski, M.J.; Hess, S.; Deshaies, R.J. Ribosomal proteins produced in excess are degraded by the ubiquitin-proteasome system. Mol. Biol. Cell 2016, 27, 2642–2652. [Google Scholar] [CrossRef] [PubMed]
- Sung, M.K.; Porras-Yakushi, T.R.; Reitsma, J.M.; Huber, F.M.; Sweredoski, M.J.; Hoelz, A.; Hess, S.; Deshaies, R.J. A conserved quality-control pathway that mediates degradation of unassembled ribosomal proteins. eLife 2016, 5, e19105. [Google Scholar] [CrossRef] [PubMed]
- Lam, Y.W.; Lamond, A.I.; Mann, M.; Andersen, J.S. Analysis of nucleolar protein dynamics reveals the nuclear degradation of ribosomal proteins. Curr. Biol. 2007, 17, 749–760. [Google Scholar] [CrossRef] [PubMed]
- Marblestone, J.G.; Edavettal, S.C.; Lim, Y.; Lim, P.; Zuo, X.; Butt, T.R. Comparison of SUMO fusion technology with traditional gene fusion systems: Enhanced expression and solubility with SUMO. Protein Sci. 2006, 15, 182–189. [Google Scholar] [CrossRef] [PubMed]
- Jäkel, S.; Mingot, J.M.; Schwarzmaier, P.; Hartmann, E.; Görlich, D. Importins fulfil a dual function as nuclear import receptors and cytoplasmic chaperones for exposed basic domains. EMBO J. 2002, 21, 377–386. [Google Scholar] [CrossRef] [PubMed]
- Pillet, B.; Mitterer, V.; Kressler, D.; Pertschy, B. Hold on to your friends: Dedicated chaperones of ribosomal proteins: Dedicated chaperones mediate the safe transfer of ribosomal proteins to their site of pre-ribosome incorporation. Bioessays 2017, 39, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Espinar-Marchena, F.J.; Babiano, R.; de la Cruz, J. Placeholder factors in ribosome biogenesis: Please, pave my way. Microbial Cell 2017, 4, 144–168. [Google Scholar] [CrossRef] [PubMed]
- Peña, C.; Hurt, E.; Panse, V.G. Eukaryotic ribosome assembly, transport and quality control. Nat. Struct. Mol. Biol. 2017, 24, 689–699. [Google Scholar] [CrossRef]
- Catic, A.; Sun, Z.Y.; Ratner, D.M.; Misaghi, S.; Spooner, E.; Samuelson, J.; Wagner, G.; Ploegh, H.L. Sequence and structure evolved separately in a ribosomal ubiquitin variant. EMBO J. 2007, 26, 3474–3483. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.; Candido, E.P. Novel ubiqutin-like ribosomal protein fusion genes from the nematodes Caenorhabditis elegans and Caenorhabditis briggsae. J. Biol. Chem. 1993, 268, 19545–19551. [Google Scholar]




| Strain a | Relevant Genotype b | Source |
|---|---|---|
| W303-1A | MATaade2-1 his3-11,15 leu2-3,112 trp1-1 ura3-1 | [37] |
| W303-1B | As W303-1A but MATα | [37] |
| TAY001 | MATaubi1::kanMX4 ubi2::kanMX4 ade3::kanMX4 [pHT4467∆-UBI1] | [36] |
| SMY106 | MATaubi1::klURA3 | This study |
| SMY113 | MATa UBI1-HA | This study |
| SMY256 | MATaUBI1-HA ubi2::kanMX4 | This study |
| SMY107 | MATaubi1∆ub-HA | This study |
| SMY257 | MATα ubi1∆ub-HA ubi2::kanMX4 | This study |
| SMY215 | MATaubi1::kanMX4 ubi2::kanMX4 ade3::kanMX4 [YCplac111-ubi1∆ub-HA] | This study |
| JDY923 | MATα ubi2::kanMX4 | [33] |
| SMY218 | MATaSMT3-S-eL40A-HA | This study |
| SMY258 | MATα SMT3-S-eL40A-HA ubi2::kanMX4 | This study |
| SMY216 | MATaubi1::kanMX4 ubi2::kanMX4 ade3::kanMX4 [YCplac111-SMT3-S-eL40A-HA] | This study |
| JDY532 | MATassb1::HIS3MX6 ssb2::natNT2 | This study |
| SMY324 | MATassb1::HIS3MX6 ssb2::natNT2 UBI3-HA::kanMX4 | This study |
| TLY56.D3 | MATaUBI3-HA::kanMX4 | [35] |
| TLY61.A2 | MATaubi3∆ub-HA::kanMX4 | [35] |
| TLY14.3C | MATα ubi3::HIS3MX6 | [35] |
| SMY315 | MATα rps12::kanMX4 | This study |
| ORY211 | MATα rpl39::natNT2 | This study |
| MS157-1A | MATα dob1-1 | [42] |
| Name (Collection Name) | Relevant Information | Source |
|---|---|---|
| pHT4467∆-UBI1 | CEN6 (instable), URA3, ADE3. Wild-type Ubi1; promoter and terminator of UBI1 | [36] |
| YCplac111 | CEN, LEU2 | [43] |
| YCplac111-UBI1-HA (pDK4131) | CEN, LEU2. C-terminally 1xHA-tagged eL40A; promoter and terminator of UBI1 | [36] |
| YCplac111-ubi1∆ub-HA (pDK4192) | CEN, LEU2. Allele ubi1∆ub-HA (M-II); promoter and terminator of UBI1 | This study |
| YCplac111-SMT3-S-eL40-HA (pDK4193) | CEN, LEU2. Allele SMT3-S-eL40A-HA (GG-S-II); promoter and terminator of UBI1 | This study |
| pADH111-Ub (pDK2253) | CEN, LEU2. Wild-type ubiquitin; promoter and terminator of ADH1 | This study |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martín-Villanueva, S.; Fernández-Pevida, A.; Kressler, D.; de la Cruz, J. The Ubiquitin Moiety of Ubi1 Is Required for Productive Expression of Ribosomal Protein eL40 in Saccharomyces cerevisiae. Cells 2019, 8, 850. https://doi.org/10.3390/cells8080850
Martín-Villanueva S, Fernández-Pevida A, Kressler D, de la Cruz J. The Ubiquitin Moiety of Ubi1 Is Required for Productive Expression of Ribosomal Protein eL40 in Saccharomyces cerevisiae. Cells. 2019; 8(8):850. https://doi.org/10.3390/cells8080850
Chicago/Turabian StyleMartín-Villanueva, Sara, Antonio Fernández-Pevida, Dieter Kressler, and Jesús de la Cruz. 2019. "The Ubiquitin Moiety of Ubi1 Is Required for Productive Expression of Ribosomal Protein eL40 in Saccharomyces cerevisiae" Cells 8, no. 8: 850. https://doi.org/10.3390/cells8080850






