Lipoic Acid Synergizes with Antineoplastic Drugs in Colorectal Cancer by Targeting p53 for Proteasomal Degradation
Abstract
1. Introduction
2. Materials and Methods
2.1. Material
2.2. Cell Culture and Treatments
2.3. Preparation of Protein Lysates and Cell Fractionation
2.4. Co-Immunoprecipitation
2.5. Transient Transfection with siRNA
2.6. SDS-PAGE and Immunoblot Analysis
2.7. Preparation of RNA and Quantitative Real Time PCR
2.8. Confocal Immunofluorescence Microscopy of p53
2.9. Flow Cytometry-Based Analysis of Autophagy Induction
2.10. Cell Death Measurement by FLOW cytometry
2.11. Determination of Reactive Oxygen Species by Flow Cytometry
2.12. Assessment of Combination Effect
2.13. Statistics
3. Results
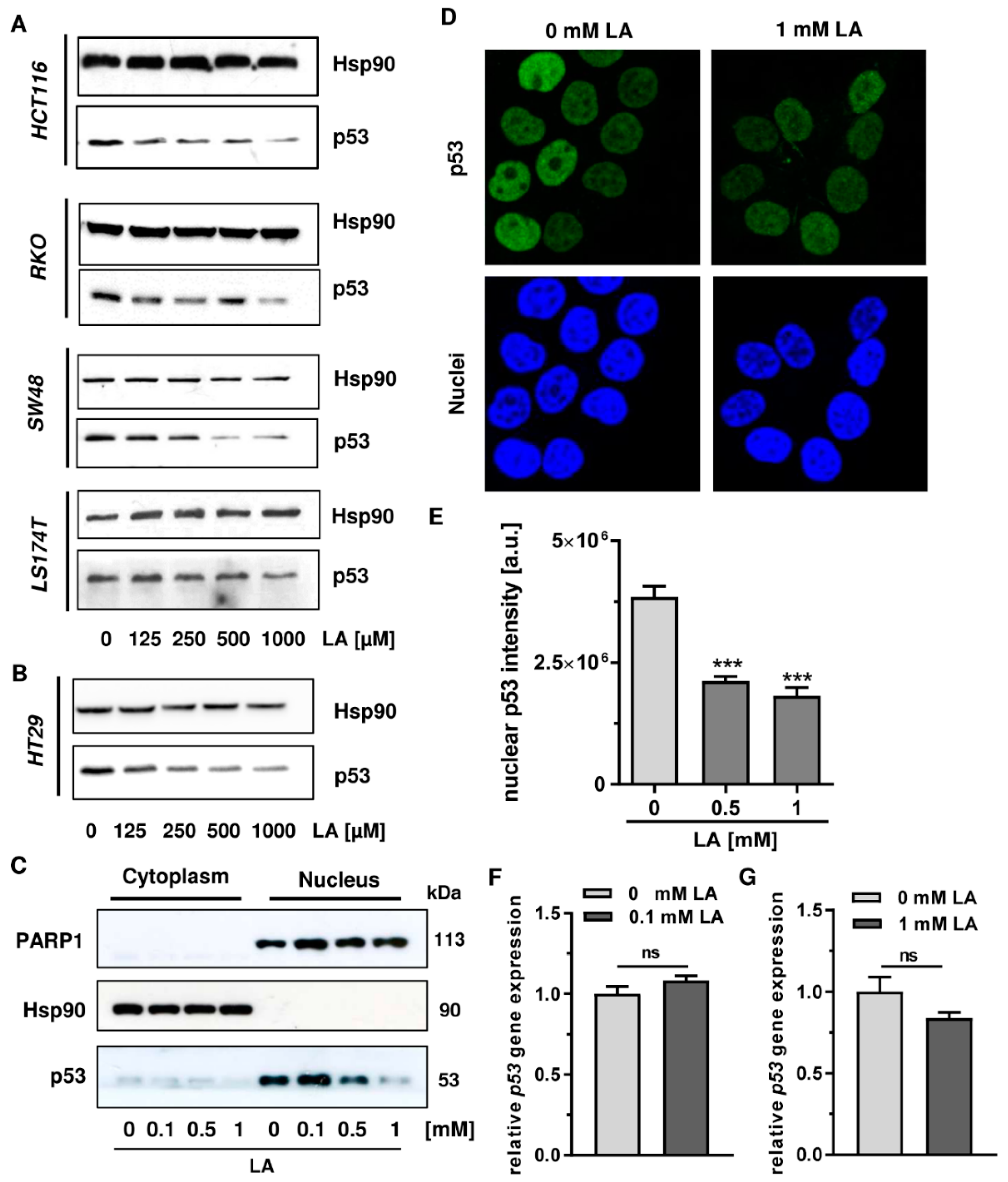
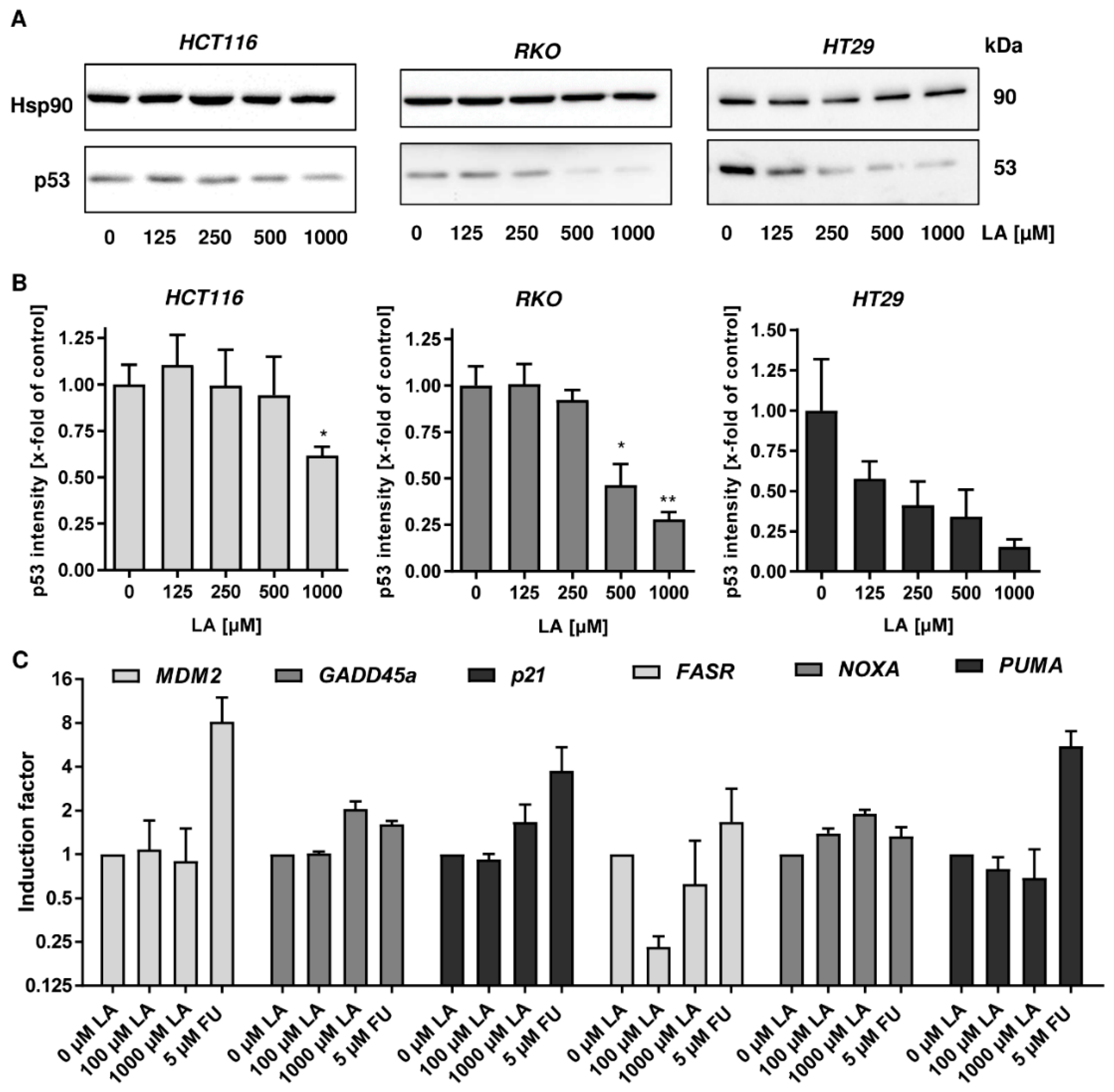
3.1. LA Leads to the Depletion of Wildtype and Mutant p53 in CRC Cell Lines
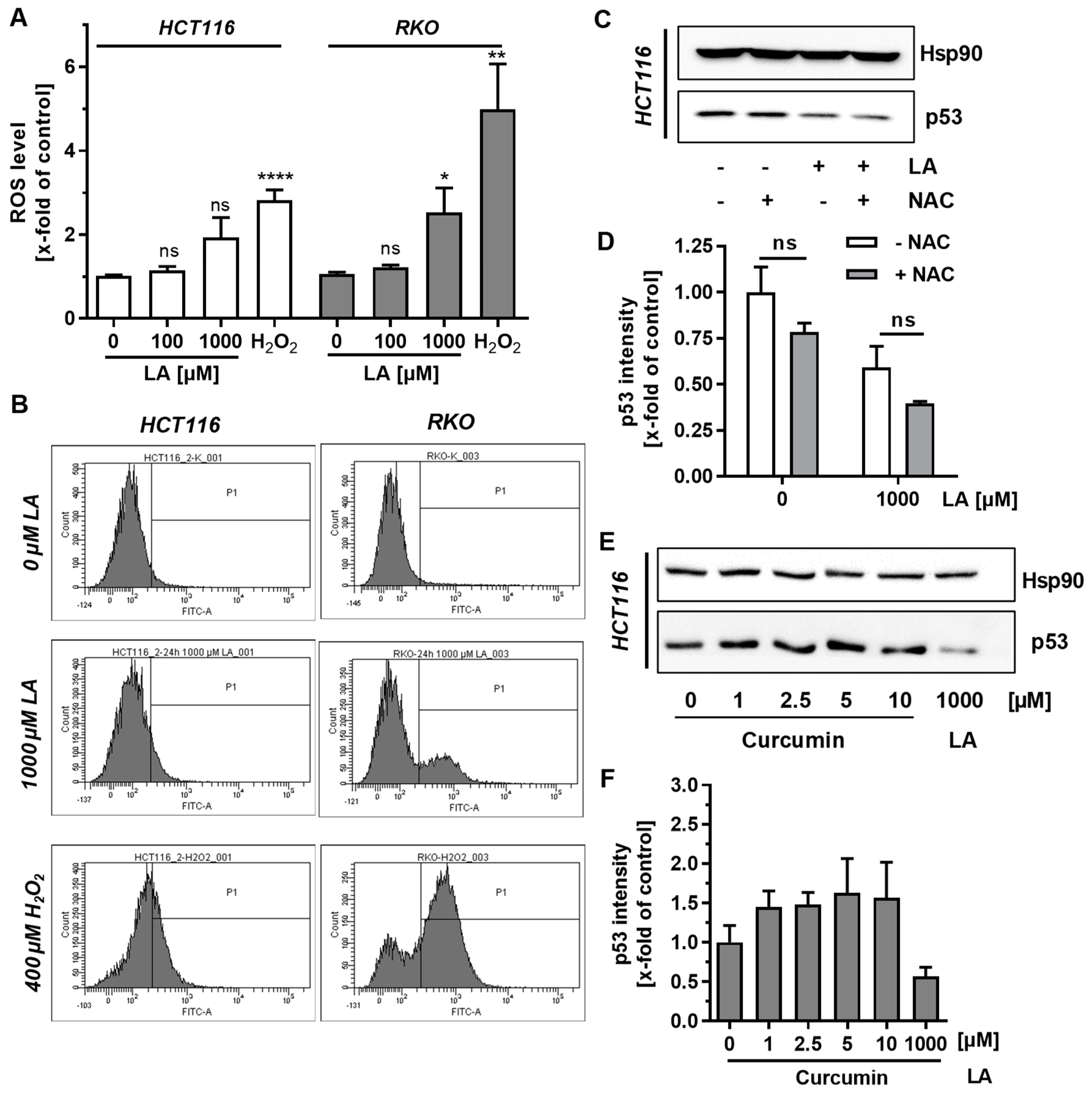
3.2. Supplementation with Antioxidants Does Not Rescue p53 Depletion
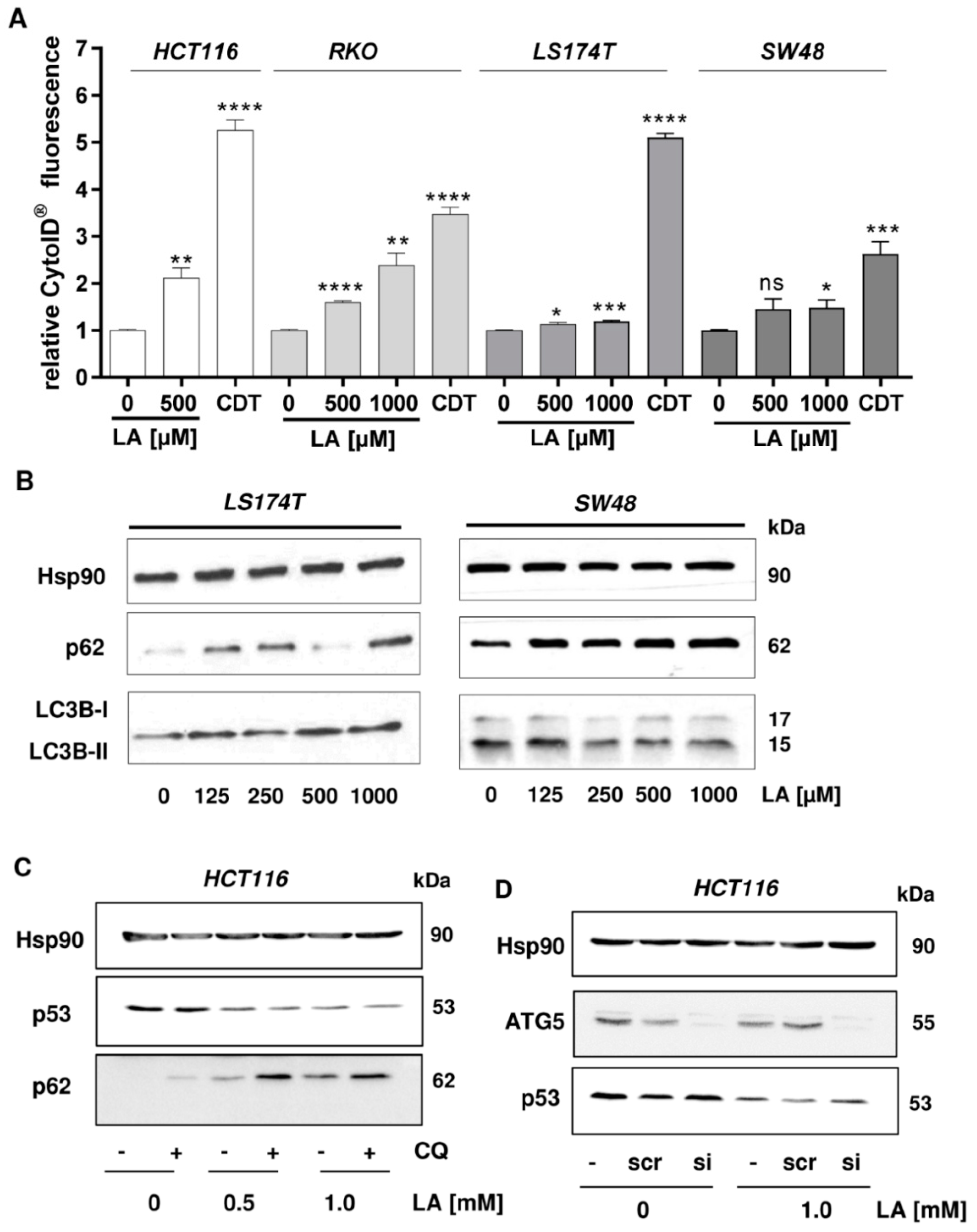
3.3. LA-Triggered Autophagy Is Not Involved in the Mechanism of p53 Depletion
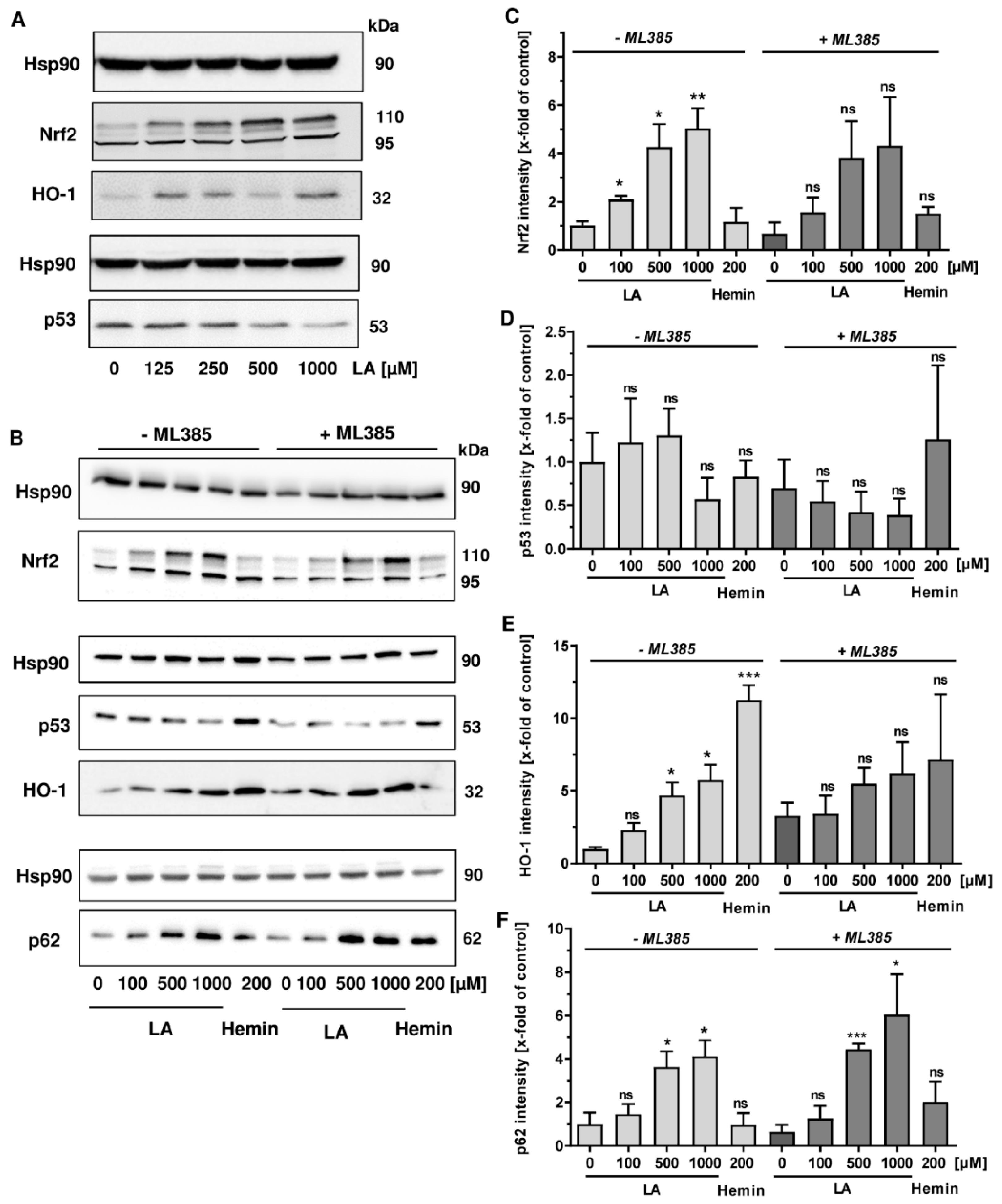
3.4. LA Causes Nrf2 Induction, Which is Dispensible for p53 Degradation
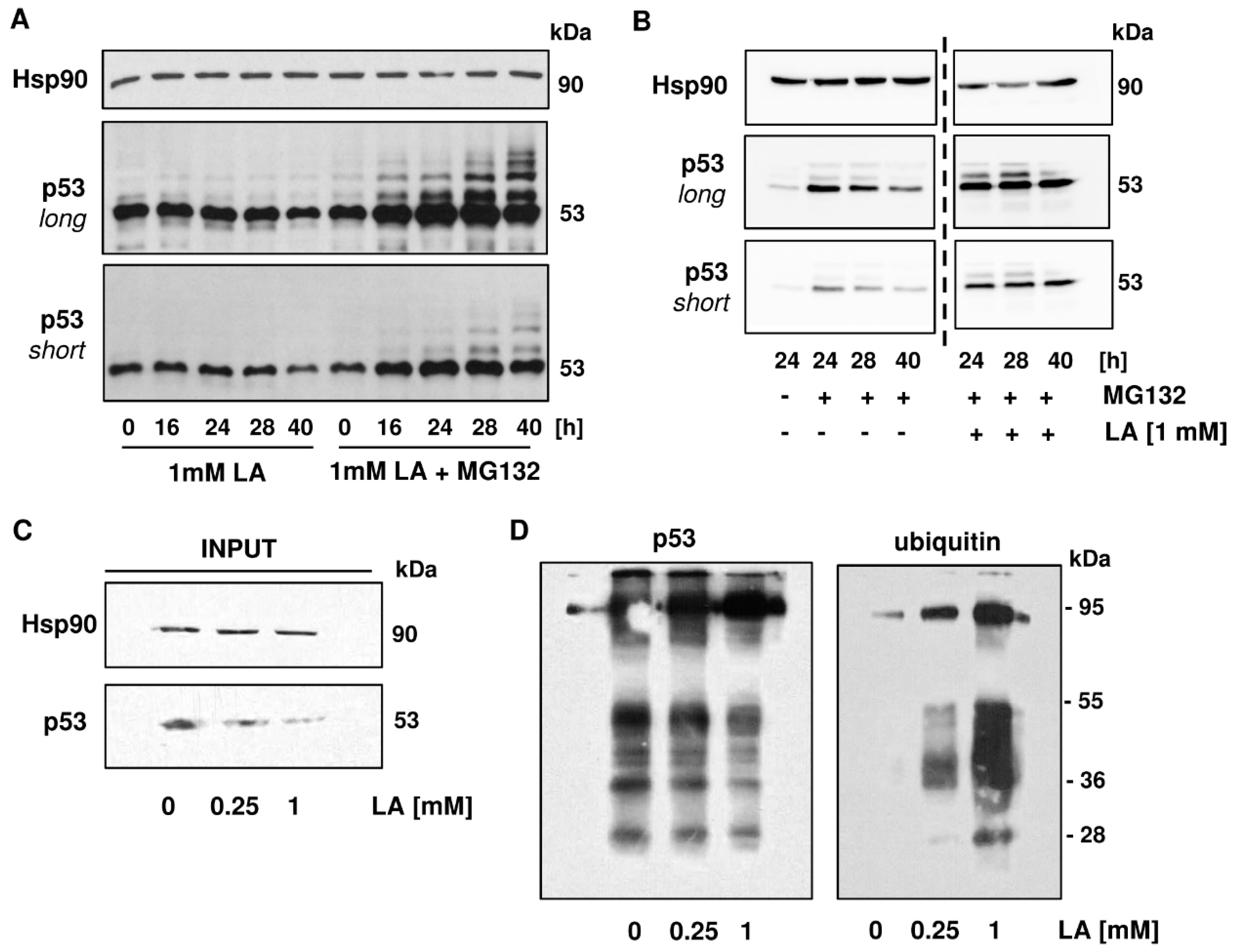
3.5. p53 is Readily Ubiquitinated upon LA Treatment
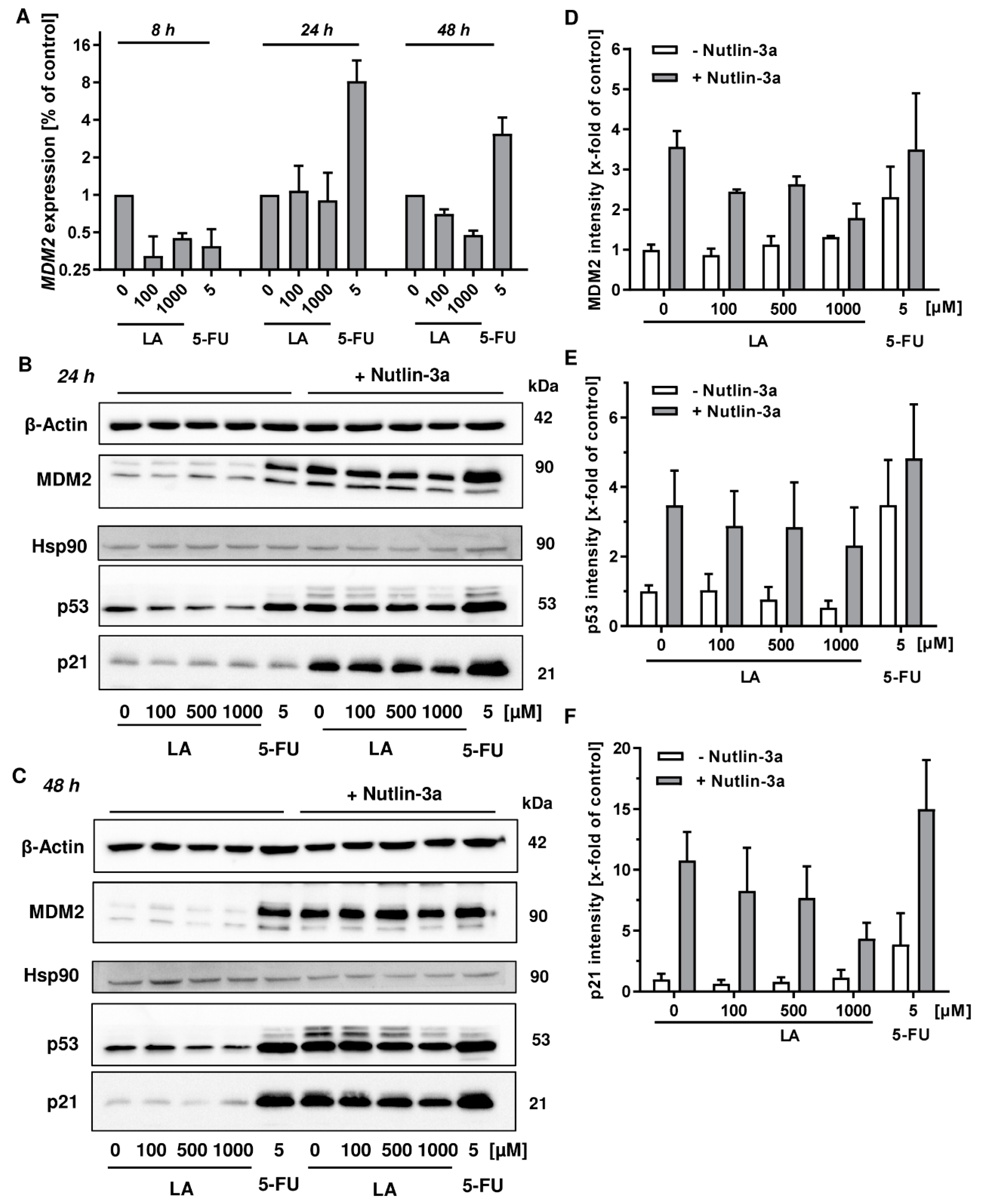
3.6. MDM2 Inhibition Does Not Rescue LA-Triggered p53 Degradation
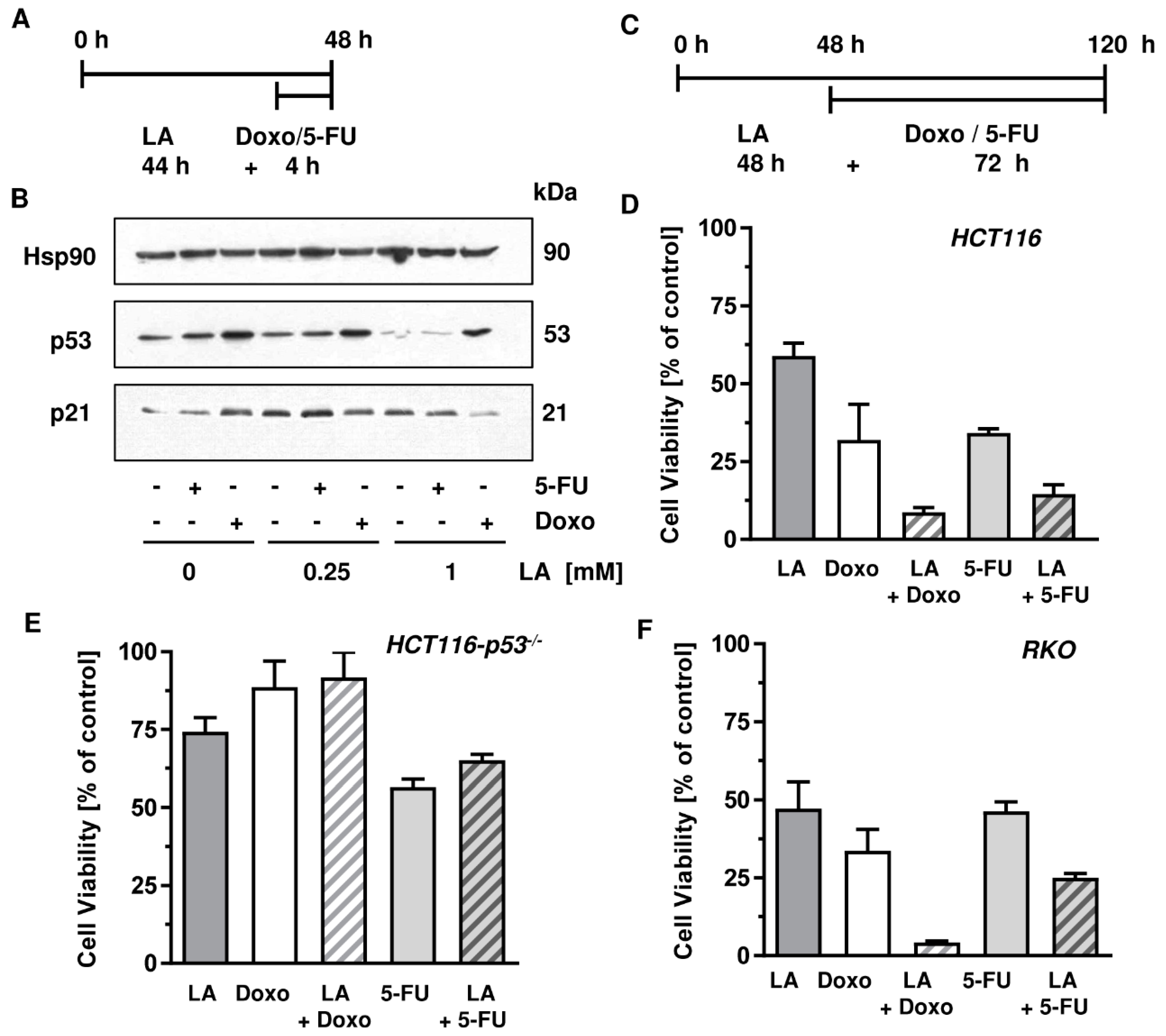
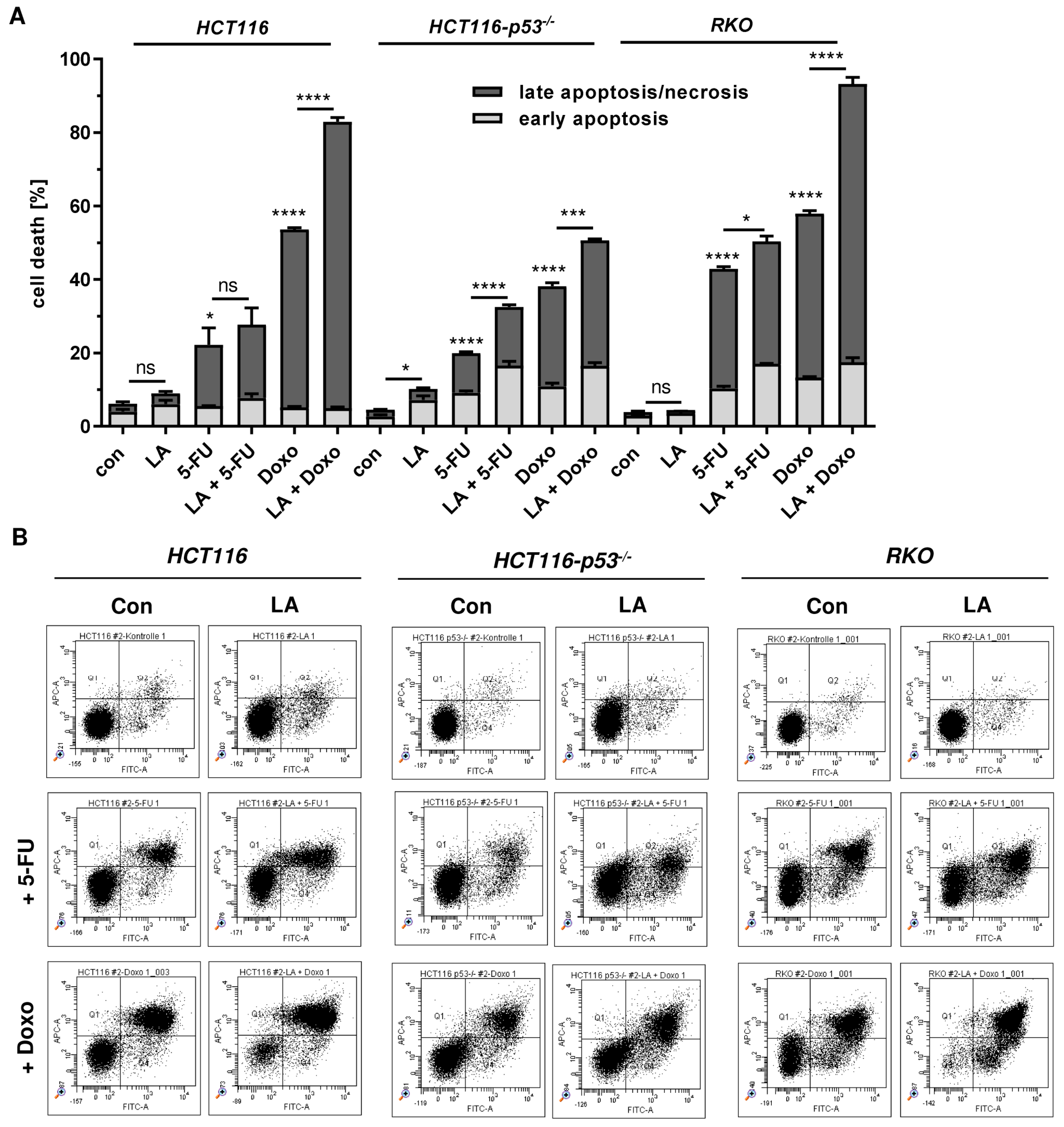
3.7. LA Synergizes with Standard Chemotherapeutics by Potentiating Cytotoxicity
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| 5-FU | 5-fluorouracil |
| ARE | antioxidant response element |
| CDT | cytolethal distending toxin |
| CI | combination index |
| CRC | colorectal cancer |
| CQ | chloroquine |
| DBD | DNA binding domain |
| DDR | DNA damage response |
| DHLA | dihydrolipoic acid |
| Doxo | doxorubicin |
| DSF | disulfiram |
| HO-1 | heme oxygenase 1 |
| LA | ɑ-lipoic acid |
| MDM2 | mouse double minute 2 homolog |
| MGMT | O6-methylguanine-DNA methyltransferase |
| NAC | N-Acetyl-cysteine |
| NEM | N-ethylmaleimide |
| NRF2 | nuclear factor erythroid 2-related factor 2 |
| PI | propidium iodide |
| ROS | reactive oxygen species |
| RT | room temperature |
| scr | scrambled |
| SEM | standard error of the means |
Appendix A
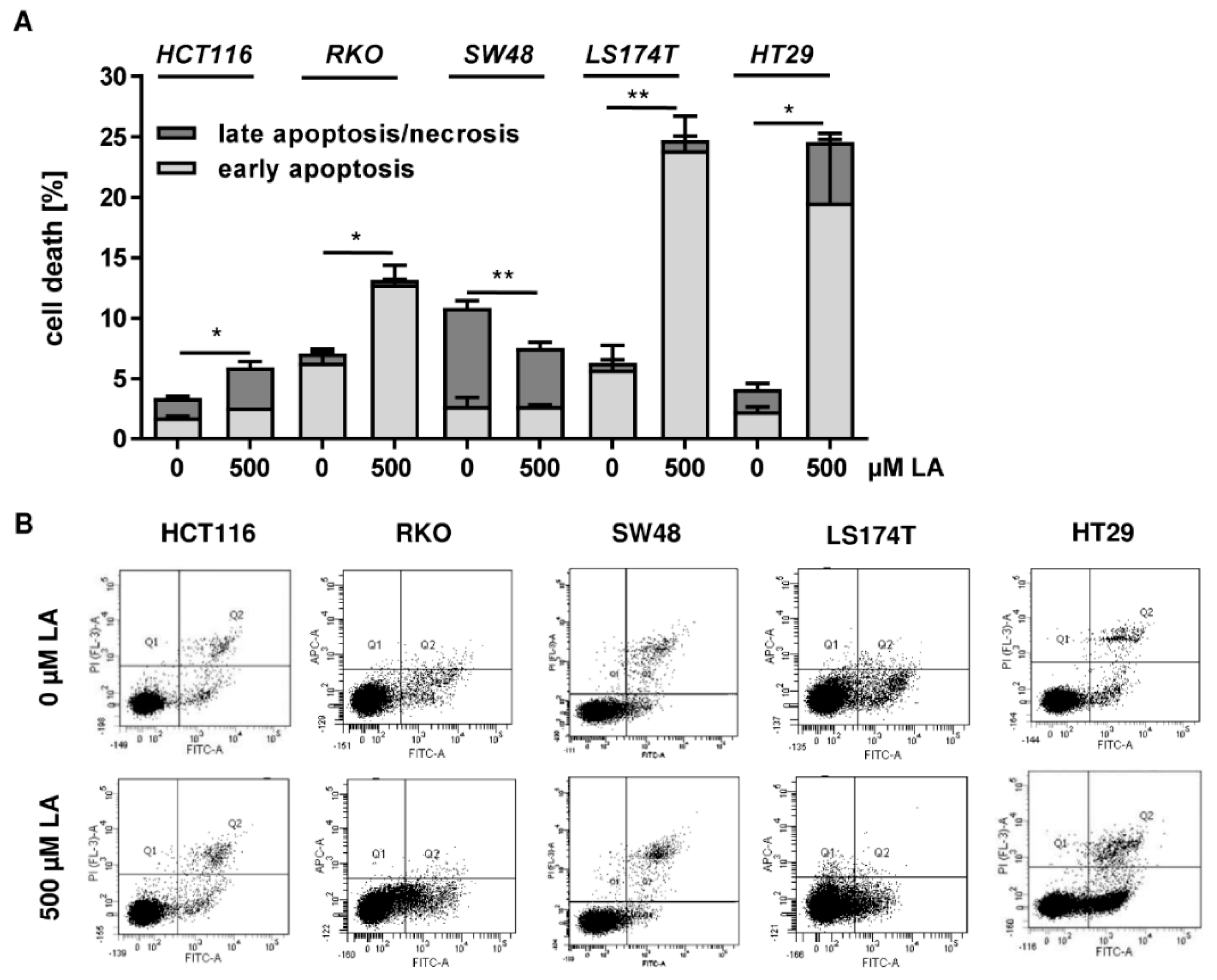
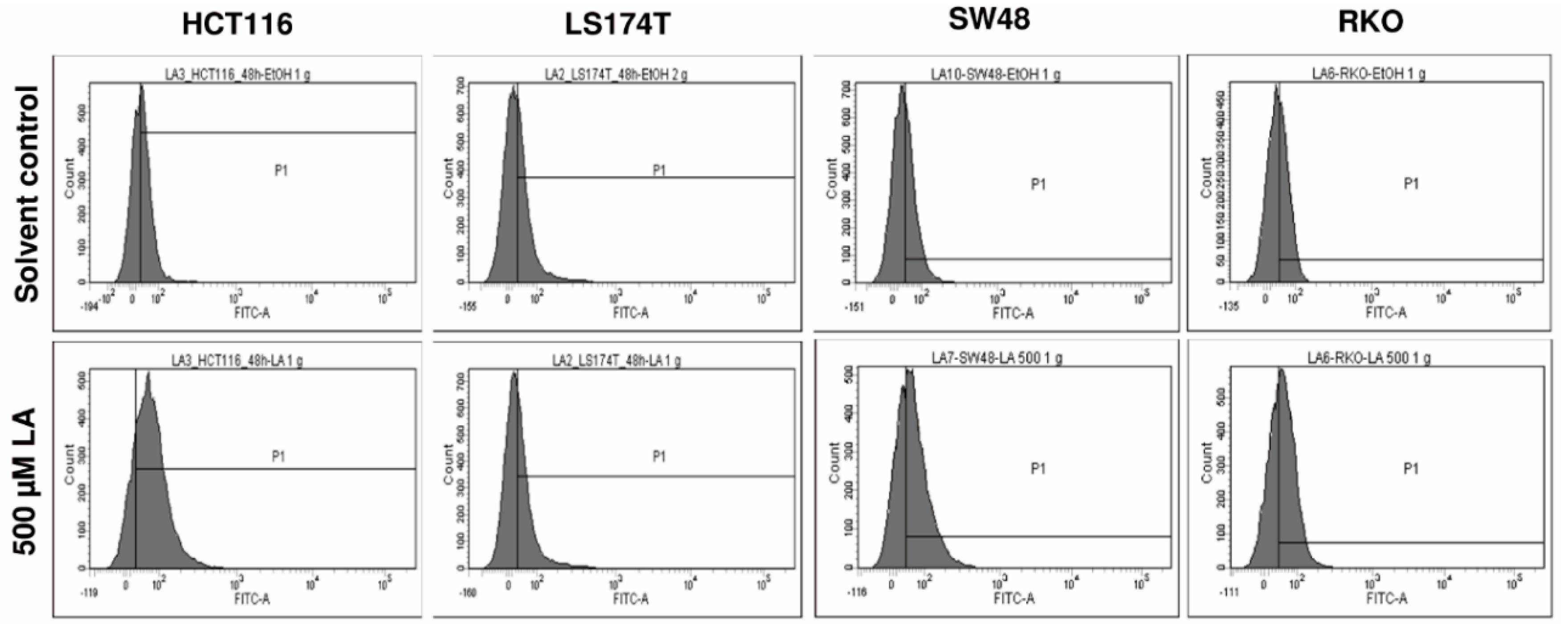
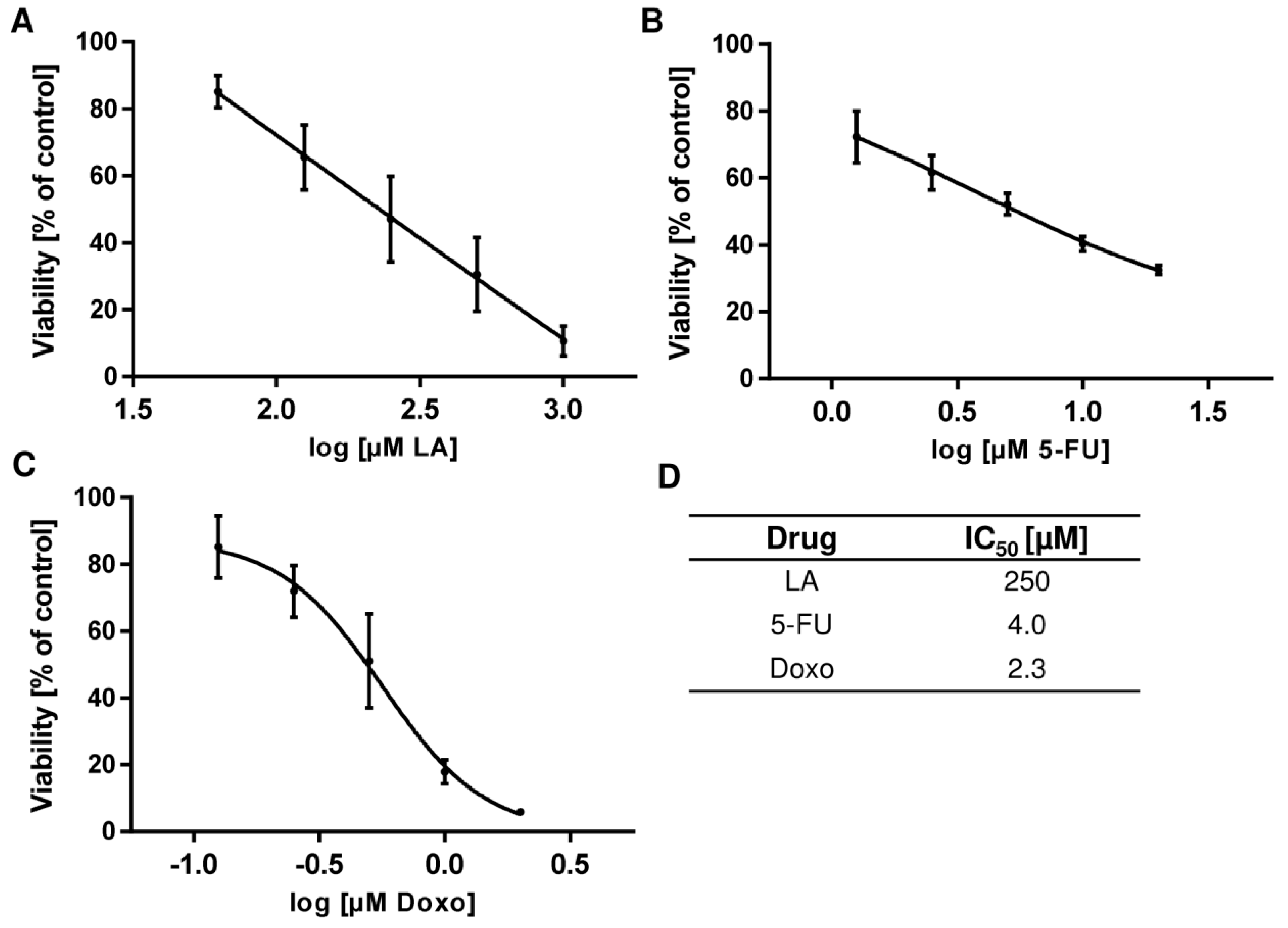
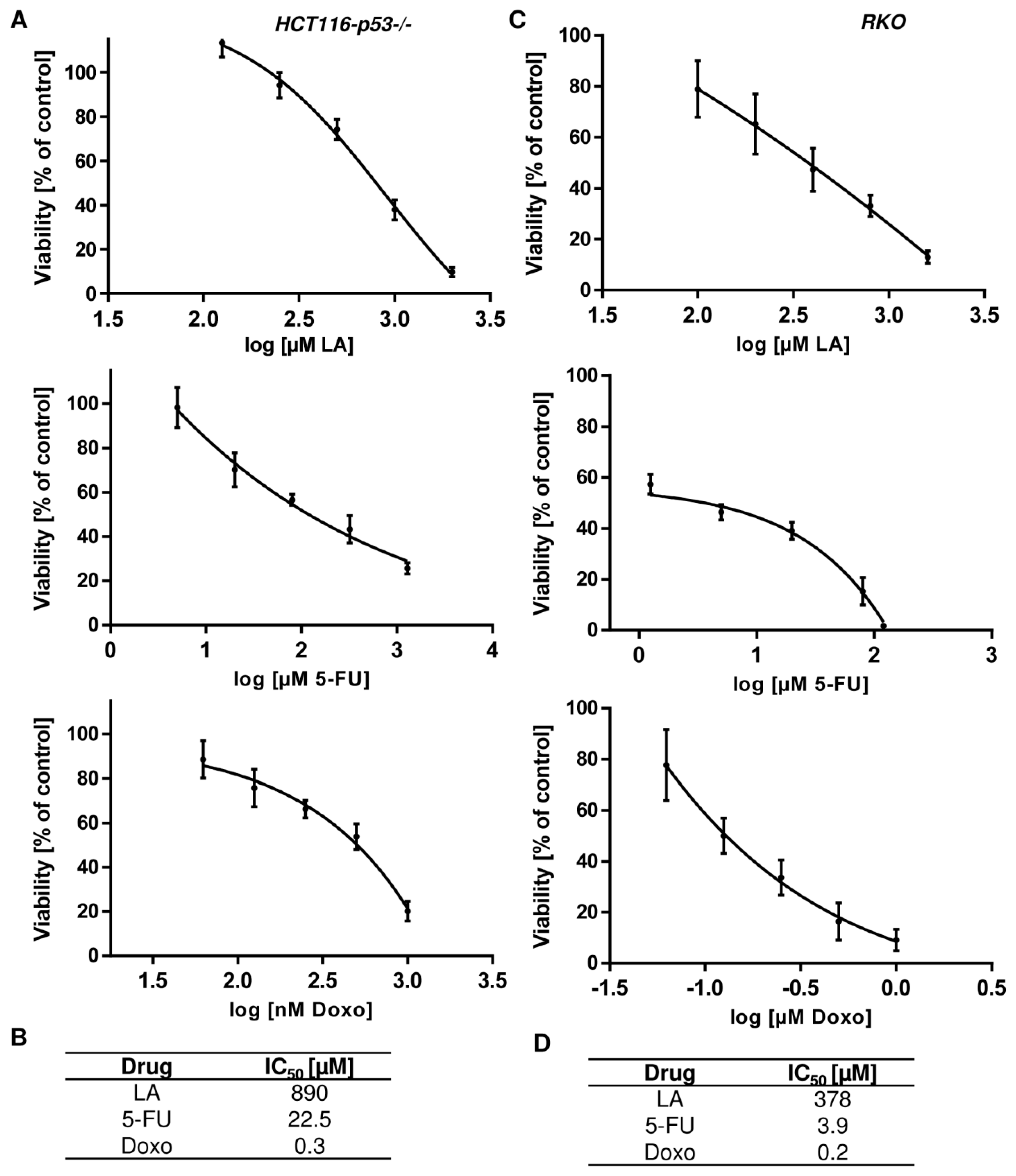
References
- Jones, W.; Li, X.; Qu, Z.-c.; Perriott, L.; Whitesell, R.R.; May, J.M. Uptake, recycling, and antioxidant actions of α-lipoic acid in endothelial cells. Free Radic. Biol. Med. 2002, 33, 83–93. [Google Scholar] [CrossRef]
- Sigel, H.; Prijs, B.; McCormick, D.B.; Shih, J.C. Stability and structure of binary and ternary complexes of α-lipoate and lipoate derivatives with Mn2+, Cu2+, and Zn2+ in solution. Arch. Biochem. Biophys. 1978, 187, 208–214. [Google Scholar] [CrossRef]
- Shay, K.P.; Moreau, R.F.; Smith, E.J.; Smith, A.R.; Hagen, T.M. Alpha-lipoic acid as a dietary supplement: Molecular mechanisms and therapeutic potential. Biochim. Biophys Acta 2009, 1790, 1149–1160. [Google Scholar] [CrossRef] [PubMed]
- Rochette, L.; Ghibu, S.; Richard, C.; Zeller, M.; Cottin, Y.; Vergely, C. Direct and indirect antioxidant properties of α-lipoic acid and therapeutic potential. Mol. Nutr. Food Res. 2013, 57, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Satoh, S.; Shindoh, M.; Min, J.Z.; Toyo’oka, T.; Fukushima, T.; Inagaki, S. Selective and sensitive determination of lipoyllysine (protein-bound alpha-lipoic acid) in biological specimens by high-performance liquid chromatography with fluorescence detection. Anal. Chim. Acta 2008, 618, 210–217. [Google Scholar] [CrossRef] [PubMed]
- Akiba, S.; Matsugo, S.; Packer, L.; Konishi, T. Assay of protein-bound lipoic acid in tissues by a new enzymatic method. Anal. Biochem. 1998, 258, 299–304. [Google Scholar] [CrossRef] [PubMed]
- Jarrett, J.T. The biosynthesis of thiol- and thioether-containing cofactors and secondary metabolites catalyzed by radical S-adenosylmethionine enzymes. J. Biol. Chem. 2015, 290, 3972–3979. [Google Scholar] [CrossRef]
- Reljanovic, M.; Reichel, G.; Rett, K.; Lobisch, M.; Schuette, K.; Möller, W.; Tritschler, H.-J.; Mehnert, H. Treatment of diabetic polyneuropathy with the antioxidant thioctic acid (α -lipoic acid): A two year multicenter randomized double-blind placebo-controlled trial (ALADIN II). Free Radic. Res. 2009, 31, 171–179. [Google Scholar] [CrossRef]
- Dörsam, B.; Fahrer, J. The disulfide compound α-lipoic acid and its derivatives: A novel class of anticancer agents targeting mitochondria. Cancer Lett. 2016, 371, 12–19. [Google Scholar] [CrossRef]
- Abolhassani, M.; Guais, A.; Sanders, E.; Campion, F.; Fichtner, I.; Bonte, J.; Baronzio, G.; Fiorentini, G.; Israël, M.; Schwartz, L. Screening of well-established drugs targeting cancer metabolism: Reproducibility of the efficacy of a highly effective drug combination in mice. Invest. New Drugs 2012, 30, 1331–1342. [Google Scholar] [CrossRef]
- Feuerecker, B.; Pirsig, S.; Seidl, C.; Aichler, M.; Feuchtinger, A.; Bruchelt, G.; Senekowitsch-Schmidtke, R. Lipoic acid inhibits cell proliferation of tumor cells in vitro and in vivo. Cancer Biol. 2012, 13, 1425–1435. [Google Scholar] [CrossRef] [PubMed]
- Jeon, M.J.; Kim, W.G.; Lim, S.; Choi, H.-J.; Sim, S.; Kim, T.Y.; Shong, Y.K.; Kim, W.B. Alpha lipoic acid inhibits proliferation and epithelial mesenchymal transition of thyroid cancer cells. Mol. Cell Endocrinol. 2016, 419, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Göder, A.; Nagel, G.; Kraus, A.; Dörsam, B.; Seiwert, N.; Kaina, B.; Fahrer, J. Lipoic acid inhibits the DNA repair protein O 6-methylguanine-DNA methyltransferase (MGMT) and triggers its depletion in colorectal cancer cells with concomitant autophagy induction. Carcinogenesis 2015, 36, 817–831. [Google Scholar] [CrossRef] [PubMed]
- Zachar, Z.; Marecek, J.; Maturo, C.; Gupta, S.; Stuart, S.D.; Howell, K.; Schauble, A.; Lem, J.; Piramzadian, A.; Karnik, S.; et al. Non-redox-active lipoate derivates disrupt cancer cell mitochondrial metabolism and are potent anticancer agents in vivo. J. Mol. Med. 2011, 89, 1137–1148. [Google Scholar] [CrossRef] [PubMed]
- Dörsam, B.; Göder, A.; Seiwert, N.; Kaina, B.; Fahrer, J. Lipoic acid induces p53-independent cell death in colorectal cancer cells and potentiates the cytotoxicity of 5-fluorouracil. Arch. Toxicol. 2015, 89, 1829–1846. [Google Scholar] [CrossRef] [PubMed]
- Moungjaroen, J.; Nimmannit, U.; Callery, P.S.; Wang, L.; Azad, N.; Lipipun, V.; Chanvorachote, P.; Rojanasakul, Y. Reactive oxygen species mediate caspase activation and apoptosis induced by lipoic acid in human lung epithelial cancer cells through Bcl-2 downregulation. J. Pharm. Exp. 2006, 319, 1062–1069. [Google Scholar] [CrossRef] [PubMed]
- Wenzel, U.; Nickel, A.; Daniel, H. alpha-Lipoic acid induces apoptosis in human colon cancer cells by increasing mitochondrial respiration with a concomitant O2-*-generation. Apoptosis 2005, 10, 359–368. [Google Scholar] [CrossRef]
- Sen, C.K.; Sashwati, R.; Packer, L. Fas mediated apoptosis of human Jurkat T-cells: Intracellular events and potentiation by redox-active alpha-lipoic acid. Cell Death Differ. 1999, 6, 481–491. [Google Scholar] [CrossRef]
- Yoo, T.-H.; Lee, J.-H.; Chun, H.-S.; Chi, S.-G. α-Lipoic acid prevents p53 degradation in colon cancer cells by blocking NF-κB induction of RPS6KA4. Anticancer. Drugs 2013, 24, 555–565. [Google Scholar] [CrossRef]
- Fahrer, J.; Kaina, B. O6-methylguanine-DNA methyltransferase in the defense against N-nitroso compounds and colorectal cancer. Carcinogenesis 2013, 34, 2435–2442. [Google Scholar] [CrossRef]
- Brady, C.A.; Attardi, L.D. p53 at a glance. J. Cell Sci. 2010, 123, 2527–2532. [Google Scholar] [CrossRef] [PubMed]
- Christmann, M.; Kaina, B. Transcriptional regulation of human DNA repair genes following genotoxic stress: Trigger mechanisms, inducible responses and genotoxic adaptation. Nucleic Acids Res. 2013, 41, 8403–8420. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.B.; Schumacher, B. p53 in the DNA-Damage-Repair Process. Cold Spring Harb Perspect Med. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Chen, J. The Cell-Cycle Arrest and Apoptotic Functions of p53 in Tumor Initiation and Progression. Cold Spring Harb Perspect Med. 2016, 6, a026104. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Gu, W. p53 post-translational modification: Deregulated in tumorigenesis. Trends Mol. Med. 2010, 16, 528–536. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.-H.; Kundu, J.K.; Surh, Y.-J. Redox modulation of p53: Mechanisms and functional significance. Mol. Carcinog 2011, 50, 222–234. [Google Scholar] [CrossRef] [PubMed]
- Muller, P.A.J.; Vousden, K.H. p53 mutations in cancer. Nat. Cell Biol 2013, 15, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.; Shirole, N.; Tian, R.; Pal, D.; Sordella, R. The Evolution of TP53 Mutations: From Loss-of-Function to Separation-of-Function Mutants. J. Cancer Biol. Res. 2016, 4. [Google Scholar]
- Olivier, M.; Hollstein, M.; Hainaut, P. TP53 mutations in human cancers: Origins, consequences, and clinical use. Cold Spring Harb Perspect Biol 2010, 2, a001008. [Google Scholar] [CrossRef]
- Gatei, M.; Shkedy, D.; Khanna, K.K.; Uziel, T.; Shiloh, Y.; Pandita, T.K.; Lavin, M.F.; Rotman, G. Ataxia-telangiectasia: Chronic activation of damage-responsive functions is reduced by alpha-lipoic acid. Oncogene 2001, 20, 289–294. [Google Scholar] [CrossRef]
- Simbula, G.; Columbano, A.; Ledda-Columbano, G.M.; Sanna, L.; Deidda, M.; Diana, A.; Pibiri, M. Increased ROS generation and p53 activation in alpha-lipoic acid-induced apoptosis of hepatoma cells. Apoptosis 2007, 12, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Choi, S.K.; Choi, Y.; Moon, H.-S. AMPK/p53 Axis Is Essential for α-Lipoic Acid-Regulated Metastasis in Human and Mouse Colon Cancer Cells. J. Investig. Med. 2015, 63, 882–885. [Google Scholar] [CrossRef] [PubMed]
- Seiwert, N.; Neitzel, C.; Stroh, S.; Frisan, T.; Audebert, M.; Toulany, M.; Kaina, B.; Fahrer, J. AKT2 suppresses pro-survival autophagy triggered by DNA double-strand breaks in colorectal cancer cells. Cell Death Dis. 2017, 8, e3019. [Google Scholar] [CrossRef] [PubMed]
- Mimmler, M.; Peter, S.; Kraus, A.; Stroh, S.; Nikolova, T.; Seiwert, N.; Hasselwander, S.; Neitzel, C.; Haub, J.; Monien, B.H.; et al. DNA damage response curtails detrimental replication stress and chromosomal instability induced by the dietary carcinogen PhIP. Nucleic Acids Res. 2016, 44, 10259–10276. [Google Scholar] [CrossRef] [PubMed]
- Christmann, M.; Boisseau, C.; Kitzinger, R.; Berac, C.; Allmann, S.; Sommer, T.; Aasland, D.; Kaina, B.; Tomicic, M.T. Adaptive upregulation of DNA repair genes following benzo(a)pyrene diol epoxide protects against cell death at the expense of mutations. Nucleic Acids Res. 2016, 44, 10727–10743. [Google Scholar] [CrossRef]
- Fahrer, J.; Huelsenbeck, J.; Jaurich, H.; Dörsam, B.; Frisan, T.; Eich, M.; Roos, W.P.; Kaina, B.; Fritz, G. Cytolethal distending toxin (CDT) is a radiomimetic agent and induces persistent levels of DNA double-strand breaks in human fibroblasts. DNA Repair (Amst) 2014, 18, 31–43. [Google Scholar] [CrossRef] [PubMed]
- Vermes, I.; Haanen, C.; Steffens-Nakken, H.; Reutelingsperger, C. A novel assay for apoptosis. Flow cytometric detection of phosphatidylserine expression on early apoptotic cells using fluorescein labeled Annexin V. J. Immunol. Methods 1995, 184, 39–51. [Google Scholar] [CrossRef]
- Rieger, A.M.; Hall, B.E.; Le Luong, T.; Schang, L.M.; Barreda, D.R. Conventional apoptosis assays using propidium iodide generate a significant number of false positives that prevent accurate assessment of cell death. J. Immunol. Methods 2010, 358, 81–92. [Google Scholar] [CrossRef]
- Dörsam, B.; Wu, C.-F.; Efferth, T.; Kaina, B.; Fahrer, J. The eucalyptus oil ingredient 1,8-cineol induces oxidative DNA damage. Arch. Toxicol. 2015, 89, 797–805. [Google Scholar] [CrossRef]
- Chou, T.-C. Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharm. Rev. 2006, 58, 621–681. [Google Scholar] [CrossRef]
- Ahmed, D.; Eide, P.W.; Eilertsen, I.A.; Danielsen, S.A.; Eknæs, M.; Hektoen, M.; Lind, G.E.; Lothe, R.A. Epigenetic and genetic features of 24 colon cancer cell lines. Oncogenesis 2013, 2, e71. [Google Scholar] [CrossRef] [PubMed]
- Subbarao Sreedhar, A.; Kalmár, É.; Csermely, P.; Shen, Y.-F. Hsp90 isoforms: Functions, expression and clinical importance. FEBS Lett. 2004, 562, 11–15. [Google Scholar] [CrossRef]
- Dörsam, B.; Seiwert, N.; Foersch, S.; Stroh, S.; Nagel, G.; Begaliew, D.; Diehl, E.; Kraus, A.; McKeague, M.; Minneker, V.; et al. PARP-1 protects against colorectal tumor induction, but promotes inflammation-driven colorectal tumor progression. Proc. Natl. Acad Sci. USA 2018, 115, E4061–E4070. [Google Scholar] [CrossRef]
- Abrahams, S.; Haylett, W.L.; Johnson, G.; Carr, J.A.; Bardien, S. Antioxidant effects of curcumin in models of neurodegeneration, aging, oxidative and nitrosative stress: A review. Neuroscience 2019, 406, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Glick, D.; Barth, S.; Macleod, K.F. Autophagy: Cellular and molecular mechanisms. J. Pathol. 2010, 221, 3–12. [Google Scholar] [CrossRef]
- Meijer, A.J.; Codogno, P. Regulation and role of autophagy in mammalian cells. Int. J. Biochem. Cell Biol. 2004, 36, 2445–2462. [Google Scholar] [CrossRef]
- Klionsky, D.J. Autophagy: From phenomenology to molecular understanding in less than a decade. Nat. Rev. Mol. Cell Biol. 2007, 8, 931–937. [Google Scholar] [CrossRef]
- Mizushima, N.; Yoshimori, T.; Ohsumi, Y. The role of Atg proteins in autophagosome formation. Annu Rev. Cell Dev. Biol. 2011, 27, 107–132. [Google Scholar] [CrossRef]
- Tebay, L.E.; Robertson, H.; Durant, S.T.; Vitale, S.R.; Penning, T.M.; Dinkova-Kostova, A.T.; Hayes, J.D. Mechanisms of activation of the transcription factor Nrf2 by redox stressors, nutrient cues, and energy status and the pathways through which it attenuates degenerative disease. Free Radic. Biol. Med. 2015, 88, 108–146. [Google Scholar] [CrossRef]
- Lau, A.; Tian, W.; Whitman, S.A.; Zhang, D.D. The predicted molecular weight of Nrf2: It is what it is not. Antioxid. Redox Signal. 2013, 18, 91–93. [Google Scholar] [CrossRef]
- Singh, A.; Venkannagari, S.; Oh, K.H.; Zhang, Y.-Q.; Rohde, J.M.; Liu, L.; Nimmagadda, S.; Sudini, K.; Brimacombe, K.R.; Gajghate, S.; et al. Small Molecule Inhibitor of NRF2 Selectively Intervenes Therapeutic Resistance in KEAP1-Deficient NSCLC Tumors. ACS Chem. Biol. 2016, 11, 3214–3225. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Xi, Z.; Xue, Y.; Ren, J.; Sun, Y.; Wang, B.; Zhong, Z.; Yang, G.-Y.; Sun, Q.; Bian, L. Hemoglobin pretreatment endows rat cortical astrocytes resistance to hemin-induced toxicity via Nrf2/HO-1 pathway. Exp. Cell Res. 2017, 361, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Bard, J.A.M.; Goodall, E.A.; Greene, E.R.; Jonsson, E.; Dong, K.C.; Martin, A. Structure and Function of the 26S Proteasome. Annu. Rev. Biochem. 2018, 87, 697–724. [Google Scholar] [CrossRef] [PubMed]
- Wade, M.; Li, Y.-C.; Wahl, G.M. MDM2, MDMX and p53 in oncogenesis and cancer therapy. Nat. Rev. Cancer 2013, 13, 83–96. [Google Scholar] [CrossRef] [PubMed]
- Vassilev, L.T.; Vu, B.T.; Graves, B.; Carvajal, D.; Podlaski, F.; Filipovic, Z.; Kong, N.; Kammlott, U.; Lukacs, C.; Klein, C.; et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 2004, 303, 844–848. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, D.; Atkin, W.; Lenz, H.-J.; Lynch, H.T.; Minsky, B.; Nordlinger, B.; Starling, N. Colorectal cancer. Lancet 2010, 375, 1030–1047. [Google Scholar] [CrossRef]
- Ju, J.; Schmitz, J.C.; Song, B.; Kudo, K.; Chu, E. Regulation of p53 expression in response to 5-fluorouracil in human cancer RKO cells. Clin. Cancer Res. 2007, 13, 4245–4251. [Google Scholar] [CrossRef]
- Rodrigues, N.R.; Rowan, A.; Smith, M.E.; Kerr, I.B.; Bodmer, W.F.; Gannon, J.V.; Lane, D.P. p53 mutations in colorectal cancer. Proc. Natl. Acad. Sci. USA 1990, 87, 7555–7559. [Google Scholar] [CrossRef]
- Rivlin, N.; Brosh, R.; Oren, M.; Rotter, V. Mutations in the p53 Tumor Suppressor Gene: Important Milestones at the Various Steps of Tumorigenesis. Genes Cancer 2011, 2, 466–474. [Google Scholar] [CrossRef]
- Kalo, E.; Kogan-Sakin, I.; Solomon, H.; Bar-Nathan, E.; Shay, M.; Shetzer, Y.; Dekel, E.; Goldfinger, N.; Buganim, Y.; Stambolsky, P.; et al. Mutant p53R273H attenuates the expression of phase 2 detoxifying enzymes and promotes the survival of cells with high levels of reactive oxygen species. J. Cell Sci. 2012, 125, 5578–5586. [Google Scholar] [CrossRef]
- Muller, P.A.J.; Caswell, P.T.; Doyle, B.; Iwanicki, M.P.; Tan, E.H.; Karim, S.; Lukashchuk, N.; Gillespie, D.A.; Ludwig, R.L.; Gosselin, P.; et al. Mutant p53 drives invasion by promoting integrin recycling. Cell 2009, 139, 1327–1341. [Google Scholar] [CrossRef] [PubMed]
- Yeudall, W.A.; Vaughan, C.A.; Miyazaki, H.; Ramamoorthy, M.; Choi, M.-Y.; Chapman, C.G.; Wang, H.; Black, E.; Bulysheva, A.A.; Deb, S.P.; et al. Gain-of-function mutant p53 upregulates CXC chemokines and enhances cell migration. Carcinogenesis 2012, 33, 442–451. [Google Scholar] [CrossRef] [PubMed]
- Kruse, J.-P.; Gu, W. Modes of p53 regulation. Cell 2009, 137, 609–622. [Google Scholar] [CrossRef] [PubMed]
- Watson, J.L.; Hill, R.; Yaffe, P.B.; Greenshields, A.; Walsh, M.; Lee, P.W.; Giacomantonio, C.A.; Hoskin, D.W. Curcumin causes superoxide anion production and p53-independent apoptosis in human colon cancer cells. Cancer Lett. 2010, 297, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Danieli, A.; Martens, S. p62-mediated phase separation at the intersection of the ubiquitin-proteasome system and autophagy. J. Cell Sci. 2018, 131. [Google Scholar] [CrossRef]
- Lamark, T.; Svenning, S.; Johansen, T. Regulation of selective autophagy: The p62/SQSTM1 paradigm. Essays Biochem. 2017, 61, 609–624. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Lamark, T.; Sjøttem, E.; Larsen, K.B.; Awuh, J.A.; Øvervatn, A.; McMahon, M.; Hayes, J.D.; Johansen, T. p62/SQSTM1 is a target gene for transcription factor NRF2 and creates a positive feedback loop by inducing antioxidant response element-driven gene transcription. J. Biol. Chem. 2010, 285, 22576–22591. [Google Scholar] [CrossRef]
- Komatsu, M.; Kurokawa, H.; Waguri, S.; Taguchi, K.; Kobayashi, A.; Ichimura, Y.; Sou, Y.-S.; Ueno, I.; Sakamoto, A.; Tong, K.I.; et al. The selective autophagy substrate p62 activates the stress responsive transcription factor Nrf2 through inactivation of Keap1. Nat. Cell Biol. 2010, 12, 213–223. [Google Scholar] [CrossRef]
- Shay, K.P.; Michels, A.J.; Li, W.; Kong, A.-N.T.; Hagen, T.M. Cap-independent Nrf2 translation is part of a lipoic acid-stimulated detoxification stress response. Biochim Biophys Acta 2012, 1823, 1102–1109. [Google Scholar] [CrossRef]
- Lii, C.-K.; Liu, K.-L.; Cheng, Y.-P.; Lin, A.-H.; Chen, H.-W.; Tsai, C.-W. Sulforaphane and alpha-lipoic acid upregulate the expression of the pi class of glutathione S-transferase through c-jun and Nrf2 activation. J. Nutr. 2010, 140, 885–892. [Google Scholar] [CrossRef]
- Pilar Valdecantos, M.; Prieto-Hontoria, P.L.; Pardo, V.; Módol, T.; Santamaría, B.; Weber, M.; Herrero, L.; Serra, D.; Muntané, J.; Cuadrado, A.; et al. Essential role of Nrf2 in the protective effect of lipoic acid against lipoapoptosis in hepatocytes. Free Radic. Biol. Med. 2015, 84, 263–278. [Google Scholar] [CrossRef] [PubMed]
- Pickering, A.M.; Linder, R.A.; Zhang, H.; Forman, H.J.; Davies, K.J.A. Nrf2-dependent induction of proteasome and Pa28αβ regulator are required for adaptation to oxidative stress. J. Biol. Chem. 2012, 287, 10021–10031. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Martin, J.D.; Zhang, H.; Auger, K.R.; Ho, T.F.; Kirkpatrick, R.B.; Grooms, M.H.; Johanson, K.O.; Tummino, P.J.; Copeland, R.A.; et al. A second p53 binding site in the central domain of Mdm2 is essential for p53 ubiquitination. Biochemistry 2006, 45, 9238–9245. [Google Scholar] [CrossRef] [PubMed]
- Chao, C.C.-K. Mechanisms of p53 degradation. Clin. Chim. Acta 2015, 438, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Scotcher, J.; Clarke, D.J.; Mackay, C.L.; Hupp, T.; Sadler, P.J.; Langridge-Smith, P.R.R. Redox regulation of tumor suppressor protein p53: Identification of the sites of hydrogen peroxide oxidation and glutathionylation. Chem. Sci. 2013, 4, 1257. [Google Scholar] [CrossRef]
- Hainaut, P.; Milner, J. Redox modulation of p53 conformation and sequence-specific DNA binding in vitro. Cancer Res. 1993, 53, 4469–4473. [Google Scholar] [PubMed]
- Nomura, T.; Kamada, R.; Ito, I.; Sakamoto, K.; Chuman, Y.; Ishimori, K.; Shimohigashi, Y.; Sakaguchi, K. Probing phenylalanine environments in oligomeric structures with pentafluorophenylalanine and cyclohexylalanine. Biopolymers 2011, 95, 410–419. [Google Scholar] [CrossRef] [PubMed]
- Scotcher, J.; Clarke, D.J.; Weidt, S.K.; Mackay, C.L.; Hupp, T.R.; Sadler, P.J.; Langridge-Smith, P.R.R. Identification of two reactive cysteine residues in the tumor suppressor protein p53 using top-down FTICR mass spectrometry. J. Am. Soc. Mass Spectrom. 2011, 22, 888–897. [Google Scholar] [CrossRef]
- Velu, C.S.; Niture, S.K.; Doneanu, C.E.; Pattabiraman, N.; Srivenugopal, K.S. Human p53 is inhibited by glutathionylation of cysteines present in the proximal DNA-binding domain during oxidative stress. Biochemistry 2007, 46, 7765–7780. [Google Scholar] [CrossRef]
- Paranjpe, A.; Srivenugopal, K.S. Degradation of NF-κB, p53 and other regulatory redox-sensitive proteins by thiol-conjugating and -nitrosylating drugs in human tumor cells. Carcinogenesis 2013, 34, 990–1000. [Google Scholar] [CrossRef]
- Puchsaka, P.; Chaotham, C.; Chanvorachote, P. α-Lipoic acid sensitizes lung cancer cells to chemotherapeutic agents and anoikis via integrin β1/β3 downregulation. Int. J. Oncol. 2016, 49, 1445–1456. [Google Scholar] [CrossRef] [PubMed]
| qPCR Primer | Sequence (5′–3′) |
|---|---|
| ACTB-real-up | TGGCATCCACGAAACTACC |
| ACTB-real-low | GTGTTGGCGTACAGGTCTT |
| FASR-real-up | TTATCTGATGTTGACTTGAGTAA |
| FASR-real-low | GGCTTCATTGACACCATT |
| GADD45a-real-up | ATCTCCCTGAACGGTGAT |
| GADD45a-real-low | TGTAATCCTTGCATCAGTGT |
| GAPDH-real-up | CATGAGAAGTATGACAACAG |
| GAPDH-real-low | ATGAGTCCTTCCACGAT |
| MDM2-real-up | ATCTTGATGCTGGTGTAA |
| MDM2-real-low | AGGCTATAATCTTCTGAGTC |
| NOXA-real-up | TCTTCGGTCACTACACAAC |
| NOXA-real-low | CCAACAGGAACACATTGAAT |
| p21-real-up | ACCATGTCAGAACCGGCTGGG |
| p21-real-low | TGGGCGGATTAGGGCTTC |
| PUMA-real-up | TAAGGATGGAAAGTGTAG |
| PUMA-real-low | TTCAGTTTCTCATTGTTAC |
| p53-real-up | AGCACTAAGCGAGCACTG |
| p53-real-low | ACGGATCTGAAGGGTGAAA |
| Cell Line/CI | LA + 5-FU | LA + Doxo |
|---|---|---|
| HCT116 | 0.67 | 0.55 |
| HCT116-p53−/− | 1.22 | 1.29 |
| RKO | 0.84 | 0.59 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Neitzel, C.; Seiwert, N.; Göder, A.; Diehl, E.; Weber, C.; Nagel, G.; Stroh, S.; Rasenberger, B.; Christmann, M.; Fahrer, J. Lipoic Acid Synergizes with Antineoplastic Drugs in Colorectal Cancer by Targeting p53 for Proteasomal Degradation. Cells 2019, 8, 794. https://doi.org/10.3390/cells8080794
Neitzel C, Seiwert N, Göder A, Diehl E, Weber C, Nagel G, Stroh S, Rasenberger B, Christmann M, Fahrer J. Lipoic Acid Synergizes with Antineoplastic Drugs in Colorectal Cancer by Targeting p53 for Proteasomal Degradation. Cells. 2019; 8(8):794. https://doi.org/10.3390/cells8080794
Chicago/Turabian StyleNeitzel, Carina, Nina Seiwert, Anja Göder, Erika Diehl, Carina Weber, Georg Nagel, Svenja Stroh, Birgit Rasenberger, Markus Christmann, and Jörg Fahrer. 2019. "Lipoic Acid Synergizes with Antineoplastic Drugs in Colorectal Cancer by Targeting p53 for Proteasomal Degradation" Cells 8, no. 8: 794. https://doi.org/10.3390/cells8080794
APA StyleNeitzel, C., Seiwert, N., Göder, A., Diehl, E., Weber, C., Nagel, G., Stroh, S., Rasenberger, B., Christmann, M., & Fahrer, J. (2019). Lipoic Acid Synergizes with Antineoplastic Drugs in Colorectal Cancer by Targeting p53 for Proteasomal Degradation. Cells, 8(8), 794. https://doi.org/10.3390/cells8080794






