Characterization of Lipid and Lipid Droplet Metabolism in Human HCC
Abstract
1. Introduction
2. Materials and Methods
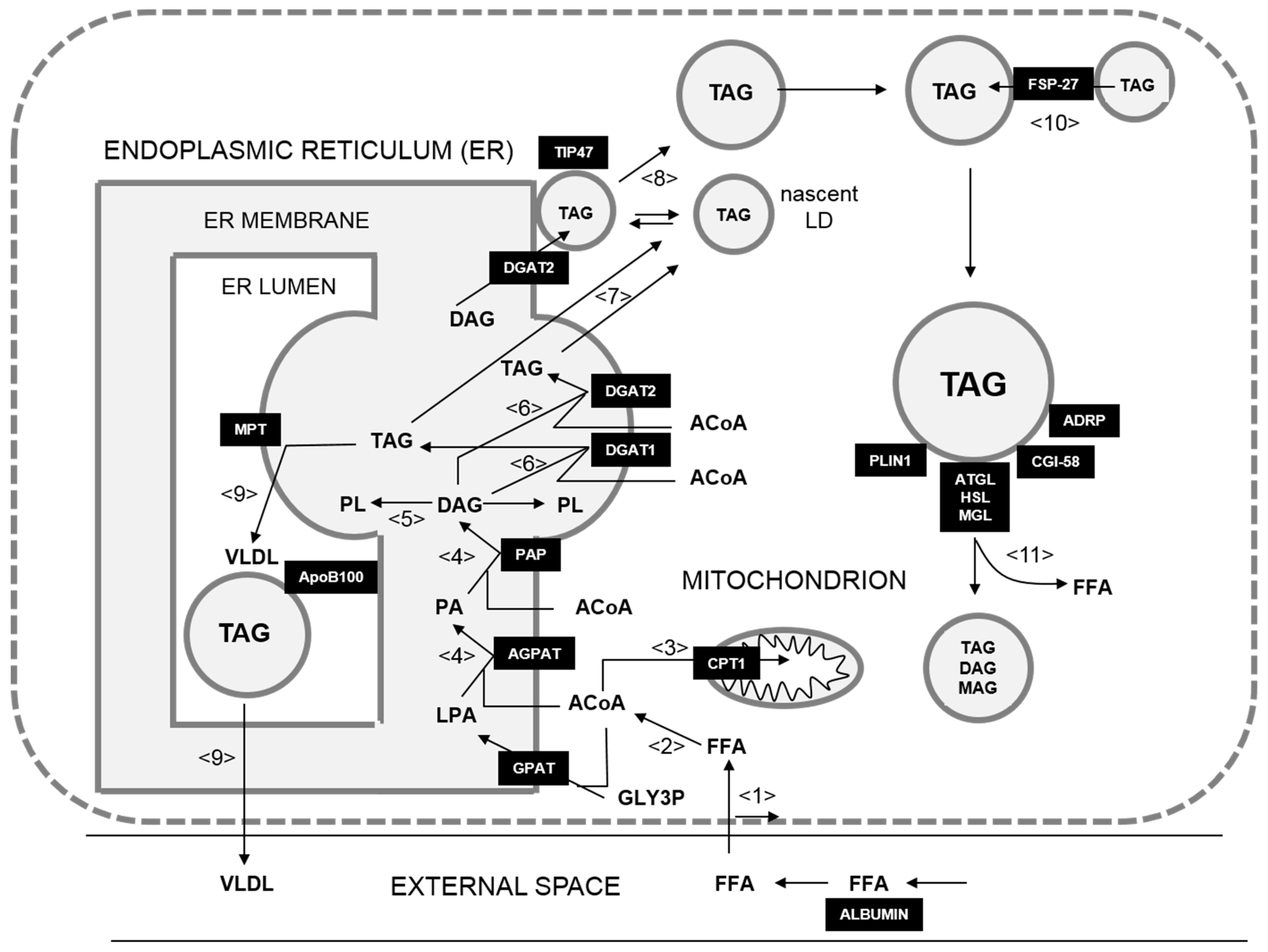
2.1. Kinetic Model of Hepatic LD Metabolism
2.2. Boundary Conditions
2.3. Calibration of the Basic Model to Tumor-Specific Instantiations
2.4. Acquisition of Tissue Samples
2.5. Quantitative Proteomics of Tissue Samples
2.6. Mapping of Protein Intensities Onto Model Processes
3. Results
3.1. Histological Characterization of Tissue Samples
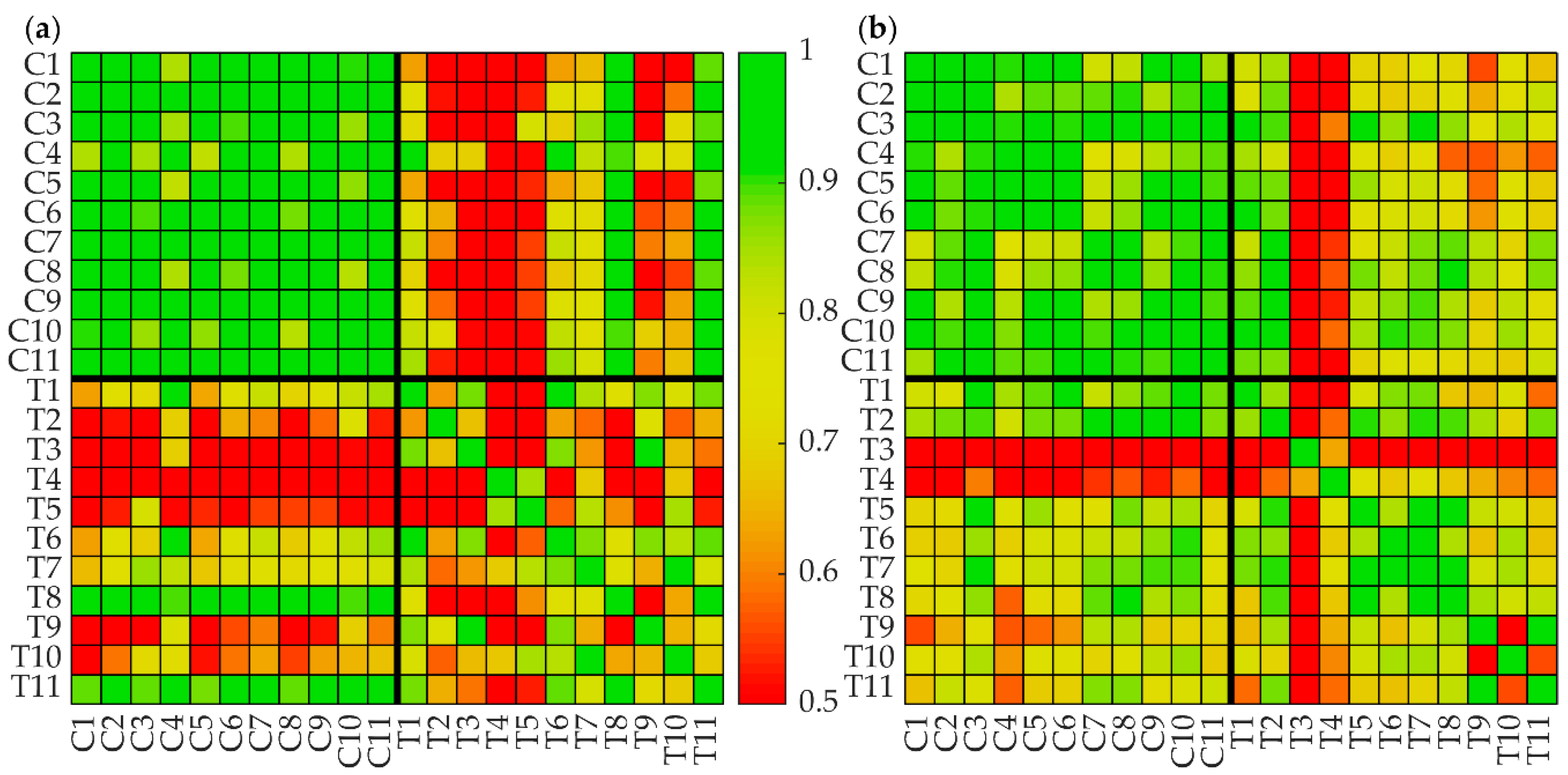
3.2. Protein Intensity Profiles of Tumors and Noncancerous Tissue
3.3. Tumor-Specific Calibration of the Kinetic Model
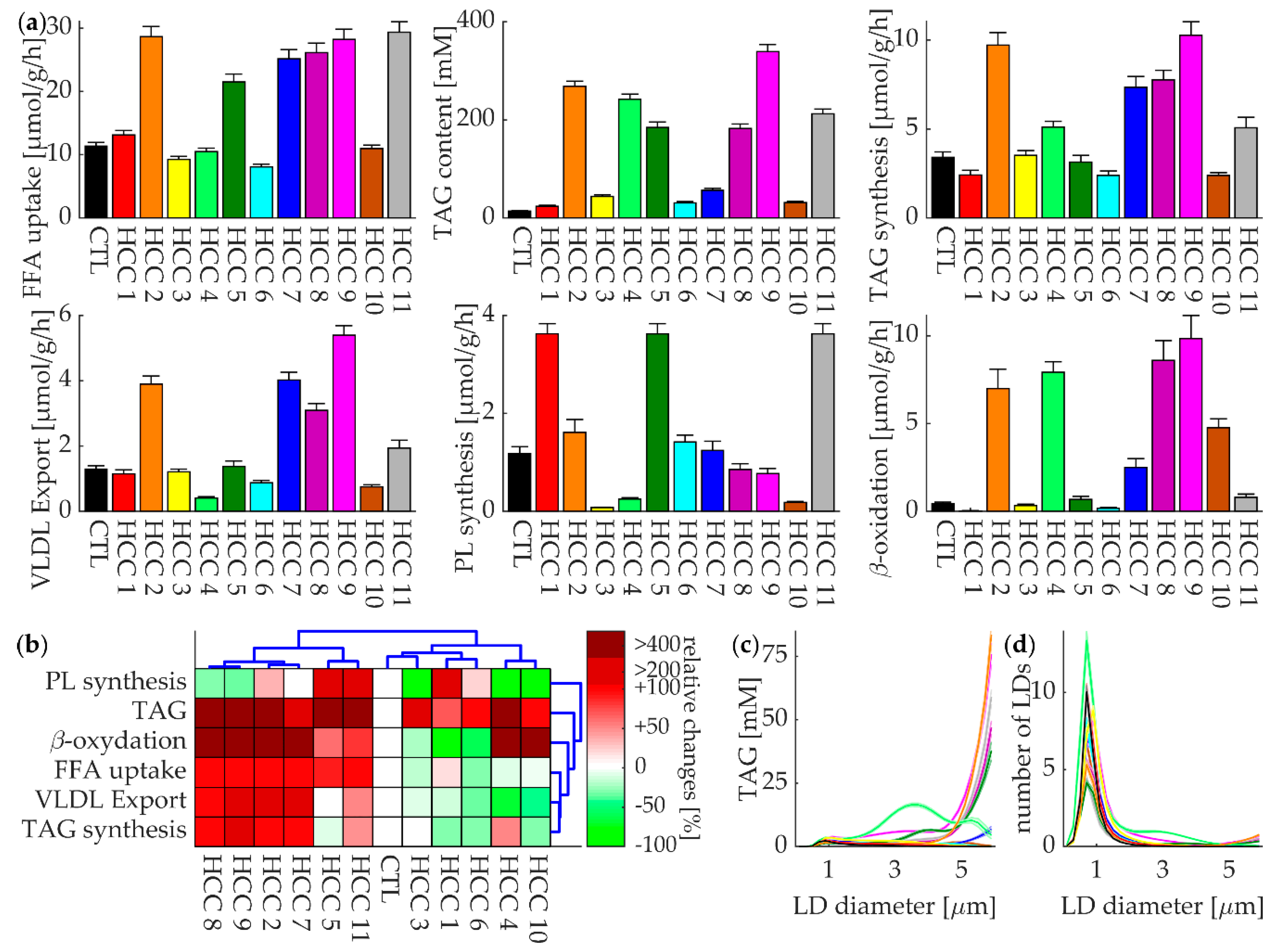
3.4. Lipid Metabolism of Individual HCCs at Normal Plasma Profile of FFAs
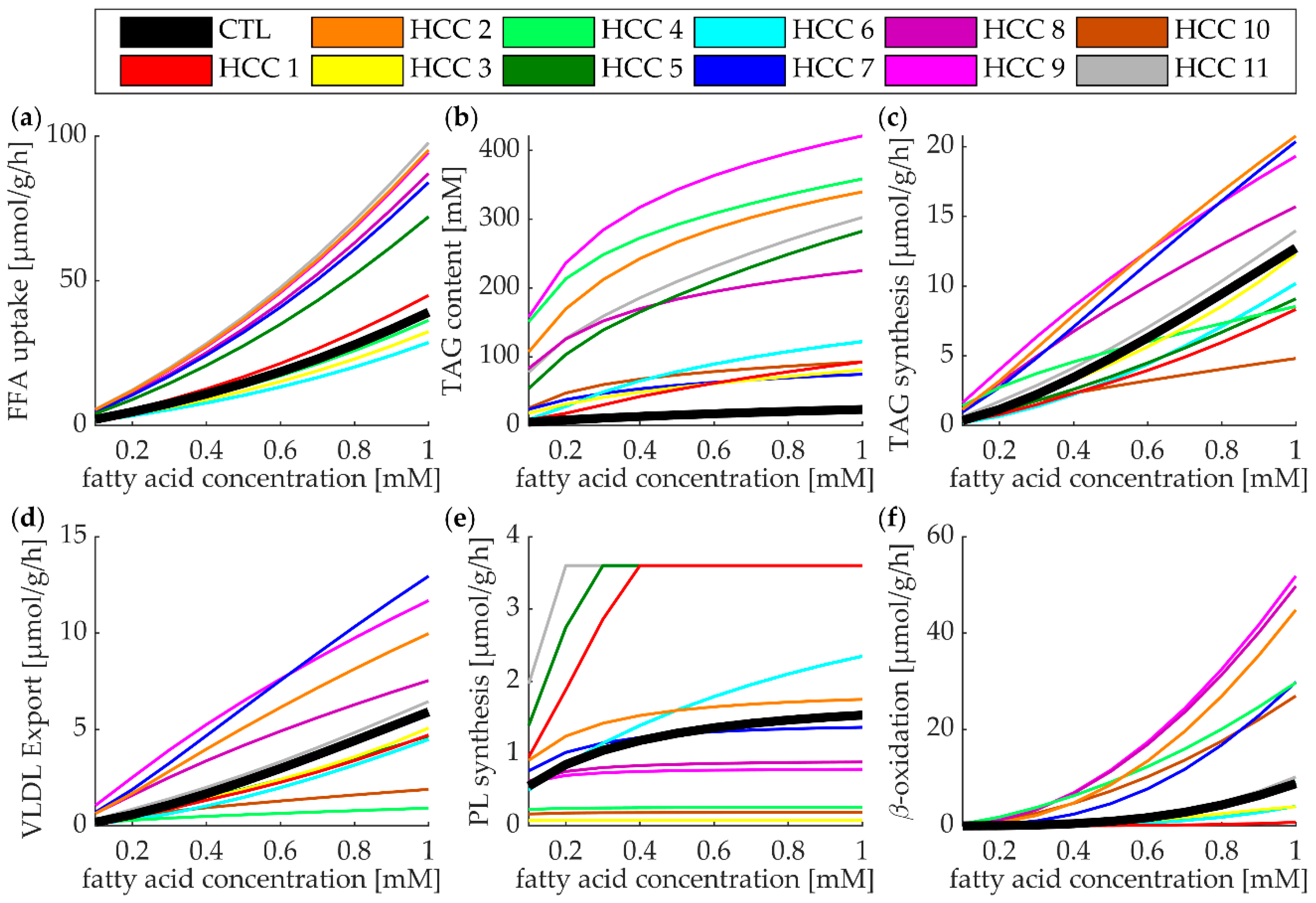
3.5. Lipid Metabolism of Individual HCCs at High Plasma Profile of FFAs
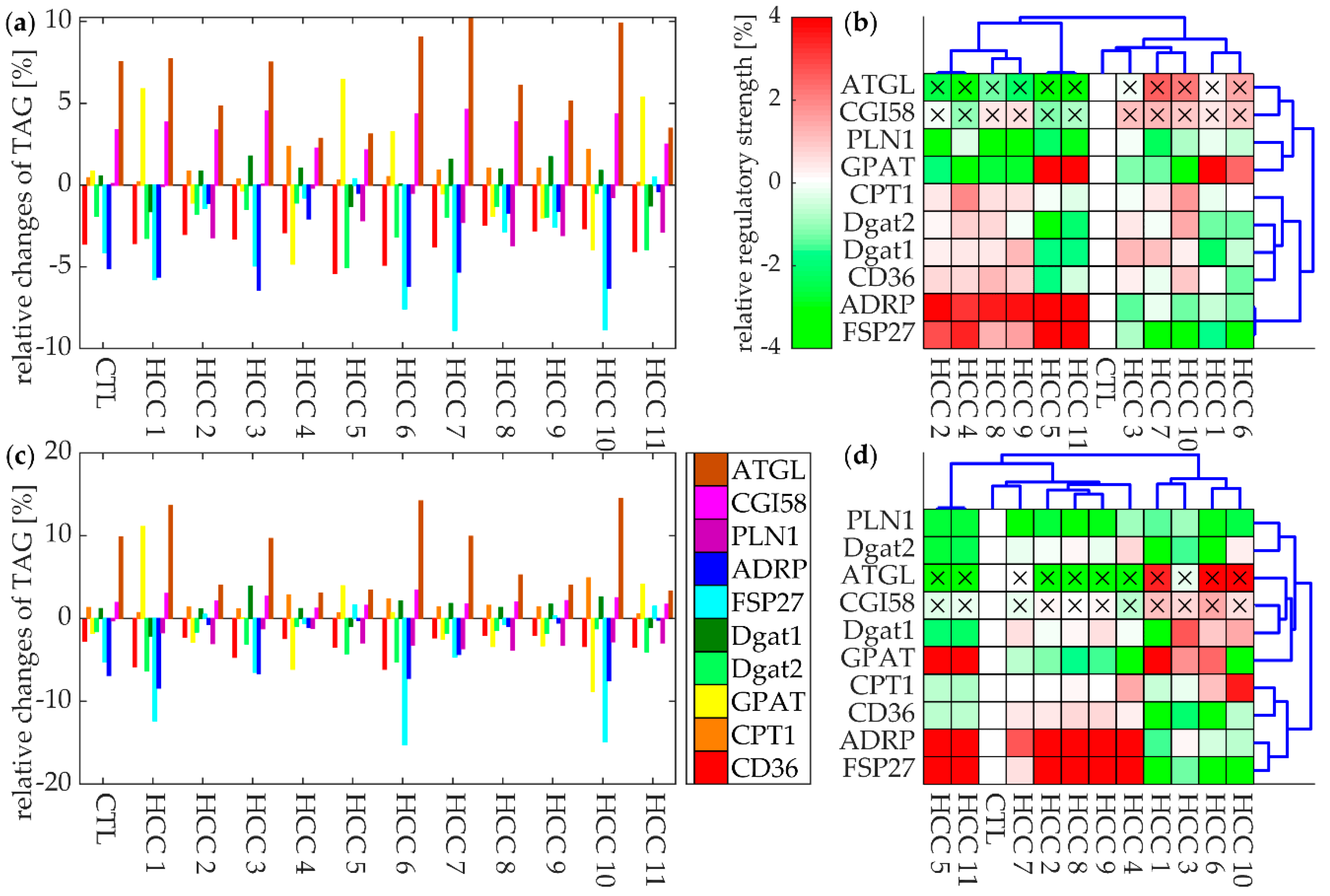
3.6. The Regulatory Impact of Enzymes and RSPs on the TAG Content of Tumors
4. Discussion
4.1. Tissue Metabolism In Vivo: Detailed Insights by Means of a Computational Approach
4.2. Main Finding
4.3. Comparing our Findings with Known Metabolic Features of HCC
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- El-Serag, H.B.; Rudolph, L. Hepatocellular carcinoma: Epidemiology and molecular carcinogenesis. Gastroenterology 2007, 132, 2557–2576. [Google Scholar] [CrossRef] [PubMed]
- Petta, S.; Craxi, A. Hepatocellular Carcinoma and Non-Alcoholic Fatty Liver Disease: From a Clinical to a Molecular Association. Curr. Pharm. Des. 2010, 16, 741–752. [Google Scholar] [CrossRef] [PubMed]
- Cholankeril, G.; Patel, R.; Khurana, S.; Satapathy, S.K. Hepatocellular carcinoma in non-alcoholic steatohepatitis: Current knowledge and implications for management. World J. Hepatol. 2017, 9, 533–543. [Google Scholar]
- Lonardo, A.; Ballestri, S.; Marchesini, G.; Angulo, P.; Loria, P. Nonalcoholic fatty liver disease: A precursor of the metabolic syndrome. Dig. Liver Dis. 2015, 47, 181–190. [Google Scholar] [CrossRef]
- Guri, Y.; Colombi, M.; Dazert, E.; Hindupur, S.K.; Roszik, J.; Moes, S.; Jenoe, P.; Heim, M.H.; Riezman, I.; Riezman, H.; et al. mTORC2 Promotes Tumorigenesis via Lipid Synthesis. Cancer Cell 2017, 32, 807–823. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, H.; Hayata, Y.; Kawamura, S.; Yamada, T.; Fujiwara, N.; Koike, K. Lipid Metabolic Reprogramming in Hepatocellular Carcinoma. Cancers 2018, 10, 447. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.D.; Han, J.; Xing, H.; Zhang, H.; Li, Z.; Liang Li, C.; Dai, S.; Wu, M.; Shen, F.; Yang, T. Dysregulated fatty acid metabolism in hepatocellular carcinoma. Hepatic Oncol. 2016, 3, 241–251. [Google Scholar] [CrossRef]
- Muto, J.; Shirabe, K.; Sugimachi, K.; Maehara, Y. Review of angiogenesis in hepatocellular carcinoma. Hepatol. Res. 2015, 45, 1–9. [Google Scholar] [PubMed]
- Bensaad, K.; Favaro, E.; Lewis, C.A.; Peck, B.; Lord, S.; Collins, J.M.; Pinnick, K.E.; Wigfield, S.; Buffa, F.M.; Li, J.L.; et al. Fatty Acid Uptake and Lipid Storage Induced by HIF-1 alpha Contribute to Cell Growth and Survival after Hypoxia-Reoxygenation. Cell Rep. 2014, 9, 349–365. [Google Scholar] [CrossRef] [PubMed]
- Qiu, B.; Ackerman, D.; Sanchez, D.J.; Li, B.; Ochocki, J.D.; Grazioli, A.; Bobrovnikova-Marjon, E.; Diehl, J.A.; Keith, B.; Simon, M.C. HIF2 alpha-Dependent Lipid Storage Promotes Endoplasmic Reticulum Homeostasis in Clear-Cell Renal Cell Carcinoma. Cancer Discov. 2015, 5, 652–667. [Google Scholar] [CrossRef]
- Koizume, S.; Miyagi, Y. Lipid Droplets: A Key Cellular Organelle Associated with Cancer Cell Survival under Normoxia and Hypoxia. Int. J. Mol. Sci. 2016, 17, 1430. [Google Scholar] [CrossRef]
- Reid, B.N.; Ables, G.P.; Otlivanchik, O.A.; Schoiswohl, G.; Zechner, R.; Blaner, W.S.; Goldberg, I.J.; Schwabe, R.F.; Chua, S.C., Jr.; Huang, L.S. Hepatic overexpression of hormone-sensitive lipase and adipose triglyceride lipase promotes fatty acid oxidation, stimulates direct release of free fatty acids, and ameliorates steatosis. J. Biol. Chem. 2008, 283, 13087–13099. [Google Scholar] [CrossRef]
- Ong, K.T.; Mashek, M.T.; Bu, S.Y.; Greenberg, A.S.; Mashek, D.G. Adipose triglyceride lipase is a major hepatic lipase that regulates triacylglycerol turnover and fatty acid signaling and partitioning. Hepatology 2011, 53, 116–126. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.W.; Wang, S.P.; Alvarez, F.; Casavant, S.; Gauthier, N.; Abed, L.; Soni, K.G.; Yang, G.; Mitchell, G.A. Deficiency of liver adipose triglyceride lipase in mice causes progressive hepatic steatosis. Hepatology 2011, 54, 122–132. [Google Scholar] [CrossRef] [PubMed]
- Gocze, P.M.; Freeman, D.A. Factors underlying the variability of lipid droplet fluorescence in MA-10 Leydig tumor cells. Cytometry 1994, 17, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Herms, A.; Bosch, M.; Ariotti, N.; Reddy, B.J.; Fajardo, A.; Fernández-Vidal, A.; Alvarez-Guaita, A.; Fernández-Rojo, M.A.; Rentero, C.; Tebar, F.; et al. Cell-to-cell heterogeneity in lipid droplets suggests a mechanism to reduce lipotoxicity. Curr. Biol. 2013, 23, 1489–1496. [Google Scholar] [CrossRef] [PubMed]
- Wallstab, C.; Eleftheriadou, D.; Schulz, T.; Damm, G.; Seehofer, D.; Borlak, J.; Holzhütter, H.G.; Berndt, N. A unifying mathematical model of lipid droplet metabolism reveals key molecular players in the development of hepatic steatosis. FEBS J. 2017, 284, 3245–3261. [Google Scholar] [CrossRef]
- Edge, S.B.; Compton, C.C. The American Joint Committee on Cancer: The 7th Edition of the AJCC Cancer Staging Manual and the Future of TNM. Ann. Surg. Oncol. 2010, 17, 1471–1474. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, S.R.; Aaltonen, L.A. Pathology and Genetics of Tumours of the Digestive System; World Health Organization Classification of Tumours; IARC Press: Lyon, France, 2000. [Google Scholar]
- Gielisch, I.; Meierhofer, D. Metabolome and Proteome Profiling of Complex I Deficiency Induced by Rotenone. J. Proteome Res. 2015, 14, 224–235. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef]
- Desmet, V.J.; Gerber, M.; Hoofnagle, J.H.; Manns, M.; Scheuer, P.J. Classification of chronic hepatitis: Diagnosis, grading and staging. Hepatology 1994, 19, 1513–1520. [Google Scholar] [CrossRef] [PubMed]
- Bulik, S.; Holzhutter, H.G.; Berndt, N. The relative importance of kinetic mechanisms and variable enzyme abundances for the regulation of hepatic glucose metabolism—Insights from mathematical modeling. BMC Biol. 2016, 14, 15. [Google Scholar] [CrossRef] [PubMed]
- Senni, N.; Savall, M.; Cabrerizo Granados, D.; Alves-Guerra, M.C.; Sartor, C.; Lagoutte, I.; Gougelet, A.; Terris, B.; Gilgenkrantz, H.; Perret, C.; et al. beta-catenin-activated hepatocellular carcinomas are addicted to fatty acids. Gut 2019, 68, 322–334. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.; Li, T.1.; Li, X.; Zhang, L.; Sun, L.; He, X.; Zhong, X.; Jia, D.; Song, L.; Semenza, G.L.; et al. HIF-1-Mediated Suppression of Acyl-CoA Dehydrogenases and Fatty Acid Oxidation Is Critical for Cancer Progression. Cell Rep. 2014, 8, 1930–1942. [Google Scholar] [CrossRef] [PubMed]
- Dewhirst, M.W.; Cao, Y.; Moeller, B. Cycling hypoxia and free radicals regulate angiogenesis and radiotherapy response. Nat. Rev. Cancer 2008, 8, 425–437. [Google Scholar] [CrossRef]
- Schlaepfer, I.R.; Nambiar, D.K.; Ramteke, A.; Kumar, R.; Dhar, D.; Agarwal, C.; Bergman, B.; Graner, M.; Maroni, P.; Singh, R.P.; et al. Hypoxia induces triglycerides accumulation in prostate cancer cells and extracellular vesicles supporting growth and invasiveness following reoxygenation. Oncotarget 2015, 6, 22836–22856. [Google Scholar] [CrossRef] [PubMed]
- Weber, G.; Cantero, A. Effect of Neoplasia and Fasting on Phospholipid Turnover Rate in Rat Liver. Biochim. Biophys. Acta 1959, 35, 257–259. [Google Scholar] [CrossRef]
- Liu, Y. Protein kinases and phospholipids in human hepatocellular carcinoma. Chin. J. Oncol. 1991, 13, 242–244. [Google Scholar]
- Lee, G.; Jeong, Y.S.; Kwak, M.J.; Koh, J.; Joo, E.W.; Lee, J.S.; Kah, S.; Sim, Y.E.; Yim, S.Y. Clinical significance of APOB inactivation in hepatocellular carcinoma. Exp. Mol. Med. 2018, 50, 147. [Google Scholar] [CrossRef]
- Itabe, H.; Yamaguchi, T.; Nimura, S.; Sasabe, N. Perilipins: A diversity of intracellular lipid droplet proteins. Lipids Health Dis. 2017, 16, 83. [Google Scholar] [CrossRef] [PubMed]
- Ploegh, H.L. A lipid-based model for the creation of an escape hatch from the endoplasmic reticulum. Nature 2007, 448, 435–438. [Google Scholar] [CrossRef] [PubMed]
- Petan, T.; Jarc, E.; Jusovic, M. Lipid Droplets in Cancer: Guardians of Fat in a Stressful World. Molecules 2018, 23, 1941. [Google Scholar] [CrossRef] [PubMed]
- Lazar, C.; Gatto, L.; Ferro, M.; Bruley, C.; Burger, T. Accounting for the Multiple Natures of Missing Values in Label-Free Quantitative Proteomics Data Sets to Compare Imputation Strategies. J. Proteome Res. 2016, 15, 1116–1125. [Google Scholar] [CrossRef] [PubMed]
- Webb-Robertson, B.J.; Wiberg, H.K.; Matzke, M.M.; Brown, J.N.; Wang, J.; McDermott, J.E.; Smith, R.D.; Rodland, K.D.; Metz, T.O.; Pounds, J.G.; et al. Review, evaluation, and discussion of the challenges of missing value imputation for mass spectrometry-based label-free global proteomics. J. Proteome Res. 2015, 14, 1993–2001. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.; Kang, D.D.; Brock, G.N.; Tseng, G.C. Biological impact of missing-value imputation on downstream analyses of gene expression profiles. Bioinformatics 2011, 27, 78–86. [Google Scholar] [CrossRef]
- Berndt, N.; Bulik, S.; Wallach, I.; Wünsch, T.; König, M.; Stockmann, M.; Meierhofer, D.; Holzhütter, H.G. HEPATOKIN1 is a biochemistry-based model of liver metabolism for applications in medicine and pharmacology. Nat. Commun. 2018, 9, 2386. [Google Scholar] [CrossRef]
| HCC# | Gender | Age | T | N | G | L | V | R | Fibrosis | Liver Disease |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | f | 70 | 2 | - | 2 | 0 | 1 | 0 | 4 | Alcohol-induced cirrhosis |
| 2 | m | 68 | 2 | 0 (0/1) | 2 | 0 | 1 | 0 | 2 | Cryptogenic fibrosis *) |
| 3 | m | 67 | 1b | - | 2 | 0 | 0 | 1 | 1 | Cryptogenic fibrosis *) |
| 4 | m | 80 | 1 | - | 3 | 0 | 0 | 0 | 1 | Alcohol-induced hepatic fibrosis |
| 5 | f | 22 | 1 | - | 1 | 0 | 0 | 0 | 0 | No chronic liver disease history |
| 6 | f | 67 | 1a | - | 2 | 0 | 0 | 0 | 4 | Cryptogenic cirrhosis *) |
| 7 | m | 73 | 2 | - | 2 | 0 | 0 | 0 | 4 | Alcohol-induced cirrhosis |
| 8 | m | 71 | 2 | - | 2 | 0 | 0 | 0 | 3 | Alcohol-induced advanced hepatic fibrosis |
| 9 | f | 66 | 1b | - | 2 | 0 | 0 | 0 | 4 | NASH cirrhosis |
| 10 | f | 59 | 2 | - | 2 | 0 | 0 | 0 | 2 | Cryptogenic fibrosis *) |
| 11 | m | 77 | 1b | - | 3 | 0 | 0 | 0 | 2 | Cryptogenic fibrosis *) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Berndt, N.; Eckstein, J.; Heucke, N.; Gajowski, R.; Stockmann, M.; Meierhofer, D.; Holzhütter, H.-G. Characterization of Lipid and Lipid Droplet Metabolism in Human HCC. Cells 2019, 8, 512. https://doi.org/10.3390/cells8050512
Berndt N, Eckstein J, Heucke N, Gajowski R, Stockmann M, Meierhofer D, Holzhütter H-G. Characterization of Lipid and Lipid Droplet Metabolism in Human HCC. Cells. 2019; 8(5):512. https://doi.org/10.3390/cells8050512
Chicago/Turabian StyleBerndt, Nikolaus, Johannes Eckstein, Niklas Heucke, Robert Gajowski, Martin Stockmann, David Meierhofer, and Hermann-Georg Holzhütter. 2019. "Characterization of Lipid and Lipid Droplet Metabolism in Human HCC" Cells 8, no. 5: 512. https://doi.org/10.3390/cells8050512
APA StyleBerndt, N., Eckstein, J., Heucke, N., Gajowski, R., Stockmann, M., Meierhofer, D., & Holzhütter, H.-G. (2019). Characterization of Lipid and Lipid Droplet Metabolism in Human HCC. Cells, 8(5), 512. https://doi.org/10.3390/cells8050512








