Comparison of the Opn-CreER and Ck19-CreER Drivers in Bile Ducts of Normal and Injured Mouse Livers
Abstract
:1. Introduction
2. Materials and Methods
2.1. Mouse Strains
2.2. Tamoxifen and CCl4 Injections
2.3. Immunostaining and Imaging
2.4. Quantitative Analysis
2.5. Statistical Analysis
3. Results
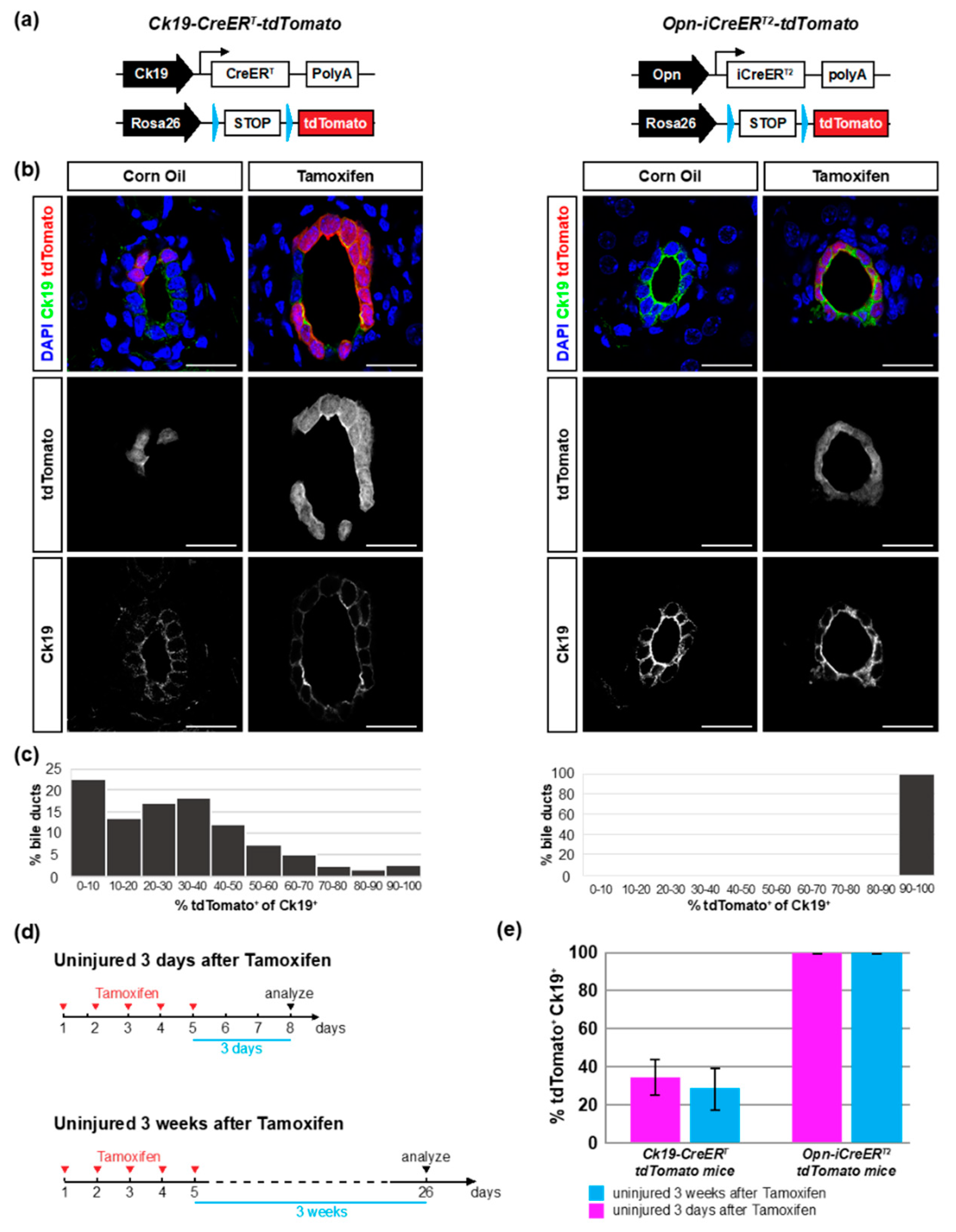
3.1. Opn-iCreERT2 Drives loxP Site Recombination more Efficiently than Ck19-CreERT
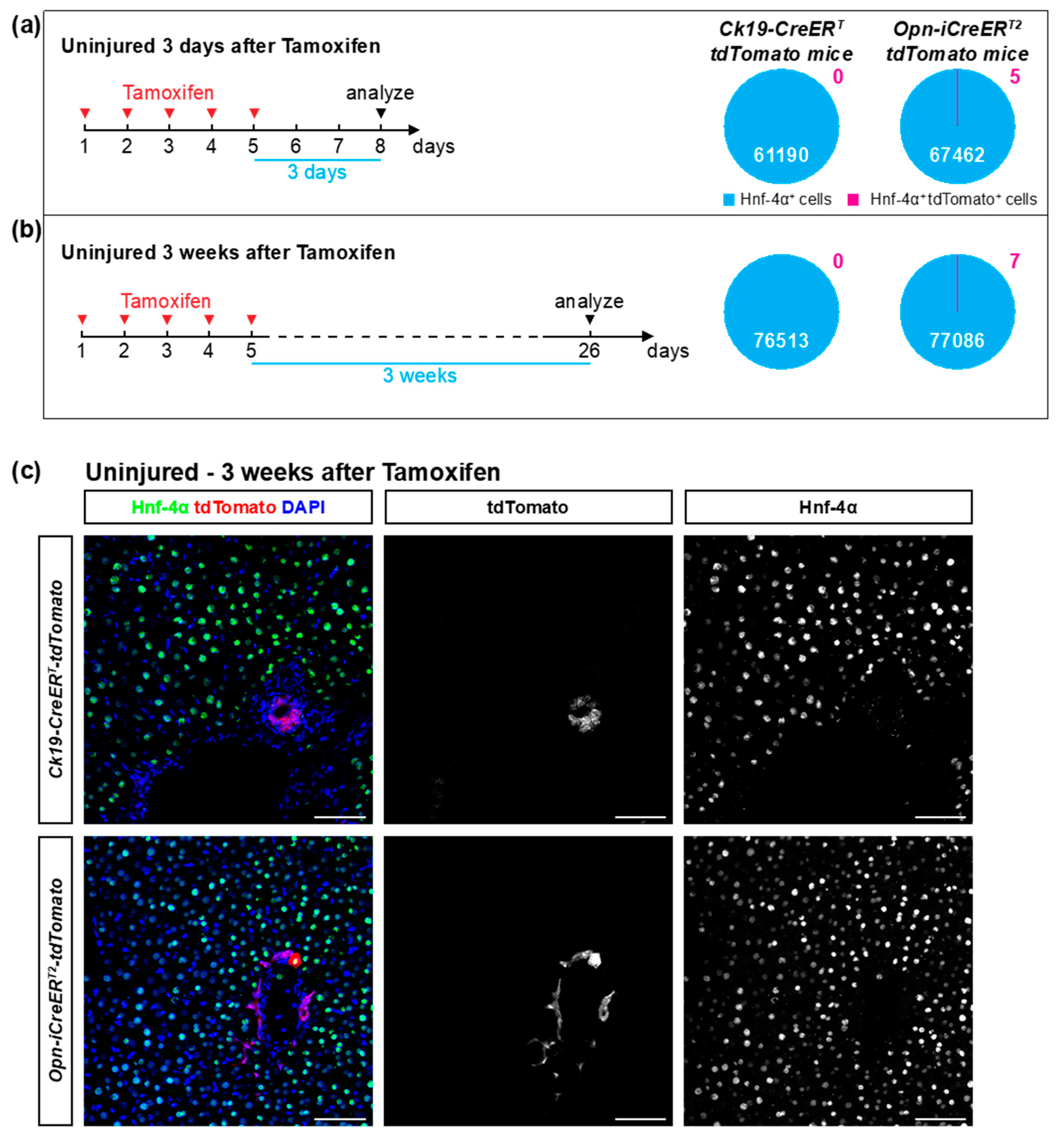
3.2. The Ck19-CreERT and Opn-iCreERT2 Drivers are Highly Specific towards Cholangiocytes
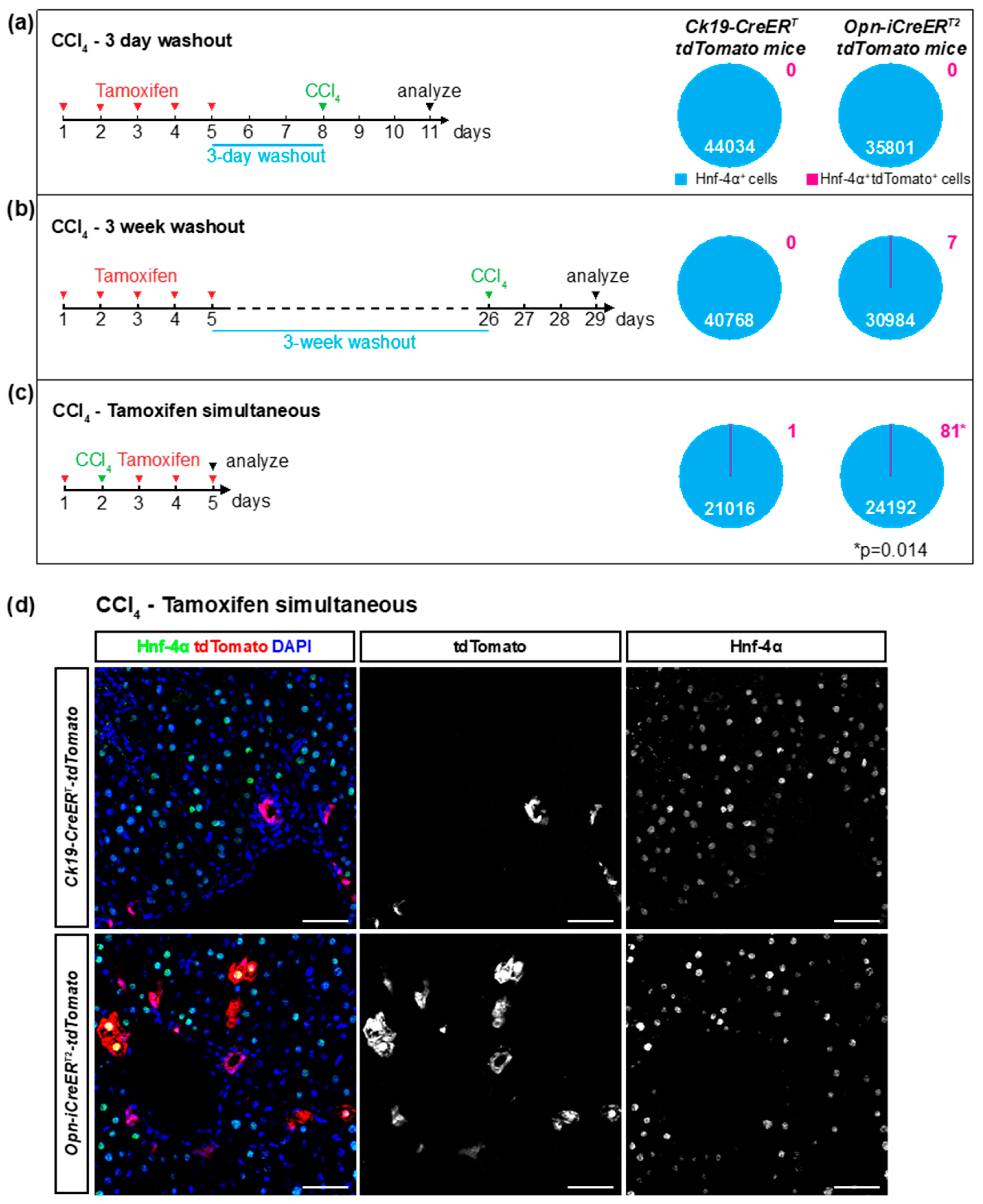
3.3. Ectopic Expression of the Cre Drivers after Liver Injury
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Lemaigre, F.P. Determining the fate of hepatic cells by lineage tracing: Facts and pitfalls. Hepatology 2015, 61, 2100–2103. [Google Scholar] [CrossRef] [Green Version]
- Means, A.L.; Xu, Y.; Zhao, A.; Ray, K.C.; Gu, G. A CK19(CreERT) knockin mouse line allows for conditional DNA recombination in epithelial cells in multiple endodermal organs. Genesis 2008, 46, 318–323. [Google Scholar] [CrossRef] [PubMed]
- Solar, M.; Cardalda, C.; Houbracken, I.; Martín, M.; Maestro, M.A.; De Medts, N.; Xu, X.; Grau, V.; Heimberg, H.; Bouwens, L.; et al. Pancreatic exocrine duct cells give rise to insulin-producing beta cells during embryogenesis but not after birth. Dev. Cell 2009, 17, 849–860. [Google Scholar] [CrossRef] [PubMed]
- Furuyama, K.; Kawaguchi, Y.; Akiyama, H.; Horiguchi, M.; Kodama, S.; Kuhara, T.; Hosokawa, S.; Elbahrawy, A.; Soeda, T.; Koizumi, M.; et al. Continuous cell supply from a Sox9-expressing progenitor zone in adult liver, exocrine pancreas and intestine. Nat. Genet. 2011, 43, 34–41. [Google Scholar] [CrossRef]
- Español-Suñer, R.; Carpentier, R.; Van Hul, N.; Legry, V.; Achouri, Y.; Cordi, S.; Jacquemin, P.; Lemaigre, F.; Leclercq, I.A. Liver progenitor cells yield functional hepatocytes in response to chronic liver injury in mice. Gastroenterology 2012, 143, 1564–1575. [Google Scholar] [CrossRef] [PubMed]
- Jörs, S.; Jeliazkova, P.; Ringelhan, M.; Thalhammer, J.; Dürl, S.; Ferrer, J.; Sander, M.; Heikenwalder, M.; Schmid, R.M.; Siveke, J.T.; et al. Lineage fate of ductular reactions in liver injury and carcinogenesis. J. Clin. Invest. 2015, 125, 2445–2457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodrigo-Torres, D.; Affò, S.; Coll, M.; Morales-Ibanez, O.; Millán, C.; Blaya, D.; Alvarez-Guaita, A.; Rentero, C.; Lozano, J.J.; Maestro, M.A.; et al. The biliary epithelium gives rise to liver progenitor cells. Hepatology 2014, 60, 1367–1377. [Google Scholar] [CrossRef] [PubMed]
- Lokmane, L.; Haumaitre, C.; Garcia-Villalba, P.; Anselme, I.; Schneider-Maunoury, S.; Cereghini, S. Crucial role of vHNF1 in vertebrate hepatic specification. Development 2008, 135, 2777–2786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kopp, J.L.; Dubois, C.L.; Schaffer, A.E.; Hao, E.; Shih, H.P.; Seymour, P.A.; Ma, J.; Sander, M. Sox9+ ductal cells are multipotent progenitors throughout development but do not produce new endocrine cells in the normal or injured adult pancreas. Development 2011, 138, 653–665. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Font-Burgada, J.; Shalapour, S.; Ramaswamy, S.; Hsueh, B.; Rossell, D.; Umemura, A.; Taniguchi, K.; Nakagawa, H.; Valasek, M.A.; Ye, L.; et al. Hybrid Periportal Hepatocytes Regenerate the Injured Liver without Giving Rise to Cancer. Cell 2015, 162, 766–779. [Google Scholar] [CrossRef] [Green Version]
- Tarlow, B.D.; Finegold, M.J.; Grompe, M. Clonal tracing of Sox9+ liver progenitors in mouse oval cell injury. Hepatology 2014, 60, 278–289. [Google Scholar] [CrossRef]
- He, L.; Li, Y.; Pu, W.; Huang, X.; Tian, X.; Wang, Y.; Zhang, H.; Liu, Q.; Zhang, L.; Zhao, H.; et al. Enhancing the precision of genetic lineage tracing using dual recombinases. Nat. Med. 2017, 23, 1488–1498. [Google Scholar] [CrossRef]
- Arriazu, E.; Ge, X.; Leung, T.M.; Magdaleno, F.; Lopategi, A.; Lu, Y.; Kitamura, N.; Urtasun, R.; Theise, N.; Antoine, D.J.; et al. Signalling via the osteopontin and high mobility group box-1 axis drives the fibrogenic response to liver injury. Gut 2017, 66, 1123–1137. [Google Scholar] [CrossRef]
- Wang, X.; Lopategi, A.; Ge, X.; Lu, Y.; Kitamura, N.; Urtasun, R.; Leung, T.M.; Fiel, M.I.; Nieto, N. Osteopontin induces ductular reaction contributing to liver fibrosis. Gut 2014, 63, 1805–1818. [Google Scholar] [CrossRef]
- Kawashima, R.; Mochida, S.; Matsui, A.; You Lu Tu, Z.Y.; Ishikawa, K.; Toshima, K.; Yamanobe, F.; Inao, M.; Ikeda, H.; Ohno, A.; et al. Expression of osteopontin in Kupffer cells and hepatic macrophages and Stellate cells in rat liver after carbon tetrachloride intoxication: A possible factor for macrophage migration into hepatic necrotic areas. Biochem. Biophys. Res. Commun. 1999, 256, 527–531. [Google Scholar] [CrossRef]
- Madisen, L.; Zwingman, T.A.; Sunkin, S.M.; Oh, S.W.; Zariwala, H.A.; Gu, H.; Ng, L.L.; Palmiter, R.D.; Hawrylycz, M.J.; Jones, A.R.; et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 2010, 13, 133–140. [Google Scholar] [CrossRef]
- Mu, X.; Español-Suñer, R.; Mederacke, I.; Affò, S.; Manco, R.; Sempoux, C.; Lemaigre, F.P.; Adili, A.; Yuan, D.; Weber, A.; et al. Hepatocellular carcinoma originates from hepatocytes and not from the progenitor/biliary compartment. J. Clin. Invest. 2015, 125, 3891–3903. [Google Scholar] [CrossRef] [Green Version]
- Lorena, D.; Darby, I.A.; Gadeau, A.P.; Leen, L.L.; Rittling, S.; Porto, L.C.; Rosenbaum, J.; Desmoulière, A. Osteopontin expression in normal and fibrotic liver. altered liver healing in osteopontin-deficient mice. J. Hepatol. 2006, 44, 383–390. [Google Scholar] [CrossRef]
- Sahai, A.; Malladi, P.; Melin-Aldana, H.; Green, R.M.; Whitington, P.F. Upregulation of osteopontin expression is involved in the development of nonalcoholic steatohepatitis in a dietary murine model. Am. J. Physiol. Gastrointest. Liver Physiol. 2004, 287, G264–G273. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Willet, S.G.; Bankaitis, E.D.; Xu, Y.; Wright, C.V.; Gu, G. Non-parallel recombination limits Cre-LoxP-based reporters as precise indicators of conditional genetic manipulation. Genesis 2013, 51, 436–442. [Google Scholar] [CrossRef] [Green Version]
- Reinert, R.B.; Kantz, J.; Misfeldt, A.A.; Poffenberger, G.; Gannon, M.; Brissova, M.; Powers, A.C. Tamoxifen-Induced Cre-loxP Recombination Is Prolonged in Pancreatic Islets of Adult Mice. PLoS ONE 2012, 7, e33529. [Google Scholar] [CrossRef]
- Carpentier, R.; Suñer, R.E.; van Hul, N.; Kopp, J.L.; Beaudry, J.B.; Cordi, S.; Antoniou, A.; Raynaud, P.; Lepreux, S.; Jacquemin, P.; et al. Embryonic ductal plate cells give rise to cholangiocytes, periportal hepatocytes, and adult liver progenitor cells. Gastroenterology 2011, 141, 1432–1438.e4. [Google Scholar] [CrossRef]
- Chen, Y.; Guldiken, N.; Spurny, M.; Mohammed, H.H.; Haybaeck, J.; Pollheimer, M.J.; Fickert, P.; Gassler, N.; Jeon, M.K.; Trautwein, C.; et al. Loss of keratin 19 favours the development of cholestatic liver disease through decreased ductular reaction. J. Pathol. 2015, 237, 343–354. [Google Scholar] [CrossRef]
| Treatment Group | Mice (n) | % tdTomato+ of Ck19+ | ||||
|---|---|---|---|---|---|---|
| Opn-Cre | Ck19-Cre | Opn-Cre | Ck19-Cre | Opn-Cre | Ck19-Cre | |
| Uninjured 3 days after TAM | 7 | 7 | 99.9 ± 0.1 | 34.8 ± 9.1 | ||
| Uninjured 3 weeks after TAM | 8 | 8 | 99.9 ± 0.1 | 27.6 ± 11.7 | ||
| CCl4 3-day washout | 4 | 5 | 100 | 38.9 ± 10.8 | ||
| CCl4 3-week washout | 3 | 4 | 99.9 ± 0.2 | 27.8 ± 5.3 | ||
| Treatment Group | Mice (n) | % tdTomato+ of Hnf-4α+ | ||||
|---|---|---|---|---|---|---|
| Opn-Cre | Ck19-Cre | Opn-Cre | Ck19-Cre | Opn-Cre | Ck19-Cre | |
| uninjured 3 days after TAM | 7 | 7 | 0.0074 ± 0.0126 | 0 | ||
| uninjured 3 weeks after TAM | 8 | 8 | 0.0091 ± 0.0098 | 0 | ||
| CCl4 3-day washout | 4 | 5 | 0 | 0 | ||
| CCl4 3-week washout | 3 | 4 | 0.0226 ± 0.0371 | 0 | ||
| CCl4 no washout † | 5 | 5 | 0.3348 ± 0.3588 | 0.0048 ± 0.0091 | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lesaffer, B.; Verboven, E.; Van Huffel, L.; Moya, I.M.; van Grunsven, L.A.; Leclercq, I.A.; Lemaigre, F.P.; Halder, G. Comparison of the Opn-CreER and Ck19-CreER Drivers in Bile Ducts of Normal and Injured Mouse Livers. Cells 2019, 8, 380. https://doi.org/10.3390/cells8040380
Lesaffer B, Verboven E, Van Huffel L, Moya IM, van Grunsven LA, Leclercq IA, Lemaigre FP, Halder G. Comparison of the Opn-CreER and Ck19-CreER Drivers in Bile Ducts of Normal and Injured Mouse Livers. Cells. 2019; 8(4):380. https://doi.org/10.3390/cells8040380
Chicago/Turabian StyleLesaffer, Bram, Elisabeth Verboven, Leen Van Huffel, Iván M. Moya, Leo A. van Grunsven, Isabelle A. Leclercq, Frédéric P. Lemaigre, and Georg Halder. 2019. "Comparison of the Opn-CreER and Ck19-CreER Drivers in Bile Ducts of Normal and Injured Mouse Livers" Cells 8, no. 4: 380. https://doi.org/10.3390/cells8040380
APA StyleLesaffer, B., Verboven, E., Van Huffel, L., Moya, I. M., van Grunsven, L. A., Leclercq, I. A., Lemaigre, F. P., & Halder, G. (2019). Comparison of the Opn-CreER and Ck19-CreER Drivers in Bile Ducts of Normal and Injured Mouse Livers. Cells, 8(4), 380. https://doi.org/10.3390/cells8040380






