Trehalose Modulates Autophagy Process to Counteract Gliadin Cytotoxicity in an In Vitro Celiac Disease Model
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture and Autophagy Modulation
2.2. Gliadin Digestion and Fluorescent Labelling
2.3. Immunoblotting Analysis
2.4. Apoptosis and Autophagy Cytofluorimetric Analysis
2.5. Multispectral Imaging Flow Cytometry (MIFC) and Spot-Count Analysis.
2.6. Statistical Analysis
3. Results
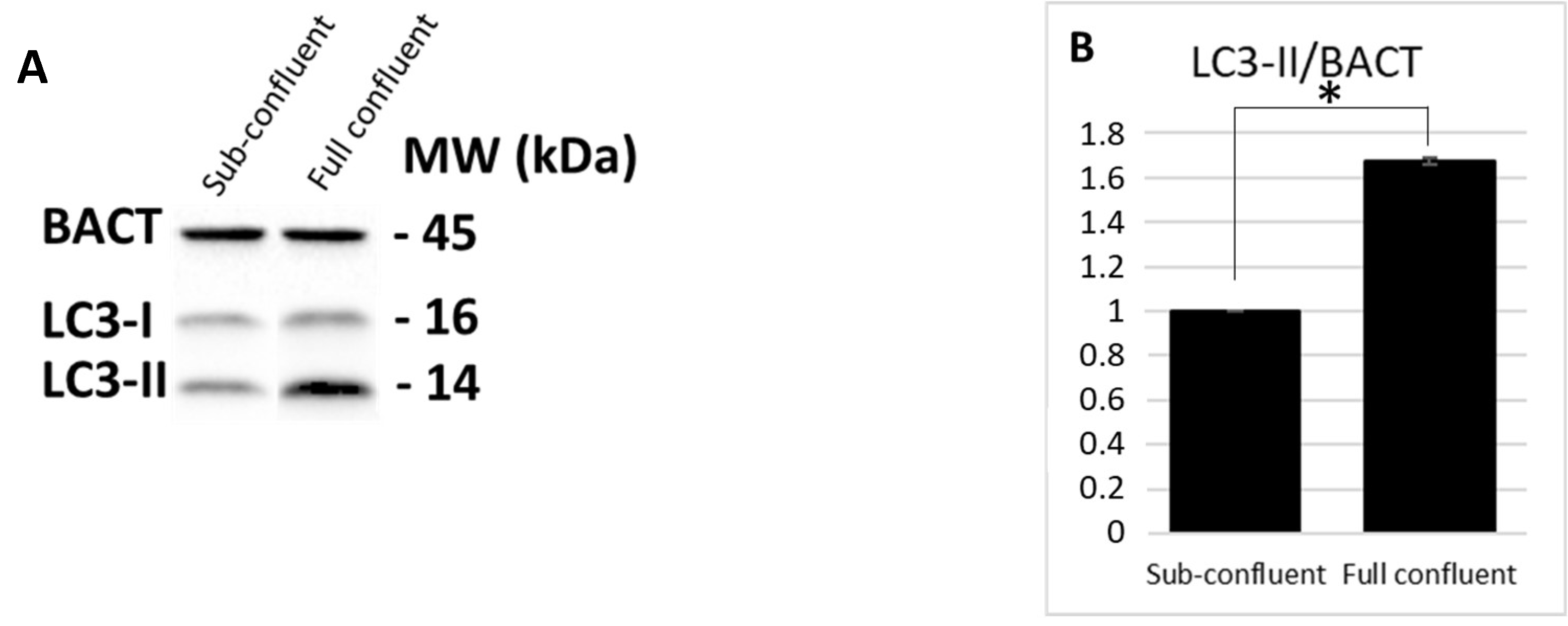
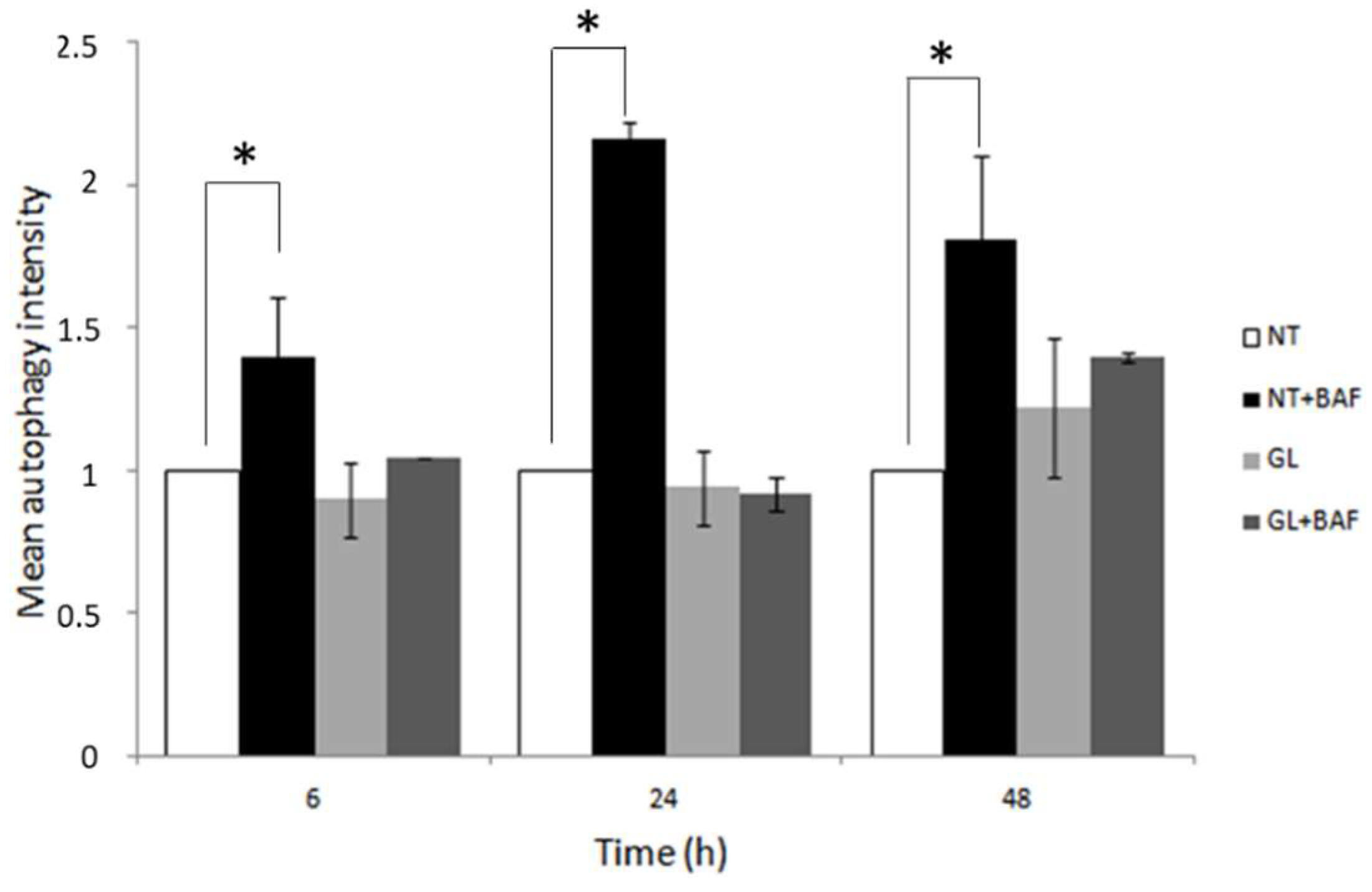
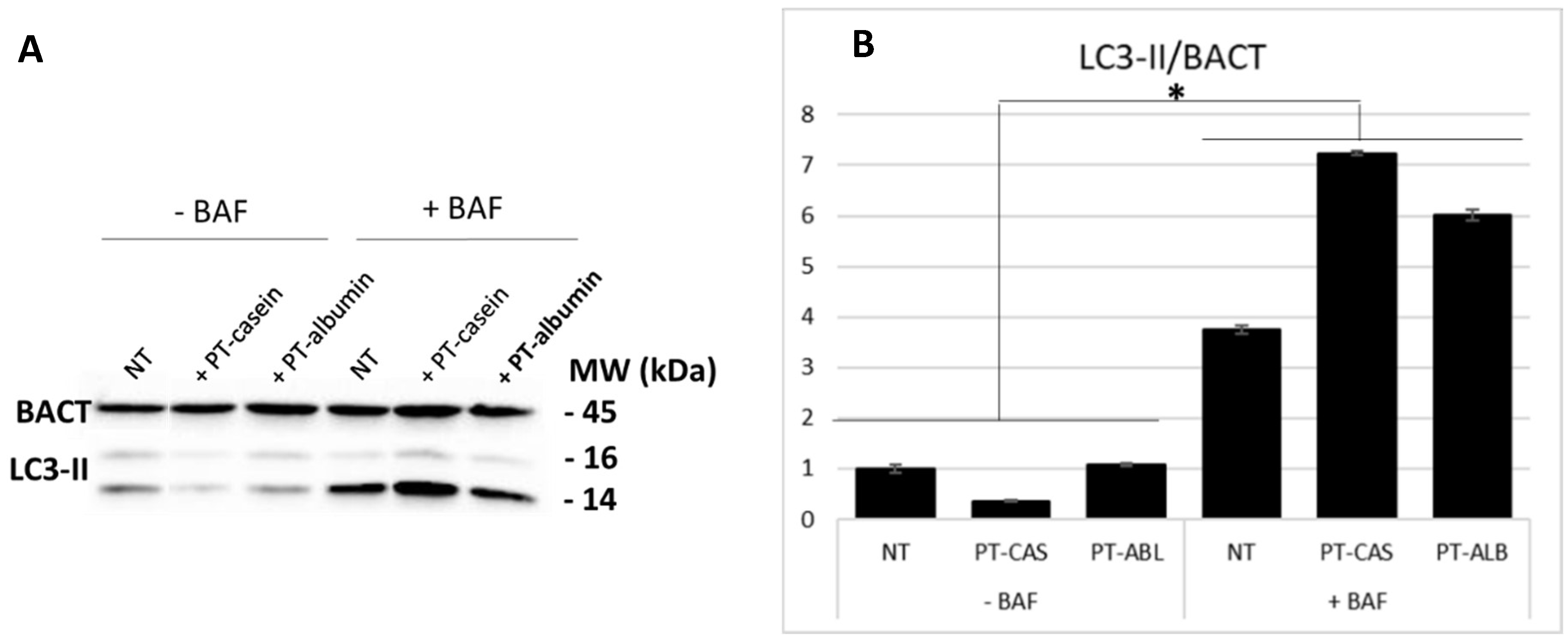
3.1. PT-Gliadin Administration Leads to Autophagy Blockage and Cell Death in Caco-2 Cells Monolayer
3.2. Single Cell Analysis Confirms Caco-2 Autophagy Blockage After PT-Gliadin Administration
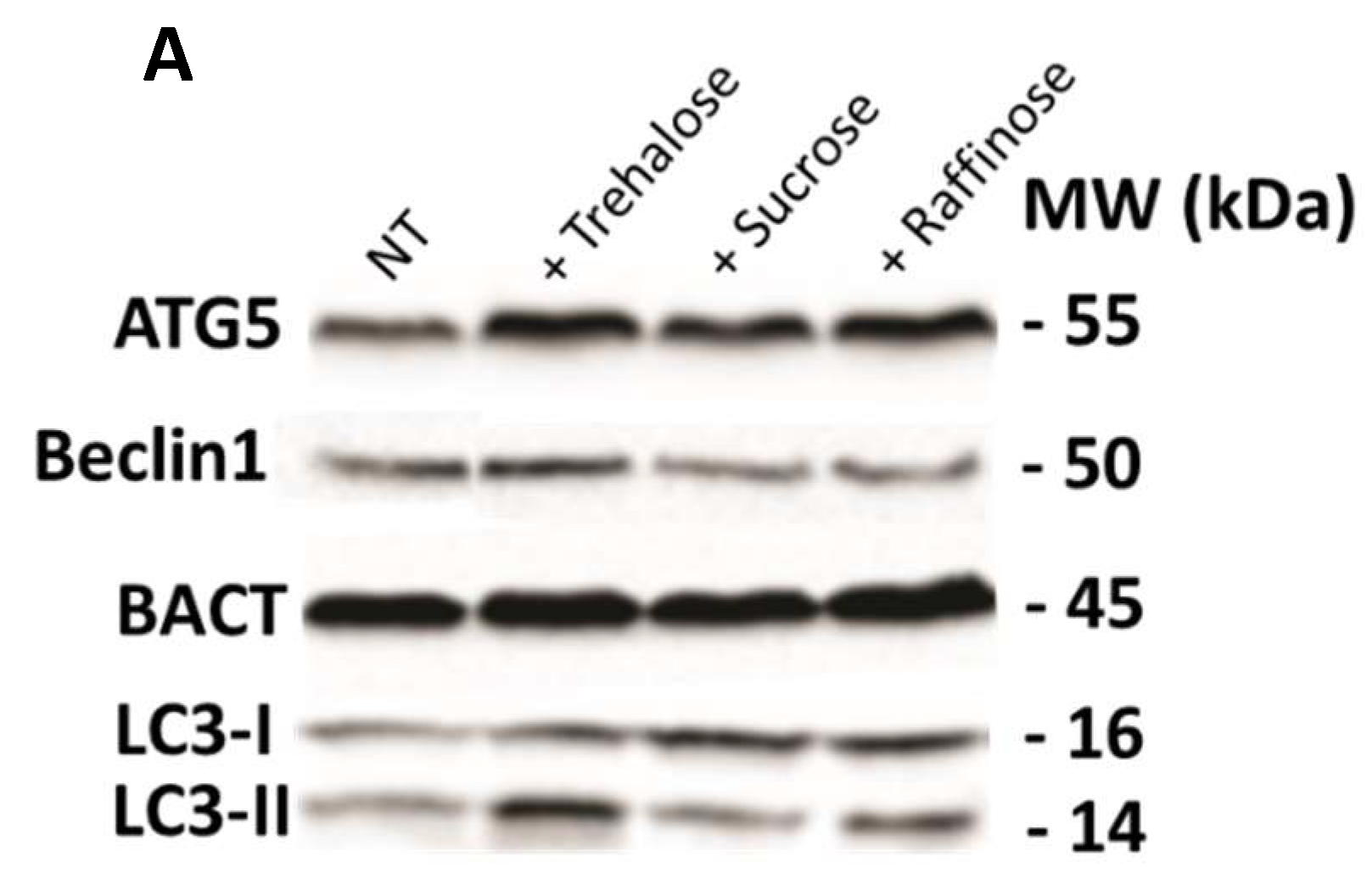
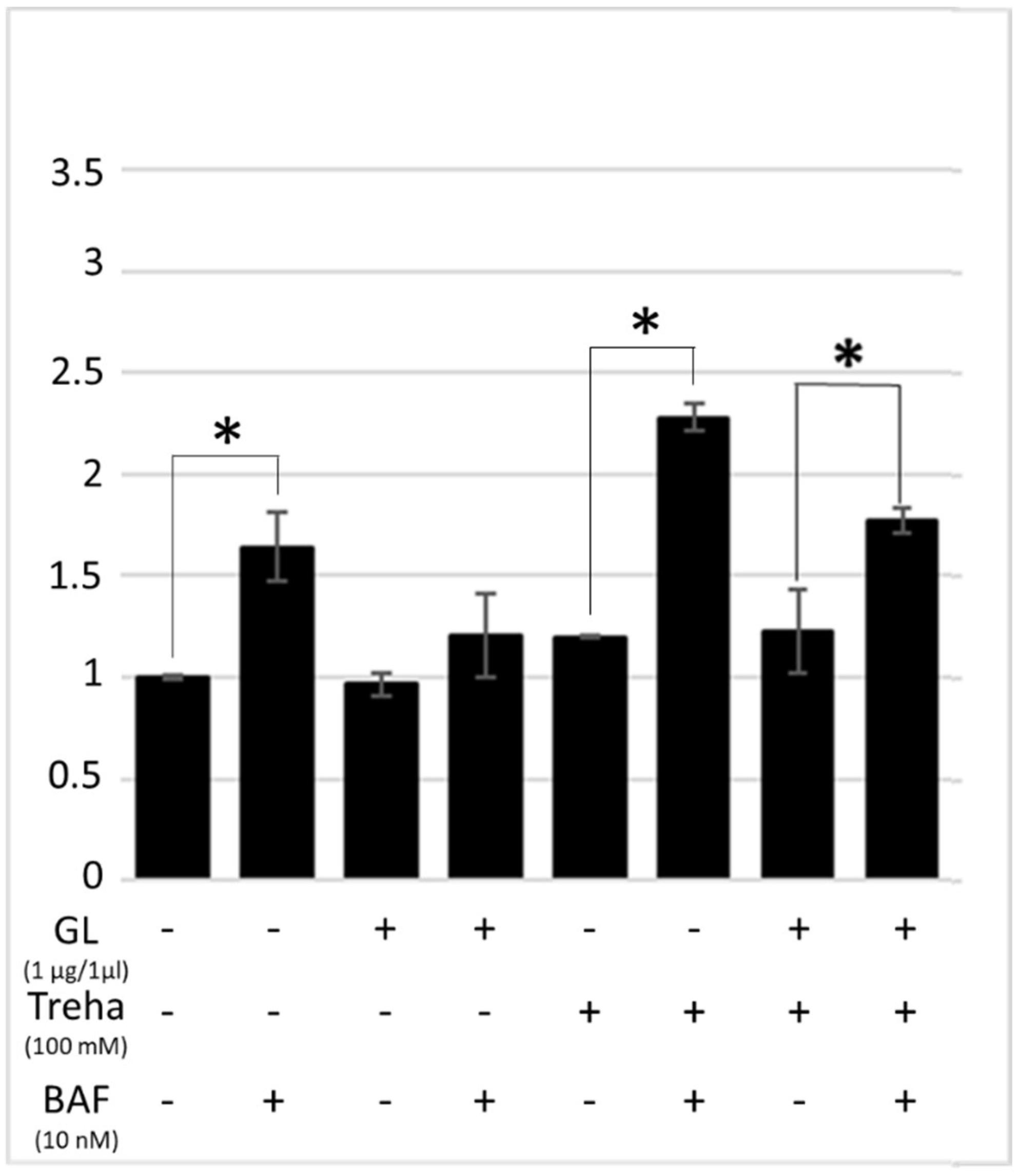
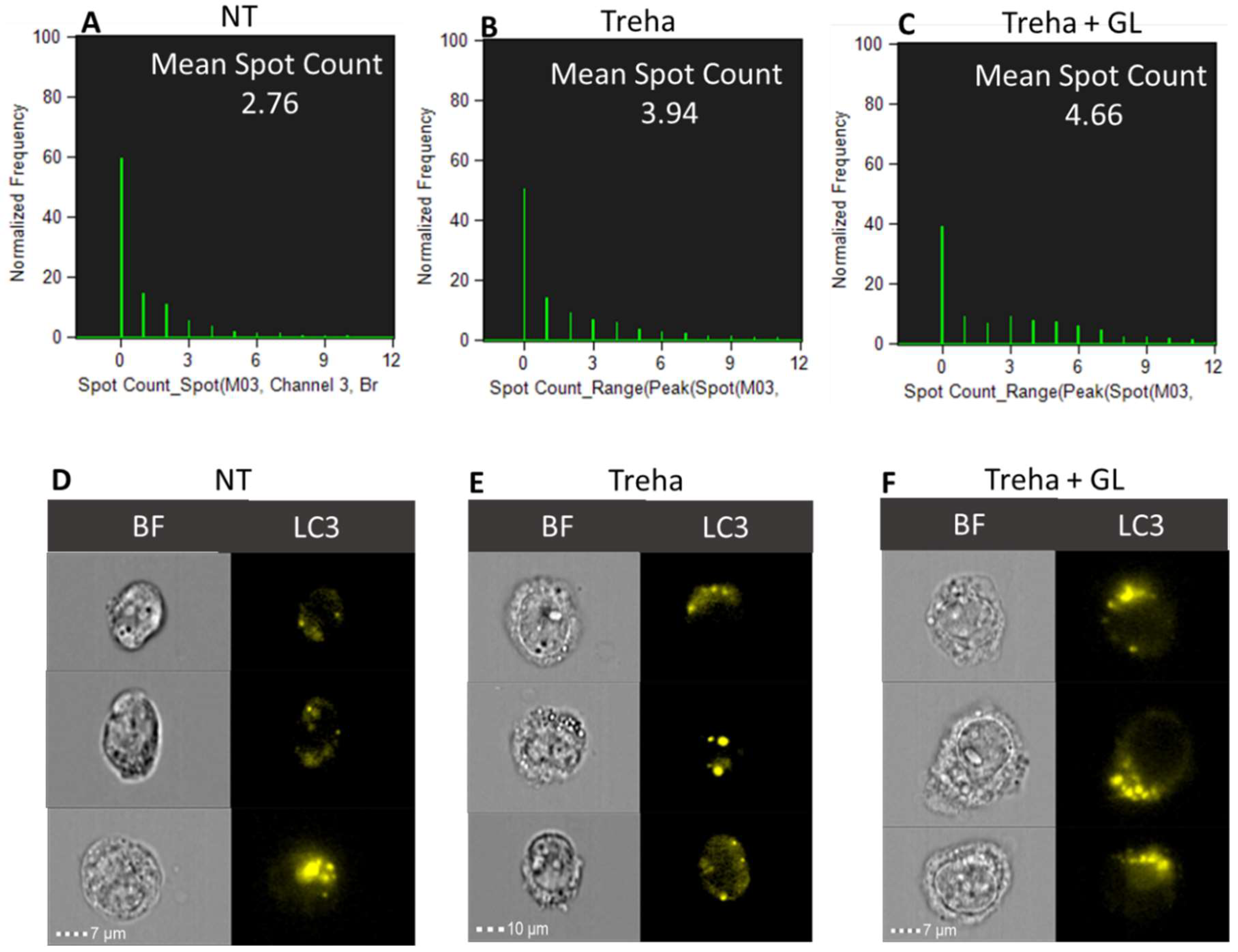
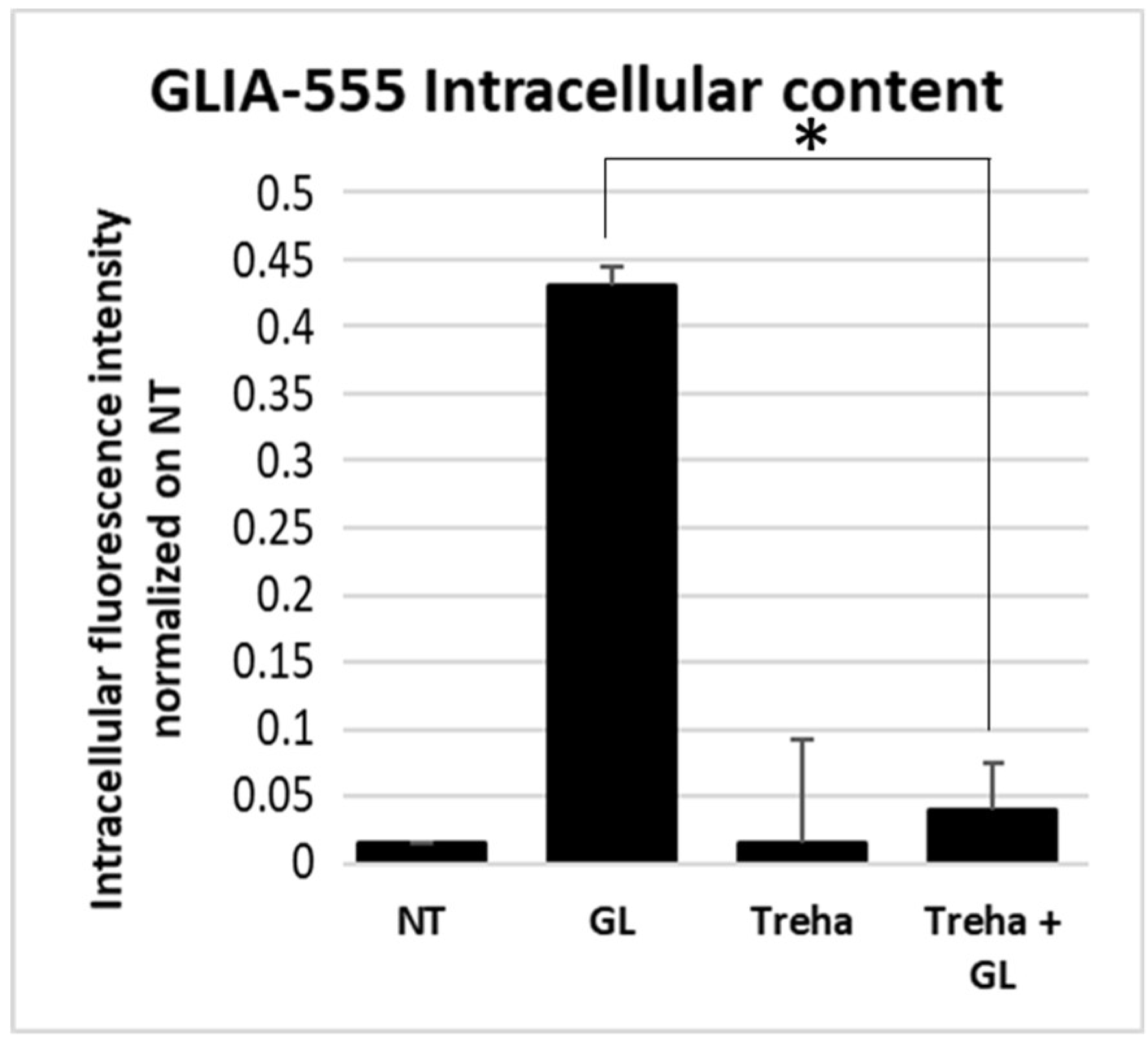
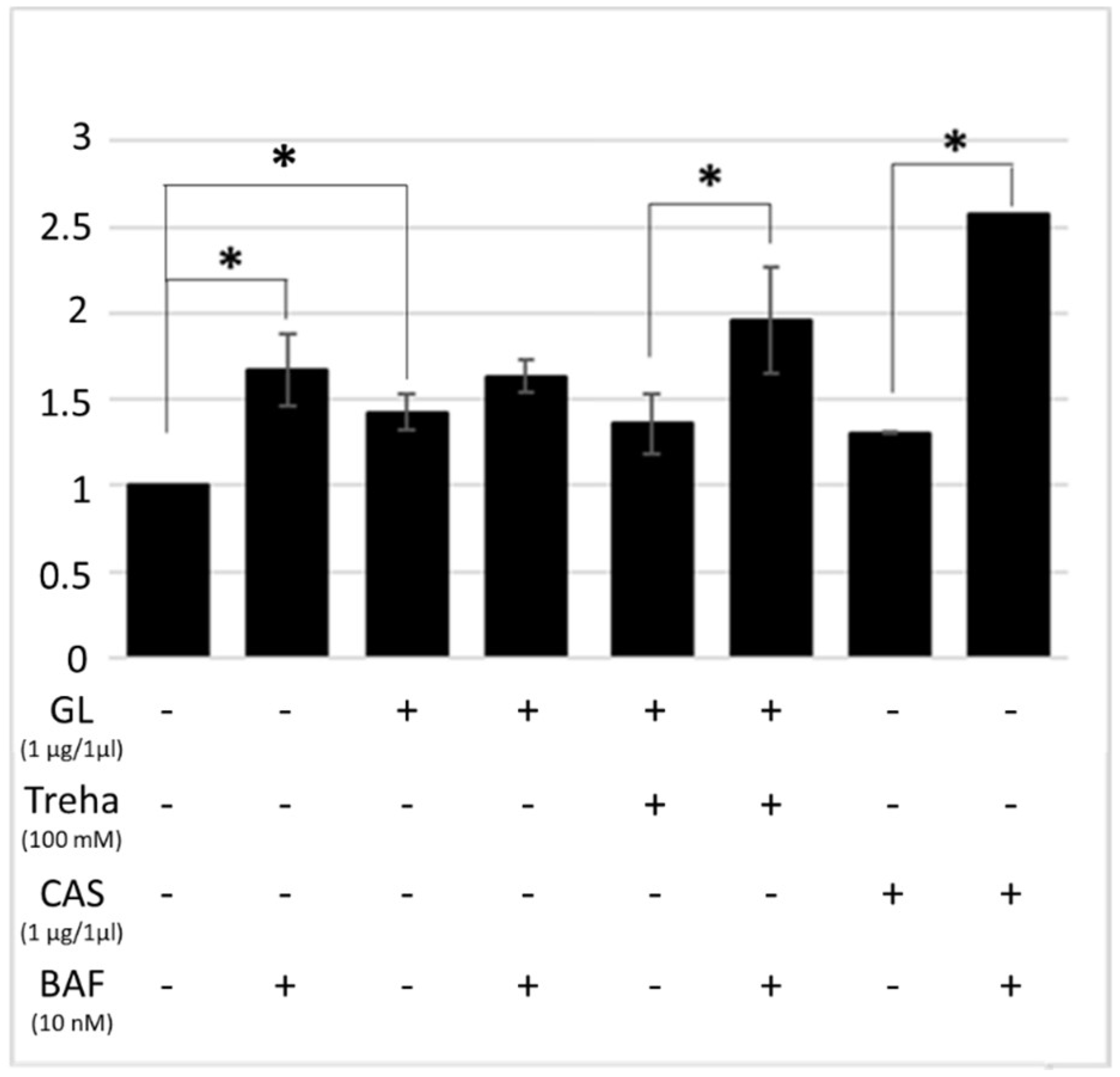
3.3. Trehalose, an mTOR-Independent Inducer of Autophagy, Increases Autophagic Flux and Counteract PT-Gliadin Cytotoxicity Reducing its Intracellular Content
3.4. Trehalose Activates Autophagy in HT-29 Cells Following PT-Gliadin Administration
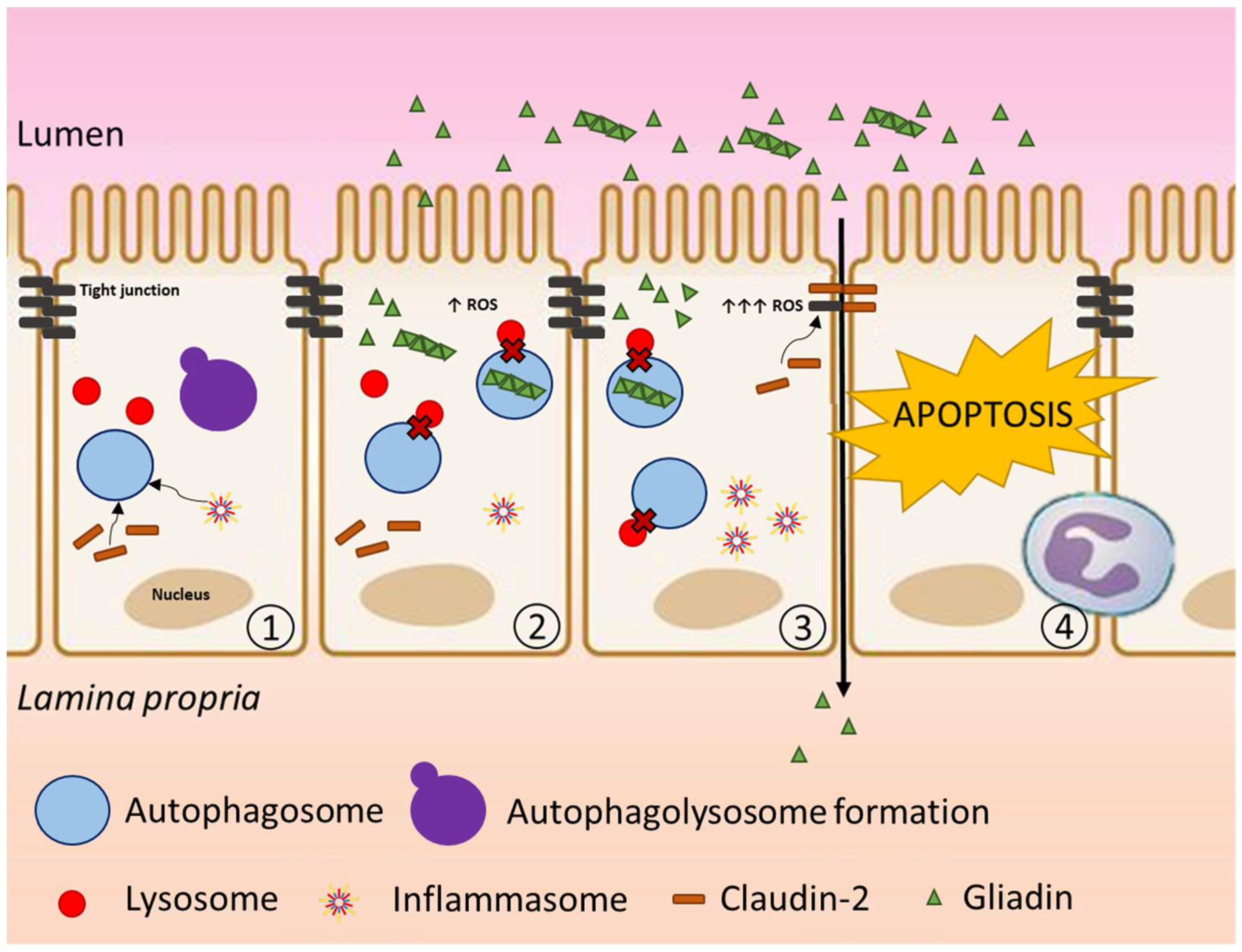
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Di Sabatino, A.; Corazza, G.R. Coeliac disease. Lancet 2009, 373, 1480–1493. [Google Scholar] [CrossRef]
- Cornell, H.J.; Townley, R.R. The toxicity of certain cereal proteins in coeliac disease. Gut 1974, 15, 862–869. [Google Scholar] [CrossRef] [PubMed]
- Wieser, H. Chemistry of gluten proteins. Food Microbiol. 2007, 24, 115–119. [Google Scholar] [CrossRef]
- Wieser, H. Relation between gliadin structure and coeliac toxicity. Acta Paediatr. Suppl. 1996, 412, 3–9. [Google Scholar] [CrossRef]
- Cornell, H.J.; Wills-Johnson, G. Structure-activity relationships in coeliac-toxic gliadin peptides. Amino Acids 2001, 21, 243–253. [Google Scholar] [CrossRef] [PubMed]
- Barone, M.V.; Nanayakkara, M.; Paolella, G.; Maglio, M.; Vitale, V.; Troiano, R.; Ribecco, M.T.; Lania, G.; Zanzi, D.; Santagata, S.; et al. Gliadin peptide P31-43 localises to endocytic vesicles and interferes with their maturation. PLoS ONE 2010, 5, e12246. [Google Scholar] [CrossRef]
- Herrera, M.G.; Benedini, L.A.; Lonez, C.; Schilardi, P.L.; Hellweg, T.; Ruysschaert, J.M.; Dodero, V.I. Self-assembly of 33-mer gliadin peptide oligomers. Soft Matter 2015, 11, 8648–8660. [Google Scholar] [CrossRef]
- Herrera, M.G.; Vazquez, D.S.; Sreij, R.; Drechsler, M.; Hertle, Y.; Hellweg, T.; Dodero, V.I. Insights into gliadin supramolecular organization at digestive pH 3.0. Colloids Surf. B Biointerfaces 2018, 165, 363–370. [Google Scholar] [CrossRef] [PubMed]
- Herrera, M.G.; Pizzuto, M.; Lonez, C.; Rott, K.; Hütten, A.; Sewald, N.; Ruysschaert, J.M.; Dodero, V.I. Large supramolecular structures of 33-mer gliadin peptide activate toll-like receptors in macrophages. Nanomedicine 2018, 14, 1417–1427. [Google Scholar] [CrossRef]
- Scharl, M.; Rogler, G. Inflammatory bowel disease: Dysfunction of autophagy? Dig. Dis. 2012, 30, 12–19. [Google Scholar] [CrossRef] [PubMed]
- Henckaerts, L.; Cleynen, I.; Brinar, M.; John, J.M.; Van Steen, K.; Rutgeerts, P.; Vermeire, S. Genetic variation in the autophagy gene ULK1 and risk of Crohn’s disease. Inflamm. Bowel Dis. 2011, 17, 1392–1397. [Google Scholar] [CrossRef]
- Manai, F.; Azzalin, A.; Gabriele, F.; Martinelli, C.; Morandi, M.; Biggiogera, M.; Bozzola, M.; Comincini, S. The in vitro effects of enzymatic digested gliadin on the functionality of the autophagy process. Int. J. Mol. Sci. 2018, 19, 635. [Google Scholar] [CrossRef]
- Comincini, S.; Manai, F.; Meazza, C.; Pagani, S.; Martinelli, C.; Pasqua, N.; Pelizzo, G.; Biggiogera, M.; Bozzola, M. Identification of autophagy-related genes and their regulatory miRNAs associated with celiac disease in children. Int. J. Mol. Sci. 2017, 18, 391. [Google Scholar] [CrossRef]
- Kroemer, G.; Mariño, G.; Levine, B. Autophagy and the integrated stress response. Mol. Cell 2010, 40, 280–293. [Google Scholar] [CrossRef]
- Lamark, T.; Johansen, T. Aggrephagy: Selective disposal of protein aggregates by macroautophagy. Int. J. Cell Biol. 2012, 2012, 736905. [Google Scholar] [CrossRef]
- Nighot, P.K.; Hu, C.A.; Ma, T.Y. Autophagy enhances intestinal epithelial tight junction barrier function by targeting claudin-2 protein degradation. J. Biol. Chem. 2015, 290, 7234–7246. [Google Scholar] [CrossRef]
- Alfaiz, A.A.; Micale, L.; Mandriani, B.; Augello, B.; Pellico, M.T.; Chrast, J.; Xenarios, I.; Zelante, L.; Merla, G.; Reymond, A. TBC1D7 mutations are associated with intellectual disability, macrocrania, patellar dislocation, and celiac disease. Hum. Mutat. 2014, 35, 447–451. [Google Scholar] [CrossRef]
- Newberry, C.; McKnight, L.; Sarav, M.; Pickett-Blakely, O. Going gluten free: The history and nutritional mplications of today’s most popular diet. Curr. Gastroenterol. Rep. 2017, 19, 54. [Google Scholar] [CrossRef]
- Chen, X.; Li, M.; Li, L.; Xu, S.; Huang, D.; Ju, M.; Huang, J.; Chen, K.; Gu, H. Trehalose, sucrose and raffinose are novel activators of autophagy in human keratinocytes through an mTOR-independent pathway. Sci. Rep. 2016, 6, 28423. [Google Scholar] [CrossRef]
- Mizunoe, Y.; Kobayashi, M.; Sudo, Y.; Watanabe, S.; Yasukawa, H.; Natori, D.; Hoshino, A.; Negishi, A.; Okita, N.; Komatsu, M.; et al. Trehalose protects against oxidative stress by regulating the Keap1-Nrf2 and autophagy pathways. Redox. Biol. 2018, 15, 115–124. [Google Scholar] [CrossRef]
- Sarkar, S.; Davies, J.E.; Huang, Z.; Tunnacliffe, A.; Rubinsztein, DC. Trehalose, a novel mTOR-independent autophagy enhancer, accelerates the clearance of mutant huntingtin and alpha-synuclein. J. Biol. Chem. 2007, 282, 5641–5652. [Google Scholar] [CrossRef]
- Drago, S.; El Asmar, R.; Di Pierro, M.; Grazia Clemente, M.; Tripathi, A.; Sapone, A.; Thakar, M.; Iacono, G.; Carroccio, A.; D’Agate, C.; et al. Gliadin, zonulin and gut permeability: Effects on celiac and non-celiac intestinal mucosa and intestinal cell lines. Scand. J. Gastroenterol. 2006, 41, 408–419. [Google Scholar] [CrossRef]
- Klionsky, D.J.; Abdelmohsen, K.; Abe, A.; Abedin, M.J.; Abeliovich, H.; Acevedo Arozena, A.; Adachi, H.; Adams, C.M.; Adams, P.D.; Adeli, K.; et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 2016, 12, 1–222. [Google Scholar] [CrossRef]
- Hidalgo, I.J.; Raub, T.J.; Borchardt, R.T. Characterization of the human colon carcinoma cell line (Caco-2) as a model system for intestinal epithelial permeability. Gastroenterology 1989, 96, 736–749. [Google Scholar] [CrossRef]
- Sambuy, Y.; De Angelis, I.; Ranaldi, G.; Scarino, M.L.; Stammati, A.; Zucco, F. The Caco-2 cell line as a model of the intestinal barrier: Influence of cell and culture-related factors on Caco-2 cell functional characteristics. Cell Biol. Toxicol. 2005, 21, 1–26. [Google Scholar] [CrossRef]
- Leontieva, O.V.; Demidenko, Z.N.; Blagosklonny, M.V. Contact inhibition and high cell density deactivate the mammalian target of rapamycin pathway, thus suppressing the senescence program. Proc. Natl. Acad. Sci. USA 2014, 111, 8832–8837. [Google Scholar] [CrossRef]
- Tunçer, S.; Banerjee, S. Determination of autophagy in the Caco-2 spontaneously differentiating gmodelof intestinal epithelial cells. Methods Mol. Biol. 2019, 1854, 55–70. [Google Scholar]
- Barteneva, N.S.; Fasler-Kan, E.; Vorobjev, I.A. Imaging flow cytometry: Coping with heterogeneity in biological systems. J. Histochem. Cytochem. 2012, 60, 723–733. [Google Scholar] [CrossRef]
- Filomeni, G.; De Zio, D.; Cecconi, F. Oxidative stress and autophagy: The clash between damage and metabolic needs. Cell Death Differ. 2015, 22, 377–388. [Google Scholar] [CrossRef]
- Booth, L.A.; Tavallai, S.; Hamed, H.A.; Cruickshanks, N.; Dent, P. The role of cell signalling in the crosstalk between autophagy and apoptosis. Cell Signal. 2014, 26, 549–555. [Google Scholar] [CrossRef]
- Menzies, F.M.; Fleming, A.; Caricasole, A.; Bento, C.F.; Andrews, S.P.; Ashkenazi, A.; Füllgrabe, J.; Jackson, A.; Jimenez Sanchez, M.; Karabiyik, C.; et al. Autophagy and neurodegeneration: Pathogenic mechanisms and therapeutic opportunities. Neuron 2017, 93, 1015–1034. [Google Scholar] [CrossRef] [PubMed]
- Thellung, S.; Scoti, B.; Corsaro, A.; Villa, V.; Nizzari, M.; Gagliani, M.C.; Porcile, C.; Russo, C.; Pagano, A.; Tacchetti, C.; et al. Pharmacological activation of autophagy favors the clearing of intracellular aggregates of misfolded prion protein peptide to prevent neuronal death. Cell Death Dis. 2018, 9, 166. [Google Scholar] [CrossRef]
- Paolella, G.; Lepretti, M.; Martucciello, S.; Nanayakkara, M.; Auricchio, S.; Esposito, C.; Barone, M.V.; Caputo, I. The toxic alpha-gliadin peptide 31-43 enters cells without a surface membrane receptor. Cell Biol. Int. 2018, 42, 112–120. [Google Scholar] [CrossRef] [PubMed]
- Baxt, L.A.; Xavier, R.J. Role of autophagy in the maintenance of intestinal homeostasis. Gastroenterology 2015, 149, 553–562. [Google Scholar] [CrossRef] [PubMed]
- Macias-Ceja, D.C.; Cosín-Roger, J.; Ortiz-Masiá, D.; Salvador, P.; Hernández, C.; Esplugues, J.V.; Calatayud, S.; Barrachina, M.D. Stimulation of autophagy prevents intestinal mucosal inflammation and ameliorates murine colitis. Br. J. Pharmacol. 2017, 174, 2501–2511. [Google Scholar] [CrossRef]
- Ballou, L.M.; Lin, R.Z. Rapamycin and mTOR kinase inhibitors. J. Chem. Biol. 2008, 1, 27–36. [Google Scholar] [CrossRef]
- Houtkooper, R.H.; Pirinen, E.; Auwerx, J. Sirtuins as regulators of metabolism and healthspan. Nat. Rev. Mol. Cell Biol. 2012, 13, 225–238. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, W.; Yan, Z.; Zhao, W.; Mi, J.; Li, J.; Yan, H. Metformin induces autophagy and G0/G1 phase cell cycle arrest in myeloma by targeting the AMPK/mTORC1 and mTORC2 pathways. J. Exp. Clin. Cancer Res. 2018, 37, 63. [Google Scholar] [CrossRef]
- Sarkar, S.; Perlstein, E.O.; Imarisio, S.; Pineau, S.; Cordenier, A.; Maglathlin, R.L.; Webster, J.A.; Lewis, T.A.; O’Kane, C.J.; Schreiber, S.L.; et al. Small molecules enhance autophagy and reduce toxicity in Huntington’s disease models. Nat. Chem. Biol. 2007, 3, 331–338. [Google Scholar] [CrossRef]
- Casarejos, M.J.; Perucho, J.; López-Sendón, J.L.; García de Yébenes, J.; Bettencourt, C.; Gómez, A.; Ruiz, C.; Heutink, P.; Rizzu, P.; Mena, M.A. Trehalose improves human fibroblast deficits in a new CHIP-mutation related ataxia. PLoS ONE 2014, 9, e106931. [Google Scholar] [CrossRef]
- Kang, Y.L.; Saleem, M.A.; Chan, K.W.; Yung, B.Y.; Law, H.K. Trehalose, an mTOR independent autophagy inducer, alleviates human podocyte injury after puromycin aminonucleoside treatment. PLoS ONE 2014, 9, e113520. [Google Scholar] [CrossRef]
- Higuchi, T.; Nishikawa, J.; Inoue, H. Sucrose induces vesicles accumulation and autophagy. J. Cell. Biochem. 2015, 116, 609–617. [Google Scholar] [CrossRef]
- Franke, S.I.R.; Molz, P.; Mai, C.; Ellwanger, J.H.; Zenkner, F.F.; Horta, J.A.; Prá, D. High consumption of sucrose induces DNA damage in male Wistar rats. Anais da Academia Brasileira de Ciências 2017, 89, 2657–2662. [Google Scholar] [CrossRef]
- Elbein, A.D.; Pan, Y.T.; Pastuszak, I.; Carroll, D. New insights on trehalose: A multifunctional molecule. Glycobiology 2003, 13, 17R–27R. [Google Scholar] [CrossRef]
- Tanaka, M.; Machida, Y.; Niu, S.; Ikeda, T.; Jana, N.R.; Doi, H.; Kurosawa, M.; Nekooki, M.; Nukina, N. Trehalose alleviates polyglutamine-mediated pathology in a mouse model of Huntington disease. Nat. Med. 2004, 10, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Netea-Maier, R.T.; Plantinga, T.S.; van de Veerdonk, F.L.; Smit, J.W.; Netea, M.G. Modulation of inflammation by autophagy: Consequences for human disease. Autophagy 2016, 12, 245–260. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, D.S.; Gaviglio, E.A.; Peralta Ramos, J.M.; Bussi, C.; Rodriguez-Galan, M.C.; Iribarren, P. Autophagy in inflammation, infection, neurodegeneration and cancer. Int. Immunopharmacol. 2014, 18, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Rossin, F.; D’Eletto, M.; Macdonald, D.; Farrace, M.G.; Piacentini, M. TG2 transamidating activity acts as a reostat controlling the interplay between apoptosis and autophagy. Amino Acids 2012, 42, 1793–1802. [Google Scholar] [CrossRef] [PubMed]








| Stages | Description | Techniques | Figures |
|---|---|---|---|
| 1 | Effects of digest gliadin on Caco-2 | IB, FC, MIFC | 1,2,3,4,5,6 |
| 2 | Evaluation of different autophagy inducers | FC | 7 |
| 3 | Identification of trehalose as the best candidate | IB | 8 |
| 4 | Beneficial effects of trehalose against digested gliadin | IB, FC, MIFC | 9,10,11,12,13 |
| 5 | Comparison with HT-29 cells | FC | 14 |
| Live | Apoptotic/dead | |
|---|---|---|
| NT | 78.17 ± 2.28 | 21.82 ± 2.28 |
| Trehalose | 75.40 ± 2.45 | 24.60 ± 2.45 |
| Raffinose | 71.97 ± 4.43 | 28.02 ± 4.43 |
| Live | Apoptotic/dead | |
|---|---|---|
| NT | 88.49 ± 3.18 | 11.51 ± 3.18 |
| PT-gliadin | 69.57 ± 5.27 * | 30.43 ± 5.27 * |
| PT-casein | 85.11 ± 2.57 | 14.89 ± 2.57 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manai, F.; Azzalin, A.; Morandi, M.; Riccardi, V.; Zanoletti, L.; Dei Giudici, M.; Gabriele, F.; Martinelli, C.; Bozzola, M.; Comincini, S. Trehalose Modulates Autophagy Process to Counteract Gliadin Cytotoxicity in an In Vitro Celiac Disease Model. Cells 2019, 8, 348. https://doi.org/10.3390/cells8040348
Manai F, Azzalin A, Morandi M, Riccardi V, Zanoletti L, Dei Giudici M, Gabriele F, Martinelli C, Bozzola M, Comincini S. Trehalose Modulates Autophagy Process to Counteract Gliadin Cytotoxicity in an In Vitro Celiac Disease Model. Cells. 2019; 8(4):348. https://doi.org/10.3390/cells8040348
Chicago/Turabian StyleManai, Federico, Alberto Azzalin, Martina Morandi, Veronica Riccardi, Lisa Zanoletti, Marco Dei Giudici, Fabio Gabriele, Carolina Martinelli, Mauro Bozzola, and Sergio Comincini. 2019. "Trehalose Modulates Autophagy Process to Counteract Gliadin Cytotoxicity in an In Vitro Celiac Disease Model" Cells 8, no. 4: 348. https://doi.org/10.3390/cells8040348
APA StyleManai, F., Azzalin, A., Morandi, M., Riccardi, V., Zanoletti, L., Dei Giudici, M., Gabriele, F., Martinelli, C., Bozzola, M., & Comincini, S. (2019). Trehalose Modulates Autophagy Process to Counteract Gliadin Cytotoxicity in an In Vitro Celiac Disease Model. Cells, 8(4), 348. https://doi.org/10.3390/cells8040348







