Autophagy: Dual Response in the Development of Hepatocellular Carcinoma
Abstract
1. Introduction
2. Role of Autophagy in Normal Liver Homeostasis
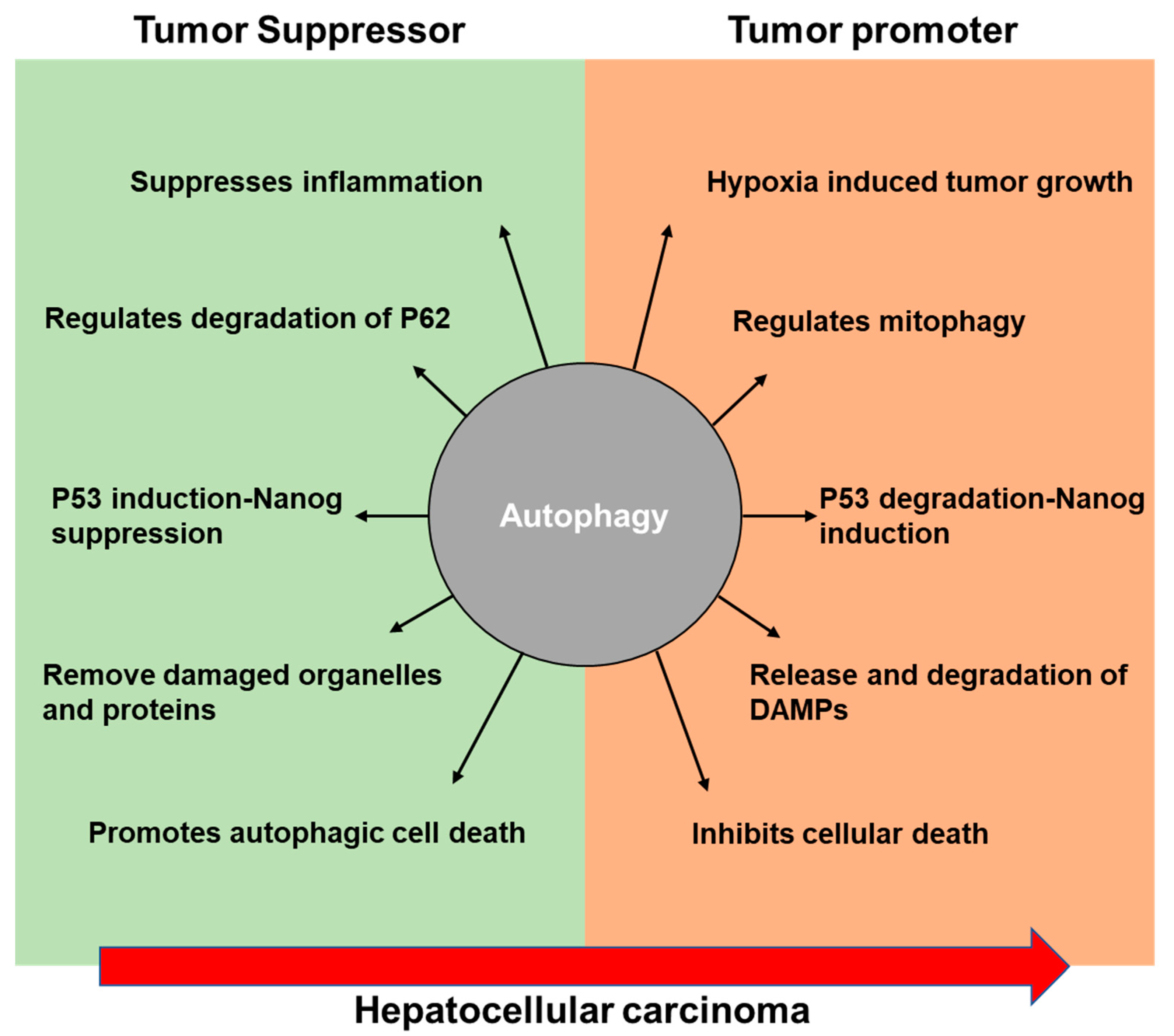
3. Role of Autophagy in Hepatocellular Carcinoma
3.1. Tumor-Suppressive Role of Autophagy
3.2. Tumor-Promoting Role of Autophagy
3.3. Autophagy in HCC Metastasis
4. Autophagy in HCC Therapy
4.1. Autophagy Inducers
4.2. Autophagy Inhibitors
5. Concluding Remarks
Funding
Acknowledgments
Conflicts of Interest
References
- Xie, Z.; Klionsky, D.J. Autophagosome formation: Core machinery and adaptations. Nat. Cell Biol. 2007, 9, 1102–1109. [Google Scholar] [CrossRef] [PubMed]
- Devenish, R.J.; Klionsky, D.J. Autophagy: Mechanism and physiological relevance “brewed” from yeast studies. Front. Biosci. 2012, 54, 1354–1363. [Google Scholar] [CrossRef]
- White, E. The role for autophagy in cancer. J. Clin. Investig. 2015, 125, 42–46. [Google Scholar] [CrossRef] [PubMed]
- Mizushima, N.; Komatsu, M. Autophagy: Renovation of Cells and Tissues. Cell 2011, 147, 728–741. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Klionsky, D.J. Autophagy and human disease. Cell Cycle 2007, 6, 1837–1849. [Google Scholar] [CrossRef] [PubMed]
- Yorimitsu, T.; Klionsky, D.J. Autophagy: Molecular machinery for self-eating. Cell Death Differ. 2005, 12, 1542–1552. [Google Scholar] [CrossRef] [PubMed]
- Kwanten, W.J.; Martinet, W.; Michielsen, P.P.; Francque, S.M. Role of autophagy in the pathophysiology of nonalcoholic fatty liver disease: A controversial issue. World J. Gastroenterol. 2014, 20, 7325–7338. [Google Scholar] [CrossRef]
- Cui, J.; Gong, Z.; Shen, H.-M. The role of autophagy in liver cancer: Molecular mechanisms and potential therapeutic targets. Biochim. Biophys. Acta Rev. Cancer 2013, 1836, 15–26. [Google Scholar] [CrossRef]
- Portolani, N.; Coniglio, A.; Ghidoni, S.; Giovanelli, M.; Benetti, A.; Tiberio, G.A.M.; Giulini, S.M. Early and Late Recurrence after Liver Resection for Hepatocellular Carcinoma. Ann. Surg. 2006, 243, 229–235. [Google Scholar] [CrossRef]
- White, E. Deconvoluting the context-dependent role for autophagy in cancer. Nat. Rev. Cancer 2012, 12, 401–410. [Google Scholar] [CrossRef]
- Janku, F.; McConkey, D.J.; Hong, D.S.; Kurzrock, R. Autophagy as a target for anticancer therapy. Nat. Rev. Clin. Oncol. 2011, 8, 528–539. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, M.; Waguri, S.; Ueno, T.; Iwata, J.; Murata, S.; Tanida, I.; Ezaki, J.; Mizushima, N.; Ohsumi, Y.; Uchiyama, Y.; et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 2005, 169, 425–434. [Google Scholar] [CrossRef] [PubMed]
- Ding, W.-X. Role of autophagy in liver physiology and pathophysiology. World J. Biol. Chem. 2010, 1, 3–12. [Google Scholar] [CrossRef]
- Takamura, A.; Komatsu, M.; Hara, T.; Sakamoto, A.; Kishi, C.; Waguri, S.; Eishi, Y.; Hino, O.; Tanaka, K.; Mizushima, N. Autophagy-deficient mice develop multiple liver tumors. Genes Dev. 2011, 25, 795–800. [Google Scholar] [CrossRef] [PubMed]
- Malhi, H.; Gores, G.J. Cellular and Molecular Mechanisms of Liver Injury. Gastroenterology 2008, 134, 1641–1654. [Google Scholar] [CrossRef] [PubMed]
- Ueno, T.; Komatsu, M. Autophagy in the liver: Functions in health and disease. Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 170–184. [Google Scholar] [CrossRef]
- Rautou, P.E.; Mansouri, A.; Lebrec, D.; Durand, F.; Valla, D.; Moreau, R. Autophagy in liver diseases. J. Hepatol. 2010, 53, 1123–1134. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Han, W.; Sui, X.; Fang, Y.; Pan, H. Autophagy: A novel therapeutic target for hepatocarcinoma (Review). Oncol. Lett. 2014, 7, 1345–1351. [Google Scholar] [CrossRef]
- Abdoli, A.; Alirezaei, M.; Mehrbod, P.; Forouzanfar, F. Autophagy: The multi-purpose bridge in viral infections and host cells. Rev. Med. Virol. 2018, 28, e1973. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Kaushik, S.; Wang, Y.; Xiang, Y.; Novak, I.; Komatsu, M.; Tanaka, K.; Cuervo, A.M.; Czaja, M.J. Autophagy regulates lipid metabolism. Nature 2009, 458, 1131–1135. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Kanasaki, M.; He, J.; Kitada, M.; Nagao, K.; Jinzu, H.; Noguchi, Y.; Maegawa, H.; Kanasaki, K.; Koya, D. Ketogenic essential amino acids replacement diet ameliorated hepatosteatosis with altering autophagy-associated molecules. Biochim. Biophys. Acta Mol. Basis Dis. 2013, 1832, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2017. CA Cancer J. Clin. 2017, 67, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Auld, S.K.J.R.; Scholefield, J.A.; Little, T.J. Genetic variation in the cellular response of Daphnia magna (Crustacea: Cladocera) to its bacterial parasite. Proc. Biol. Sci. 2010, 277, 3291–3297. [Google Scholar] [CrossRef] [PubMed]
- Zhong, J.-H.; Huang, D.-H.; Chen, Z.-Y. Prognostic role of systemic immune-inflammation index in solid tumors: A systematic review and meta-analysis. Oncotarget 2017, 8, 75381–75388. [Google Scholar] [CrossRef] [PubMed]
- Arzumanyan, A.; Reis, H.M.; Feitelson, M.A. Pathogenic mechanisms in HBV- and HCV-associated hepatocellular carcinoma. Nat. Rev. Cancer 2013, 13, 123–135. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Sung, J.J.Y.; Yu, J.; Ng, S.C.; Wong, S.H.; Cho, C.H.; Ng, S.S.M.; Chan, F.K.L.; Wu, W.K.K. Xenophagy in Helicobacter pylori- and Epstein-Barr virus-induced gastric cancer. J. Pathol. 2014, 233, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Shibutani, S.T.; Saitoh, T.; Nowag, H.; Münz, C.; Yoshimori, T. Autophagy and autophagy-related proteins in the immune system. Nat. Immunol. 2015, 16, 1014–1024. [Google Scholar] [CrossRef] [PubMed]
- Saitoh, T.; Fujita, N.; Jang, M.H.; Uematsu, S.; Yang, B.-G.; Satoh, T.; Omori, H.; Noda, T.; Yamamoto, N.; Komatsu, M.; et al. Loss of the autophagy protein Atg16L1 enhances endotoxin-induced IL-1β production. Nature 2008, 456, 264–268. [Google Scholar] [CrossRef]
- Zhou, R.; Yazdi, A.S.; Menu, P.; Tschopp, J. A role for mitochondria in NLRP3 inflammasome activation. Nature 2011, 469, 221–225. [Google Scholar] [CrossRef]
- Bujak, A.L.; Crane, J.D.; Lally, J.S.; Ford, R.J.; Kang, S.J.; Rebalka, I.A.; Green, A.E.; Kemp, B.E.; Hawke, T.J.; Schertzer, J.D.; et al. AMPK activation of muscle autophagy prevents fasting-induced hypoglycemia and myopathy during aging. Cell Metab. 2015, 21, 883–890. [Google Scholar] [CrossRef]
- Zhong, Z.; Sanchez-Lopez, E.; Karin, M. Autophagy, Inflammation, and Immunity: A Troika Governing Cancer and Its Treatment. Cell 2016, 166, 288–298. [Google Scholar] [CrossRef] [PubMed]
- Mridha, A.R.; Wree, A.; Robertson, A.A.B.; Yeh, M.M.; Johnson, C.D.; Van Rooyen, D.M.; Haczeyni, F.; Teoh, N.C.H.; Savard, C.; Ioannou, G.N.; et al. NLRP3 inflammasome blockade reduces liver inflammation and fibrosis in experimental NASH in mice. J. Hepatol. 2017, 66, 1037–1046. [Google Scholar] [CrossRef] [PubMed]
- Petrasek, J.; Bala, S.; Csak, T.; Lippai, D.; Kodys, K.; Menashy, V.; Barrieau, M.; Min, S.-Y.; Kurt-Jones, E.A.; Szabo, G. IL-1 receptor antagonist ameliorates inflammasome-dependent alcoholic steatohepatitis in mice. J. Clin. Investig. 2012, 122, 3476–3489. [Google Scholar] [CrossRef] [PubMed]
- Nakahira, K.; Haspel, J.A.; Rathinam, V.A.K.; Lee, S.-J.; Dolinay, T.; Lam, H.C.; Englert, J.A.; Rabinovitch, M.; Cernadas, M.; Kim, H.P.; et al. Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat. Immunol. 2011, 12, 222–230. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, H.; Zhen, Z.; Ma, X.; Yu, W.; Zeng, H.; Li, L. CXCL17 promotes cell metastasis and inhibits autophagy via the LKB1-AMPK pathway in hepatocellular carcinoma. Gene 2019, 690, 129–136. [Google Scholar] [CrossRef]
- Kotsafti, A.; Farinati, F.; Cardin, R.; Cillo, U.; Nitti, D.; Bortolami, M. Autophagy and apoptosis-related genes in chronic liver disease and hepatocellular carcinoma. BMC Gastroenterol. 2012, 12, 118. [Google Scholar] [CrossRef]
- Yue, Z.; Jin, S.; Yang, C.; Levine, A.J.; Heintz, N. Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc. Natl. Acad. Sci. USA 2003, 100, 15077–15082. [Google Scholar] [CrossRef]
- Wu, F.-Q.; Fang, T.; Yu, L.-X.; Lv, G.-S.; Lv, H.-W.; Liang, D.; Li, T.; Wang, C.-Z.; Tan, Y.-X.; Ding, J.; et al. ADRB2 signaling promotes HCC progression and sorafenib resistance by inhibiting autophagic degradation of HIF1α. J. Hepatol. 2016, 65, 314–324. [Google Scholar] [CrossRef]
- Pursiheimo, J.-P.; Rantanen, K.; Heikkinen, P.T.; Johansen, T.; Jaakkola, P.M. Hypoxia-activated autophagy accelerates degradation of SQSTM1/p62. Oncogene 2009, 28, 334–344. [Google Scholar] [CrossRef]
- Komatsu, M.; Waguri, S.; Koike, M.; Sou, Y.; Ueno, T.; Hara, T.; Mizushima, N.; Iwata, J.; Ezaki, J.; Murata, S.; et al. Homeostatic Levels of p62 Control Cytoplasmic Inclusion Body Formation in Autophagy-Deficient Mice. Cell 2007, 131, 1149–1163. [Google Scholar] [CrossRef]
- Yoshii, S.R.; Mizushima, N. Monitoring and Measuring Autophagy. Int. J. Mol. Sci. 2017, 18, 1865. [Google Scholar] [CrossRef] [PubMed]
- Kessler, S.M.; Laggai, S.; Barghash, A.; Schultheiss, C.S.; Lederer, E.; Artl, M.; Helms, V.; Haybaeck, J.; Kiemer, A.K. IMP2/p62 induces genomic instability and an aggressive hepatocellular carcinoma phenotype. Cell Death Dis. 2015, 6, e1894. [Google Scholar] [CrossRef] [PubMed]
- Umemura, A.; He, F.; Taniguchi, K.; Nakagawa, H.; Yamachika, S.; Font-Burgada, J.; Zhong, Z.; Subramaniam, S.; Raghunandan, S.; Duran, A.; et al. p62, Upregulated during Preneoplasia, Induces Hepatocellular Carcinogenesis by Maintaining Survival of Stressed HCC-Initiating Cells. Cancer Cell 2016, 29, 935–948. [Google Scholar] [CrossRef] [PubMed]
- Aigelsreiter, A.; Neumann, J.; Pichler, M.; Halasz, J.; Zatloukal, K.; Berghold, A.; Douschan, P.; Rainer, F.; Stauber, R.; Haybaeck, J.; et al. Hepatocellular carcinomas with intracellular hyaline bodies have a poor prognosis. Liver Int. 2017, 37, 600–610. [Google Scholar] [CrossRef]
- Moscat, J.; Diaz-Meco, M.T. p62 at the Crossroads of Autophagy, Apoptosis, and Cancer. Cell 2009, 137, 1001–1004. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhang, Y.; Zhu, X.; Chen, C.; Zhang, C.; Xia, Y.; Zhao, Y.; Andrisani, O.; Kong, L. DEAD Box Protein 5 Inhibits Liver Tumorigenesis by Stimulating Autophagy via Interaction with p62/SQSTM1. Hepatology 2018. [Google Scholar] [CrossRef] [PubMed]
- Nicol, S.M.; Bray, S.E.; Derek Black, H.; Lorimore, S.A.; Wright, E.G.; Lane, D.P.; Meek, D.W.; Coates, P.J.; Fuller-Pace, F.V. The RNA helicase p68 (DDX5) is selectively required for the induction of p53-dependent p21 expression and cell-cycle arrest after DNA damage. Oncogene 2013, 32, 3461–3469. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Du, Q.; Cao, Z.; Guo, Z.; Evankovich, J.; Yan, W.; Chang, Y.; Shao, L.; Stolz, D.B.; Tsung, A.; et al. Interferon-gamma induces autophagy with growth inhibition and cell death in human hepatocellular carcinoma (HCC) cells through interferon-regulatory factor-1 (IRF-1). Cancer Lett. 2012, 314, 213–222. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Wu, H.; Liu, X.; Li, B.; Chen, Y.; Ren, X.; Liu, C.-G.; Yang, J.-M. Regulation of autophagy by a beclin 1-targeted microRNA, miR-30a, in cancer cells. Autophagy 2009, 5, 816–823. [Google Scholar] [CrossRef] [PubMed]
- Frankel, L.B.; Wen, J.; Lees, M.; Høyer-Hansen, M.; Farkas, T.; Krogh, A.; Jäättelä, M.; Lund, A.H. microRNA-101 is a potent inhibitor of autophagy. EMBO J. 2011, 30, 4628–4641. [Google Scholar] [CrossRef]
- Chang, Y.; Yan, W.; He, X.; Zhang, L.; Li, C.; Huang, H.; Nace, G.; Geller, D.A.; Lin, J.; Tsung, A. MiR-375 inhibits autophagy and reduces viability of hepatocellular carcinoma cells under hypoxic conditions. Gastroenterology 2012, 143, 177–187. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, Q.; Song, J. Inhibition of autophagy potentiates the proliferation inhibition activity of microRNA-7 in human hepatocellular carcinoma cells. Oncol. Lett. 2017, 14, 3566–3572. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Liu, S.; Han, M.; Feng, S.; Liang, J.; Li, Z.; Li, Y.; Lu, H.; Liu, T.; Ma, Y.; et al. MicroRNA-185 induces potent autophagy via AKT signaling in hepatocellular carcinoma. Tumour Biol. 2017, 39, 1010428317694313. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Lin, J.; Tsung, A. Manipulation of autophagy by MIR375 generates antitumor effects in liver cancer. Autophagy 2012, 8, 1833–1834. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.C.-L.; Kai, A.K.-L.; Ng, I.O.-L. The impact of hypoxia in hepatocellular carcinoma metastasis. Front. Med. 2014, 8, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Wilson, G.K.; Tennant, D.A.; McKeating, J.A. Hypoxia inducible factors in liver disease and hepatocellular carcinoma: Current understanding and future directions. J. Hepatol. 2014, 61, 1397–1406. [Google Scholar] [CrossRef] [PubMed]
- Eales, K.L.; Hollinshead, K.E.R.; Tennant, D.A. Hypoxia and metabolic adaptation of cancer cells. Oncogenesis 2016, 61, 1397–1406. [Google Scholar] [CrossRef]
- Tohme, S.; Yazdani, H.O.; Liu, Y.; Loughran, P.; van der Windt, D.J.; Huang, H.; Simmons, R.L.; Shiva, S.; Tai, S.; Tsung, A. Hypoxia mediates mitochondrial biogenesis in hepatocellular carcinoma to promote tumor growth through HMGB1 and TLR9 interaction. Hepatology 2017, 66, 182–197. [Google Scholar] [CrossRef]
- Schito, L.; Rey, S. Cell-Autonomous Metabolic Reprogramming in Hypoxia. Trends Cell Biol. 2018, 28, 128–142. [Google Scholar] [CrossRef]
- Bellot, G.; Garcia-Medina, R.; Gounon, P.; Chiche, J.; Roux, D.; Pouyssegur, J.; Mazure, N.M. Hypoxia-Induced Autophagy Is Mediated through Hypoxia-Inducible Factor Induction of BNIP3 and BNIP3L via Their BH3 Domains. Mol. Cell. Biol. 2009, 29, 2570–2581. [Google Scholar] [CrossRef]
- Sijtsema, W.Y. Sport voor mensen met een handicap. Tijdschrift voor Ziekenverpleging 1977, 30, 115–125. [Google Scholar] [PubMed]
- Peng, W.X.; Xiong, E.M.; Ge, L.; Wan, Y.Y.; Zhang, C.L.; Du, F.Y.; Xu, M.; Bhat, R.A.; Jin, J.; Gong, A.H. Egr-1 promotes hypoxia-induced autophagy to enhance chemo-resistance of hepatocellular carcinoma cells. Exp. Cell Res. 2016, 340, 62–70. [Google Scholar] [CrossRef] [PubMed]
- Bjelland, S.; Seeberg, E. Mutagenicity, toxicity and repair of DNA base damage induced by oxidation. Mutat. Res. 2003, 531, 37–80. [Google Scholar] [CrossRef]
- Scherz-Shouval, R.; Elazar, Z. ROS, mitochondria and the regulation of autophagy. Trends Cell Biol. 2007, 17, 422–427. [Google Scholar] [CrossRef] [PubMed]
- Han, D.; Williams, E.; Cadenas, E. Mitochondrial respiratory chain-dependent generation of superoxide anion and its release into the intermembrane space. Biochem. J. 2001, 353, 411–416. [Google Scholar] [CrossRef]
- Muller, F. The nature and mechanism of superoxide production by the electron transport chain: Its relevance to aging. Age 2000, 23, 227–253. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Lamark, T.; Sjøttem, E.; Larsen, K.B.; Awuh, J.A.; Øvervatn, A.; McMahon, M.; Hayes, J.D.; Johansen, T. p62/SQSTM1 is a target gene for transcription factor NRF2 and creates a positive feedback loop by inducing antioxidant response element-driven gene transcription. J. Biol. Chem. 2010, 285, 22576–22591. [Google Scholar] [CrossRef] [PubMed]
- Copple, I.M.; Lister, A.; Obeng, A.D.; Kitteringham, N.R.; Jenkins, R.E.; Layfield, R.; Foster, B.J.; Goldring, C.E.; Park, B.K. Physical and functional interaction of sequestosome 1 with Keap1 regulates the Keap1-Nrf2 cell defense pathway. J. Biol. Chem. 2010, 285, 16782–16788. [Google Scholar] [CrossRef] [PubMed]
- Katsuragi, Y.; Ichimura, Y.; Komatsu, M. Regulation of the Keap1–Nrf2 pathway by p62/SQSTM1. Curr. Opin. Toxicol. 2016, 1, 54–61. [Google Scholar] [CrossRef]
- Bartolini, D.; Dallaglio, K.; Torquato, P.; Piroddi, M.; Galli, F. Nrf2-p62 autophagy pathway and its response to oxidative stress in hepatocellular carcinoma. Transl. Res. 2018, 193, 54–71. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yu, Y.; Ji, T.; Ma, R.; Chen, M.; Li, G.; Li, F.; Ding, Q.; Kang, Q.; Huang, D.; et al. Clinical implication of Keap1 and phosphorylated Nrf2 expression in hepatocellular carcinoma. Cancer Med. 2016, 5, 2678–2687. [Google Scholar] [CrossRef] [PubMed]
- Lemasters, J.J. Selective Mitochondrial Autophagy, or Mitophagy, as a Targeted Defense against Oxidative Stress, Mitochondrial Dysfunction, and Aging. Rejuv. Res. 2005, 8, 3–5. [Google Scholar] [CrossRef] [PubMed]
- Wild, P.; Dikic, I. Mitochondria get a Parkin’ ticket. Nat. Cell Biol. 2010, 12, 104–106. [Google Scholar] [CrossRef] [PubMed]
- Bedard, K.; Krause, K.-H. The NOX Family of ROS-Generating NADPH Oxidases: Physiology and Pathophysiology. Physiol. Rev. 2007, 87, 245–313. [Google Scholar] [CrossRef] [PubMed]
- Lambeth, J.D. NOX enzymes and the biology of reactive oxygen. Nat. Rev. Immunol. 2004, 4, 181–189. [Google Scholar] [CrossRef] [PubMed]
- Ma, Q. Role of Nrf2 in Oxidative Stress and Toxicity. Annu. Rev. Pharmacol. Toxicol. 2013, 53, 401–426. [Google Scholar] [CrossRef] [PubMed]
- Twig, G.; Elorza, A.; Molina, A.J.A.; Mohamed, H.; Wikstrom, J.D.; Walzer, G.; Stiles, L.; Haigh, S.E.; Katz, S.; Las, G.; et al. Fission and selective fusion govern mitochondrial segregation and elimination by autophagy. EMBO J. 2008, 27, 433–446. [Google Scholar] [CrossRef]
- Liu, K.; Lee, J.; Kim, J.Y.; Wang, L.; Tian, Y.; Chan, S.T.; Cho, C.; Machida, K.; Chen, D.; Ou, J.-H.J. Mitophagy Controls the Activities of Tumor Suppressor p53 to Regulate Hepatic Cancer Stem Cells. Mol. Cell 2017, 68, 281–292. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.; Cai, Y.; Li, Y.; Li, Y.; Hu, N.; Ma, S.; Hu, S.; Zhu, P.; Wang, W.; Zhou, H. Yap promotes hepatocellular carcinoma metastasis and mobilization via governing cofilin/F-actin/lamellipodium axis by regulation of JNK/Bnip3/SERCA/CaMKII pathways. Redox Biol. 2018, 14, 59–71. [Google Scholar] [CrossRef]
- Inbal, B.; Bialik, S.; Sabanay, I.; Shani, G.; Kimchi, A. DAP kinase and DRP-1 mediate membrane blebbing and the formation of autophagic vesicles during programmed cell death. J. Cell Biol. 2002, 157, 455–468. [Google Scholar] [CrossRef] [PubMed]
- Dasari, S.K.; Bialik, S.; Levin-Zaidman, S.; Levin-Salomon, V.; Merrill, A.H.; Futerman, A.H.; Kimchi, A. Signalome-wide RNAi screen identifies GBA1 as a positive mediator of autophagic cell death. Cell Death Differ. 2017, 24, 1288–1302. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, S.; Konishi, A.; Nishida, Y.; Mizuta, T.; Nishina, H.; Yamamoto, A.; Tsujimoto, Y. Involvement of JNK in the regulation of autophagic cell death. Oncogene 2010, 29, 2070–2082. [Google Scholar] [CrossRef] [PubMed]
- Arakawa, S.; Tsujioka, M.; Yoshida, T.; Tajima-Sakurai, H.; Nishida, Y.; Matsuoka, Y.; Yoshino, I.; Tsujimoto, Y.; Shimizu, S. Role of Atg5-dependent cell death in the embryonic development of Bax/Bak double-knockout mice. Cell Death Differ. 2017, 24, 1598–1608. [Google Scholar] [CrossRef] [PubMed]
- Reef, S.; Zalckvar, E.; Shifman, O.; Bialik, S.; Sabanay, H.; Oren, M.; Kimchi, A. A short mitochondrial form of p19ARF induces autophagy and caspase-independent cell death. Mol. Cell 2006, 22, 463–475. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, H.; Miyashita, T.; Nakano, Y.; Yamamoto, D. HSpin1, a transmembrane protein interacting with Bcl-2/Bcl-xL, induces a caspase-independent autophagic cell death. Cell Death Differ. 2003, 10, 798–807. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Liu, Y.; Varley, P.; Chang, Y.; He, X.X.; Huang, H.; Tang, D.; Lotze, M.T.; Lin, J.; Tsung, A. High-mobility group box 1 promotes hepatocellular carcinoma progression through MIR-21-mediated matrix metalloproteinase activity. Cancer Res. 2015, 75, 1645–1656. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Nace, G.W.; McDonald, K.-A.; Tai, S.; Klune, J.R.; Rosborough, B.R.; Ding, Q.; Loughran, P.; Zhu, X.; Beer-Stolz, D.; et al. Hepatocyte-specific high-mobility group box 1 deletion worsens the injury in liver ischemia/reperfusion: A role for intracellular high-mobility group box 1 in cellular protection. Hepatology 2014, 59, 1984–1997. [Google Scholar] [CrossRef]
- Pan, B.; Chen, D.; Huang, J.; Wang, R.; Feng, B.; Song, H.; Chen, L. HMGB1-mediated autophagy promotes docetaxel resistance in human lung adenocarcinoma. Mol. Cancer 2014, 13, 165. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Yang, M.; Yang, L.; Yu, Y.; Xie, M.; Zhu, S.; Kang, R.; Tang, D.; Jiang, Z.; Yuan, W.; et al. HMGB1 regulates autophagy through increasing transcriptional activities of JNK and ERK in human myeloid leukemia cells. BMB Rep. 2011, 44, 601–606. [Google Scholar] [CrossRef]
- Liu, Y.; Yan, W.; Tohme, S.; Chen, M.; Fu, Y.; Tian, D.; Lotze, M.; Tang, D.; Tsung, A. Hypoxia induced HMGB1 and mitochondrial DNA interactions mediate tumor growth in hepatocellular carcinoma through Toll-like receptor 9. J. Hepatol. 2015, 63, 114–121. [Google Scholar] [CrossRef] [PubMed]
- Debnath, J. Detachment-induced autophagy during anoikis and lumen formation in epithelial acini. Autophagy 2008, 4, 351–353. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.-F.; Shi, Y.-H.; Shen, Y.-H.; Ding, Z.-B.; Ke, A.-W.; Zhou, J.; Qiu, S.-J.; Fan, J. Promoting colonization in metastatic HCC cells by modulation of autophagy. PLoS ONE 2013, 8, e74407. [Google Scholar] [CrossRef] [PubMed]
- Seyfried, T.N.; Huysentruyt, L.C. On the Origin of Cancer Metastasis. Crit. Rev. Oncog. 2013, 18, 43–73. [Google Scholar] [CrossRef] [PubMed]
- Avivar-Valderas, A.; Salas, E.; Bobrovnikova-Marjon, E.; Diehl, J.A.; Nagi, C.; Debnath, J.; Aguirre-Ghiso, J.A. PERK Integrates Autophagy and Oxidative Stress Responses To Promote Survival during Extracellular Matrix Detachment. Mol. Cell. Biol. 2011, 31, 3616–3629. [Google Scholar] [CrossRef] [PubMed]
- Sharifi, M.N.; Mowers, E.E.; Drake, L.E.; Collier, C.; Chen, H.; Zamora, M.; Mui, S.; Macleod, K.F. Autophagy Promotes Focal Adhesion Disassembly and Cell Motility of Metastatic Tumor Cells through the Direct Interaction of Paxillin with LC3. Cell Rep. 2016, 15, 1660–1672. [Google Scholar] [CrossRef]
- Li, J.; Yang, B.; Zhou, Q.; Wu, Y.; Shang, D.; Guo, Y.; Song, Z.; Zheng, Q.; Xiong, J. Autophagy promotes hepatocellular carcinoma cell invasion through activation of epithelial–mesenchymal transition. Carcinogenesis 2013, 34, 1343–1351. [Google Scholar] [CrossRef]
- Peng, Y.F.; Shi, Y.H.; Ding, Z.B; Ke, A.W.; Gu, C.Y.; Hui, B.; Zhou, J.; Qiu, S.J.; Dai, Z.; Fan, J. Autophagy inhibition suppresses pulmonary metastasis of HCC in mice via impairing anoikis resistance and colonization of HCC cells. Autophagy 2013, 9, 2056–2068. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, Y.; Feng, T.; Su, G.; He, J.; Gao, W.; Shen, Y.; Liu, X. Fluid shear stress promotes autophagy in hepatocellular carcinoma cells. Int. J. Biol. Sci. 2018, 14, 1277–1290. [Google Scholar] [CrossRef]
- Shen, H.-M.; Codogno, P. Autophagic cell death: Loch Ness monster or endangered species? Autophagy 2011, 7, 457–465. [Google Scholar] [CrossRef]
- Boya, P.; Gonzalez-Polo, R.-A.; Casares, N.; Perfettini, J.-L.; Dessen, P.; Larochette, N.; Metivier, D.; Meley, D.; Souquere, S.; Yoshimori, T.; et al. Inhibition of Macroautophagy Triggers Apoptosis. Mol. Cell. Biol. 2005, 25, 1025–1040. [Google Scholar] [CrossRef]
- Cheng, A.-L.; Kang, Y.-K.; Chen, Z.; Tsao, C.-J.; Qin, S.; Kim, J.S.; Luo, R.; Feng, J.; Ye, S.; Yang, T.-S.; et al. Efficacy and safety of sorafenib in patients in the Asia-Pacific region with advanced hepatocellular carcinoma: A phase III randomised, double-blind, placebo-controlled trial. Lancet Oncol. 2009, 10, 25–34. [Google Scholar] [CrossRef]
- Abdel-Rahman, O.; Fouad, M. Sorafenib-based combination as a first line treatment for advanced hepatocellular carcinoma: A systematic review of the literature. Crit. Rev. Oncol. Hematol. 2014, 91, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Decaens, T.; Luciani, A.; Itti, E.; Hulin, A.; Roudot-Thoraval, F.; Laurent, A.; Zafrani, E.S.; Mallat, A.; Duvoux, C. Phase II study of sirolimus in treatment-naive patients with advanced hepatocellular carcinoma. Dig. Liver Dis. 2012, 44, 610–616. [Google Scholar] [CrossRef]
- Adnane, L.; Trail, P.A.; Taylor, I.; Wilhelm, S.M. Sorafenib (BAY 43-9006, Nexavar®), a Dual-Action Inhibitor That Targets RAF/MEK/ERK Pathway in Tumor Cells and Tyrosine Kinases VEGFR/PDGFR in Tumor Vasculature. Methods Enzymol. 2006, 407, 597–612. [Google Scholar] [PubMed]
- Shimizu, S.; Takehara, T.; Hikita, H.; Kodama, T.; Tsunematsu, H.; Miyagi, T.; Hosui, A.; Ishida, H.; Tatsumi, T.; Kanto, T.; et al. Inhibition of autophagy potentiates the antitumor effect of the multikinase inhibitor sorafenib in hepatocellular carcinoma. Int. J. Cancer 2012, 131, 548–557. [Google Scholar] [CrossRef] [PubMed]
- Gedaly, R.; Angulo, P.; Hundley, J.; Daily, M.F.; Chen, C.; Evers, B.M. PKI-587 and sorafenib targeting PI3K/AKT/mTOR and Ras/Raf/MAPK pathways synergistically inhibit HCC cell proliferation. J. Surg. Res. 2012, 176, 542–548. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Hernández, M.A.; González, R.; de la Rosa, Á.J.; Gallego, P.; Ordóñez, R.; Navarro-Villarán, E.; Contreras, L.; Rodríguez-Arribas, M.; González-Gallego, J.; Álamo-Martínez, J.M.; et al. Molecular characterization of autophagic and apoptotic signaling induced by sorafenib in liver cancer cells. J. Cell. Physiol. 2019, 234, 692–708. [Google Scholar] [CrossRef] [PubMed]
- Nishida, N.; Kitano, M.; Sakurai, T.; Kudo, M. Molecular Mechanism and Prediction of Sorafenib Chemoresistance in Human Hepatocellular Carcinoma. Dig. Dis. 2015, 33, 771–779. [Google Scholar] [CrossRef]
- Lachenmayer, A.; Toffanin, S.; Cabellos, L.; Alsinet, C.; Hoshida, Y.; Villanueva, A.; Minguez, B.; Tsai, H.W.; Ward, S.C.; Thung, S.; et al. Combination therapy for hepatocellular carcinoma: Additive preclinical efficacy of the HDAC inhibitor panobinostat with sorafenib. J. Hepatol. 2012, 56, 1343–1350. [Google Scholar] [CrossRef]
- Liang, Y.; Zheng, T.; Song, R.; Wang, J.; Yin, D.; Wang, L.; Liu, H.; Tian, L.; Fang, X.; Meng, X.; et al. Hypoxia-mediated sorafenib resistance can be overcome by EF24 through Von Hippel-Lindau tumor suppressor-dependent HIF-1α inhibition in hepatocellular carcinoma. Hepatology 2013, 57, 1847–1857. [Google Scholar] [CrossRef]
- Prieto-Domínguez, N.; Méndez-Blanco, C.; Carbajo-Pescador, S.; Fondevila, F.; García-Palomo, A.; González-Gallego, J.; Mauriz, J.L. Melatonin enhances sorafenib actions in human hepatocarcinoma cells by inhibiting mTORC1/p70S6K/HIF-1α and hypoxia-mediated mitophagy. Oncotarget 2017, 8, 91402–91414. [Google Scholar] [CrossRef] [PubMed]
- Prieto-Domínguez, N.; Ordóñez, R.; Fernández, A.; García-Palomo, A.; Muntané, J.; González-Gallego, J.; Mauriz, J.L. Modulation of Autophagy by Sorafenib: Effects on Treatment Response. Front. Pharmacol. 2016, 7, 151. [Google Scholar] [CrossRef] [PubMed]
- Zhi, X.; Zhong, Q. Autophagy in cancer. F1000Prime Rep. 2015, 7, 18. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.-B.; Hui, B.; Shi, Y.-H.; Zhou, J.; Peng, Y.-F.; Gu, C.-Y.; Yang, H.; Shi, G.-M.; Ke, A.-W.; Wang, X.-Y.; et al. Autophagy Activation in Hepatocellular Carcinoma Contributes to the Tolerance of Oxaliplatin via Reactive Oxygen Species Modulation. Clin. Cancer Res. 2011, 17, 6229–6238. [Google Scholar] [CrossRef] [PubMed]
- Sheng, J.; Qin, H.; Zhang, K.; Li, B.; Zhang, X. Targeting autophagy in chemotherapy-resistant of hepatocellular carcinoma. Am. J. Cancer Res. 2018, 8, 354–365. [Google Scholar] [PubMed]
- Toshima, T.; Shirabe, K.; Matsumoto, Y.; Yoshiya, S.; Ikegami, T.; Yoshizumi, T.; Soejima, Y.; Ikeda, T.; Maehara, Y. Autophagy enhances hepatocellular carcinoma progression by activation of mitochondrial β-oxidation. J. Gastroenterol. 2014, 49, 907–916. [Google Scholar] [CrossRef] [PubMed]
- Hu, P.; Cheng, B.; He, Y.; Wei, Z.; Wu, D.; Meng, Z. Autophagy suppresses proliferation of HepG2 cells via inhibiting glypican-3/wnt/β-catenin signaling. Oncotargets Ther. 2018, 11, 193–200. [Google Scholar] [CrossRef]
- Da Silva, E.F.G.; Krause, G.C.; Lima, K.G.; Haute, G.V.; Pedrazza, L.; Mesquita, F.C.; Basso, B.S.; Velasquez, A.C.; Nunes, F.B.; De Oliveira, J.R. Rapamycin and fructose-1,6-bisphosphate reduce the HEPG2 cell proliferation via increase of free radicals and apoptosis. Oncol. Rep. 2016, 36, 2647–2652. [Google Scholar] [CrossRef]
- Engl, T.; Rutz, J.; Maxeiner, S.; Juengel, E.; Roos, F.; Khoder, W.; Bechstein, W.O.; Nelson, K.; Tsaur, I.; Haferkamp, A.; et al. mTOR inhibition reduces growth and adhesion of hepatocellular carcinoma cells in vitro. Mol. Med. Rep. 2017, 16, 7064–7071. [Google Scholar] [CrossRef]
- Kim, J.-O.; Kim, K.-H.; Song, I.S.; Cheon, K.-S.; Kim, O.-H.; Lee, S.C.; Lee, S.K.; Kim, S.-J. Potentiation of the anticancer effects of everolimus using a dual mTORC1/2 inhibitor in hepatocellular carcinoma cells. Oncotarget 2017, 8, 2936–2948. [Google Scholar] [CrossRef]
- Tong, Y.; Huang, H.; Pan, H. Inhibition of MEK/ERK activation attenuates autophagy and potentiates pemetrexed-induced activity against HepG2 hepatocellular carcinoma cells. Biochem. Biophys. Res. Commun. 2015, 456, 86–91. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.-Y.; Tai, W.-T.; Hsieh, C.-Y.; Hsu, W.-M.; Lai, Y.-J.; Chen, L.-J.; Shiau, C.-W.; Chen, K.-F. A sorafenib derivative and novel SHP-1 agonist, SC-59, acts synergistically with radiotherapy in hepatocellular carcinoma cells through inhibition of STAT3. Cancer Lett. 2014, 349, 136–143. [Google Scholar] [CrossRef] [PubMed]
- Tai, W.-T.; Shiau, C.-W.; Chen, H.-L.; Liu, C.-Y.; Lin, C.-S.; Cheng, A.-L.; Chen, P.-J.; Chen, K.-F. Mcl-1-dependent activation of Beclin 1 mediates autophagic cell death induced by sorafenib and SC-59 in hepatocellular carcinoma cells. Cell Death Dis. 2013, 4, e485. [Google Scholar] [CrossRef] [PubMed]
- Chang, Z.; Shi, G.; Jin, J.; Guo, H.; Guo, X.; Luo, F.; Song, Y.; Jia, X. Dual PI3K/mTOR inhibitor NVP-BEZ235-induced apoptosis of hepatocellular carcinoma cell lines is enhanced by inhibitors of autophagy. Int. J. Mol. Med. 2013, 31, 1449–1456. [Google Scholar] [CrossRef] [PubMed]
- Simioni, C.; Martelli, A.M.; Cani, A.; Cetin-Atalay, R.; McCubrey, J.A.; Capitani, S.; Neri, L.M. The AKT Inhibitor MK-2206 is Cytotoxic in Hepatocarcinoma Cells Displaying Hyperphosphorylated AKT-1 and Synergizes with Conventional Chemotherapy. Oncotarget 2013, 4, 1496–1506. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Chen, G.G.; Zhang, Z.; Chun, S.; Leung, B.C.S.; Lai, P.B.S. Induction of autophagy in hepatocellular carcinoma cells by SB203580 requires activation of AMPK and DAPK but not p38 MAPK. Apoptosis 2012, 17, 325–334. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandro, R.; Refolo, M.G.; Lippolis, C.; Messa, C.; Cavallini, A.; Rossi, R.; Resta, L.; Di Carlo, A.; Carr, B.I. Reversibility of regorafenib effects in hepatocellular carcinoma cells. Cancer Chemother. Pharmacol. 2013, 72, 869–877. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.C.; Lin, C.S.; Tai, W.T.; Liu, C.Y.; Shiau, C.W.; Chen, K.F. Nilotinib induces autophagy in hepatocellular carcinoma through AMPK activation. J. Biol. Chem. 2013, 288, 18249–18259. [Google Scholar] [CrossRef]
- Ni, Z.; Wang, B.; Dai, X.; Ding, W.; Yang, T.; Li, X.; Lewin, S.; Xu, L.; Lian, J.; He, F. HCC cells with high levels of Bcl-2 are resistant to ABT-737 via activation of the ROS-JNK-autophagy pathway. Free Radic. Biol. Med. 2014, 70, 194–203. [Google Scholar] [CrossRef]
- Gao, M.; Yeh, P.Y.; Lu, Y.S.; Hsu, C.H.; Chen, K.F.; Lee, W.C.; Feng, W.C.; Chen, C.S.; Kuo, M.L.; Cheng, A.L. OSU-03012, a novel celecoxib derivative, induces reactive oxygen species-related autophagy in hepatocellular carcinoma. Cancer Res. 2008, 68, 9348–9357. [Google Scholar] [CrossRef]
- Saeki, I.; Terai, S.; Fujisawa, K.; Takami, T.; Yamamoto, N.; Matsumoto, T.; Hirose, Y.; Murata, Y.; Yamasaki, T.; Sakaida, I. Bortezomib induces tumor-specific cell death and growth inhibition in hepatocellular carcinoma and improves liver fibrosis. J. Gastroenterol. 2013, 48, 738–750. [Google Scholar] [CrossRef] [PubMed]
- Du, H.; Yang, W.; Chen, L.; Shi, M.; Seewoo, V.; Wang, J.; Lin, A.; Liu, Z.; Qiu, W. Role of autophagy in resistance to oxaliplatin in hepatocellular carcinoma cells. Oncol. Rep. 2012, 27, 143–150. [Google Scholar] [PubMed]
- Chen, L.H.; Loong, C.C.; Su, T.L.; Lee, Y.J.; Chu, P.M.; Tsai, M.L.; Tsai, P.H.; Tu, P.H.; Chi, C.W.; Lee, H.C.; et al. Autophagy inhibition enhances apoptosis triggered by BO-1051, an N-mustard derivative, and involves the ATM signaling pathway. Biochem. Pharmacol. 2011, 81, 594–605. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.J.; Dong, L.W.; Tan, Y.X.; Yang, G.Z.; Pan, Y.F.; Li, Z.; Tang, L.; Wang, M.; Wang, Q.; Wang, H.Y. Inhibition of active autophagy induces apoptosis and increases chemosensitivity in cholangiocarcinoma. Lab. Invest. 2011, 91, 1146–1157. [Google Scholar] [CrossRef] [PubMed]
- Rong, L.-W.; Wang, R.-X.; Zheng, X.-L.; Feng, X.-Q.; Zhang, L.; Zhang, L.; Lin, Y.; Li, Z.-P.; Wang, X. Combination of wogonin and sorafenib effectively kills human hepatocellular carcinoma cells through apoptosis potentiation and autophagy inhibition. Oncol. Lett. 2017, 13, 5028–5034. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Jiang, K.; Liu, P.; Zhang, X.; Dong, X.; Gao, J.; Liu, Q.; Barr, M.P.; Zhang, Q.; Hou, X.; et al. Bafilomycin A1 induces caspase-independent cell death in hepatocellular carcinoma cells via targeting of autophagy and MAPK pathways. Sci. Rep. 2016, 6, 37052. [Google Scholar] [CrossRef] [PubMed]
- Thoen, L.F.R.; Guimarães, E.L.M.; Dollé, L.; Mannaerts, I.; Najimi, M.; Sokal, E.; van Grunsven, L.A. A role for autophagy during hepatic stellate cell activation. J. Hepatol. 2011, 55, 1353–1360. [Google Scholar] [CrossRef] [PubMed]
- Eisenberg-Lerner, A.; Bialik, S.; Simon, H.-U.; Kimchi, A. Life and death partners: Apoptosis, autophagy and the cross-talk between them. Cell Death Differ. 2009, 16, 966–975. [Google Scholar] [CrossRef]
- Booth, L.A.; Tavallai, S.; Hamed, H.A.; Cruickshanks, N.; Dent, P. The role of cell signalling in the crosstalk between autophagy and apoptosis. Cell. Signal. 2014, 26, 549–555. [Google Scholar] [CrossRef] [PubMed]
- McFarlane, S.; Nicholl, M.J.; Sutherland, J.S.; Preston, C.M. Interaction of the human cytomegalovirus particle with the host cell induces hypoxia-inducible factor 1 alpha. Virology 2011, 414, 83–90. [Google Scholar] [CrossRef]
| Reagents | Autophagy Target | Cells Treated | Result | Ref. |
|---|---|---|---|---|
| Sirolimus (Rapamycin) | mTOR | HepG2 | Upregulation of autophagy-cell death | [117,118] |
| Temsirolimus (CCI-779) | mTOR | HepG2, Huh7 | Upregulation of autophagy-cell death | [119] |
| Everolimus (RAD001) | mTOR | Hep3B, HepG2, Huh7 | Upregulation of autophagy-cell death | [120] |
| Pemetrexed | mTORC1 | HepG2 | Upregulation of autophagy-cell death | [121] |
| SC-59 | mTORC1 | PLC5, Sk-Hep1, HepG2 and Hep3B | Upregulation of autophagy-cell death | [122,123] |
| BEZ235 | PI3K/mTOR | Hep3B, PLC/PRF/5 | Upregulation of autophagy-cell death | [124] |
| MK-2206 | AKT | SNU449, SNU378, SNU475 | Upregulation of autophagy-cell death | [125] |
| SB203580 | MAPK | HepG2, Hep3B, PLC/PRF/5, Huh-7 | Upregulation of autophagy-cell death | [126] |
| Regorafenib | Tyrosine-kinase inhibitor | HepG2 and Hep3B | Upregulation of autophagy-cell death | [127] |
| Sorafenib | Tyrosine-kinase inhibitor | Hep3B, HepG2, Huh7 | Upregulation of autophagy-cell death | [105,112] |
| Nilotinib | Tyrosine kinase inhibitor | PLC5, Huh-7, Hep3B | Upregulation of autophagy-cell death | [128] |
| ABT-737 | JNK pathway | Huh7, HepG2 | Upregulation of autophagy-cell death | [129] |
| OSU-03012 | PDK1/AKT | Huh7, Hep3B, and HepG2 | Upregulation of autophagy-cell death | [130] |
| 5-FU | Induce ER stress | HepG2, SMMC-7721, Hep3B, BEL-7402 | Upregulation of autophagy-cell survival | [114] |
| Bortezomib | Proteasome inhibitor | Huh7, HepG2, HuH7, human hepatocytes, rat hepatocytes | Upregulation of autophagy-cell death | [131] |
| Chloroquine | Lysosome | HepG2, Huh7, HA22T/VGH, Mahlavu | Downregulation of autophagy-cell death | [132,133] |
| 3-MA | PI3K III | H22, HepG2, PLC/PRF/5, SMMC7721 | Downregulation of autophagy-cell death and cell survival | [134,135] |
| Bafilomycin A1 | Lysosome | BEL7402, HepG2, Huh7, SMMC-7721 | Downregulation of autophagy-cell death | [136] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yazdani, H.O.; Huang, H.; Tsung, A. Autophagy: Dual Response in the Development of Hepatocellular Carcinoma. Cells 2019, 8, 91. https://doi.org/10.3390/cells8020091
Yazdani HO, Huang H, Tsung A. Autophagy: Dual Response in the Development of Hepatocellular Carcinoma. Cells. 2019; 8(2):91. https://doi.org/10.3390/cells8020091
Chicago/Turabian StyleYazdani, Hamza O., Hai Huang, and Allan Tsung. 2019. "Autophagy: Dual Response in the Development of Hepatocellular Carcinoma" Cells 8, no. 2: 91. https://doi.org/10.3390/cells8020091
APA StyleYazdani, H. O., Huang, H., & Tsung, A. (2019). Autophagy: Dual Response in the Development of Hepatocellular Carcinoma. Cells, 8(2), 91. https://doi.org/10.3390/cells8020091





