18F-FDG PET-Based Imaging of Myocardial Inflammation Predicts a Functional Outcome Following Transplantation of mESC-Derived Cardiac Induced Cells in a Mouse Model of Myocardial Infarction
Abstract
1. Introduction
2. Materials and Methods
2.1. Stem Cell Culture and Cardiovascular Differentiation
2.2. Animal Model
- (1)
- no intervention, isofluorane (n = 2)
- (2)
- no intervention, ketamine/xylazine (n = 2)
- (3)
- MI group, isofluorane (n = 4)
- (4)
- MI group, ketamine/xylazine (n = 6)
- (5)
- MIC group, ketamine/xylazine (n = 7)
- (6)
- MI only (n = 6)
- (7)
- MIC (n = 6)
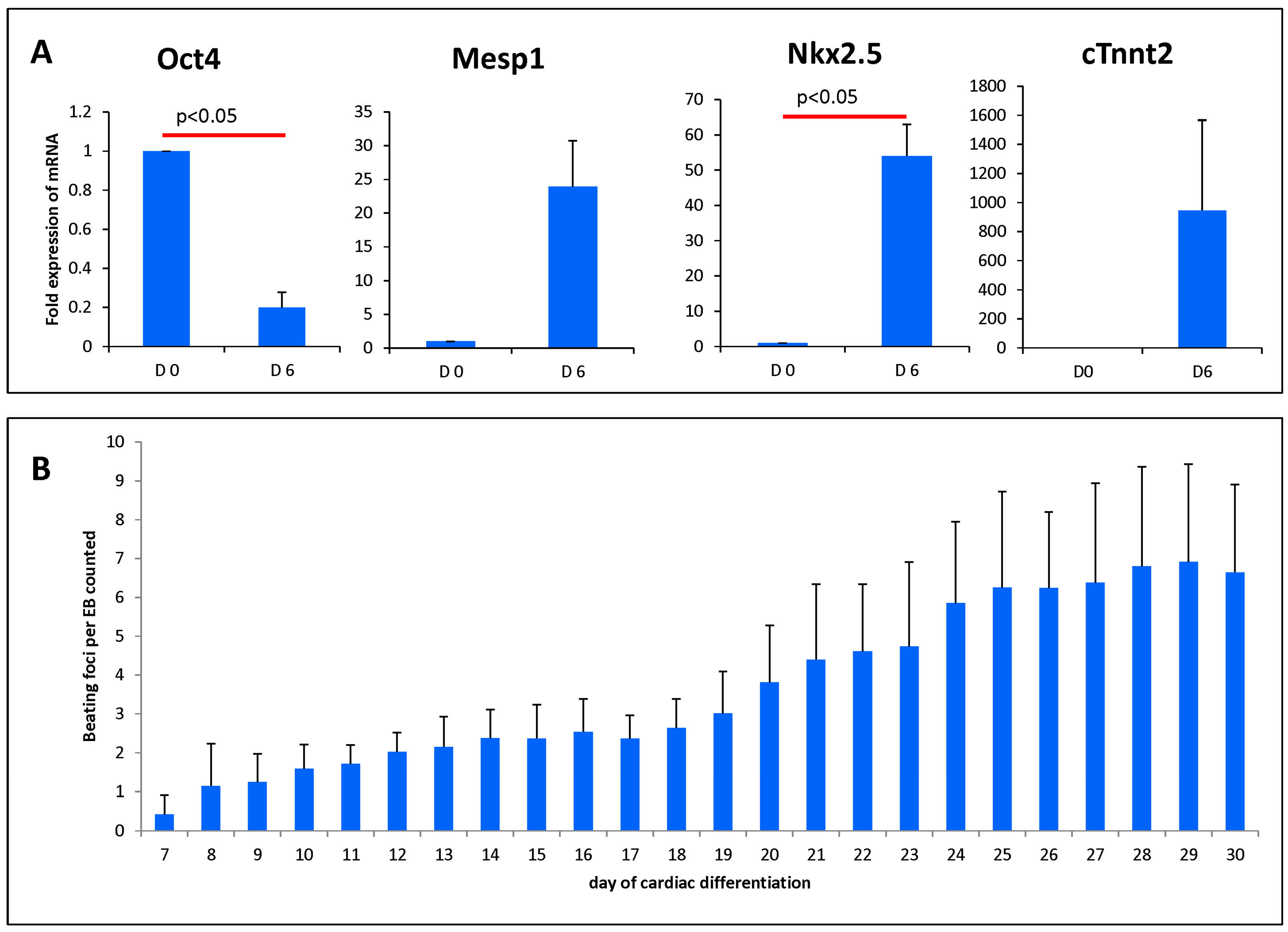
2.3. qRT-PCR
2.4. Beating Foci Analysis
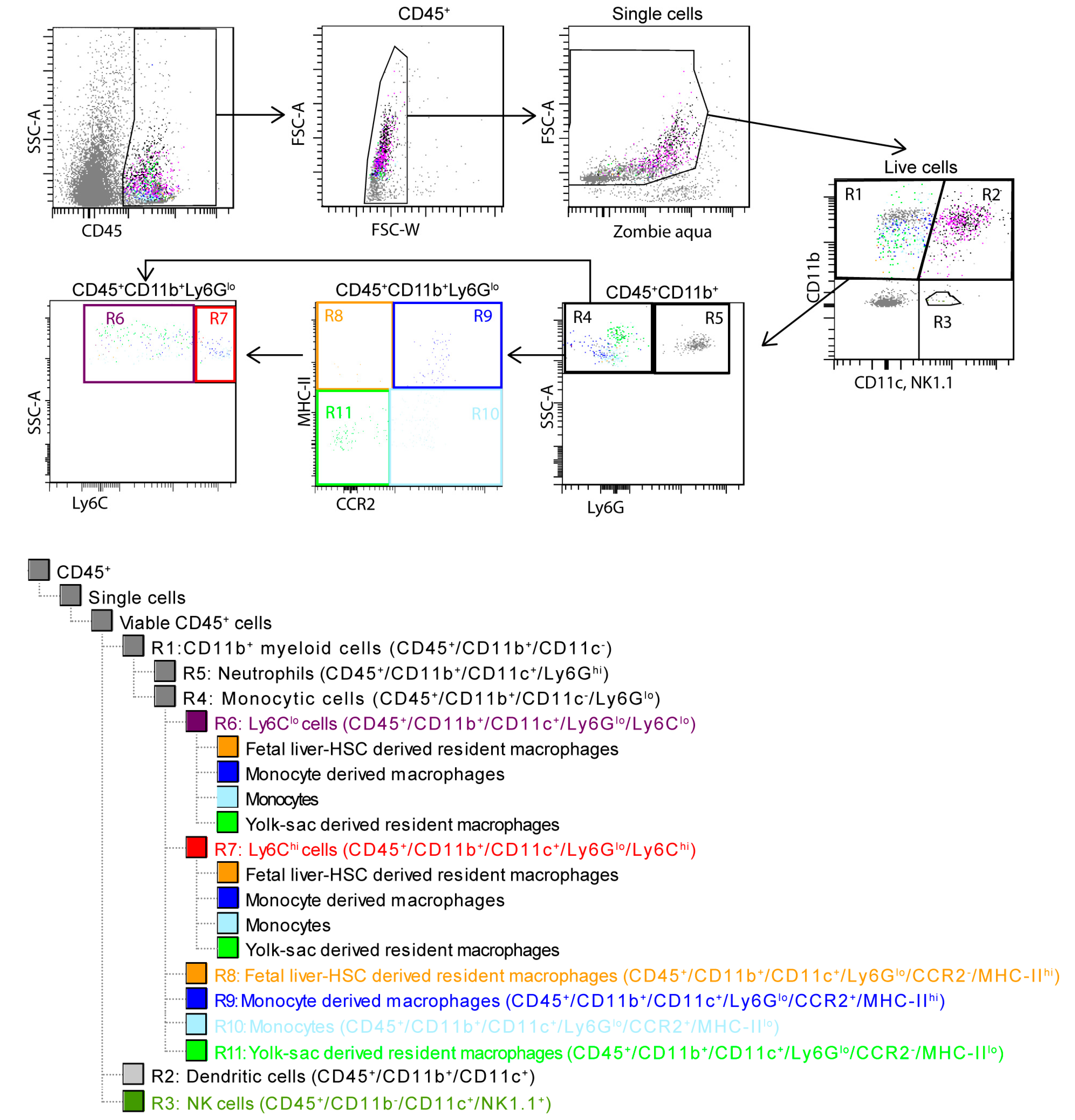
2.5. Flow Cytometry
2.6. PET Imaging
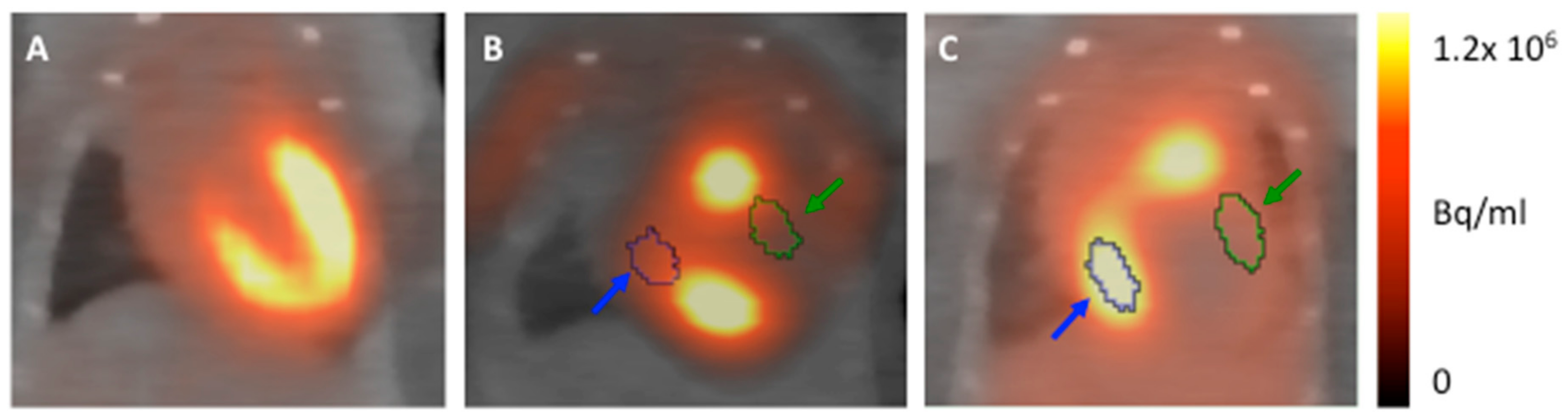
2.7. PET Image Analysis
2.8. Cardiac Magnetic Resonance Imaging
2.9. Cardiac Magnetic Resonance Analysis
2.10. Statistics
3. Results
3.1. Cardiac Induced Cells Show Increased Cardiac Markers and Beating Activity During Differentiation
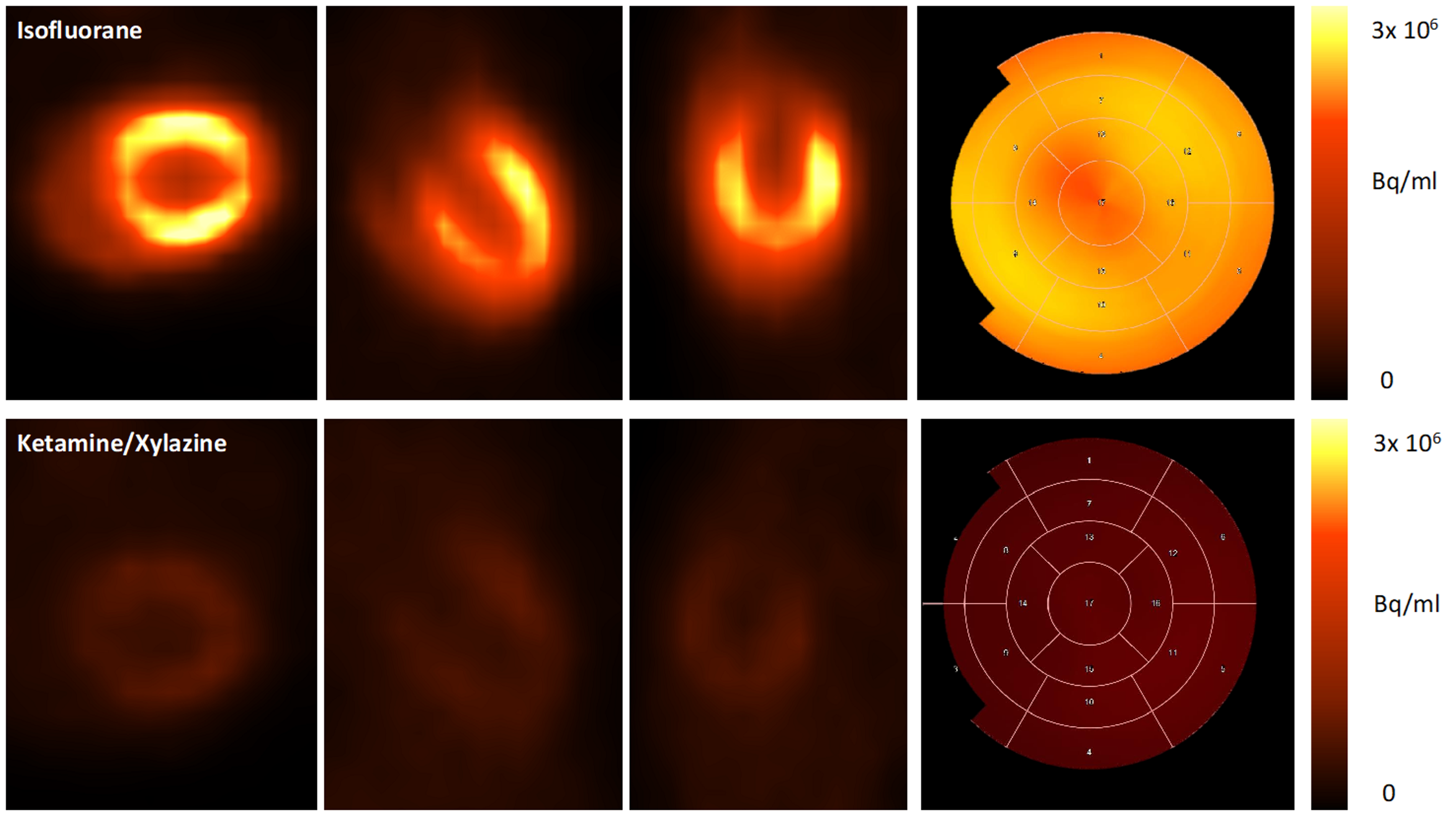
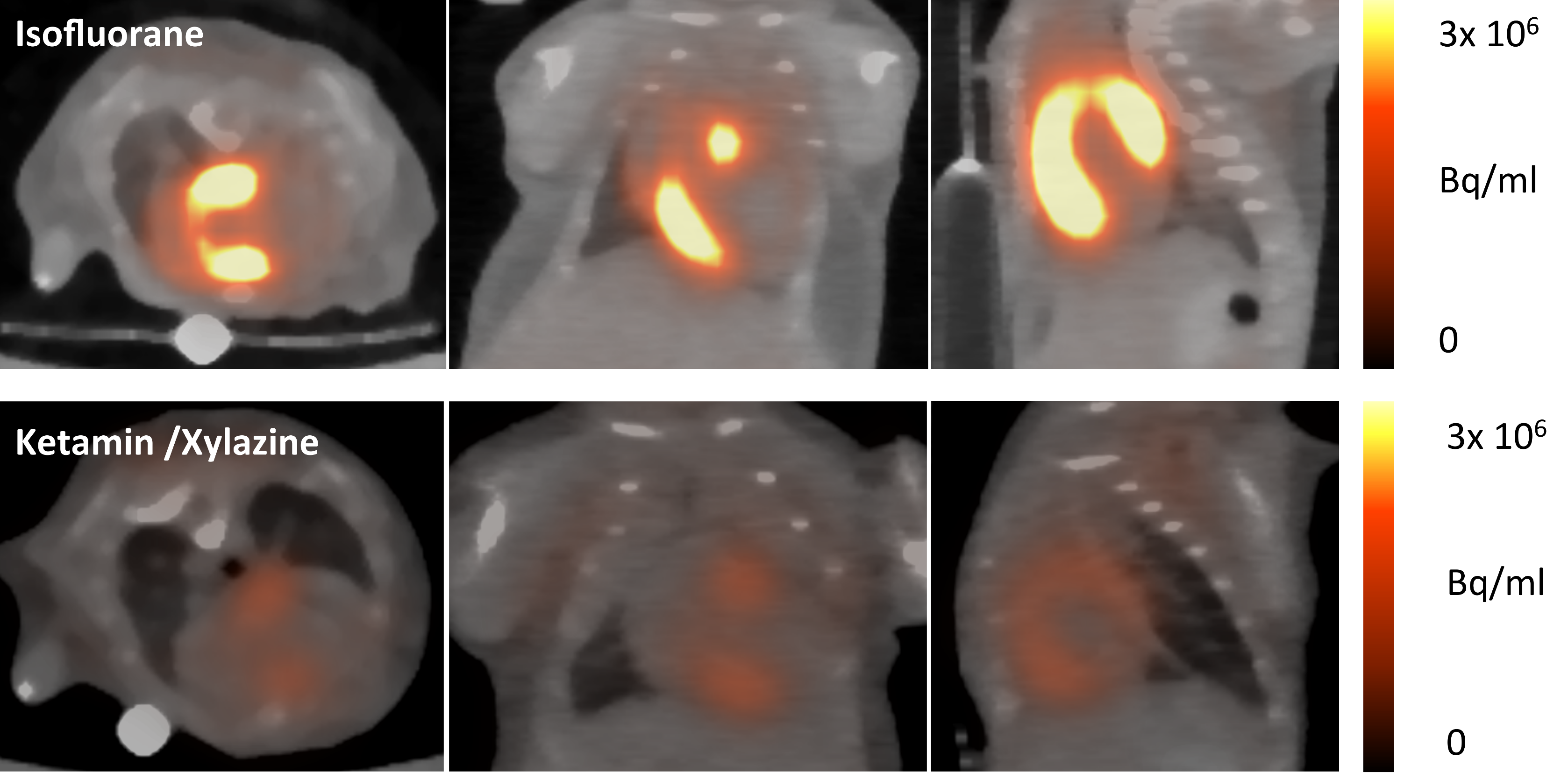
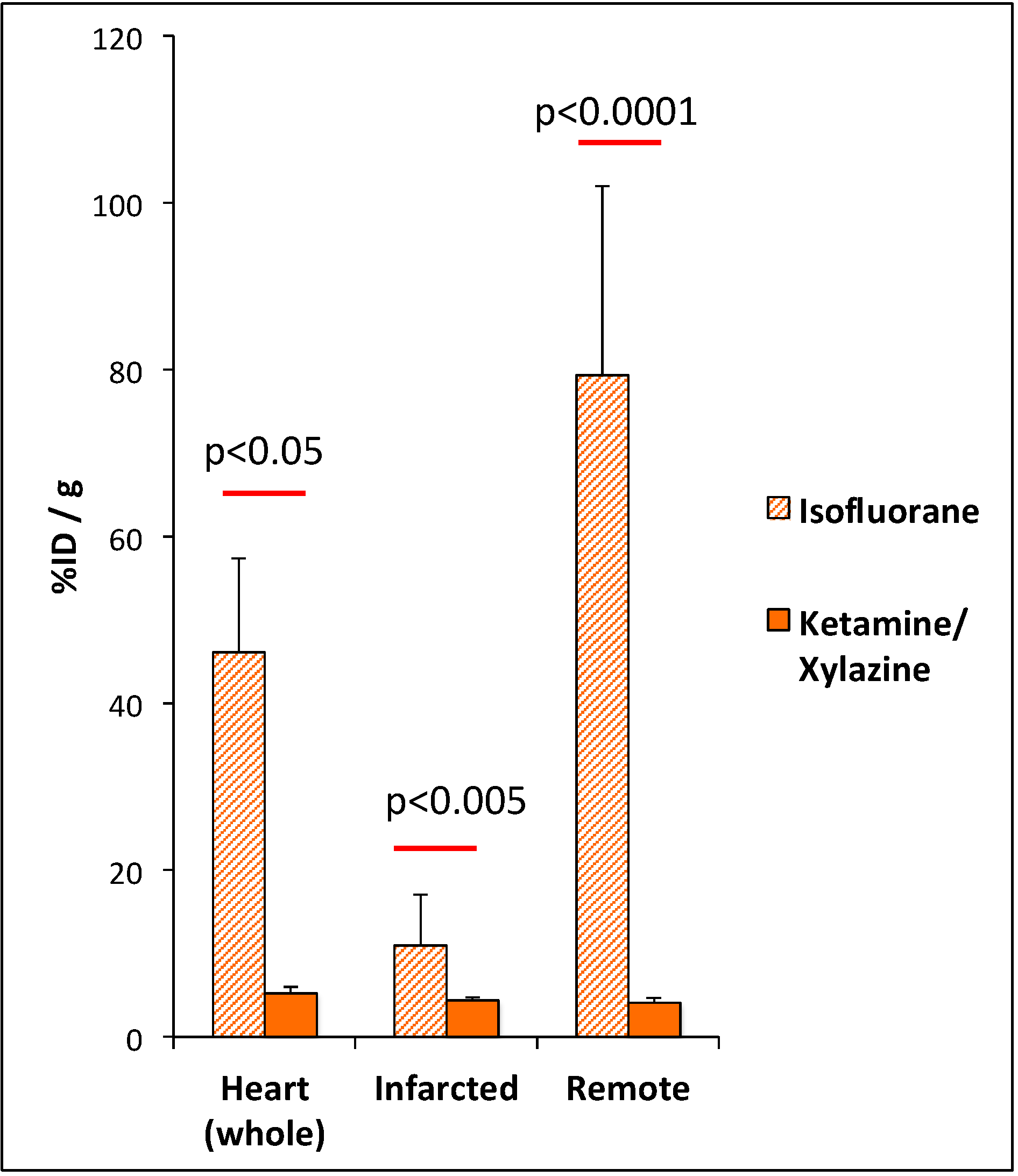
3.2. Impact of Anaesthesia on the FDG-Uptake Pattern in the Infarcted Heart
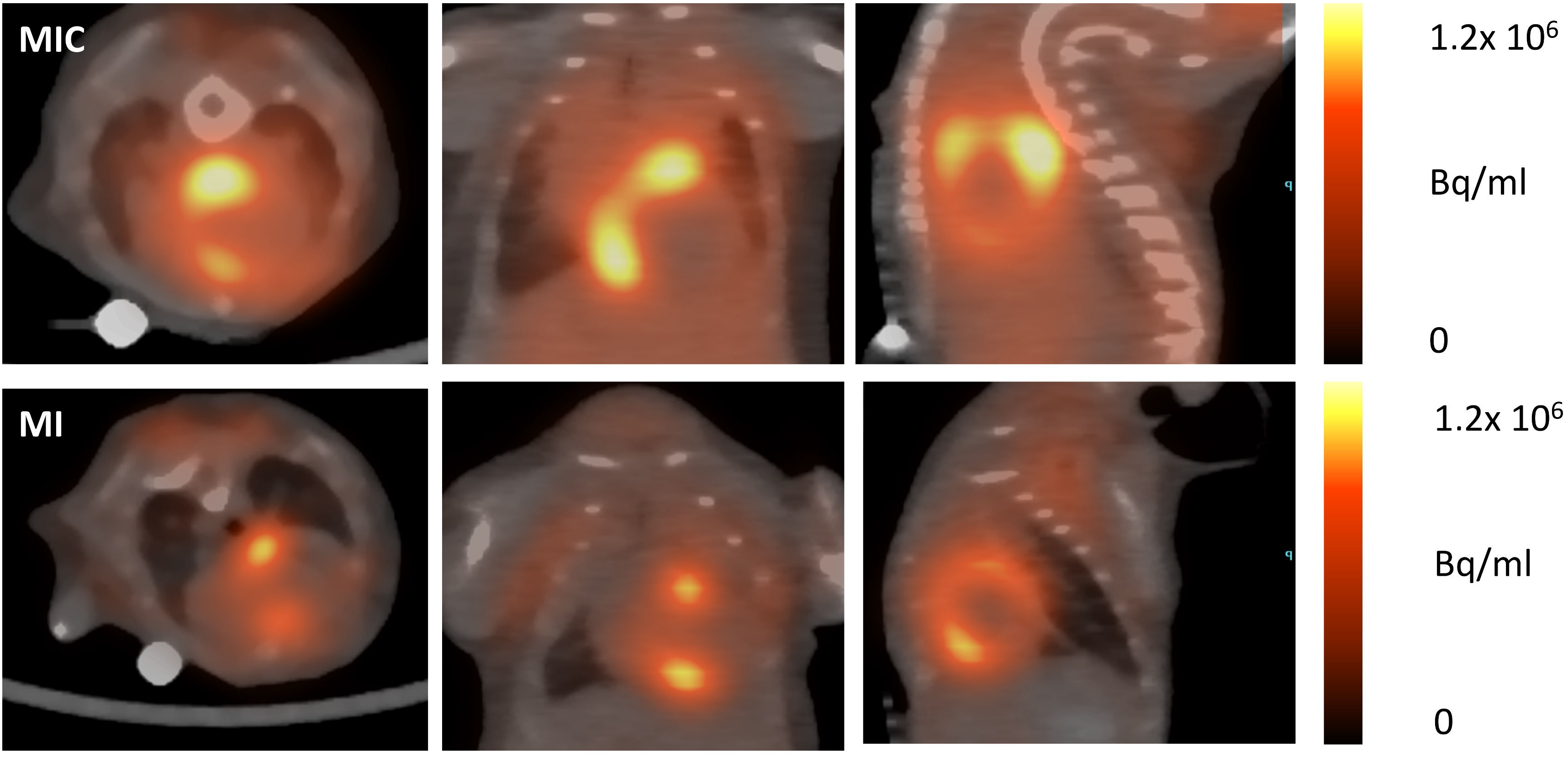
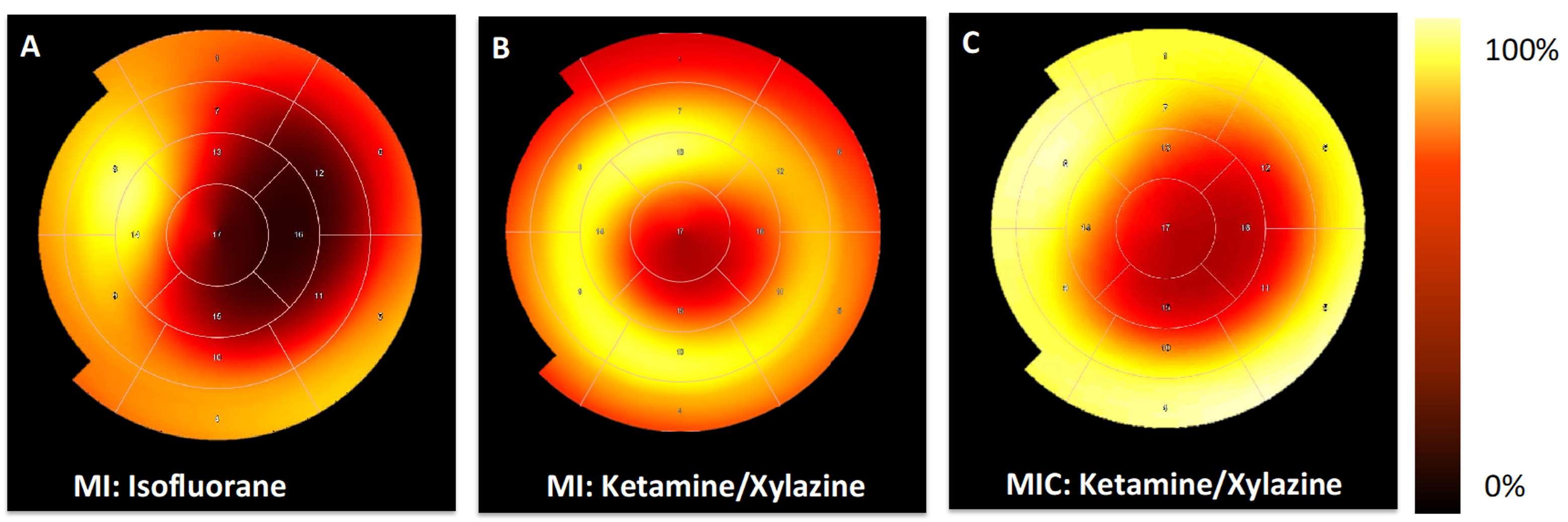
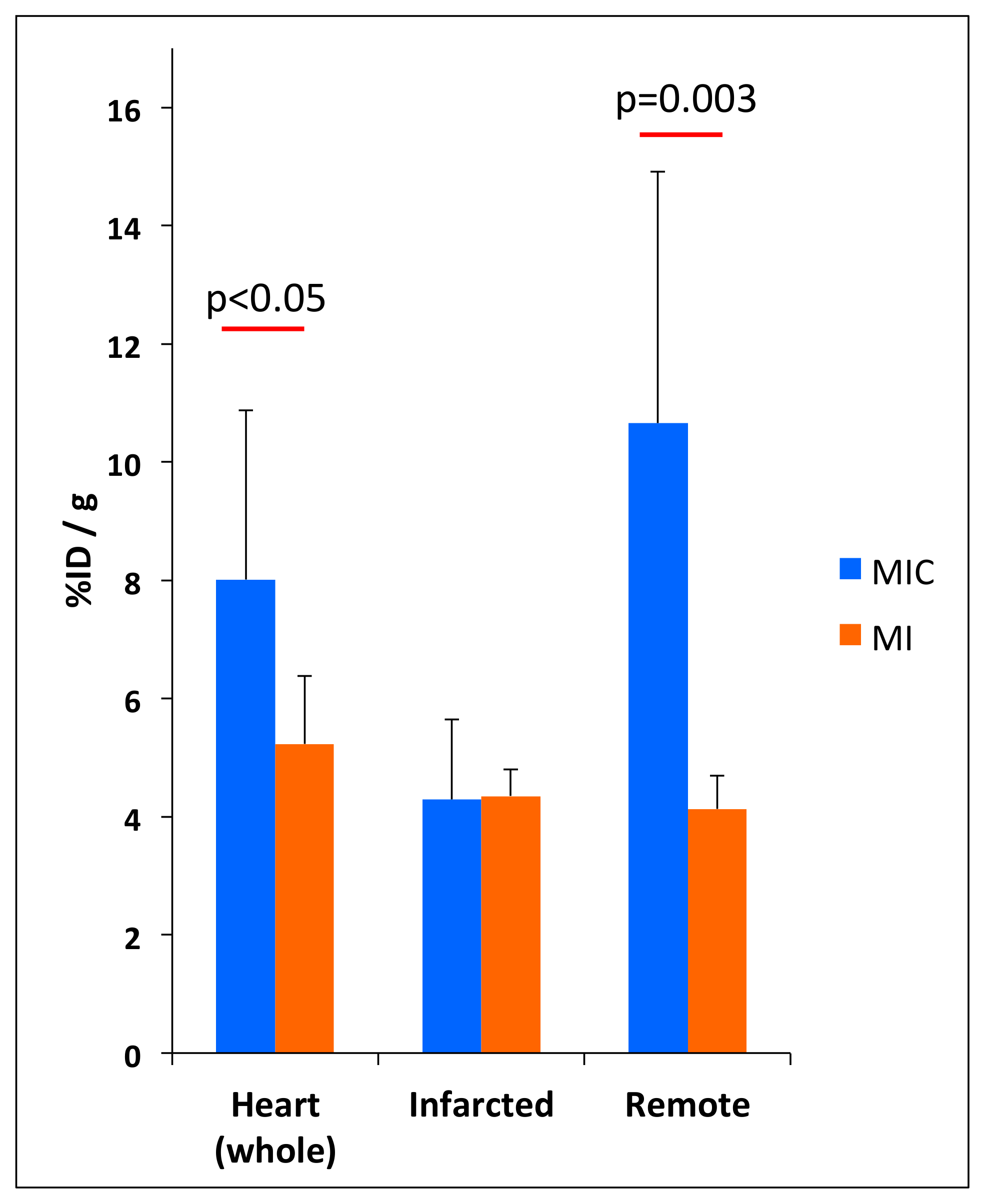
3.3. FDG-Uptake Pattern is Fundamentally Changed by Cell Transplantation
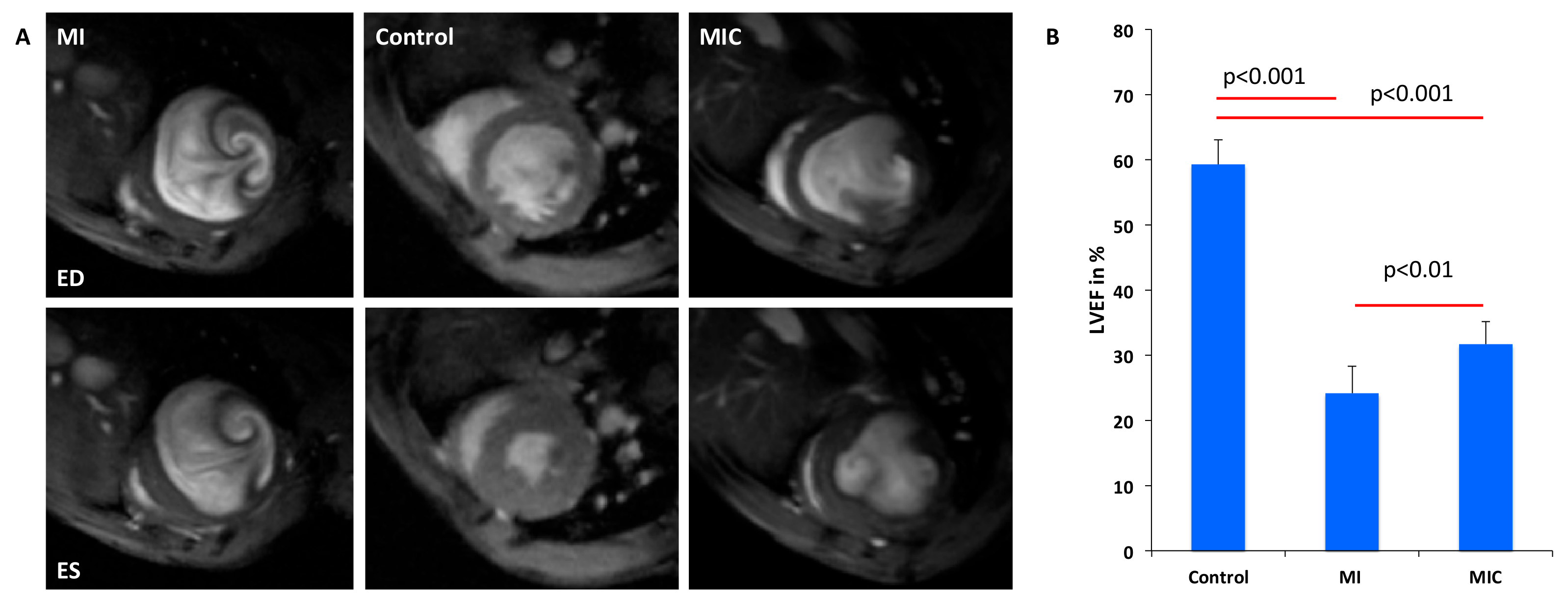
3.4. Improvement of Cardiac Function Through Cell Therapy Assessed by CMR
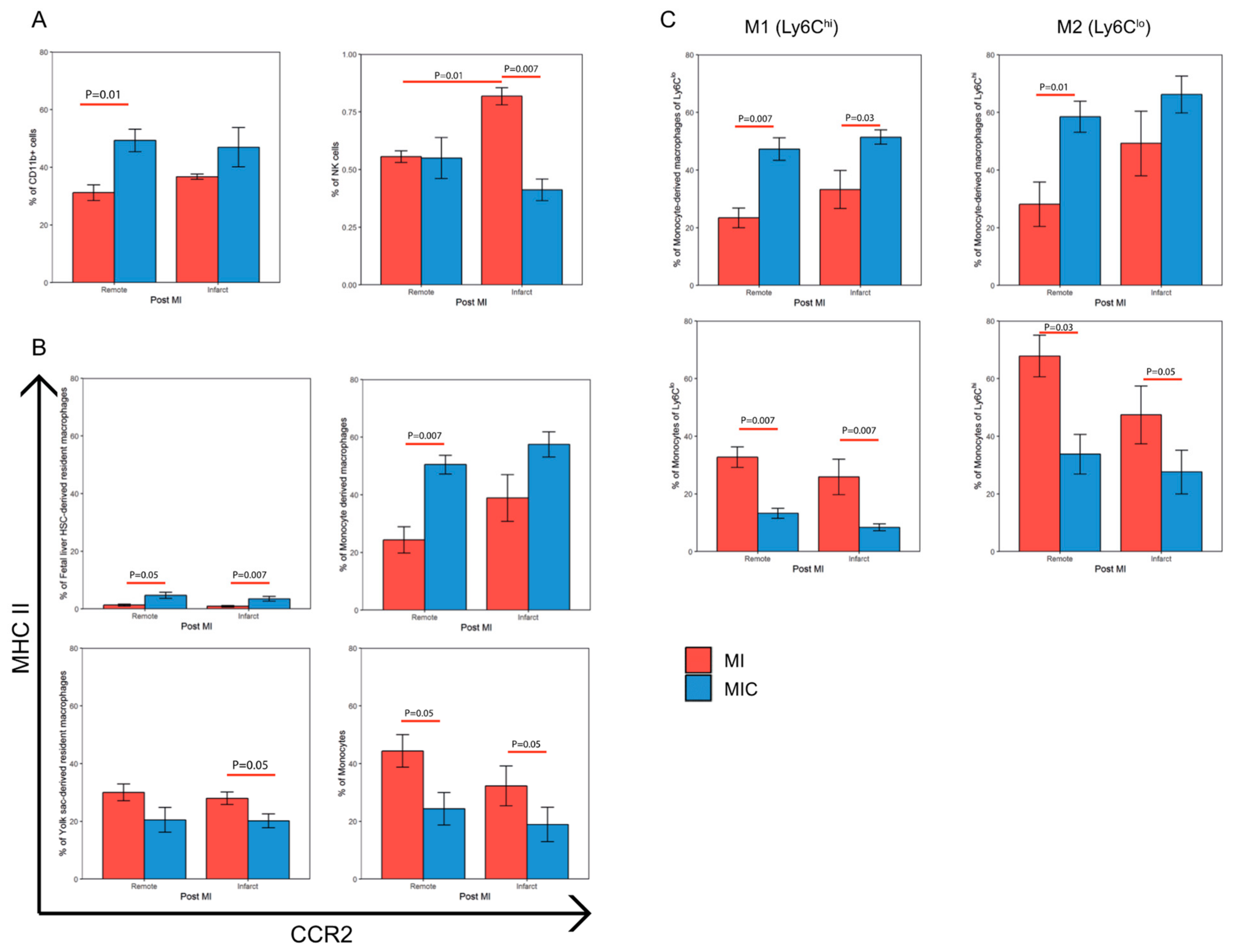
3.5. Modulation of the Immune Response Through Cell Therapy Assessed by Flow Cytometry
4. Discussion
- (A)
- Distribution pattern of CD11b+ myeloid cells at day five after MI induction can be modified by intramyocardial transplantation of CiC;
- (B)
- This change can be visualized and quantified by 18F-FDG PET using ketamine/xylazine anaesthesia;
- (C)
- The specific 18F-FDG uptake pattern correlates with functional outcome as measured by cardiac magnetic resonance imaging.
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| CiC | cardiac induced cells |
| mESCs | murine embryonic stem cells |
| MI | myocardial infarction |
| EB | embryoid bodies |
| LAD | left anterior descending coronary artery |
| FOV | field of view |
| VOI | volumes of interest |
| CMR | cardiac magnetic resonance |
| LV | left ventricle |
| LVEF | left ventricular ejection fraction |
| SD | standard deviation |
| SEM | standard error of mean |
| MNC | mononuclear cell |
| KX | ketamine/xylazine |
| PET | positron emission tomography |
| CT | computerized tomography |
References
- Lee, W.W.; Marinelli, B.; van der Laan, A.M.; Sena, B.F.; Gorbatov, R.; Leuschner, F.; Dutta, P.; Iwamoto, Y.; Ueno, T.; Begieneman, M.P.V.; et al. PET/MRI of inflammation in myocardial infarction. J. Am. Coll. Cardiol. 2012, 59, 153–163. [Google Scholar] [CrossRef] [PubMed]
- Van Hout, G.P.; Jansen of Lorkeers, S.J.; Wever, K.E.; Sena, E.S.; Kouwenberg, L.H.; van Solinge, W.W.; Macleod, M.R.; Doevendans, P.A.; Pasterkamp, G.; Chamuleau, S.A.J.; et al. Translational failure of anti-inflammatory compounds for myocardial infarction: A meta-analysis of large animal models. Cardiovasc. Res. 2016, 109, 240–248. [Google Scholar] [CrossRef] [PubMed]
- Rischpler, C.; Dirschinger, R.J.; Nekolla, S.G.; Kossmann, H.; Nicolosi, S.; Hanus, F.; van Marwick, S.; Kunze, K.P.; Meinicke, A.; Götze, K.; et al. Prospective Evaluation of 18F-Fluorodeoxyglucose Uptake in Postischemic Myocardium by Simultaneous Positron Emission Tomography/Magnetic Resonance Imaging as a Prognostic Marker of Functional Outcome. Circ. Cardiovasc. Imaging 2016, 9, e004316. [Google Scholar] [CrossRef] [PubMed]
- Thackeray, J.T.; Bankstahl, J.P.; Wang, Y.; Wollert, K.C.; Bengel, F.M. Clinically relevant strategies for lowering cardiomyocyte glucose uptake for 18F-FDG imaging of myocardial inflammation in mice. Eur. J. Nucl. Med. Mol. Imaging 2015, 42, 771–780. [Google Scholar] [CrossRef]
- Thackeray, J.T.; Bankstahl, J.P.; Wang, Y.; Korf-Klingebiel, M.; Walte, A.; Wittneben, A.; Wollert, K.C.; Bengel, F.M. Targeting post-infarct inflammation by PET imaging: Comparison of (68)Ga-citrate and (68)Ga-DOTATATE with (18)F-FDG in a mouse model. Eur. J. Nucl. Med. Mol. Imaging 2015, 42, 317–327. [Google Scholar] [CrossRef]
- Auerbach, W.; Dunmore, J.H.; Fairchild-Huntress, V.; Fang, Q.; Auerbach, A.B.; Huszar, D.; Joyner, A.L. Establishment and chimera analysis of 129/SvEv- and C57BL/6-derived mouse embryonic stem cell lines. Biotechniques 2000, 29, 1024–1028. [Google Scholar] [CrossRef]
- Thiele, F.; Voelkner, C.; Krebs, V.; Muller, P.; Jung, J.J.; Rimmbach, C.; Steinhoff, G.; Noack, T.; David, R.; Lemcke, H. Nkx2.5 Based Ventricular Programming of Murine ESC-Derived Cardiomyocytes. Cell. Physiol. Biochem. 2019, 53, 337–354. [Google Scholar]
- Cao, N.; Liu, Z.; Chen, Z.; Wang, J.; Chen, T.; Zhao, X.; Ma, Y.; Qin, L.; Kang, J.; Wei, B.; et al. Ascorbic acid enhances the cardiac differentiation of induced pluripotent stem cells through promoting the proliferation of cardiac progenitor cells. Cell Res. 2012, 22, 219–236. [Google Scholar] [CrossRef]
- Rimmbach, C.; Jung, J.J.; David, R. Generation of murine cardiac pacemaker cell aggregates based on ES-cell-programming in combination with Myh6-promoter-selection. J. Vis. Exp. 2015, e52465. [Google Scholar] [CrossRef]
- Muller, P.; Gaebel, R.; Lemcke, H.; Wiekhorst, F.; Hausburg, F.; Lang, C.; Zarniko, N.; Westphal, B.; Steinhoff, G.; David, R.; et al. Intramyocardial fate and effect of iron nanoparticles co-injected with MACS((R)) purified stem cell products. Biomaterials 2017, 135, 74–84. [Google Scholar] [CrossRef]
- Klopsch, C.; Gaebel, R.; Lemcke, H.; Beyer, M.; Vasudevan, P.; Fang, H.Y.; Quante, M.; Vollmar, B.; Skorska, A.; David, R.; et al. Vimentin-Induced Cardiac Mesenchymal Stem Cells Proliferate in the Acute Ischemic Myocardium. Cells Tissues Organs 2018, 206, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Bao, Q.; Newport, D.; Chen, M.; Stout, D.B.; Chatziioannou, A.F. Performance evaluation of the inveon dedicated PET preclinical tomograph based on the NEMA NU-4 standards. J. Nucl. Med. 2009, 50, 401–408. [Google Scholar] [CrossRef] [PubMed]
- Lang, C.; Lehner, S.; Todica, A.; Boening, G.; Franz, W.M.; Bartenstein, P.; Hacker, M.; David, R. Positron emission tomography based in-vivo imaging of early phase stem cell retention after intramyocardial delivery in the mouse model. Eur. J. Nucl. Med. Mol. Imaging 2013, 40, 1730–1738. [Google Scholar] [CrossRef] [PubMed]
- Heiberg, E.; Sjogren, J.; Ugander, M.; Carlsson, M.; Engblom, H.; Arheden, H. Design and validation of Segment—Freely available software for cardiovascular image analysis. BMC Med. Imaging 2010, 10, 1. [Google Scholar] [CrossRef]
- Aurora, A.B.; Olson, E.N. Immune modulation of stem cells and regeneration. Cell Stem Cell 2014, 15, 14–25. [Google Scholar] [CrossRef]
- Ben-Mordechai, T.; Palevski, D.; Glucksam-Galnoy, Y.; Elron-Gross, I.; Margalit, R.; Leor, J. Targeting macrophage subsets for infarct repair. J. Cardiovasc. Pharmacol. Ther. 2015, 20, 36–51. [Google Scholar] [CrossRef]
- Lavine, K.J.; Epelman, S.; Uchida, K.; Weber, K.J.; Nichols, C.G.; Schilling, J.D.; Ornitz, D.M.; Randolph, G.J.; Mann, D.L. Distinct macrophage lineages contribute to disparate patterns of cardiac recovery and remodeling in the neonatal and adult heart. Proc. Natl. Acad. Sci. USA 2014, 111, 16029–16034. [Google Scholar] [CrossRef]
- Ben-Mordechai, T.; Holbova, R.; Landa-Rouben, N.; Harel-Adar, T.; Feinberg, M.S.; Abd Elrahman, I.; Blum, G.; Epstein, F.H.; Silman, Z.; Cohen, S.; et al. Macrophage subpopulations are essential for infarct repair with and without stem cell therapy. J. Am. Coll. Cardiol. 2013, 62, 1890–1901. [Google Scholar] [CrossRef]
- Hasan, A.S.; Luo, L.; Yan, C.; Zhang, T.X.; Urata, Y.; Goto, S.; Mangoura, S.A.; Abdel-Raheem, M.H.; Zhang, S.; Li, T.-S. Cardiosphere-Derived Cells Facilitate Heart Repair by Modulating M1/M2 Macrophage Polarization and Neutrophil Recruitment. PLoS ONE 2016, 11, e0165255. [Google Scholar] [CrossRef]
- Czapla, J.; Matuszczak, S.; Wisniewska, E.; Jarosz-Biej, M.; Smolarczyk, R.; Cichon, T.; Głowala-Kosińska, M.; Śliwka, J.; Garbacz, M.; Szczypior, M.; et al. Human Cardiac Mesenchymal Stromal Cells with CD105+CD34- Phenotype Enhance the Function of Post-Infarction Heart in Mice. PLoS ONE 2016, 11, e0158745. [Google Scholar] [CrossRef]
- Lang, C.I.; Wolfien, M.; Langenbach, A.; Muller, P.; Wolkenhauer, O.; Yavari, A.; Ince, H.; Steinhoff, G.; Krause, B.; David, R.; et al. Cardiac Cell Therapies for the Treatment of Acute Myocardial Infarction: A Meta-Analysis from Mouse Studies. Cell. Physiol. Biochem. 2017, 42, 254–268. [Google Scholar] [CrossRef] [PubMed]
- Sonnenberg, G.F.; Hepworth, M.R. Functional interactions between innate lymphoid cells and adaptive immunity. Nat. Rev. Immunol. 2019, 19, 599–613. [Google Scholar] [CrossRef] [PubMed]
- Love, C.; Tomas, M.B.; Tronco, G.G.; Palestro, C.J. FDG PET of infection and inflammation. Radiographics 2005, 25, 1357–1368. [Google Scholar] [CrossRef] [PubMed]
- Peterson, L.R.; Gropler, R.J. Radionuclide imaging of myocardial metabolism. Circ. Cardiovasc. Imaging 2010, 3, 211–222. [Google Scholar] [CrossRef]
- Saha, J.K.; Xia, J.; Grondin, J.M.; Engle, S.K.; Jakubowski, J.A. Acute hyperglycemia induced by ketamine/xylazine anesthesia in rats: Mechanisms and implications for preclinical models. Exp. Biol. Med. (Maywood) 2005, 230, 777–784. [Google Scholar] [CrossRef]
- Ahmed, N.; Kansara, M.; Berridge, M.V. Acute regulation of glucose transport in a monocyte-macrophage cell line: Glut-3 affinity for glucose is enhanced during the respiratory burst. Biochem. J. 1997, 327 Pt 2, 369–375. [Google Scholar] [CrossRef]
- Schatka, I.; Bengel, F.M. Advanced imaging of cardiac sarcoidosis. J. Nucl. Med. 2014, 55, 99–106. [Google Scholar] [CrossRef]
| Target | Clone | Source |
|---|---|---|
| CD45 | 30-F11 | Biolegend |
| CD11b | M1/70 | Biolegend |
| CD11c | N418 | Biolegend |
| NK1.1 | PK136 | Biolegend |
| Ly6G | 1A8 | Biolegend |
| Ly6C | Hk1.4 | Biolegend |
| CCR2 | 475301 | R and D |
| MHC-II | AF6-120.1 | Biolegend |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vasudevan, P.; Gaebel, R.; Doering, P.; Mueller, P.; Lemcke, H.; Stenzel, J.; Lindner, T.; Kurth, J.; Steinhoff, G.; Vollmar, B.; et al. 18F-FDG PET-Based Imaging of Myocardial Inflammation Predicts a Functional Outcome Following Transplantation of mESC-Derived Cardiac Induced Cells in a Mouse Model of Myocardial Infarction. Cells 2019, 8, 1613. https://doi.org/10.3390/cells8121613
Vasudevan P, Gaebel R, Doering P, Mueller P, Lemcke H, Stenzel J, Lindner T, Kurth J, Steinhoff G, Vollmar B, et al. 18F-FDG PET-Based Imaging of Myocardial Inflammation Predicts a Functional Outcome Following Transplantation of mESC-Derived Cardiac Induced Cells in a Mouse Model of Myocardial Infarction. Cells. 2019; 8(12):1613. https://doi.org/10.3390/cells8121613
Chicago/Turabian StyleVasudevan, Praveen, Ralf Gaebel, Piet Doering, Paula Mueller, Heiko Lemcke, Jan Stenzel, Tobias Lindner, Jens Kurth, Gustav Steinhoff, Brigitte Vollmar, and et al. 2019. "18F-FDG PET-Based Imaging of Myocardial Inflammation Predicts a Functional Outcome Following Transplantation of mESC-Derived Cardiac Induced Cells in a Mouse Model of Myocardial Infarction" Cells 8, no. 12: 1613. https://doi.org/10.3390/cells8121613
APA StyleVasudevan, P., Gaebel, R., Doering, P., Mueller, P., Lemcke, H., Stenzel, J., Lindner, T., Kurth, J., Steinhoff, G., Vollmar, B., Krause, B. J., Ince, H., David, R., & Lang, C. I. (2019). 18F-FDG PET-Based Imaging of Myocardial Inflammation Predicts a Functional Outcome Following Transplantation of mESC-Derived Cardiac Induced Cells in a Mouse Model of Myocardial Infarction. Cells, 8(12), 1613. https://doi.org/10.3390/cells8121613






