Rewiring of Lipid Metabolism and Storage in Ovarian Cancer Cells after Anti-VEGF Therapy
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture and Treatments
2.2. In Vivo Experiments
2.3. Histology and Immunohistochemistry
2.4. Lipid Extraction and LC-MS Measurements
2.5. NMR Tissue Metabolomics
2.6. Lipid Droplets (LD) Evaluation
2.7. Proliferation Assay
2.8. Annexin-V Apoptosis Assay
2.9. Microarray Expression Analysis
2.10. Western Immunoblotting
- Fatty Acid Synthase (FAS) (C20G5) #3180, 1:1000 (Cell Signaling Technology; Denvers, Massachusetts, USA)
- CD36/SR-B3 (CD36) NB400-144, 1:500 (NOVUSBIO; Minneapolis, Minnesota, USA)
2.11. Statistical Analysis
3. Results
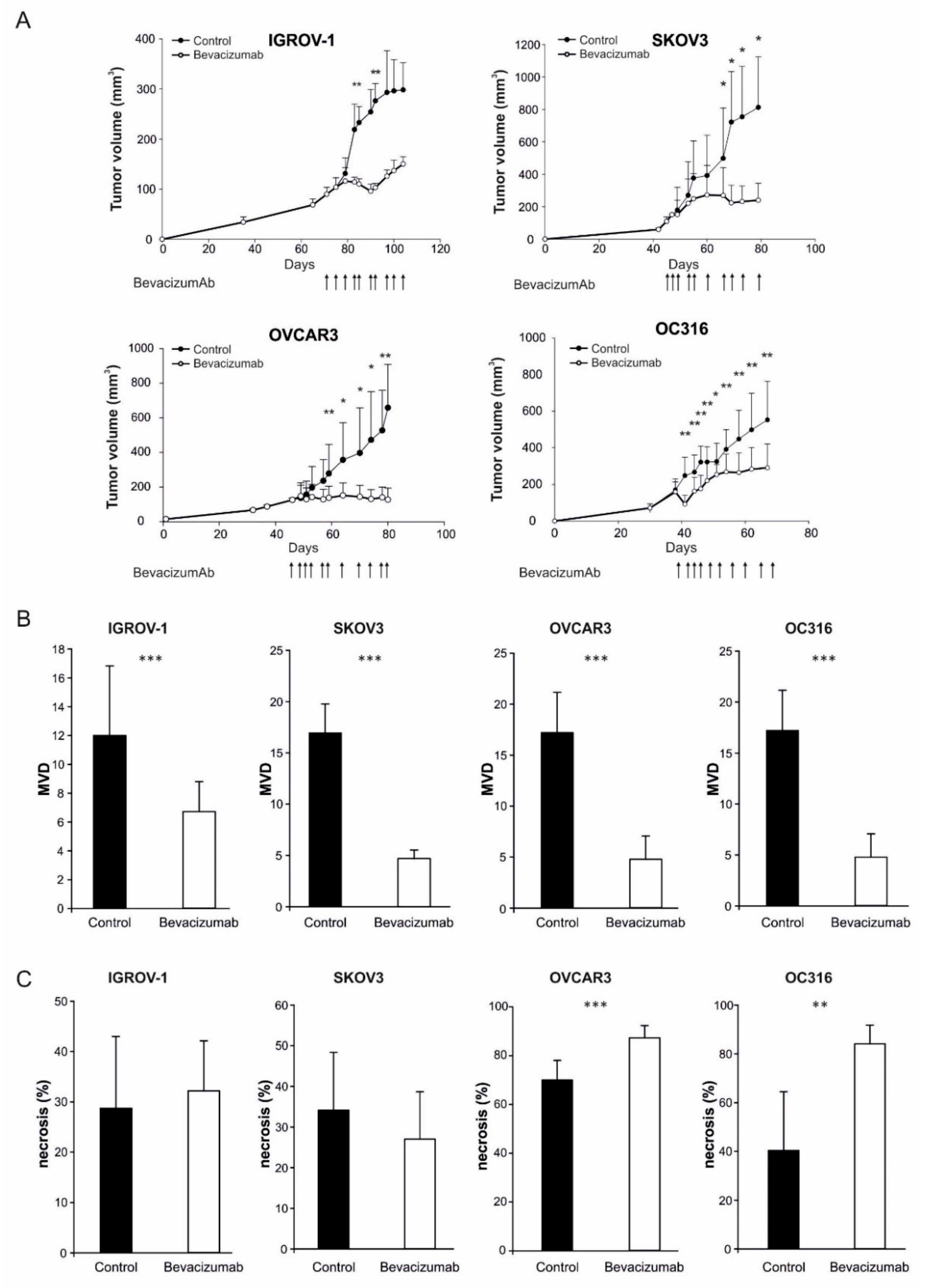
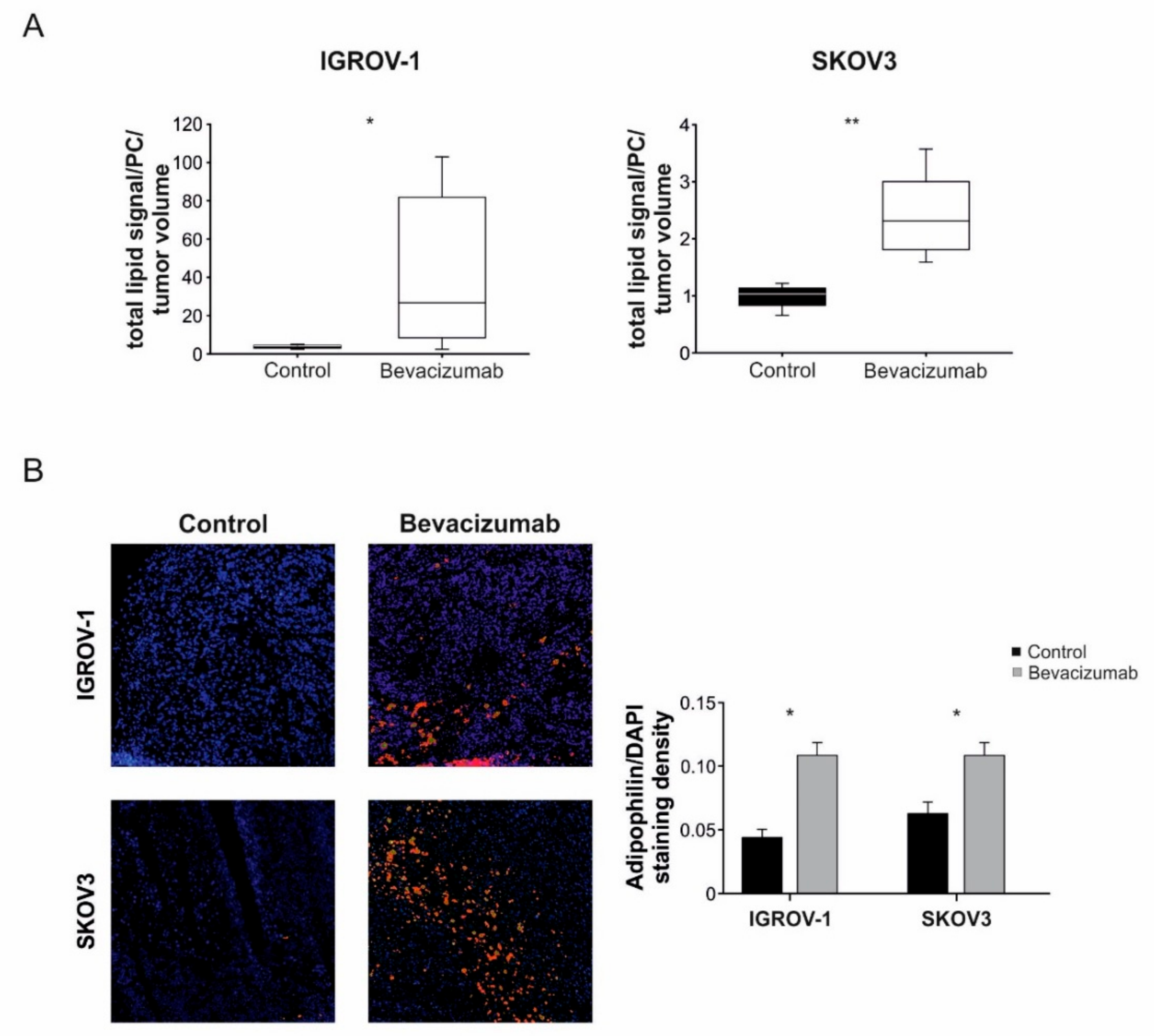
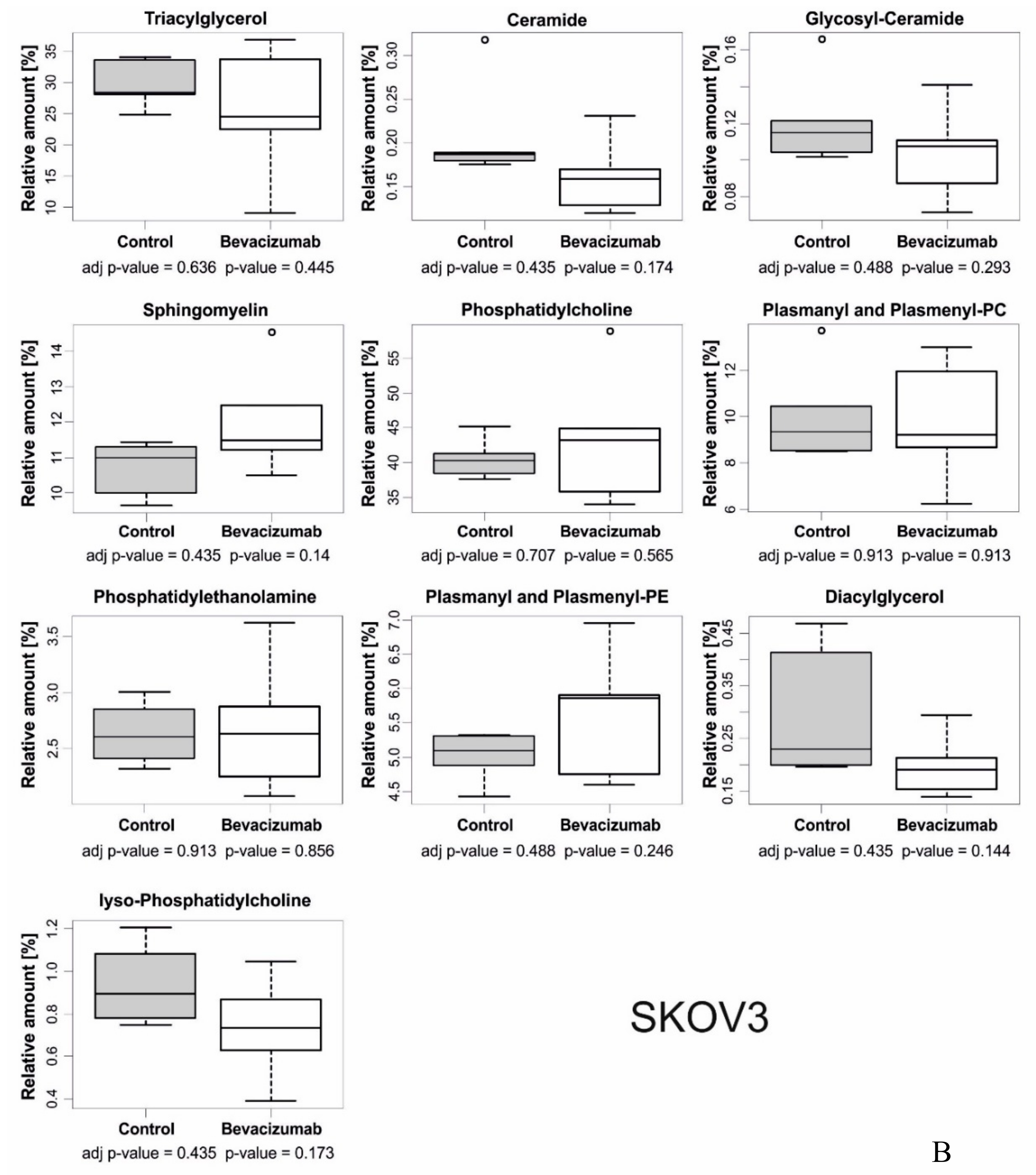
3.1. Anti-VEGF Therapy Causes Heterogeneous, Model-Dependent Effects on the Tumor Lipidome
3.2. Anti-VEGF Therapy is Associated with Increased Expression of Components of TAG and Glycerophospholipid Biosynthesis
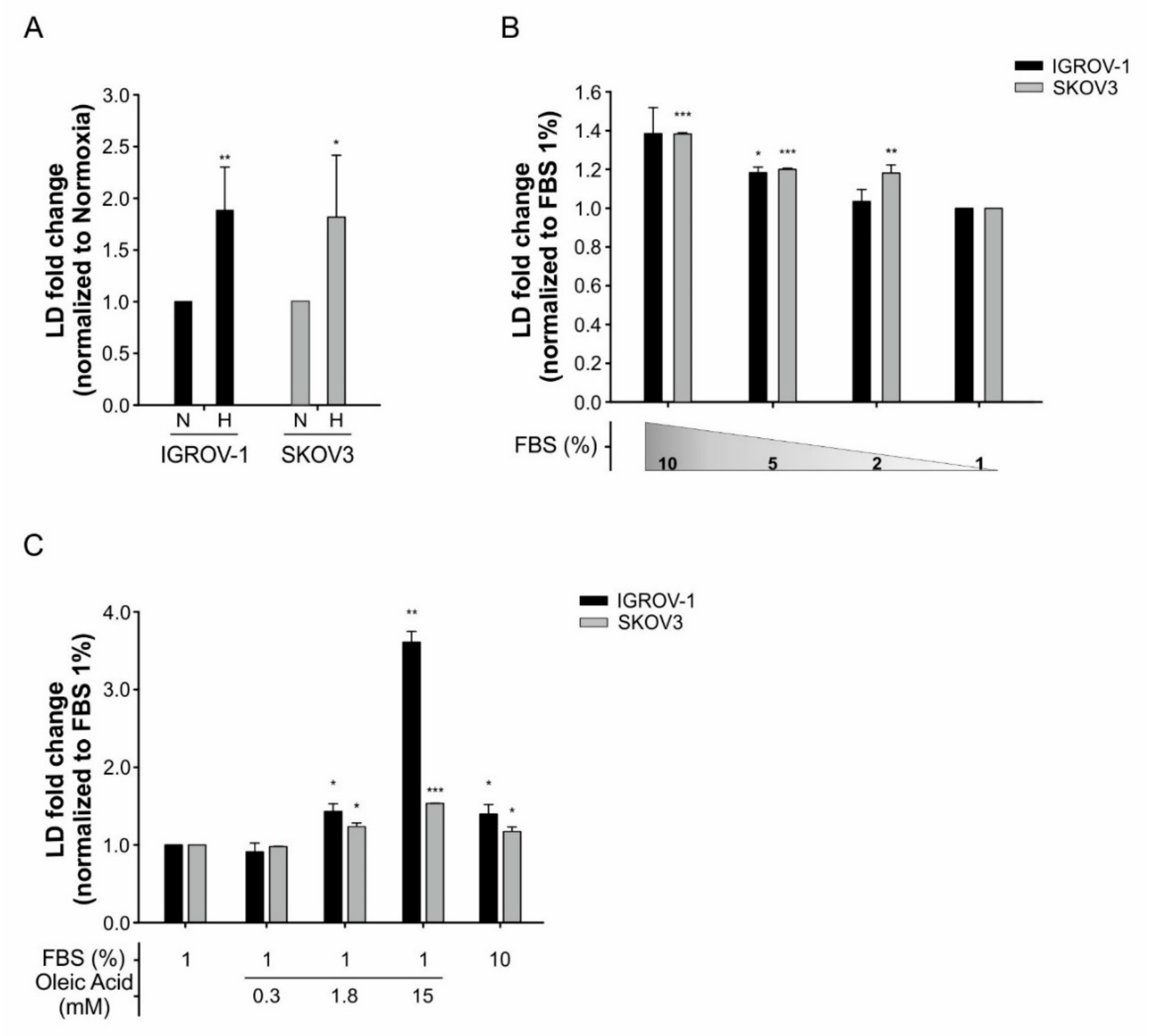
3.3. Hypoxia Accounts for Increased LD Accumulation and Expression of Cancer Stem Cells (CSCs) Markers in Tumor Cells
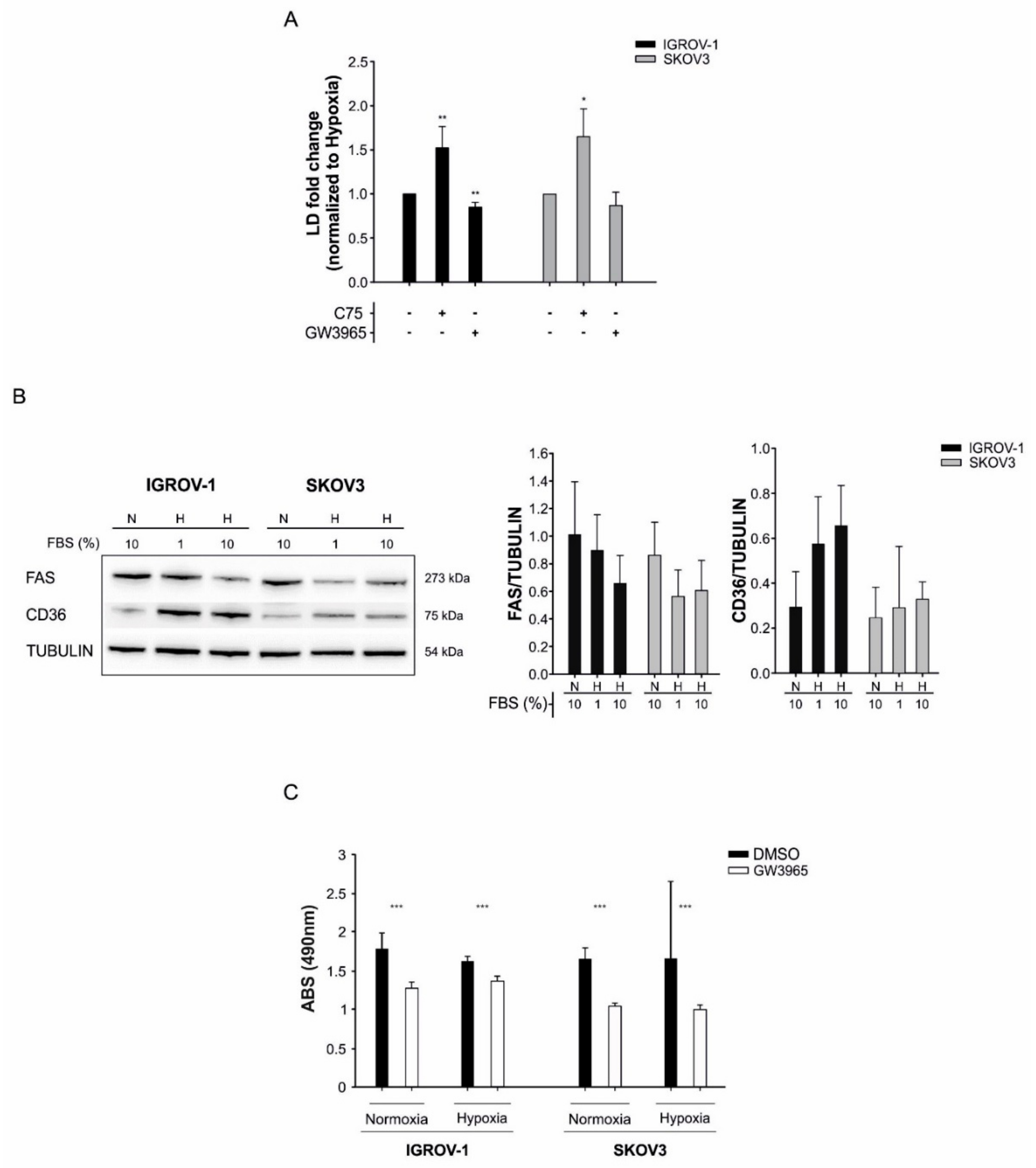
3.4. LD Accumulation in Tumor Cells is Counteracted by the LXR-Agonist GW3965
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hu, J.; Locasale, J.W.; Bielas, J.H.; O’Sullivan, J.; Sheahan, K.; Cantley, L.C.; Vander Heiden, M.G.; Vitkup, D. Heterogeneity of tumor-induced gene expression changes in the human metabolic network. Nat. Biotechnol. 2013, 31, 522–529. [Google Scholar] [CrossRef] [PubMed]
- Morandi, A.; Indraccolo, S. Linking metabolic reprogramming to therapy resistance in cancer. Biochim. Biophys. Acta 2017, 1868, 1–6. [Google Scholar] [CrossRef] [PubMed]
- McIntyre, A.; Harris, A.L. Metabolic and hypoxic adaptation to anti-angiogenic therapy: A target for induced essentiality. EMBO Mol. Med. 2015, 7, 368–379. [Google Scholar] [CrossRef] [PubMed]
- Pignata, S.; C-Cecere, S.; Du Bois, A.; Harter, P.; Heitz, F. Treatment of recurrent ovarian cancer. Ann. Oncol. 2017, 28, viii51–viii56. [Google Scholar] [CrossRef]
- Rossi, L.; Verrico, M.; Zaccarelli, E.; Papa, A.; Colonna, M.; Strudel, M.; Vici, P.; Bianco, V.; Tomao, F. Bevacizumab in ovarian cancer: A critical review of phase III studies. Oncotarget 2017, 8, 12389–12405. [Google Scholar] [CrossRef]
- Curtarello, M.; Zulato, E.; Nardo, G.; Valtorta, S.; Guzzo, G.; Rossi, E.; Esposito, G.; Msaki, A.; Pasto, A.; Rasola, A.; et al. VEGF-targeted therapy stably modulates the glycolytic phenotype of tumor cells. Cancer Res. 2015, 75, 120–133. [Google Scholar] [CrossRef]
- Nardo, G.; Favaro, E.; Curtarello, M.; Moserle, L.; Zulato, E.; Persano, L.; Rossi, E.; Esposito, G.; Crescenzi, M.; Casanovas, O.; et al. Glycolytic phenotype and AMP kinase modify the pathologic response of tumor xenografts to VEGF neutralization. Cancer Res. 2011, 71, 4214–4225. [Google Scholar] [CrossRef]
- Corbet, C.; Feron, O. Tumour acidosis: From the passenger to the driver’s seat. Nat. Rev. Cancer 2017, 17, 577–593. [Google Scholar] [CrossRef]
- Corbet, C.; Pinto, A.; Martherus, R.; Santiago de Jesus, J.P.; Polet, F.; Feron, O. Acidosis Drives the Reprogramming of Fatty Acid Metabolism in Cancer Cells through Changes in Mitochondrial and Histone Acetylation. Cell Metab. 2016, 24, 311–323. [Google Scholar] [CrossRef]
- Hattingen, E.; Jurcoane, A.; Bahr, O.; Rieger, J.; Magerkurth, J.; Anti, S.; Steinbach, J.P.; Pilatus, U. Bevacizumab impairs oxidative energy metabolism and shows antitumoral effects in recurrent glioblastomas: A 31P/1H MRSI and quantitative magnetic resonance imaging study. Neuro Oncol. 2011, 13, 1349–1363. [Google Scholar] [CrossRef]
- Keunen, O.; Johansson, M.; Oudin, A.; Sanzey, M.; Rahim, S.A.; Fack, F.; Thorsen, F.; Taxt, T.; Bartos, M.; Jirik, R.; et al. Anti-VEGF treatment reduces blood supply and increases tumor cell invasion in glioblastoma. Proc. Natl. Acad. Sci. USA 2011, 108, 3749–3754. [Google Scholar] [CrossRef]
- Kumar, K.; Wigfield, S.; Gee, H.E.; Devlin, C.M.; Singleton, D.; Li, J.L.; Buffa, F.; Huffman, M.; Sinn, A.L.; Silver, J.; et al. Dichloroacetate reverses the hypoxic adaptation to bevacizumab and enhances its antitumor effects in mouse xenografts. J. Mol. Med. (Berl.) 2013, 91, 749–758. [Google Scholar] [CrossRef]
- Menendez, J.A.; Lupu, R. Fatty acid synthase and the lipogenic phenotype in cancer pathogenesis. Nat. Rev. Cancer 2007, 7, 763–777. [Google Scholar] [CrossRef]
- Ackerman, D.; Simon, M.C. Hypoxia, lipids, and cancer: Surviving the harsh tumor microenvironment. Trends Cell Biol. 2014, 24, 472–478. [Google Scholar] [CrossRef]
- Zaidi, N.; Lupien, L.; Kuemmerle, N.B.; Kinlaw, W.B.; Swinnen, J.V.; Smans, K. Lipogenesis and lipolysis: The pathways exploited by the cancer cells to acquire fatty acids. Prog. Lipid Res. 2013, 52, 585–589. [Google Scholar] [CrossRef] [PubMed]
- Bensaad, K.; Favaro, E.; Lewis, C.A.; Peck, B.; Lord, S.; Collins, J.M.; Pinnick, K.E.; Wigfield, S.; Buffa, F.M.; Li, J.L.; et al. Fatty acid uptake and lipid storage induced by HIF-1alpha contribute to cell growth and survival after hypoxia-reoxygenation. Cell Rep. 2014, 9, 349–365. [Google Scholar] [CrossRef] [PubMed]
- Sounni, N.E.; Cimino, J.; Blacher, S.; Primac, I.; Truong, A.; Mazzucchelli, G.; Paye, A.; Calligaris, D.; Debois, D.; De Tullio, P.; et al. Blocking lipid synthesis overcomes tumor regrowth and metastasis after antiangiogenic therapy withdrawal. Cell Metab. 2014, 20, 280–294. [Google Scholar] [CrossRef]
- Jayson, G.C.; Kerbel, R.; Ellis, L.M.; Harris, A.L. Antiangiogenic therapy in oncology: Current status and future directions. Lancet 2016, 388, 518–529. [Google Scholar] [CrossRef]
- Ye, W. The Complexity of Translating Anti-angiogenesis Therapy from Basic Science to the Clinic. Dev. Cell 2016, 37, 114–125. [Google Scholar] [CrossRef]
- Favaro, E.; Nardo, G.; Persano, L.; Masiero, M.; Moserle, L.; Zamarchi, R.; Rossi, E.; Esposito, G.; Plebani, M.; Sattler, U.; et al. Hypoxia inducible factor-1alpha inactivation unveils a link between tumor cell metabolism and hypoxia-induced cell death. Am. J. Pathol. 2008, 173, 1186–1201. [Google Scholar] [CrossRef]
- Zulato, E.; Bergamo, F.; De Paoli, A.; Griguolo, G.; Esposito, G.; De Salvo, G.L.; Mescoli, C.; Rugge, M.; Nardin, M.; Di Grazia, L.; et al. Prognostic significance of AMPK activation in advanced stage colorectal cancer treated with chemotherapy plus bevacizumab. Br. J. Cancer 2014, 111, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Bligh, E.G.; Dyer, W.J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 1959, 37, 911–917. [Google Scholar] [CrossRef] [PubMed]
- Beckonert, O.; Keun, H.C.; Ebbels, T.M.; Bundy, J.; Holmes, E.; Lindon, J.C.; Nicholson, J.K. Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat. Protoc. 2007, 2, 2692–2703. [Google Scholar] [CrossRef] [PubMed]
- Canese, R.; Palombelli, G.; Chirico, M.; Sestili, P.; Bagnoli, M.; Canevari, S.; Mezzanzanica, D.; Podo, F.; Iorio, E. Integration of MRI and MRS approaches to monitor molecular imaging and metabolomic effects of trabectedin on a preclinical ovarian cancer model. NMR Biomed. 2018, 32, e4016. [Google Scholar] [CrossRef]
- Iorio, E.; Ricci, A.; Bagnoli, M.; Pisanu, M.E.; Castellano, G.; Di Vito, M.; Venturini, E.; Glunde, K.; Bhujwalla, Z.M.; Mezzanzanica, D.; et al. Activation of phosphatidylcholine cycle enzymes in human epithelial ovarian cancer cells. Cancer Res. 2010, 70, 2126–2135. [Google Scholar] [CrossRef]
- Gentleman, R.C.; Carey, V.J.; Bates, D.M.; Bolstad, B.; Dettling, M.; Dudoit, S.; Ellis, B.; Gautier, L.; Ge, Y.; Gentry, J.; et al. Bioconductor: Open software development for computational biology and bioinformatics. Genome Biol. 2004, 5, R80. [Google Scholar] [CrossRef]
- Irizarry, R.A.; Bolstad, B.M.; Collin, F.; Cope, L.M.; Hobbs, B.; Speed, T.P. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 2003, 31, e15. [Google Scholar] [CrossRef]
- Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3. [Google Scholar] [CrossRef]
- Ramakrishnan, S.; Subramanian, I.V.; Yokoyama, Y.; Geller, M. Angiogenesis in normal and neoplastic ovaries. Angiogenesis 2005, 8, 169–182. [Google Scholar] [CrossRef]
- Koizume, S.; Miyagi, Y. Lipid Droplets: A Key Cellular Organelle Associated with Cancer Cell Survival under Normoxia and Hypoxia. Int. J. Mol. Sci. 2016, 17. [Google Scholar] [CrossRef]
- Li, J.; Condello, S.; Thomes-Pepin, J.; Ma, X.; Xia, Y.; Hurley, T.D.; Matei, D.; Cheng, J.X. Lipid Desaturation Is a Metabolic Marker and Therapeutic Target of Ovarian Cancer Stem Cells. Cell Stem Cell 2017, 20, 303–314. [Google Scholar] [CrossRef] [PubMed]
- Lo Re, O.; Douet, J.; Buschbeck, M.; Fusilli, C.; Pazienza, V.; Panebianco, C.; Castracani, C.C.; Mazza, T.; Li Volti, G.; Vinciguerra, M. Histone variant macroH2A1 rewires carbohydrate and lipid metabolism of hepatocellular carcinoma cells towards cancer stem cells. Epigenetics 2018, 13, 829–845. [Google Scholar] [CrossRef] [PubMed]
- Pasto, A.; Bellio, C.; Pilotto, G.; Ciminale, V.; Silic-Benussi, M.; Guzzo, G.; Rasola, A.; Frasson, C.; Nardo, G.; Zulato, E.; et al. Cancer stem cells from epithelial ovarian cancer patients privilege oxidative phosphorylation, and resist glucose deprivation. Oncotarget 2014, 5, 4305–4319. [Google Scholar] [CrossRef] [PubMed]
- Lo Sasso, G.; Bovenga, F.; Murzilli, S.; Salvatore, L.; Di Tullio, G.; Martelli, N.; D’Orazio, A.; Rainaldi, S.; Vacca, M.; Mangia, A.; et al. Liver X receptors inhibit proliferation of human colorectal cancer cells and growth of intestinal tumors in mice. Gastroenterology 2013, 144, 1497–1507. [Google Scholar] [CrossRef]
- Bovenga, F.; Sabba, C.; Moschetta, A. Uncoupling nuclear receptor LXR and cholesterol metabolism in cancer. Cell Metab. 2015, 21, 517–526. [Google Scholar] [CrossRef]
- Fang, H.; He, J.; Ran, T.; Chen, H.; Jin, W.; Tang, B.; Hong, Z.; Fang, M. Synthesis, biological activities, and docking studies of d-pantolactone derivatives as novel FAS inhibitors. Bioorg. Med. Chem. 2019, 27, 115069. [Google Scholar] [CrossRef]
- Scoles, D.R.; Xu, X.; Wang, H.; Tran, H.; Taylor-Harding, B.; Li, A.; Karlan, B.Y. Liver X receptor agonist inhibits proliferation of ovarian carcinoma cells stimulated by oxidized low density lipoprotein. Gynecol. Oncol. 2010, 116, 109–116. [Google Scholar] [CrossRef]
- Schug, Z.T.; Peck, B.; Jones, D.T.; Zhang, Q.; Grosskurth, S.; Alam, I.S.; Goodwin, L.M.; Smethurst, E.; Mason, S.; Blyth, K.; et al. Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell 2015, 27, 57–71. [Google Scholar] [CrossRef]
- Garcia, D.; Shaw, R.J. AMPK: Mechanisms of Cellular Energy Sensing and Restoration of Metabolic Balance. Mol. Cell 2017, 66, 789–800. [Google Scholar] [CrossRef]
- Du, W.; Zhang, L.; Brett-Morris, A.; Aguila, B.; Kerner, J.; Hoppel, C.L.; Puchowicz, M.; Serra, D.; Herrero, L.; Rini, B.I.; et al. HIF drives lipid deposition and cancer in ccRCC via repression of fatty acid metabolism. Nat. Commun. 2017, 8, 1769. [Google Scholar] [CrossRef]
- Mayers, J.R.; Vander Heiden, M.G. Famine versus feast: Understanding the metabolism of tumors in vivo. Trends Biochem. Sci. 2015, 40, 130–140. [Google Scholar] [CrossRef]
- Boren, J.; Brindle, K.M. Apoptosis-induced mitochondrial dysfunction causes cytoplasmic lipid droplet formation. Cell Death Differ. 2012, 19, 1561–1570. [Google Scholar] [CrossRef] [PubMed]
- Penno, A.; Hackenbroich, G.; Thiele, C. Phospholipids and lipid droplets. Biochim. Biophys. Acta 2013, 1831, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Qiu, B.; Ackerman, D.; Sanchez, D.J.; Li, B.; Ochocki, J.D.; Grazioli, A.; Bobrovnikova-Marjon, E.; Diehl, J.A.; Keith, B.; Simon, M.C. HIF2alpha-Dependent Lipid Storage Promotes Endoplasmic Reticulum Homeostasis in Clear-Cell Renal Cell Carcinoma. Cancer Discov. 2015, 5, 652–667. [Google Scholar] [CrossRef]
- Cotte, A.K.; Aires, V.; Fredon, M.; Limagne, E.; Derangere, V.; Thibaudin, M.; Humblin, E.; Scagliarini, A.; de Barros, J.P.; Hillon, P.; et al. Lysophosphatidylcholine acyltransferase 2-mediated lipid droplet production supports colorectal cancer chemoresistance. Nat. Commun. 2018, 9, 322. [Google Scholar] [CrossRef]
- Cao, Y. Adipocyte and lipid metabolism in cancer drug resistance. J. Clin. Invest. 2019, 129, 3006–3017. [Google Scholar] [CrossRef]
- Fessler, M.B. The challenges and promise of targeting the Liver X Receptors for treatment of inflammatory disease. Pharmacol. Ther. 2018, 181, 1–12. [Google Scholar] [CrossRef]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Curtarello, M.; Tognon, M.; Venturoli, C.; Silic-Benussi, M.; Grassi, A.; Verza, M.; Minuzzo, S.; Pinazza, M.; Brillo, V.; Tosi, G.; et al. Rewiring of Lipid Metabolism and Storage in Ovarian Cancer Cells after Anti-VEGF Therapy. Cells 2019, 8, 1601. https://doi.org/10.3390/cells8121601
Curtarello M, Tognon M, Venturoli C, Silic-Benussi M, Grassi A, Verza M, Minuzzo S, Pinazza M, Brillo V, Tosi G, et al. Rewiring of Lipid Metabolism and Storage in Ovarian Cancer Cells after Anti-VEGF Therapy. Cells. 2019; 8(12):1601. https://doi.org/10.3390/cells8121601
Chicago/Turabian StyleCurtarello, Matteo, Martina Tognon, Carolina Venturoli, Micol Silic-Benussi, Angela Grassi, Martina Verza, Sonia Minuzzo, Marica Pinazza, Valentina Brillo, Giovanni Tosi, and et al. 2019. "Rewiring of Lipid Metabolism and Storage in Ovarian Cancer Cells after Anti-VEGF Therapy" Cells 8, no. 12: 1601. https://doi.org/10.3390/cells8121601
APA StyleCurtarello, M., Tognon, M., Venturoli, C., Silic-Benussi, M., Grassi, A., Verza, M., Minuzzo, S., Pinazza, M., Brillo, V., Tosi, G., Ferrazza, R., Guella, G., Iorio, E., Godfroid, A., Sounni, N. E., Amadori, A., & Indraccolo, S. (2019). Rewiring of Lipid Metabolism and Storage in Ovarian Cancer Cells after Anti-VEGF Therapy. Cells, 8(12), 1601. https://doi.org/10.3390/cells8121601





