Subcutaneous Maturation of Neural Stem Cell-Loaded Hydrogels Forms Region-Specific Neuroepithelium
Abstract
:1. Introduction
2. Materials and Methods
2.1. Methacrylamide Chitosan (MAC) Preparation
2.2. MAC-DBCO Preparation for Azide-Tagged IFN-γ (azIFN-γ) Immobilization
2.3. Chitosan Conduit Synthesis
2.4. Photoreactive Laminin
2.5. aNSC Harvest and Culture
2.6. NSC Encapsulation in MAC Hydrogels
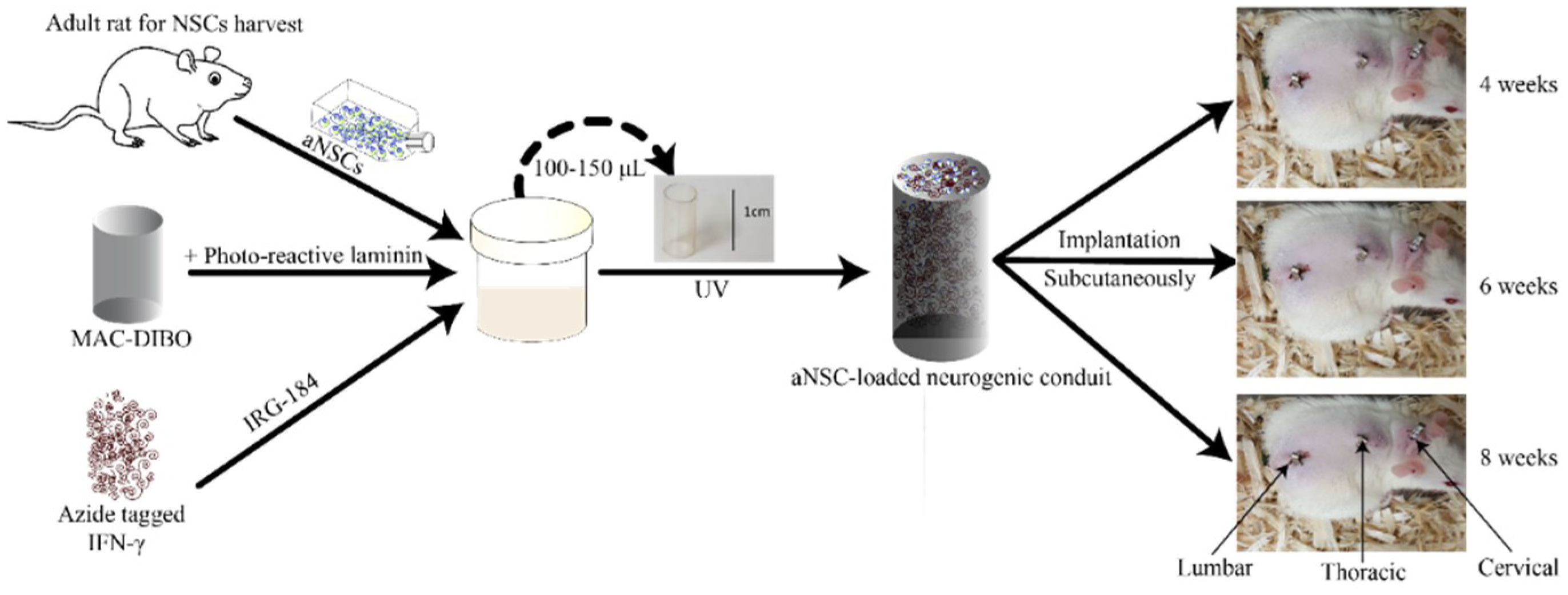
2.7. Implantation of the aNSC-MAC System Subcutaneously
2.8. Immunohistochemistry (IHC)
2.9. RT-qPCR
2.9.1. Absolute Quantification Analysis of Target Genes
2.9.2. Relative Expression qPCR Assays
2.10. Statistics
3. Results
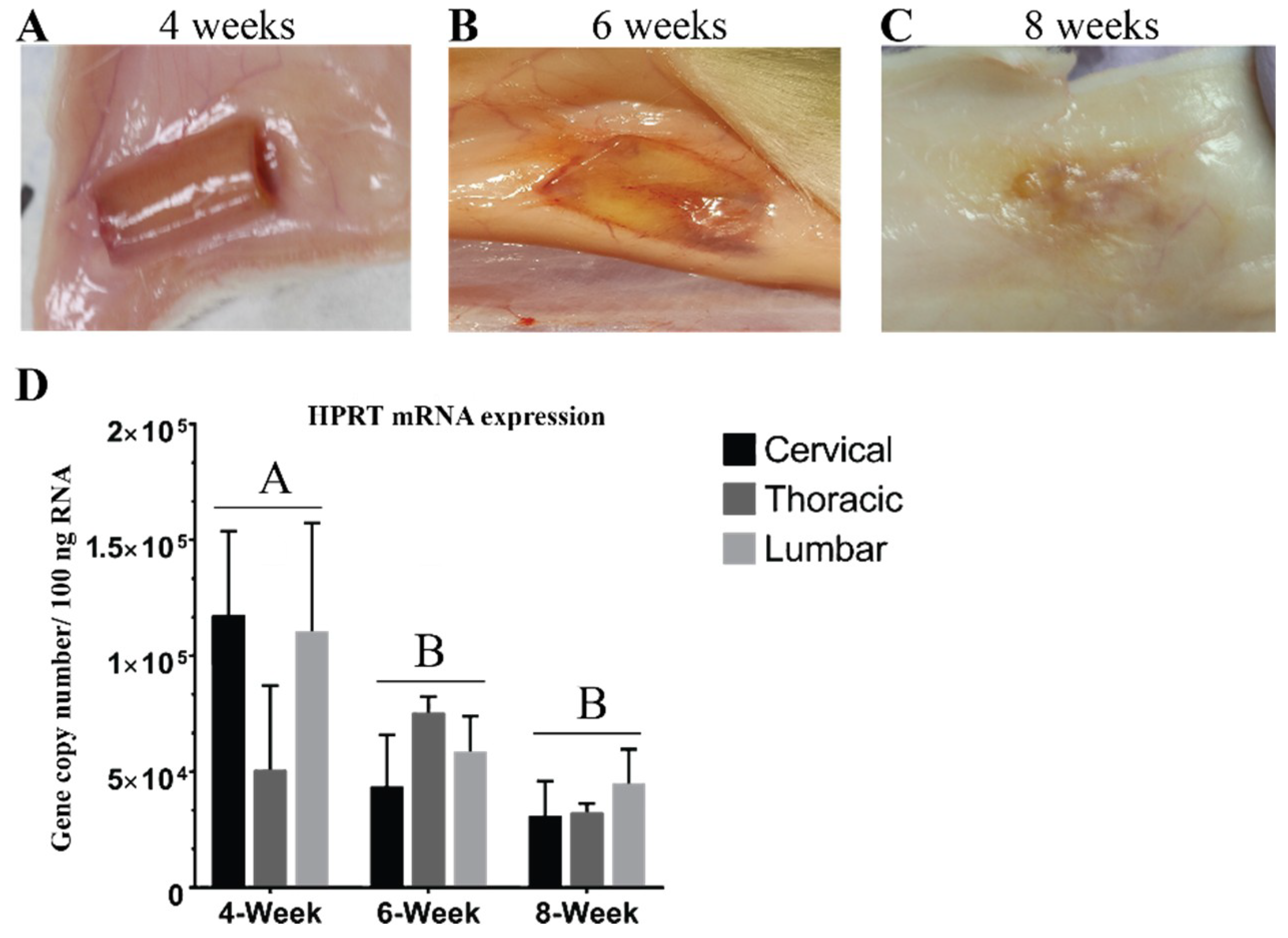
3.1. Hydrogel Degradation Corresponds to Loss in Gene Expression
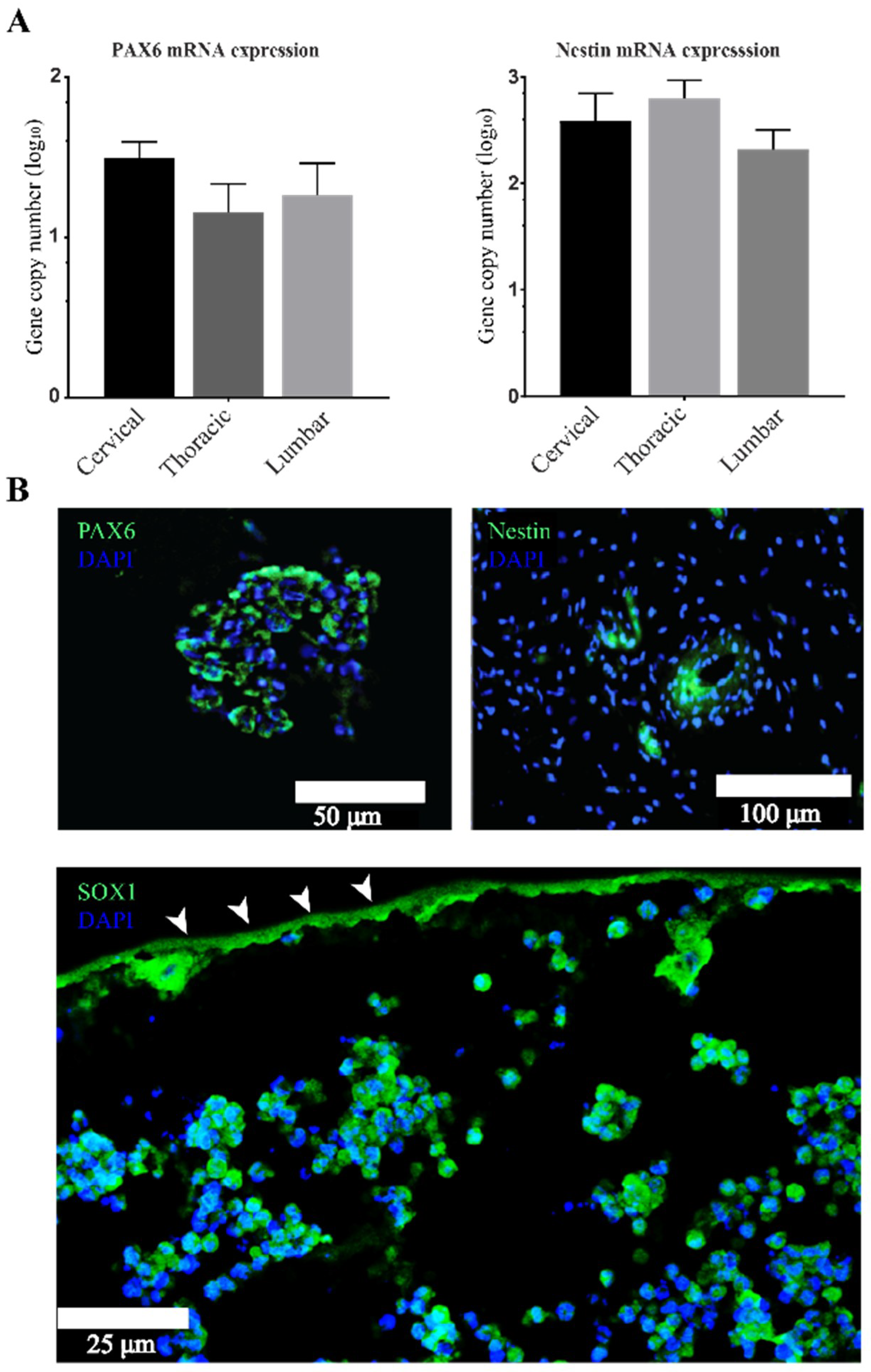
3.2. Stemness Expression Profiles
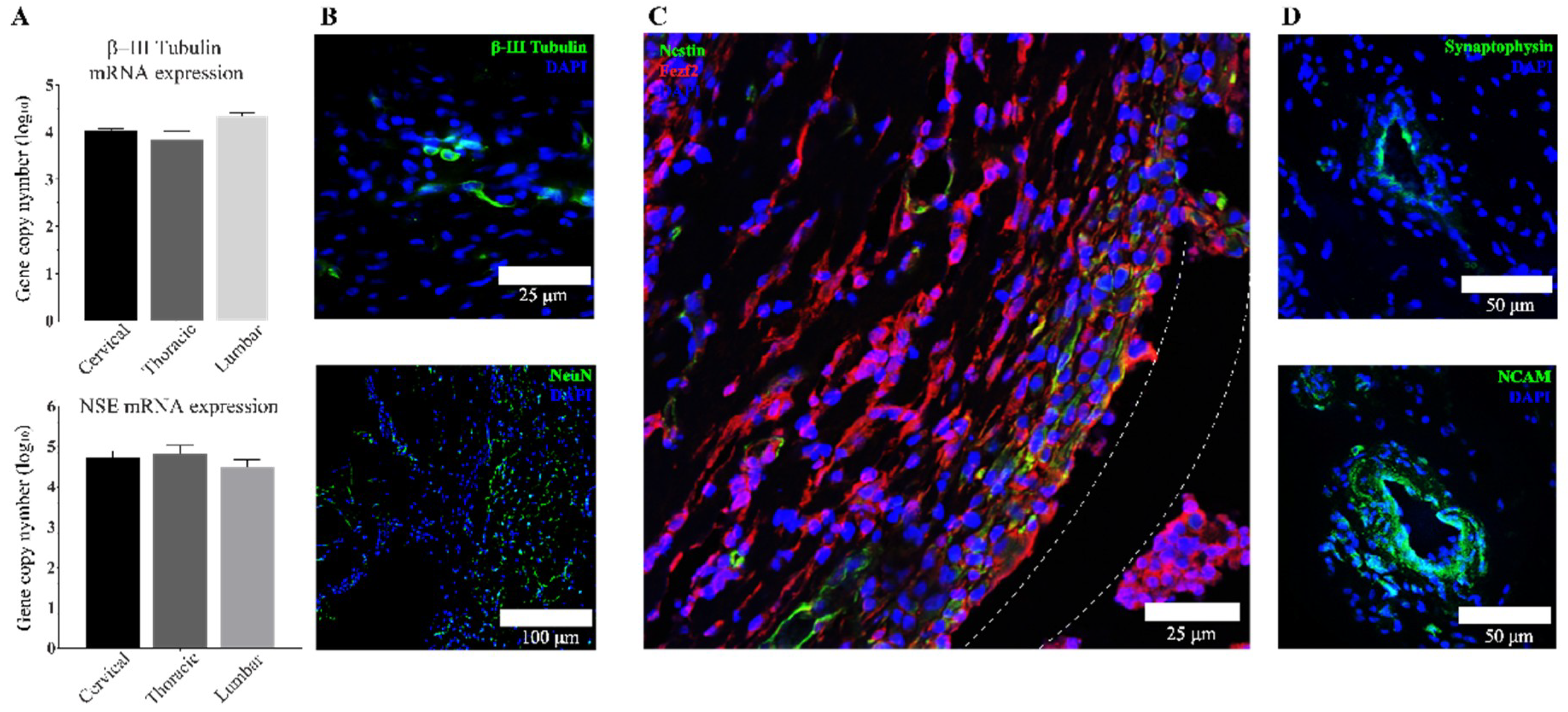
3.3. Transplanted aNSCs Express Neuronal Markers
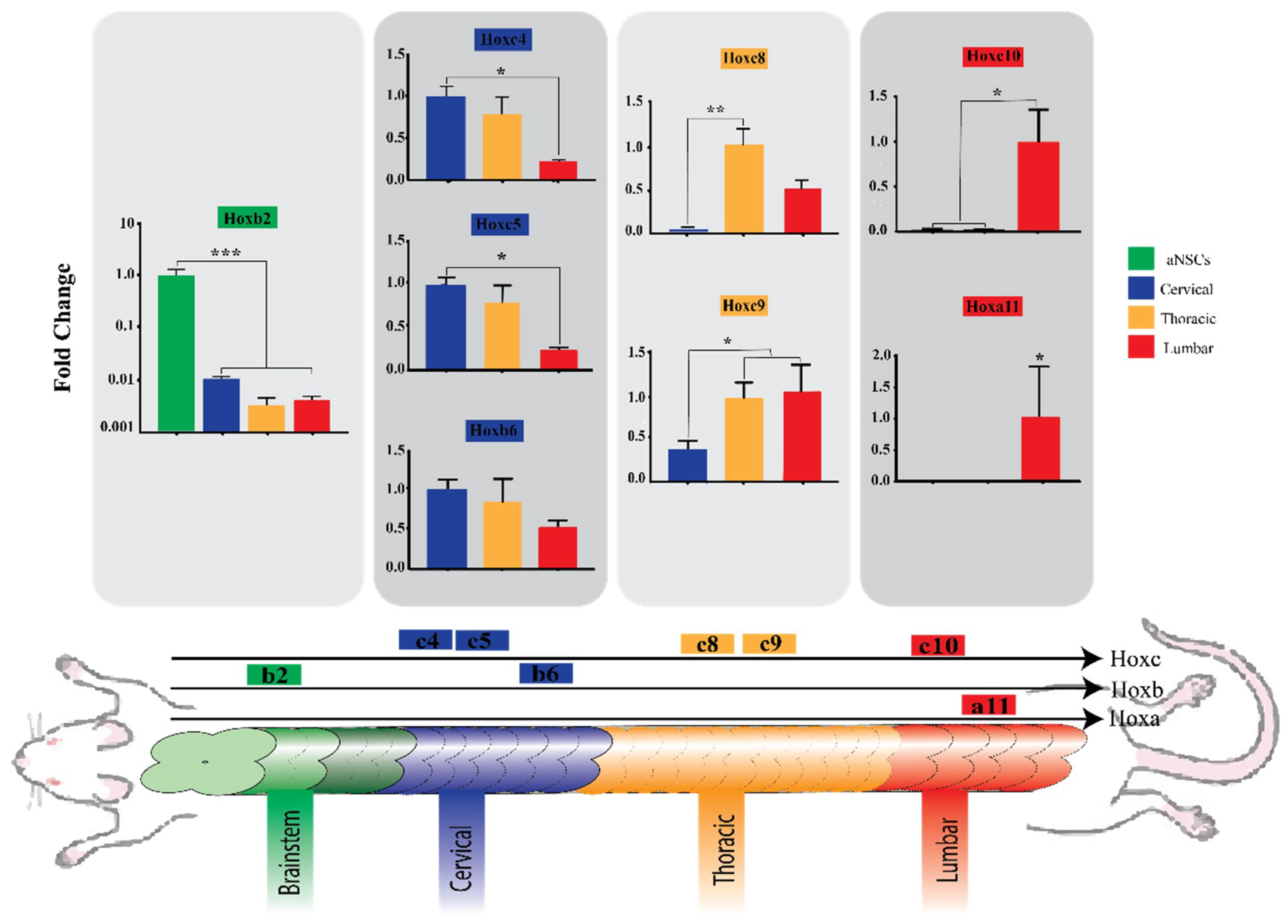
3.4. Region-Specific Differentiation of aNSCs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ahuja, C.S.; Fehlings, M. Concise Review: Bridging the gap: Novel neuroregenerative and neuroprotective strategies in spinal cord injury. Stem Cells Transl. Med. 2016, 5, 914–924. [Google Scholar] [CrossRef] [PubMed]
- Ahuja, C.S.; Martin, A.R.; Fehlings, M. Recent advances in managing a spinal cord injury secondary to trauma. F1000Research 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Adami, R.; Scesa, G.; Bottai, D. Stem cell transplantation in neurological diseases: Improving effectiveness in animal models. Front. Cell Dev. Biol 2014, 2, 2183–2200. [Google Scholar] [CrossRef] [PubMed]
- Okano, H. Neural stem cells and strategies for the regeneration of the central nervous system. Proc. Jpn. Acad. Ser. B 2010, 86, 438–450. [Google Scholar] [CrossRef] [Green Version]
- Leipzig, N.D.; Xu, C.; Zahir, T.; Shoichet, M.S. Functional immobilization of interferon-gamma induces neuronal differentiation of neural stem cells. J. Biomed. Mater. Res. A 2010, 93, 625–633. [Google Scholar] [CrossRef] [PubMed]
- Leipzig, N.D.; Shoichet, M.S. The effect of substrate stiffness on adult neural stem cell behavior. Biomaterials 2009, 30, 6867–6878. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Koenig, A.M.; Sloan, P.; Leipzig, N.D. In vivo assessment of guided neural stem cell differentiation in growth factor immobilized chitosan-based hydrogel scaffolds. Biomaterials 2014, 35, 9049–9057. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wijekoon, A.; Leipzig, N.D. 3D differentiation of neural stem cells in macroporous photopolymerizable hydrogel scaffolds. PLoS ONE 2012, 7, e48824. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Ham, T.R.; Neill, N.; Farrag, M.; Mohrman, A.E.; Koenig, A.M.; Leipzig, N.D. A hydrogel bridge incorporating immobilized growth factors and neural stem/progenitor cells to treat spinal cord injury. Adv. Healthc. Mater. 2016, 5, 802–812. [Google Scholar] [CrossRef] [PubMed]
- Ham, T.R.; Farrag, M.; Leipzig, N.D. Covalent growth factor tethering to direct neural stem cell differentiation and self-organization. Acta Biomater. 2017, 53, 140–151. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pluchino, S.; Zanotti, L.; Brini, E.; Ferrari, S.; Martino, G. Regeneration and repair in multiple sclerosis: The role of cell transplantation. Neurosci. Lett. 2009, 456, 101–106. [Google Scholar] [CrossRef] [PubMed]
- Pluchino, S.; Quattrini, A.; Brambilla, E.; Gritti, A.; Salani, G.; Dina, G.; Galli, R.; Del Carro, U.; Amadio, S.; Bergami, A.; et al. Injection of adult neurospheres induces recovery in a chronic model of multiple sclerosis. Nature 2003, 422, 688–694. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Z.; Zhu, W.; Cao, K.; Wu, F.; Li, J.; Wang, G.; Li, H.; Lu, M.; Ren, Y.; He, X. Anti-inflammatory mechanism of neural stem cell transplantation in spinal cord injury. Int. J. Mol. Sci. 2016, 17, 1380. [Google Scholar] [CrossRef] [PubMed]
- Xiong, L.-L.; Zou, Y.; Shi, Y.; Zhang, P.; Zhang, R.-P.; Dai, X.-J.; Liu, B.; Wang, T.H. Tree shrew neural stem cell transplantation promotes functional recovery of tree shrews with a hemi-sectioned spinal cord injury by upregulating nerve growth factor expression. Int. J. Mol. Med. 2018, 41, 3267–3277. [Google Scholar] [CrossRef] [PubMed]
- Lu, P.; Jones, L.L.; Snyder, E.Y.; Tuszynski, M.H. Neural stem cells constitutively secrete neurotrophic factors and promote extensive host axonal growth after spinal cord injury. Exp. Neurol. 2003, 181, 115–129. [Google Scholar] [CrossRef]
- Cusimano, M.; Biziato, D.; Brambilla, E.; Donegà, M.; Alfaro-Cervello, C.; Snider, S.; Salani, G.; Pucci, F.; Comi, G.; Garcia-Verdugo, J.M.; et al. Transplanted neural stem/precursor cells instruct phagocytes and reduce secondary tissue damage in the injured spinal cord. Brain 2012, 135, 447–460. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Assinck, P.; Duncan, G.J.; Hilton, B.J.; Plemel, J.R.; Tetzlaff, W. Cell transplantation therapy for spinal cord injury. Nat. Neurosci. 2017, 20, 637–647. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Shi, J.; van Ginkel, F.W.; Lan, L.; Niemeyer, G.; Martin, D.R.; Snyder, E.Y.; Cox, N.R. Neural stem/progenitor cells modulate immune responses by suppressing T lymphocytes with nitric oxide and prostaglandin E2. Exp. Neurol. 2009, 216, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Nam, H.; Lee, K.-H.; Nam, D.-H.; Joo, K.M. Adult human neural stem cell therapeutics: Current developmental status and prospect. World J. Stem Cells 2015, 7, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Crook, J.M.; Tomaskovic-Crook, E. Culturing and Cryobanking Human Neural Stem Cells. Methods Mol. Biol. 2017, 1590, 199–206. [Google Scholar] [PubMed]
- Elkabetz, Y.; Panagiotakos, G.; Al Shamy, G.; Socci, N.D.; Tabar, V.; Studer, L. Human ES cell-derived neural rosettes reveal a functionally distinct early neural stem cell stage. Genes Dev. 2008, 22, 152–165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cardozo, M.J.; Mysiak, K.S.; Becker, T.; Becker, C.G. Reduce, reuse, recycle—Developmental signals in spinal cord regeneration. Dev. Biol. 2017, 432, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Hayakawa, K.; Okazaki, R.; Morioka, K.; Nakamura, K.; Tanaka, S.; Ogata, T. Lipopolysaccharide preconditioning facilitates M2 activation of resident microglia after spinal cord injury. J. Neurosci. Res. 2014, 92, 1647–1658. [Google Scholar] [CrossRef] [PubMed]
- Saxena, S.; Shukla, D.; Bansal, A. Augmentation of aerobic respiration and mitochondrial biogenesis in skeletal muscle by hypoxia preconditioning with cobalt chloride. Toxicol. Appl. Pharmacol. 2012, 264, 324–334. [Google Scholar] [CrossRef] [PubMed]
- Shafiq, M.; Jung, Y.; Kim, S.H. Insight on stem cell preconditioning and instructive biomaterials to enhance cell adhesion, retention, and engraftment for tissue repair. Biomaterials 2016, 90, 85–115. [Google Scholar] [CrossRef] [PubMed]
- Pei, M. Environmental preconditioning rejuvenates adult stem cells’ proliferation and chondrogenic potential. Biomaterials 2017, 117, 10–23. [Google Scholar] [CrossRef] [PubMed]
- Sakata, H.; Niizuma, K.; Yoshioka, H.; Kim, G.S.; Jung, J.E.; Katsu, M.; Narasimhan, P.; Maier, C.M.; Nishiyama, Y.; Chan, P.H. Minocycline-preconditioned neural stem cells enhance neuroprotection after ischemic stroke in rats. J. Neurosci. 2012, 32, 3462–3473. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Yang, X.; Liu, T.; Shao, J.; Fu, N.; Yan, A.; Geng, K.; Xia, W. Adjudin-preconditioned neural stem cells enhance neuroprotection after ischemia reperfusion in mice. Stem Cell Res. Ther. 2017, 8, 248. [Google Scholar] [CrossRef] [PubMed]
- Bernstock, J.D.; Peruzzotti-Jametti, L.; Ye, D.; Gessler, F.A.; Maric, D.; Vicario, N.; Lee, Y.J.; Pluchino, S.; Hallenbeck, J.M. Neural stem cell transplantation in ischemic stroke: A role for preconditioning and cellular engineering. J. Cereb. Blood Flow Metab. 2017, 37, 2314–2319. [Google Scholar] [CrossRef] [PubMed]
- Göritz, C.; Frisén, J. Neural Stem Cells and Neurogenesis in the Adult. Cell Stem Cell 2012, 10, 657–659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wiese, S.; Faissner, A. The role of extracellular matrix in spinal cord development. Exp. Neurol. 2015, 274, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, D.R.; Dours-Zimmermann, M.T. Extracellular matrix of the central nervous system: From neglect to challenge. Histochem. Cell Biol. 2008, 130, 635–653. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berry, D.C.; Stenesen, D.; Zeve, D.; Graff, J.M. The developmental origins of adipose tissue. Development 2013, 140, 3939–3949. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liedtke, S.; Buchheiser, A.; Bosch, J.; Bosse, F.; Kruse, F.; Zhao, X.; Santourlidis, S.; Kögler, G. The HOX Code as a “biological fingerprint” to distinguish functionally distinct stem cell populations derived from cord blood. Stem Cell Res. 2010, 5, 40–50. [Google Scholar] [CrossRef] [PubMed]
- Lippmann, E.S.; Williams, C.E.; Ruhl, D.A.; Estevez-Silva, M.C.; Chapman, E.R.; Coon, J.J.; Ashton, R.S. Deterministic HOX patterning in human pluripotent stem cell-derived neuroectoderm. Stem Cell Rep. 2015, 4, 632–644. [Google Scholar] [CrossRef] [PubMed]
- Nolte, C.; Krumlauf, R. Expression of Hox Genes in the Nervous System of Vertebrates; Landes Bioscience: Austin, TX, USA, 2013. [Google Scholar]
- Yamamoto, M.; Takai, D.; Yamamoto, F.; Yamamoto, F. Comprehensive expression profiling of highly homologous 39 hox genes in 26 different human adult tissues by the modified systematic multiplex rt-pcr method reveals tissue-specific expression pattern that suggests an important role of chromosomal structure. Gene Expr. 2003, 11, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Hejnol, A.; Martindale, M.Q. Coordinated spatial and temporal expression of Hox genes during embryogenesis in the acoel convolutriloba longifissura. BMC Biol. 2009, 7, 65. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lacombe, J.; Hanley, O.; Jung, H.; Philippidou, P.; Surmeli, G.; Grinstein, J.; Dasen, J.S. genetic and functional modularity of Hox activities in the specification of limb-innervating motor neurons. PLoS Genet. 2013, 9, e1003184. [Google Scholar] [CrossRef] [PubMed]
- Catela, C.; Shin, M.M.; Lee, D.H.; Liu, J.-P.; Dasen, J.S. Hox proteins coordinate motor neuron differentiation and connectivity programs through Ret/Gfrα genes. Cell Rep. 2016, 14, 1901–1915. [Google Scholar] [CrossRef] [PubMed]
- Seifert, A.; Werheid, D.F.; Knapp, S.M.; Tobiasch, E. Role of Hox genes in stem cell differentiation. World J. Stem Cells 2015, 7, 583–595. [Google Scholar] [CrossRef] [PubMed]
- Toshner, M.; Dunmore, B.J.; McKinney, E.F.; Southwood, M.; Caruso, P.; Upton, P.D.; Waters, J.P.; Ormiston, M.L.; Skepper, J.N.; Nash, G.; et al. Transcript analysis reveals a specific HOX signature associated with positional identity of human endothelial cells. PLoS ONE 2014, 9, e91334. [Google Scholar] [CrossRef] [PubMed]
- Molino, Y.; Jabès, F.; Bonnet, A.; Gaudin, N.; Bernard, A.; Benech, P.; Khrestchatisky, M. Gene expression comparison reveals distinct basal expression of HOX members and differential TNF-induced response between brain- and spinal cord-derived microvascular endothelial cells. J. Neuroinflamm. 2016, 13, 290. [Google Scholar] [CrossRef] [PubMed]
- Iyer, N.R.; Wilems, T.S.; Sakiyama-Elbert, S.E. Stem cells for spinal cord injury: Strategies to inform differentiation and transplantation. Biotechnol. Bioeng. 2017, 114, 245–259. [Google Scholar] [CrossRef] [PubMed]
- Raspa, A.; Pugliese, R.; Maleki, M.; Gelain, F. Recent therapeutic approaches for spinal cord injury. Biotechnol. Bioeng. 2016, 113, 253–259. [Google Scholar] [CrossRef] [PubMed]
- Agbay, A.; Edgar, J.M.; Robinson, M.; Styan, T.; Wilson, K.; Schroll, J.; Ko, J.; Khadem Mohtaram, N.; Jun, M.B.; Willerth, S.M. Biomaterial strategies for delivering stem cells as a treatment for spinal cord injury. Cells Tissues Organs 2016, 202, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Führmann, T.; Anandakumaran, P.N.; Shoichet, M.S. Combinatorial therapies after spinal cord injury: How can biomaterials help? Adv. Healthc. Mater. 2017, 6, 1601130. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Schackel, T.; Weidner, N.; Puttagunta, R. Biomaterial-supported cell transplantation treatments for spinal cord injury: Challenges and perspectives. Front. Cell Neurosci. 2018, 11, 430. [Google Scholar] [CrossRef] [PubMed]
- Sakiyama-Elbert, S.; Johnson, P.J.; Hodgetts, S.I.; Plant, G.W.; Harvey, A.R. Scaffolds to promote spinal cord regeneration. Handb. Clin. Neurol. 2012, 109, 575–594. [Google Scholar] [PubMed]
- Wong, F.; Chan, B.; Lo, A. Carriers in cell-based therapies for neurological disorders. Int. J. Mol. Sci. 2014, 15, 10669–10723. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Monteiro, S.; Roque, S.; Marques, F.; Correia-Neves, M.; Cerqueira, J.J. Brain interference: Revisiting the role of IFNγ in the central nervous system. Prog. Neurobiol. 2017, 156, 149–163. [Google Scholar] [CrossRef] [PubMed]
- Mondello, S.; Loane, D.J.; Clausen, F.; Roselli, F.; Chandrasekar, A.; Morganti-Kossmann, M.C. Interferons in traumatic brain and spinal cord injury: Current evidence for translational application. Front. Neurol. 2018, 9, 458. [Google Scholar]
- Nomura, H.; Zahir, T.; Kim, H.; Katayama, Y.; Kulbatski, I.; Morshead, C.M.; Shoichet, M.S.; Tator, C.H. Extramedullary chitosan channels promote survival of transplanted neural stem and progenitor cells and create a tissue bridge after complete spinal cord transection. Tissue Eng. Part A 2008, 14, 649–665. [Google Scholar] [CrossRef] [PubMed]
- Freier, T.; Montenegro, R.; Koh, H.S.; Shoichet, M.S. Chitin-based tubes for tissue engineering in the nervous system. Biomaterials 2005, 26, 4624–4632. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Tanapat, P. Neuronal Cell Markers. Mater. Methods 2013, 3, 196. [Google Scholar] [CrossRef]
- Kim, D.S.; Lee, D.R.; Kim, H.S.; Yoo, J.E.; Jung, S.J.; Lim, B.Y.; Jang, J.; Kang, H.C.; You, S.; Hwang, D.Y.; et al. Highly Pure and Expandable PSA-NCAM-Positive Neural Precursors from Human ESC and iPSC-Derived Neural Rosettes. PLoS ONE 2012, 7, e39715. [Google Scholar] [CrossRef] [PubMed]
- Kwon, S.E.; Chapman, E.R. Synaptophysin regulates the kinetics of synaptic vesicle endocytosis in central neurons. Neuron 2011, 70, 847–854. [Google Scholar] [CrossRef] [PubMed]
- Kadoya, K.; Lu, P.; Nguyen, K.; Lee-Kubli, C.; Kumamaru, H.; Yao, L.; Knackert, J.; Poplawski, G.; Dulin, J.N.; Strobl, H.; et al. Spinal cord reconstitution with homologous neural grafts enables robust corticospinal regeneration. Nat. Med. 2016, 22, 479–487. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Movassaghi, K.; Ver Halen, J.; Ganchi, P.; Amin-Hanjani, S.; Mesa, J.; Yaremchuk, M.J. Cranioplasty with subcutaneously preserved autologous bone grafts. Plast. Reconstr. Surg. 2006, 117, 202–206. [Google Scholar] [CrossRef] [PubMed]
- Shoakazemi, A.; Flannery, T.; McConnell, R.S. Long-term outcome of subcutaneously preserved autologous cranioplasty. Neurosurgery 2009, 65, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Morina, A.; Kelmendi, F.; Morina, Q.; Dragusha, S.; Ahmeti, F.; Morina, D.; Gashi, K. Cranioplasty with subcutaneously preserved autologous bone grafts in abdominal wall-Experience with 75 cases in a post-war country Kosova. Surg. Neurol. Int. 2011, 2, 72. [Google Scholar] [CrossRef] [PubMed]
- Amini, A.R.; Laurencin, C.T.; Nukavarapu, S.P. Bone tissue engineering: Recent advances and challenges. Crit. Rev. Biomed. Eng. 2012, 40, 363–408. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-T.; Kim, I.-S.; Lee, I.-S.; Lee, J.-P.; Snyder, E.Y.; In Park, K. Human neurospheres derived from the fetal central nervous system are regionally and temporally specified but are not committed. Exp. Neurol. 2006, 199, 222–235. [Google Scholar] [CrossRef] [PubMed]
- Stelnicki, E.J.; Kömüves, L.G.; Kwong, A.O.; Holmes, D.; Klein, P.; Rozenfeld, S.; Lawrence, H.J.; Adzick, N.S.; Harrison, M.; Largman, C. HOX homeobox genes exhibit spatial and temporal changes in expression during human skin development. J. Investig. Dermatol. 1998, 110, 110–115. [Google Scholar] [CrossRef] [PubMed]
- Philippidou, P.; Dasen, J. Hox genes: Choreographers in neural development, architects of circuit organization. Neuron 2013, 80, 12–34. [Google Scholar] [CrossRef] [PubMed]
- Wilson, P.G.; Stice, S.S. Development and differentiation of neural rosettes derived from human embryonic stem cells. Stem Cell Rev. 2006, 2, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Cristina Valensisi, A.; Andrus, C.; Buckberry, S.; Doni Jayavelu, N.; Lund, R.J.; Lister, R.; Hawkins, R.D. Epigenomic landscapes of hesc-derived neural rosettes: Modeling neural tube formation and diseases. Cell Rep. 2017, 20, 1448–1462. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.M.Y.; Kazazian, K.; Shoichet, M.S. Peptide surface modification of methacrylamide chitosan for neural tissue engineering applications. J. Biomed. Mater. Res. A 2007, 82, 243–255. [Google Scholar] [CrossRef] [PubMed]
- Bozkurt, G.; Mothe, A.J.; Zahir, T.; Kim, H.; Shoichet, M.S.; Tator, C.H. chitosan channels containing spinal cord-derived stem/progenitor cells for repair of subacute spinal cord injury in the rat. Neurosurgery 2010, 67, 1733–1744. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Farrag, M.; Leipzig, N.D. Subcutaneous Maturation of Neural Stem Cell-Loaded Hydrogels Forms Region-Specific Neuroepithelium. Cells 2018, 7, 173. https://doi.org/10.3390/cells7100173
Farrag M, Leipzig ND. Subcutaneous Maturation of Neural Stem Cell-Loaded Hydrogels Forms Region-Specific Neuroepithelium. Cells. 2018; 7(10):173. https://doi.org/10.3390/cells7100173
Chicago/Turabian StyleFarrag, Mahmoud, and Nic D. Leipzig. 2018. "Subcutaneous Maturation of Neural Stem Cell-Loaded Hydrogels Forms Region-Specific Neuroepithelium" Cells 7, no. 10: 173. https://doi.org/10.3390/cells7100173
APA StyleFarrag, M., & Leipzig, N. D. (2018). Subcutaneous Maturation of Neural Stem Cell-Loaded Hydrogels Forms Region-Specific Neuroepithelium. Cells, 7(10), 173. https://doi.org/10.3390/cells7100173




