Monte Carlo Study Elucidates the Type 1/Type 2 Choice in Apoptotic Death Signaling in Healthy and Cancer Cells
Abstract
1. Introduction
2. Experimental Section
2.1. The Signaling Model for Apoptotic Cell Death

2.2. Monte Carlo Simulation of Cell Death Signaling
2.3. Estimation of Parameter Values (as Used in Our Monte Carlo Simulation)
3. Results and Discussion
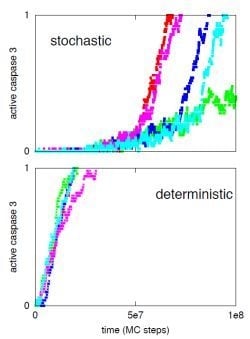
3.1. Increased Active Caspase 8 Concentration Switches the Activation from Stochastic Type 2 to Deterministic Type 1
3.2. Systems Level Regulatory Mechanisms of Type 1/Type 2 Choice
3.2.1. Effect of Small (~3-Fold) Bcl2 Variation on Cyto C Release and Type 1/Type 2 Choice
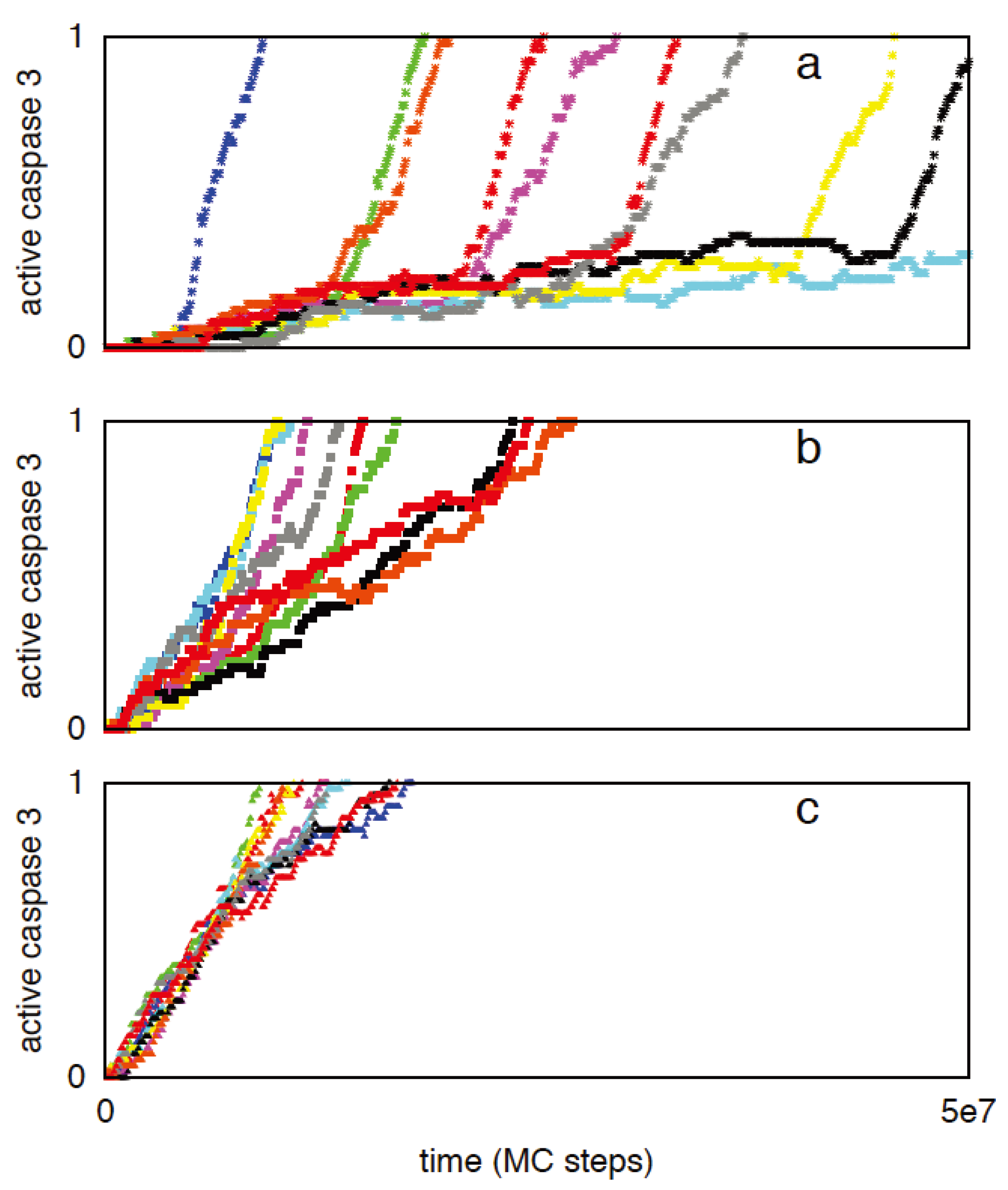
3.2.2. XIAP to Smac Ratio Is a Key Regulator of Type 1/Type 2 Choice: Inhibition of the Type 1 Activation by XIAP

| XIAP/Smac combinations | Average Caspase 3 activation (SD) | ||
|---|---|---|---|
| T = 2.5 × 107 (MC steps) | T = 5 × 107 (MC steps) | T = 1 × 108 (MC steps) | |
| XIAP = 30 nM Smac = 50 nM (ratio 0.6) | 0.65 (0.38) | 0.92 (0.20) | 1.0 (0.0) |
| XIAP = 30 nM Smac = 10 nM (ratio 3) | 0.47 (0.38) | 0.81 (0.32) | 0.98 (0.08) |
| XIAP = 90 nM Smac = 50 nM (ratio 1.8) | 0.01 (0.01) | 0.1 (0.08) | 1.0 (0.0) |
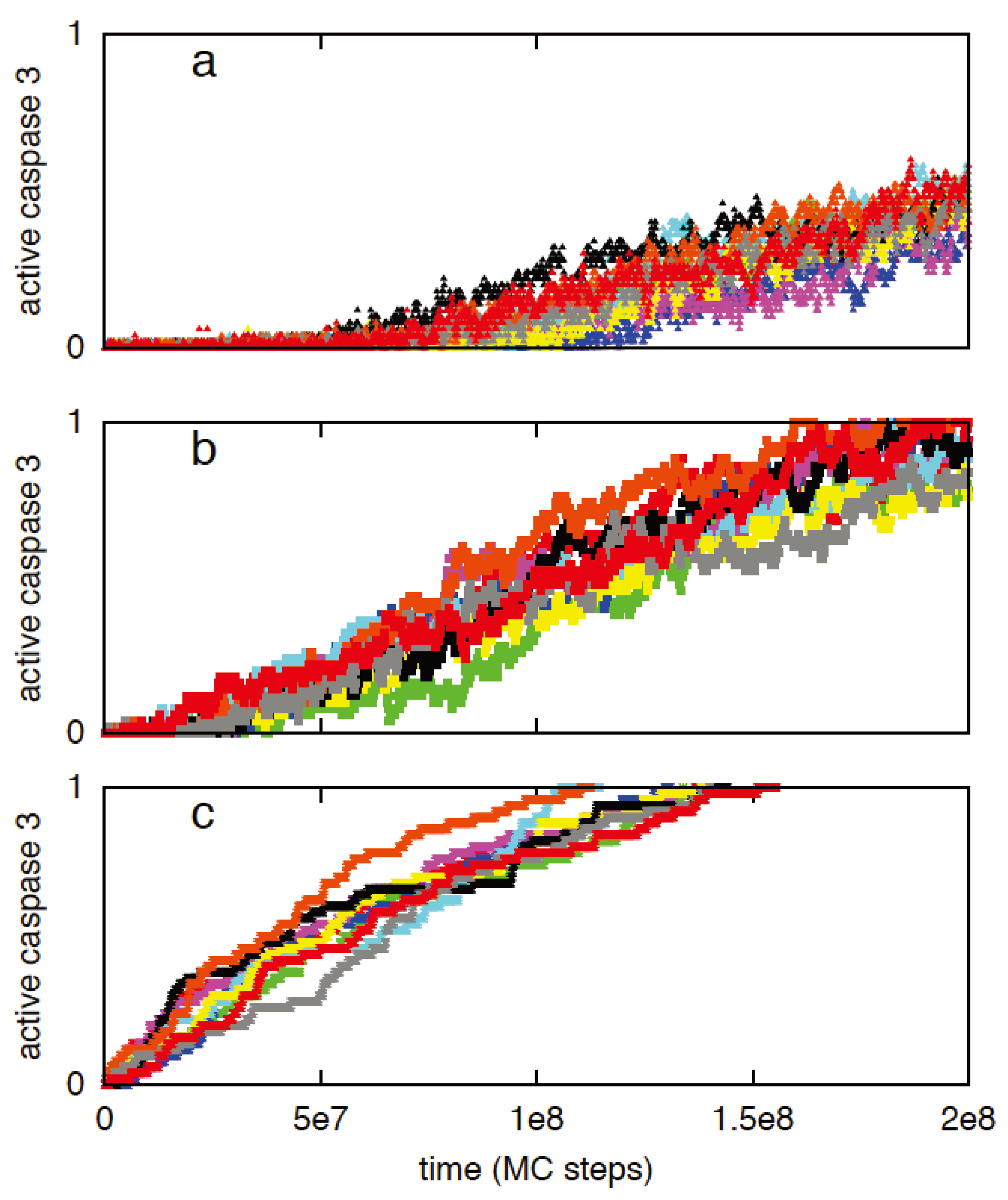
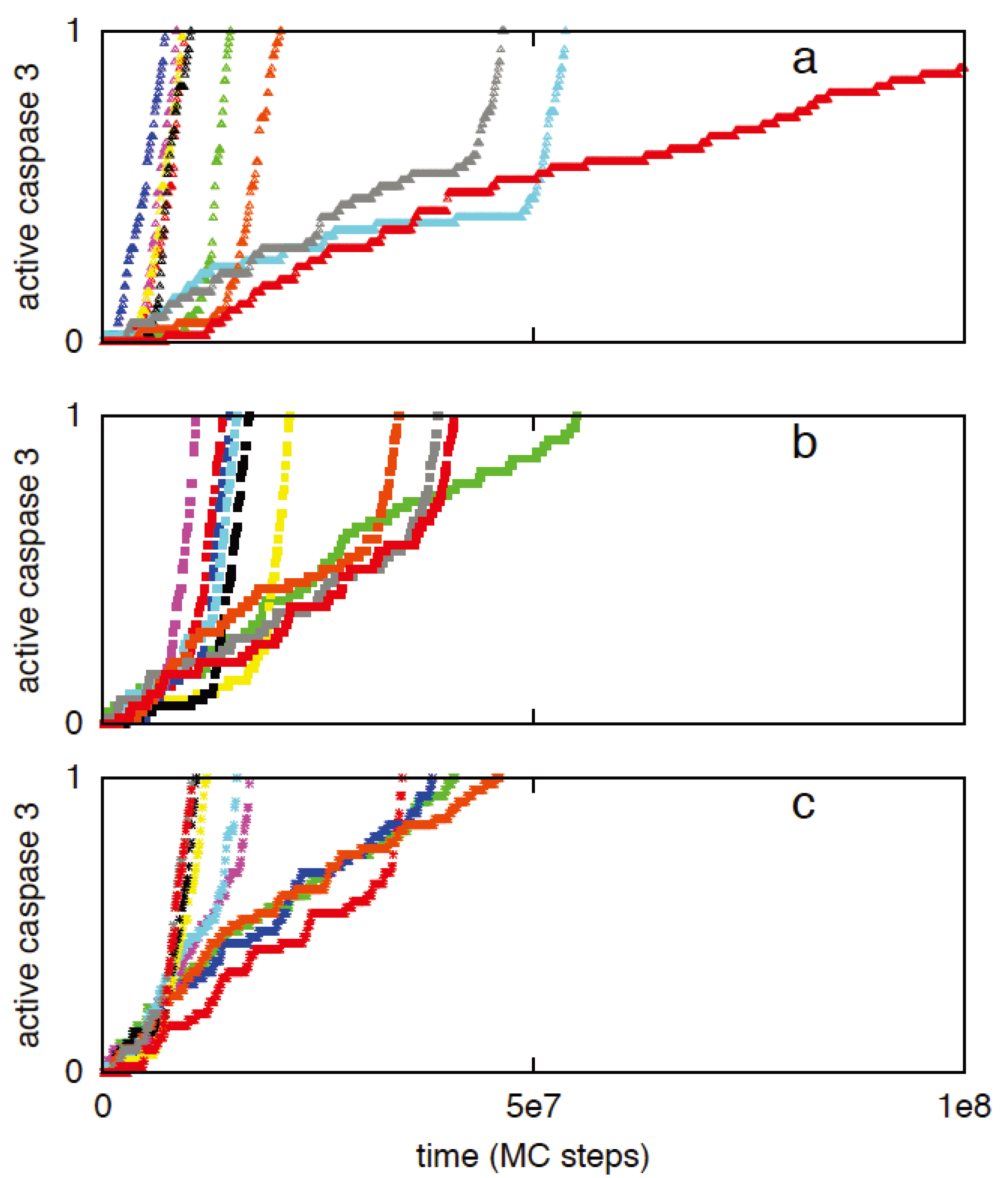
3.2.3. Increased Rate of Apoptosome Formation Favors the Type 2 Pathway
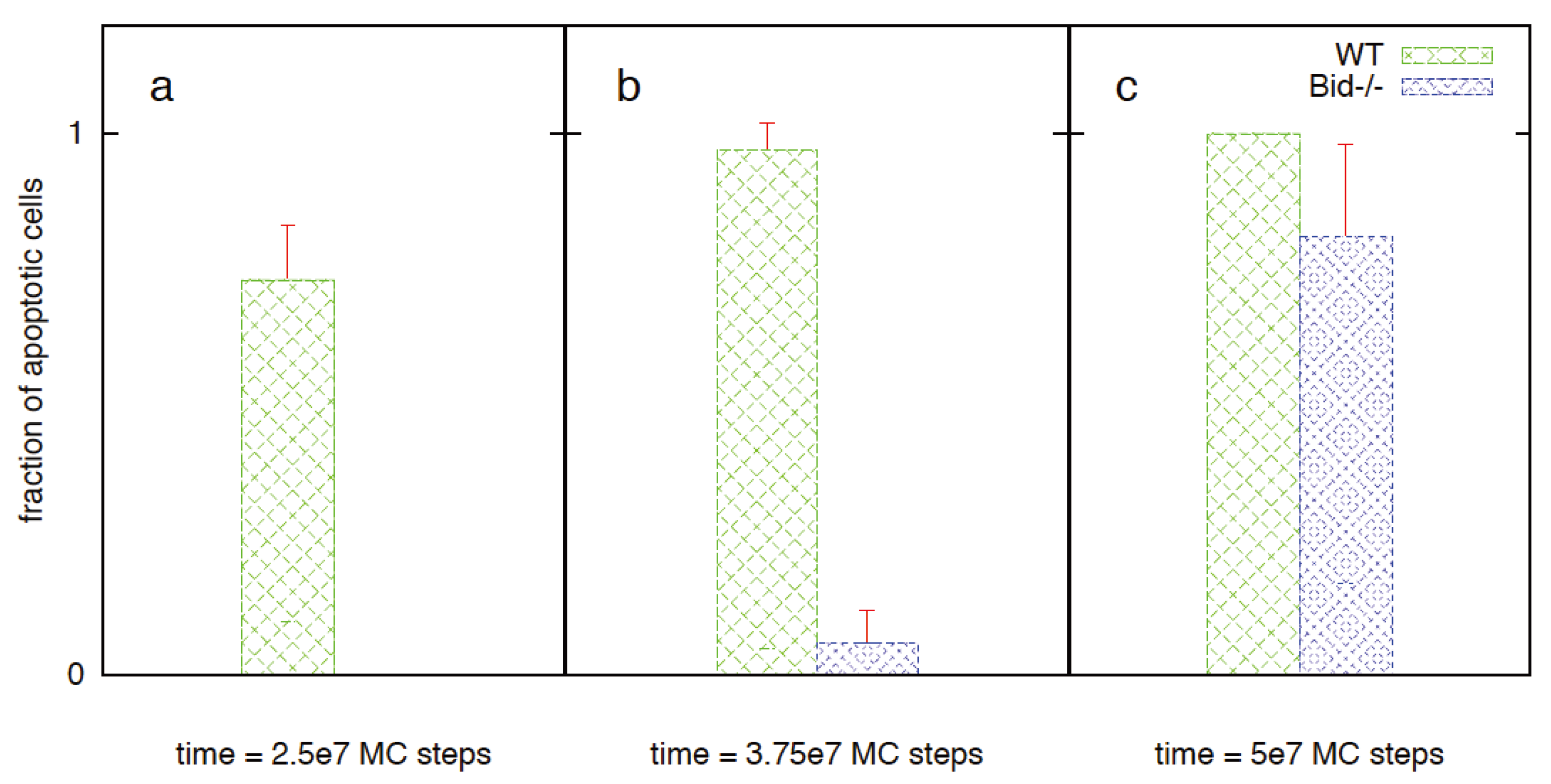
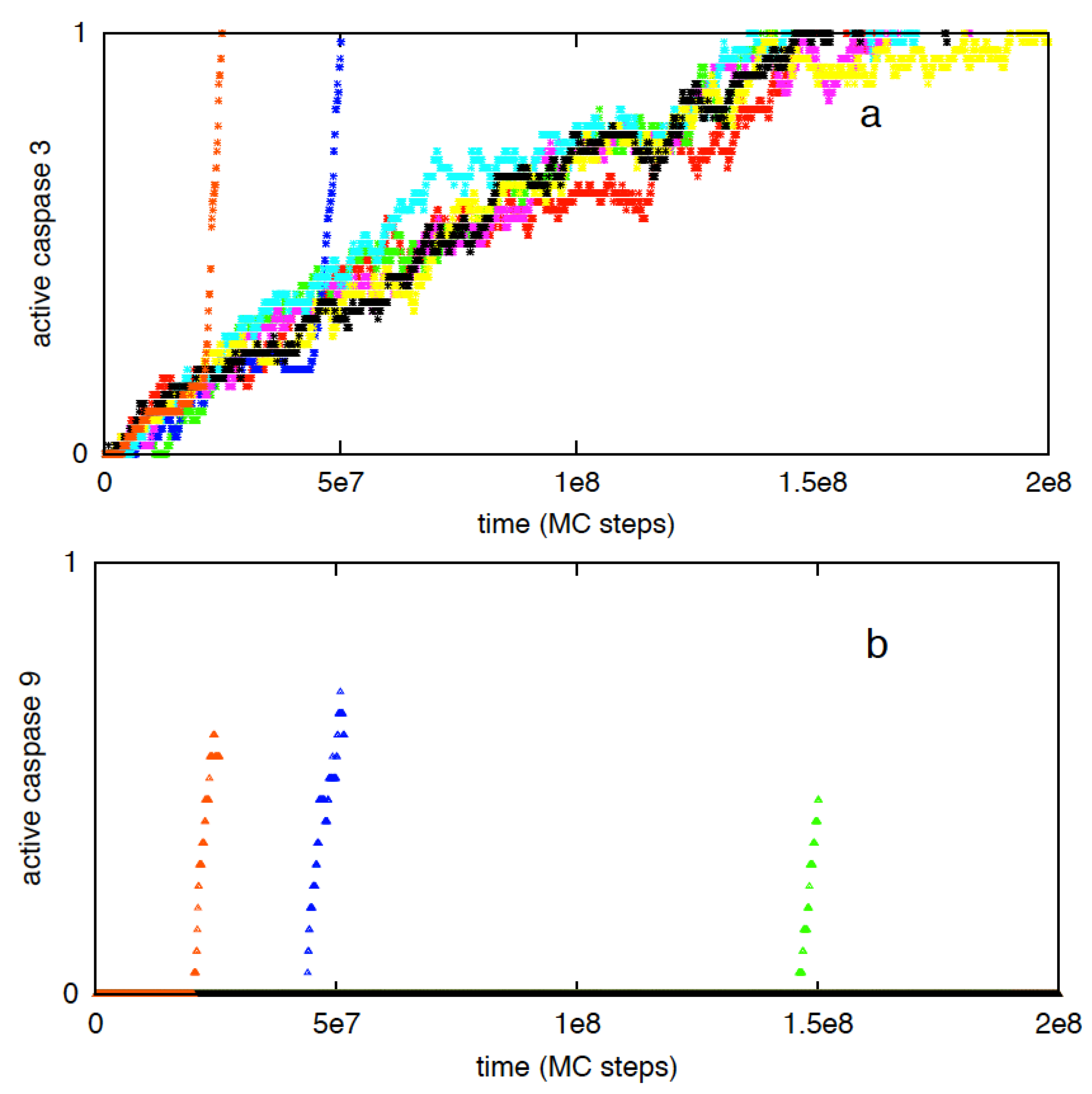
| Kinetic rate constant for Cyto c–Apaf binding | Time to (first) apoptosome formation (MC steps) |
|---|---|
| 2-fold high | 1.03 × 107 |
| normal | 2.47 × 107 |
| 2-fold low | 3.90 × 107 * |
3.3. Robustness of Type 1/Type 2 Choice in Apoptosis Regulation

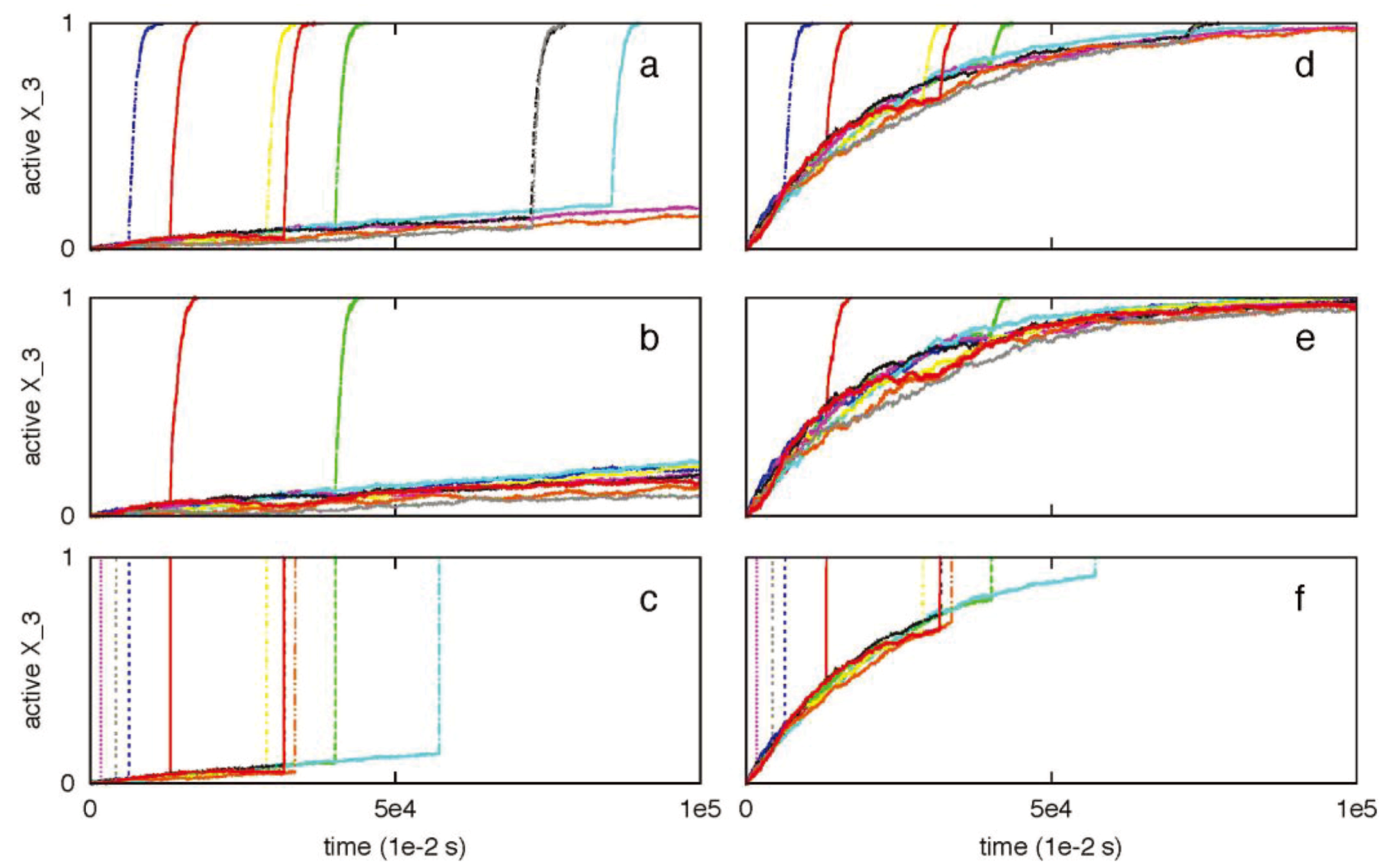

3.4. Regulation of Type 1/Type 2 Choice in Cancer Cells Having Over-Expressed Anti-Apoptotic Proteins Bcl2 and XIAP
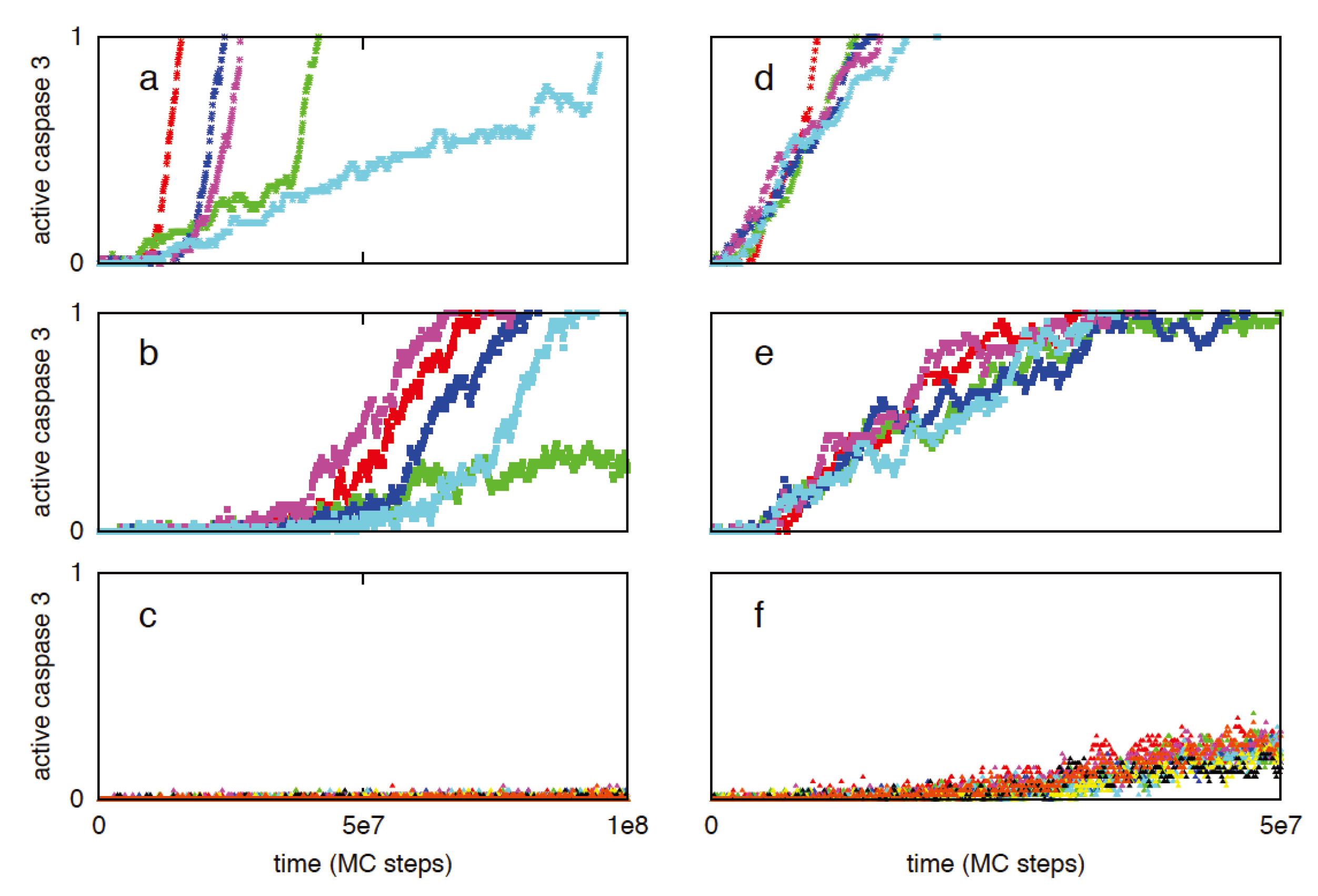
| Cell-to-cell variability in caspase 3 activation | |||
|---|---|---|---|
| Time-to-Cyto c /Smac release | Time-to-apoptosome formation | Time-to-death | |
| Cell 1 | 6.2 × 106 | 8.6 × 106 | 1.6 × 107 |
| Cell 2 | 7.6 × 106 | 9.1 × 107 | 9.5 × 107 |
| Cell 3 | 7.0 × 106 | 4.2 × 107 | 4.7 × 107 |


3.5. Apoptosis in Cancer Cells Having Heightened Sensitivity to Death Receptor Activation: A Mechanism for Selective Targeting of Cancer Cells
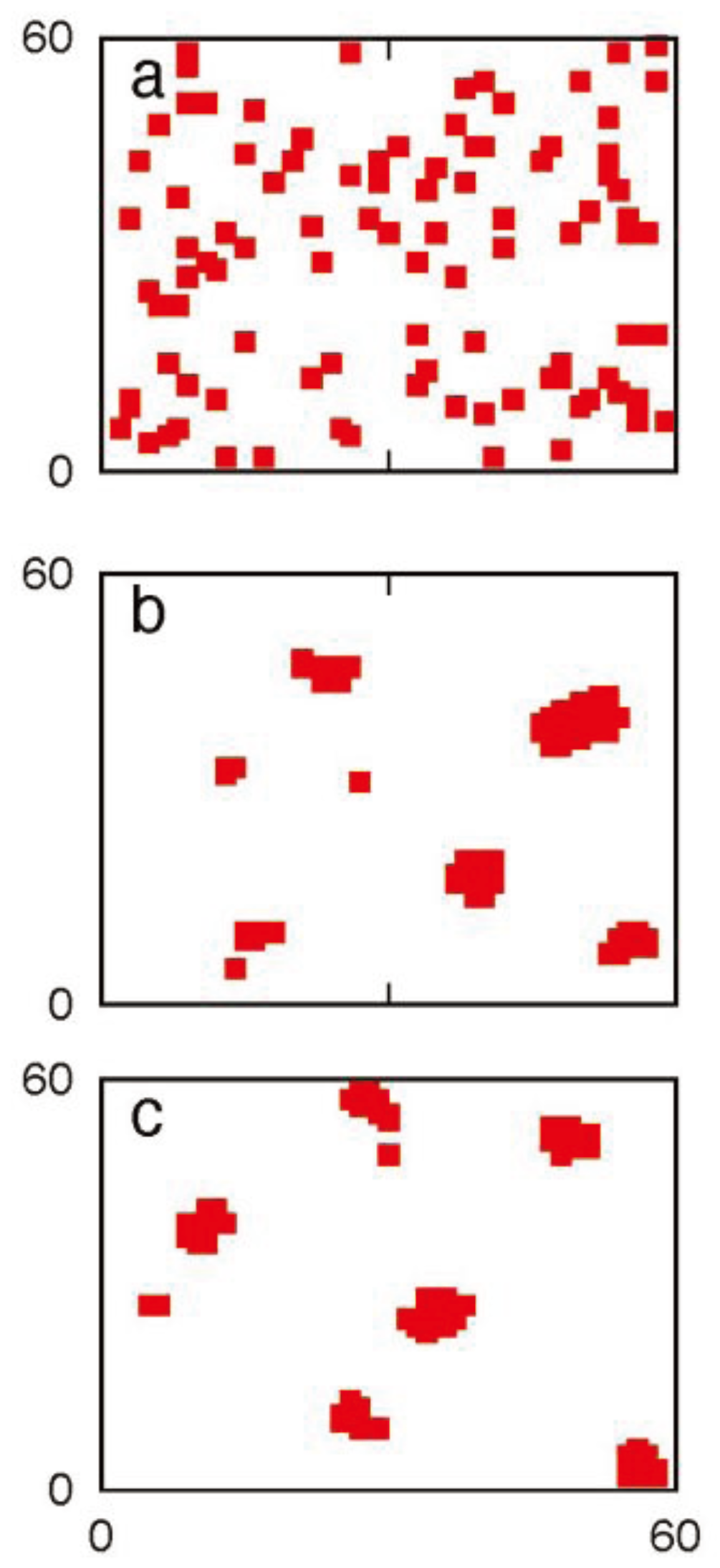

3.6. Combined Death Ligand Induction and XIAP Inhibition can Be an Optimal Strategy to Kill Cancer Cells: Maximizing Specificity and Minimizing Cell-to-Cell Stochastic Variability

4. Conclusions
Appendix I
| Apoptosis signaling reactions | Kinetic rate constants |
|---|---|
| DISC + ProCaspase8 ⇄ Disc-ProCaspase8 | kon: 3.5 × 10−3 nM−1 s−1; koff: 1.8 × 10−2 s−1 [6] |
| Disc-ProCaspase8 + ProCaspase8 ⇄ Disc-ProCaspase8_2 | kon: 3.5 × 10−3 nM−1 s−1; koff: 1.8 × 10−2 s−1 [6] |
| Disc-ProCaspase8_2 → DISC + 2 P43/41 | kcat: 3.0 × 10−1 s−1 [6] |
| P43/41→ Caspase 8 | kcat: 1.0 × 10−1 s−1 [6] |
| Caspase 8 + Bid ⇄ Caspase8-Bid | kon: 5.0 × 10−3 nM−1 s−1; koff: 5.0 × 10−3 s−1 [6] |
| Caspase8-Bid → Caspase 8 + tBid | kcat: 1.0 × 10−1 s−1 [6] |
| Caspase 8 + ProCaspase 3 ⇄ Caspase8-ProCaspase 3 | kon: 1.0 × 10−5 nM−1 s−1; koff: 6.0 × 10−2 s−1 [6] |
| Caspase8-ProCaspase 3 → Caspase 8 + Caspase 3 | kcat: 1.0 × 10−1 s−1 [6] |
| Bid + Bax ⇄ Bid-Bax | kon: 1.0 × 10−6 nM−1 s−1; koff: 1.0 × 10−2 s−1 [12] |
| Bid-Bax + Bax ⇄ Bid-Bax_2 | kon: 1.0 × 10−6 nM−1 s−1; koff: 1.0 × 10−2 s−1 [12] |
| Bid-Bax_2 → Bid + Bax_2 | kcat: 1.0 × 10−2 s−1 [6] |
| Bid + Bcl2⇄ Bid-Bcl2 | kon: 1.0 × 10−5 nM−1 s−1; koff: 1.0 × 10−2 s−1 [12] |
| tBid + Bax ⇄ tBid-Bax | kon: 2.0 × 10−4 nM−1 s−1; koff: 2.0 × 10−2 s−1 [6] |
| tBid-Bax + Bax ⇄ tBid-Bax_2 | kon: 2.0 × 10−4 nM−1 s−1; koff: 2.0 × 10−2 s−1 [6] |
| tBid-Bax_2 → tBid + Bax_2 | kcat: 2.0 × 10−2 s−1 (taken as ~koff) |
| Bcl2 + tBid ⇄ Bcl2-tBid | kon: 2.0 × 10−3 nM−1 s−1; koff: 2.0 × 10−2 s−1 [6,67] |
| Bcl2 + Bax ⇄ Bcl2-Bax | kon: 2.0 × 10−3 nM−1 s−1; koff: 2.0 × 10−2 s−1 [6,67] |
| Cytochrome C + Apaf ⇄ Cytochrome C-Apaf | kon: 2.8 × 10−7 nM−1 s−1; koff: 5.7 × 10−3 s−1 [6] |
| Apaf + Apaf ⇄ Apaf-Apaf | kon: 1.0 × 10−2 nM−1 s−1; koff: 10 s−1 (μM affinity based on CARD-CARD interaction [7]) |
| XIAP + Smac ⇄ XIAP-Smac | kon: 7.0 × 10−3 nM−1 s−1; koff: 2.2 × 10−3 s−1 [6,68] |
| Apoptosome + ProCaspase9 ⇄ Apoptosome-ProCaspase 9 | kon: 2.8 × 10−4 nM−1 s−1; koff: 7.5 × 10−2 s−1 [6] |
| Apoptosome-ProCaspase9 + ProCaspase9 ⇄ Apoptosome-ProCaspase 9_2 | kon: 2.8 × 10−4 nM−1 s−1; koff: 7.5 × 10−2 s−1 [6] |
| Apoptosome-ProCaspase 9_2 → Apoptosome + 2 Caspase 9 | kcat: 7.0 × 10−1 s−1 [6] |
| XIAP + ProCaspase 9 ⇄ XIAP-ProCaspase 9 | kon: 5.0 × 10−3 nM−1 s−1; koff: 3.5 × 10−3 s−1 (taken to be same as that for Caspase 9) |
| XIAP + Caspase 9 ⇄ XIAP-Caspase 9 | kon: 5.0 × 10−3 nM−1 s−1; koff: 3.5 × 10−3 s−1 [25,69] |
| Caspase 9 + ProCaspase 3 ⇄ Caspase 9-ProCaspase 3 | kon: 2.0 × 10−5 nM−1 s−1 [6,26]; koff: 5.7 × 10−1 s−1 [6,25] |
| Caspase 9-ProCaspase 3 → Caspase 9 + Caspase 3 | kcat: 4.8 s−1 [6,70] |
| XIAP + Caspase 3 ⇄ XIAP-Caspase 3 | kon: 2.5 × 10−3 nM−1 s−1; koff: 2.4 × 10−3 s−1 [6,71] |
| Molecule type | Concentration (nM) |
|---|---|
| DISC (adaptor such as FADD/TRADD) | 100 [32] |
| ProCaspase 8 | 30 [3,6,72] |
| Bid | 33 [6,25] |
| Bax | 83 [6] |
| Bcl-2 | 75 [6] |
| Cytochrome C | 100 [6] |
| Smac | 50 (~100 nM in [6]) |
| Apaf | 100 [6]; 20 [26] |
| XIAP | 30 [3,6] |
| ProCaspase 9 | 20 [6,70,73] |
| ProCaspase 3 | 100 [72] (also see [3,6,23]) |
| Apoptosis signaling reactions | Probabilistic parameters used in the Monte Carlo simulation of apoptosis signaling (Pon/Poff/Pcat) |
|---|---|
| DISC + ProCaspase8 ⇄ Disc-ProCaspase8 | Pon: 1.0; Poff: 1.8 × 10−6 s−1 (Pon taken as 1.0 instead of 0.35 as an approximation) |
| Disc-ProCaspase8 + ProCaspase8 ⇄ Disc-ProCaspase8_2 | Pon: 1.0; Poff:1.8 × 10−6 s−1 |
| Disc-ProCaspase8_2 → DISC + 2 P43/41 | Pcat: 3.0 × 10−5 |
| P43/41→ Caspase 8 | Pcat: 1.0 × 10−5 |
| Caspase 8 + Bid ⇄ Caspase8-Bid | Pon: 5.0 × 10−1; Poff: 5.0 × 10−7 |
| Caspase8-Bid → Caspase 8 + tBid | Pcat: 1.0 × 10−5 |
| Caspase 8 + ProCaspase 3 ⇄ Caspase8-ProCaspase 3 | Pon: 1.0 × 10−3; Poff: 6.0 × 10−6 |
| Caspase8-ProCaspase 3 → Caspase 8 + Caspase 3 | Pcat: 1.0 × 10−5 |
| Bid + Bax ⇄ Bid-Bax | Pon: 1.0 × 10−4; Poff: 1.0 × 10−6 |
| Bid-Bax + Bax ⇄ Bid-Bax_2 | Pon: 1.0 × 10−4; Poff: 1.0 × 10−6 |
| Bid-Bax_2 → Bid + Bax_2 | Pcat: 1.0 × 10−6 s−1 |
| Bid + Bcl2⇄ Bid-Bcl2 | Pon: 1.0 × 10−3; Poff: 1.0 × 10−6 |
| tBid + Bax ⇄ tBid-Bax | Pon: 2.0 × 10−2; Poff: 2.0 × 10−2 s−1 |
| tBid-Bax + Bax ⇄ tBid-Bax_2 | Pon: 2.0 × 10−4 nM−1 s−1; Poff: 2.0 × 10−2 s−1 |
| tBid-Bax_2 → tBid + Bax_2 | Pcat: 2.0 × 10−2 s−1 |
| Bcl2 + tBid ⇄ Bcl2-tBid | Pon: 2.0 × 10−1; Poff: 2.0 × 10−6 |
| Bcl2 + Bax ⇄ Bcl2-Bax | Pon: 2.0 × 10−1; Poff: 2.0 × 10−6 |
| Cytochrome C + Apaf ⇄ Cytochrome C-Apaf | Pon: 2.8 × 10−5; Poff: 5.7 × 10−7 |
| Apaf + Apaf ⇄ Apaf-Apaf | Pon: 1.0 × 100; Poff: 1.0 × 10−3 |
| XIAP + Smac ⇄ XIAP-Smac | Pon: 7.0 × 10−1; Poff: 2.2 × 10−7 |
| Apoptosome + ProCaspase9 ⇄ Apoptosome-ProCaspase 9 | Pon: 2.8 × 10−2; Poff: 7.5 × 10−6 |
| Apoptosome-ProCaspase9 + ProCaspase9 ⇄ Apoptosome-ProCaspase 9_2 | Pon: 2.8 × 10−2; Poff: 7.5 × 10−6 |
| Apoptosome-ProCaspase 9_2 → Apoptosome + 2 Caspase 9 | Pcat: 7.0 × 10−5 s−1 |
| XIAP + ProCaspase 9 ⇄ XIAP-ProCaspase 9 | Pon: 5.0 × 10−1; Poff: 3.5 × 10−7 |
| XIAP + Caspase 9 ⇄ XIAP-Caspase 9 | Pon: 5.0 × 10−1; Poff: 3.5 × 10−7 |
| Caspase 9 + ProCaspase 3 ⇄ Caspase 9-ProCaspase 3 | Pon: 2.0 × 10−3; Poff: 5.7 × 10−5 |
| Caspase 9-ProCaspase 3 → Caspase 9 + Caspase 3 | Pcat: 4.8 × 10−4 |
| XIAP + Caspase 3 ⇄ XIAP-Caspase 3 | Pon: 2.5 × 10−1; Poff: 2.4 × 10−7 |
Appendix II











Acknowledgments
Conflict of Interest
References
- Scaffidi, C.; Fulda, S.; Srinivasan, A.; Friesen, C.; Li, F.; Tomaselli, K.J.; Debatin, K.M.; Krammer, P.H.; Peter, M.E. Two cd95 (apo-1/fas) signaling pathways. EMBO J. 1998, 17, 1675–1687. [Google Scholar] [CrossRef]
- Scaffidi, C.; Schmitz, I.; Zha, J.; Korsmeyer, S.J.; Krammer, P.H.; Peter, M.E. Differential modulation of apoptosis sensitivity in cd95 type i and type ii cells. J. Biol. Chem. 1999, 274, 22532–22538. [Google Scholar]
- Sun, X.M.; Bratton, S.B.; Butterworth, M.; MacFarlane, M.; Cohen, G.M. Bcl-2 and bcl-xl inhibit cd95-mediated apoptosis by preventing mitochondrial release of smac/diablo and subsequent inactivation of x-linked inhibitor-of-apoptosis protein. J. Biol. Chem. 2002, 277, 11345–11351. [Google Scholar]
- Jost, P.J.; Grabow, S.; Gray, D.; McKenzie, M.D.; Nachbur, U.; Huang, D.C.; Bouillet, P.; Thomas, H.E.; Borner, C.; Silke, J.; et al. Xiap discriminates between type i and type ii fas-induced apoptosis. Nature 2009, 460, 1035–1039. [Google Scholar] [CrossRef]
- Meng, X.W.; Peterson, K.L.; Dai, H.; Schneider, P.; Lee, S.H.; Zhang, J.S.; Koenig, A.; Bronk, S.; Billadeau, D.D.; Gores, G.J.; et al. High cell surface death receptor expression determines type i versus type ii signaling. J. Biol. Chem. 2011, 286, 35823–35833. [Google Scholar] [CrossRef]
- Hua, F.; Cornejo, M.G.; Cardone, M.H.; Stokes, C.L.; Lauffenburger, D.A. Effects of bcl-2 levels on fas signaling-induced caspase-3 activation: Molecular genetic tests of computational model predictions. J. Immunol. 2005, 175, 985–995. [Google Scholar]
- Raychaudhuri, S.; Willgohs, E.; Nguyen, T.N.; Khan, E.M.; Goldkorn, T. Monte carlo simulation of cell death signaling predicts large cell-to-cell stochastic fluctuations through the type 2 pathway of apoptosis. Biophys. J. 2008, 95, 3559–3562. [Google Scholar] [CrossRef]
- Raychaudhuri, S. A minimal model of signaling network elucidates cell-to-cell stochastic variability in apoptosis. PLoS One 2010, 5, e11930. [Google Scholar] [CrossRef]
- Raychaudhuri, S.; Skommer, J.; Henty, K.; Birch, N.; Brittain, T. Neuroglobin protects nerve cells from apoptosis by inhibiting the intrinsic pathway of cell death. Apoptosis 2010, 15, 401–411. [Google Scholar] [CrossRef]
- Skommer, J.; Brittain, T.; Raychaudhuri, S. Bcl-2 inhibits apoptosis by increasing the time-to-death and intrinsic cell-to-cell variations in the mitochondrial pathway of cell death. Apoptosis 2010, 15, 1223–1233. [Google Scholar] [CrossRef]
- Albeck, J.G.; Burke, J.M.; Spencer, S.L.; Lauffenburger, D.A.; Sorger, P.K. Modeling a snap-action, variable-delay switch controlling extrinsic cell death. PLoS Biol. 2008, 6, 2831–2852. [Google Scholar]
- Raychaudhuri, S.; Das, S.C. Low probability activation of bax/bak can induce selective killing of cancer cells by generating heterogeneoity in apoptosis. J. Healthc. Eng. 2013, 4, 47–66. [Google Scholar] [CrossRef]
- Skommer, J.; Das, S.C.; Nair, A.; Brittain, T.; Raychaudhuri, S. Nonlinear regulation of commitment to apoptosis by simultaneous inhibition of bcl-2 and xiap in leukemia and lymphoma cells. Apoptosis 2011, 16, 619–626. [Google Scholar] [CrossRef]
- Skommer, J.; Raychaudhuri, S.; Wlodkowic, D. Timing is everything: Stochastic origins of cell-to-cell variability in cancer cell death. Front. Biosci. 2011, 16, 307–314. [Google Scholar] [CrossRef]
- Erwig, L.P.; Henson, P.M. Clearance of apoptotic cells by phagocytes. Cell Death Differ. 2008, 15, 243–250. [Google Scholar] [CrossRef]
- Raychaudhuri, S. How can we kill cancer cells: Insights from the computational models of apoptosis. World J. Clin. Oncol. 2010, 1, 24–28. [Google Scholar] [CrossRef]
- Certo, M.; Del Gaizo Moore, V.; Nishino, M.; Wei, G.; Korsmeyer, S.; Armstrong, S.A.; Letai, A. Mitochondria primed by death signals determine cellular addiction to antiapoptotic bcl-2 family members. Cancer Cell 2006, 9, 351–365. [Google Scholar] [CrossRef]
- Kurita, S.; Mott, J.L.; Cazanave, S.C.; Fingas, C.D.; Guicciardi, M.E.; Bronk, S.F.; Roberts, L.R.; Fernandez-Zapico, M.E.; Gores, G.J. Hedgehog inhibition promotes a switch from type ii to type i cell death receptor signaling in cancer cells. PLoS One 2011, 6, e18330. [Google Scholar] [CrossRef]
- Shirley, S.; Morizot, A.; Micheau, O. Regulating trail receptor-induced cell death at the membrane: A deadly discussion. Recent Pat. Anticancer Drug Discov. 2011, 6, 311–323. [Google Scholar] [CrossRef]
- Del Gaizo Moore, V.; Letai, A. Bh3 profiling—Measuring integrated function of the mitochondrial apoptotic pathway to predict cell fate decisions. Cancer Lett. 2012, 332, 202–205. [Google Scholar] [CrossRef]
- Fussenegger, M.; Bailey, J.E.; Varner, J. A mathematical model of caspase function in apoptosis. Nat. Biotechnol. 2000, 18, 768–774. [Google Scholar] [CrossRef]
- Bentele, M.; Lavrik, I.; Ulrich, M.; Stosser, S.; Heermann, D.W.; Kalthoff, H.; Krammer, P.H.; Eils, R. Mathematical modeling reveals threshold mechanism in cd95-induced apoptosis. J. Cell Biol. 2004, 166, 839–851. [Google Scholar] [CrossRef]
- Eissing, T.; Conzelmann, H.; Gilles, E.D.; Allgower, F.; Bullinger, E.; Scheurich, P. Bistability analyses of a caspase activation model for receptor-induced apoptosis. J. Biol. Chem. 2004, 279, 36892–36897. [Google Scholar]
- Stucki, J.W.; Simon, H.U. Mathematical modeling of the regulation of caspase-3 activation and degradation. J. Theor. Biol. 2005, 234, 123–131. [Google Scholar] [CrossRef]
- Bagci, E.Z.; Vodovotz, Y.; Billiar, T.R.; Ermentrout, G.B.; Bahar, I. Bistability in apoptosis: Roles of bax, bcl-2, and mitochondrial permeability transition pores. Biophys. J. 2006, 90, 1546–1559. [Google Scholar] [CrossRef]
- Legewie, S.; Bluthgen, N.; Herzel, H. Mathematical modeling identifies inhibitors of apoptosis as mediators of positive feedback and bistability. PLoS Comput. Biol. 2006, 2, e120. [Google Scholar] [CrossRef]
- Rehm, M.; Huber, H.J.; Dussmann, H.; Prehn, J.H. Systems analysis of effector caspase activation and its control by x-linked inhibitor of apoptosis protein. EMBO J. 2006, 25, 4338–4349. [Google Scholar] [CrossRef]
- Chen, C.; Cui, J.; Lu, H.; Wang, R.; Zhang, S.; Shen, P. Modeling of the role of a bax-activation switch in the mitochondrial apoptosis decision. Biophys. J. 2007, 92, 4304–4315. [Google Scholar] [CrossRef]
- Ryu, S.; Lin, S.C.; Ugel, N.; Antoniotti, M.; Mishra, B. Mathematical modeling of the formation of apoptosome in intrinsic pathway of apoptosis. Syst. Synth. Biol. 2008, 2, 49–66. [Google Scholar] [CrossRef]
- Rehm, M.; Huber, H.J.; Hellwig, C.T.; Anguissola, S.; Dussmann, H.; Prehn, J.H. Dynamics of outer mitochondrial membrane permeabilization during apoptosis. Cell Death Differ. 2009, 16, 613–623. [Google Scholar] [CrossRef]
- Spencer, S.L.; Gaudet, S.; Albeck, J.G.; Burke, J.M.; Sorger, P.K. Non-genetic origins of cell-to-cell variability in trail-induced apoptosis. Nature 2009, 459, 428–432. [Google Scholar] [CrossRef]
- Fricker, N.; Beaudouin, J.; Richter, P.; Eils, R.; Krammer, P.H.; Lavrik, I.N. Model-based dissection of cd95 signaling dynamics reveals both a pro- and antiapoptotic role of c-flipl. J. Cell Biol. 2010, 190, 377–389. [Google Scholar] [CrossRef]
- Aldridge, B.B.; Gaudet, S.; Lauffenburger, D.A.; Sorger, P.K. Lyapunov exponents and phase diagrams reveal multi-factorial control over trail-induced apoptosis. Mol. Syst. Biol. 2011, 7, 553. [Google Scholar]
- Gu, C.; Zhang, J.; Chen, Y.; Lei, J. A trigger model of apoptosis induced by tumor necrosis factor signaling. BMC Syst. Biol. 2011, 5, S13. [Google Scholar]
- Hong, J.Y.; Kim, G.H.; Kim, J.W.; Kwon, S.S.; Sato, E.F.; Cho, K.H.; Shim, E.B. Computational modeling of apoptotic signaling pathways induced by cisplatin. BMC Syst. Biol. 2012, 6, 122. [Google Scholar] [CrossRef]
- Peter, M.E.; Krammer, P.H. The cd95(apo-1/fas) disc and beyond. Cell Death Differ. 2003, 10, 26–35. [Google Scholar] [CrossRef]
- Reddy, A.S.; Chilukuri, S.; Raychaudhuri, S. The network of receptors characterize b cell receptor micro- and macroclustering in a monte carlo model. J. Phys. Chem. B 2010, 114, 487–494. [Google Scholar] [CrossRef]
- Raychaudhuri, S. Kinetic Monte Carlo simulation in biophysics and systems biology. In Theory and Applications of Monte Carlo Simulations; Chan, W.K.V., Ed.; InTech: Rijeka, Croatia, 2013. [Google Scholar] [Green Version]
- Goldstein, J.C.; Waterhouse, N.J.; Juin, P.; Evan, G.I.; Green, D.R. The coordinate release of cytochrome c during apoptosis is rapid, complete and kinetically invariant. Nat. Cell Biol. 2000, 2, 156–162. [Google Scholar] [CrossRef]
- Dussmann, H.; Rehm, M.; Concannon, C.G.; Anguissola, S.; Wurstle, M.; Kacmar, S.; Voller, P.; Huber, H.J.; Prehn, J.H. Single-cell quantification of bax activation and mathematical modelling suggest pore formation on minimal mitochondrial bax accumulation. Cell Death Differ. 2010, 17, 278–290. [Google Scholar] [CrossRef]
- Lee, J.K.; Lu, S.; Madhukar, A. Real-time dynamics of Ca2+, caspase-3/7, and morphological changes in retinal ganglion cell apoptosis under elevated pressure. PLoS One 2010, 5, e13437. [Google Scholar]
- Newman, M.E.J.; Barkema, G.T. Monte Carlo Methods in Statistical Physics; Oxford University Press: New York, NY, USA, 1999. [Google Scholar]
- Lopez-Araiza, H.; Ventura, J.L.; Lopez-Diazguerrero, N.E.; Gonzalez-Marquez, H.; Gutierrez-Ruiz, M.C.; Zentella, A.; Konigsberg, M. Organ- and tissue-specific alterations in the anti-apoptotic protein bcl-2 in cd1 female mice of different ages. Biogerontology 2006, 7, 63–67. [Google Scholar] [CrossRef]
- Maas, C.; Verbrugge, I.; de Vries, E.; Savich, G.; van de Kooij, L.W.; Tait, S.W.; Borst, J. Smac/diablo release from mitochondria and xiap inhibition are essential to limit clonogenicity of type i tumor cells after trail receptor stimulation. Cell Death Differ. 2010, 17, 1613–1623. [Google Scholar] [CrossRef]
- Yakovlev, A.G.; Ota, K.; Wang, G.; Movsesyan, V.; Bao, W.L.; Yoshihara, K.; Faden, A.I. Differential expression of apoptotic protease-activating factor-1 and caspase-3 genes and susceptibility to apoptosis during brain development and after traumatic brain injury. J. Neurosci. 2001, 21, 7439–7446. [Google Scholar]
- Johnson, C.E.; Huang, Y.Y.; Parrish, A.B.; Smith, M.I.; Vaughn, A.E.; Zhang, Q.; Wright, K.M.; van Dyke, T.; Wechsler-Reya, R.J.; Kornbluth, S.; et al. Differential apaf-1 levels allow cytochrome c to induce apoptosis in brain tumors but not in normal neural tissues. Proc. Natl. Acad. Sci. USA 2007, 104, 20820–20825. [Google Scholar] [CrossRef]
- Matsuyama, S.; Reed, J.C. Mitochondria-dependent apoptosis and cellular ph regulation. Cell Death Differ. 2000, 7, 1155–1165. [Google Scholar] [CrossRef]
- Kang, Z.; Sun, S.Y.; Cao, L. Activating death receptor dr5 as a therapeutic strategy for rhabdomyosarcoma. ISRN Oncol. 2012, 2012, 395952. [Google Scholar]
- Weller, M.; Kleihues, P.; Dichgans, J.; Ohgaki, H. Cd95 ligand: Lethal weapon against malignant glioma? Brain Pathol. 1998, 8, 285–293. [Google Scholar]
- Pan, G.; Ni, J.; Wei, Y.F.; Yu, G.; Gentz, R.; Dixit, V.M. An antagonist decoy receptor and a death domain-containing receptor for trail. Science 1997, 277, 815–818. [Google Scholar] [CrossRef]
- Sheridan, J.P.; Marsters, S.A.; Pitti, R.M.; Gurney, A.; Skubatch, M.; Baldwin, D.; Ramakrishnan, L.; Gray, C.L.; Baker, K.; Wood, W.I.; et al. Control of trail-induced apoptosis by a family of signaling and decoy receptors. Science 1997, 277, 818–821. [Google Scholar] [CrossRef]
- Ho, I.A.; Ng, W.H.; Lam, P.Y. Fasl and fadd delivery by a glioma-specific and cell cycle-dependent hsv-1 amplicon virus enhanced apoptosis in primary human brain tumors. Mol. Cancer 2010, 9, 270. [Google Scholar] [CrossRef]
- Scott, F.L.; Stec, B.; Pop, C.; Dobaczewska, M.K.; Lee, J.J.; Monosov, E.; Robinson, H.; Salvesen, G.S.; Schwarzenbacher, R.; Riedl, S.J. The fas-fadd death domain complex structure unravels signalling by receptor clustering. Nature 2009, 457, 1019–1022. [Google Scholar] [CrossRef]
- Song, J.H.; Tse, M.C.; Bellail, A.; Phuphanich, S.; Khuri, F.; Kneteman, N.M.; Hao, C. Lipid rafts and nonrafts mediate tumor necrosis factor related apoptosis-inducing ligand induced apoptotic and nonapoptotic signals in non small cell lung carcinoma cells. Cancer Res. 2007, 67, 6946–6955. [Google Scholar] [CrossRef]
- Legembre, P.; Daburon, S.; Moreau, P.; Ichas, F.; de Giorgi, F.; Moreau, J.F.; Taupin, J.L. Amplification of fas-mediated apoptosis in type ii cells via microdomain recruitment. Mol. Cell. Biol. 2005, 25, 6811–6820. [Google Scholar] [CrossRef]
- Picone, P.; Carrotta, R.; Montana, G.; Nobile, M.R.; San Biagio, P.L.; di Carlo, M. Abeta oligomers and fibrillar aggregates induce different apoptotic pathways in lan5 neuroblastoma cell cultures. Biophys. J. 2009, 96, 4200–4211. [Google Scholar]
- Degterev, A.; Yuan, J. Expansion and evolution of cell death programmes. Nat. Rev. Mol. Cell Biol. 2008, 9, 378–390. [Google Scholar] [CrossRef]
- Walsh, C.M.; Edinger, A.L. The complex interplay between autophagy, Apoptosis, And necrotic signals promotes t-cell homeostasis. Immunol. Rev. 2010, 236, 95–109. [Google Scholar] [CrossRef]
- Mollinedo, F.; Gajate, C. Fas/cd95 death receptor and lipid rafts: New targets for apoptosis-directed cancer therapy. Drug Resist. Updates 2006, 9, 51–73. [Google Scholar] [CrossRef]
- Gulbins, E.; Kolesnick, R. Raft ceramide in molecular medicine. Oncogene 2003, 22, 7070–7077. [Google Scholar] [CrossRef]
- Quintana, E.; Shackleton, M.; Sabel, M.S.; Fullen, D.R.; Johnson, T.M.; Morrison, S.J. Efficient tumour formation by single human melanoma cells. Nature 2008, 456, 593–598. [Google Scholar] [CrossRef]
- Takeda, K.; Hayakawa, Y.; Smyth, M.J.; Kayagaki, N.; Yamaguchi, N.; Kakuta, S.; Iwakura, Y.; Yagita, H.; Okumura, K. Involvement of tumor necrosis factor-related apoptosis-inducing ligand in surveillance of tumor metastasis by liver natural killer cells. Nat. Med. 2001, 7, 94–100. [Google Scholar] [CrossRef]
- Safa, A.R.; Pollok, K.E. Targeting the anti-apoptotic protein c-flip for cancer therapy. Cancers 2011, 3, 1639–1671. [Google Scholar] [CrossRef]
- Picarda, G.; Trichet, V.; Teletchea, S.; Heymann, D.; Redini, F. Trail receptor signaling and therapeutic option in bone tumors: The trap of the bone microenvironment. Am. J. Cancer Res. 2012, 2, 45–64. [Google Scholar]
- Accordi, B.; Pillozzi, S.; Dell’Orto, M.C.; Cazzaniga, G.; Arcangeli, A.; Kronnie, G.T.; Basso, G. Hepatocyte growth factor receptor c-met is associated with fas and when activated enhances drug-induced apoptosis in pediatric b acute lymphoblastic leukemia with tel-aml1 translocation. J. Biol. Chem. 2007, 282, 29384–29393. [Google Scholar]
- Zhao, Y.; Difrancesca, D.; Wang, X.; Zarnegar, R.; Michalopoulos, G.K.; Yin, X.M. Promotion of fas-mediated apoptosis in type ii cells by high doses of hepatocyte growth factor bypasses the mitochondrial requirement. J. Cell. Physiol. 2007, 213, 556–563. [Google Scholar] [CrossRef]
- Letai, A.; Bassik, M.C.; Walensky, L.D.; Sorcinelli, M.D.; Weiler, S.; Korsmeyer, S.J. Distinct bh3 domains either sensitize or activate mitochondrial apoptosis, serving as prototype cancer therapeutics. Cancer Cell 2002, 2, 183–192. [Google Scholar] [CrossRef]
- Huang, Y.; Rich, R.L.; Myszka, D.G.; Wu, H. Requirement of both the second and third bir domains for the relief of x-linked inhibitor of apoptosis protein (xiap)-mediated caspase inhibition by smac. J. Biol. Chem. 2003, 278, 49517–49522. [Google Scholar] [CrossRef]
- Asthagiri, A.R.; Lauffenburger, D.A. A computational study of feedback effects on signal dynamics in a mitogen-activated protein kinase (mapk) pathway model. Biotechnol. Progress 2001, 17, 227–239. [Google Scholar] [CrossRef]
- Renatus, M.; Stennicke, H.R.; Scott, F.L.; Liddington, R.C.; Salvesen, G.S. Dimer formation drives the activation of the cell death protease caspase 9. Proc. Natl. Acad. Sci. USA 2001, 98, 14250–14255. [Google Scholar]
- Riedl, S.J.; Renatus, M.; Schwarzenbacher, R.; Zhou, Q.; Sun, C.; Fesik, S.W.; Liddington, R.C.; Salvesen, G.S. Structural basis for the inhibition of caspase-3 by xiap. Cell 2001, 104, 791–800. [Google Scholar] [CrossRef]
- Stennicke, H.R.; Jurgensmeier, J.M.; Shin, H.; Deveraux, Q.; Wolf, B.B.; Yang, X.; Zhou, Q.; Ellerby, H.M.; Ellerby, L.M.; Bredesen, D.; et al. Pro-caspase-3 is a major physiologic target of caspase-8. J. Biol. Chem. 1998, 273, 27084–27090. [Google Scholar] [CrossRef]
- Stennicke, H.R.; Deveraux, Q.L.; Humke, E.W.; Reed, J.C.; Dixit, V.M.; Salvesen, G.S. Caspase-9 can be activated without proteolytic processing. J. Biol. Chem. 1999, 274, 8359–8362. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Raychaudhuri, S.; Raychaudhuri, S.C. Monte Carlo Study Elucidates the Type 1/Type 2 Choice in Apoptotic Death Signaling in Healthy and Cancer Cells. Cells 2013, 2, 361-392. https://doi.org/10.3390/cells2020361
Raychaudhuri S, Raychaudhuri SC. Monte Carlo Study Elucidates the Type 1/Type 2 Choice in Apoptotic Death Signaling in Healthy and Cancer Cells. Cells. 2013; 2(2):361-392. https://doi.org/10.3390/cells2020361
Chicago/Turabian StyleRaychaudhuri, Subhadip, and Somkanya C Raychaudhuri. 2013. "Monte Carlo Study Elucidates the Type 1/Type 2 Choice in Apoptotic Death Signaling in Healthy and Cancer Cells" Cells 2, no. 2: 361-392. https://doi.org/10.3390/cells2020361
APA StyleRaychaudhuri, S., & Raychaudhuri, S. C. (2013). Monte Carlo Study Elucidates the Type 1/Type 2 Choice in Apoptotic Death Signaling in Healthy and Cancer Cells. Cells, 2(2), 361-392. https://doi.org/10.3390/cells2020361