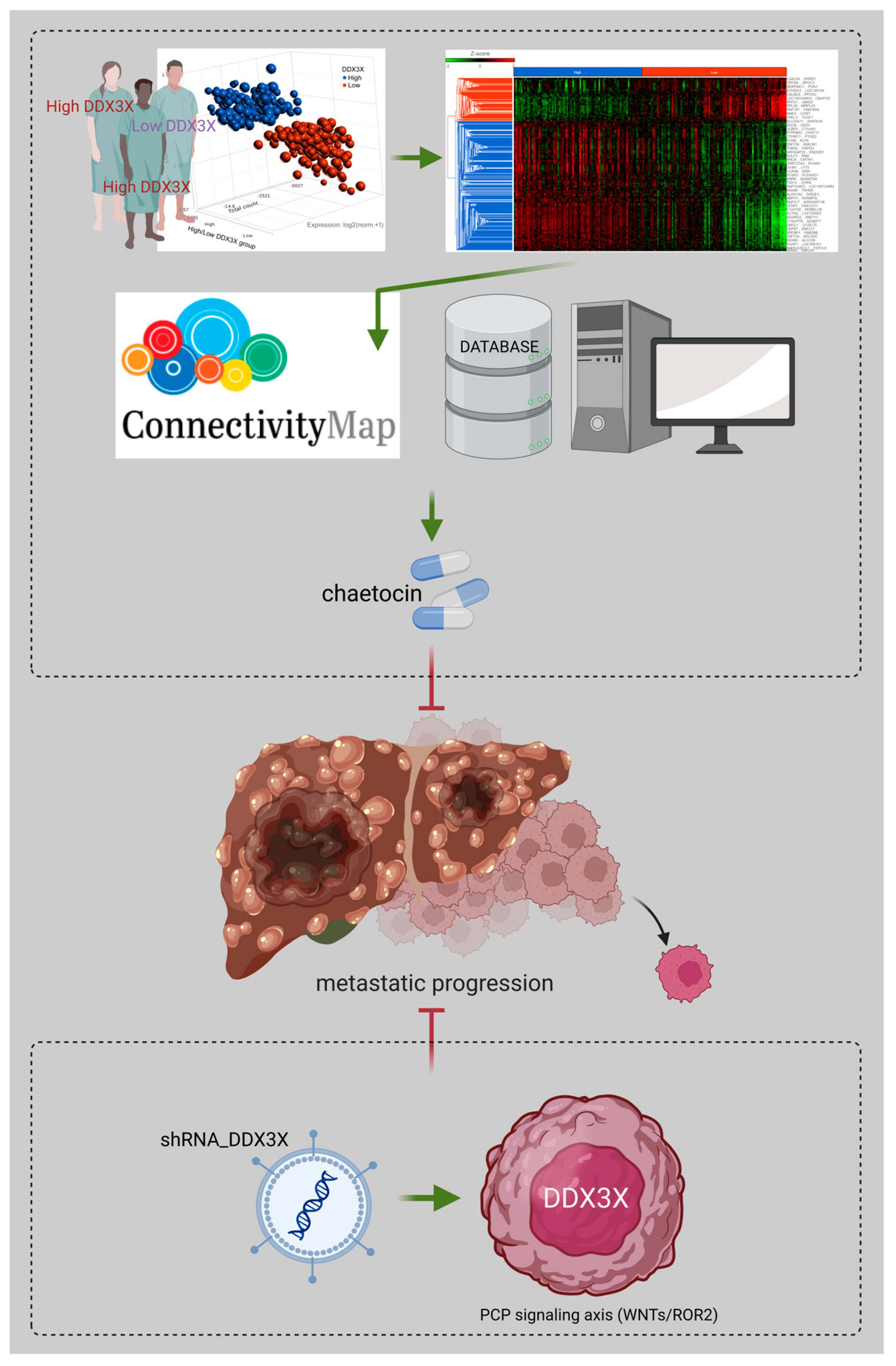
Perturbation Analysis of a Prognostic DDX3X-Mediated Gene Expression Signature Identifies the Antimetastatic Potential of Chaetocin in Hepatocellular Carcinoma
Abstract
:1. Introduction
2. Materials and Methods
2.1. TCGA Dataset Analysis
2.2. Ingenuity Pathway Analysis
2.3. Cell Culture
2.4. Lentivirus-Based shRNA Production and Infection
2.5. Cell Migration and Invasion Assay
2.6. Wound Healing Assay
2.7. Western Blot Analysis
2.8. Statistical Analysis
3. Results
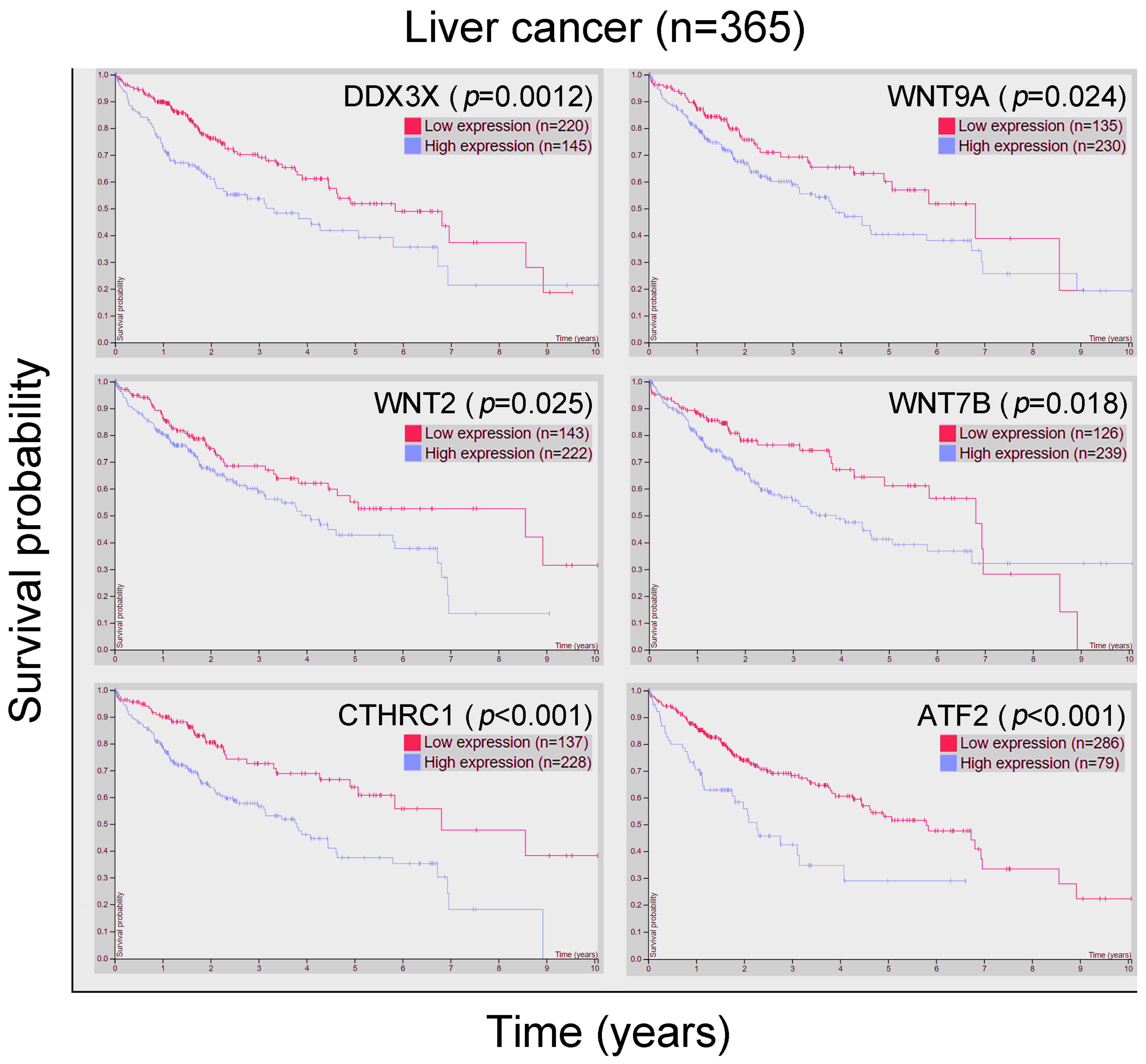
3.1. DDX3X Expression Is Significantly Correlated with Poor Outcomes in Liver Cancer, and the Significantly Differential Gene Signature Was Identified after Comparison between Patient Groups with High and Low DDX3X Levels
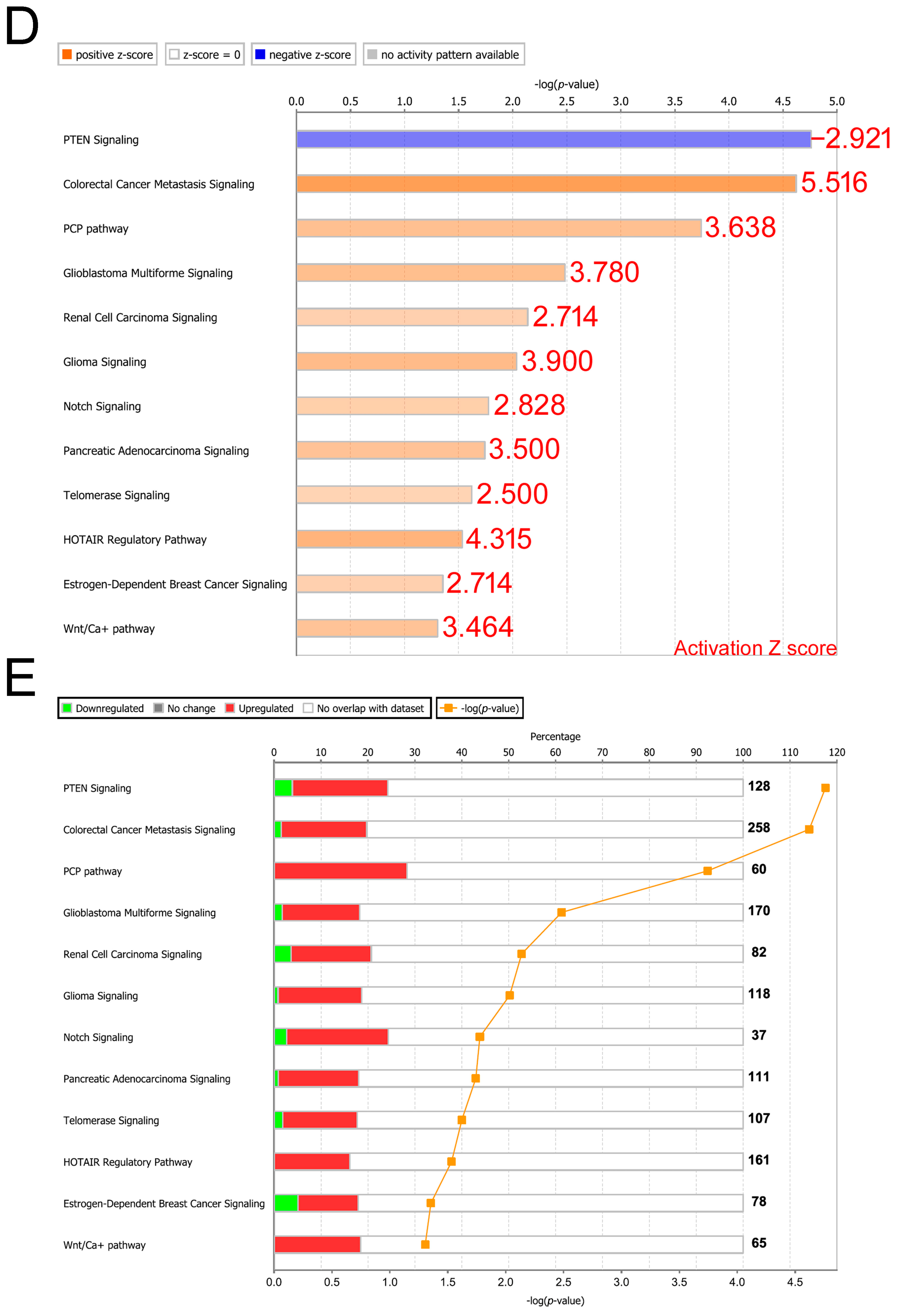
3.2. Knowledge-Based Analysis of Gene Signatures Reveals the Potential for Triggering Cancer Metastasis and Progression
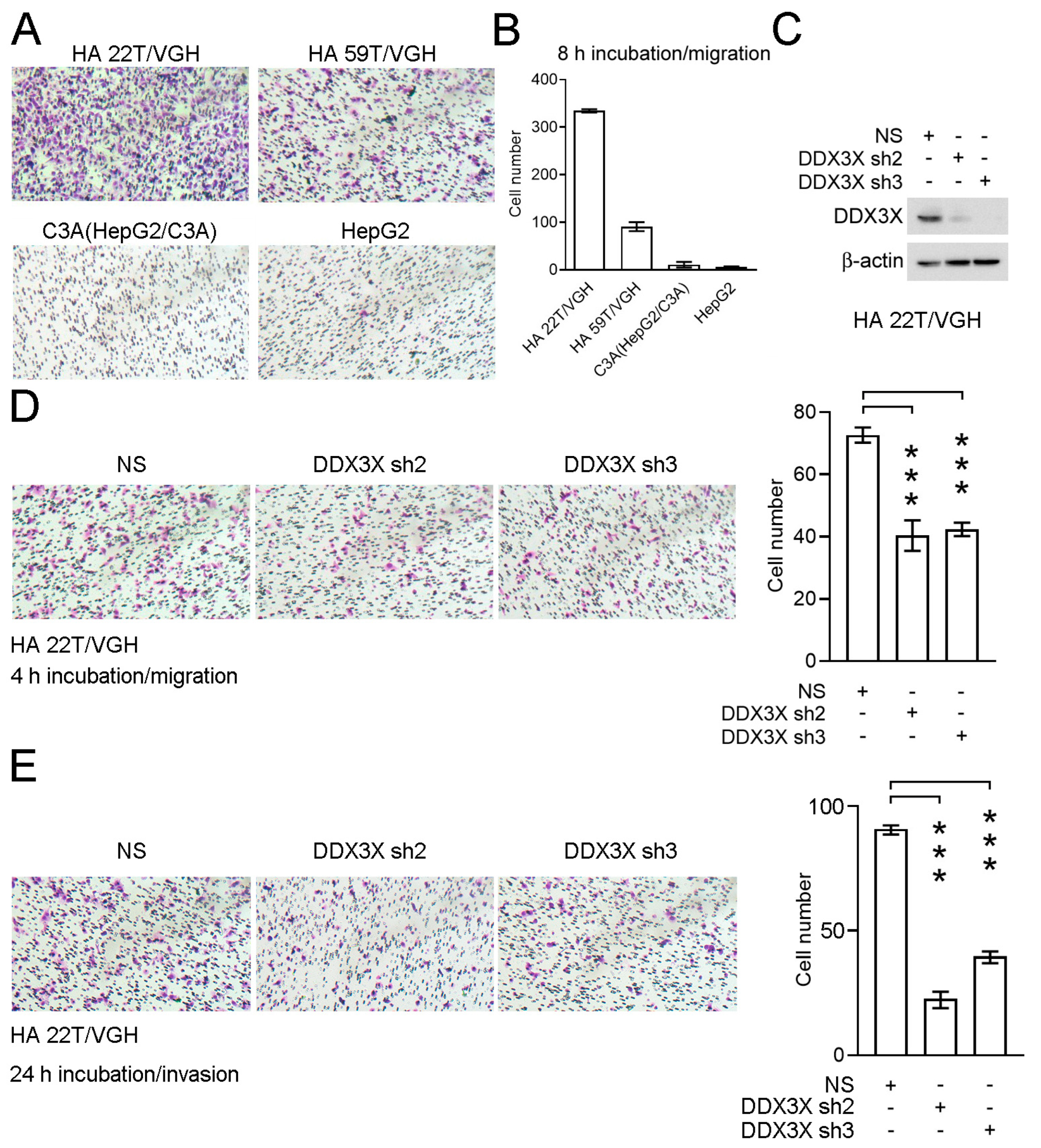
3.3. Knockdown of DDX3X Expression Inhibits Cell Migration and Invasion in a Liver Cancer Cell Line
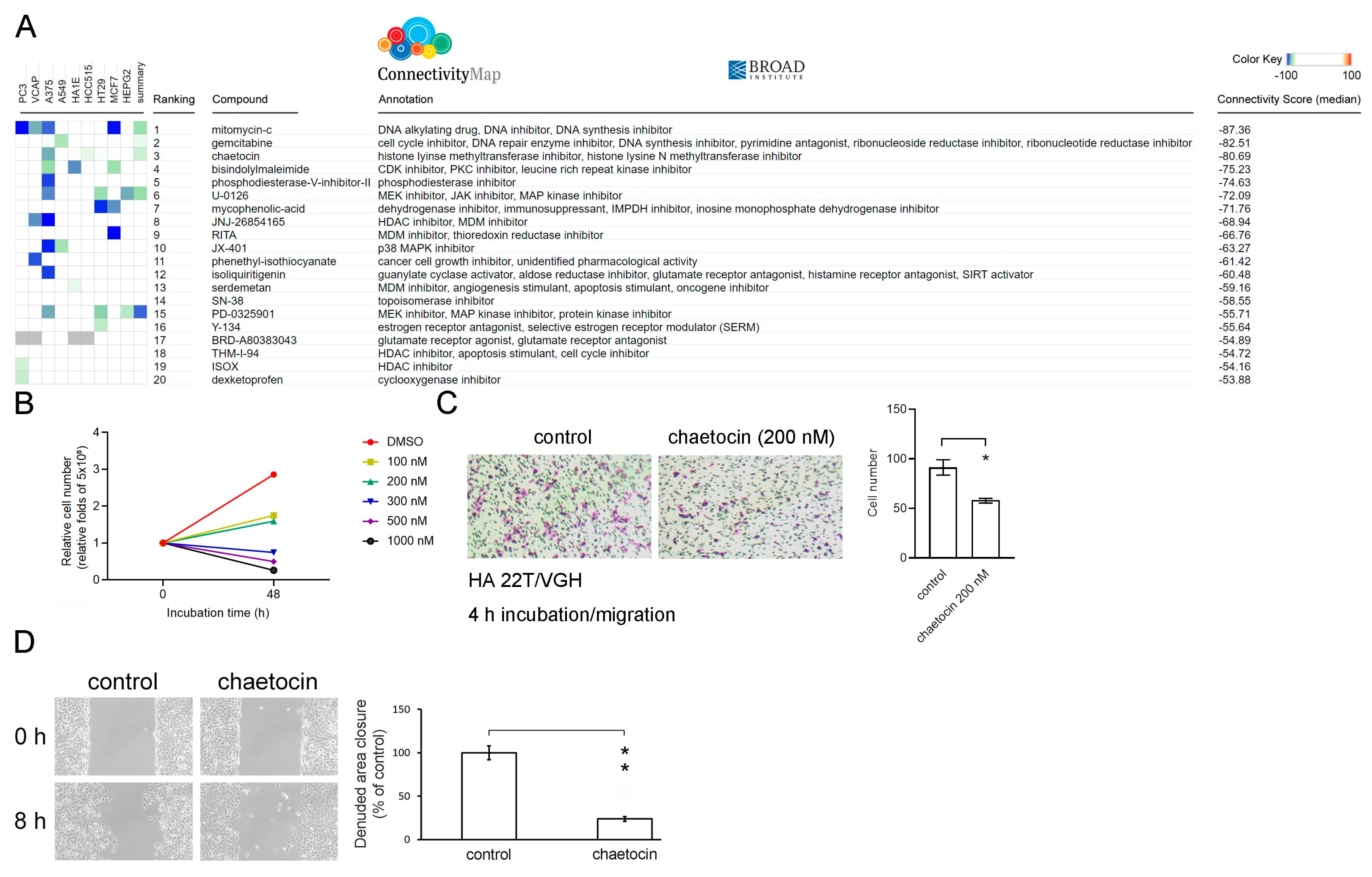
3.4. Chaetocin Reverses the High DDX3X Expression-Mediated Gene Signature and Suppresses Liver Cancer Cell Migration
4. Conclusions
5. Discussion
Supplementary Materials
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhou, Z.; Licklider, L.J.; Gygi, S.P.; Reed, R. Comprehensive proteomic analysis of the human spliceosome. Nature 2002, 419, 182–185. [Google Scholar] [CrossRef] [PubMed]
- Yedavalli, V.S.; Neuveut, C.; Chi, Y.H.; Kleiman, L.; Jeang, K.T. Requirement of DDX3 DEAD box RNA helicase for HIV-1 Rev-RRE export function. Cell 2004, 119, 381–392. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chao, C.H.; Chen, C.M.; Cheng, P.L.; Shih, J.W.; Tsou, A.P.; Lee, Y.H. DDX3, a DEAD box RNA helicase with tumor growth-suppressive property and transcriptional regulation activity of the p21waf1/cip1 promoter, is a candidate tumor suppressor. Cancer Res. 2006, 66, 6579–6588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chuang, R.Y.; Weaver, P.L.; Liu, Z.; Chang, T.H. Requirement of the DEAD-Box protein ded1p for messenger RNA translation. Science 1997, 275, 1468–1471. [Google Scholar] [CrossRef]
- Shih, J.W.; Tsai, T.Y.; Chao, C.H.; Wu Lee, Y.H. Candidate tumor suppressor DDX3 RNA helicase specifically represses cap-dependent translation by acting as an eIF4E inhibitory protein. Oncogene 2008, 27, 700–714. [Google Scholar] [CrossRef] [Green Version]
- Lee, C.S.; Dias, A.P.; Jedrychowski, M.; Patel, A.H.; Hsu, J.L.; Reed, R. Human DDX3 functions in translation and interacts with the translation initiation factor eIF3. Nucleic Acids Res. 2008, 36, 4708–4718. [Google Scholar] [CrossRef] [Green Version]
- Rocak, S.; Linder, P. DEAD-box proteins: The driving forces behind RNA metabolism. Nat. Rev. Mol. Cell Biol. 2004, 5, 232–241. [Google Scholar] [CrossRef]
- Fuller-Pace, F.V. DEAD box RNA helicase functions in cancer. RNA Biol. 2013, 10, 121–132. [Google Scholar] [CrossRef] [Green Version]
- Huang, J.S.; Chao, C.C.; Su, T.L.; Yeh, S.H.; Chen, D.S.; Chen, C.T.; Chen, P.J.; Jou, Y.S. Diverse cellular transformation capability of overexpressed genes in human hepatocellular carcinoma. Biochem. Biophys. Res. Commun. 2004, 315, 950–958. [Google Scholar] [CrossRef]
- Chan, C.H.; Chen, C.M.; Lee, Y.W.; You, L.R. DNA Damage, Liver Injury, and Tumorigenesis: Consequences of DDX3X Loss. Mol. Cancer Res. 2019, 17, 555–566. [Google Scholar] [CrossRef] [Green Version]
- Chang, P.C.; Chi, C.W.; Chau, G.Y.; Li, F.Y.; Tsai, Y.H.; Wu, J.C.; Wu Lee, Y.H. DDX3, a DEAD box RNA helicase, is deregulated in hepatitis virus-associated hepatocellular carcinoma and is involved in cell growth control. Oncogene 2006, 25, 1991–2003. [Google Scholar] [CrossRef] [Green Version]
- Bol, G.M.; Xie, M.; Raman, V. DDX3, a potential target for cancer treatment. Mol. Cancer 2015, 14, 188. [Google Scholar] [CrossRef] [Green Version]
- Subramanian, A.; Narayan, R.; Corsello, S.M.; Peck, D.D.; Natoli, T.E.; Lu, X.; Gould, J.; Davis, J.F.; Tubelli, A.A.; Asiedu, J.K.; et al. A Next Generation Connectivity Map: L1000 Platform and the First 1,000,000 Profiles. Cell 2017, 171, 1437–1452 e1417. [Google Scholar] [CrossRef]
- Uhlen, M.; Bjorling, E.; Agaton, C.; Szigyarto, C.A.; Amini, B.; Andersen, E.; Andersson, A.C.; Angelidou, P.; Asplund, A.; Asplund, C.; et al. A human protein atlas for normal and cancer tissues based on antibody proteomics. Mol. Cell. Proteom. 2005, 4, 1920–1932. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Uhlen, M.; Fagerberg, L.; Hallstrom, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, A.; Kampf, C.; Sjostedt, E.; Asplund, A.; et al. Proteomics. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef] [PubMed]
- Thul, P.J.; Akesson, L.; Wiking, M.; Mahdessian, D.; Geladaki, A.; Ait Blal, H.; Alm, T.; Asplund, A.; Bjork, L.; Breckels, L.M.; et al. A subcellular map of the human proteome. Science 2017, 356, eaal3321. [Google Scholar] [CrossRef]
- Uhlen, M.; Oksvold, P.; Fagerberg, L.; Lundberg, E.; Jonasson, K.; Forsberg, M.; Zwahlen, M.; Kampf, C.; Wester, K.; Hober, S.; et al. Towards a knowledge-based Human Protein Atlas. Nat. Biotechnol. 2010, 28, 1248–1250. [Google Scholar] [CrossRef]
- Uhlen, M.; Zhang, C.; Lee, S.; Sjostedt, E.; Fagerberg, L.; Bidkhori, G.; Benfeitas, R.; Arif, M.; Liu, Z.; Edfors, F.; et al. A pathology atlas of the human cancer transcriptome. Science 2017, 357, eaan2507. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gyorffy, B.; Schafer, R. Meta-analysis of gene expression profiles related to relapse-free survival in 1079 breast cancer patients. Breast Cancer Res. Treat. 2009, 118, 433–441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, M.; Zhou, Q.; Liu, X.; Wang, C.; Liu, G. CTHRC1 overexpression promotes cervical carcinoma progression by activating the Wnt/PCP signaling pathway. Oncol. Rep. 2019, 41, 1531–1538. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, X.M.; You, H.Y.; Li, Q.; Ma, H.; Wang, Y.H.; Zhang, Y.L.; Zhu, L.; Nie, H.Z.; Qin, W.X.; Zhang, Z.G.; et al. CTHRC1 promotes human colorectal cancer cell proliferation and invasiveness by activating Wnt/PCP signaling. Int. J. Clin. Exp. Pathol. 2015, 8, 12793–12801. [Google Scholar]
- Monch, D.; Koch, J.; Maass, A.; Janssen, N.; Murdter, T.; Renner, P.; Fallier-Becker, P.; Solass, W.; Schwab, M.; Dahlke, M.H.; et al. A human ex vivo coculture model to investigate peritoneal metastasis and innovative treatment options. Pleura. Peritoneum. 2021, 6, 121–129. [Google Scholar] [CrossRef]
- He, J.; Lin, X.; Meng, F.; Zhao, Y.; Wang, W.; Zhang, Y.; Chai, X.; Zhang, Y.; Yu, W.; Yang, J.; et al. A Novel Small Molecular Prostaglandin Receptor EP4 Antagonist, L001, Suppresses Pancreatic Cancer Metastasis. Molecules 2022, 27, 1209. [Google Scholar] [CrossRef]
- Jiang, H.; Li, Y.; Xiang, X.; Tang, Z.; Liu, K.; Su, Q.; Zhang, X.; Li, L. Chaetocin: A review of its anticancer potentials and mechanisms. Eur. J. Pharmacol. 2021, 910, 174459. [Google Scholar] [CrossRef]
- Yang, Z.; Wang, H.; Zhang, N.; Xing, T.; Zhang, W.; Wang, G.; Li, C.; Yu, C. Chaetocin Abrogates the Self-Renewal of Bladder Cancer Stem Cells via the Suppression of the KMT1A-GATA3-STAT3 Circuit. Front. Cell Dev. Biol. 2020, 8, 424. [Google Scholar] [CrossRef] [PubMed]
- Ozyerli-Goknar, E.; Sur-Erdem, I.; Seker, F.; Cingoz, A.; Kayabolen, A.; Kahya-Yesil, Z.; Uyulur, F.; Gezen, M.; Tolay, N.; Erman, B.; et al. The fungal metabolite chaetocin is a sensitizer for pro-apoptotic therapies in glioblastoma. Cell Death Dis. 2019, 10, 894. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kang, L.J.; Nguyen, K.V.A.; Eom, S.; Choi, Y.J.; Nguyen, C.N.; Lee, J.; Kim, C.; Lee, S.; Lee, S.G.; Lee, J.H. Stimulating DDX3 expression by serotonin 5-HT receptor 7 through phosphorylation of p53 via the AC-PKA-ERK signaling pathway. J. Cell Biochem. 2019, 120, 18193–18208. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Lei, A.; Xu, L.; Chen, L.; Chen, Y.; Zhang, X.; Gao, Y.; Yang, X.; Zhang, M.; Cao, Y. Similarity in gene-regulatory networks suggests that cancer cells share characteristics of embryonic neural cells. J. Biol. Chem. 2017, 292, 12842–12859. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, M.; Vesuna, F.; Tantravedi, S.; Bol, G.M.; Heerma van Voss, M.R.; Nugent, K.; Malek, R.; Gabrielson, K.; van Diest, P.J.; Tran, P.T.; et al. RK-33 Radiosensitizes Prostate Cancer Cells by Blocking the RNA Helicase DDX3. Cancer Res. 2016, 76, 6340–6350. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bol, G.M.; Vesuna, F.; Xie, M.; Zeng, J.; Aziz, K.; Gandhi, N.; Levine, A.; Irving, A.; Korz, D.; Tantravedi, S.; et al. Targeting DDX3 with a small molecule inhibitor for lung cancer therapy. EMBO Mol. Med. 2015, 7, 648–669. [Google Scholar] [CrossRef]
- Brai, A.; Ronzini, S.; Riva, V.; Botta, L.; Zamperini, C.; Borgini, M.; Trivisani, C.I.; Garbelli, A.; Pennisi, C.; Boccuto, A.; et al. Synthesis and Antiviral Activity of Novel 1,3,4-Thiadiazole Inhibitors of DDX3X. Molecules 2019, 24, 3988. [Google Scholar] [CrossRef] [Green Version]
- Radi, M.; Falchi, F.; Garbelli, A.; Samuele, A.; Bernardo, V.; Paolucci, S.; Baldanti, F.; Schenone, S.; Manetti, F.; Maga, G.; et al. Discovery of the first small molecule inhibitor of human DDX3 specifically designed to target the RNA binding site: Towards the next generation HIV-1 inhibitors. Bioorg. Med. Chem. Lett. 2012, 22, 2094–2098. [Google Scholar] [CrossRef]
- Botlagunta, M.; Kollapalli, B.; Kakarla, L.; Gajarla, S.P.; Gade, S.P.; Dadi, C.L.; Penumadu, A.; Javeed, S. In vitro anti-cancer activity of doxorubicin against human RNA helicase, DDX3. Bioinformation 2016, 12, 347–353. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Samal, S.K.; Routray, S.; Veeramachaneni, G.K.; Dash, R.; Botlagunta, M. Ketorolac salt is a newly discovered DDX3 inhibitor to treat oral cancer. Sci. Rep. 2015, 5, 9982. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tiemann, K.; Garri, C.; Lee, S.B.; Malihi, P.D.; Park, M.; Alvarez, R.M.; Yap, L.P.; Mallick, P.; Katz, J.E.; Gross, M.E.; et al. Loss of ER retention motif of AGR2 can impact mTORC signaling and promote cancer metastasis. Oncogene 2019, 38, 3003–3018. [Google Scholar] [CrossRef] [PubMed]
- Orecchia, A.; Mettouchi, A.; Uva, P.; Simon, G.C.; Arcelli, D.; Avitabile, S.; Ragone, G.; Meneguzzi, G.; Pfenninger, K.H.; Zambruno, G.; et al. Endothelial cell adhesion to soluble vascular endothelial growth factor receptor-1 triggers a cell dynamic and angiogenic phenotype. FASEB J. 2014, 28, 692–704. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.J.; Kang, L.J.; Lee, S.G. Stimulation of DDX3 expression by ginsenoside Rg3 through the Akt/p53 pathway activates the innate immune response via TBK1/IKKepsilon/IRF3 signalling. Curr. Med. Chem. 2014, 21, 1050–1060. [Google Scholar] [CrossRef]



| Symbol | Cancer Type | Prognosis | Endpoint | p Value | Case | Dataset | Method | Probe ID |
|---|---|---|---|---|---|---|---|---|
| DDX3X | Glioma | - | Overall survival | N.S. | 153 | TCGA | RNA Seq | |
| DDX3X | Thyroid Cancer | - | Overall survival | N.S. | 501 | TCGA | RNA Seq | |
| DDX3X | Lung Cancer | - | Overall survival | N.S. | 994 | TCGA | RNA Seq | |
| DDX3X | Colorectal Cancer | Good | Overall survival | 0.008 | 597 | TCGA | RNA Seq | |
| DDX3X | Head and Neck Cancer | - | Overall survival | N.S. | 499 | TCGA | RNA Seq | |
| DDX3X | Stomach Cancer | - | Overall survival | N.S. | 354 | TCGA | RNA Seq | |
| DDX3X | Liver Cancer | Poor | Overall survival | 0.0012 | 365 | TCGA | RNA Seq | |
| DDX3X | Pancreatic Cancer | Poor | Overall survival | 0.033 | 176 | TCGA | RNA Seq | |
| DDX3X | Renal Cancer | - | Overall survival | N.S. | 877 | TCGA | RNA Seq | |
| DDX3X | Urothelial Cancer | Good | Overall survival | 0.029 | 406 | TCGA | RNA Seq | |
| DDX3X | Prostate Cancer | - | Overall survival | N.S. | 494 | TCGA | RNA Seq | |
| DDX3X | Testis Cancer | - | Overall survival | N.S. | 134 | TCGA | RNA Seq | |
| DDX3X | Breast cancer | Poor | Overall survival | 0.024 | 1075 | TCGA | RNA Seq | |
| DDX3X | Cervical Cancer | - | Overall survival | N.S. | 291 | TCGA | RNA Seq | |
| DDX3X | Endometrial Cancer | - | Overall survival | N.S. | 541 | TCGA | RNA Seq | |
| DDX3X | Ovarian Cancer | - | Overall survival | N.S. | 373 | TCGA | RNA Seq | |
| DDX3X | Melanoma | - | Overall survival | N.S. | 102 | TCGA | RNA Seq | |
| DDX3X | Breast cancer | Poor | Relapse-free survival | <0.001 | 4929 | E-MTAB-365, E-TABM-43, GSE: 11121, 12093, | Array | 201210_at |
| 12276, 1456, 16391, 16446, 16716, 17705, 17907, | ||||||||
| 18728, 19615, 20194, 20271, 2034, 20685, 20711, | ||||||||
| 21653, 22093, 25066, 2603, 26971, 29044, 2990, | ||||||||
| 31448, 31519, 32646, 3494, 36771, 37946, 41998, | ||||||||
| 42568, 43358, 43365, 45255, 4611, 46184, 48390, | ||||||||
| 50948, 5327, 58812, 61304, 65194, 6532, 69031, | ||||||||
| 7390, 76275, 78958, 9195 | ||||||||
| DDX3X | Ovarian cancer | Poor | Progression-free survival | <0.001 | 1435 | GSE: 14764, 15622, 18520, 19829, 23554, 26193, | Array | 201210_at |
| 26712, 27651, 30161, 3149, 51373, 63885, 65986, | RNA Seq | |||||||
| 9891, TCGA (N = 565) | ||||||||
| DDX3X | Lung cancer | Good | Overall survival | <0.001 | 1925 | CAARRAY, GSE: 14814, 19188, 29013, 30219, | Array | 201210_at |
| 31210, 3141, 31908, 37745, 43580, 4573, 50081, | RNA Seq | |||||||
| 8894, TCGA (N = 133) | ||||||||
| DDX3X | Gastric cancer | Good | Post progression survival | <0.001 | 498 | GSE: 14210, 15459, 22377, 29272, 51105, 62254 | Array | 201210_at |
| Variable | Comparison | HR (95% CI) | p Value |
|---|---|---|---|
| Sex | M:F | 0.800 (0.562–1.141) | 0.218 |
| Stage | 3–4:1–2 | 2.485 (1.714–3.603) | <0.001 |
| T | T3–4:T1–2 | 2.578 (1.812–3.667) | <0.001 |
| N | N1:N0 | 2.012 (0.493–8.212) | 0.330 |
| M | M1:M0 | 4.055 (1.274–12.906) | 0.018 |
| DDX3X | High:Low | 1.485 (1.050–2.099) | 0.025 |
| Ranking | Categories | Diseases/Functions Annotation | Predicted Activation State | Activation z-Score |
|---|---|---|---|---|
| 1 | Cellular Movement | Cell movement | Increased | 9.208 |
| 2 | Cellular Movement | Migration of cells | Increased | 8.506 |
| 3 | Cellular Assembly and Organization, Cellular Function and Maintenance | Organization of cytoplasm | Increased | 8.472 |
| 4 | Cellular Assembly and Organization, Cellular Function and Maintenance | Organization of cytoskeleton | Increased | 8.43 |
| 5 | Cellular Movement | Cell movement of tumor cell lines | Increased | 8.254 |
| 6 | Tissue Morphology | Quantity of cells | Increased | 8.25 |
| 7 | Cell Morphology, Cellular Assembly and Organization, Cellular Function and Maintenance | Formation of cellular protrusions | Increased | 8.077 |
| 8 | Cellular Assembly and Organization, Cellular Function and Maintenance | Microtubule dynamics | Increased | 7.699 |
| 9 | Cellular Development, Cellular Growth and Proliferation, Nervous System Development and Function, Tissue Development | Development of neurons | Increased | 6.967 |
| 10 | Cellular Movement | Homing of cells | Increased | 6.705 |
| 11 | Cellular Function and Maintenance | Cellular homeostasis | Increased | 6.55 |
| 12 | Cellular Movement | Invasion of cells | Increased | 6.461 |
| 13 | Cardiovascular System Development and Function | Development of vasculature | Increased | 6.43 |
| 14 | Cardiovascular System Development and Function, Organismal Development | Angiogenesis | Increased | 6.418 |
| 15 | Cellular Movement | Chemotaxis | Increased | 6.411 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, T.-C. Perturbation Analysis of a Prognostic DDX3X-Mediated Gene Expression Signature Identifies the Antimetastatic Potential of Chaetocin in Hepatocellular Carcinoma. Cells 2023, 12, 1628. https://doi.org/10.3390/cells12121628
Lin T-C. Perturbation Analysis of a Prognostic DDX3X-Mediated Gene Expression Signature Identifies the Antimetastatic Potential of Chaetocin in Hepatocellular Carcinoma. Cells. 2023; 12(12):1628. https://doi.org/10.3390/cells12121628
Chicago/Turabian StyleLin, Tsung-Chieh. 2023. "Perturbation Analysis of a Prognostic DDX3X-Mediated Gene Expression Signature Identifies the Antimetastatic Potential of Chaetocin in Hepatocellular Carcinoma" Cells 12, no. 12: 1628. https://doi.org/10.3390/cells12121628
APA StyleLin, T.-C. (2023). Perturbation Analysis of a Prognostic DDX3X-Mediated Gene Expression Signature Identifies the Antimetastatic Potential of Chaetocin in Hepatocellular Carcinoma. Cells, 12(12), 1628. https://doi.org/10.3390/cells12121628






