A Comprehensive Review of miRNAs and Their Epigenetic Effects in Glioblastoma
Abstract
1. Introduction
The CNS5 Classification and Glioblastoma
2. DNA Methylation
2.1. DNA Methylation in GBM
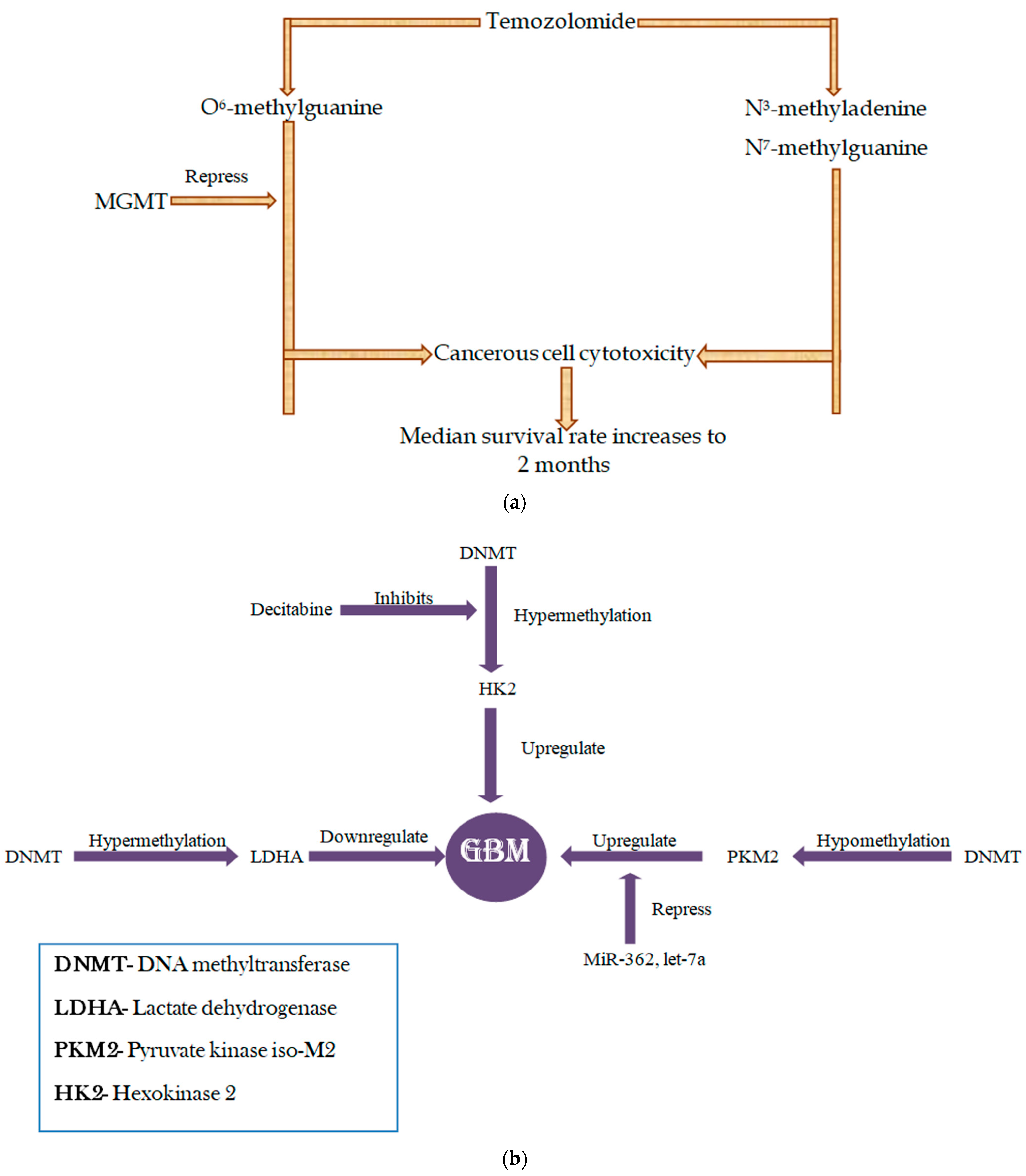
2.2. DNA Methylation: Its Role in Regulation of Metabolism in GBM
2.3. PKM2
2.4. LDHA
2.5. HK2
3. miRNA
3.1. miRNA Biogenesis
3.2. miRNA in GBM
3.3. Upregulated miRNA
miR-21
3.4. miR-10b and miR-10a
3.5. miR-10b-miR-21
3.5.1. miR-10b and miR-222
3.5.2. miR-9
3.5.3. miR-221/222
3.5.4. miR-26a
3.5.5. miR-17-92 Cluster
3.5.6. miR-148a
3.5.7. Other Upregulated miRNA in GBM
3.6. Downregulated miRNAs
3.6.1. miR-31
3.6.2. miR-124
3.6.3. miR-34
3.6.4. miR-302-367 Cluster
3.6.5. miR-181
3.6.6. miR-219-5p and miR-219-1-3p
3.6.7. miR-1
3.6.8. miR-370-3p
3.6.9. miR-328
3.6.10. miR-375
3.6.11. miR-137
3.6.12. miR-128
3.6.13. miR-7
| miRNA | Target | Expression | Function | Reference |
|---|---|---|---|---|
| miR-21 | PTEN, p53, VH1, PPARa, TIMP3, RECK, SPOCK1, RB1CC1 | Up | Tumor Growth (+), regulate EGFR/AKT signaling, Cell invasion (+), Cell proliferation (+), Apoptosis (−) | [7,24,29,30] |
| 10b | p-53, CDKN1A, CDKN2A, BIM, BCL2, TEAP2C, HOXD1O, uPAR, R4OC | Up | Promotes cell cycle, Cell invasion (+) | [14,18,31] |
| miR-10b/222 | p53/PTEN, BIM | Up | Apoptosis (+), Cell Proliferation (+) | [18,34] |
| miR-9 | NF1, PTCH1P | Up | Cell proliferation (+), Cell migration (+), Inflammation (+), Resistance to chemotherapy (+), Apoptosis (−) | [14,35,36] |
| miR-221/222 | PTEN, MMP2, MMP3, BEGF, PUMA, E2F3, TIMP3, P27KiP1 | Up | Tumor growth (+), Apoptosis (−), Proliferation (+), Angiogenesis (+), Migration (+), Invasion (+) | [7,12,18,30,37] |
| miR-26a | PTEN | Up | Tumor growth (+) | [7,14,29] |
| miR- 148a | CADM1 | Up | Cell proliferation (+), Metastasis (+) | [18,40] |
| miR-125 | BMF | Up | Apoptosis (−) | [29] |
| 182 | USPI5, TNIP1, CMTLD | Up | GBM aggressiveness (+), Disrupt negative feedback loop of NF-KB | [29] |
| miR-196 | Up | Cell proliferation (+), Poor survival | [29] | |
| miR-30 | SOCS3, JAK/STAT3, TRAIL Protein | Up | GSC differentiation (+), Apoptosis (−) | [41] |
| miR-143 | HKII | Up | Cell differentiation (+) | [14] |
| miR-145 | LKB 39- AMPK pathway | Up | Tumor growth (+) | [14] |
| miR-495-3p | PTEN/AKT pathway | Up | Migration (+), Proliferation (+), Invasion (+) | [13] |
| miR-503 | PACDA | Up | Apoptosis (+) | [18] |
| miR-93 | Up | Angiogenesis (+), Tumor growth (+) | [13] | |
| miR-378 | VEGFR2 | Up | Angiogenesis (+), Tumor growth (+) | [18,20] |
| miR-201 | HIF1, HIF2 | Up | Apoptosis (−), Cell proliferation (+) | [18,42] |
| miRNA | Target | Expression | Function | Reference |
|---|---|---|---|---|
| miR-31 | Radixin | Down | Invasion (−), Migration (−) | [29] |
| miR-124 | SNA12, CDKA, CDK6, Cyclin D | Down | Cell cycle arrest (+), GSCs invasiveness (−) | [29,43] |
| miR-34a | Notch 1, Notch 2, CDK6, EGFR, C-met, BCI-2 | Down | Cell proliferation (−), Invasion (−), GSCs differentiation (−), Cell cycle arrest (+) | [13,44,45,46] |
| miR-34c-3p | Notch 2 | Down | S-phase arrest (+), Proliferation (−), Apoptosis (+) | [44] |
| miR-34c-5p | Notch 1, Notch 2, CDK6, EGFR | Down | Cell proliferation (−) | [44] |
| miR-302-367 cluster | GIC, CXRC4, PI3K/AKT pathway, STAT3 pathway, SALL2, OLIG2, SOX2, CMyC, KLF4, OCT3/4, UCH1, MYBBP1A, PEAL5 | Down | Tumor growth (−), GSC stemness (−) | [14,47] |
| miR-181 (a) miR-181a (c) miR-181d | CDI33, BMI1, WNT signaling pathway, CCR1, IL-1b, BCI-2, K-Ra5 | Down Down | GSC stemness (−) Tumor growth (+) | [21] [48,49] |
| (b) miR-181c | TGFBR1 TGFBR2, TGFBRAP1 | Down | Cell invasion (−), Proliferation (−) | [56] |
| miR-219-5p miR-219-1-3p | Down | Tumor growth (−), Proliferation (−) | [13] | |
| miR-1 | Down | Sensitize GBM to TMZ, Apoptosis (+) | [50] | |
| miR-328 | Down | Proliferation (−) | [52] | |
| miR-375 | Down | Proliferation (−), Invasion (−), Migration (−) | [53] | |
| miR-137 | EZH2 | Down | Angiogenesis (−), Proliferation (−) | [18,20] |
| miR-128 | WNT, BRK, EGFR, IGF1R, BCL2, 5UZI2, BIM1, EZF3, PDGFRA | Down | Apoptosis (+), Proliferation (−), Metastasis (−), Angiogenesis (−), GSCs Renewability (−) | [13,24,54,55] |
| miR-7 | PKM2, EGFR, AKT/PI3K pathway | Down | Tumor Growth (−) | [13,14] |
4. miRNA and DNA Methylation: An Epigenetic Interplay in GBM
5. miRNA and Epigenetic Modifications in TMZ Response and Drug Resistance
5.1. Epigenetic Modulation and TMZ Response
5.2. miRNA and TMZ Response
- miR-21 in an oncogenic miRNA that contributes to drug resistance. Its downregulation enhances chemotherapy efficacy against human GBM cells [63]. miR-21 is often considered a potential biomarker for TMZ resistance [18]. Therefore, silencing miR-21 with simultaneous TMZ treatment can markedly enhance the apoptosis of cancer cells and, therefore, increase the median survival time of patients with TMZ-resistant GBM [18].
- miR-181d has also been identified as a predictor of TMZ response and patient survival [49]. It was experimentally proved that transfecting miR-181d into GBM cells caused MGMT expression decay, which is associated with good prognosis and overcoming of resistance. So, miR-181d positively associates with TMZ response and patient survival [18,63]. Another miRNA of the same family, miR-181c, is involved in TMZ resistance, as it is suppressed in a patient with GBM who showed a positive response to radiotherapy/TMZ treatment [64].
- miR-195 and miR-10a are reported to be overexpressed in GBM cells having low sensitivity to TMZ; therefore, downregulation of these miRNAs can significantly improve TMZ response and survival chance [55].
- miR-124, miR-134, and miR-128 induce their antitumor activity synergistically by inhibiting GSC proliferation and promoting an effective response of radiotherapy and chemotherapy against GBM [63].
- miR-370-3p, a negative regulator of MGMT, has been reported to be highly downregulated in TMZ-resistant GBM cells. miR-370-3p suppresses MGMT expression in GBM cells and sensitive glioma cells to TMZ [51,65], inducing apoptosis of tumor cells [51,65]. Thus, miR-370-3p can have a potential therapeutic role in the treatment of recurring GBM if used to improve TMZ response [65].
- miR-128 and miR-149 overexpression sensitize glioma cells to TMZ, especially in the case of non-stem GBM cells and, therefore, contribute to better prognosis [54].
- miR-125b overexpression confers chemoresistance of GSCs to TMZ treatment. The combined inhibition of PI3K and miR-125b significantly enhances TMZ-induced inhibition of GSC proliferation and invasiveness [18]. miR-100 overexpression in glioma cells sensitized them to ionizing radiation by downregulating the ataxia telangiectasia mutated (ATM) gene [29].
- miR-328 sensitizes GSC to TMZ by directly suppressing ABCG2 expression [29]. miR-218 and miR-1268a are associated with enhanced TMZ response in GBM patients [18,29]. miR-1268a is downregulated in a patient with recurrent GBM, and its overexpression promotes TMZ sensitivity to GBM cells via inhibition of translation of the ABCCL gene [18].
- miR-299-5p enhances TMZ sensitivity to GBM cells by inhibiting cell proliferation via regulation of the ERK signaling pathway [18].
- Overexpression of miR-423-5p and miR-223 promotes GBM cell survival by decreasing TMZ response [18]. miR-223 expression suppresses TMZ, inducing the inhibition of cell proliferation as well as the miR-223/PAX6 axis that further contributes to chemoresistance and decreases in TMZ response by regulating the PI3K/AKT signaling pathway.
- miR-318, miR-381, and miR-20a overexpression also result in increased TMZ resistance [18]. Apart from the aforementioned miRNAs, new miRNAs are continuously being discovered that can be used as potential therapeutic tools in combination with established chemotherapy and radiation therapy.
| miRNA/Epigenetic Modulator | Expression | Effect on TMZ Response | Reference |
|---|---|---|---|
| MGMT | High | Induces TMZ resistance | [62] |
| miR-21 | Up | Induces TMZ resistance | [18] |
| miR-181d | Up | Sensitizes glioma cells to TMZ | [18,63] |
| miR-195 and miR-10a | Up | Confers TMZ resistance to glioma cells | [55] |
| miR-124, miR-134, and miR-128 | Up | Promotes TMZ-induced cytotoxicity of glioma cells | [63] |
| miR-370-3p | Down | TMZ resistance | [51,65] |
| miR-125b | Up | Confers chemoresistance to GSCs against TMZ | [18] |
| miR-128 and miR-149 | Up | Sensitizes non-stem glioma cells to TMZ | [54] |
| miR-328 | Up | Enhances TMZ response to GBM by targeting ABCG2 | [29] |
| miR-1268a and miR-218 | Up | Inhibits translation of ABCCL and enhances TMZ sensitivity | [18] |
| miR-299-5p | Up | Inhibits cell proliferation and enhances TMZ sensitivity by regulating ERK signaling pathway | [18] |
| miR-423-5p and miR-223 | Up | Decreases TMZ response and promotes GBM response | [18] |
| miR-318, miR-381, and miR-209 | Up | Increases TMZ resistance | [18] |
6. Diagnostic and Prognostic Molecular Tools in GBM: Do miRNAs Play a Role?
- miR-21 is a potential biomarker of GBM with 90% sensitivity and 100% specificity [63]. It has been observed to have low expression in the post-operation serum of GBM patients, suggesting its potential as a serum-derived miRNA biomarker in GBM [38]. High levels of miR-21 have been reported in the plasma of GBM patients, and these levels get lower once the tumor is removed [21]. miR-21 might be used to discriminate between different WHO grades as well as to predict overall survival time in GBM patients [63].
- miR-26a and miR-21 are both circulatory miRNAs that are upregulated in GBM, and their serum expression levels have been observed to be reduced after surgery [38], suggesting their importance as candidate serum-based biomarkers in the diagnosis of GBM as well as in monitoring disease progression [38]. Additionally, reduced post-operative serum level of miR-26a also indicates the humoral origin of miR-26a.
- miR-10b is upregulated in GBM, and its overexpression promotes GBM progression and correlates with poor prognosis [63]. Its expression level positively correlates with WHO grades of gliomas as well as with tumor invasiveness [21]. Therefore, miR-10b might be used as a biomarker to evaluate glioma invasiveness and, subsequently, in the sub-classification of different tumor grades. Additionally, the combined assessment of miR-10b and miR-21 in the serum of GBM patients can aid in predicting the therapeutic effect of bevacizumab (BVZ) because miR-10b and 21 serum levels have been reported to be very high in GBM patients and associated with increased tumor diameter in BVZ treated patients.
- miR-328 is downregulated in GBM and acts as a tumor suppressor. The low expression level of miR-328 correlates with poor survival rate, thus it might be used as a candidate prognostic biomarker in GBM [52].
- High plasma levels of miR-21 and low plasma levels of miR-128 and miR-342-3p act as candidate biomarkers in distinguishing GBM patients from healthy individuals with remarkably high sensitivity and specificity [67]. miR-342-3p expression is reduced in the plasma of glioma patients, and it is increased after surgery or chemotherapy. Therefore, miR-342-3p might be a candidate biomarker for the diagnosis and discrimination of glioma [67].
- miR-320a is a tumor suppressor miRNA, and its suppression correlates with excessive cell proliferation, invasion, and tumor growth [31]. Therefore, it might be used as a prognostic biomarker [31]. miR-146b and miR-4492 can be useful as novel biomarkers in predicting and monitoring GBM progression [31]. miR-146b is an oncogenic miRNA, and its major target is TRAF6. Downregulation of miR-146b and upregulation of TRAF6 correlate with inhibition of cell proliferation as well as apoptosis of tumor cells due to a decrease in Ki-67 expression. Hence, miR-146b might be suggested as a candidate biomarker for understanding GBM prognosis as well as in discriminating different grades of glioma [31].
- miR-29 plasma level serves as a potential biomarker to indicate malignancy and glioma progression from grades I-II to grades III-IV [68]. miR-454-3p serum expression levels have been found markedly increased in GBM patients, and its upregulation correlates with poor prognosis. Therefore, it can be used as a candidate prognostic biomarker [68].
- Sometimes, single miRNA profiling is not sufficient enough to predict glioma outcomes. In such cases, profiles of several miRNAs are suggested. Seven miRNAs, including miR-15b, miR-23a, miR-133a, miR-150, miR-197, miR-497, and miR-548b-5p, are all downregulated in grades II-IV glioma patients, and the combined expression profiling of these miRNAs might be taken as a candidate biomarker in the prediction of GBM malignancy [68].
- miR-181 is widely reported to be downregulated in GBM, especially in the early stages of this tumor [68]. Therefore, miR-181 might be used as a candidate biomarker for early prediction as well as in the identification of tumor grade. miR-181b and miR-181c act as predictive biomarkers of TMZ response in GBM [68] and may also help in choosing patients who are suitable for adjuvant therapy [30].
- miR-221/222 is found to be significantly upregulated in plasma samples of glioma patients [30,69], and its overexpression contributes to poor prognosis and low survival rates [69]. The study conducted by Zhang R et al. has confirmed that miR-221 and miR-222 might be used as potential diagnostic and prognostic biomarkers [69].
| Over-Expressed | Under-Expressed | Source Reference |
|---|---|---|
| miR-21 | Serum [38] Plasma [63] | |
| miR-26a and miR-21 | Serum [38] | |
| miR-21 | miR-128 and miR-342-3p | Plasma [67] |
| miR-10b and miR-21 | Serum [56] | |
| miR-320a | ||
| miR-146b | [31] | |
| miR-454-3p | Serum [67] | |
| miR-29 | Plasma [67] | |
| miR-23a, miR-133a, miR-150, miR-197, miR-497, and miR-548b-5p | [67] | |
| miR-221/222 | Plasma [30,69] | |
| miR-181 | [68] |
7. GBM Therapy
7.1. miRNA Based Glioma Therapy
7.1.1. miRNA-Based Replacement Therapy
miR-34a
miRNA-7
7.1.2. Oligonucleotide Therapy
7.2. Epigenetic Therapy
7.2.1. DNMT Inhibitors
7.2.2. Histone Deacetylase Inhibitors (HDACIs)
7.3. Molecular Target Therapy
7.4. Adjuvant Therapy
8. Challenges and Limitations
9. Perspective and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Louis, D.N.; Perry, A.; Wesseling, P.; Brat, D.J.; Cree, I.A.; Figarella-Branger, D.; Hawkins, C.; Ng, H.K.; Pfister, S.M.; Reifenberger, G.; et al. The 2021 WHO Classification of Tumors of the Central Nervous System: A summary. Neuro Oncol. 2021, 23, 1231–1251. [Google Scholar] [CrossRef] [PubMed]
- Sejda, A.; Grajkowska, W.; Trubicka, J.; Szutowicz, E.; Wojdacz, T.; Kloc, W.; Iżycka-Świeszewska, E. WHO CNS5 2021 classification of gliomas: A practical review and road signs for diagnosing pathologists and proper patho-clinical and neuro-oncological cooperation. Folia Neuropathol. 2022, 60, 137–152. [Google Scholar] [CrossRef]
- Wilson, T.A.; Karajannis, M.A.; Harter, D.H. Glioblastoma multiforme: State of the art and future therapeutics. Surg. Neurol. Int. 2014, 5, 64. [Google Scholar] [CrossRef] [PubMed]
- Lombardi, M.Y.; Assem, M. Glioblastoma Genomics: A Very Complicated Story. In Glioblastoma; De Vleeschouwer, S., Ed.; Codon Publications: Brisbane, Australia, 2017. [Google Scholar] [CrossRef]
- Mansouri, A.; Karamchandani, J.; Das, S. Molecular Genetics of Secondary Glioblastoma. In Glioblastoma; De Vleeschouwer, S., Ed.; Codon Publications: Brisbane, Australia, 2017. [Google Scholar] [CrossRef]
- Burgess, R.; Jenkins, R.; Zhang, Z. Epigenetic changes in gliomas. Cancer Biol. 2008, 7, 1326–1334. [Google Scholar] [CrossRef]
- Kreth, S.; Thon, N.; Kreth, F.W. Epigenetics in human gliomas. Cancer Lett. 2014, 342, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Ghasemi, A.; Mohammadi, A.; Fallah, S. Epigenetic Modification of MicroRNA-219-1 and Its Association with Glioblastoma Multiforme. Biochemistry 2021, 86, 420–432. [Google Scholar] [CrossRef] [PubMed]
- Ayala-Ortega, E.; Arzate-Mejia, R.; Perez-Molina, R.; Gonzalez-Buendia, E.; Meier, K.; Guerrero, G.; Recillas-Targa, F. Epigenetic silencing of miR-181c by DNA methylation in glioblastoma cell lines. BMC Cancer 2016, 16, 226. [Google Scholar] [CrossRef]
- Marzese, D.M.; Hoon, D.S. Emerging technologies for studying DNA methylation for the molecular diagnosis of cancer. Expert Rev. Mol. Diagn. 2015, 15, 647–664. [Google Scholar] [CrossRef]
- Quintavalle, C.; Mangani, D.; Roscigno, G.; Romano, G.; Diaz-Lagares, A.; Iaboni, M.; Donnarumma, E.; Fiore, D.; De Marinis, P.; Soini, Y.; et al. MiR-221/222 target the DNA methyltransferase MGMT in glioma cells. PLoS ONE 2013, 8, e74466. [Google Scholar] [CrossRef]
- Kirstein, A.; Schmid, T.E.; Combs, S.E. The Role of miRNA for the Treatment of MGMT Unmethylated Glioblastoma Multiforme. Cancers 2020, 12, 1099. [Google Scholar] [CrossRef]
- Uddin, M.S.; Mamun, A.A.; Alghamdi, B.S.; Tewari, D.; Jeandet, P.; Sarwar, M.S.; Ashraf, G.M. Epigenetics of glioblastoma multiforme: From molecular mechanisms to therapeutic approaches. Semin. Cancer Biol. 2022, 83, 100–120. [Google Scholar] [CrossRef]
- Dong, Z.; Cui, H. Epigenetic modulation of metabolism in glioblastoma. Semin. Cancer Biol. 2019, 57, 45–51. [Google Scholar] [CrossRef]
- Chesnelong, C.; Chaumeil, M.M.; Blough, M.D.; Al-Najjar, M.; Stechishin, O.D.; Chan, J.A.; Pieper, R.O.; Ronen, S.M.; Weiss, S.; Luchman, H.A.; et al. Lactate dehydrogenase A silencing in IDH mutant gliomas. Neuro Oncol. 2014, 16, 686–695. [Google Scholar] [CrossRef]
- Di, H.; Zhang, X.; Guo, Y.; Shi, Y.; Fang, C.; Yuan, Y.; Wang, J.; Shang, C.; Guo, W.; Li, C. Silencing LDHA inhibits proliferation, induces apoptosis and increases chemosensitivity to temozolomide in glioma cells. Oncol. Lett. 2018, 15, 5131–5136. [Google Scholar] [CrossRef] [PubMed]
- Blakeway, D.; Karakoula, K.; Morris, M.; Rowther, F.; Eagles, L.; Darling, J.; Warr, T. Overexpression of Hexokinase 2 is epigenetically regulated by frequent hypomethylation in glioblastoma multiforme. Neuro Oncol. 2018, 20, i12. [Google Scholar] [CrossRef]
- Rezaei, O.; Honarmand, K.; Nateghinia, S.; Taheri, M.; Ghafouri-Fard, S. miRNA signature in glioblastoma: Potential biomarkers and therapeutic targets. Exp. Mol. Pathol. 2020, 117, 104550. [Google Scholar] [CrossRef] [PubMed]
- Henriksen, M.; Johnsen, K.B.; Andersen, H.H.; Pilgaard, L.; Duroux, M. MicroRNA expression signatures determine prognosis and survival in glioblastoma multiforme—A systematic overview. Mol. Neurobiol. 2014, 50, 896–913. [Google Scholar] [CrossRef]
- Balandeh, E.; Mohammadshafie, K.; Mahmoudi, Y.; Hossein Pourhanifeh, M.; Rajabi, A.; Bahabadi, Z.R.; Mohammadi, A.H.; Rahimian, N.; Hamblin, M.R.; Mirzaei, H. Roles of Non-coding RNAs and Angiogenesis in Glioblastoma. Front. Cell Dev. Biol. 2021, 9, 716462. [Google Scholar] [CrossRef]
- Kalkan, R.; Atli, E.I. The Impacts of miRNAs in Glioblastoma Progression. Crit. Rev. Eukaryot. Gene Expr. 2016, 26, 137–142. [Google Scholar] [CrossRef]
- Huang, D.; Qiu, S.; Ge, R.; He, L.; Li, M.; Li, Y.; Peng, Y. miR-340 suppresses glioblastoma multiforme. Oncotarget 2015, 6, 9257–9270. [Google Scholar] [CrossRef]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef]
- Marumoto, T.; Saya, H. Molecular biology of glioma. Adv. Exp. Med. Biol. 2012, 746, 2–11. [Google Scholar] [CrossRef] [PubMed]
- Michlewski, G.; Caceres, J.F. Post-transcriptional control of miRNA biogenesis. RNA 2019, 25, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; He, J.; Pu, W.; Peng, Y. The Role of Exportin-5 in MicroRNA Biogenesis and Cancer. Genom. Proteom. Bioinform. 2018, 16, 120–126. [Google Scholar] [CrossRef]
- Bovell, L.C.; Putcha, B.D.; Samuel, T.; Manne, U. Clinical implications of microRNAs in cancer. Biotech. Histochem. 2013, 88, 388–396. [Google Scholar] [CrossRef] [PubMed]
- Alfardus, H.; McIntyre, A.; Smith, S. MicroRNA Regulation of Glycolytic Metabolism in Glioblastoma. Biomed. Res. Int. 2017, 2017, 9157370. [Google Scholar] [CrossRef]
- Hassan, A.; Mosley, J.; Singh, S.; Zinn, P.O. A Comprehensive Review of Genomics and Noncoding RNA in Gliomas. Top. Magn. Reson. Imaging 2017, 26, 3–14. [Google Scholar] [CrossRef]
- Barciszewska, A.M. MicroRNAs as efficient biomarkers in high-grade gliomas. Folia Neuropathol. 2016, 54, 369–374. [Google Scholar] [CrossRef]
- de Menezes, M.R.; Acioli, M.E.A.; da Trindade, A.C.L.; da Silva, S.P.; de Lima, R.E.; da Silva Teixeira, V.G.; Vasconcelos, L.R.S. Potential role of microRNAs as biomarkers in human glioblastoma: A mini systematic review from 2015 to 2020. Mol. Biol. Rep. 2021, 48, 4647–4658. [Google Scholar] [CrossRef]
- Junior, L.G.D.; Baroni, M.; Lira, R.C.P.; Teixeira, S.; Fedatto, P.F.; Silveira, V.S.; Suazo, V.K.; Veronez, L.C.; Panepucci, R.A.; Antonio, D.S.M.; et al. High-throughput microRNA profile in adult and pediatric primary glioblastomas: The role of miR-10b-5p and miR-630 in the tumor aggressiveness. Mol. Biol. Rep. 2020, 47, 6949–6959. [Google Scholar] [CrossRef]
- Malhotra, M.; Sekar, T.V.; Ananta, J.S.; Devulapally, R.; Afjei, R.; Babikir, H.A.; Paulmurugan, R.; Massoud, T.F. Targeted nanoparticle delivery of therapeutic antisense microRNAs presensitizes glioblastoma cells to lower effective doses of temozolomide in vitro and in a mouse model. Oncotarget 2018, 9, 21478–21494. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sun, B.; Zhao, X.; Ming, J.; Liu, X.; Liu, D.; Jiang, C. Stepwise detection and evaluation reveal miR-10b and miR-222 as a remarkable prognostic pair for glioblastoma. Oncogene 2019, 38, 6142–6157. [Google Scholar] [CrossRef] [PubMed]
- Coolen, M.; Katz, S.; Bally-Cuif, L. miR-9: A versatile regulator of neurogenesis. Front. Cell. Neurosci. 2013, 7, 220. [Google Scholar] [CrossRef]
- Munoz, J.L.; Rodriguez-Cruz, V.; Rameshwar, P. High expression of miR-9 in CD133(+) glioblastoma cells in chemoresistance to temozolomide. J. Cancer Stem Cell Res. 2015, 3, e1003. [Google Scholar] [CrossRef]
- Krichevsky, A.M.; Uhlmann, E.J. Oligonucleotide Therapeutics as a New Class of Drugs for Malignant Brain Tumors: Targeting mRNAs, Regulatory RNAs, Mutations, Combinations, and Beyond. Neurotherapeutics 2019, 16, 319–347. [Google Scholar] [CrossRef]
- ParvizHamidi, M.; Haddad, G.; Ostadrahimi, S.; Ostadrahimi, N.; Sadeghi, S.; Fayaz, S.; Fard-Esfahani, P. Circulating miR-26a and miR-21 as biomarkers for glioblastoma multiform. Biotechnol. Appl. Biochem. 2019, 66, 261–265. [Google Scholar] [CrossRef]
- Ernst, A.; Campos, B.; Meier, J.; Devens, F.; Liesenberg, F.; Wolter, M.; Reifenberger, G.; Herold-Mende, C.; Lichter, P.; Radlwimmer, B. De-repression of CTGF via the miR-17-92 cluster upon differentiation of human glioblastoma spheroid cultures. Oncogene 2010, 29, 3411–3422. [Google Scholar] [CrossRef]
- Cai, Q.; Zhu, A.; Gong, L. Exosomes of glioma cells deliver miR-148a to promote proliferation and metastasis of glioblastoma via targeting CADM1. Bull. Cancer 2018, 105, 643–651. [Google Scholar] [CrossRef] [PubMed]
- Che, S.; Sun, T.; Wang, J.; Jiao, Y.; Wang, C.; Meng, Q.; Qi, W.; Yan, Z. miR-30 overexpression promotes glioma stem cells by regulating Jak/STAT3 signaling pathway. Tumour Biol. 2015, 36, 6805–6811. [Google Scholar] [CrossRef]
- Lai, N.S.; Wu, D.G.; Fang, X.G.; Lin, Y.C.; Chen, S.S.; Li, Z.B.; Xu, S.S. Serum microRNA-210 as a potential noninvasive biomarker for the diagnosis and prognosis of glioma. Br. J. Cancer 2015, 112, 1241–1246. [Google Scholar] [CrossRef]
- Liu, X.; Kang, J.; Sun, S.; Luo, Y.; Ji, X.; Zeng, X.; Zhao, S. iASPP, a microRNA-124 target, is aberrantly expressed in astrocytoma and regulates malignant glioma cell migration and viability. Mol. Med. Rep. 2018, 17, 1970–1978. [Google Scholar] [CrossRef]
- Bazzoni, R.; Bentivegna, A. Role of Notch Signaling Pathway in Glioblastoma Pathogenesis. Cancers 2019, 11, 292. [Google Scholar] [CrossRef]
- Li, Y.; Guessous, F.; Zhang, Y.; Dipierro, C.; Kefas, B.; Johnson, E.; Marcinkiewicz, L.; Jiang, J.; Yang, Y.; Schmittgen, T.D.; et al. MicroRNA-34a inhibits glioblastoma growth by targeting multiple oncogenes. Cancer Res. 2009, 69, 7569–7576. [Google Scholar] [CrossRef] [PubMed]
- Janaki Ramaiah, M.; Divyapriya, K.; Kartik Kumar, S.; Rajesh, Y. Drug-induced modifications and modulations of microRNAs and long non-coding RNAs for future therapy against Glioblastoma Multiforme. Gene 2020, 723, 144126. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.M.; Chiba, T.; Brill, B.; Delis, N.; von Manstein, V.; Vafaizadeh, V.; Oellerich, T.; Groner, B. Expression of the miR-302/367 cluster in glioblastoma cells suppresses tumorigenic gene expression patterns and abolishes transformation related phenotypes. Int. J. Cancer 2015, 137, 2296–2309. [Google Scholar] [CrossRef]
- Allen, B.K.; Stathias, V.; Maloof, M.E.; Vidovic, D.; Winterbottom, E.F.; Capobianco, A.J.; Clarke, J.; Schurer, S.; Robbins, D.J.; Ayad, N.G. Epigenetic pathways and glioblastoma treatment: Insights from signaling cascades. J. Cell. Biochem. 2015, 116, 351–363. [Google Scholar] [CrossRef] [PubMed]
- Ho, K.H.; Chen, P.H.; Hsi, E.; Shih, C.M.; Chang, W.C.; Cheng, C.H.; Lin, C.W.; Chen, K.C. Identification of IGF-1-enhanced cytokine expressions targeted by miR-181d in glioblastomas via an integrative miRNA/mRNA regulatory network analysis. Sci. Rep. 2017, 7, 732. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.H.; Wang, Y.; Sims, M.; Cai, C.; Pfeffer, L.M. MicroRNA-1 suppresses glioblastoma in preclinical models by targeting fibronectin. Cancer Lett. 2019, 465, 59–67. [Google Scholar] [CrossRef]
- Nadaradjane, A.; Briand, J.; Bougras-Cartron, G.; Disdero, V.; Vallette, F.M.; Frenel, J.S.; Cartron, P.F. miR-370-3p Is a Therapeutic Tool in Anti-glioblastoma Therapy but Is Not an Intratumoral or Cell-free Circulating Biomarker. Mol. Nucleic Acids 2018, 13, 642–650. [Google Scholar] [CrossRef]
- Wu, Z.; Sun, L.; Wang, H.; Yao, J.; Jiang, C.; Xu, W.; Yang, Z. MiR-328 expression is decreased in high-grade gliomas and is associated with worse survival in primary glioblastoma. PLoS ONE 2012, 7, e47270. [Google Scholar] [CrossRef]
- Li, G.F.; Cheng, Y.Y.; Li, B.J.; Zhang, C.; Zhang, X.X.; Su, J.; Wang, C.; Chang, L.; Zhang, D.Z.; Tan, C.L.; et al. miR-375 inhibits the proliferation and invasion of glioblastoma by regulating Wnt5a. Neoplasma 2019, 66, 350–356. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, A.M.; Morais, C.M.; Pena, F.; Marante, T.; Cunha, P.P.; Jurado, A.S.; Pedroso de Lima, M.C. Differentiation of glioblastoma stem cells promoted by miR-128 or miR-302a overexpression enhances senescence-associated cytotoxicity of axitinib. Hum. Mol. Genet. 2021, 30, 160–171. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Medarova, Z.; Moore, A. Role of microRNAs in glioblastoma. Oncotarget 2021, 12, 1707–1723. [Google Scholar] [CrossRef] [PubMed]
- Sasayama, T.; Tanaka, K.; Kohmura, E. The Roles of MicroRNAs in Glioblastoma Biology and Biomarker. In Neurooncology: Newer Developments; Agrawal, A., Ed.; IntechOpen: London, UK, 2016. [Google Scholar] [CrossRef]
- Kelly, T.K.; De Carvalho, D.D.; Jones, P.A. Epigenetic modifications as therapeutic targets. Nat. Biotechnol. 2010, 28, 1069–1078. [Google Scholar] [CrossRef]
- Banelli, B.; Forlani, A.; Allemanni, G.; Morabito, A.; Pistillo, M.P.; Romani, M. MicroRNA in glioblastoma: An overview. Int. J. Genom. 2017, 2017, 7639084. [Google Scholar] [CrossRef]
- Morita, S.; Horii, T.; Kimura, M.; Ochiya, T.; Tajima, S.; Hatada, I. miR-29 represses the activities of DNA methyltransferases and DNA demethylases. Int. J. Mol. Sci. 2013, 14, 14647–14658. [Google Scholar] [CrossRef]
- Bier, A.; Giladi, N.; Kronfeld, N.; Lee, H.K.; Cazacu, S.; Finniss, S.; Xiang, C.; Poisson, L.; deCarvalho, A.C.; Slavin, S.; et al. MicroRNA-137 is downregulated in glioblastoma and inhibits the stemness of glioma stem cells by targeting RTVP-1. Oncotarget 2013, 4, 665–676. [Google Scholar] [CrossRef]
- Hiddingh, L.; Raktoe, R.S.; Jeuken, J.; Hulleman, E.; Noske, D.P.; Kaspers, G.J.; Vandertop, W.P.; Wesseling, P.; Wurdinger, T. Identification of temozolomide resistance factors in glioblastoma via integrative miRNA/mRNA regulatory network analysis. Sci. Rep. 2014, 4, 5260. [Google Scholar] [CrossRef]
- Sun, J.; Ma, Q.; Li, B.; Wang, C.; Mo, L.; Zhang, X.; Tang, F.; Wang, Q.; Yan, X.; Yao, X.; et al. RPN2 is targeted by miR-181c and mediates glioma progression and temozolomide sensitivity via the wnt/β-catenin signaling pathway. Cell Death Dis. 2020, 11, 890. [Google Scholar] [CrossRef]
- Buruiana, A.; Florian, S.I.; Florian, A.I.; Timis, T.L.; Mihu, C.M.; Miclaus, M.; Osan, S.; Hrapsa, I.; Cataniciu, R.C.; Farcas, M.; et al. The Roles of miRNA in Glioblastoma Tumor Cell Communication: Diplomatic and Aggressive Negotiations. Int. J. Mol. Sci. 2020, 21, 1950. [Google Scholar] [CrossRef]
- Gao, Y.T.; Chen, X.B.; Liu, H.L. Up-regulation of miR-370-3p restores glioblastoma multiforme sensitivity to temozolomide by influencing MGMT expression. Sci. Rep. 2016, 6, 32972. [Google Scholar] [CrossRef] [PubMed]
- Areeb, Z.; Stylli, S.S.; Koldej, R.; Ritchie, D.S.; Siegal, T.; Morokoff, A.P.; Kaye, A.H.; Luwor, R.B. MicroRNA as potential biomarkers in Glioblastoma. J. Neuro-Oncol. 2015, 125, 237–248. [Google Scholar] [CrossRef]
- Wang, Q.; Li, P.; Li, A.; Jiang, W.; Wang, H.; Wang, J.; Xie, K. Plasma specific miRNAs as predictive biomarkers for diagnosis and prognosis of glioma. J. Exp. Clin. Cancer Res. 2012, 31, 97. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Cruickshanks, N.; Pahuski, M.; Yuan, F.; Dutta, A.; Schiff, D.; Purow, B.; Abounader, R. Noncoding RNAs in Glioblastoma. In Glioblastoma; De Vleeschouwer, S., Ed.; Exon Publications: Brisbane, Australia, 2017. [Google Scholar] [CrossRef]
- Yang, L.; Ma, Y.; Xin, Y.; Han, R.; Li, R.; Hao, X. Role of the microRNA 181 family in glioma development. Mol. Med. Rep. 2018, 17, 322–329. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Pang, B.; Xin, T.; Guo, H.; Xing, Y.; Xu, S.; Feng, B.; Liu, B.; Pang, Q. Plasma miR-221/222 Family as Novel Descriptive and Prognostic Biomarkers for Glioma. Mol. Neurobiol. 2016, 53, 1452–1460. [Google Scholar] [CrossRef] [PubMed]
- Mollaei, H.; Safaralizadeh, R.; Rostami, Z. MicroRNA replacement therapy in cancer. J. Cell. Physiol. 2019, 234, 12369–12384. [Google Scholar] [CrossRef] [PubMed]
- Alamdari-Palangi, V.; Karami, Z.; Karami, H.; Baazm, M. MiRNA-7 Replacement Effect on Proliferation and Tarceva-Sensitivity in U373-MG Cell Line. Asian Pac. J. Cancer Prev. 2020, 21, 1747–1753. [Google Scholar] [CrossRef] [PubMed]
- Xiong, H.; Veedu, R.N.; Diermeier, S.D. Recent Advances in Oligonucleotide Therapeutics in Oncology. Int. J. Mol. Sci. 2021, 22, 3295. [Google Scholar] [CrossRef] [PubMed]
- Romani, M.; Pistillo, M.P.; Banelli, B. Epigenetic Targeting of Glioblastoma. Front. Oncol. 2018, 8, 448. [Google Scholar] [CrossRef]
- Stupp, R.; Brada, M.; van den Bent, M.J.; Tonn, J.C.; Pentheroudakis, G. High-grade glioma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2014, 25 (Suppl. S3), iii93–iii101. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hasan, H.; Afzal, M.; Castresana, J.S.; Shahi, M.H. A Comprehensive Review of miRNAs and Their Epigenetic Effects in Glioblastoma. Cells 2023, 12, 1578. https://doi.org/10.3390/cells12121578
Hasan H, Afzal M, Castresana JS, Shahi MH. A Comprehensive Review of miRNAs and Their Epigenetic Effects in Glioblastoma. Cells. 2023; 12(12):1578. https://doi.org/10.3390/cells12121578
Chicago/Turabian StyleHasan, Hera, Mohammad Afzal, Javier S. Castresana, and Mehdi H. Shahi. 2023. "A Comprehensive Review of miRNAs and Their Epigenetic Effects in Glioblastoma" Cells 12, no. 12: 1578. https://doi.org/10.3390/cells12121578
APA StyleHasan, H., Afzal, M., Castresana, J. S., & Shahi, M. H. (2023). A Comprehensive Review of miRNAs and Their Epigenetic Effects in Glioblastoma. Cells, 12(12), 1578. https://doi.org/10.3390/cells12121578







